Submitted:
24 February 2026
Posted:
25 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Material and Methods
2.1. Soil Sample Collection
2.2. Isolation and Screening of Nitrogen-Fixing Bacteria
2.3. Molecular Identification and Phylogenetic Analysis
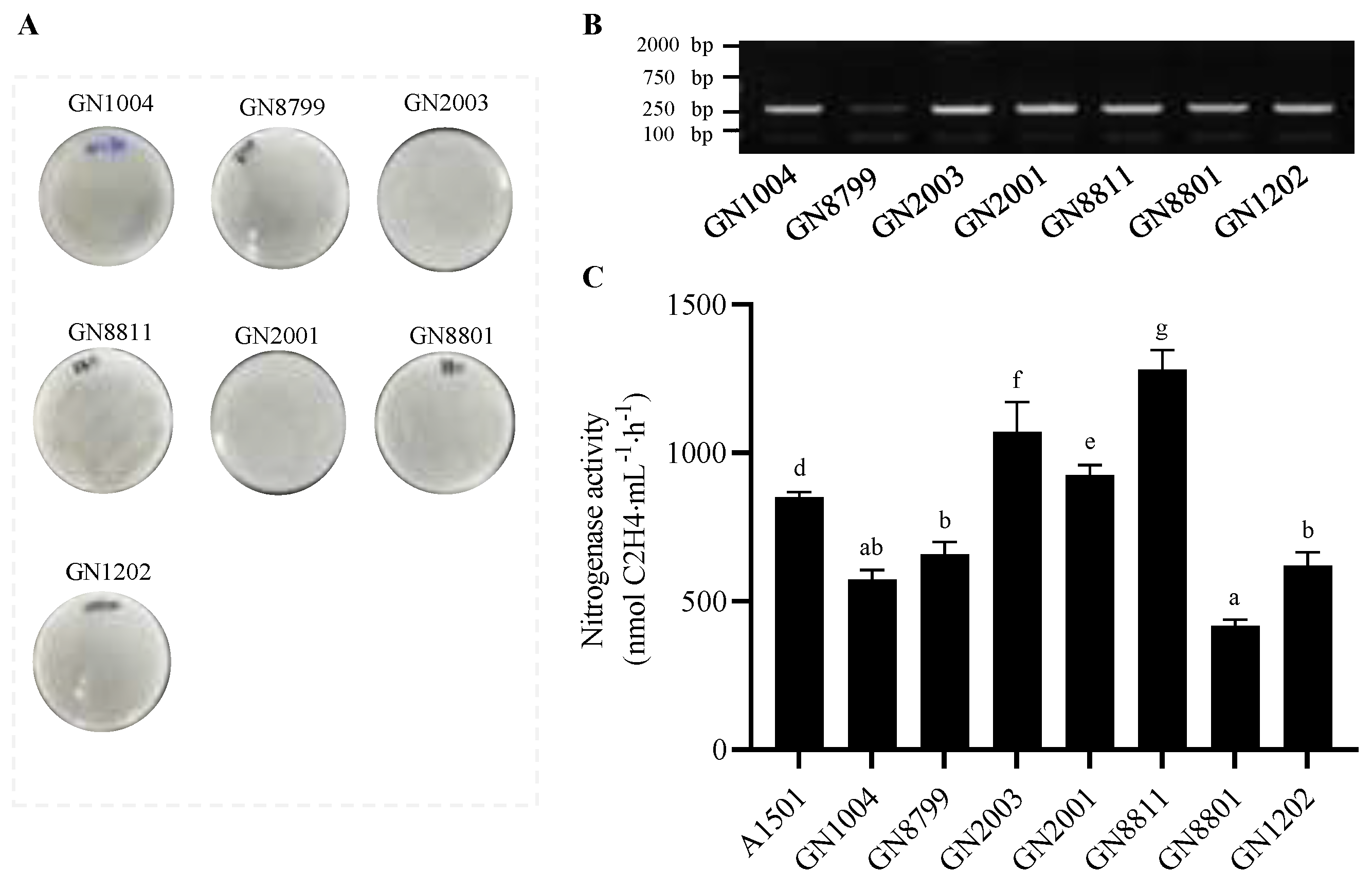
2.4. Detection of the Nifh Gene and Assay of Nitrogenase Activity
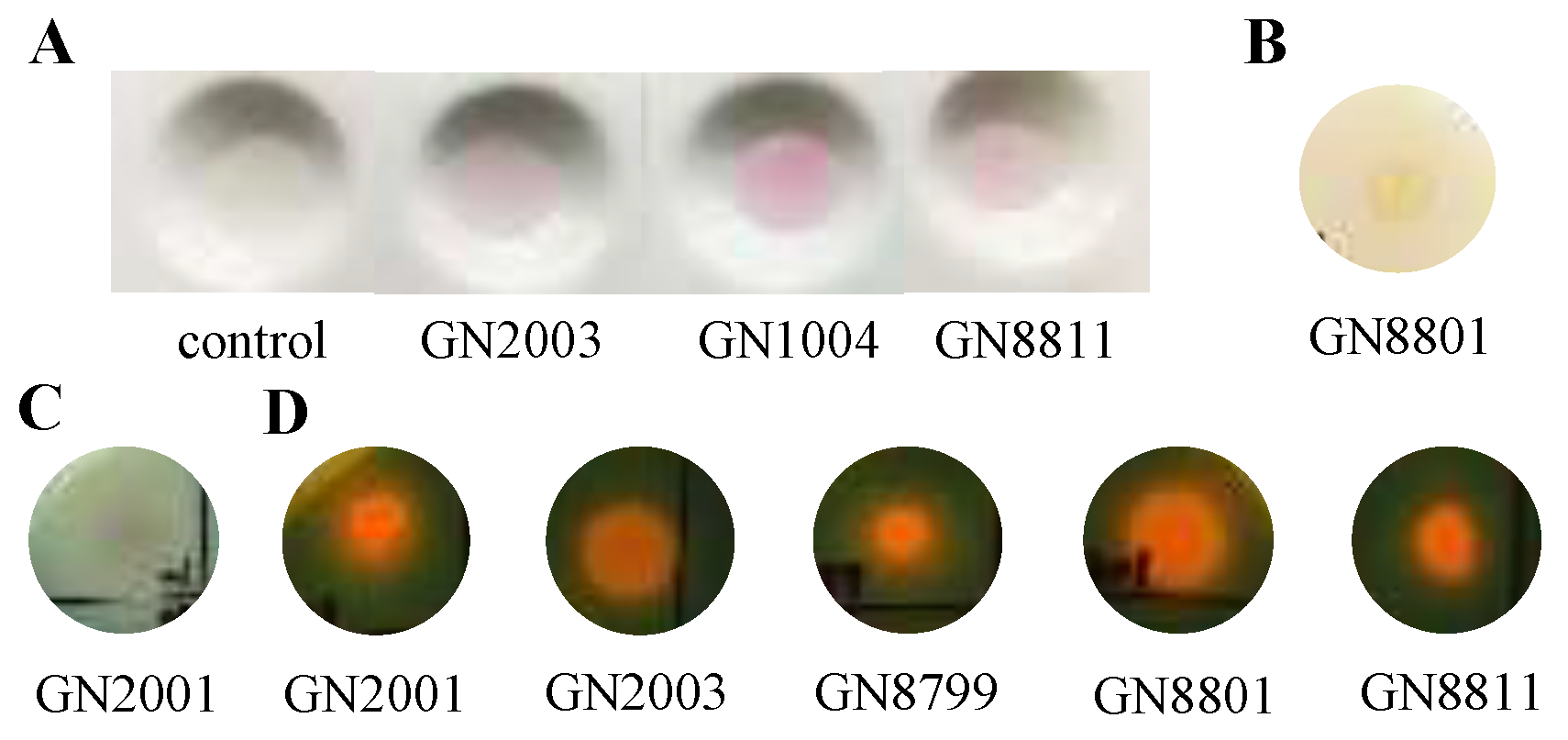
2.5. In Vitro Evaluation of Plant Growth-Promoting (pgp) Traits
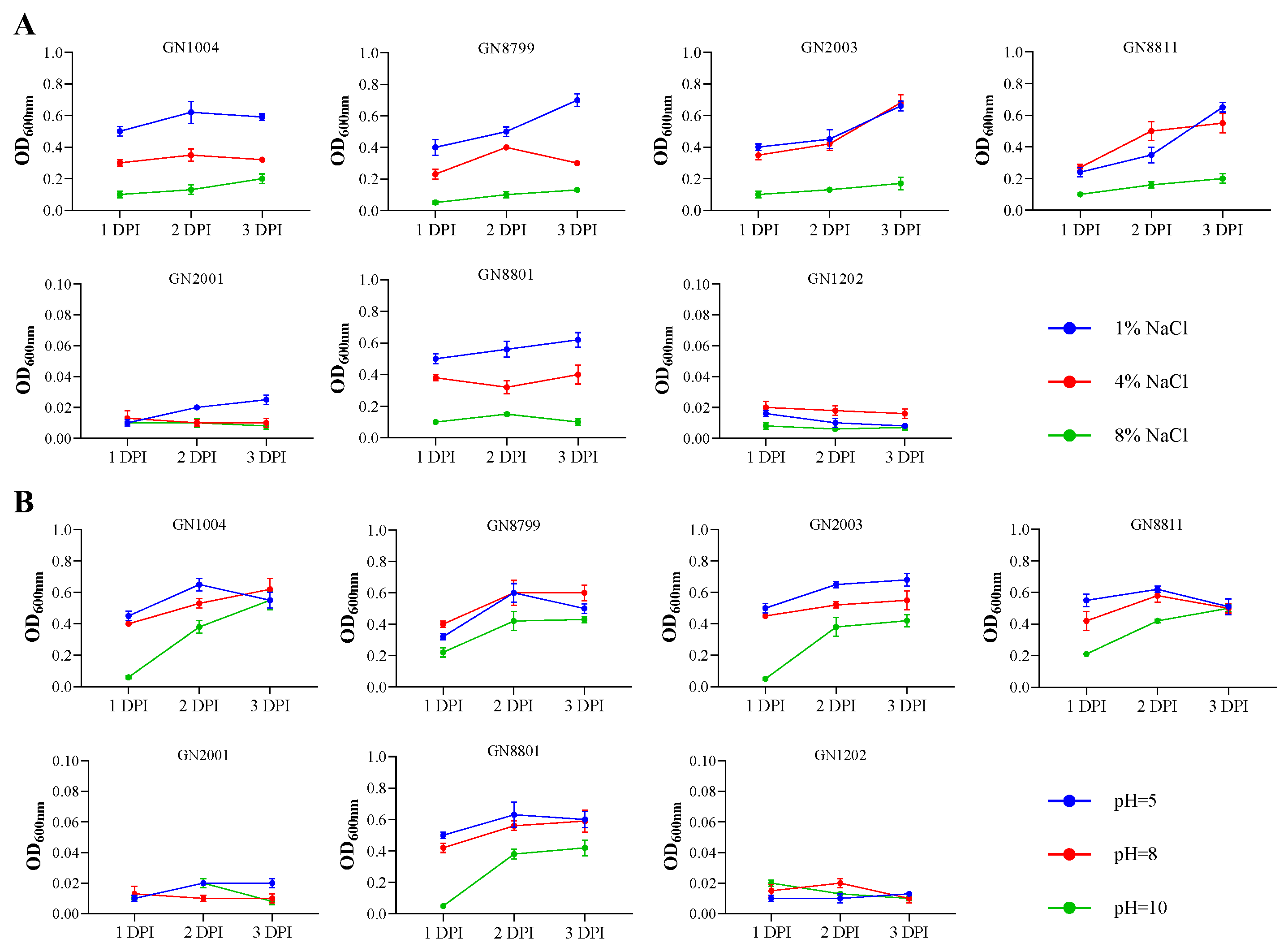
2.6. Evaluation of Environmental Stress Tolerance
2.7. Genetic Modification
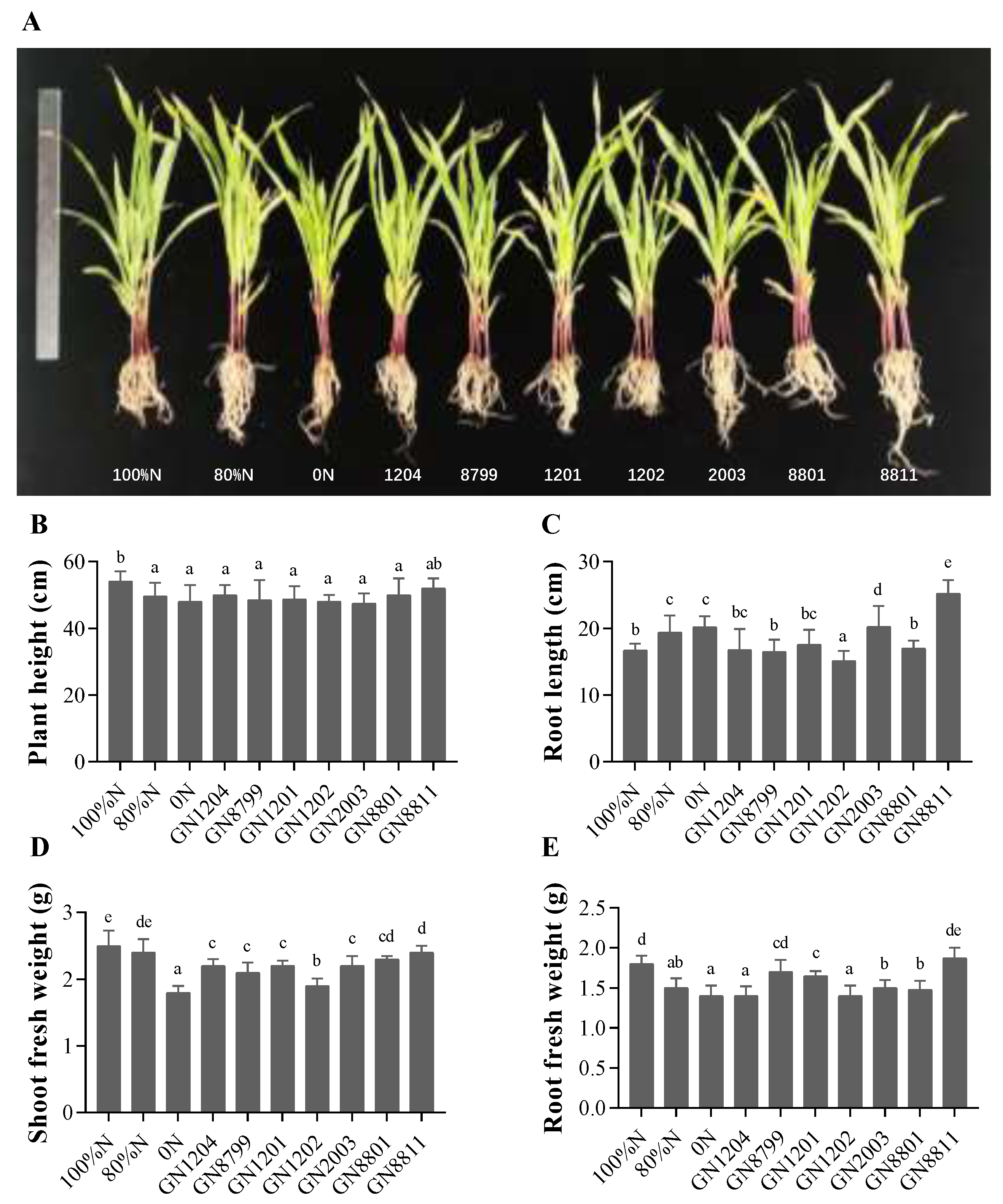
2.8. Pot Experiment to Evaluate Plant Growth Promotion
2.9. Data Visualization and Statistical Analysis
3. Results
3.1. Isolation and Nitrogen-Fixing Activity Assessment of Nitrogen-Fixing Bacteria from Maize Rhizosphere
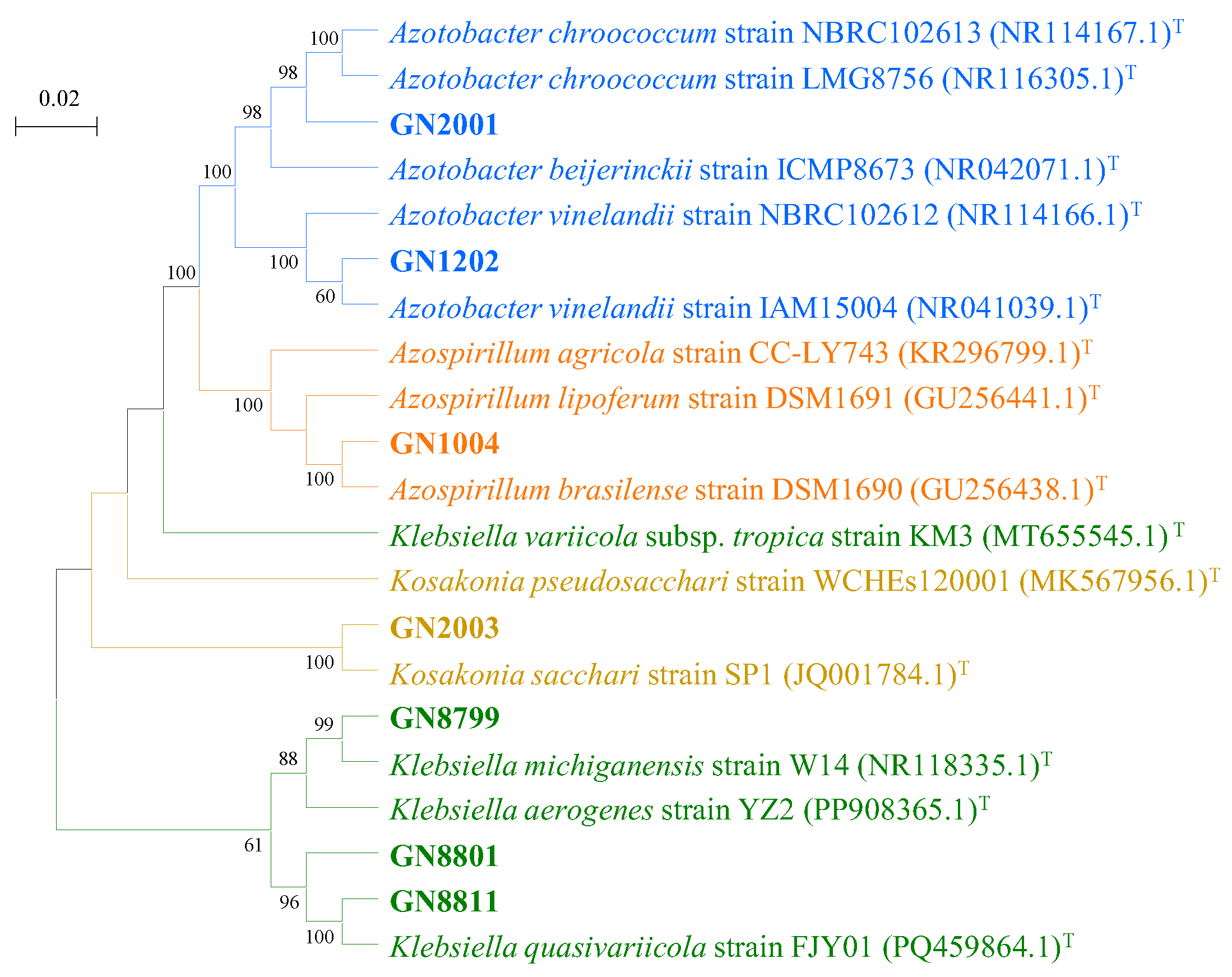
3.2. Phylogenetic Analysis of Isolated Nitrogen-Fixing Bacteria
3.3. Evaluation of Plant Growth-Promoting Traits of the Isolated Strains
3.4. Physiological Characterization of Acid, Alkali, and Salt Tolerance in the Test Isolates
3.5. Inoculation Effects of Nitrogen-Fixing Bacterial Strains on Maize Growth
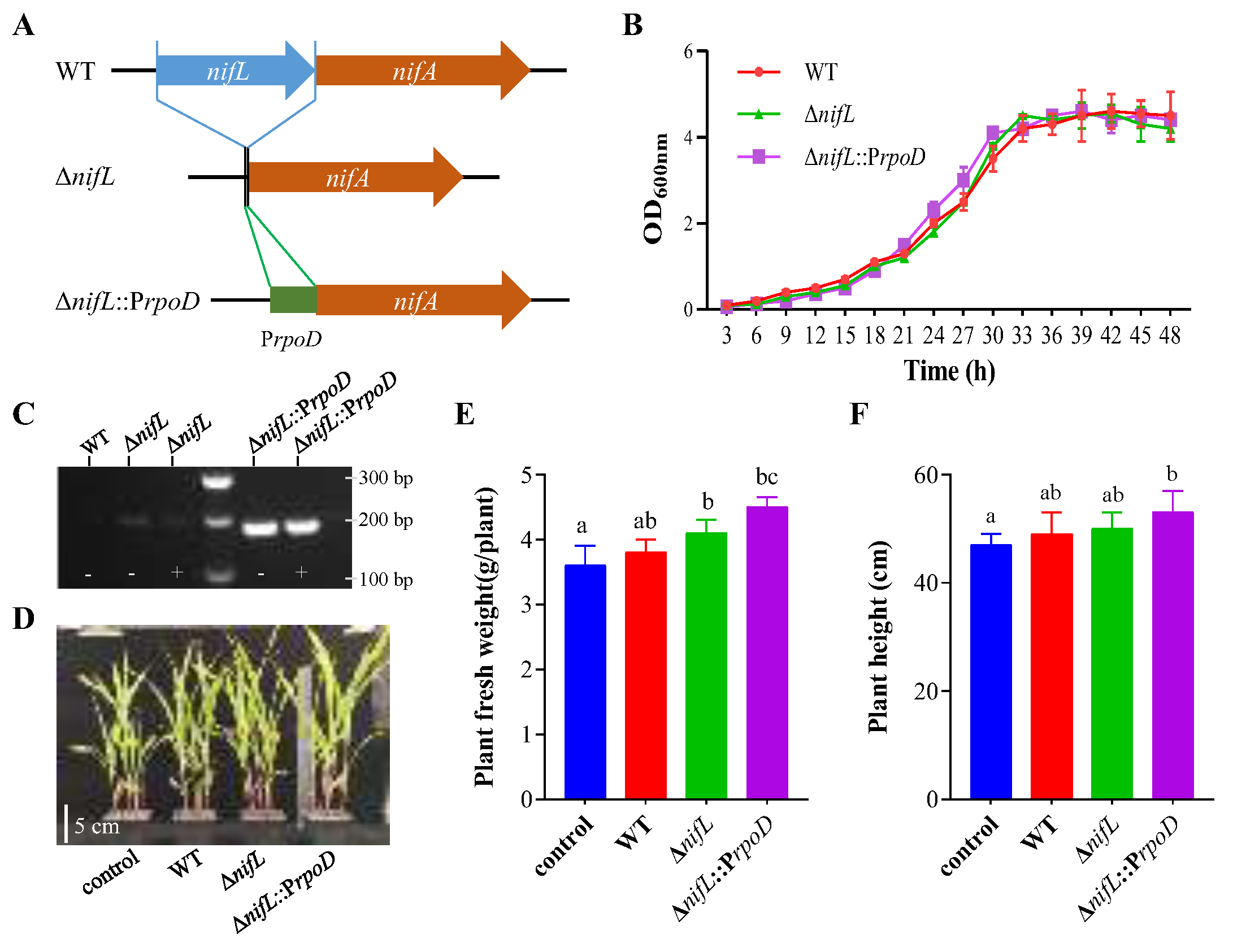
3.6. Genetic Modification of GN8811 Enhanced Plant-Growth Promotion
4. Discussion
5. Conclusion
Supplementary Materials
Acknowledgments
Conflicts of Interest
References
- Shahwar, D., et al., Role of microbial inoculants as bio fertilizers for improving crop productivity: A review. Heliyon, 2023. 9(6). [CrossRef]
- Dikir, W., Effect of long-term chemical fertilizer application on soil chemical properties: a review. J Biol Agric Healthc, 2023. 13: p. 11-18.
- Kim, H.-W., et al., Screening and isolated of plant growth promoting rhizobacteria Azotobacter sp. from maize rhizosphere. Probe - Plant & Animal Sciences, 2025. [CrossRef]
- Barron, S., F. Mus, and J.W. Peters, Nitrogen-Fixing Gamma Proteobacteria Azotobacter vinelandii—A Blueprint for Nitrogen-Fixing Plants? Microorganisms, 2024. 12(10): p. 2087. [CrossRef]
- Hossain, M.E., S. Shahrukh, and S.A. Hossain, Chemical fertilizers and pesticides: impacts on soil degradation, groundwater, and human health in Bangladesh, in Environmental degradation: challenges and strategies for mitigation. 2022, Springer. p. 63-92.
- Penuelas, J., F. Coello, and J. Sardans, A better use of fertilizers is needed for global food security and environmental sustainability. Agriculture & Food Security, 2023. 12(1): p. 1-9. [CrossRef]
- Sahoo, R.K., et al., Phenotypic and molecular characterisation of efficient nitrogen-fixing Azotobacter strains from rice fields for crop improvement. Protoplasma, 2013. 251(3): p. 511-523. [CrossRef]
- Macias-Bobadilla, I., et al., Hormetic and xenohormetic potential in the phytobiome of the center of origin. Genetic Resources and Crop Evolution, 2020. 67(5): p. 1331-1344. [CrossRef]
- Yang, J., et al., Mechanisms underlying legume–rhizobium symbioses. Journal of Integrative Plant Biology, 2022. 64(2): p. 244-267. [CrossRef]
- Pedraza, R.O., Recent advances in nitrogen-fixing acetic acid bacteria. International Journal of Food Microbiology, 2008. 125(1): p. 25-35. [CrossRef]
- Saranraj, P., et al. Isolation and nitrogen fixing efficiency of Gluconacetobacter diazotrophicus associated with sugarcane: A review. in IOP Conference Series: Earth and Environmental Science. 2021. IOP Publishing.
- Madhaiyan, M., et al., Improvement of plant growth and seed yield in Jatropha curcas by a novel nitrogen-fixing root associated Enterobacter species. Biotechnology for biofuels, 2013. 6(1): p. 140. [CrossRef]
- Li, H.-B., et al., Genetic diversity of nitrogen-fixing and plant growth promoting Pseudomonas species isolated from sugarcane rhizosphere. Frontiers in Microbiology, 2017. 8: p. 1268. [CrossRef]
- Jalal, A., et al., Plant growth-promoting bacteria and nitrogen fixing bacteria: sustainability of non-legume crops, in Nitrogen fixing bacteria: sustainable growth of non-legumes. 2022, Springer. p. 233-275.
- Bhattacharjee, R.B., A. Singh, and S. Mukhopadhyay, Use of nitrogen-fixing bacteria as biofertiliser for non-legumes: prospects and challenges. Applied microbiology and biotechnology, 2008. 80(2): p. 199-209. [CrossRef]
- Kuan, K.B., et al., Plant growth-promoting rhizobacteria inoculation to enhance vegetative growth, nitrogen fixation and nitrogen remobilisation of maize under greenhouse conditions. PloS one, 2016. 11(3): p. e0152478. [CrossRef]
- Abadi, V., et al., Diversity and abundance of culturable nitrogen-fixing bacteria in the phyllosphere of maize. Journal of Applied Microbiology, 2021. 131(2): p. 898-912. [CrossRef]
- Yuan, L., C. San-Feng, and L. Ji-Lun, Colonization pattern of Azospirillum brasilense Yu62 on maize roots. Journal of Integrative Plant Biology, 2003. 45(6): p. 748.
- Takahashi, W.Y., et al., Tracking maize colonization and growth promotion by Azospirillum reveals strain-specific behavior and the influence of inoculation method. Plant Physiology and Biochemistry, 2024. 215: p. 108979. [CrossRef]
- Monteiro, R.A., et al., Herbaspirillum-plant interactions: microscopical, histological and molecular aspects. Plant and Soil, 2012. 356(1): p. 175-196. [CrossRef]
- Di Salvo, L.P., et al., Plant growth-promoting rhizobacteria inoculation and nitrogen fertilization increase maize (Zea mays L.) grain yield and modified rhizosphere microbial communities. Applied Soil Ecology, 2018. 126: p. 113-120.
- Van Deynze, A., et al., Nitrogen fixation in a landrace of maize is supported by a mucilage-associated diazotrophic microbiota. PLoS biology, 2018. 16(8): p. e2006352.
- Sun, W., M.H. Shahrajabian, and Q. Cheng, Nitrogen fixation and diazotrophs–a review. Rom Biotechnol Lett, 2021. 26(4): p. 2834-2845. [CrossRef]
- Aira, M., et al., Plant genotype strongly modifies the structure and growth of maize rhizosphere microbial communities. Soil Biology and Biochemistry, 2010. 42(12): p. 2276-2281. [CrossRef]
- Zhao, M., et al., Zonal soil type determines soil microbial responses to maize cropping and fertilization. MSystems, 2016. 1(4): p. 10.1128/msystems. 00075-16. [CrossRef]
- Pittol, M., et al., Agronomic and environmental aspects of diazotrophic bacteria in rice fields. Annals of Microbiology, 2016. 66(2): p. 511-527. [CrossRef]
- Namsaraev, Z., et al., Effect of salinity on diazotrophic activity and microbial composition of phototrophic communities from Bitter-1 soda lake (Kulunda Steppe, Russia). Extremophiles, 2018. 22(4): p. 651-663. [CrossRef]
- Wang, Y., et al., Soil pH is a major driver of soil diazotrophic community assembly in Qinghai-Tibet alpine meadows. Soil Biology and Biochemistry, 2017. 115: p. 547-555. [CrossRef]
- Chukwudi, U.P., et al., Field application of beneficial microbes to ameliorate drought stress in maize. Plant and Soil, 2025: p. 1-20. [CrossRef]
- Wang, R., et al., Carbon allocation to the rhizosphere is affected by drought and nitrogen addition. Journal of Ecology, 2021. 109(10): p. 3699-3709. [CrossRef]
- Dixon, R. and D. Kahn, Genetic regulation of biological nitrogen fixation. Nature Reviews Microbiology, 2004. 2(8): p. 621-631. [CrossRef]
- Wu, S.C., et al., Effects of biofertilizer containing N-fixer, P and K solubilizers and AM fungi on maize growth: a greenhouse trial. Geoderma, 2005. 125(1-2): p. 155-166. [CrossRef]
- Wu, Z.H., et al., A synthetic bacterial consortium improved the phytoremediation efficiency of ryegrass on polymetallic contaminated soil. Ecotoxicol Environ Saf, 2024. 282: p. 116691. [CrossRef]
- Brtnicky, M., et al., Biodegradation of poly-3-hydroxybutyrate after soil inoculation with microbial consortium: Soil microbiome and plant responses to the changed environment. Sci Total Environ, 2024. 946: p. 174328. [CrossRef]
- Akashdeep, S. Kumari, and N. Rani, Novel cereal bran based low-cost liquid medium for enhanced growth, multifunctional traits and shelf life of consortium biofertilizer containing Azotobacter chroococcum, Bacillus subtilis and Pseudomonas sp. J Microbiol Methods, 2024. 222: p. 106952. [CrossRef]
- Rosenblueth, M., et al., Nitrogen Fixation in Cereals. Front Microbiol, 2018. 9: p. 1794.
- Zhao, Z., A.R. Fernie, and Y. Zhang, Engineering nitrogen and carbon fixation for next-generation plants. Curr Opin Plant Biol, 2025. 85: p. 102699. [CrossRef]
- Mus, F., et al., Genetic Determinants of Ammonium Excretion in nifL Mutants of Azotobacter vinelandii. Appl Environ Microbiol, 2022. 88(6): p. e0187621. [CrossRef]
- Lan, P., et al., Modification of nifL-nifA operon of Klebsiella oxytoca improves N(2)-fixation efficiency, ammonium-excretion and plant-growth promotion ability. Arch Microbiol, 2025. 207(10): p. 253. [CrossRef]
- Wu, W., et al., Harnessing biological nitrogen fixation: Multi-scale engineering for self-sustaining agroecosystems. Biotechnol Adv, 2025. 84: p. 108687. [CrossRef]
- Soumare, A., et al., Exploiting Biological Nitrogen Fixation: A Route Towards a Sustainable Agriculture. Plants (Basel), 2020. 9(8). [CrossRef]
- Kim, M., M. Morrison, and Z. Yu, Evaluation of different partial 16S rRNA gene sequence regions for phylogenetic analysis of microbiomes. Journal of microbiological methods, 2011. 84(1): p. 81-87. [CrossRef]
- Kumar, S., et al., MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol, 2018. 35(6): p. 1547-1549. [CrossRef]
- Deslippe, J.R., K.N. Egger, and G.H. Henry, Impacts of warming and fertilization on nitrogen-fixing microbial communities in the Canadian High Arctic. FEMS microbiology ecology, 2005. 53(1): p. 41-50. [CrossRef]
- Widmer, F., et al., Analysis of nifH gene pool complexity in soil and litter at a Douglas fir forest site in the Oregon Cascade Mountain Range. Applied and Environmental Microbiology, 1999. 65(2): p. 374-380. [CrossRef]
- Lv, F., et al., Regulation of hierarchical carbon substrate utilization, nitrogen fixation, and root colonization by the Hfq/Crc/CrcZY genes in Pseudomonas stutzeri. Iscience, 2022. 25(12). [CrossRef]
- Patel, D., et al., A resourceful methodology to profile indolic auxins produced by rhizo-fungi using spectrophotometry and HPTLC. 3 Biotech, 2018. 8(10): p. 413. [CrossRef]
- Muleta, A., et al., Phosphate solubilization and multiple plant growth promoting properties of Mesorhizobium species nodulating chickpea from acidic soils of Ethiopia. Arch Microbiol, 2021. 203(5): p. 2129-2137. [CrossRef]
- Himpsl, S.D. and H.L.T. Mobley, Siderophore Detection Using Chrome Azurol S and Cross-Feeding Assays. Methods Mol Biol, 2019. 2021: p. 97-108.
- Sen, A., et al., Assessing the effectiveness of indigenous phosphate-solubilizing bacteria in mitigating phosphorus fixation in acid soils. 3 Biotech, 2024. 14(9): p. 197. [CrossRef]
- Cui, W.J., et al., Lineage-Specific Rewiring of Core Pathways Predating Innovation of Legume Nodules Shapes Symbiotic Efficiency. mSystems, 2021. 6(2). [CrossRef]
- Sumbul, A., et al., Azotobacter: A potential bio-fertilizer for soil and plant health management. Saudi J Biol Sci, 2020. 27(12): p. 3634-3640. [CrossRef]
- Fukami, J., P. Cerezini, and M. Hungria, Azospirillum: benefits that go far beyond biological nitrogen fixation. AMB Express, 2018. 8(1): p. 73. [CrossRef]
- Bao, Y.Q., et al., Construction, Characterization, and Application of an Ammonium Transporter (AmtB) Deletion Mutant of the Nitrogen-Fixing Bacterium Kosakonia radicincitans GXGL-4A in Cucumis sativus L. Seedlings. Curr Microbiol, 2023. 80(2): p. 58. [CrossRef]
- Yoshidome, D., et al., Glutamate production from aerial nitrogen using the nitrogen-fixing bacterium Klebsiella oxytoca. Commun Biol, 2024. 7(1): p. 443. [CrossRef]
- Wang, J., et al., Beneficial bacteria activate nutrients and promote wheat growth under conditions of reduced fertilizer application. BMC Microbiol, 2020. 20(1): p. 38. [CrossRef]
- Ayyaz, K., et al., Isolation and identification by 16S rRNA sequence analysis of plant growth-promoting azospirilla from the rhizosphere of wheat. Braz J Microbiol, 2016. 47(3): p. 542-50. [CrossRef]
- Liang, P., et al., Klebsiella michiganensis: a nitrogen-fixing endohyphal bacterium from Ustilago maydis. AMB Express, 2023. 13(1): p. 146. [CrossRef]
- Bloch, S.E., et al., Biological nitrogen fixation in maize: optimizing nitrogenase expression in a root-associated diazotroph. J Exp Bot, 2020. 71(15): p. 4591-4603.
- Ma, H., et al., Synthetic Microbial Community Isolated from Intercropping System Enhances P Uptake in Rice. Int J Mol Sci, 2024. 25(23). [CrossRef]
- Wang, Z., et al., A Synthetic Microbial Community of Plant Core Microbiome Can Be a Potential Biocontrol Tool. J Agric Food Chem, 2023. 71(13): p. 5030-5041. [CrossRef]
- Sharma, S., J. Kulkarni, and B. Jha, Halotolerant Rhizobacteria Promote Growth and Enhance Salinity Tolerance in Peanut. Front Microbiol, 2016. 7: p. 1600. [CrossRef]
- Ishaq, S., et al., Plant growth-promoting bacteria from Uzungöl forest stimulate rice growth via seed biopriming and root inoculation: isolation and functional characterization of potent PGPR strains from rhizosphere soils of different trees. Front Plant Sci, 2025. 16: p. 1622951. [CrossRef]
- Zheng, Y., et al., Effect of planting salt-tolerant legumes on coastal saline soil nutrient availability and microbial communities. J Environ Manage, 2023. 345: p. 118574. [CrossRef]
- Huang, K., et al., Soil acidification and salinity: the importance of biochar application to agricultural soils. Front Plant Sci, 2023. 14: p. 1206820. [CrossRef]
- Hungria, M., et al., Improving maize sustainability with partial replacement of N fertilizers by inoculation with Azospirillum brasilense. Agronomy Journal, 2022. 114(5): p. 2969-2980. [CrossRef]
- dos Santos, C.L.R., et al., Contribution of a mixed inoculant containing strains of Burkholderia spp. and Herbaspirillum ssp. to the growth of three sorghum genotypes under increased nitrogen fertilization levels. Applied Soil Ecology, 2017. 113: p. 96-106.
- Schmidlin, K., et al., Environment-by-Environment Interactions Differ Across Genetic Backgrounds. Mol Biol Evol, 2025. 42(11). [CrossRef]
- Gimmi, E. and C. Vorburger, Strong genotype-by-genotype interactions between aphid-defensive symbionts and parasitoids persist across different biotic environments. J Evol Biol, 2021. 34(12): p. 1944-1953. [CrossRef]
- Gerhardt, E.C.M. and K.A. Selim, Unlocking nitrogen regulation: structural insights into the NifL-NifA complex and prospects for engineered diazotrophs. Febs j, 2025. [CrossRef]
- Poorter, H., et al., Pot size matters: a meta-analysis of the effects of rooting volume on plant growth. Funct Plant Biol, 2012. 39(11): p. 839-850. [CrossRef]
- Leontidou, K., et al., Plant growth promoting rhizobacteria isolated from halophytes and drought-tolerant plants: genomic characterisation and exploration of phyto-beneficial traits. Sci Rep, 2020. 10(1): p. 14857. [CrossRef]
- Smets, B.F. and T. Barkay, Horizontal gene transfer: perspectives at a crossroads of scientific disciplines. Nat Rev Microbiol, 2005. 3(9): p. 675-8. [CrossRef]
- Atieno, M., et al., Assessment of biofertilizer use for sustainable agriculture in the Great Mekong Region. J Environ Manage, 2020. 275: p. 111300. [CrossRef]
- Vejan, P., et al., Role of Plant Growth Promoting Rhizobacteria in Agricultural Sustainability-A Review. Molecules, 2016. 21(5). [CrossRef]
- Singh, R.P. and K. Kumari, Bacterial type VI secretion system (T6SS): an evolved molecular weapon with diverse functionality. Biotechnol Lett, 2023. 45(3): p. 309-331. [CrossRef]
- Bhagwat, A., et al., Bacterial metabolism in the host and its association with virulence. Virulence, 2025. 16(1): p. 2459336. [CrossRef]
- do Amaral, F.P., et al., Diverse Bacterial Genes Modulate Plant Root Association by Beneficial Bacteria. mBio, 2020. 11(6). [CrossRef]
- Das, H.K., Azotobacters as biofertilizer. Adv Appl Microbiol, 2019. 108: p. 1-43.
- Gómez-Godínez, L.J., et al., A Look at Plant-Growth-Promoting Bacteria. Plants (Basel), 2023. 12(8).
- Ercole, T.G., et al., The role of endophytic bacteria in enhancing plant growth and health for sustainable agriculture. Antonie Van Leeuwenhoek, 2025. 118(7): p. 88. [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



