Submitted:
21 February 2026
Posted:
28 February 2026
You are already at the latest version
Abstract
As semi-autonomous organelles, mitochondria function through the coordinated regulation of nuclear genomes and their own genetic material, primarily providing energy for eukaryotic organisms. Currently, high-throughput sequencing technologies have been used to resolve the mitochondrial genomes of various edible fungi. With advances in sequencing technology, species genome characterization has evolved from single genomes to pan-genomes. However, the application of pan-genomes for the analysis of edible mushroom mitochondrial genomes remains unexplored. In this study, we conducted a comparative mitochondrial genome analysis of 31 Hypsizygus marmoreus strains (4 newly sequenced monotypes and 27 public datasets). The results revealed that the mitochondrial genome sizes ranged from 98,284 to 111,087 bp, exhibiting significant structural diversity. This variation is primarily driven by dynamic changes in non-coding regions, particularly intronic polymorphisms in the cox1 gene. This study revealed that tRNA secondary structures exhibit atypical globular and elongated conformations alongside copy number variations. Additionally, codon usage showed a pronounced A/T bias, whereas core respiratory chain genes demonstrated an evolutionary pattern of strong purifying selection. Furthermore, the 31 mitochondrial genomes of H. marmoreus were identified 8 gene rearrangement patterns and 5 genetic clusters, and the pan-genome (220,364 bp, 217 nodes) captured abundant SNPs, InDels and structural variations. This study provides breeding-relevant genetic markers and a genomic framework for germplasm classification, genetic improvement and stress-resilient variety molecular breeding of Hypsizygus marmoreus.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Experimental Strains and Data Sources
2.2. Mycelium Collection
2.3. Genome Sequencing, Assembly, and Annotation
2.4. Repeat Sequence Analysis
2.5. Codon Preference Analysis
2.6. Non-Synonymous and Synonymous Substitution (Ka/Ks) Analysis
2.7. Single Nucleotide Polymorphism Analysis
2.8. Population Structure Analysis
2.9. Mitochondrial Pan-Genome Analysis of H. marmoreus
3. Results
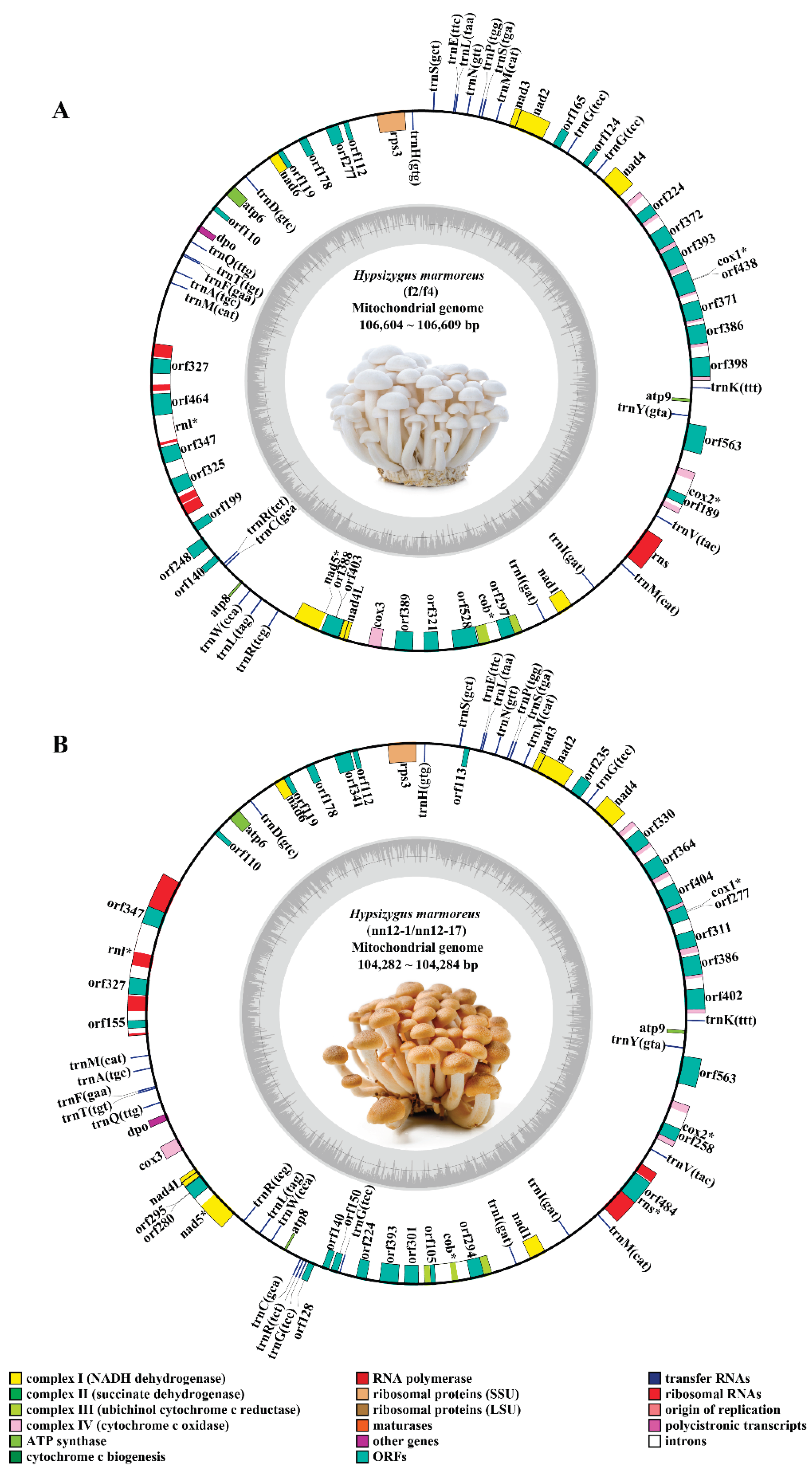
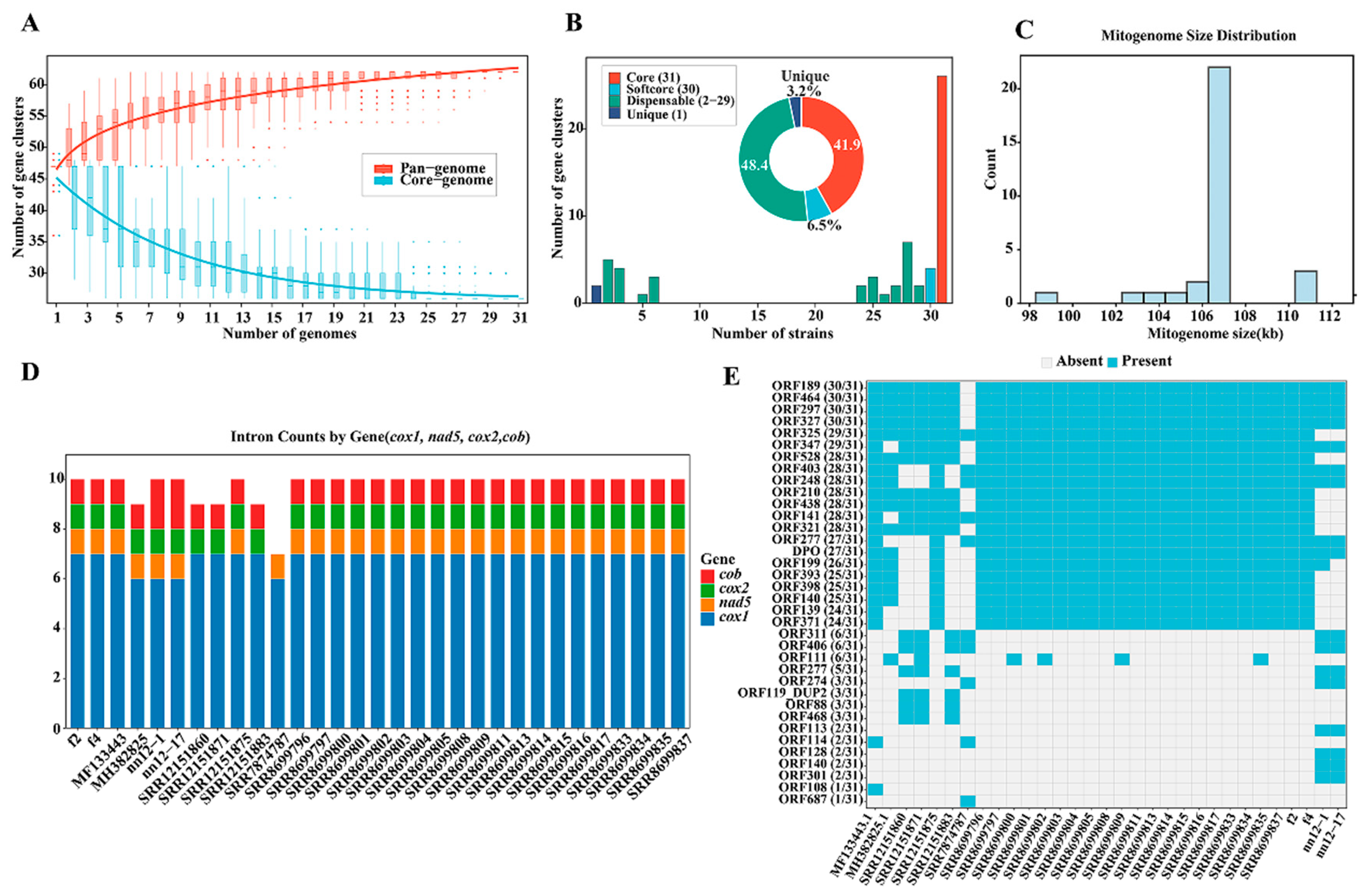
3.1. Sequence Characteristics of the Mitochondrial Pan-Genome of H. marmoreus
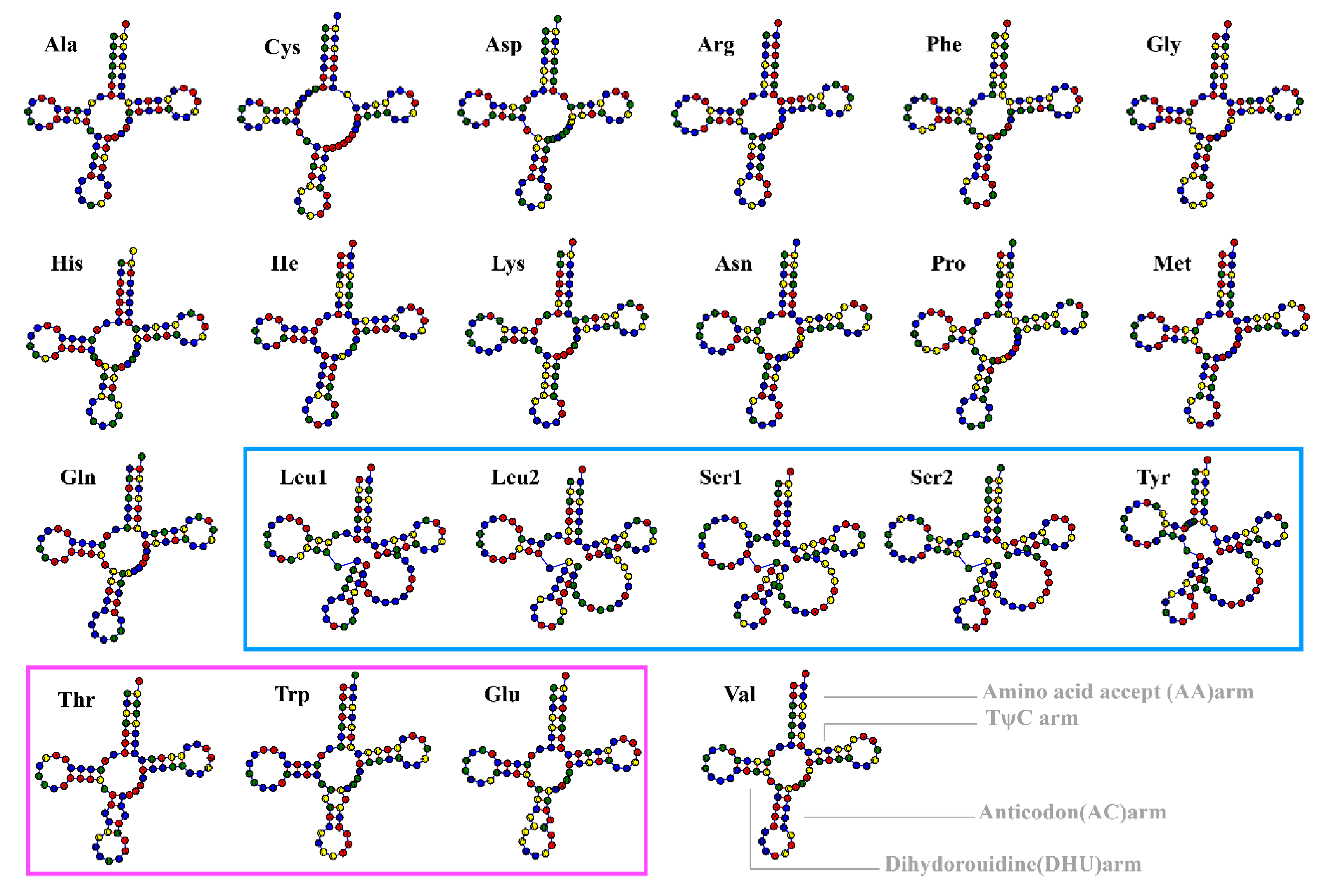
3.2. Secondary Structure Analysis of tRNAs in the Mitochondrial Genome of H. marmoreus
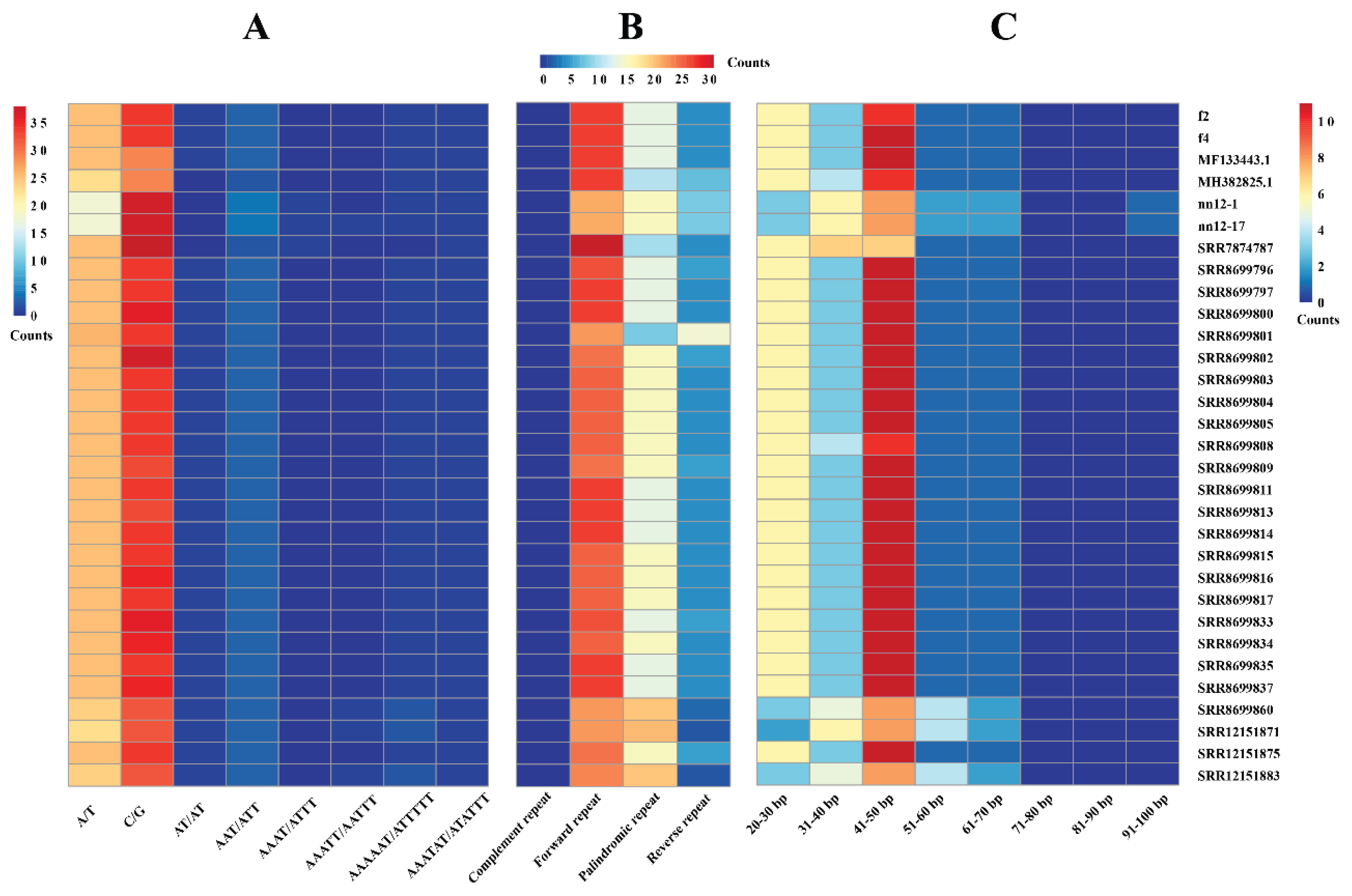
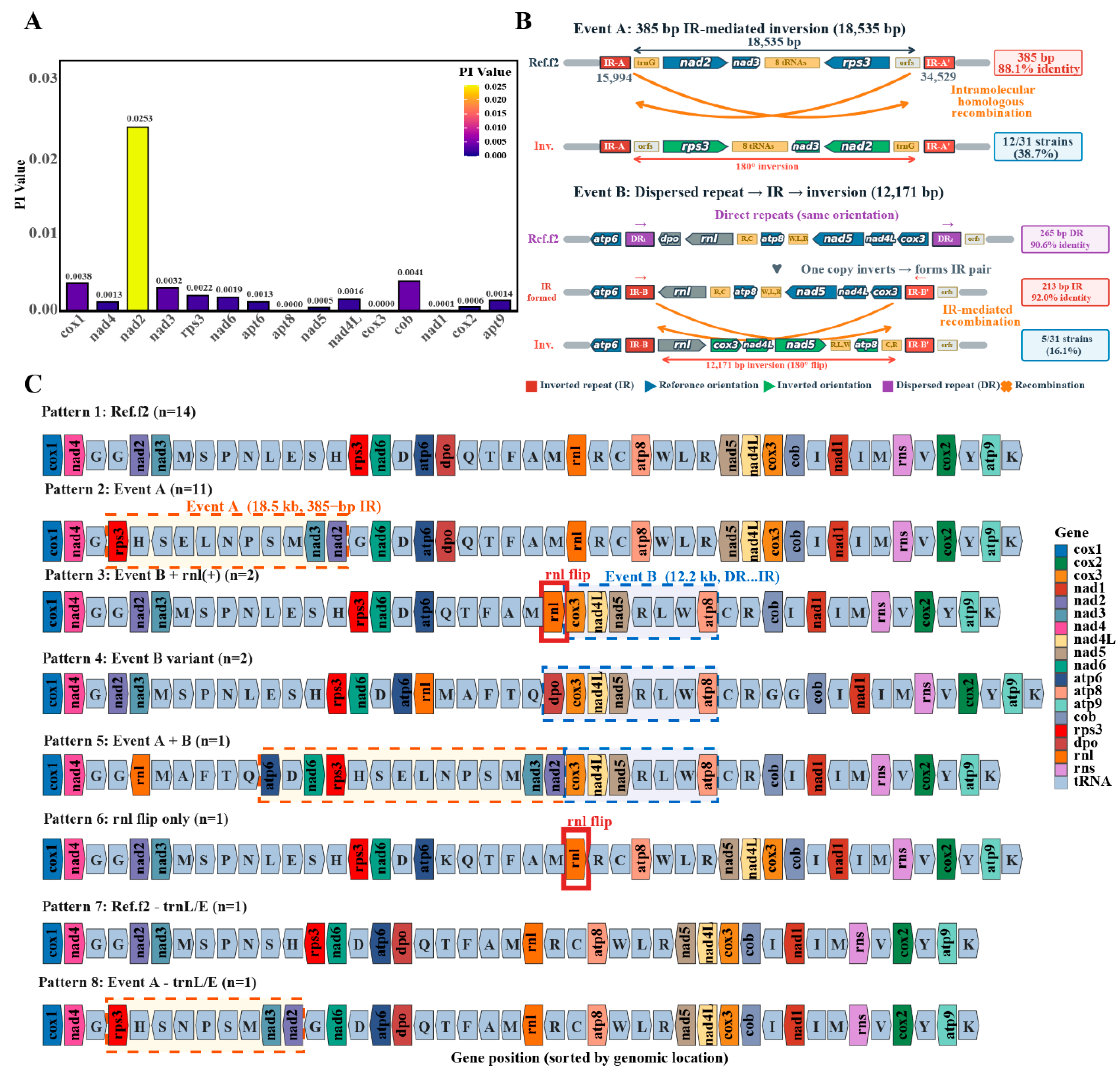
3.3. Analysis of Repeat Sequence Characteristics in the Mitochondria of H. marmoreus
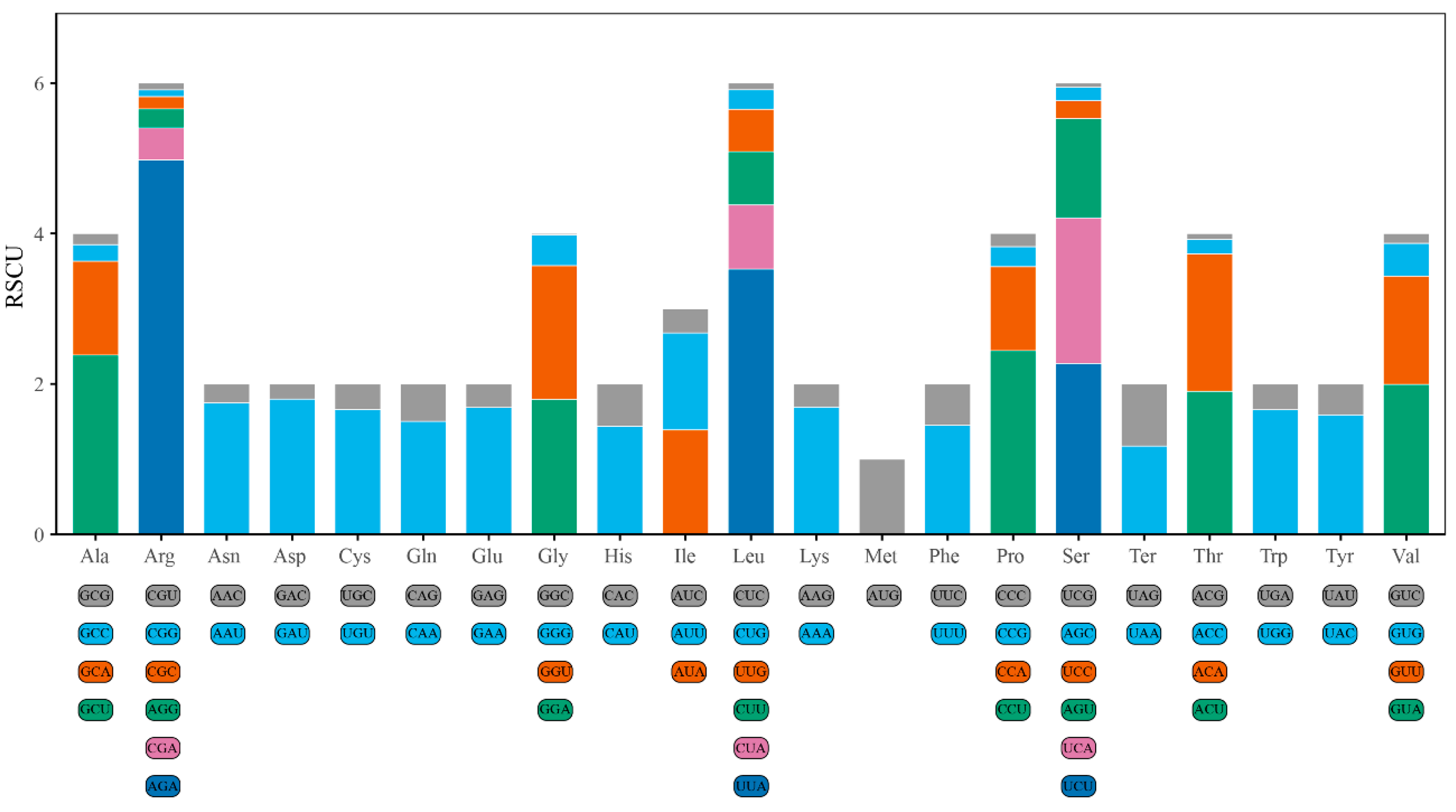
3.4. Analysis of Codon Usage Preferences in the Mitochondrial Genome of H. marmoreus
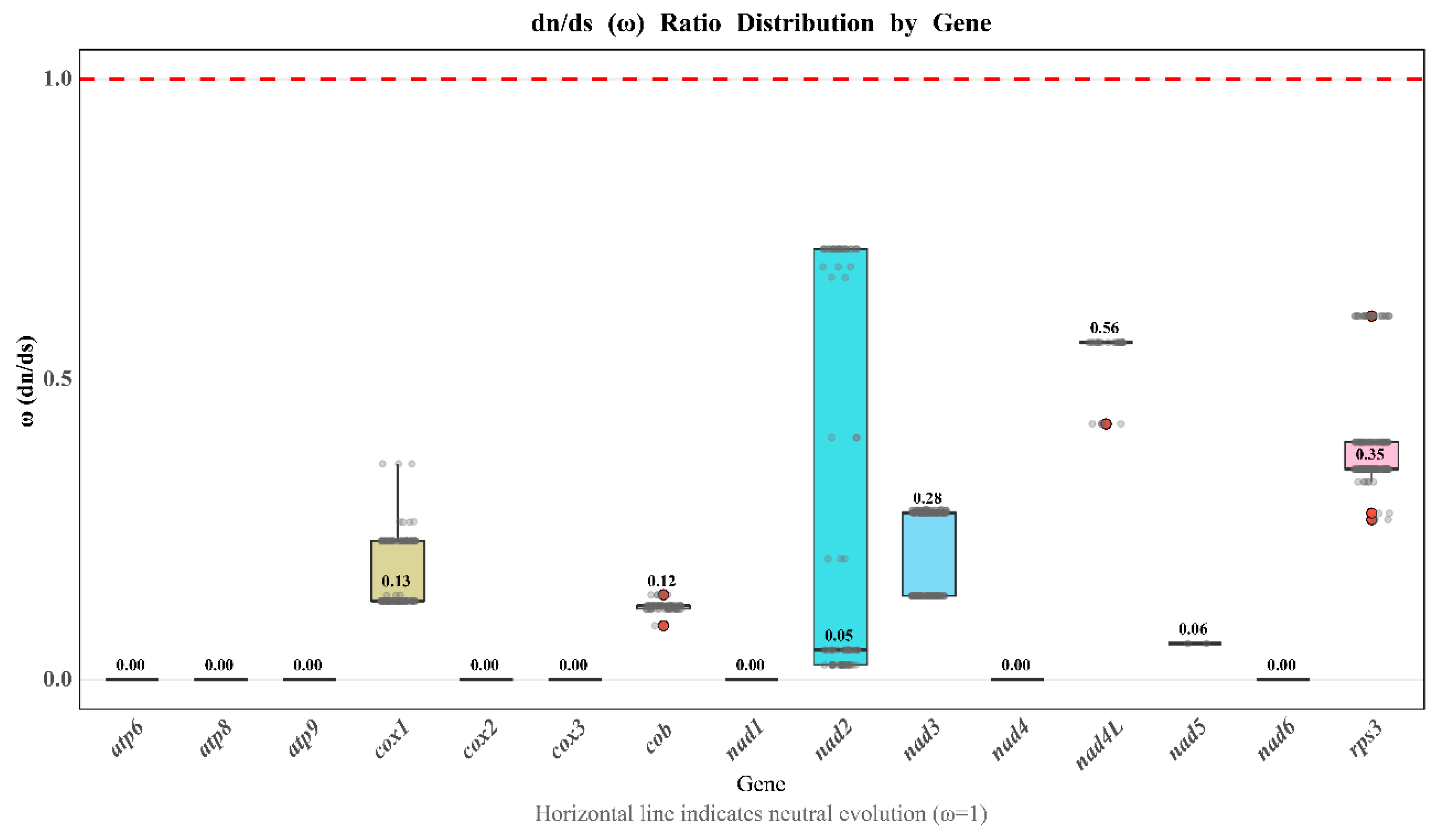
3.5. Analysis of Substitution Rates in the Mitochondrial PCGs of H. marmoreus
3.6. Analysis of Nucleic Acid Diversity in the Mitochondria of H. marmoreus
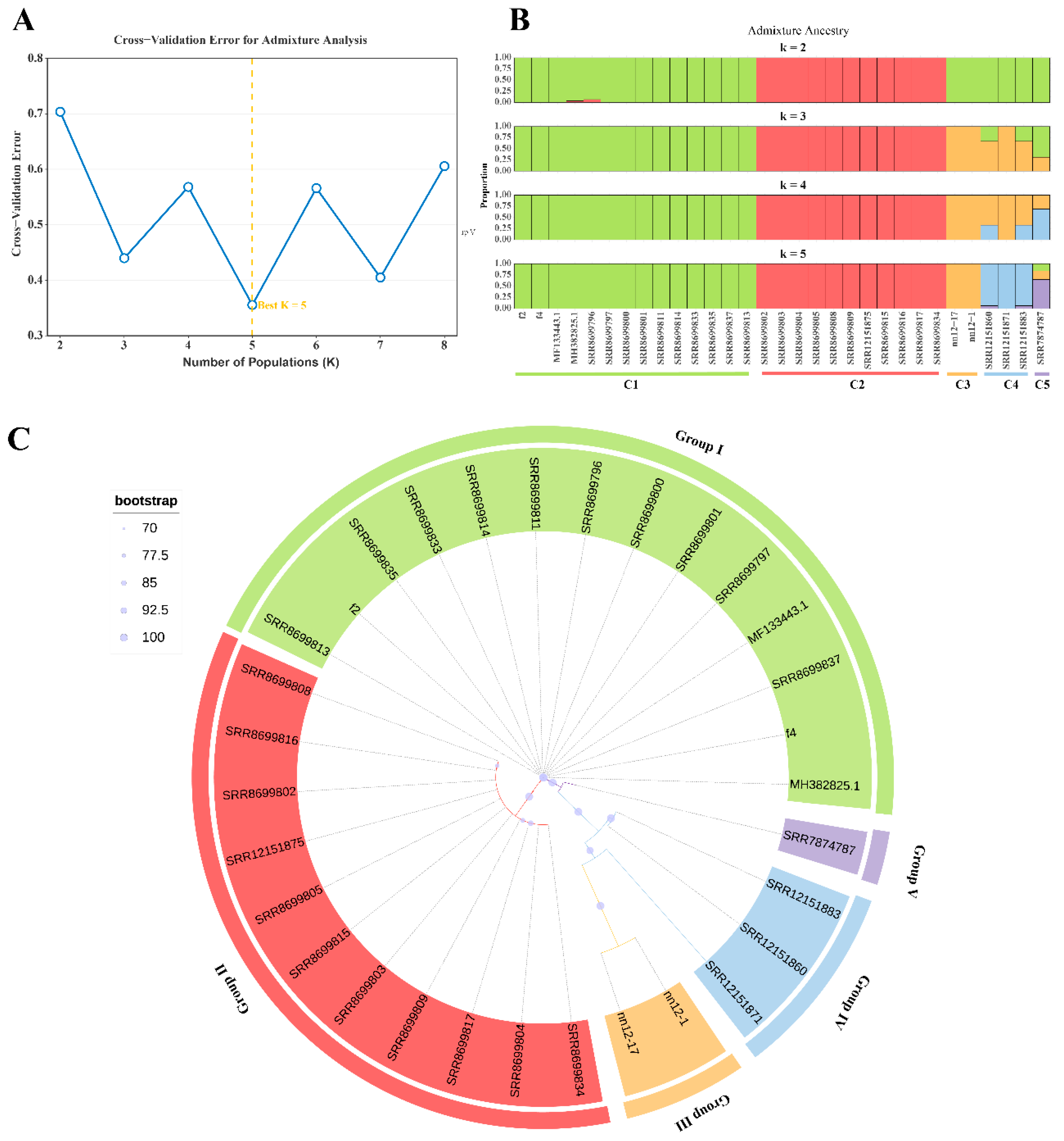
3.7. Phylogenetic and Population Structure Analysis of H. marmoreus
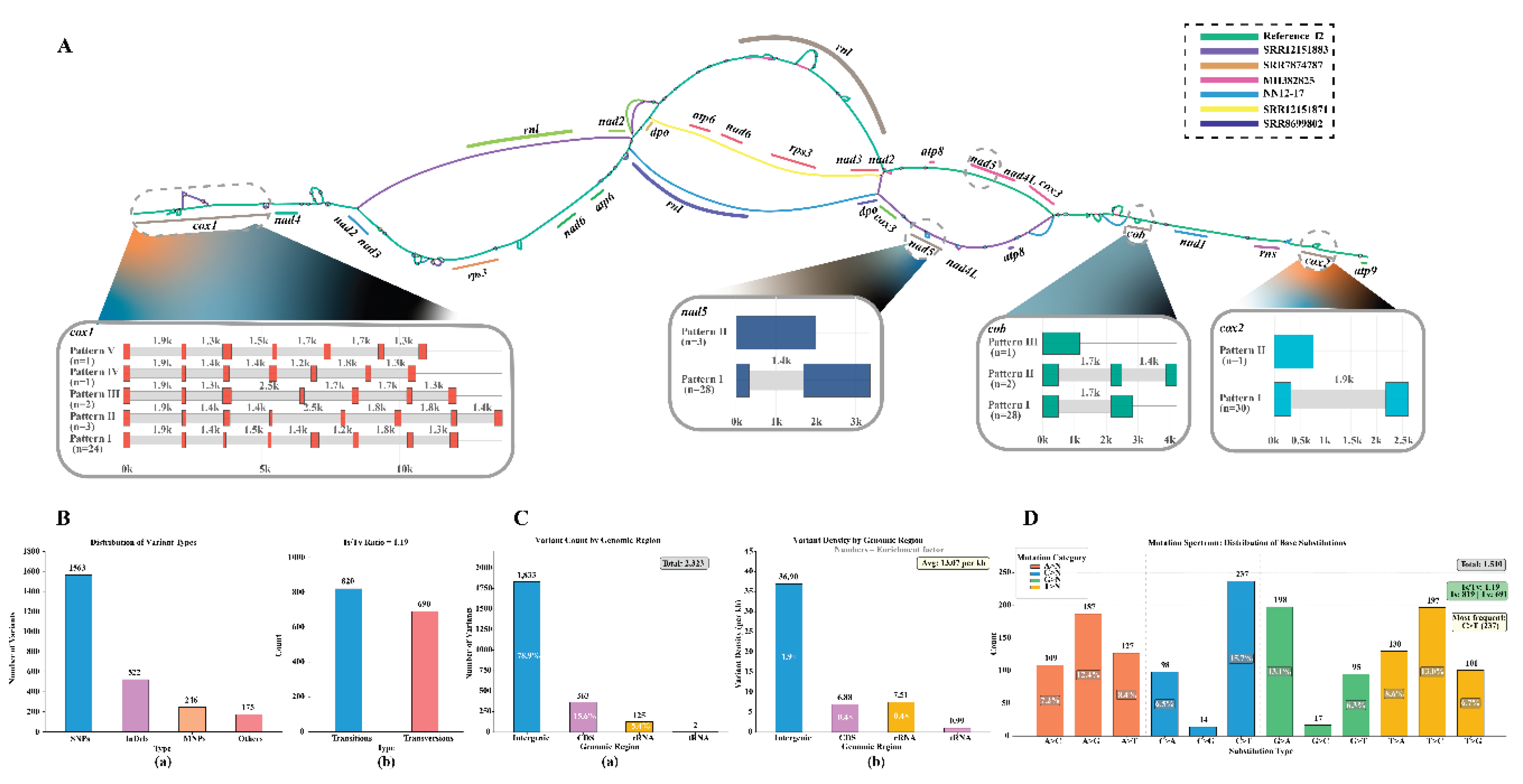
3.8. Analysis of the Pan-Genome of the Mitochondrial DNA of H. marmoreus
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wu, Y.; Peng, Q.; Kang, Q.; Zhou, C.; Li, Y.; Li, J.; Chen, H.; Bao, D. Comparative profiling of volatile compounds in white and brown Hypsizygus marmoreus during fruiting body development and postharvest storage. J. Food Compos. Anal. 2024, 136, 106800. [Google Scholar] [CrossRef]
- Min, B.; Kim, S.; Oh, Y.L.; Kong, W.S.; Park, H.; Cho, H.; Jang, K.Y.; Kim, J.G.; Choi, I.G. Genomic discovery of the hypsin gene and biosynthetic pathways for terpenoids in Hypsizygus marmoreus. BMC Genomics 2018, 19, 789. [Google Scholar] [CrossRef]
- Wang, G.; Chen, L.; Tang, W.; Wang, Y.; Zhang, Q.; Wang, H.; Zhou, X.; Wu, H.; Guo, L.; Dou, M.; et al. Identifying a melanogenesis-related candidate gene by a high-quality genome assembly and population diversity analysis in Hypsizygus marmoreus. J Genet Genomics 2021, 48, 75–87. [Google Scholar] [CrossRef]
- Wang, G.; Wang, Y.; Chen, L.; Wang, H.; Guo, L.; Zhou, X.; Dou, M.; Wang, B.; Lin, J.; Liu, L.; et al. Genetic structure and evolutionary diversity of mating-type (MAT) loci in Hypsizygus marmoreus. IMA Fungus 2021, 12, 35. [Google Scholar] [CrossRef]
- Wu, Y.Y.; Shang, J.J.; Li, Y.; Zhou, C.L.; Hou, D.; Li, J.L.; Tan, Q.; Bao, D.P.; Yang, R.H. The complete mitochondrial genome of the Basidiomycete edible fungus Hypsizygus marmoreus. Mitochondrial DNA B Resour 2018, 3, 1241–1243. [Google Scholar] [CrossRef]
- Wang, G.; Lin, J.; Shi, Y.; Chang, X.; Wang, Y.; Guo, L.; Wang, W.; Dou, M.; Deng, Y.; Ming, R.; et al. Mitochondrial genome in Hypsizygus marmoreus and its evolution in Dikarya. BMC Genomics 2019, 20, 765. [Google Scholar] [CrossRef] [PubMed]
- Sandor, S.; Zhang, Y.; Xu, J. Fungal mitochondrial genomes and genetic polymorphisms. Appl. Microbiol. Biotechnol. 2018, 102, 9433–9448. [Google Scholar] [CrossRef] [PubMed]
- Aguileta, G.; de Vienne, D.M.; Ross, O.N.; Hood, M.E.; Giraud, T.; Petit, E.; Gabaldón, T. High variability of mitochondrial gene order among fungi. Genome Biol Evol 2014, 6, 451–465. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Song, Y.; Ha, B.; Moon, Y.J.; Kim, M.; Ryu, H.; Ro, H.S. Variable Number Tandem Repeats in the Mitochondrial DNA of Lentinula edodes. Genes (Basel) 2019, 10, 542. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Zhang, M.; Chen, M.; Shang, X.; Zhou, F.; Yu, H.; Song, C.; Tan, Q. Transcriptomic Communication between Nucleus and Mitochondria during the Browning Process of Lentinula edodes. J. Agric. Food. Chem. 2024, 72, 23592–23605. [Google Scholar] [CrossRef]
- Wu, P.; Bao, Z.; Tu, W.; Li, L.; Xiong, C.; Jin, X.; Li, P.; Gui, M.; Huang, W.; Li, Q. The mitogenomes of two saprophytic Boletales species (Coniophora) reveals intron dynamics and accumulation of plasmid-derived and non-conserved genes. Computational and Structural Biotechnology Journal 2021, 19, 401–414. [Google Scholar] [CrossRef]
- Tan, H.; Yu, Y.; Fu, Y.; Liu, T.; Wang, Y.; Peng, W.; Wang, B.; Chen, J. Comparative analyses of Flammulina filiformis mitochondrial genomes reveal high length polymorphism in intergenic regions and multiple intron gain/loss in cox1. Int. J. Biol. Macromol. 2022, 221, 1593–1605. [Google Scholar] [CrossRef]
- Zhang, M.-Z.; Xu, J.-P.; Callac, P.; Chen, M.-Y.; Wu, Q.; Wach, M.; Mata, G.; Zhao, R.-L. Insight into the evolutionary and domesticated history of the most widely cultivated mushroom Agaricus bisporus via mitogenome sequences of 361 global strains. BMC Genomics 2023, 24, 182. [Google Scholar] [CrossRef]
- Brankovics, B.; Kulik, T.; Sawicki, J.; Bilska, K.; Zhang, H.; de Hoog, G.S.; van der Lee, T.A.; Waalwijk, C.; van Diepeningen, A.D. First steps towards mitochondrial pan-genomics: detailed analysis of Fusarium graminearum mitogenomes. PeerJ 2018, 6, e5963. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Li, C.; Kuang, L.; Wu, X.; Xie, K.; Zhu, A.; Xu, Q.; Larkin, R.M.; Zhou, Y.; Deng, X.; et al. Pan-mitogenomics reveals the genetic basis of cytonuclear conflicts in citrus hybridization, domestication, and diversification. Proc Natl Acad Sci U S A 2022, 119, e2206076119. [Google Scholar] [CrossRef]
- van Westerhoven, A.C.; Dijkstra, J.; Aznar Palop, J.L.; Wissink, K.; Bell, J.; Kema, G.H.J.; Seidl, M.F. Frequent genetic exchanges revealed by a pan-mitogenome graph of a fungal plant pathogen. mBio 2024, e0275824. [Google Scholar] [CrossRef] [PubMed]
- Kolmogorov, M.; Yuan, J.; Lin, Y.; Pevzner, P.A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 2019, 37, 540–546. [Google Scholar] [CrossRef]
- Wick, R.R.; Schultz, M.B.; Zobel, J.; Holt, K.E. Bandage: interactive visualization of de novo genome assemblies. Bioinformatics 2015, 31, 3350–3352. [Google Scholar] [CrossRef]
- Jin, J.J.; Yu, W.B.; Yang, J.B.; Song, Y.; dePamphilis, C.W.; Yi, T.S.; Li, D.Z. GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol 2020, 21, 241. [Google Scholar] [CrossRef] [PubMed]
- Lang, B.F.; Beck, N.; Prince, S.; Sarrasin, M.; Rioux, P.; Burger, G. Mitochondrial genome annotation with MFannot: a critical analysis of gene identification and gene model prediction. Front Plant Sci 2023, 14, 1222186. [Google Scholar] [CrossRef]
- Bernt, M.; Donath, A.; Jühling, F.; Externbrink, F.; Florentz, C.; Fritzsch, G.; Pütz, J.; Middendorf, M.; Stadler, P.F. MITOS: improved de novo metazoan mitochondrial genome annotation. Mol Phylogenet Evol 2013, 69, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A Program for Improved Detection of Transfer RNA Genes in Genomic Sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Ponty, Y.; Leclerc, F. Drawing and Editing the Secondary Structure(s) of RNA. In RNA Bioinformatics; Picardi, E., Ed.; Springer New York: New York, NY, 2015; pp. 63–100. [Google Scholar]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M. MISA-web: a web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef]
- Xiang, C.-Y.; Gao, F.; Jakovlić, I.; Lei, H.-P.; Hu, Y.; Zhang, H.; Zou, H.; Wang, G.-T.; Zhang, D. Using PhyloSuite for molecular phylogeny and tree-based analyses. iMeta 2023, 2, e87. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Xu, B.; Yang, Z. PAMLX: a graphical user interface for PAML. Mol Biol Evol 2013, 30, 2723–2724. [Google Scholar] [CrossRef]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol Biol Evol 2017, 34, 3299–3302. [Google Scholar] [CrossRef]
- Page, A.J.; Taylor, B.; Delaney, A.J.; Soares, J.; Seemann, T.; Keane, J.A.; Harris, S.R. SNP-sites: rapid efficient extraction of SNPs from multi-FASTA alignments. Microb Genom 2016, 2, e000056. [Google Scholar] [CrossRef]
- Alexander, D.H.; Novembre, J.; Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res 2009, 19, 1655–1664. [Google Scholar] [CrossRef]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol Biol Evol 2020, 37, 1530–1534. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 2024, 52, w78–w82. [Google Scholar] [CrossRef]
- Emms, D.M.; Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol 2019, 20, 238. [Google Scholar] [CrossRef]
- Hickey, G.; Monlong, J.; Ebler, J.; Novak, A.M.; Eizenga, J.M.; Gao, Y.; Abel, H.J.; Antonacci-Fulton, L.L.; Asri, M.; Baid, G.; et al. Pangenome graph construction from genome alignments with Minigraph-Cactus. Nat. Biotechnol. 2024, 42, 663–673. [Google Scholar] [CrossRef] [PubMed]
- Hickey, G.; Heller, D.; Monlong, J.; Sibbesen, J.A.; Sirén, J.; Eizenga, J.; Dawson, E.T.; Garrison, E.; Novak, A.M.; Paten, B. Genotyping structural variants in pangenome graphs using the vg toolkit. Genome Biology 2020, 21, 35. [Google Scholar] [CrossRef] [PubMed]
- Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 2011, 27, 2987–2993. [Google Scholar] [CrossRef] [PubMed]
- Guarracino, A.; Heumos, S.; Nahnsen, S.; Prins, P.; Garrison, E. ODGI: understanding pangenome graphs. Bioinformatics 2022, 38, 3319–3326. [Google Scholar] [CrossRef]
- Zeb, U.; Aziz, T.; Azizullah, A.; Zan, X.Y.; Khan, A.A.; Bacha, S.A.S.; Cui, F.J. Complete mitochondrial genomes of edible mushrooms: features, evolution, and phylogeny. Physiol. Plant. 2024, 176, e14363. [Google Scholar] [CrossRef]
- Tang, J.; Zhang, L.; Su, J.; Ye, Q.; Li, Y.; Liu, D.; Cui, H.; Zhang, Y.; Ye, Z. Insights into Fungal Mitochondrial Genomes and Inheritance Based on Current Findings from Yeast-like Fungi. J Fungi (Basel) 2024, 10, 441. [Google Scholar] [CrossRef]
- Kim, S.; Eom, H.; Nandre, R.; Choi, Y.J.; Lee, H.; Ryu, H.; Ro, H. S. Comparative structural analysis on the mitochondrial DNAs from various strains of Lentinula edodes. Frontiers in Microbiology 2022, 13, 1034387. [Google Scholar] [CrossRef]
- Gao, W.; Chen, X.; He, J.; Sha, A.; Luo, Y.; Xiao, W.; Xiong, Z.; Li, Q. Intraspecific and interspecific variations in the synonymous codon usage in mitochondrial genomes of 8 pleurotus strains. BMC Genomics 2024, 25, 456. [Google Scholar] [CrossRef]
- Li, Q.; Xiang, D.; Wan, Y.; Wu, Q.; Wu, X.; Ma, C.; Song, Y.; Zhao, G.; Huang, W. The complete mitochondrial genomes of five important medicinal Ganoderma species: Features, evolution, and phylogeny. Int. J. Biol. Macromol. 2019, 139, 397–408. [Google Scholar] [CrossRef]
- Liu, C.; Li, W.Y.; Zheng, L.X.; Dao, M.; Chen, H.H.; Han, L.H. Comparative mitogenomic analysis reveals variations and evolution of ectomycorrhizal fungal Strobilomyces. IMA Fungus 2025, 16, e141848. [Google Scholar] [CrossRef]
- Kang, M.; Wu, H.; Liu, H.; Liu, W.; Zhu, M.; Han, Y.; Liu, W.; Chen, C.; Song, Y.; Tan, L.; et al. The pan-genome and local adaptation of Arabidopsis thaliana. Nat. Commun. 2023, 14, 6259. [Google Scholar] [CrossRef] [PubMed]
- Waneka, G.; Svendsen, J.M.; Havird, J.C.; Sloan, D.B. Mitochondrial mutations in Caenorhabditis elegans show signatures of oxidative damage and an AT-bias. Genetics 2021, 219, iyab116. [Google Scholar] [CrossRef] [PubMed]
- Boore, J.L. Animal mitochondrial genomes. Nucleic Acids Res. 1999, 27, 1767–1780. [Google Scholar] [CrossRef]
- Gualberto, J.M.; Newton, K.J. Plant Mitochondrial Genomes: Dynamics and Mechanisms of Mutation. Annu Rev Plant Biol 2017, 68, 225–252. [Google Scholar] [CrossRef] [PubMed]
- Tao, G.; Ahrendt, S.; Miyauchi, S.; Zhu, X.; Peng, H.; Labutti, K.; Clum, A.; Hayes, R.; Chain, P.S.G.; Grigoriev, I.V.; et al. Characterisation and comparative analysis of mitochondrial genomes of false, yellow, black and blushing morels provide insights on their structure and evolution. IMA Fungus 2025, 16, e138363. [Google Scholar] [CrossRef]
- Al-Reedy, R.M.; Malireddy, R.; Dillman, C.B.; Kennell, J.C. Comparative analysis of Fusarium mitochondrial genomes reveals a highly variable region that encodes an exceptionally large open reading frame. Fungal genetics and biology: FG & B 2012, 49, 2–14. [Google Scholar] [CrossRef]
- Basse, C.W. Mitochondrial inheritance in fungi. Current opinion in microbiology 2010, 13, 712–719. [Google Scholar] [CrossRef]
- Deng, Y.; Hsiang, T.; Li, S.; Lin, L.; Wang, Q.; Chen, Q.; Xie, B.; Ming, R. Comparison of the Mitochondrial Genome Sequences of Six Annulohypoxylon stygium Isolates Suggests Short Fragment Insertions as a Potential Factor Leading to Larger Genomic Size. Front Microbiol 2018, 9, 2079. [Google Scholar] [CrossRef] [PubMed]
- Kolesnikova, A.I.; Putintseva, Y.A.; Simonov, E.P.; Biriukov, V.V.; Oreshkova, N.V.; Pavlov, I.N.; Sharov, V.V.; Kuzmin, D.A.; Anderson, J.B.; Krutovsky, K.V. Mobile genetic elements explain size variation in the mitochondrial genomes of four closely-related Armillaria species. BMC Genomics 2019, 20, 351. [Google Scholar] [CrossRef]
- Barroso, G.; Bois, F.; Labarère, J. Duplication of a truncated paralog of the family B DNA polymerase gene Aa-polB in the Agrocybe aegerita mitochondrial genome. Appl. Environ. Microbiol. 2001, 67, 1739–1743. [Google Scholar] [CrossRef]
- Adams, K.L.; Palmer, J.D. Evolution of mitochondrial gene content: gene loss and transfer to the nucleus. Molecular phylogenetics and evolution 2003, 29, 380–395. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wei, S.; Wu, S.; Tang, J.; Wei, J.; Liu, Z.; Qi, L. Characterization and phylogenetic analysis of the complete mitochondrial genome sequence of Auricularia villosula, an edible wild mushroom in China. Biologia 2023, 78, 3713–3723. [Google Scholar] [CrossRef]
- Li, Q.; Zhang, T.; Li, L.; Bao, Z.; Tu, W.; Xiang, P.; Wu, Q.; Li, P.; Cao, M.; Huang, W. Comparative Mitogenomic Analysis Reveals Intraspecific, Interspecific Variations and Genetic Diversity of Medical Fungus Ganoderma. Journal of Fungi 2022, 8, 781. [Google Scholar] [CrossRef]
- Wang, X.; Wang, G.; Tao, J.; Guo, Z.; Xu, G.; Li, J.; Kang, J.; Zuo, Q.; Liu, H.; Li, Q. Comparative analysis of mitochondrial genomes in lycoperdaceae fungi reveals intron dynamics and phylogenetic relationships. BMC Genomics 2025, 26, 742. [Google Scholar] [CrossRef]
- Goddard, M.R.; Burt, A. Recurrent invasion and extinction of a selfish gene. Proceedings of the National Academy of Sciences 1999, 96, 13880–13885. [Google Scholar] [CrossRef]
- Mukhopadhyay, J.; Wai, A.; Lang, B.F.; Hausner, G. Characterization of the mitochondrial genomes for Ophiostomaips and related taxa from various geographic origins and related species: large intron-rich genomes and complex intron arrangements. IMA Fungus 2025, 16, e159349. [Google Scholar] [CrossRef] [PubMed]
| Number | Name of Species | Size(bp) | Introns | Exons | Total Genes | Genes for Protein Coding | No. of rRNA | No. of tRNA |
GC Contents % | Accession No. | Source information |
| 1 | H. marmoreus | 104,282 | 10 | 14 | 78 | 48 | 2 | 28 | 31.52 | nn12-1 | This study |
| 2 | H. marmoreus | 104,284 | 10 | 14 | 74 | 46 | 2 | 28 | 31.52 | nn12-17 | |
| 3 | H. marmoreus | 106,605 | 10 | 14 | 75 | 47 | 2 | 27 | 31.72 | f2 | |
| 4 | H. marmoreus | 106,609 | 10 | 14 | 76 | 47 | 2 | 27 | 31.72 | f4 | |
| 5 | H. marmoreus | 98,284 | 7 | 9 | 65 | 39 | 2 | 28 | 31.75 | SRR7874787 | [2] |
| 6 | H. marmoreus | 102,752 | 9 | 13 | 72 | 44 | 2 | 27 | 31.86 | MH382825.1 | [6] |
| 7 | H. marmoreus | 106,518 | 10 | 14 | 77 | 49 | 2 | 27 | 31.67 | MF133443.1 | |
| 8 | H. marmoreus | 106,608 | 10 | 14 | 75 | 47 | 2 | 27 | 31.72 | SRR8699805 | |
| 9 | H. marmoreus | 106,718 | 10 | 14 | 75 | 47 | 2 | 27 | 31.77 | SRR8699833 | |
| 10 | H. marmoreus | 105,706 | 10 | 14 | 74 | 47 | 2 | 25 | 31.64 | SRR8699796 | |
| 11 | H. marmoreus | 106,610 | 10 | 14 | 75 | 47 | 2 | 27 | 31.72 | SRR8699803 | |
| 12 | H. marmoreus | 106,773 | 10 | 14 | 75 | 47 | 2 | 27 | 31.75 | SRR8699801 | |
| 13 | H. marmoreus | 106,699 | 10 | 14 | 76 | 48 | 2 | 27 | 31.77 | SRR8699802 | |
| 14 | H. marmoreus | 106,677 | 10 | 14 | 75 | 47 | 2 | 27 | 31.75 | SRR8699837 | |
| 15 | H. marmoreus | 106,604 | 10 | 14 | 75 | 47 | 2 | 27 | 31.72 | SRR8699817 | |
| 16 | H. marmoreus | 106,606 | 10 | 14 | 75 | 47 | 2 | 27 | 31.73 | SRR8699808 | |
| 17 | H. marmoreus | 106,608 | 10 | 14 | 75 | 47 | 2 | 27 | 31.72 | SRR8699813 | |
| 18 | H. marmoreus | 106,711 | 10 | 14 | 76 | 48 | 2 | 27 | 31.75 | SRR8699800 | |
| 19 | H. marmoreus | 106,612 | 10 | 14 | 76 | 48 | 2 | 27 | 31.73 | SRR8699835 | |
| 20 | H. marmoreus | 106,699 | 10 | 14 | 75 | 47 | 2 | 27 | 31.74 | SRR8699816 | |
| 21 | H. marmoreus | 106,691 | 10 | 14 | 75 | 47 | 2 | 27 | 31.75 | SRR8699834 | |
| 22 | H. marmoreus | 106,609 | 10 | 14 | 75 | 47 | 2 | 27 | 31.72 | SRR8699814 | |
| 23 | H. marmoreus | 106,620 | 10 | 14 | 75 | 47 | 2 | 27 | 31.73 | SRR8699815 | |
| 24 | H. marmoreus | 105,589 | 10 | 14 | 74 | 48 | 2 | 25 | 31.61 | SRR8699809 | |
| 25 | H. marmoreus | 106,609 | 10 | 14 | 69 | 41 | 2 | 27 | 31.72 | SRR8699804 | |
| 26 | H. marmoreus | 106,603 | 10 | 14 | 75 | 47 | 2 | 27 | 31.72 | SRR8699797 | |
| 27 | H. marmoreus | 106,602 | 10 | 14 | 69 | 41 | 2 | 27 | 31.72 | SRR8699811 | |
| 28 | H. marmoreus | 111,087 | 9 | 12 | 74 | 46 | 2 | 27 | 32.12 | SRR12151860 | [4] |
| 29 | H. marmoreus | 106,670 | 10 | 14 | 75 | 47 | 2 | 27 | 31.74 | SRR12151875 | |
| 30 | H. marmoreus | 111,031 | 9 | 12 | 75 | 47 | 2 | 27 | 32.11 | SRR12151871 | |
| 31 | H. marmoreus | 111,037 | 9 | 12 | 74 | 46 | 2 | 27 | 32.11 | SRR12151883 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





