Submitted:
24 September 2025
Posted:
25 September 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Transcriptome-Wide Association Study (TWAS)
2.2. Linkage Disequilibrium Analysis
2.3. Postmortem Expression Analysis (BrainSeq)
2.4. Developmental Expression Trajectories (BrainSpan)
2.5. UCSC Genome Browser Validation
2.6. Functional Predictions (lncHUB)
3. Results
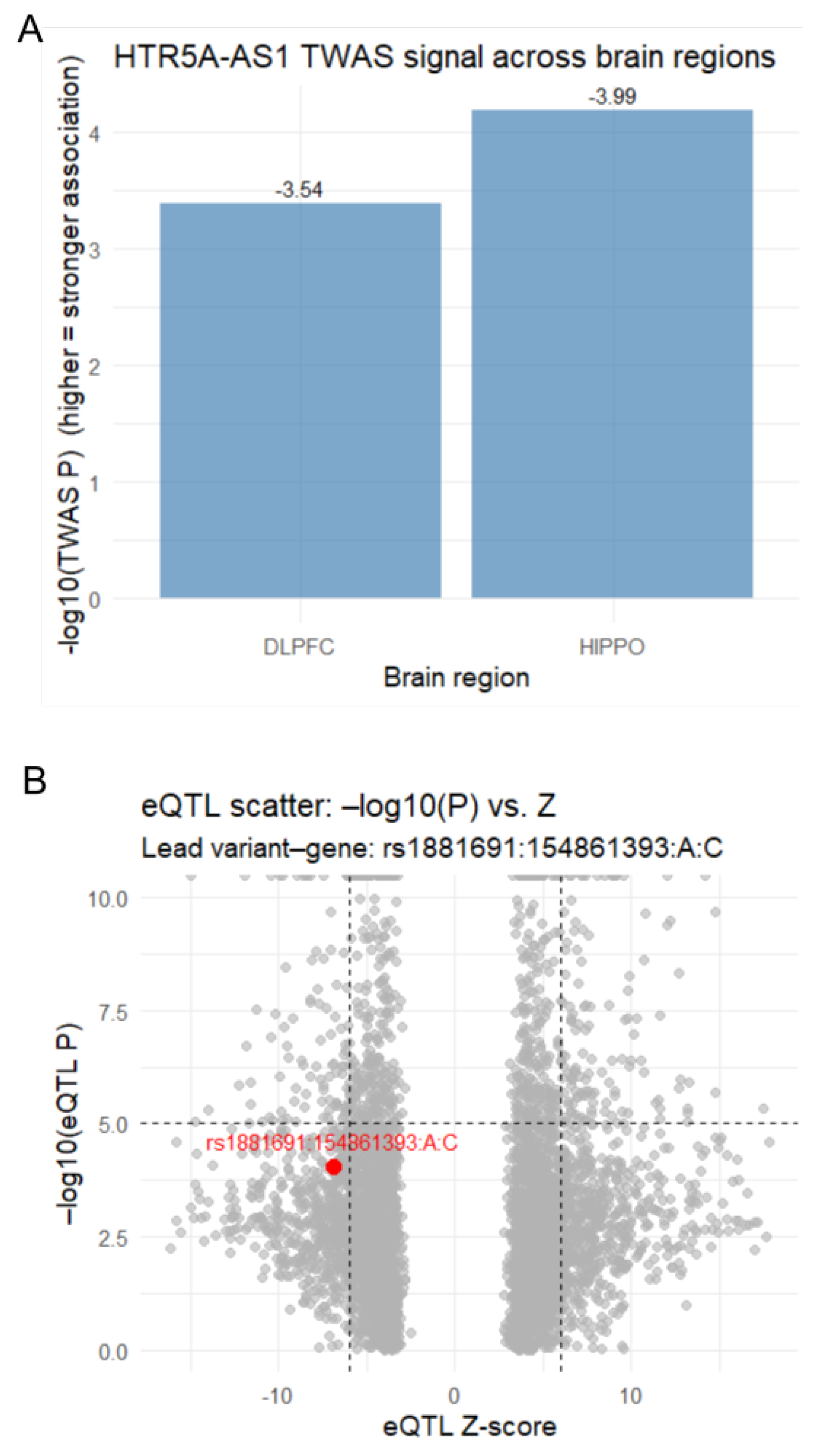
3.1. TWAS Identifies HTR5A-AS1 Association with Schizophrenia
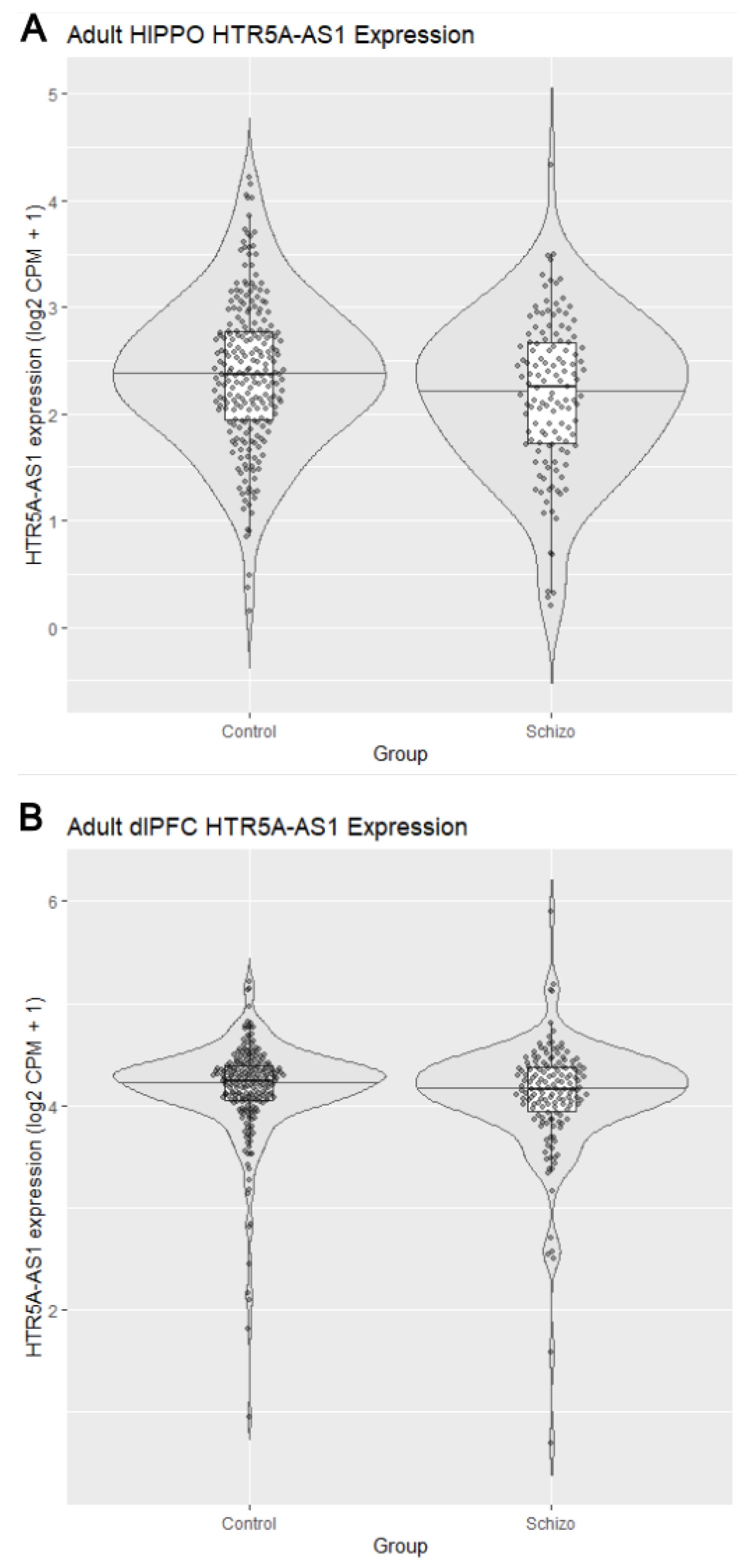
3.2. Reduced HTR5A-AS1 Expression in the Hippocampus
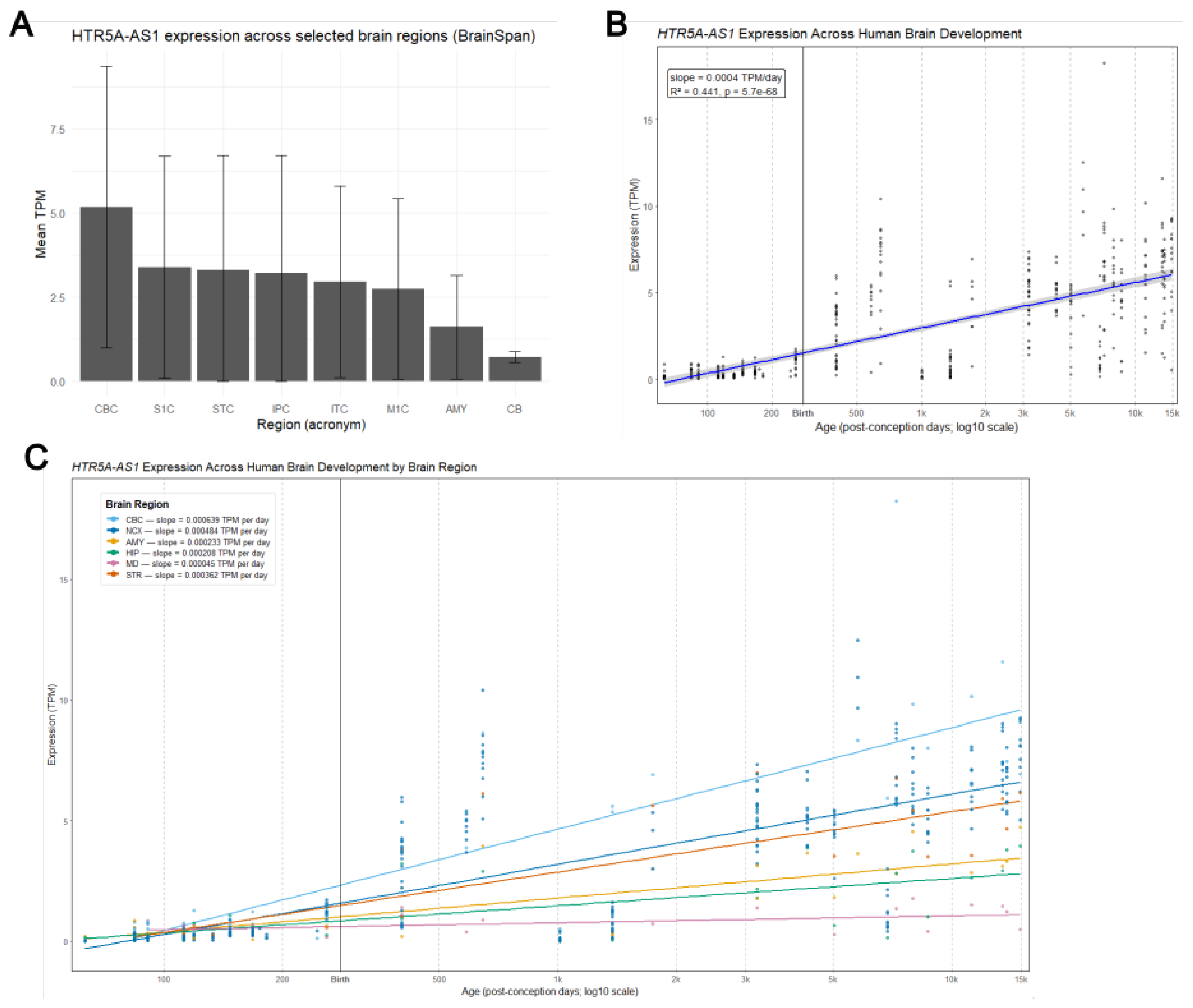
3.3. Region-Specific Expression in the Human Brain
3.4. Developmental Trajectory of HTR5A-AS1
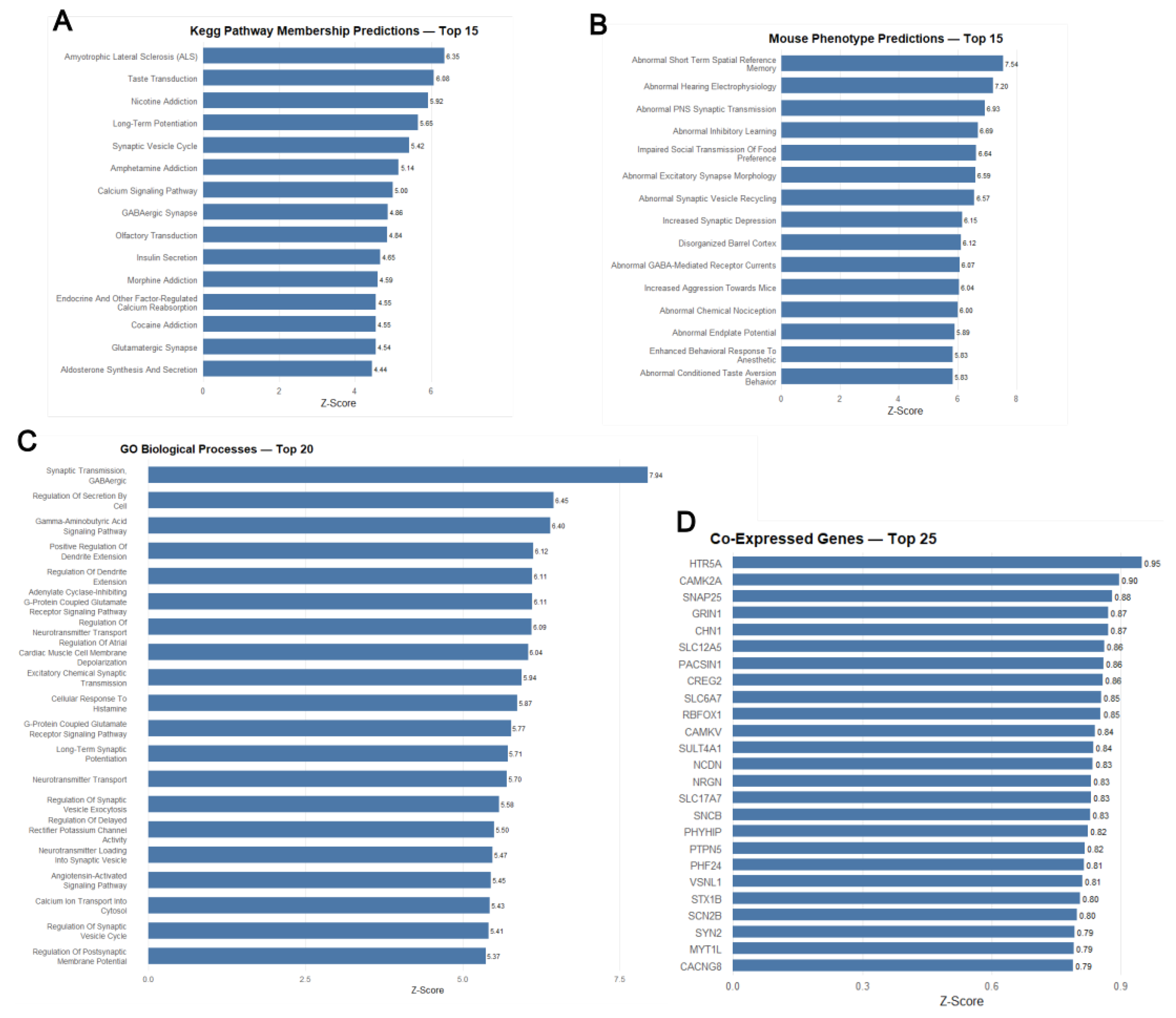
3.5. Transcript Validation via UCSC Genome Browser and Co-Expression Analysis
3.6. Predicted Functional Associations of HTR5A-AS1
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Schizophrenia. Available online: https://www.who.int/news-room/fact-sheets/detail/schizophrenia (accessed on 8 September 2025).
- Solmi, M.; Seitidis, G.; Mavridis, D.; Correll, C.U.; Dragioti, E.; Guimond, S.; Tuominen, L.; Dargél, A.; Carvalho, A.F.; Fornaro, M.; et al. Incidence, Prevalence, and Global Burden of Schizophrenia—Data, with Critical Appraisal, from the Global Burden of Disease (GBD) 2019. Mol. Psychiatry 2023, 28, 5319–5327. [Google Scholar] [CrossRef] [PubMed]
- Hany, M.; Rizvi, A. Schizophrenia. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2025. [Google Scholar]
- Marder, S.R.; Cannon, T.D. Schizophrenia. N. Engl. J. Med. 2019. [Google Scholar] [CrossRef] [PubMed]
- Chick, S.L.; Holmans, P.; Cameron, D.; Grozeva, D.; Sims, R.; Williams, J.; Bray, N.J.; Owen, M.J.; O’Donovan, M.C.; Walters, J.T.R.; Rees, E. Whole-Exome Sequencing Analysis Identifies Risk Genes for Schizophrenia. Nat. Commun. 2025, 16, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Functional Genomics Reveal Gene Regulatory Mechanisms Underlying Schizophrenia Risk. Nat. Commun. 2019. Available online: https://www.nature.com/articles/s41467-019-08666-4 (accessed on 16 August 2025).
- Birnbaum, R.; Weinberger, D.R. Genetic Insights into the Neurodevelopmental Origins of Schizophrenia. Nat. Rev. Neurosci. 2017, 18, 727–740. [Google Scholar] [CrossRef] [PubMed]
- Karlsgodt, K.H.; Sun, D.; Cannon, T.D. Structural and Functional Brain Abnormalities in Schizophrenia. Curr. Dir. Psychol. Sci. 2010, 19, 226–231. [Google Scholar] [CrossRef] [PubMed]
- Quednow, B.B.; Geyer, M.A.; Halberstadt, A.L. Serotonin and Schizophrenia. In Handbook of Behavioral Neuroscience; Müller, C.P., Cunningham, K.A., Eds.; Elsevier: Amsterdam, The Netherlands, 2020; Volume 31, pp. 711–743. [Google Scholar] [CrossRef]
- Li, P.; Snyder, G.L.; Vanover, K.E. Dopamine Targeting Drugs for the Treatment of Schizophrenia: Past, Present and Future. Curr. Top. Med. Chem. 2016, 16, 3385–3403. [Google Scholar] [CrossRef] [PubMed]
- Meltzer, H.Y.; Li, Z.; Kaneda, Y.; Ichikawa, J. Serotonin Receptors: Their Key Role in Drugs to Treat Schizophrenia. Prog. Neuropsychopharmacol. Biol. Psychiatry 2003, 27, 1159–1172. [Google Scholar] [CrossRef] [PubMed]
- Grubor, M.; Zivkovic, M.; Sagud, M.; Nikolac Perkovic, M.; Mihaljevic-Peles, A.; Pivac, N.; Muck-Seler, D.; Svob Strac, D. HTR1A, HTR1B, HTR2A, HTR2C and HTR6 Gene Polymorphisms and Extrapyramidal Side Effects in Haloperidol-Treated Patients with Schizophrenia. Int. J. Mol. Sci. 2020, 21, 2345. [Google Scholar] [CrossRef] [PubMed]
- Thomas, D.R. 5-ht5A Receptors as a Therapeutic Target. Pharmacol. Ther. 2006, 111, 707–714. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Smith, E.M.; Baye, T.M.; Eckert, J.V.; Abraham, L.J.; Moses, E.K.; Kissebah, A.H.; Martin, L.J.; Olivier, M. Serotonin (5-HT) Receptor 5A Sequence Variants Affect Human Plasma Triglyceride Levels. Physiol. Genomics 2010, 42, 168–176. [Google Scholar] [CrossRef] [PubMed]
- Gusev, A.; Ko, A.; Shi, H.; Bhatia, G.; Chung, W.; Penninx, B.W.J.H.; Jansen, R.; de Geus, E.J.; Boomsma, D.I.; Wright, F.A.; et al. Integrative Approaches for Large-Scale Transcriptome-Wide Association Studies. Nat. Genet. 2016, 48, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Collado-Torres, L.; Burke, E.E.; Peterson, A.; Shin, J.; Straub, R.E.; Rajpurohit, A.; Semick, S.A.; Ulrich, W.S.; Price, A.J.; Valencia, C.; et al. Regional Heterogeneity in Gene Expression, Regulation, and Coherence in the Frontal Cortex and Hippocampus across Development and Schizophrenia. Neuron 2019, 103, 203–216.e8. [Google Scholar] [CrossRef] [PubMed]
- National Center for Biotechnology Information. Available online: https://www.ncbi.nlm.nih.gov/ (accessed on 17 September 2025).
- GeneCards. Available online: https://www.genecards.org/ (accessed on 17 September 2025).
- BrainSpan Atlas of the Developing Human Brain. Available online: https://www.brainspan.org/ (accessed on 8 September 2025).
- UCSC Genome Browser. Available online: https://genome.ucsc.edu/ (accessed on 17 September 2025).
- lncHUB. Available online: https://maayanlab.cloud/lnchub/ (accessed on 17 September 2025).
- R: The R Project for Statistical Computing. Available online: https://www.r-project.org/ (accessed on 17 September 2025).
- readr: Read Rectangular Text Data. Available online: https://CRAN.R-project.org/package=readr (accessed on 17 September 2025).
- dplyr: A Grammar of Data Manipulation. Available online: https://CRAN.R-project.org/package=dplyr (accessed on 17 September 2025).
- Wickham, H. ggplot2; Springer International Publishing: Cham, Switzerland, 2016. [Google Scholar] [CrossRef]
- stringr: Simple, Consistent Wrappers for Common String Operations. Available online: https://CRAN.R-project.org/package=stringr (accessed on 17 September 2025).
- Myers, T.A.; Chanock, S.J.; Machiela, M.J. LDlinkR: An R Package for Rapidly Calculating Linkage Disequilibrium Statistics in Diverse Populations. Front. Genet. 2020, 11, 157. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.-H.; Thakur, R.; Machiela, M.J. LDexpress: An Online Tool for Integrating Population-Specific Linkage Disequilibrium Patterns with Tissue-Specific Expression Data. BMC Bioinformatics 2021, 22, 608. [Google Scholar] [CrossRef] [PubMed]
- SummarizedExperiment: Summarized Experiment Container. Available online: https://bioconductor.org/packages/SummarizedExperiment (accessed on 17 September 2025).
- forcats: Tools for Working with Categorical Variables (Factors). Available online: https://CRAN.R-project.org/package=forcats (accessed on 17 September 2025).
- ggbeeswarm: Categorical Scatter (Violin Point) Plots. Available online: https://CRAN.R-project.org/package=ggbeeswarm (accessed on 17 September 2025).
- broom: Convert Statistical Objects into Tidy Tibbles. Available online: https://CRAN.R-project.org/package=broom (accessed on 17 September 2025).
- tibble: Simple Data Frames. Available online: https://CRAN.R-project.org/package=tibble (accessed on 17 September 2025).
- rlang: Functions for Base Types and Core R and Tidyverse Features. Available online: https://CRAN.R-project.org/package=rlang (accessed on 17 September 2025).
- R Core Team. grid: The Grid Graphics Package. R package version bundled with R. Available online: https://stat.ethz.ch/R-manual/R-devel/library/grid/doc/grid.pdf (accessed on 17 September 2025).

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



