Submitted:
28 February 2026
Posted:
02 March 2026
You are already at the latest version
Abstract
Raw milk cheeses represent complex microbial ecosystems that may act as reservoirs of antimicrobial-resistant bacteria. This study investigated the microbiological characteristics, bacterial community structure, and antimicrobial resistance (AMR) profiles of artisanal goat and sheep cheeses produced by regional manufacturers in Poland. A total of ten cheeses, including five goat cheeses and five sheep oscypek-type cheeses, were analysed. Culture-dependent enumeration and isolation were combined with molecular identification via 16S rRNA gene sequencing and antimicrobial susceptibility testing by disk diffusion. Total viable bacterial counts ranged from 105 to 108 CFU/mL, revealing considerable variability among individual samples. Microbiological profiling indicated the predominance of lactic acid bacteria, with Lactococcus, Lactobacillus, and Streptococcus representing the dominant genera. Multivariate analysis (PCoA) demonstrated substantial intra-group dispersion and overlapping clustering patterns between goat and sheep cheeses, suggesting that sample-specific ecological factors exerted a stronger influence on microbial composition than milk origin. Among 150 bacterial isolates, multidrug resistance (MDR) was detected in 28.7% of strains. MDR prevalence varied markedly between bacterial groups, reaching 100.0% in Enterobacterales, 73.3% in Enterococcus spp., and 16.2% in lactic acid bacteria. Resistance was most frequently observed for aminoglycosides and β-lactam antibiotics, particularly streptomycin and gentamicin. The results indicate that artisanal cheeses constitute heterogeneous microbial niches and may serve as potential reservoirs of antimicrobial-resistant bacteria. Integrating microbiological and AMR analyses provides valuable insight into the ecological determinants of resistance in traditional dairy products. These findings indicate that artisanal cheeses may represent heterogeneous microbial niches and potential reservoirs of AMR bacteria. The integration of microbiome profiling and phenotypic AMR assessment provides valuable insight into the ecological drivers of resistance in traditional dairy products.
Keywords:
1. Introduction
2. Results
1.1. Microbiological Counts in Goat and Sheep Cheese Samples
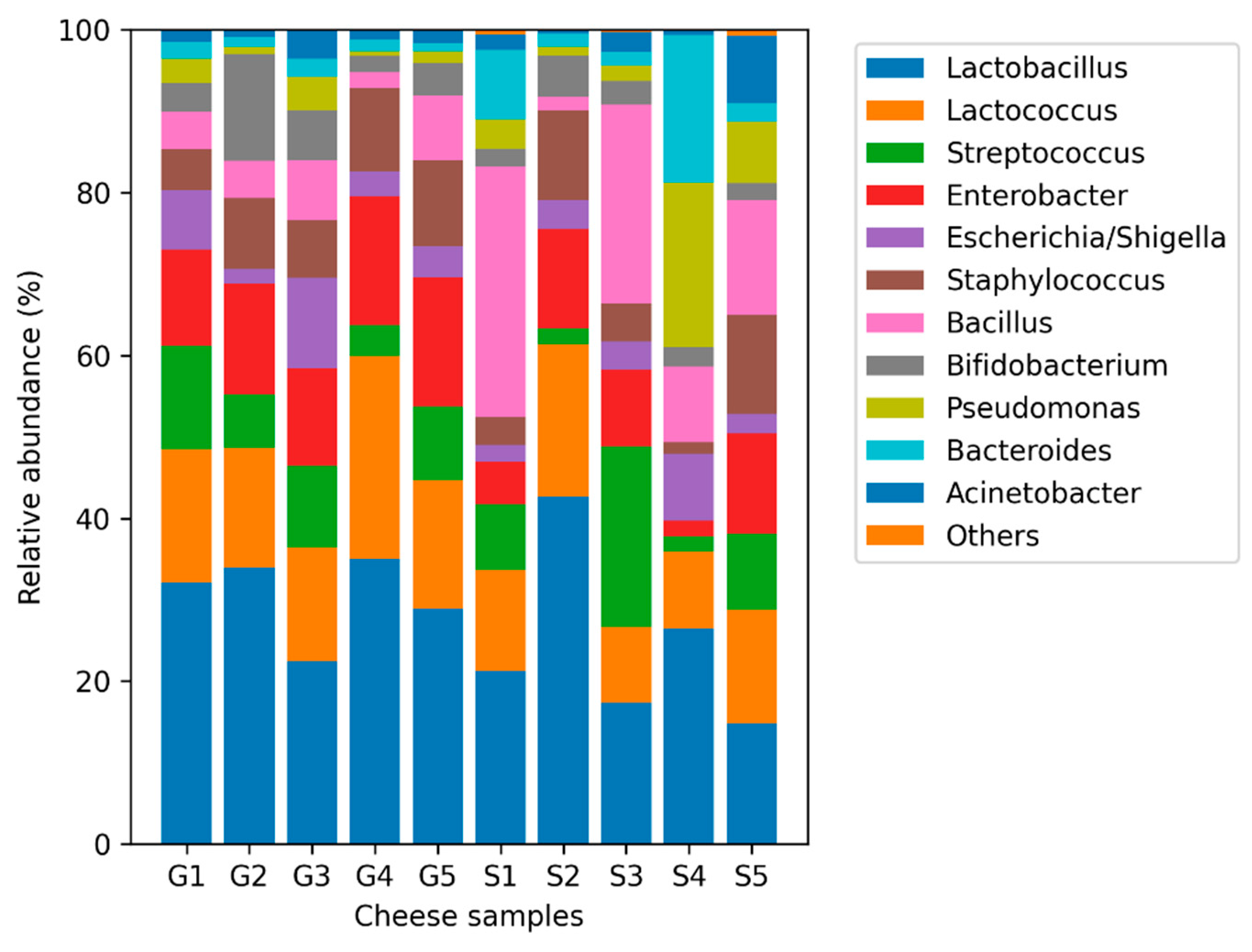
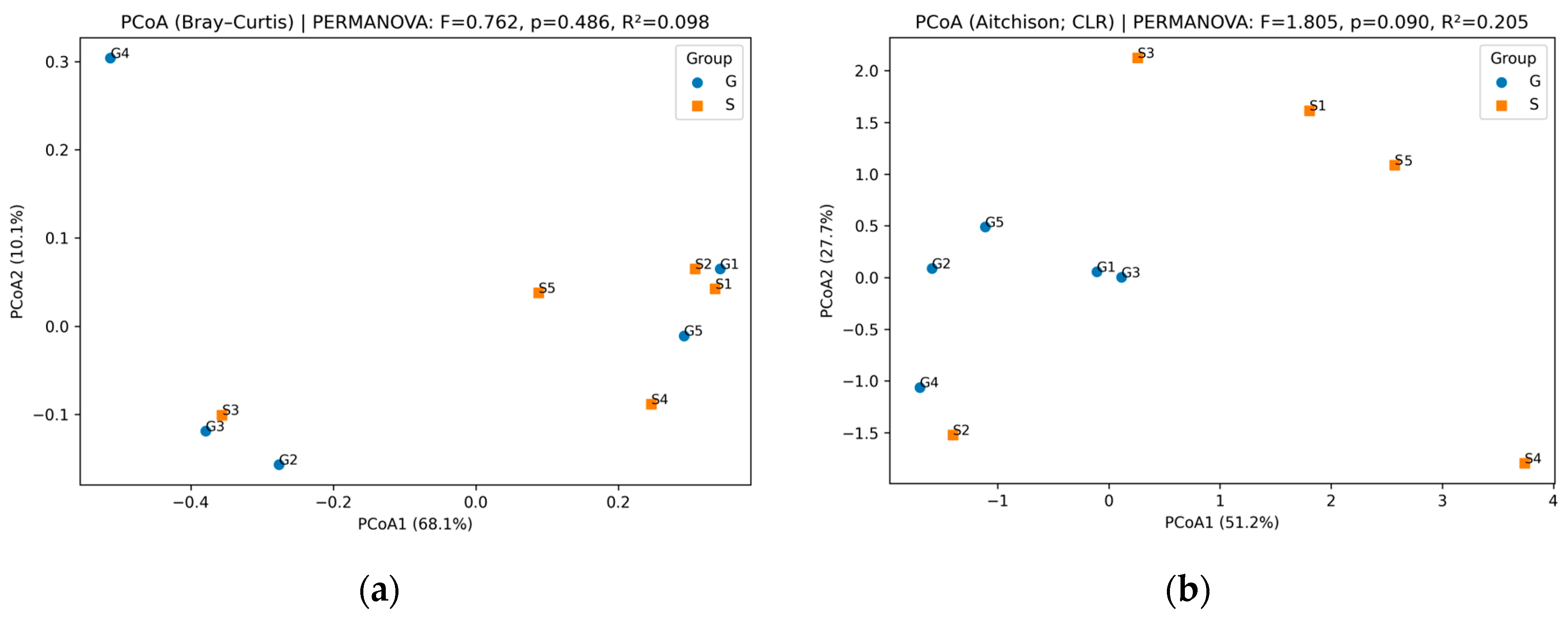
1.2. Microbial Community Composition of Individual Goat and Sheep Cheese Samples
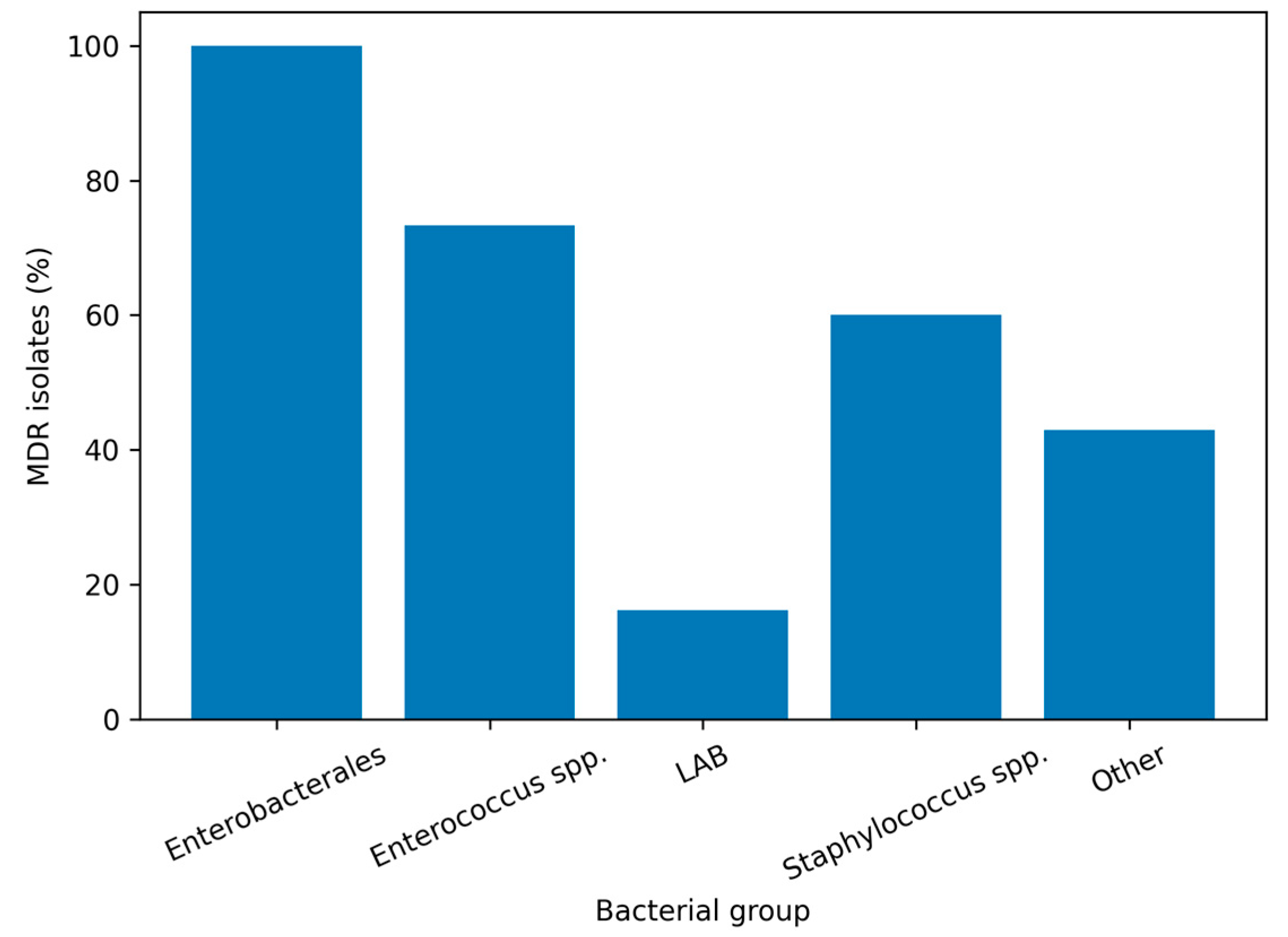
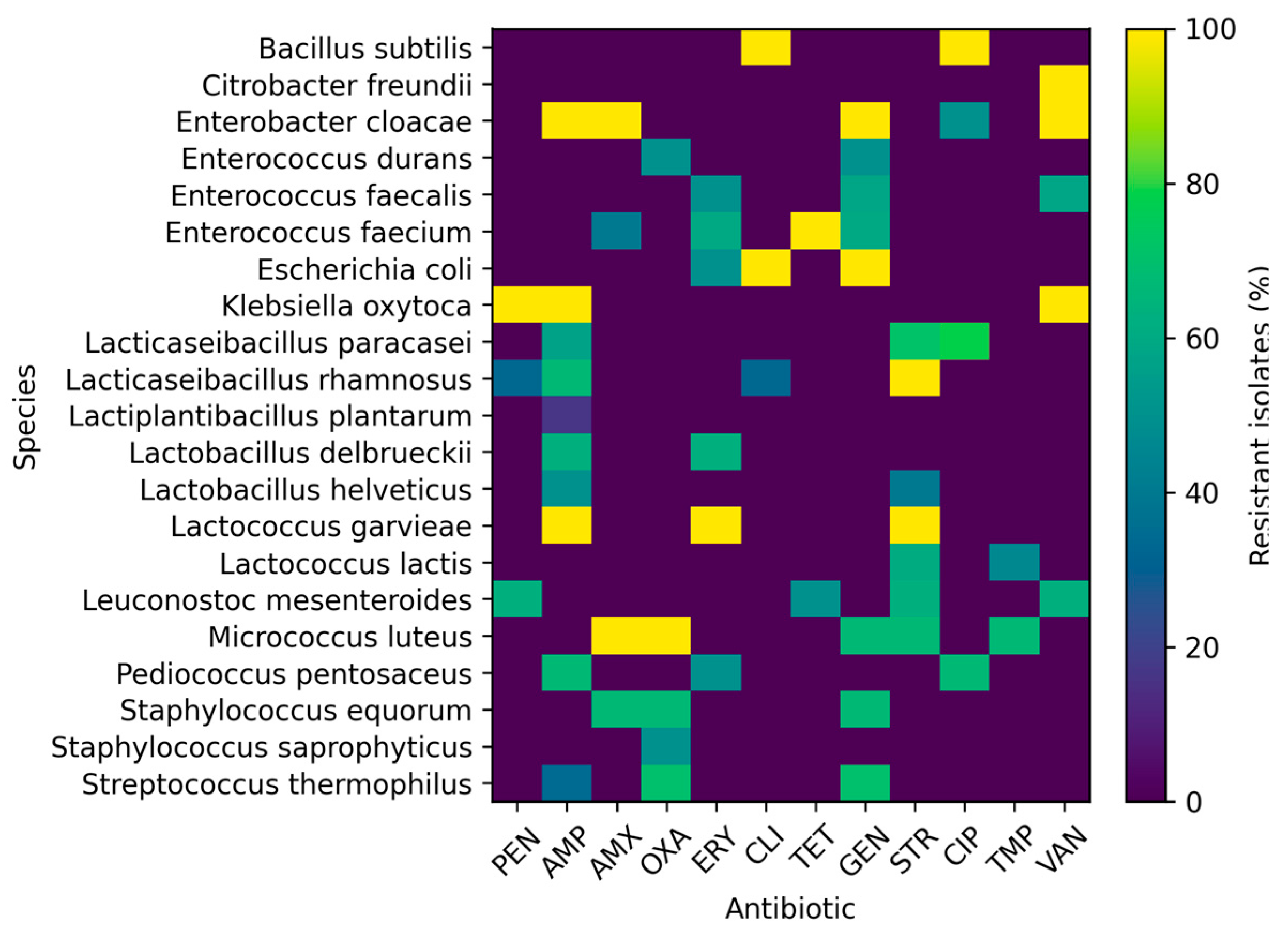
1.3. Antibiotic Susceptibility of Bacterial Isolates (Disk Diffusion Method)
3. Discussion
4. Materials and Methods
4.1. Cheese Samples
4.2. Microbiological Analyses
4.3. Antibiotic Panel and Materials
4.4. Genomic DNA Extraction
4.5. PCR Amplification of the 16S rRNA Gene
4.6. DNA Amplification
4.7. PCR Product Purification and Quantification
4.8. Library Preparation and MinION Sequencing
4.9. Flow Cell Loading and Sequencing
4.10. Bioinformatic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Conflicts of Interest
Abbreviations
| AMR | Antimicrobial Resistance |
| AR | Antibiotic Resistance |
| ARB | Antibiotic-Resistant Bacteria |
| LAB | Lactic Acid Bacteria |
| MDR | Multidrug-Resistant |
| MIC | Minimum Inhibitory Concentration |
| PCoA | Principal Coordinates Analysis |
| PERMANOVA | Permutational Multivariate Analysis of Variance |
| SIMPER | Similarity Percentage Analysis |
| CLR | Centered Log-Ratio |
| CFU | Colony Forming Units |
| PCR | Polymerase Chain Reaction |
| DNA | Deoxyribonucleic Acid |
| rRNA | Ribosomal Ribonucleic Acid |
| OTU | Operational Taxonomic Unit |
| BLAST | Basic Local Alignment Search Tool |
| NCBI | National Center for Biotechnology Information |
| AMP | Ampicillin |
| AMX | Amoxicillin |
| PEN | Penicillin |
| OXA | Oxacillin |
| GEN | Gentamicin |
| STR | Streptomycin |
| CIP | Ciprofloxacin |
| TET | Tetracycline |
| ERY | Erythromycin |
| VAN | Vancomycin |
| TMP | Trimethoprim |
| CLI | Clindamycin |
References
- Nayik, G.A.; Jagdale, Y.D.; Gaikwad, S.A.; Devkatte, A.N.; Dar, A.H.; Ansari, M.J. Bioactive components, health benefits and processing of goat milk: A review. Front. Nutr. 2021, 8, 789100. [Google Scholar] [CrossRef] [PubMed]
- AlKaisy, Q.H.; et al. Exploring the health benefits and functional properties of goat milk proteins. Food Sci. Nutr. 2023, 11, 5641–5656. [Google Scholar] [CrossRef] [PubMed]
- Kawęcka, A.; Pasternak, M. Nutritional and dietetic quality of milk and traditional cheese made from the milk of native breeds of sheep and goats. J. Appl. Anim. Res. 2022. [Google Scholar] [CrossRef]
- Montel, M.C.; Buchin, S.; Mallet, A.; et al. Traditional cheeses: Rich and diverse microbiota with associated benefits. Int. J. Food Microbiol. 2014, 177, 136–154. [Google Scholar] [CrossRef]
- Campedelli, I.; Mathur, H.; Salvetti, E.; Clarke, S.; Rea, M.C.; Torriani, S.; Ross, R.P.; Hill, C.; O’Toole, P.W. Genus-wide assessment of antibiotic resistance in Lactobacillus spp. Appl. Environ. Microbiol. 2019, 85, e01738-18. [Google Scholar] [CrossRef]
- Arias, C.A.; Murray, B.E. The rise of the Enterococcus: Beyond vancomycin resistance. Nat. Rev. Microbiol. 2012, 10, 266–278. [Google Scholar] [CrossRef]
- Ombarak, R.A.; Hinenoya, A.; Elbagory, A.-R.M.; Yamasaki, S. Prevalence and molecular characterization of antimicrobial resistance in Escherichia coli isolated from raw milk and raw milk cheese in Egypt. J. Food Prot. 2018, 81, 226–232. [Google Scholar] [CrossRef] [PubMed]
- Devirgiliis, C.; Zinno, P.; Perozzi, G. Update on antibiotic resistance in foodborne Lactobacillus and Lactococcus species. Front. Microbiol. 2013, 4, 301. [Google Scholar] [CrossRef]
- Duche, R.T.; Singh, A.; Wandhare, A.G.; Sangwan, V.; Sihag, M.K.; Nwagu, T.N.T.; Panwar, H.; et al. Antibiotic resistance in potential probiotic lactic acid bacteria of fermented foods and human origin from Nigeria. BMC Microbiol. 2023, 23. [Google Scholar] [CrossRef]
- Liu, C.M.; Stegger, M.; Aziz, M.; et al. The resistome in raw versus processed milk. Microbiome 2020, 8, 34. [Google Scholar]
- Dabour, N.; Kheadr, E.; Fliss, I.; LaPointe, G. Bacteriocin-producing lactic acid bacteria in dairy ecosystems. Int. Dairy J. 2020, 104, 104646. [Google Scholar]
- European Food Safety Authority; European Centre for Disease Prevention and Control. The European Union Summary Report on antimicrobial resistance in zoonotic and indicator bacteria from humans, animals and food in 2020/2021. EFSA J. 2023, 21, e07867. [Google Scholar] [CrossRef]
- Montel, M.C.; Buchin, S.; Mallet, A.; Delbes-Paus, C.; Vuitton, D.A.; Desmasures, N.; Berthier, F. Traditional cheeses: Rich and diverse microbiota with associated benefits. Int. J. Food Microbiol. 2014, 177, 136–154. [Google Scholar] [CrossRef]
- Parente, E.; Guidone, A.; Zotta, T.; Di Luccia, A. Microbial community dynamics in traditional cheeses. Trends Food Sci. Technol. 2020, 101, 21–35. [Google Scholar]
- Walsh, A.M.; Crispie, F.; Kilcawley, K.; O’Sullivan, O.; Claesson, M.J.; Cotter, P.D. Microbial succession in artisanal cheeses. Nat. Rev. Microbiol. 2020, 18, 351–368. [Google Scholar]
- Wolfe, B.E.; Dutton, R.J. Fermented foods as microbial ecosystems. Cell 2015, 161, 49–55. [Google Scholar] [CrossRef] [PubMed]
- De Filippis, F.; Parente, E.; Ercolini, D. Metagenomics insights into food fermentations. ISME J. 2022, 16, 1884–1896. [Google Scholar] [CrossRef]
- Rantsiou, K.; Kathariou, S.; Winkler, A.; Skandamis, P.; Cocolin, L. Food microbiomes and antimicrobial resistance. Front. Microbiol. 2022, 13, 850276. [Google Scholar]
- Ombarak, R.A.; Hinenoya, A.; Elbagory, A.-R.M.; Yamasaki, S. AMR in E. coli from raw milk and cheese. J. Food Prot. 2018, 81, 226–232. [Google Scholar] [CrossRef]
- Arias, C.A.; Murray, B.E. The rise of the Enterococcus. Nat. Rev. Microbiol. 2012, 10, 266–278. [Google Scholar] [CrossRef]
- Hollenbeck, B.L.; Rice, L.B. Resistance mechanisms in Enterococcus. Virulence 2012, 3, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Lebreton, F.; van Schaik, W.; McGuire, A.M.; et al. Emergence of MDR Enterococcus faecium. Nat. Rev. Microbiol. 2017, 15, 347–361. [Google Scholar]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile genetic elements and AMR. Clin. Microbiol. Rev. 2018, 31, e00088-17. [Google Scholar] [CrossRef]
- EFSA; ECDC. EU Summary Report on AMR. EFSA J. 2023, 21, e07867. [Google Scholar]
- Devirgiliis, C.; Zinno, P.; Perozzi, G. Antibiotic resistance in LAB. Front. Microbiol. 2013, 4, 301. [Google Scholar]
- Campedelli, I.; Mathur, H.; Salvetti, E.; et al. Antibiotic resistance in Lactobacillus. Appl. Environ. Microbiol. 2019, 85, e01738-18. [Google Scholar] [CrossRef]
- Sharma, C.; Gulhane, R.D.; Shinde, T.; et al. Antibiotic resistance in LAB. Front. Microbiol. 2022, 13, 861689. [Google Scholar]
- De Sant'Anna, F.M.; Chakrawarti, A.; Haley, B.J.; et al. The resistome of pasteurized and raw milk cheeses. Int. J. Food Microbiol. 2025, 441, 111333. [Google Scholar] [CrossRef]
- Bonilla-Luque, O.M.; Possas, A.; Cabo, M.L.; et al. Tracking microbial quality and contamination in cheesemaking facilities. Food Control 2023, 148, 104301. [Google Scholar]
- Piras, F.; Siddi, G.; Le Guern, A-S.; et al. Traceability, virulence and antimicrobial resistance of Yersinia enterocolitica in two industrial cheese-making plants. Int. J. Food Microbiol. 2023, 398, 110225. [Google Scholar] [CrossRef] [PubMed]
- Bayrakal, G.M.; Aydin, A. Investigation of various toxigenic genes and antibiotic and disinfectant resistance profiles of Staphylococcus aureus originating from raw milk. Foods 2024, 13, 3448. [Google Scholar] [CrossRef] [PubMed]
- Hassan, H.F.; Haddad, R.; Saidy, L.; et al. Tracking of enrofloxacin antibiotic in the making of common Middle Eastern cheeses. Appl. Food Res. 2021, 1, 100004. [Google Scholar] [CrossRef]
| Milk Type | Sample | LAB (log10 CFU/g) | Mesophilic Bacteria (log10 CFU/g) | Enterobacteriaceae (log10 CFU/g) |
|---|---|---|---|---|
| Goat | G1 | 8.56 ± 0.21 | 8.19 ± 0.13 | 2.72 ± 0.26 |
| Goat | G2 | 8.74 ± 0.30 | 8.39 ± 0.20 | 3.41 ± 0.18 |
| Goat | G3 | 8.34 ± .0.17 | 8.05 ± 0.03 | 1.83 ± 0.08 |
| Goat | G4 | 7.85 ± 0.11 | 7.68 ± 0.24 | 2.49 ± 0.21 |
| Goat | G5 | 7.65 ± 0.14 | 7.51 ± 0.31 | 1.77 ± 0.33 |
| Sheep | S1 | 4.95 ± 0.19 | 5.93 ± 0.05 | 1.94 ± 0.06 |
| Sheep | S2 | 5.04 ± 0.02 | 5.26 ± 0.14 | 2.20 ± 0.15 |
| Sheep | S3 | 5.48 ± 0.21 | 5.85 ± 0.09 | 3.27 ± 0.10 |
| Sheep | S4 | 6.15 ± 0.06 | 6.34 ± 0,12 | 3.11 ± 0.05 |
| Sheep | S5 | 5.48 ± 0.11 | 5.66 ± 0,21 | 3.74 ± 0.27 |
| Species | n isolates | MDR (%) | MDR (n) | Resistance pattern |
|---|---|---|---|---|
| Lactococcus lactis | 28 | 0.0 | 0 | STR-TMP |
| Streptococcus thermophilus | 20 | 0.0 | 0 | AMP-OXA-GEN |
| Lactiplantibacillus plantarum | 18 | 0.0 | 0 | AMP |
| Lacticaseibacillus paracasei | 14 | 0.0 | 0 | AMP-STR-CIP |
| Enterococcus faecalis | 12 | 100.0 | 12 | ERY-GEN-VAN |
| Lactobacillus helveticus | 10 | 0.0 | 0 | AMP-STR |
| Enterococcus faecium | 10 | 100.0 | 10 | AMX-ERY-TET-GEN |
| Lactobacillus delbrueckii | 8 | 0.0 | 0 | AMP-ERY |
| Leuconostoc mesenteroides | 8 | 100.0 | 8 | PEN-TET-STR-VAN |
| Enterococcus durans | 8 | 0.0 | 0 | OXA-GEN |
| Pediococcus pentosaceus | 6 | 100.0 | 6 | AMP-ERY-CIP |
| Staphylococcus equorum | 6 | 100.0 | 6 | AMX-OXA-GEN |
| Staphylococcus saprophyticus | 4 | 0.0 | 0 | OXA |
| Bacillus subtilis | 4 | 0.0 | 0 | CLI-CIP |
| Lacticaseibacillus rhamnosus | 3 | 100.0 | 3 | PEN-AMP-CLI-STR |
| Micrococcus luteus | 3 | 100.0 | 3 | AMX-OXA-GEN-STR-TMP |
| Lactococcus garvieae | 2 | 100.0 | 2 | AMP-ERY-STR |
| Enterobacter cloacae | 2 | 100.0 | 2 | AMP-AMX-GEN-CIP-VAN |
| Escherichia coli | 2 | 100.0 | 2 | ERY-CLI-GEN |
| Citrobacter freundii | 1 | 100.0 | 1 | VAN |
| Klebsiella oxytoca | 1 | 100.0 | 1 | PEN-AMP-VAN |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





