Submitted:
04 March 2026
Posted:
04 March 2026
You are already at the latest version
Abstract
Rust, caused by Puccinia arachidis, is one among the most destructive fungal diseases constraining global groundnut (Arachis hypogaea L.) production. While the development of disease-resistant varieties stands as the most effective approach to preventing substantial yield losses, the genetic mechanisms underlying resistance to rust is not yet well understood, emphasizing the necessity for further detailed research. In this study, 184 accessions from the ICRISAT groundnut mini-core collection were evaluated for rust resistance at Dharwad, India, across multiple seasons, as well as in Vietnam for one season. Whole-genome resequencing-based genome-wide association study (GWAS) identified five highly significant marker trait associations (MTAs) for rust resistance (p = 5.22 × 10-13 to 7.21 × 10-08). Among these, two robust rust-associated kompetitive allele specific PCR (KASP) markers, snpAH00607 at chromosome Ah01 and snpAH00609 at chromosome Ah17, were validated across diverse set of breeding and pre-breeding lines. These markers were linked to candidate genes encoding sterol C4-methyl oxidase 1-2, implicated in brassinosteroid-mediated salicylic acid signalling, and MYB transcription factor known to be associated with defense responses. The identified SNPs, validated markers, and candidate genes will serve as important resources for marker-assisted breeding of rust disease resistant groundnut varieties.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Material and Phenotyping for Rust
| Trait | Source of Variation | Df | Sum Sq | Mean Sq | F value | Pr(>F) |
|---|---|---|---|---|---|---|
| Rust | Replication | 2 | 0.115 | 0.058 | 0.346 | 0.707 |
| Genotype ** | 156 | 243.033 | 1.558 | 9.358 | 0 | |
| Environment ** | 3 | 2406.136 | 802.045 | 4817.742 | 0 | |
| Genotype x Environment ** | 468 | 216.48 | 0.463 | 2.779 | 0 | |
| Error | 1254 | 208.763 | 0.166 |
| SNP | Type of variant | Chr. No | Position | Allele (Ref/Alt) | P - value | Models | PVE (%) | Gene ID | Annotation |
|---|---|---|---|---|---|---|---|---|---|
| Arahy.01_30014046 | intergenic | 1 | 30014046 | C/T | 5.22 x 10-13 | FarmCPU, BLINK | 19.7 | P9U8RJ | Ubiquitin family protein |
| N67QXJ | Protein kinase superfamily protein | ||||||||
| Arahy.01_48511619 | intergenic | 1 | 48511619 | A/G | 7.06 x 10-09 | FarmCPU | 17.6 | 696GFM | Uncharacterized protein At4g22758-like |
| 11UTVE | Sterol C4-methyl oxidase 1-2 | ||||||||
| Arahy.08_28112898 | upstream | 8 | 28112898 | T/G | 4.57 x 10-11 | Blink | 4.2 | CC6MMP | 40s ribosomal protein SA |
| Arahy.15_142635616 | intergenic | 15 | 142635616 | G/A | 7.08 x 10-08 | FarmCPU | 22.71 | 5E7A70 | Winged-helix DNA-binding transcription factor family protein, putative isoform |
| V5GBYV | Protein kinase superfamily protein | ||||||||
| Arahy.17_133493906 | intergenic | 17 | 133493906 | A/G | 3.54 x 10-09 | FarmCPU | 0.53 | ES2NTH | Ferrochelatase 1 |
| H8IXR5 | MYB transcription factor MYB127 [Glycine max] |
2.2. Statistical Analysis of Phenotypic Data
2.3. DNA Isolation, Sequencing and SNP Calling for WGRS Data
2.4. Genome Wide Studies for Identifying MTAs with Rust Resistance
2.5. Gene Predictions Within the Candidate Interval and Pathway Analysis
2.6. Kompetitive Allele-Specific PCR (KASP) Marker Development and Validation
3. Results
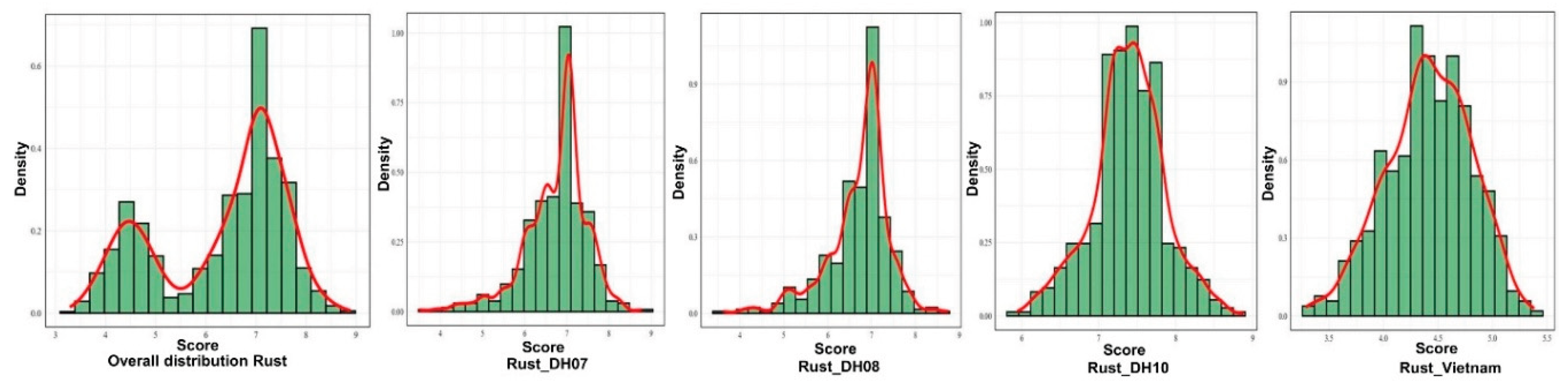
3.1. Phenotype Variation and Descriptive Statistics for Rust Across Environments
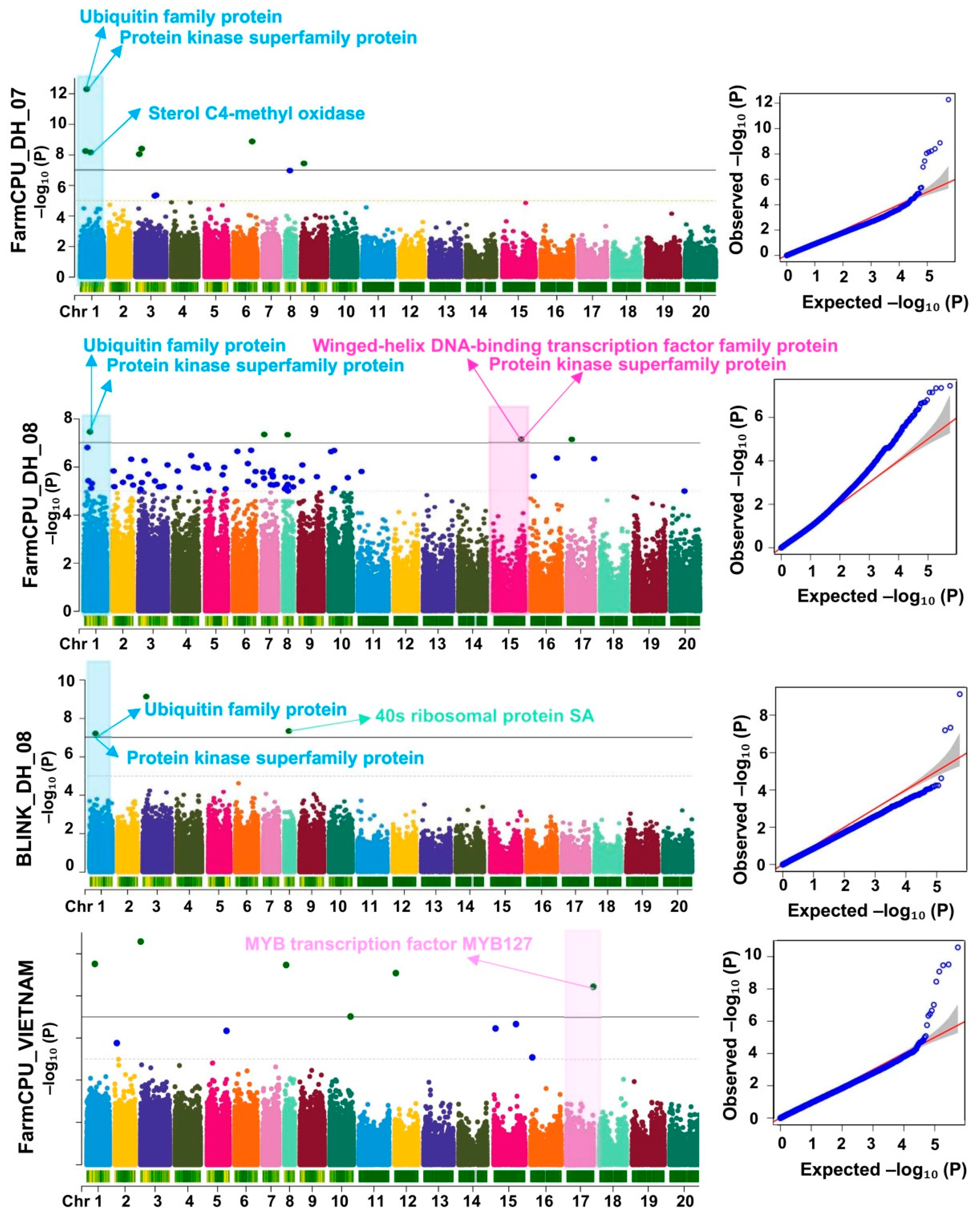
3.2. Genome- Wide Association Studies for Rust
3.2.1. Significant MTAs Associated with Rust
3.2.3. Identification of Candidate Genes for Resistance to Rust
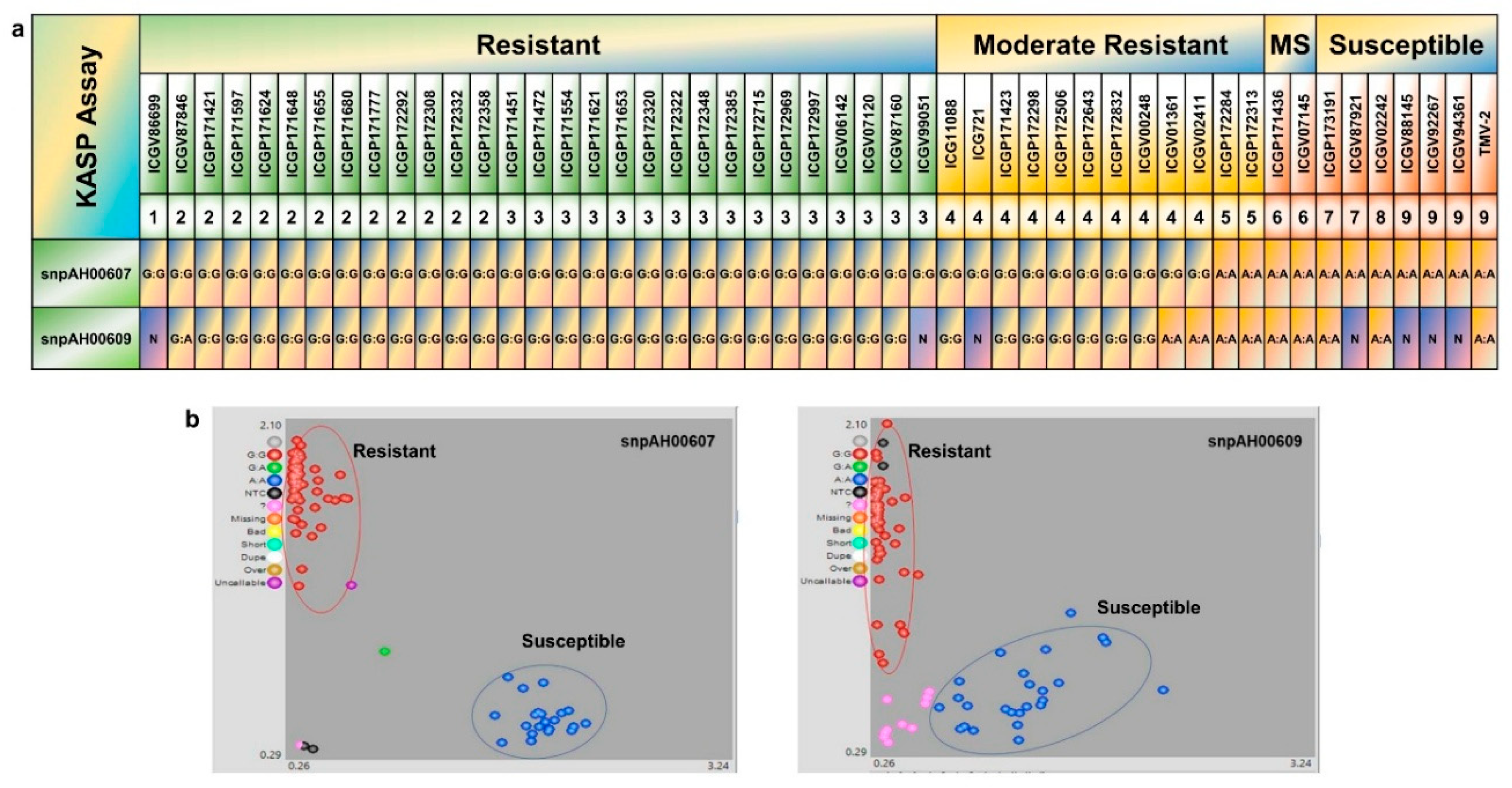
3.3. Development and Validation of KASP Markers Associated with Rust
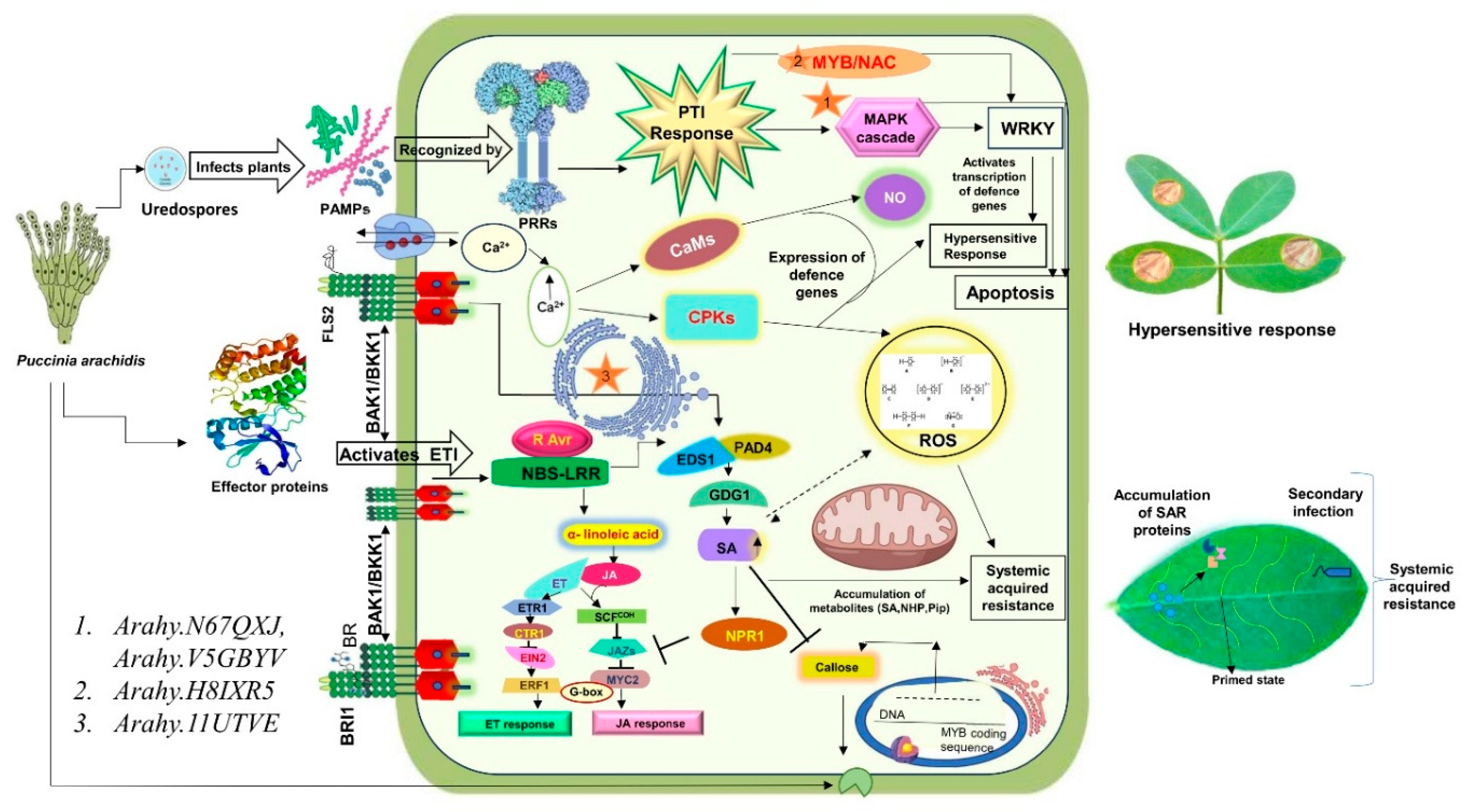
3.4. Gene Pathway for Rust Resistance
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Stalker, H.T. Utilizing wild species for peanut improvement. Crop Science 2017, 57(3), 1102–1120. [Google Scholar] [CrossRef]
- Krapovickas, A.; Gregory, W.C.; Williams, D.E.; Simpson, C.E. Taxonomy of the genus Arachis (Leguminosae). Bonplandia 2007, 16, 7–205. [Google Scholar] [CrossRef]
- Knauft, D.A.; Gorbet, D.W. Genetic diversity among peanut cultivars. Crop Science 1989, 29(6), 1417–1422. [Google Scholar] [CrossRef]
- Chamberlin, K.D.C.; Melouk, H.A.; Payton, M.E. Evaluation of the US peanut mini core collection using a molecular marker for resistance to Sclerotinia minor Jagger. Euphytica 2010, 172(1), 109–115. [Google Scholar] [CrossRef]
- Anderson, W.F.; Holbrook, C.C.; Culbreath, A.K. Screening the peanut core collection for resistance to tomato spotted wilt virus. Peanut Science 1996, 23(1), 57–61. [Google Scholar] [CrossRef]
- Sudini, H.; Upadhyaya, H.D.; Reddy, S.V.; Mangala, U.N.; Rathore, A.; Kumar, K.V.K. Resistance to late leaf spot and rust diseases in ICRISAT’s mini core collection of peanut (Arachis hypogaea L.). Australasian Plant Pathology 2015, 44(5), 557–566. [Google Scholar] [CrossRef]
- Toomer, O.T. Nutritional chemistry of the peanut (Arachis hypogaea). Critical Reviews in Food Science and Nutrition 2018, 58(17), 3042–3053. [Google Scholar] [CrossRef]
- Zhang, H.; Chu, Y.; Dang, P.; Tang, Y.; Jiang, T.; Clevenger, J.P.; Ozias-Akins, P.; Holbrook, C.; Wang, M.L.; Campbell, H.; Hagan, A. Identification of QTLs for resistance to leaf spots in cultivated peanut (Arachis hypogaea L.) through GWAS analysis. Theoretical and Applied Genetics 2020, 133(7), 2051–2061. [Google Scholar] [CrossRef]
- Anco, D.J.; Thomas, J.S.; Jordan, D.L.; Shew, B.B.; Monfort, W.S.; Mehl, H.L.; Small, I.M.; Wright, D.L.; Tillman, B.L.; Dufault, N.S.; Hagan, A.K. Peanut yield loss in the presence of defoliation caused by late or early leaf spot. Plant Disease 2020, 104(5), 1390–1399. [Google Scholar] [CrossRef] [PubMed]
- Pandey, M.K.; Gangurde, S.S.; Shasidhar, Y.; Sharma, V.; Kale, S.M.; Khan, A.W.; Shah, P.; Joshi, P.; Bhat, R.S.; Janila, P.; Bera, S.K. High-throughput diagnostic markers for foliar fungal disease resistance and high oleic acid content in groundnut. BMC Plant Biology 2024, 24(1), pp.262. [Google Scholar] [PubMed]
- Mondal, S.; Badigannavar, A.M. Peanut rust (Puccinia arachidis Speg.) disease: its background and recent accomplishments towards disease resistance breeding. Protoplasma 2015, 252(6), 1409–1420. [Google Scholar] [CrossRef] [PubMed]
- Riley, R.; Chapman, V.; Johnson, R.O.Y. Introduction of yellow rust resistance of Aegilops comosa into wheat by genetically induced homoeologous recombination. Nature 1968, 217(5126), 383–384. [Google Scholar] [CrossRef]
- Chaudhari, S.; Khare, D.; Patil, S.C.; Sundravadana, S.; Variath, M.T.; Sudini, H.K.; Manohar, S.S.; Bhat, R.S.; Pasupuleti, J. Genotype × environment studies on resistance to late leaf spot and rust in genomic selection training population of peanut (Arachis hypogaea L.). Frontiers in Plant Science 2019, 10, 1–14. [Google Scholar] [CrossRef]
- Luo, D.; Shi, L.; Sun, Z.; Qi, F.; Liu, H.; Xue, L.; Li, X.; Liu, H.; Qu, P.; Zhao, H.; Dai, X. Genome-wide association studies of embryogenic callus induction rate in peanut (Arachis hypogaea L.). Genes 2024, 15(2), 1–12. [Google Scholar] [CrossRef]
- Pandey, M.K.; Khan, A.W.; Singh, V.K.; Vishwakarma, M.K.; Shasidhar, Y.; Kumar, V.; Garg, V.; Bhat, R.S.; Chitikineni, A.; Janila, P.; Guo, B. QTL-seq approach identified genomic regions and diagnostic markers for rust and late leaf spot resistance in groundnut (Arachis hypogaea L.). Plant Biotechnology Journal 2017, 15(8), 927–941. [Google Scholar] [CrossRef]
- Sujay, V.; Gowda, M.V.C.; Pandey, M.K.; Bhat, R.S.; Khedikar, Y.P.; Nadaf, H.L.; Gautami, B.; Sarvamangala, C.; Lingaraju, S.; Radhakrishan, T.; Knapp, S.J. Quantitative trait locus analysis and construction of consensus genetic map for foliar disease resistance based on two recombinant inbred line populations in cultivated groundnut (Arachis hypogaea L.). Molecular Breeding 2012, 30(2), 773–788. [Google Scholar] [CrossRef]
- Cui, C.; Mei, H.; Liu, Y.; Zhang, H.; Zheng, Y. Genetic diversity, population structure, and linkage disequilibrium of an association-mapping panel revealed by genome-wide SNP markers in sesame. Frontiers in Plant Science 2017, 8, 1189. [Google Scholar]
- Liu, X.; Huang, M.; Fan, B.; Buckler, E.S.; Zhang, Z. Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS Genetics 2016, 12(2), e1005767. [Google Scholar] [CrossRef]
- Parmar, S.; Sharma, V.; Bomireddy, D.; Soni, P.; Joshi, P.; Gangurde, S.S.; Wang, J.; Bera, S.K.; Bhat, R.S.; Desmae, H.; Shirasawa, K. Recent advances in genetics, genomics, and breeding for nutritional quality in groundnut. Accelerated Plant Breeding 4 2022, 111–137. [Google Scholar]
- Berhe, M.; Dossa, K.; You, J.; Mboup, P.A.; Diallo, I.N.; Diouf, D.; Zhang, X.; Wang, L. Genome-wide association study and its applications in the non-model crop Sesamum indicum. BMC Plant Biology 2021, 21(1), 283. [Google Scholar] [CrossRef] [PubMed]
- Karikari, B.; Wang, Z.; Zhou, Y.; Yan, W.; Feng, J.; Zhao, T. Identification of quantitative trait nucleotides and candidate genes for soybean seed weight by multiple models of genome-wide association study. BMC Plant Biology 2020, 20(1), 404. [Google Scholar] [CrossRef] [PubMed]
- Osuna-Caballero, S.; Cobos, M.J.; Ruiz, C.M.; Wohor, O.Z.; Rispail, N.; Rubiales, D. Genome-wide association studies on resistance to pea weevil: identification of novel sources of resistance and associated markers. International Journal of Molecular Sciences 2024, 25(14), 7920. [Google Scholar] [CrossRef]
- Vikas, V.K.; Pradhan, A.K.; Budhlakoti, N.; Mishra, D.C.; Chandra, T.; Bhardwaj, S.C.; Kumar, S.; Sivasamy, M.; Jayaprakash, P.; Nisha, R.; Shajitha, P. Multi-locus genome-wide association studies (ML-GWAS) reveal novel genomic regions associated with seedling and adult plant stage leaf rust resistance in bread wheat (Triticum aestivum L.). Heredity 2022, 128(6), 434–449. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Bramel, P.J.; Ortiz, R.; Singh, S. Developing a mini core of peanut for utilization of genetic resources. Crop Science 2002, 42(6), 2150–2156. [Google Scholar] [CrossRef]
- Subrahmanyam, P.; McDonald, D.; Waliyar, F.; Reddy, L.J.; Nigam, S.N.; Gibbons, R.W.; Rao, V.R.; Singh, A.K.; Pande, S.; Reddy, P.M.; Rao, P.S. Screening methods and sources of resistance to rust and late leaf spot of groundnut. In Information Bulletin no. 47; International Crops Research Institute for the Semi-Arid Tropics, 1995. [Google Scholar]
- Burton, G.W. Quantitative inheritance in grasses; 1952; pp. 277–283. [Google Scholar]
- Johnson, H.W.; Robinson, H.F.; Comstock, R. Estimates of genetic and environmental variability in soybeans. Agronomy Journal 1955, 47, 314–18. [Google Scholar] [CrossRef]
- Tang, Y.; Liu, X.; Wang, J.; Li, M.; Wang, Q.; Tian, F.; Su, Z.; Pan, Y.; Liu, D.; Lipka, A.E.; Buckler, E.S. GAPIT version 2: an enhanced integrated tool for genomic association and prediction. The Plant Genome 2016, 9(2), 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M. Toward pathway engineering: a new database of genetic and molecular pathways. Sci. Technol. Jap. 1996, 59, 34–38. [Google Scholar]
- Pandey, M.K.; Upadhyaya, H.D.; Rathore, A.; Vadez, V.; Sheshshayee, M.S.; Sriswathi, M.; Govil, M.; Kumar, A.; Gowda, M.V.C.; Sharma, S.; Hamidou, F. Genome wide association studies for 50 agronomic traits in peanut using the ‘reference set’ comprising 300 genotypes from 48 countries of the semi-arid tropics of the world. PLoS One 2014, 9(8), e105228. [Google Scholar] [CrossRef]
- Clevenger, J.; Chu, Y.; Chavarro, C.; Botton, S.; Culbreath, A.; Isleib, T.G.; Holbrook, C.C.; Ozias-Akins, P. Mapping late leaf spot resistance in peanut (Arachis hypogaea L.) using QTL-seq reveals markers for marker-assisted selection. Frontiers in Plant Science 2018, 9, 83. [Google Scholar] [CrossRef]
- Pandey, M.K.; Agarwal, G.; Kale, S.M.; Clevenger, J.; Nayak, S.N.; Sriswathi, M.; Chitikineni, A.; Chavarro, C.; Chen, X.; Upadhyaya, H.D.; Vishwakarma, M.K. Development and evaluation of a high density genotyping ‘Axiom_ Arachis’ array with 58 K SNPs for accelerating genetics and breeding in groundnut. Scientific Reports 2017, 7(1), 40577. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, W.; Chen, H.; Yang, M.; Wang, J.; Pandey, M.K.; Zhang, C.; Chang, W.C.; Zhang, L.; Zhang, X.; Tang, R.; Garg, V. The genome of cultivated peanut provides insight into legume karyotypes, polyploid evolution and crop domestication. Nature Genetics 2019, 51(5), 865–876. [Google Scholar] [CrossRef]
- Bertioli, D.J.; Jenkins, J.; Clevenger, J.; Dudchenko, O.; Gao, D.; Seijo, G.; Leal-Bertioli, S.C.; Ren, L.; Farmer, A.D.; Pandey, M.K.; Samoluk, S.S. The genome sequence of segmental allotetraploid peanut Arachis hypogaea. Nature Genetics 2019, 51(5), 877–884. [Google Scholar] [CrossRef] [PubMed]
- Varshney, R.K.; Pandey, M.K.; Janila, P.; Nigam, S.N.; Sudini, H.; Gowda, M.V.C.; Sriswathi, M.; Radhakrishnan, T.; Manohar, S.S.; Nagesh, P. Marker-assisted introgression of a QTL region to improve rust resistance in three elite and popular varieties of peanut (Arachis hypogaea L.). Theoretical and Applied Genetics 2014, 127(8), 1771–1781. [Google Scholar] [CrossRef]
- Khedikar, Y.P.; Gowda, M.V.C.; Sarvamangala, C.; Patgar, K.V.; Upadhyaya, H.D.; Varshney, R.K. A QTL study on late leaf spot and rust revealed one major QTL for molecular breeding for rust resistance in groundnut (Arachis hypogaea L.). Theoretical and Applied Genetics 2010, 121(5), 971–984. [Google Scholar] [CrossRef]
- Shi, X.; Sun, Z.; Qi, F.; Han, S.; Zheng, Y.; Dong, W.; Zhang, M.; Zhang, X. Identification of genomic regions associated with peanut rust resistance by genome-wide association studies. Plants 2025, 14(8), 1219. [Google Scholar] [CrossRef]
- Rao, A.S.; McDonald, D.; Reddy, K.R. Effect of temperature on rust and leaf spot disease development in groundnut. Journal of Oilseeds Research 1997, 14(2), 249–255. [Google Scholar]
- Daudi, H.; Shimelis, H.; Mwadzingeni, L.; Laing, M.; Okori, P. Breeding groundnut for rust resistance: A review. legume research (TSI) 2019, 42(3), 291–299. [Google Scholar] [CrossRef]
- Mondal, S.; Shafi, K.M.; Raizada, A.; Song, H.; Badigannavar, A.M.; Sowdhamini, R. Development of candidate gene-based markers and map-based cloning of a dominant rust resistance gene in cultivated groundnut (Arachis hypogaea L.). Gene 2022, 827, 146474. [Google Scholar] [CrossRef] [PubMed]
- Rathod, V.; Hamid, R.; Tomar, R.S.; Patel, R.; Padhiyar, S.; Kheni, J.; Thirumalaisamy, P.P.; Munshi, N.S. Comparative RNA-Seq profiling of a resistant and susceptible peanut (Arachis hypogaea) genotypes in response to leaf rust infection caused by Puccinia arachidis. 3 Biotech 2020, 10(6), 284. [Google Scholar] [CrossRef]
- Mondal, S.; Badigannavar, A.M. Mapping of a dominant rust resistance gene revealed two R genes around the major Rust_QTL in cultivated peanut (Arachis hypogaea L.). Theoretical and Applied Genetics 2018, 131(8), 1671–1681. [Google Scholar] [CrossRef]
- Gao, X.; Cox, K.L., Jr.; He, P. Functions of calcium-dependent protein kinases in plant innate immunity. Plants 2014, 3(1), 160–176. [Google Scholar] [CrossRef]
- Asai, T.; Tena, G.; Plotnikova, J.; Willmann, M.R.; Chiu, W.L.; Gomez-Gomez, L.; Boller, T.; Ausubel, F.M.; Sheen, J. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 2002, 415(6875), 977–983. [Google Scholar] [CrossRef]
- Heil, M.; Bostock, R.M. Induced systemic resistance (ISR) against pathogens in the context of induced plant defences. Annals of Botany 2002, 89(5), 503–512. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Zhou, H.; Zhang, Y.; Zhao, Y.; Zhang, Y.; Feng, X.; Lin, H. Diverse roles of MYB transcription factors in plants. Journal of Integrative Plant Biology 2025, 67(3), 539–562. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Jing, W.; Zheng, J.; Jin, Y.; Wu, D.; Cao, C.; Dong, Y.; Shi, X.; Zhang, W. The ATP-binding cassette transporter OsPDR1 regulates plant growth and pathogen resistance by affecting jasmonates biosynthesis in rice. Plant Science 2020, 298, 110582. [Google Scholar] [CrossRef]
- Yang, J.; Duan, G.; Li, C.; Liu, L.; Han, G.; Zhang, Y.; Wang, C. The crosstalks between jasmonic acid and other plant hormone signaling highlight the involvement of jasmonic acid as a core component in plant response to biotic and abiotic stresses. Frontiers in Plant Science 2019, 10, 1349. [Google Scholar] [CrossRef] [PubMed]
- Manghwar, H.; Hussain, A.; Ali, Q.; Liu, F. Brassinosteroids (BRs) role in plant development and coping with different stresses. International Journal of Molecular Sciences 2022, 23(3), 1012. [Google Scholar] [CrossRef]
- Miao, R.; Li, C.; Liu, Z.; Zhou, X.; Chen, S.; Zhang, D.; Luo, J.; Tang, W.; Wang, C.; Wu, J.; Chen, Z. The role of endogenous brassinosteroids in the mechanisms regulating plant reactions to various abiotic stresses. Agronomy 2024, 14(2), 356. [Google Scholar] [CrossRef]
- Fujioka, S.; Sakurai, A. Biosynthesis and metabolism of brassinosteroids. Physiologia Plantarum 1997, 100(3), 710–715. [Google Scholar] [CrossRef]
- Fujioka, S.; Yokota, T. Biosynthesis and metabolism of brassinosteroids. Annual Review of Plant Biology 2003, 54(1), 137–164. [Google Scholar] [CrossRef]
- Kim, Y.W.; Youn, J.H.; Roh, J.; Kim, J.M.; Kim, S.K.; Kim, T.W. Brassinosteroids enhance salicylic acid-mediated immune responses by inhibiting BIN2 phosphorylation of clade I TGA transcription factors in Arabidopsis. Molecular Plant 2022, 15(6), 991–1007. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





