1. Introduction
Furunculosis is a bacterial disease that is caused by
Aeromonas salmonicida, which affects various species of salmonid and non-salmonid fish, in both freshwater and marine environments and is characterized by high mortality and morbidity.
A. salmonicida is a Gram-negative, facultatively anaerobic, and nonmotile bacterium, which is classified within the genus
Aeromonas and belongs to the class Gammaproteobacteria and the family Aeromonadaceae [
1]. The bacterium exhibits a rod-shaped morphology, measures approximately 1.3–2.0 by 0.8–1.3 μm, and displays optimal growth at temperatures ranging from 22 to 25 °C. Moreover, it possesses the ability to both ferment and oxidize glucose and exhibits positive results in catalase and cytochrome oxidase tests. It particularly affects salmonids and trout, thereby making it a significant concern in aquaculture.
A. salmonicida can be divided into five subspecies:
salmonicida,
smithia,
achromogenes,
masoucida, and
pectinolytica [
2,
3,
4,
5,
6,
7,
8,
9]. The
A. salmonicida subspecies
salmonicida represents the classical causative agent of furunculosis in salmonids, although atypical strains of
A. salmonicida such as
masoucida,
achromogenes, and
smithia can cause a variant in non-salmonids and warm-water fish known as atypical furunculosis [
10,
11,
12]. These strains show optimal growth at temperatures ranging from 22 to 25 °C. Notably, among the known subspecies of
A. salmonicida, only
A. salmonicida pectinolytica has been observed growing at temperatures exceeding 35 °C.
The first report of
A. salmonicida atypica isolates in Atlantic salmon (
S. salar) cultured in Chilean seawater was [
13]. Subsequently,
atypical furunculosis spread rapidly in estuarine, marine, and freshwater fish farms in 2000, with the freshwater stage of the disease only being reported in the localities of Osorno and Punta Arenas in the fall of 2009 [
14]. In 2001, an injectable intraperitoneal vaccination containing inactivated
A. salmonicida atypica was initiated in Atlantic salmon (
S. salar), which reduced disease prevalence. However, despite vaccination becoming an established strategy in salmon farms, sporadic outbreaks continue to occur in Atlantic salmon (
S. salar) in lakes associated with vaccination management, estuary fattening, and endemic fish farms. Moreover, between 2021 and 2023, the prevalence of clinical cases in fish farms using recirculating aquaculture technology (RAS) increased among unvaccinated fish groups.
Recently, various aspects contributing to the genomic diversity of
A. salmonicida have been described. These distinctive patterns appear to be primarily associated with the accessory genome, thereby enabling differentiation between
psychrophilic and
mesophilic clades. The detection of a novel genomic entity, denoted as AsaGEI1a, within the 01-B526 strain has been a pivotal revelation since it encompasses a 51 kb chromosomal island, which contains prophage genes. This entity exhibits distinct variants, AsaGEI1b and AsaGEI2a, which are characterized by specific geographical distributions. Thus, 139 isolates were screened via PCR, and revealing a noteworthy geographical association, whereby AsaGEI1a and AsaGEI2a were predominantly identified in North American isolates. In contrast, AsaGEI1b was prevalent, while there was a conspicuous absence of other common genomic elements (GEIs) in European isolates. The identification of an additional prophage (prophage 3) intricately linked to AsaGEI2a further enriched the understanding of genomic dynamics by demonstrating an exclusive geographic prevalence in North America, with a single exception also found in a European isolate [
15]. Moreover, the establishment and implementation of a vapA-centric typing system facilitated the subtyping of 675 isolates of
A. salmonicida. This innovative approach led to the identification of nine novel subtypes (15–23), with certain subtypes exhibiting a discernible affinity for specific fish species, notably, Cyprinidae and Pleuronectidae [
16]. Furthermore, the sequencing and comparative analyses of entire genomes from
A. salmonicida strains provided further insights into the genomic distinctions between typical and atypical strains. The genomic disparities were scrutinized by evaluating the insertion sequences (ISs), along with identifying variations in the presence and absence of genes associated with virulence factors, transcriptional regulators, and non-coding RNAs. Additionally, a discernible role of the plasmidome in both the virulence and genomic adaptability of
A. salmonicida has been highlighted [
17]. Similarly, a comparative examination of 25 complete
A. salmonicida genomes revealed gene clusters exclusive to the
psychrophilic clades, which are associated with the lateral flagella, outer membrane proteins (including A-layer and T2SS proteins), and insertion sequences (specifically ISAs4, ISAs7, and ISAs29). In contrast, the
mesophilic clade exhibited the exclusive presence of MSH type IV pili, which suggests that discernible genetic factors are linked to the specific lifestyle of
A. salmonicida within this particular clade [
18]. Furthermore, the comprehensive analysis of extensive genomic data compiled from hundreds of
Aeromonas strains in aquaculture has enabled the detailed profiling of resistome patterns and can offer insights into associations by isolation year, country of origin, and species characteristics. Indeed, over 100 antibiotic resistance genes (ARGs) have been identified within a dataset encompassing approximately 400
A. salmonicida,
A. hydrophila,
A. veronii,
A. media, and
A. sobria genomes, which display either species-specific or non-species-specific patterns. Notably, species such as
A. salmonicida and
A. media exhibit higher proportions of species-specific ARGs, indicating distinctive patterns in the acquisition of antibiotic resistance. The prevalence of genes such as sul1, tet(A), and tet(D) has also been determined in specific regions of Asia and North America. Recently, there is a noteworthy surge in ARGs these aquatic environmental strains. This underscores the importance of regulating and controlling the excessive use of antibiotics in aquaculture to mitigate the escalation of antibiotic resistance [
19].
Prokaryotic defense mechanisms against foreign genomes involve crucial components known as restriction-modification (R-M) systems, which are present in a diverse range of unicellular organisms, encompassing both eubacteria and archaea [
20,
21]. These systems include two distinct enzymatic functions: restriction endonuclease (REase) and methyltransferase (MTase) functions. The REase targets and cleaves foreign DNA sequences at specific locations, while MTase activity maintains a distinction between the DNA of the host and any external DNA by adding methyl groups to the same specified DNA sequences. R-M systems exhibit diversity in their classifications, with distinctions based on cleavage position, co-factor requirements, sequence recognition, subunit composition, and substrate specificity, thus resulting in four primary types [
22]. However, R-M systems seem to play multifaceted roles in bacterial biology, and their functions extend beyond merely protecting against foreign genetic material, and include modulating horizontal gene transfer (HGT) and differentiating among bacterial strains. Due to the rampant impact of HGT on prokaryotic species boundaries, bacterial species are sustained through genetic isolation. REases operate as regulators by restricting foreign DNA with non-native methylation patterns to control the genetic influx. This creates a barrier, which could also serve to preserve the bacterial species [
23]. In this study, we describe the histopathological and genomic profile of re-emergent
A. salmonicida outbreaks in Chile. We corroborate recent findings regarding genomic variation patterns among
typical psychrophilic,
atypical psychrophilic, and
mesophilic strains of
A. salmonicida by comparing the genomes described in this study with 25 closed genomes available on NCBI [
24]. Finally, we describe the R-M landscape among
A. salmonicida strains.
4. Discussion
Atypical furunculosis represents an endemic bacterial disease with significant morbidity and mortality, which has traditionally been controlled through vaccination strategies. Notably, a substantial increase in the incidence of this disease was observed between 2022 and 2023, with a widespread geographical distribution in various freshwater production systems. This clinical and histopathological manifestation of cases is consistent with patterns described for furunculosis observed in salmonid fish. The histopathological findings are characterized by the presence of basophilic coccobacillary bacterial colonies in tissues, with inflammatory reactions being scant or absent. Other specific histological findings can be identified, such as cardiac inflammation, and hepatic and pancreatic necrosis, which are associated with co-infections with PRV and IPNV.
The genomic analysis of re-emergent
A. salmonicida outbreaks in Atlantic salmon (
S. salar) has provided significant insights into the pathogenesis and genomic diversity of this bacterial pathogen. The identified distinctive genomic patterns provide a comprehensive understanding of the complex dynamics associated with furunculosis, emphasizing its relevance in both salmonid and non-salmonid fish species across diverse aquatic environments. This study comprehensively examined clinical cases of atypical furunculosis in Atlantic salmon (
S. salar) aquaculture in Chile, offering valuable insights into the epidemiological and clinical aspects of this disease. The 4.66% observed mortality rate and an outbreak duration of 8 weeks serve as indicators of the disease’s significant impact on the aquaculture industry. The selection of bacterial isolates from clinically diseased fish, characterized by the highest frequency of clinical signs, enabled a detailed genomic investigation, and shed light on the pathogenesis and antibiotic resistance profiles of this bacterium in Chile. The gross pathology and histopathological findings corroborate previous observations [
14], which noted a range of clinical signs, including periorbital hemorrhages, abdominal hemorrhages, and the presence of furuncles in affected fish. The histopathological analysis underscores extensive tissue colonization by
A. salmonicida, primarily characterized by bacterial colonies that replace tissues with limited inflammatory responses. The prevalence of necrotic lesions in the epidermis and dermis is a key pathological feature, which aligns with the observed clinical manifestations.
A pivotal contribution of this study is the genomic exploration of
A. salmonicida, particularly in the distinctions between the mesophilic and psychrophilic strains, to confirm and extend recent findings in the field. The phylogenomic analysis revealed three distinct clusters, typical psychrophilic, atypical psychrophilic, and mesophilic, which demonstrates the genetic diversity within this species and validates previous observations [
17,
18]. The strains investigated in this study are classified within the atypical psychrophilic cluster. Five subspecies of
Aeromonas salmonicida have been described; however, the assignment of different isolates to each of these subspecies has been debated, primarily through a comparative analysis of the complete genomes. Such analyses have revealed that this subspecies classification does not reflect divergence at the core genome level. In contrast to the classification into typical psychrophilic, atypical psychrophilic, and mesophilic groups, the last appears to have a genomic basis that is evident at the phylogenetic level and in the distribution of ISs and coding sequences in the R-M system.
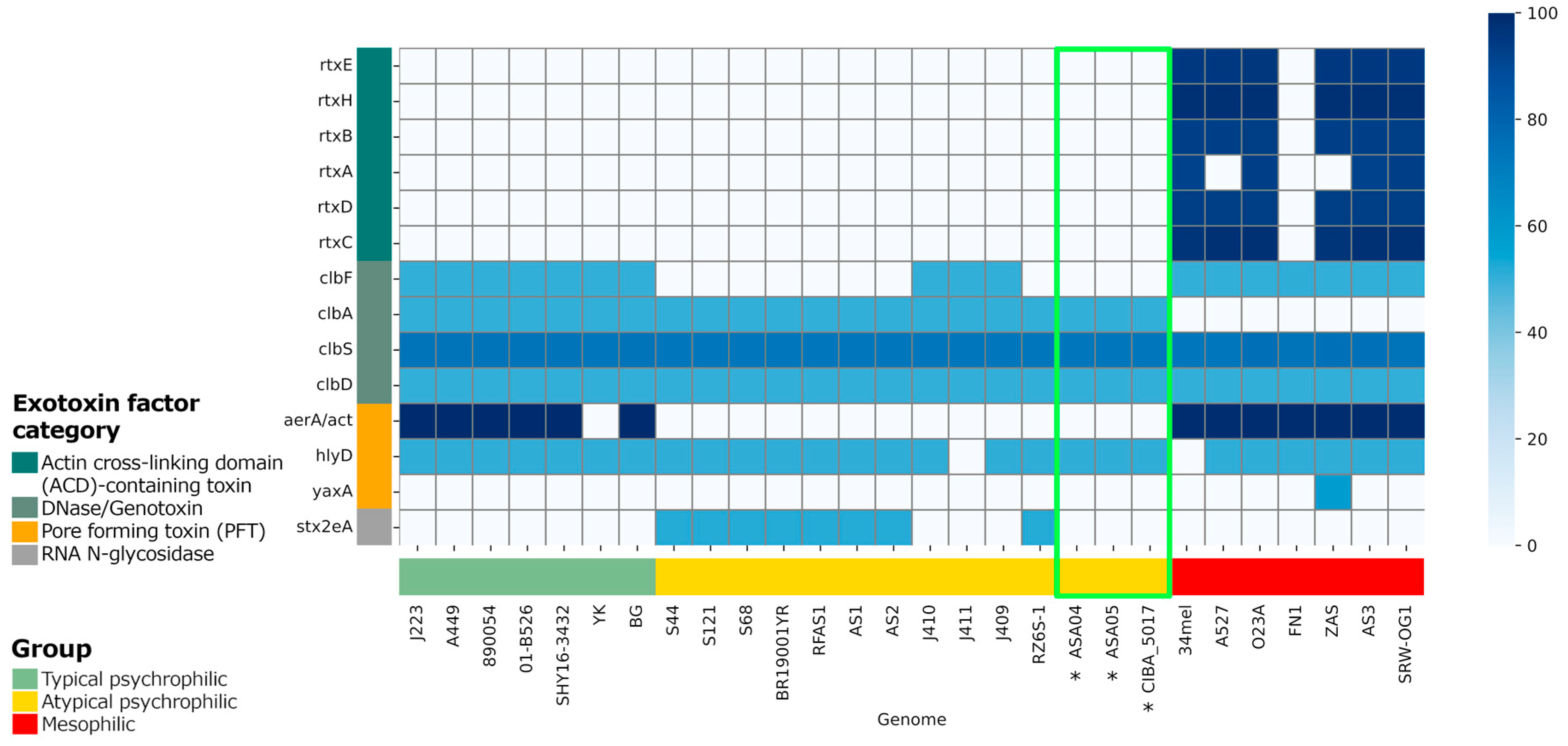
The unique gene families found in these strains highlight the genomic complexity within atypical psychrophilic A. salmonicida and further emphasize the need for tailored genomic characterization. Thus, the analysis of virulence factors is of paramount importance. Specifically, since variations in exotoxin factors, particularly the actin cross-linking domain (ACD)-containing toxins, are present in the mesophilic strains but not in the psychrophilic strains, the potential role of these factors in pathogenicity is underscored. Distinctions in the presence of virulence genes such as clbF and aerA/act further accentuate the genetic diversity within typical and atypical psychrophilic strains. The selective presence of the RNA N-glycosidase factor stx2eA among atypical psychrophilic strains serves as an essential marker for distinguishing between subgroups within this classification. Plasmid characterization reveals differences in plasmid content, emphasizing the contribution of each plasmid to virulence and genome plasticity. The presence of antibiotic resistance factors within these genomes raises concerns regarding the development of antibiotic resistance and necessitates careful management strategies in Chilean aquaculture settings.
Furthermore, the diverse patterns in the insertion sequences and restriction-modification system coding sequences underscore the genomic structural differences among strains. REases function as regulatory agents, limiting the entry of foreign DNA with non-native methylation patterns, to modulate genetic influx. This impediment, in effect, may also play a role in preserving bacterial species. Corroborating this, many
E. coli and
Salmonella enterica serovar Typhimurium strains possess specific genomic loci, referred to as “immigration control regions”, which are abundant in R-M systems [
49]. Therefore, R-M systems could achieve greater genetic isolation by regulating the uptake of environmental DNA. This would allow distinctive R-M patterns to be established among closely related strains, further genetically isolating them. As several genetic variants distinctly accumulate, the related bacterial strains gradually evolve and become more distant, until eventually becoming different species entirely. Additionally, studies on type III-like R-M enzymes have uncovered the function of these REases, which acted as a significant obstacle to the HGT in clinical variants of methicillin-resistant
S. aureus [
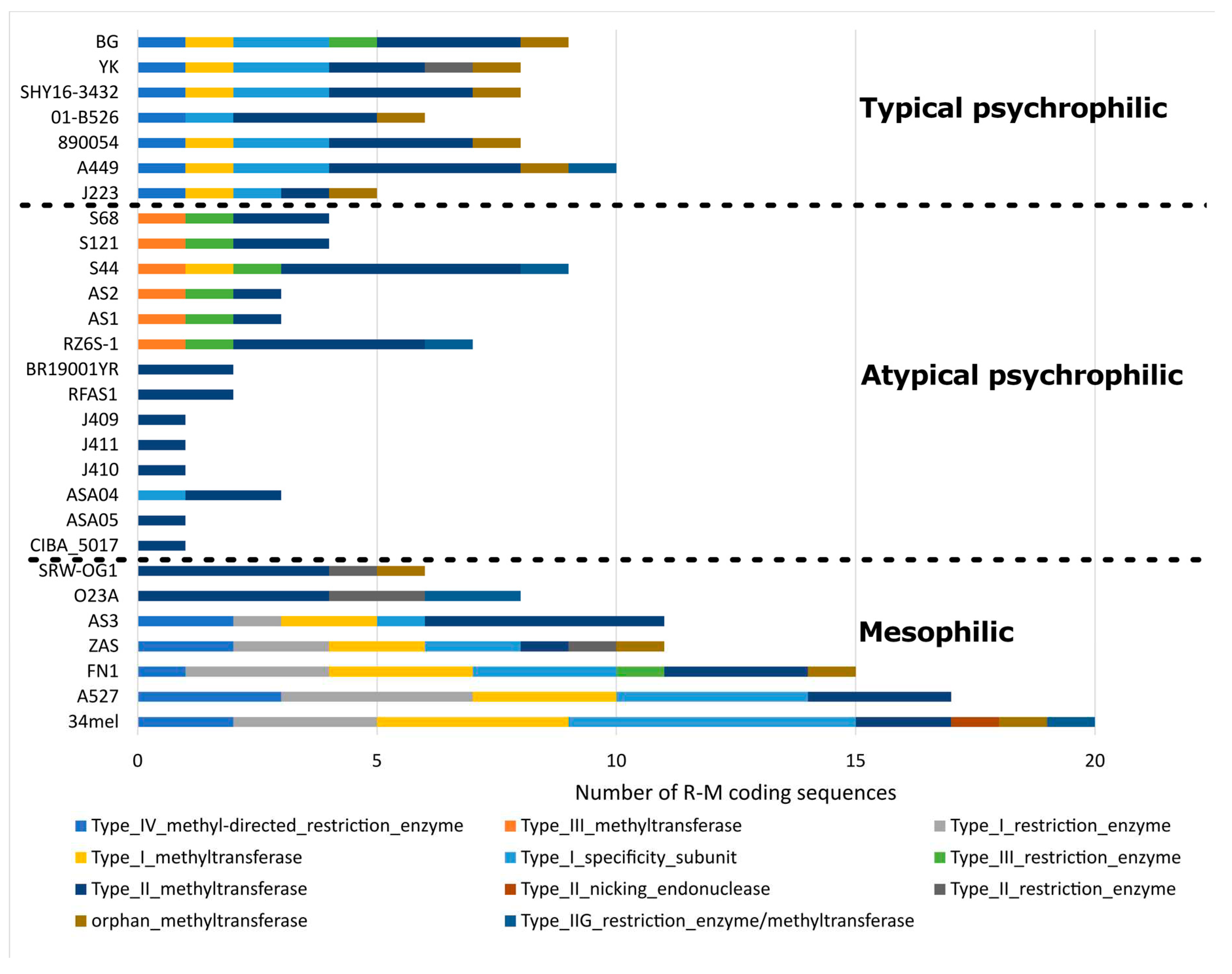
50]. The distinctive R-M coding sequence patterns observed among typical psychrophilic, atypical psychrophilic, and mesophilic
A. salmonicida strains highlight their genomic diversity. Further, two distinctive patterns of R-M coding sequences are observed within the atypical psychrophilic and mesophilic strains. However, the impact that the R-M system may have on the genomic diversity of
A. salmonicida requires further investigation.
Author Contributions
Conceptualization, M.G., A.M, R.S, and H.B; formal analysis, M.M.O., D.C, K.K, H.B; writing—original draft preparation, M.G., M.M.O, R.S, J.P.P, D.C, K.K.; writing—review and editing, M.G., M.M.O, A.M, J.P.P, H.B, M.K, F.K, S.B; funding acquisition, M.G. All authors have read and agreed to the published version of the manuscript.
Figure 1.
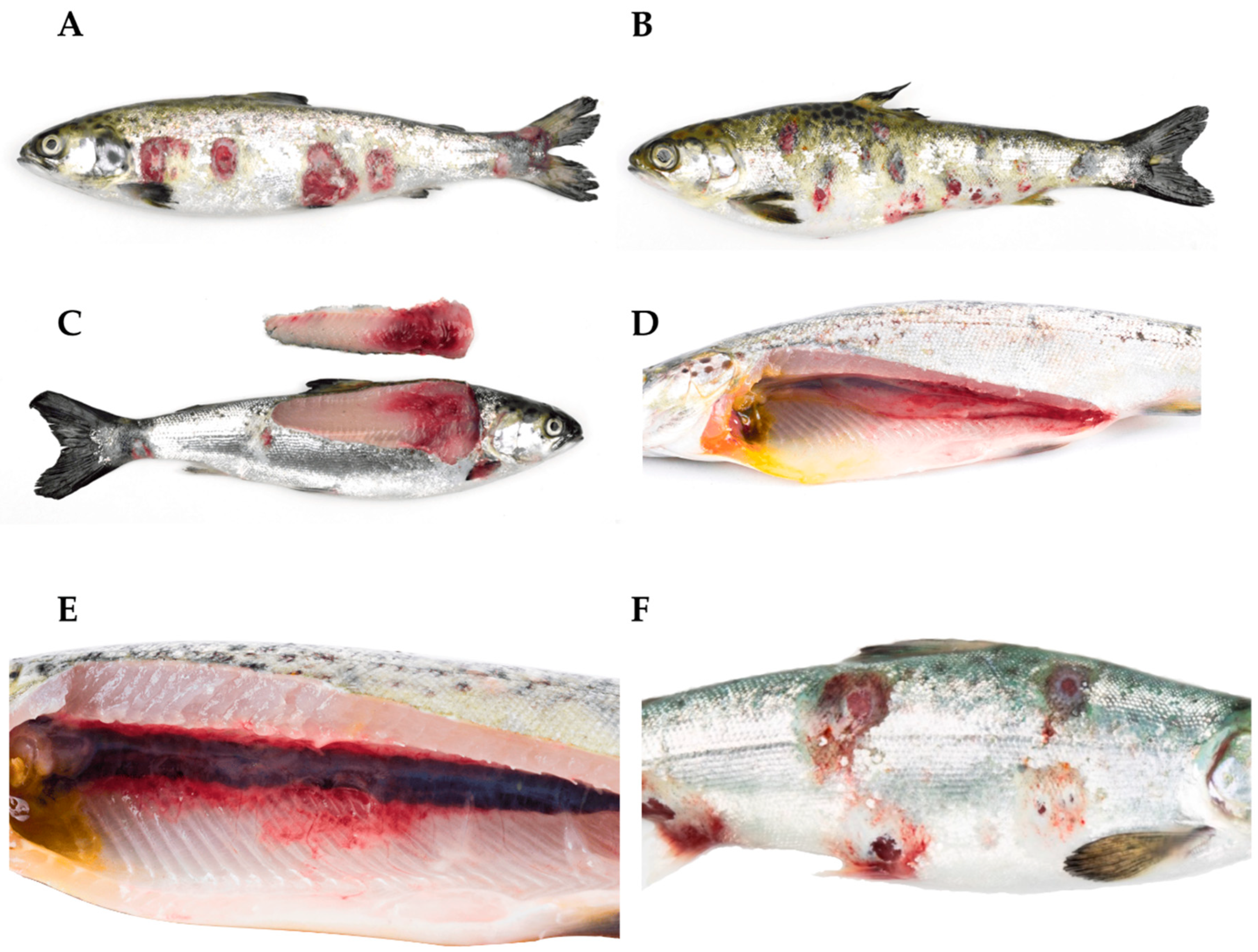
Atlantic salmon (Salmo salar) in the freshwater phase are affected by a clinical presentation of atypical furunculosis caused by atypical Aeromonas salmonicida. (A,B) Multiple ulcers are observed, with some surrounded by a white-colored border. (C) Loss of structure and muscle hemorrhages are observed. (D) Petechial hemorrhages are observed in the swim bladder. (E) Atlantic salmon (S. salar) affected by a clinical picture of atypical furunculosis (atypical A. salmonicida). The presence of perirenal muscle hemorrhages is observed. (F) Atlantic salmon (S. salar) affected by a clinical picture of atypical furunculosis (atypical A. salmonicida). Periocular hemorrhage, exophthalmia, and multiple ulcers are noted, some of which have coalesced.
Figure 1.
Atlantic salmon (Salmo salar) in the freshwater phase are affected by a clinical presentation of atypical furunculosis caused by atypical Aeromonas salmonicida. (A,B) Multiple ulcers are observed, with some surrounded by a white-colored border. (C) Loss of structure and muscle hemorrhages are observed. (D) Petechial hemorrhages are observed in the swim bladder. (E) Atlantic salmon (S. salar) affected by a clinical picture of atypical furunculosis (atypical A. salmonicida). The presence of perirenal muscle hemorrhages is observed. (F) Atlantic salmon (S. salar) affected by a clinical picture of atypical furunculosis (atypical A. salmonicida). Periocular hemorrhage, exophthalmia, and multiple ulcers are noted, some of which have coalesced.
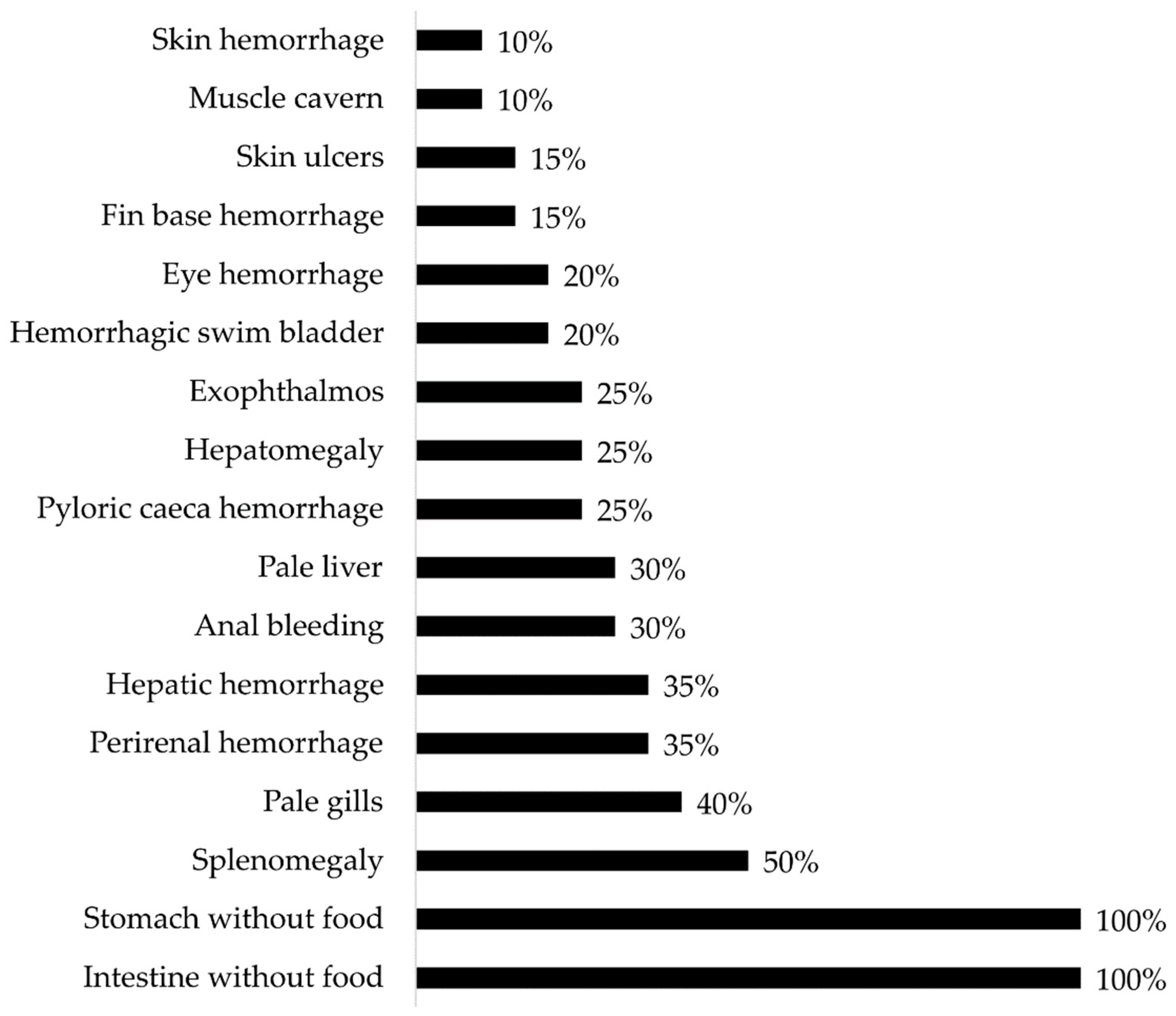
Figure 2.
Gross pathology findings were observed in Atlantic salmon (Salmo salar) presenting clinical cases of atypical furunculosis between 2022 and 2023, in a total sample size (n) of 294.
Figure 2.
Gross pathology findings were observed in Atlantic salmon (Salmo salar) presenting clinical cases of atypical furunculosis between 2022 and 2023, in a total sample size (n) of 294.
Figure 3.
(A) Atlantic salmon (Salmo salar), skin, and red muscle (H & E). Colonies of bacteria with coccobacillary morphology are observed in red muscle, consistent with atypical Aeromonas salmonicida infection (arrow). (B) Atlantic salmon (S. salar), red muscle (H & E). Observation of diffused infection of bacteria with coccobacillary morphology, consistent with atypical A. salmonicida infection. Additionally, hemorrhage and necrosis of the muscle tissue are observed. (C) Surrounding the muscular tissue, the presence of coccobacillary bacterial colonies consistent with infection by atypical A. salmonicida is observed (arrow). Additionally, muscular necrosis and hemorrhage are observed. (D) Atlantic salmon (S. salar), spleen (H & E). In the splenic parenchyma, multiple colonies of bacteria with coccobacillary morphology are observed, consistent with atypical A. salmonicida infection (arrows). (E) Colonies of bacteria with coccobacillary morphology are observed in the auricle, consistent with atypical A. salmonicida infection (arrow). (F) Atlantic salmon (S. salar), gill (H & E). Bacterial infiltration of the center gill filament with coccobacillary morphology is observed, consistent with atypical A. salmonicida infection (arrow).
Figure 3.
(A) Atlantic salmon (Salmo salar), skin, and red muscle (H & E). Colonies of bacteria with coccobacillary morphology are observed in red muscle, consistent with atypical Aeromonas salmonicida infection (arrow). (B) Atlantic salmon (S. salar), red muscle (H & E). Observation of diffused infection of bacteria with coccobacillary morphology, consistent with atypical A. salmonicida infection. Additionally, hemorrhage and necrosis of the muscle tissue are observed. (C) Surrounding the muscular tissue, the presence of coccobacillary bacterial colonies consistent with infection by atypical A. salmonicida is observed (arrow). Additionally, muscular necrosis and hemorrhage are observed. (D) Atlantic salmon (S. salar), spleen (H & E). In the splenic parenchyma, multiple colonies of bacteria with coccobacillary morphology are observed, consistent with atypical A. salmonicida infection (arrows). (E) Colonies of bacteria with coccobacillary morphology are observed in the auricle, consistent with atypical A. salmonicida infection (arrow). (F) Atlantic salmon (S. salar), gill (H & E). Bacterial infiltration of the center gill filament with coccobacillary morphology is observed, consistent with atypical A. salmonicida infection (arrow).
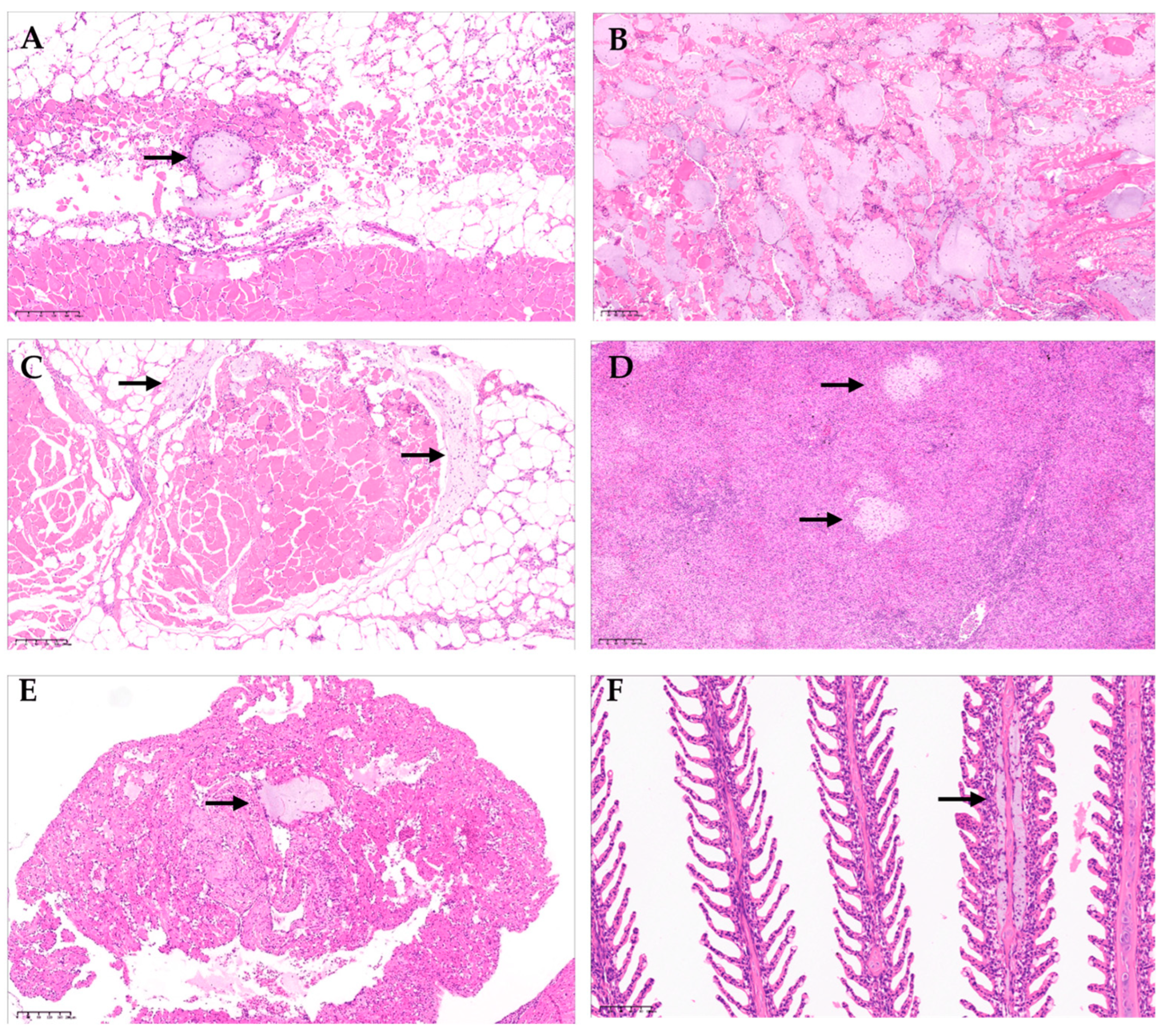
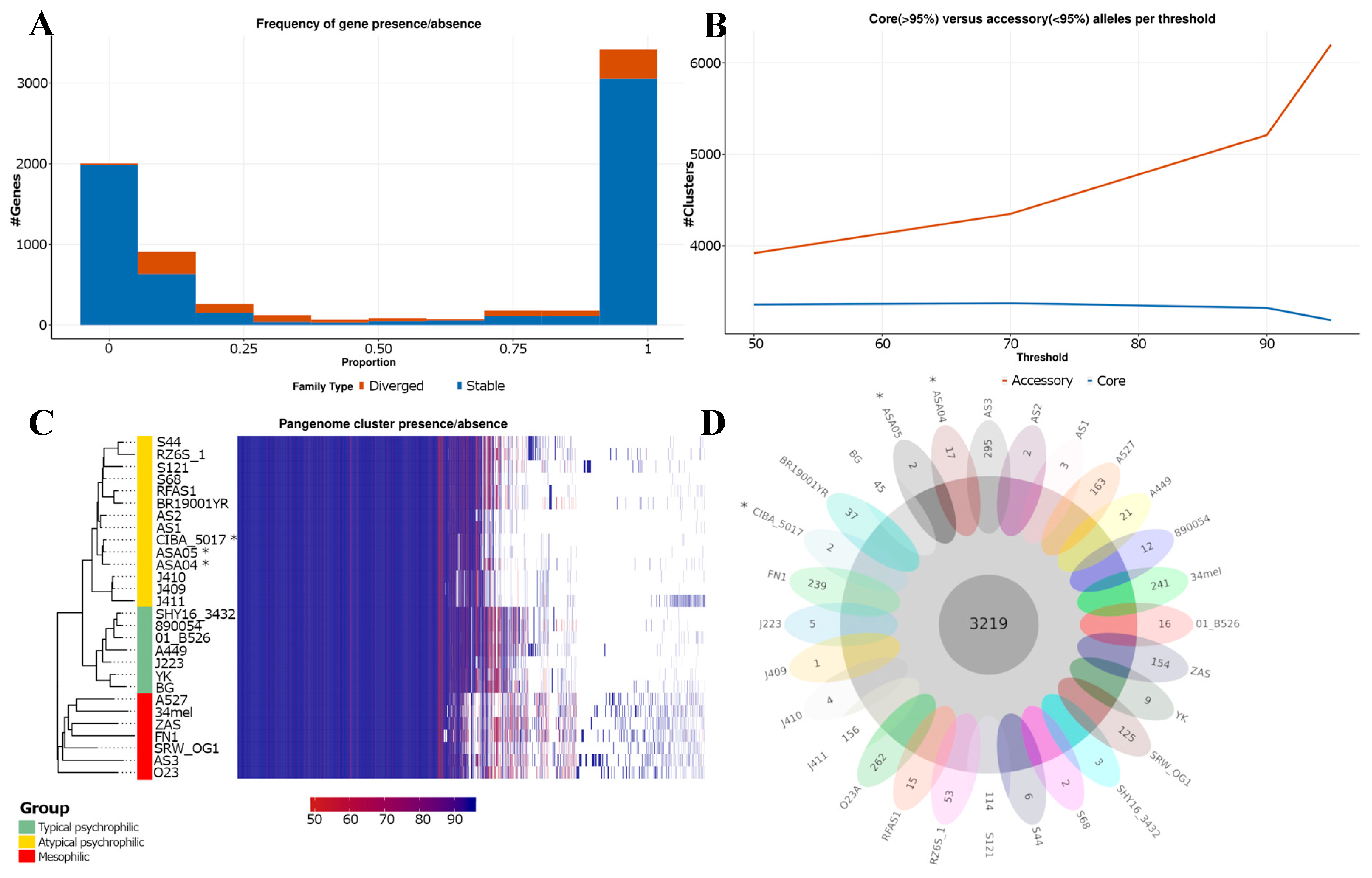
Figure 4.
Pan-genome analysis of Aeromonas salmonicida strains. (A) The proportion of genomes with gene families present. Gene families are classified as stable (in blue) when they contain only a single allele with a 98% similarity in their amino acid sequence, and as divergent (in red) when they have more than one allele. (B) Core and accessory gene/allele estimates. The estimates represent “allelic” variation. Core gene families are characterized as being present in > 95% of genomes. (C) Shared gene presence per isolate, ordered alongside the phylogenetic tree. Gene family presence is indicated by a blue block per column. (D) Core and unique gene families per genome. *: Genomes described in this study.
Figure 4.
Pan-genome analysis of Aeromonas salmonicida strains. (A) The proportion of genomes with gene families present. Gene families are classified as stable (in blue) when they contain only a single allele with a 98% similarity in their amino acid sequence, and as divergent (in red) when they have more than one allele. (B) Core and accessory gene/allele estimates. The estimates represent “allelic” variation. Core gene families are characterized as being present in > 95% of genomes. (C) Shared gene presence per isolate, ordered alongside the phylogenetic tree. Gene family presence is indicated by a blue block per column. (D) Core and unique gene families per genome. *: Genomes described in this study.
Figure 5.
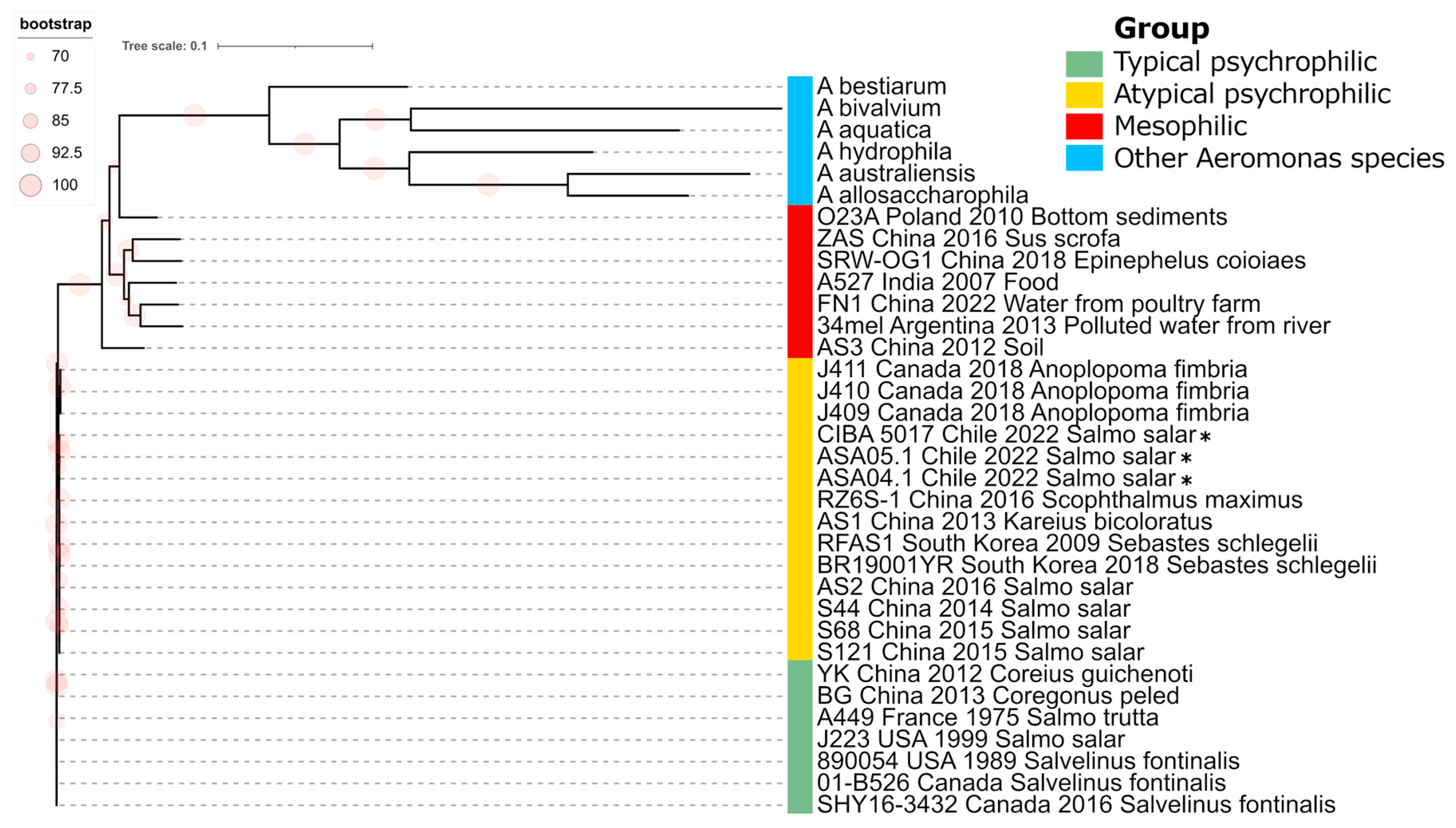
Phylogenomic analysis of Aeromonas salmonicida strains. Maximum likelihood phylogenic tree constructed from a core genome alignment encompassing 89,915 SNPs. *: Genomes described in this study. Bootstrap frequencies above 70% are represented as red circles. Other representative Aeromonas species are included as outgroup isolates.
Figure 5.
Phylogenomic analysis of Aeromonas salmonicida strains. Maximum likelihood phylogenic tree constructed from a core genome alignment encompassing 89,915 SNPs. *: Genomes described in this study. Bootstrap frequencies above 70% are represented as red circles. Other representative Aeromonas species are included as outgroup isolates.
Figure 6.
Distribution of restriction-modification (R-M) coding sequences.
Figure 6.
Distribution of restriction-modification (R-M) coding sequences.
Figure 7.
Profile of exotoxin virulence factors in complete Aeromonas salmonicida genomes. The presence and absence of exotoxin factors were predicted based on local alignments with the VFDB entries. The similarity percentage with the identified factors is shown. *: Genomes reported in this study, with their results highlighted in the green box.
Figure 7.
Profile of exotoxin virulence factors in complete Aeromonas salmonicida genomes. The presence and absence of exotoxin factors were predicted based on local alignments with the VFDB entries. The similarity percentage with the identified factors is shown. *: Genomes reported in this study, with their results highlighted in the green box.
Figure 8.
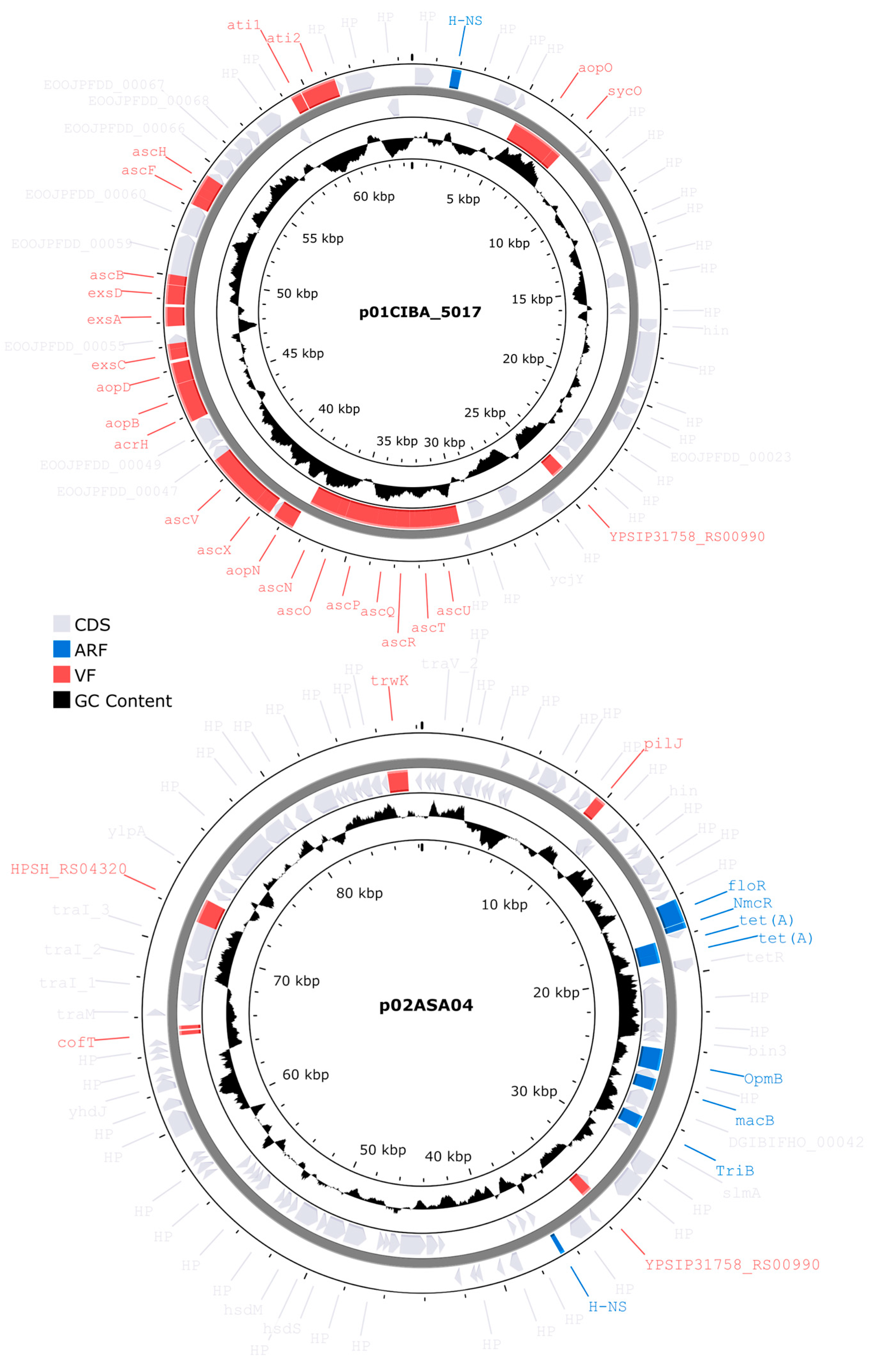
Virulence and antibiotic resistance factors identified in Aeromonas salmonicida plasmids.
Figure 8.
Virulence and antibiotic resistance factors identified in Aeromonas salmonicida plasmids.
Figure 9.
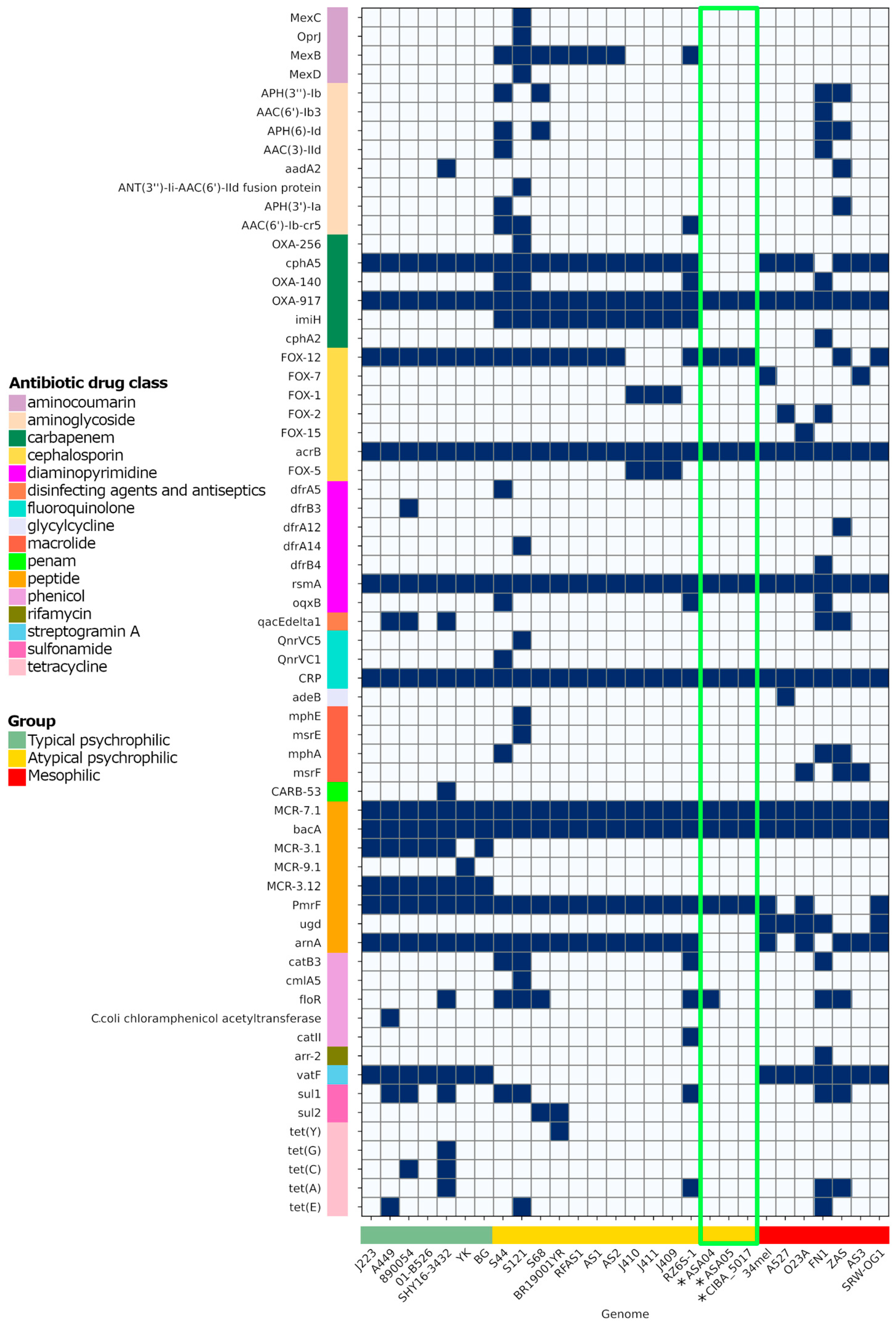
Antibiotic resistance repertoire in Aeromonas salmonicida complete genomes. The presence and absence of the factor are represented in blue and white, respectively. *: Genomes reported in this study, with their results highlighted in the green box.
Figure 9.
Antibiotic resistance repertoire in Aeromonas salmonicida complete genomes. The presence and absence of the factor are represented in blue and white, respectively. *: Genomes reported in this study, with their results highlighted in the green box.
Table 1.
Epidemiological background and laboratory results for the cases analyzed in this study.
Table 1.
Epidemiological background and laboratory results for the cases analyzed in this study.
| Case ID |
4978 |
4988 |
5131 |
7003 |
7031 |
7081 |
7088 |
7124 |
7167 |
7175 |
7174 |
7215 |
7216 |
7214 |
| Date |
10-Mar |
14-Mar |
11-May |
23-May |
03-Jun |
23-Jun |
30-Jun |
21-Jul |
08-Aug |
12-Aug |
12-Aug |
18-Aug |
24-Aug |
25-Aug |
| Region |
X |
X |
X |
X |
X |
XII |
XII |
XII |
X |
X |
X |
XII |
XII |
X |
| Salinity |
FW |
FW |
BW |
FW |
FW |
BW |
BW |
BW |
BW |
BW |
FW |
BW |
BW |
BW |
| Development stage |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
| Fish sampled |
20 |
7 |
10 |
30 |
6 |
7 |
7 |
10 |
10 |
11 |
24 |
1 |
1 |
1 |
| Average ASA Ct |
22.8 |
26.1 |
29.2 |
28.4 |
23.8 |
27.5 |
26.2 |
24.4 |
27.1 |
29.8 |
21.4 |
26.1 |
32.8 |
22.5 |
| Coinfection |
PRV |
IPNV |
IPNV |
R. salmoninarum |
PRV |
IPNV-F.psy |
IPNV-PRV |
IPNV-PRV |
- |
- |
IPNV-PRV |
- |
- |
- |
| Bacterial isolation |
Positive |
Positive |
Positive |
- |
- |
- |
- |
- |
Positive |
- |
- |
- |
- |
- |
| GenBank accession |
- |
- |
- |
- |
- |
- |
- |
- |
- |
- |
- |
- |
- |
- |
Table 1.
(continued) Epidemiological background and laboratory results for the cases analyzed in this study.
Table 1.
(continued) Epidemiological background and laboratory results for the cases analyzed in this study.
| Case ID |
7198 |
7213 |
7217 |
7385 |
7388 |
7387 |
7412 |
7416 |
7435 |
CIBA-5017 |
ASA04 |
ASA05 |
| Date |
26-Aug |
26-Aug |
06-Sept |
20-Oct |
21-Oct |
21-Oct |
27-Oct |
28-Oct |
04-Nov |
25-Mar |
22-Nov |
23-Nov |
| Region |
X |
X |
X |
X |
X |
X |
X |
X |
X |
X |
X |
X |
| Salinity |
FW |
FW |
FW |
FW |
BW |
FW |
FW |
FW |
BW |
FW |
FW |
FW |
| Development stage |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Pre-smolt |
Smolt |
Smolt |
Smolt |
| Fish sampled |
10 |
1 |
7 |
24 |
24 |
26 |
15 |
10 |
15 |
20 |
25 |
11 |
| Average ASA Ct |
29.6 |
26.2 |
23.2 |
30.6 |
26.6 |
24.3 |
30.8 |
29.1 |
27.0 |
28.8 |
29.7 |
26.1 |
| Coinfection |
IPNV |
- |
IPNV |
- |
- |
- |
- |
PRV |
PRV |
- |
IPNV-PRV |
IPNV-PRV |
| Bacterial isolation |
Positive |
- |
- |
Negative |
- |
Positive |
Negative |
Negative |
- |
Positive |
Positive |
Positive |
| GenBank accession |
- |
- |
- |
- |
- |
- |
- |
- |
- |
- |
- |
- |