Submitted:
15 December 2024
Posted:
17 December 2024
Read the latest preprint version here
Abstract
Keywords:
Introduction
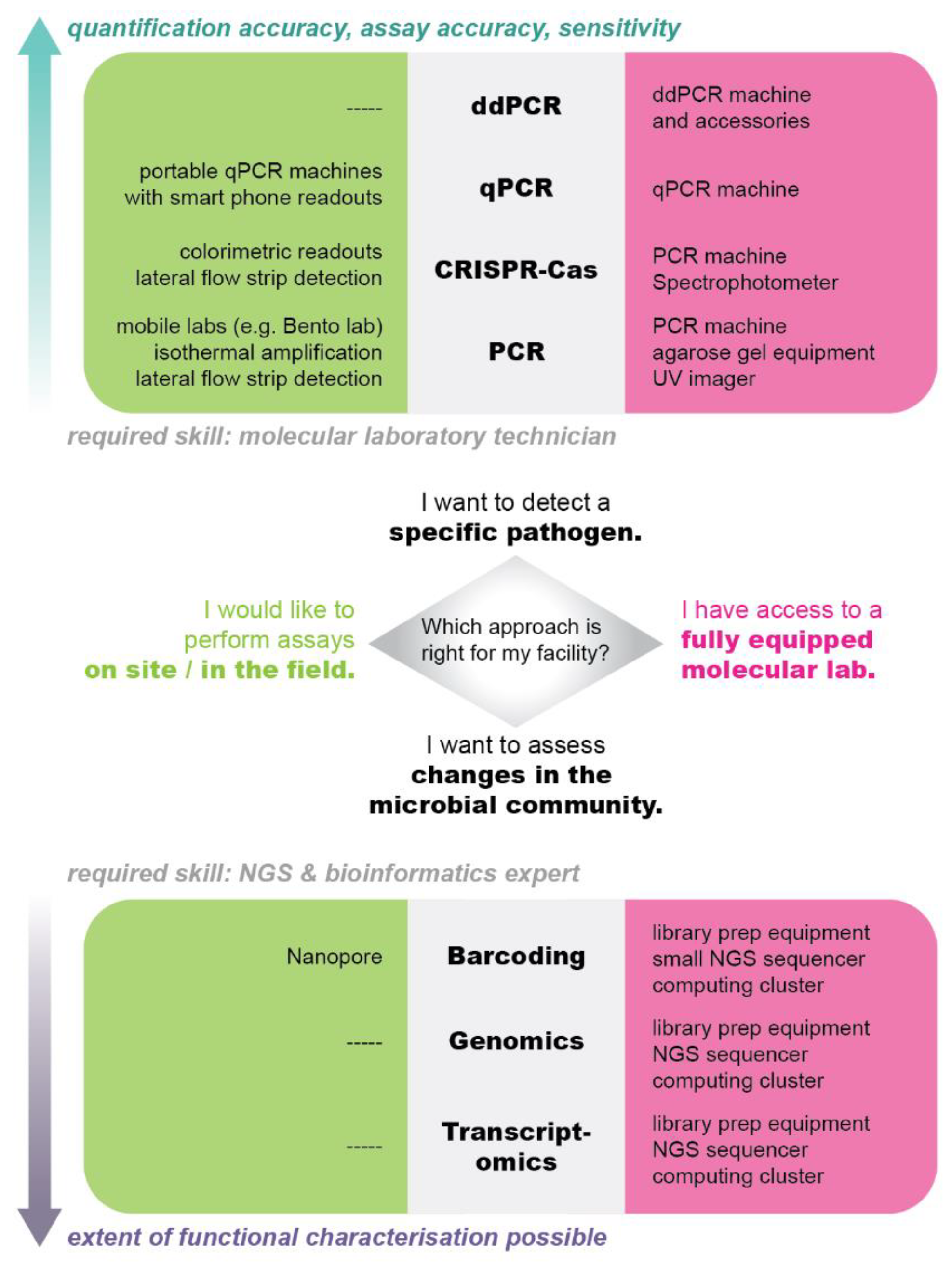
Bringing the Lab to the Farm
Independence from Controlled Environments
Moving Away from Fish Handling
Transfer of CRISPR Technology to Aquaculture
| METHOD | PROTEINS | AMPLIFICATION | DETECTION | TARGET | SENSITIVITY | TIME | REF |
| SHERLOCKV1 | Cas13a | RPA | Fluorescence, Colorimetry |
DNA/ RNA |
1.06 copies (10 copies /colorimetry) |
60 mins | Sullivan, 2019 |
| SHERLOCKV2 | Cas12b | LAMP | Fluorescence | DNA/ RNA |
100 copies | 30- 60 mins | Major, 2023 |
| DETECTR | Cas12a | RPA | Fluorescence, Colorimetry | DNA | 40 copies (200 copies/ colorimetry) |
Li, 2022 | |
| RAA-CRISPR/ CAS12A |
Cas12a | RAA | Fluorescence | DNA | 2 copies | 40 mins | Xiao, 2021 |
Transfer of Microbial Community Ecology to Aquaculture
Conclusion
Ethics and integrity statement
Author contributions
Acknowledgements
References
- Almeida, D. B., Magalhães, C., Sousa, Z., Borges, M. T., Silva, E., Blanquet, I., & Mucha, A. P. (2021). Microbial community dynamics in a hatchery recirculating aquaculture system (RAS) of sole (Solea senegalensis). Aquaculture, 539. [CrossRef]
- Anupama, K. P. , Nayak, A., Karunasagar, I., & Maiti, B. (2020). Rapid visual detection of Vibrio parahaemolyticus in seafood samples by loop-mediated isothermal amplification with hydroxynaphthol blue dye. World Journal of Microbiology and Biotechnology, 36(5), Article 5. [CrossRef]
- Bakke, I. , Åm, A. L., Kolarevic, J., Ytrestøyl, T., Vadstein, O., Attramadal, K. J. K., & Terjesen, B. F. (2017). Microbial community dynamics in semi-commercial RAS for production of Atlantic salmon post-smolts at different salinities. Aquacultural Engineering, 78, 42–49. [CrossRef]
- Bass, D. , Christison, K. W., Stentiford, G. D., Cook, L. S. J., & Hartikainen, H. (2023). Environmental DNA/RNA for pathogen and parasite detection, surveillance, and ecology. Trends in Parasitology, 39(4), Article 4. [CrossRef]
- Bastos Gomes, G., Hutson, K. S., Domingos, J. A., Chung, C., Hayward, S., Miller, T. L., & Jerry, D. R. (2017). Use of environmental DNA (eDNA) and water quality data to predict protozoan parasites outbreaks in fish farms. Aquaculture, 479, 467–473. [CrossRef]
- Bentzon-Tilia, M. , Sonnenschein, E. C., & Gram, L. (2016). Monitoring and managing microbes in aquaculture – Towards a sustainable industry. Microbial Biotechnology, 9(5), Article 5. [CrossRef]
- Bloemen, B. , Gand, M., Vanneste, K., Marchal, K., Roosens, N. H. C., & De Keersmaecker, S. C. J. (2023). Development of a portable on-site applicable metagenomic data generation workflow for enhanced pathogen and antimicrobial resistance surveillance. Scientific Reports, 13(1), Article 1. [CrossRef]
- Bohara, K. , Yadav, A. K., & Joshi, P. (2022). Detection of Fish Pathogens in Freshwater Aquaculture Using eDNA Methods. Diversity, 14(12), Article 12. [CrossRef]
- Cain, K. (2022). The many challenges of disease management in aquaculture. Journal of the World Aquaculture Society, 53(6), 1080–1083. [CrossRef]
- Chang, W.-H. , Yang, S.-Y., Wang, C.-H., Tsai, M.-A., Wang, P.-C., Chen, T.-Y., Chen, S.-C., & Lee, G.-B. (2013). Rapid isolation and detection of aquaculture pathogens in an integrated microfluidic system using loop-mediated isothermal amplification. Sensors and Actuators B: Chemical, 180, 96–106. [CrossRef]
- De Schryver, P. , & Vadstein, O. (2014). Ecological theory as a foundation to control pathogenic invasion in aquaculture. The ISME Journal, 8(12), Article 12. [CrossRef]
- Delamare-Deboutteville, J. (2021). Uncover aquaculture pathogens identity using Nanopore MinION.pdf. https://digitalarchive.worldfishcenter.org/bitstream/handle/20.500.12348/4931/eb0411d2c9e634acfae22ee2db9e7391.pdf?sequence2=.
- Ellison, A. R. , Uren Webster, T. M., Rey, O., Garcia de Leaniz, C., Consuegra, S., Orozco-terWengel, P., & Cable, J. (2018). Transcriptomic response to parasite infection in Nile tilapia (Oreochromis niloticus) depends on rearing density. BMC Genomics, 19(1), Article 1. [CrossRef]
- Ellison, A. R. , Uren Webster, T. M., Rodriguez-Barreto, D., de Leaniz, C. G., Consuegra, S., Orozco-terWengel, P., & Cable, J. (2020). Comparative transcriptomics reveal conserved impacts of rearing density on immune response of two important aquaculture species. Fish & Shellfish Immunology, 104, 192–201. [CrossRef]
- El-Saadony, M. T. , Alagawany, M., Patra, A. K., Kar, I., Tiwari, R., Dawood, M. A. O., Dhama, K., & Abdel-Latif, H. M. R. (2021). The functionality of probiotics in aquaculture: An overview. Fish & Shellfish Immunology, 117, 36–52. [CrossRef]
- FAO. (2020). The State of World Fisheries and Aquaculture 2020. In brief. FAO. [CrossRef]
- FAO. (2022). The State of World Fisheries and Aquaculture 2022. FAO. [CrossRef]
- Fossøy, F. , Brandsegg, H., Sivertsgård, R., Pettersen, O., Sandercock, B. K., Solem, Ø., Hindar, K., & Mo, T. A. (2020). Monitoring presence and abundance of two gyrodactylid ectoparasites and their salmonid hosts using environmental DNA. Environmental DNA, 2(1), 53–62. [CrossRef]
- Gallagher, M. D. , Matejusova, I., Nguyen, L., Ruane, N. M., Falk, K., & Macqueen, D. J. (2018). Nanopore sequencing for rapid diagnostics of salmonid RNA viruses. Scientific Reports, 8(1), Article 1. [CrossRef]
- Gao, F. , Jiang, J.-Z., Wang, J.-Y., & Wei, H.-Y. (2018). Real-time isothermal detection of Abalone herpes-like virus and red-spotted grouper nervous necrosis virus using recombinase polymerase amplification. Journal of Virological Methods, 251, 92–98. [CrossRef]
- Geng, Y. , Tan, K., Liu, L., Sun, X. X., Zhao, B., & Wang, J. (2019). Development and evaluation of a rapid and sensitive RPA assay for specific detection of Vibrio parahaemolyticus in seafood. BMC Microbiology, 19(1), Article 1. [CrossRef]
- Giuffrida, M. C. , & Spoto, G. (2017). Integration of isothermal amplification methods in microfluidic devices: Recent advances. Biosensors and Bioelectronics, 90, 174–186. [CrossRef]
- Gootenberg, J. S. , Abudayyeh, O. O., Lee, J. W., Essletzbichler, P., Dy, A. J., Joung, J., Verdine, V., Donghia, N., Daringer, N. M., Freije, C. A., Myhrvold, C., Bhattacharyya, R. P., Livny, J., Regev, A., Koonin, E. V., Hung, D. T., Sabeti, P. C., Collins, J. J., & Zhang, F. (2017). Nucleic acid detection with CRISPR-Cas13a/C2c2. Science (New York, N.Y.), 356(6336), 438–442. [CrossRef]
- Gorgannezhad, L. , Stratton, H., & Nguyen, N.-T. (2019). Microfluidic-Based Nucleic Acid Amplification Systems in Microbiology. Micromachines, 10(6), Article 6. [CrossRef]
- Guptha Yedire, S. , Khan, H., AbdelFatah, T., Moakhar, R. S., & Mahshid, S. (2023). Microfluidic-based colorimetric nucleic acid detection of pathogens. Sensors & Diagnostics. [CrossRef]
- Hook, S. E., White, C., & Ross, D. J. (2021). A metatranscriptomic analysis of changing dynamics in the plankton communities adjacent to aquaculture leases in southern Tasmania, Australia. Marine Genomics, 59, 100858. [CrossRef]
- Jaroenram, W. , & Owens, L. (2014). Recombinase polymerase amplification combined with a lateral flow dipstick for discriminating between infectious Penaeus stylirostris densovirus and virus-related sequences in shrimp genome. Journal of Virological Methods, 208, 144–151. [CrossRef]
- Jin, R. , Zhai, L., Zhu, Q., Feng, J., & Pan, X. (2020). Naked-eyes detection of Largemouth bass ranavirus in clinical fish samples using gold nanoparticles as colorimetric sensor. Aquaculture, 528, 735554. [CrossRef]
- Kanitchinda, S. , Srisala, J., Suebsing, R., Prachumwat, A., & Chaijarasphong, T. (2020). CRISPR-Cas fluorescent cleavage assay coupled with recombinase polymerase amplification for sensitive and specific detection of Enterocytozoon hepatopenaei. Biotechnology Reports, 27, e00485. [CrossRef]
- Kant, K. , Shahbazi, M.-A., Dave, V. P., Ngo, T. A., Chidambara, V. A., Than, L. Q., Bang, D. D., & Wolff, A. (2018). Microfluidic devices for sample preparation and rapid detection of foodborne pathogens. Biotechnology Advances, 36(4), Article 4. [CrossRef]
- Kellner, M. J. , Koob, J. G., Gootenberg, J. S., Abudayyeh, O. O., & Zhang, F. (2019). SHERLOCK: Nucleic acid detection with CRISPR nucleases. Nature Protocols, 14(10), Article 10. [CrossRef]
- Khan, R., Petersen, F. C., & Shekhar, S. (2019). Commensal Bacteria: An Emerging Player in Defense Against Respiratory Pathogens. Frontiers in Immunology, 10. https://www.frontiersin.org/articles/10.3389/fimmu.2019.01203.
- Kulkarni, M. B. , & Goel, S. (2020). Advances in continuous-flow based microfluidic PCR devices—A review. Engineering Research Express, 2(4), Article 4. [CrossRef]
- Lee, M. D., Pedroso, A. A., & Maurer, J. J. (2023). Bacterial composition of a competitive exclusion product and its correlation with product efficacy at reducing Salmonella in poultry. Frontiers in Physiology, 13. https://www.frontiersin.org/articles/10.3389/fphys.2022.1043383.
- Li, C., Lin, N., Feng, Z., Lin, M., Guan, B., Chen, K., Liang, W., Wang, Q., Li, M., You, Y., & Chen, Q. (2022). CRISPR/Cas12a Based Rapid Molecular Detection of Acute Hepatopancreatic Necrosis Disease in Shrimp. Frontiers in Veterinary Science, 8. https://www.frontiersin.org/articles/10.3389/fvets.2021.819681.
- Li, H. , Zhang, L., Yu, Y., Ai, T., Zhang, Y., & Su, J. (2022). Rapid detection of Edwardsiella ictaluri in yellow catfish (Pelteobagrus fulvidraco) by real-time RPA and RPA-LFD. Aquaculture, 552, 737976. [CrossRef]
- Liu, R. , Han, G., Li, Z., Cun, S., Hao, B., Zhang, J., & Liu, X. (2022). Bacteriophage therapy in aquaculture: Current status and future challenges. Folia Microbiologica, 67(4), 573–590. [CrossRef]
- Mabrok, M. , Elayaraja, S., Chokmangmeepisarn, P., Jaroenram, W., Arunrut, N., Kiatpathomchai, W., Debnath, P. P., Delamare-Deboutteville, J., Mohan, C. V., Fawzy, A., & Rodkhum, C. (2021). Rapid visualization in the specific detection of Flavobacterium columnare, a causative agent of freshwater columnaris using a novel recombinase polymerase amplification (RPA) combined with lateral flow dipstick (LFD) assay. Aquaculture, 531, 735780. [CrossRef]
- Major, S. R. , Harke, M. J., Cruz-Flores, R., Dhar, A. K., Bodnar, A. G., & Wanamaker, S. A. (2023). Rapid Detection of DNA and RNA Shrimp Viruses Using CRISPR-Based Diagnostics. Applied and Environmental Microbiology, 89(6), e02151-22. [CrossRef]
- Martínez-Porchas, M. , & Vargas-Albores, F. (2017). Microbial metagenomics in aquaculture: A potential tool for a deeper insight into the activity. Reviews in Aquaculture, 9(1), 42–56. [CrossRef]
- Mérou, N. , Lecadet, C., Pouvreau, S., & Arzul, I. (2020). An eDNA/eRNA-based approach to investigate the life cycle of non-cultivable shellfish micro-parasites: The case of Bonamia ostreae, a parasite of the European flat oyster Ostrea edulis. Microbial Biotechnology, 13(6), 1807–1818. [CrossRef]
- Morsy, M. K., Zór, K., Kostesha, N., Alstrøm, T. S., Heiskanen, A., El-Tanahi, H., Sharoba, A., Papkovsky, D., Larsen, J., Khalaf, H., Jakobsen, M. H., & Emnéus, J. (2016). Development and validation of a colorimetric sensor array for fish spoilage monitoring. Food Control, 60, 346–352. [CrossRef]
- Naylor, R. L. , Hardy, R. W., Buschmann, A. H., Bush, S. R., Cao, L., Klinger, D. H., Little, D. C., Lubchenco, J., Shumway, S. E., & Troell, M. (2021). A 20-year retrospective review of global aquaculture. Nature, 591(7851), Article 7851. [CrossRef]
- Nguyen, P. L. , Sudheesh, P. S., Thomas, A. C., Sinnesael, M., Haman, K., & Cain, K. D. (2018). Rapid Detection and Monitoring of Flavobacterium psychrophilum in Water by Using a Handheld, Field-Portable Quantitative PCR System. Journal of Aquatic Animal Health, 30(4), 302–311. [CrossRef]
- Nogueira, T. , & Botelho, A. (2021). Metagenomics and other omics approaches to bacterial communities and antimicrobial resistance assessment in aquacultures. Antibiotics, 10(7). [CrossRef]
- Notomi, T. , Okayama, H., Masubuchi, H., Yonekawa, T., Watanabe, K., Amino, N., & Hase, T. (2000). Loop-mediated isothermal amplification of DNA. Nucleic Acids Research, 28(12), Article 12. [CrossRef]
- Novoslavskij, A. , Terentjeva, M., Eizenberga, I., Valciņa, O., Bartkevičs, V., & Bērziņš, A. (2016). Major foodborne pathogens in fish and fish products: A review. Annals of Microbiology, 66(1), Article 1. [CrossRef]
- Phelps, M. (2019). Increasing eDNA capabilities with CRISPR technology for real-time monitoring of ecosystem biodiversity. Molecular Ecology Resources, 19(5), 1103–1105. [CrossRef]
- Prescott, M. A. , Reed, A. N., Jin, L., & Pastey, M. K. (2016). Rapid Detection of Cyprinid Herpesvirus 3 in Latently Infected Koi by Recombinase Polymerase Amplification. Journal of Aquatic Animal Health, 28(3), Article 3. [CrossRef]
- Rey-Campos, M. , Ríos-Castro, R., Gallardo-Escárate, C., Novoa, B., & Figueras, A. (2022). Exploring the Potential of Metatranscriptomics to Describe Microbial Communities and Their Effects in Molluscs. International Journal of Molecular Sciences, 23(24), 16029. [CrossRef]
- Rieder, J., Kapopoulou, A., Bank, C., & Adrian-Kalchhauser, I. (2023). Metagenomics and metabarcoding experimental choices and their impact on microbial community characterization in freshwater recirculating aquaculture systems. Environmental Microbiome, 18(1), 8. [CrossRef]
- Rieder, J. , Martin-Sanchez, P. M., Osman, O. A., Adrian-Kalchhauser, I., & Eiler, A. (2022). Detecting aquatic pathogens with field-compatible dried qPCR assays. Journal of Microbiological Methods, 202, 106594. [CrossRef]
- Rupp, M. , Pilo, P., Müller, B., Knüsel, R., von Siebenthal, B., Frey, J., Sindilariu, P.-D., & Schmidt-Posthaus, H. (2019). Systemic infection in European perch with thermoadapted virulent Aeromonas salmonicida (Perca fluviatilis). Journal of Fish Diseases, 42(5), Article 5. [CrossRef]
- Samacoits, A. , Nimsamer, P., Mayuramart, O., Chantaravisoot, N., Sitthi-amorn, P., Nakhakes, C., Luangkamchorn, L., Tongcham, P., Zahm, U., Suphanpayak, S., Padungwattanachoke, N., Leelarthaphin, N., Huayhongthong, H., Pisitkun, T., Payungporn, S., & Hannanta-anan, P. (2021). Machine Learning-Driven and Smartphone-Based Fluorescence Detection for CRISPR Diagnostic of SARS-CoV-2. ACS Omega, 6(4), 2727–2733. [CrossRef]
- Shahin, K. , Gustavo Ramirez-Paredes, J., Harold, G., Lopez-Jimena, B., Adams, A., & Weidmann, M. (2018). Development of a recombinase polymerase amplification assay for rapid detection of Francisella noatunensis subsp. Orientalis. PloS One, 13(2), Article 2. [CrossRef]
- Sieber, N. , Hartikainen, H., & Vorburger, C. (2020). Validation of an eDNA-based method for the detection of wildlife pathogens in water. Diseases of Aquatic Organisms, 141, 171–184. [CrossRef]
- Soliman, H. , Kumar, G., & El-Matbouli, M. (2018). Recombinase polymerase amplification assay combined with a lateral flow dipstick for rapid detection of Tetracapsuloides bryosalmonae, the causative agent of proliferative kidney disease in salmonids. Parasites & Vectors, 11(1), Article 1. [CrossRef]
- Song, X. , Coulter, F. J., Yang, M., Smith, J. L., Tafesse, F. G., Messer, W. B., & Reif, J. H. (2022). A lyophilized colorimetric RT-LAMP test kit for rapid, low-cost, at-home molecular testing of SARS-CoV-2 and other pathogens. Scientific Reports, 12(1), Article 1. [CrossRef]
- Stentiford, G. D. , Peeler, E. J., Tyler, C. R., Bickley, L. K., Holt, C. C., Bass, D., Turner, A. D., Baker-Austin, C., Ellis, T., Lowther, J. A., Posen, P. E., Bateman, K. S., Verner-Jeffreys, D. W., van Aerle, R., Stone, D. M., Paley, R., Trent, A., Katsiadaki, I., Higman, W. A., … Hartnell, R. E. (2022). A seafood risk tool for assessing and mitigating chemical and pathogen hazards in the aquaculture supply chain. Nature Food, 3(2), Article 2. [CrossRef]
- Suebsing, R. , Kampeera, J., Sirithammajak, S., Withyachumnarnkul, B., Turner, W., & Kiatpathomchai, W. (2015). Colorimetric Method of Loop-Mediated Isothermal Amplification with the Pre-Addition of Calcein for Detecting Flavobacterium columnare and its Assessment in Tilapia Farms. Journal of Aquatic Animal Health, 27(1), Article 1. [CrossRef]
- Sukonta, T. , Senapin, S., Taengphu, S., Hannanta-anan, P., Kitthamarat, M., Aiamsa-at, P., & Chaijarasphong, T. (2022). An RT-RPA-Cas12a platform for rapid and sensitive detection of tilapia lake virus. Aquaculture, 560, 738538. [CrossRef]
- Sullivan, T. J. , Dhar, A. K., Cruz-Flores, R., & Bodnar, A. G. (2019). Rapid, CRISPR-Based, Field-Deployable Detection Of White Spot Syndrome Virus In Shrimp. Scientific Reports, 9(1), Article 1. [CrossRef]
- Sutherland, B. J. G. , Finke, J. F., Saunders, R., Warne, S., Schulze, A. D., Strohm, J. H. T., Chan, A. M., Suttle, C. A., & Miller, K. M. (2022). Metatranscriptomics reveals a shift in microbial community composition and function during summer months in a coastal marine environment. Environmental DNA. [CrossRef]
- Tian, Y., Liu, T., Liu, C., Xu, Q., & Liu, Q. (2022). Pathogen detection strategy based on CRISPR. Microchemical Journal, 174, 107036. [CrossRef]
- Tian, Z. , Yang, L., Qi, X., Zheng, Q., Shang, D., & Cao, J. (2022). Visual LAMP method for the detection of Vibrio vulnificus in aquatic products and environmental water. BMC Microbiology, 22(1), Article 1. [CrossRef]
- United Nations. (2016). The 2030 Agenda for Sustainable Development.
- Urban, L. , Holzer, A., Baronas, J. J., Hall, M. B., Braeuninger-Weimer, P., Scherm, M. J., Kunz, D. J., Perera, S. N., Martin-Herranz, D. E., Tipper, E. T., Salter, S. J., & Stammnitz, M. R. (2021). Freshwater monitoring by nanopore sequencing. eLife, 10, e61504. [CrossRef]
- Verschuere, L. , Rombaut, G., Sorgeloos, P., & Verstraete, W. (2000). Probiotic Bacteria as Biological Control Agents in Aquaculture. Microbiology and Molecular Biology Reviews, 64(4), Article 4.
- Wang, P. , Guo, B., Zhang, X., Wang, Y., Yang, G., Shen, H., Gao, S., & Zhang, L. (2023). One-Pot Molecular Diagnosis of Acute Hepatopancreatic Necrosis Disease by Recombinase Polymerase Amplification and CRISPR/Cas12a with Specially Designed crRNA. Journal of Agricultural and Food Chemistry, 71(16), 6490–6498. [CrossRef]
- Williams, M. A. , de Eyto, E., Caestecker, S., Regan, F., & Parle-McDermott, A. (2022). Development and field validation of RPA-CRISPR-Cas environmental DNA assays for the detection of brown trout (Salmo trutta) and Arctic char (Salvelinus alpinus). Environmental DNA. [CrossRef]
- Williams, M.-A. , O’Grady, J., Ball, B., Carlsson, J., de Eyto, E., McGinnity, P., Jennings, E., Regan, F., & Parle-McDermott, A. (2019). The application of CRISPR-Cas for single species identification from environmental DNA. Molecular Ecology Resources, 19(5), 1106–1114. [CrossRef]
- World Bank. (2014). Reducing Disease Risk in Aquaculture. http://hdl.handle.net/10986/18936.
- Wright, A. , Li, X., Yang, X., Soto, E., & Gross, J. (2023). Disease prevention and mitigation in US finfish aquaculture: A review of current approaches and new strategies. Reviews in Aquaculture, n/a(n/a), Article n/a. [CrossRef]
- Xia, X. , Yu, Y., Hu, L., Weidmann, M., Pan, Y., Yan, S., & Wang, Y. (2015). Rapid detection of infectious hypodermal and hematopoietic necrosis virus (IHHNV) by real-time, isothermal recombinase polymerase amplification assay. Archives of Virology, 160(4), Article 4. [CrossRef]
- Xiao, X., Lin, Z., Huang, X., Lu, J., Zhou, Y., Zheng, L., & Lou, Y. (2021). Rapid and Sensitive Detection of Vibrio vulnificus Using CRISPR/Cas12a Combined With a Recombinase-Aided Amplification Assay. Frontiers in Microbiology, 12. https://www.frontiersin.org/articles/10.3389/fmicb.2021.767315.
- Yang, W., Zheng, Z., Lu, K., Zheng, C., Du, Y., Wang, J., & Zhu, J. (2020). Manipulating the phytoplankton community has the potential to create a stable bacterioplankton community in a shrimp rearing environment. Aquaculture, 520, 734789. [CrossRef]
| 1 | https://www.biokeyqpcr.com/supplier-3083726-portable-qpcr-machine. |
| 2 | https://www.biokeyqpcr.com/quality-27511152-shrimp-tissues-vpa-vibrio-parahaemolyticus-real-time-pcr-detection-kit. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




