1. Introduction
Sesame (
Sesamum indicum L.) is among the oldest oilseed crops known to humanity and is often referred to as the “queen of oilseed crops.” It is a diploid species (2n = 2x = 26 chromosomes), although variability in chromosome numbers has been identified in other species within the genus
Sesamum [
1]. Species such as
S. calycinum,
S. latifolium, and
S. angolense are also diploids (2n = 2x) but with 32 chromosomes [
1]. Sesame’s seeds are rich in edible oil, containing more than 50% oil content. Furthermore, they are a rich source of tocopherol, sterol, and sesamol, compounds known for their medicinal properties. Seed meal contains 35–50% protein and is a rich source of amino acids such as methionine and tryptophan. Despite its usefulness as an oilseed crop, or wide use in confectionary products, little efforts were made for its varietal improvement to enhance its yield potential or low research priorities were given by scientific academia and industry. It is considered an orphan crop that grows widely from 40°N to 40°S latitude, adapting to various climates. However, the most intensive cultivation occurs in the northern hemisphere, particularly between the equator and 30°N. This distribution is influenced by suitable growing conditions, historical practices, and market demands [
2]. In addition, it is grown on an area of 12.81 million ha and 6.55 million tons were produced worldwide in 2022 [3, 4]. It was thought to be originated in Africa which is the center of diversity, and several species were identified in the wild. It is still grown on a large scale in Africa with Sudan as its largest producer in the world followed by Myanmar and India [3, 4].
Sesame is a self-pollinated, semi-erect plant that grows to a height of 0.6 to 1.2 meters. Its ovate or lanceolate leaves vary, with basal leaves being tri-lobed and upper leaves irregular and serrated. As a drought-tolerant crop, sesame prefers well-drained soils with a pH of 5.5 to 6.8 and is suited to temperate tropical climates. The bell-shaped flowers have petal colours ranging from pale yellow to purple. However, sesame has a low yield potential and typically responds poorly to farm inputs, thus making it an unattractive option for growers compared to other oilseed crops. Continuous cultivation of sesame in the same field leads to declining yield, protein, and mineral content over time [
5]. Additionally, the crop faces challenges such as shattering at maturity, making harvesting difficult, and insect and disease infestations causing significant yield losses globally. Concrete efforts in breeding and genetics are needed to develop elite, high-yielding sesame cultivars with resistance to disease and insects. Current developments in breeding, genomics, and marker-assisted selection, therefore, highlight the importance of this crop.
2. Germplasm Resources Collection, and Diversity Study in Sesame
2.1. Germplasm Resources Collection and Conservation
The primary breeding objectives for sesame are to increase seed yield, improve plant morphology, enhance tolerance to biotic and abiotic stresses, develop indehiscent capsules, and improve oil quality. These varied breeding goals highlight the critical importance of germplasm collection and conservation. A robust and diverse genetic repository is essential for breeders, providing access to a wide range of traits and alleles needed to develop improved cultivars that meet evolving agricultural challenges and consumer demands. Maintaining a comprehensive germplasm collection ensures the preservation of genetic diversity, which is crucial for the long-term sustainability and advancement of sesame breeding programs. This genetic resource is the foundation for future crop improvement innovations, climate change adaptation, and food security. Therefore, germplasm collection and conservation is a continuous process to protect wild resources for sustainable agriculture. Evidence suggests that sesame originated from wild populations native to the Indian subcontinent or parts of Pakistan or Afghanistan. Several wild species of the Sesamum genus are also native to Africa, but no study has specifically classified any species as potential progenitors of cultivated sesame.
Sesamum indicum germplasm is conserved at the International Crops Research Institute for the Semi-Arid Tropics (ICRISAT, India) and the National Bureau of Plant Genetic Resources (NBPGR, India). ICRISAT conserves 1,302 accessions of sesame germplasm, including wild accessions and those collected from international germplasm exchange. Similarly, NBPGR conserves 5,288 accessions of sesame germplasm [
6]. Germplasm resources from other countries are also being conserved, with 11,116 accessions at the USDA-ARS repository, 9,221 accessions in the Vavilov Institute of Plant Industry (VIR, Russia), and 3,317 accessions at the National Institute of Agricultural Science and Technology (NIAST, South Korea). Other collections include the Brazilian Gene Bank (BGB) with 1,455 accessions and the Nigerian National Gene Bank (NNGB) with 1,039 accessions [
6]. Evaluation and characterization of these accessions for morphological, agronomical, and molecular traits have been carried out, resulting in the identification of unique genotypes with high yield, disease resistance, and quality traits. For example, the FGsesame (
http://ncgr.ac.cn/SesameFG/) database includes core collections of 705 sesame accessions from 29 countries, each identified based on origin, ecotype, sequencing coverage, and group information [
7].
Germplasm resources are also crucial for ecosystem sustainability and nutritional security. The utilization of these resources in breeding programs, therefore, can lead to the development of improved sesame cultivars. The wild relatives of sesame contain unique genes that are not found in cultivated varieties [
8]. It's therefore, important to protect them from over-exploitation by local communities for culinary, medicinal, and cosmetic uses to prevent their extinction and maintain ecosystem balance.
2.2. Diversity Studies in Sesame
Studies have demonstrated high genetic diversity among wild sesame relatives and cultivated germplasm from various regions. Wild relatives of sesame offer valuable traits for breeding programs, such as cytoplasmic male sterility, heat tolerance, disease resistance, and fatty acid modification. For instance,
Sesamum radiatum has more linolenic acid than other species [
9]. The wild population of
Sesamum orientale closely resembles cultivated types (
S. indicum) in both morphology and cytogenetics. Both species are diploid with 26 chromosomes and produce fertile hybrids under spontaneous or artificial reciprocal hybridization. CpDNA-based fingerprinting also supports a close relationship between
S. orientale and
S. indicum [
10]. Some authors suggest its origin from the African population of
Sesamum latifolium.
Crosses between wild and cultivated species have successfully introgressed desirable traits, such as charcoal rot resistance [
11]. However, crosses between
S. indicum and
S. latifolium were largely unsuccessful, producing only a few seeds due to chromosomal number and structural differences (
S. latifolium has 2n = 32 chromosomes). Both species also show qualitative differences, such as the absence of sesamolin in
S. latifolium, indicating both cytological and morphological differentiation between the species.
Molecular markers have been used to assess diversity among sesame accessions. Isozyme analysis and morphological trait comparisons have differentiated wild and cultivated species [1, 12]. The presence of polymorphic co-dominant cathodic and anodic peroxidase isozymes, Co-Px-2 and An-Px-2, is unique to wild sesame germplasm. Additionally, the Sh-1 marker distinguishes S. indicum from wild S. mulayanum. Similarly, the Sh-1 marker can distinguish S. orientale from other wild accessions, confirming genetic diversity in wild sesame.
The development of molecular markers, such as simple sequence repeats (SSR) and single nucleotide polymorphisms (SNP), has also been instrumental in genetic diversity analysis and mapping of quantitative trait loci (QTL) in sesame [
13]. SSR-based research have shown genetic diversity indices and divergence among Indian accessions [
14]. EST databases were exploited for the development and characterization of gene-derived SSR markers using the computer-based software (MISA), while SNPs were detected and visualized using the QualitySNPng software tool (
http://www.bioinformatics.nl/QualitySNPng). SNP variants were identified through a genome comparison of sesame. The sesame genome survey identified SSR markers that could be exploited to determine genetic diversity and QTL mapping of economic traits. There were 23,438 SSRs (five repeats), with dinucleotides being the most prevalent repeat pattern (84.24%), followed by trinucleotides (13.53%), tetranucleotides (1.65%), pentanucleotides (0.3%), and hexanucleotides (0.28%) [
13]. Identifying microsatellites in the sesame genome led to the development of 218 polymorphic SSR markers for screening sesame germplasm for genetic diversity analysis [
13]. The sesame cultivar “Sweeta” genome sequences were obtained through Illumina paired-end sequencing and 454 shotgun sequence technologies. Improve the assembly and annotation of the sesame genome by high-quality sesame genome assembly using PacBio and Hi-C technology, improving continuity and chromosome anchoring. The new assembly identifies 24,345 protein-coding genes and reveals ancient whole-genome duplication events. These advancements will support future genetic studies and breeding efforts in sesame [
15]. The “GINMicrosatDb” database of microsatellites was developed, containing five sets of primer pairs for each microsatellite locus [
16]. The database also included GC content, melting point, and flanking sequences [
16].
Transcriptomic profiling of sesame genes revealed 57 MADS-box genes across 14 linkage groups that are essential for sesame development and growth [
7]. Structural analysis and functional differences among MADS-box genes showed similarities and differences with model species
Arabidopsis thaliana and
Solanum lycopersicum [
7]. The MIKCC-type MADS-box genes are also shown related to sesame flower initiation and seed development [
7]. Transcriptomic analysis of interspecific hybrids has also identified genes related to disease resistance [
17]. Studies of 4.4 Mbps related to fatty acid metabolism show genetic diversity loss during domestication of sesame [
18]. Expression profiles of traits such as capsule, seed leaf, and root development in variety “Zhongzhi13” and the transcriptomic profile of genes related to waterlogging tolerance in “ZZM2541” and oil content in high oil-yielding variety “Zhongfengzhi1” were identified [
7]. Candidate genes related to seed yield, lipid metabolism, phenological traits, or disease resistance are available in the database [
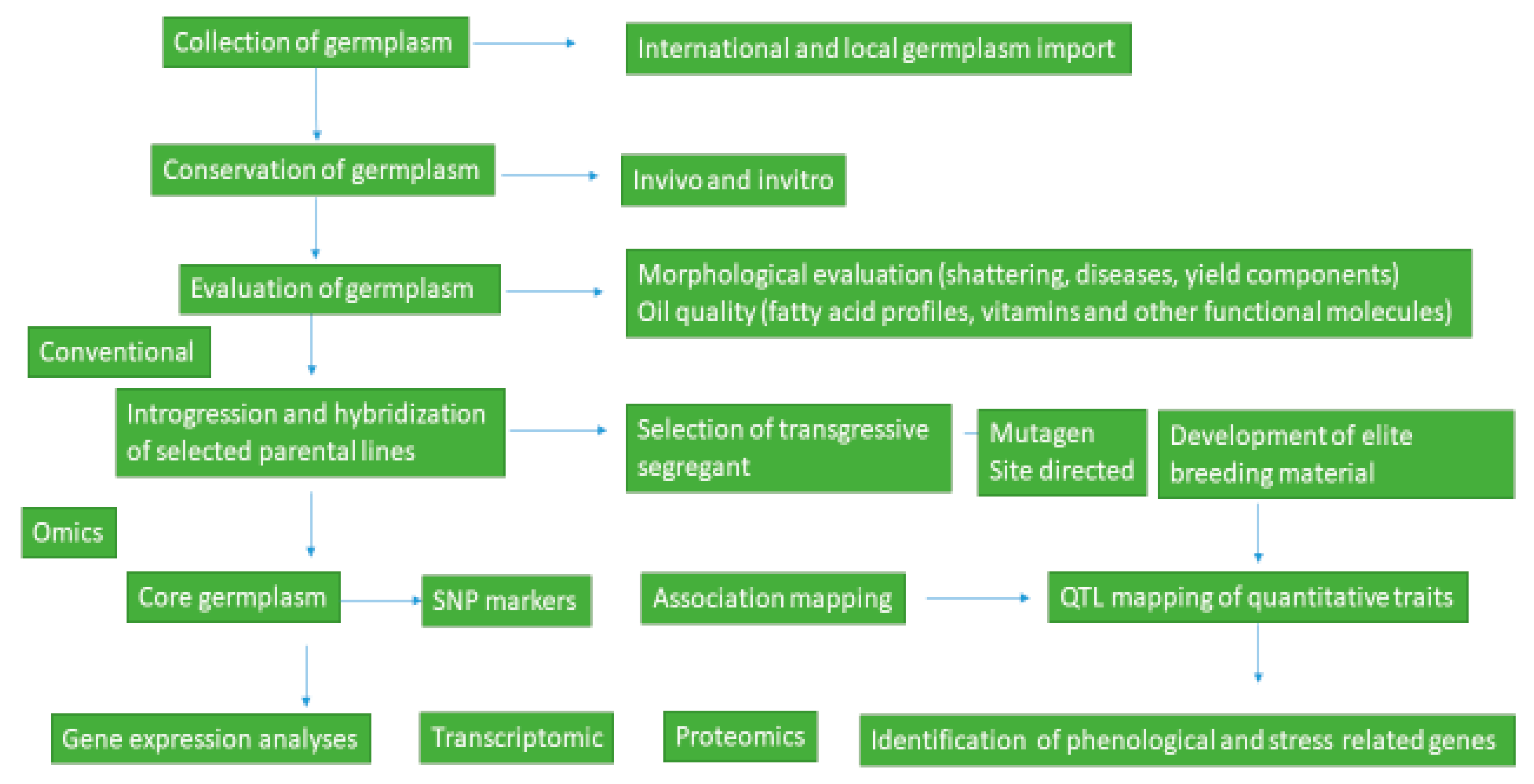
7]. Hence, further evaluation and characterization of accessions and wild relatives for morphological, agronomical, and molecular traits should be done to develop improved sesame genotypes with high yield, disease resistance, and quality traits. A comprehensive summary illustrating the concepts discussed in the text on sesame breeding and conservation is illustrated in
Figure 1 below.
3. Oil Content and Fatty Acids Composition Analysis in Sesame Germplasm
Oil content in sesame accessions varies depending on genotype and environmental interaction, ranging from 45% to 53% among accessions from Turkey and 11 other countries [
19]. The predominant fatty acids in sesame oil are oleic acid (36–43%), linoleic acid (39–46%), and palmitic acid (8–10%) [
19]. Sesame oil is rich in unsaturated fatty acids, making it suitable for culinary and industrial applications. A recent study utilized genome-wide association analysis (GWAS) and transcriptome analysis to identify novel loci and regulatory genes involved in fatty acid biosynthesis [
20]. The study discovered that specific gene clusters, including FAD2,
LOC10515945,
LOC105161564, and
LOC105162196, play crucial roles in regulating unsaturated fatty acid biosynthesis. Additionally, a regulatory co-expression network was constructed based on differentially expressed genes, aiding in understanding the transcriptional and translational regulation of oil biosynthesis in sesame seeds. These findings provide a molecular platform for improving the quality and yield of sesame oil, potentially benefiting breeding programs aimed at developing sesame cultivars with higher levels of unsaturated fatty acids.
Major lignans in sesame seeds include sesamin and sesamolin, with higher concentrations found in white-seeded cultivars [
21]. Sesame seeds are also a rich source of minerals, with calcium being the predominant cation, followed by potassium and magnesium [
22]. The seeds contain various phenolic compounds that impart antioxidant properties, which may help prevent disease [
23].
Table 1.
Natural variation for oil content (%) and fatty acid composition (%) in sesame germplasm.
Table 1.
Natural variation for oil content (%) and fatty acid composition (%) in sesame germplasm.
| Accession |
Oil
Content
|
Protein |
Fatty acids |
Reference |
| |
16:0 |
18:0 |
18:1 |
18:2 |
18.3 |
22:1 |
22 cultivars, 4 landraces of S. mulayanum and
7 accessions of 4 wild species |
– |
– |
9–15 |
– |
37–52 |
30-52 |
– |
0–8 |
[18] |
| 4 Pakistani sesame varieties |
50–54 |
19–23 |
3–19 |
5–22 |
10 |
5–13 |
– |
16 |
[24] |
| Market sample |
48 |
|
5 |
13 |
43 |
36 |
– |
– |
[25] |
| Brown non roasted vs roasted sesame |
49 vs 51 |
– |
5.6 vs 7.1 |
5.3 vs 6.8 |
40.3 vs 58.7 |
46.1 vs 25 |
0.4 vs 0.2 |
– |
[26] |
| 103 sesame landraces |
41-63 |
– |
8-10 |
– |
29-41 |
41-49 |
– |
– |
[27] |
| Collection of 12 countries |
45-53 |
– |
5-6 |
8-10 |
36-44 |
39-46 |
0.3-04 |
|
[19] |
6 cultivated (C)
3 wild spp. (W) |
53-55 (C)
54-59 (W) |
– |
4-9 (C)
4-8 (W) |
4.5-10 (C)
4-5 (W) |
33-38 (C)
35-37 (W) |
42-52 (C)
43-46 (W) |
4-9.5 (C)
5-10 (W) |
– |
[9] |
| Indian sesame cultivar |
42-54 |
16-27 |
8-12 |
4-8 |
41.5-50 |
32-43 |
0.2-0.35 |
– |
[28] |
5. Sesame Wilting Disease
Sudden wilting in sesame is a common production constraint in hot and dry regions of the world. Initially, it was known that the disease was caused by the
Fusarium oxysporum [
38]. However, it has been observed that sudden wilting syndrome was caused by cumulative biotic and abiotic factors. In Pakistan, sudden wilting syndrome was one of the major production constraints due to the hot climate and caused 50-100% yield losses across the belt of central Punjab, Pakistan. Sudden wilting is observed in hot clime (40°C) after delayed irrigation, and plants experience sudden wilting within 24 hours. Infected plants contained infection of
Fusarium oxysporium and
Macrophomina phaseolina [
39]. However, many of the samples did not carry any of the two pathogens showing abiotic factors such as heat and drought stress injuries leading to the sesame wilt. The incidence of wilting increased with delayed irrigation [
39].
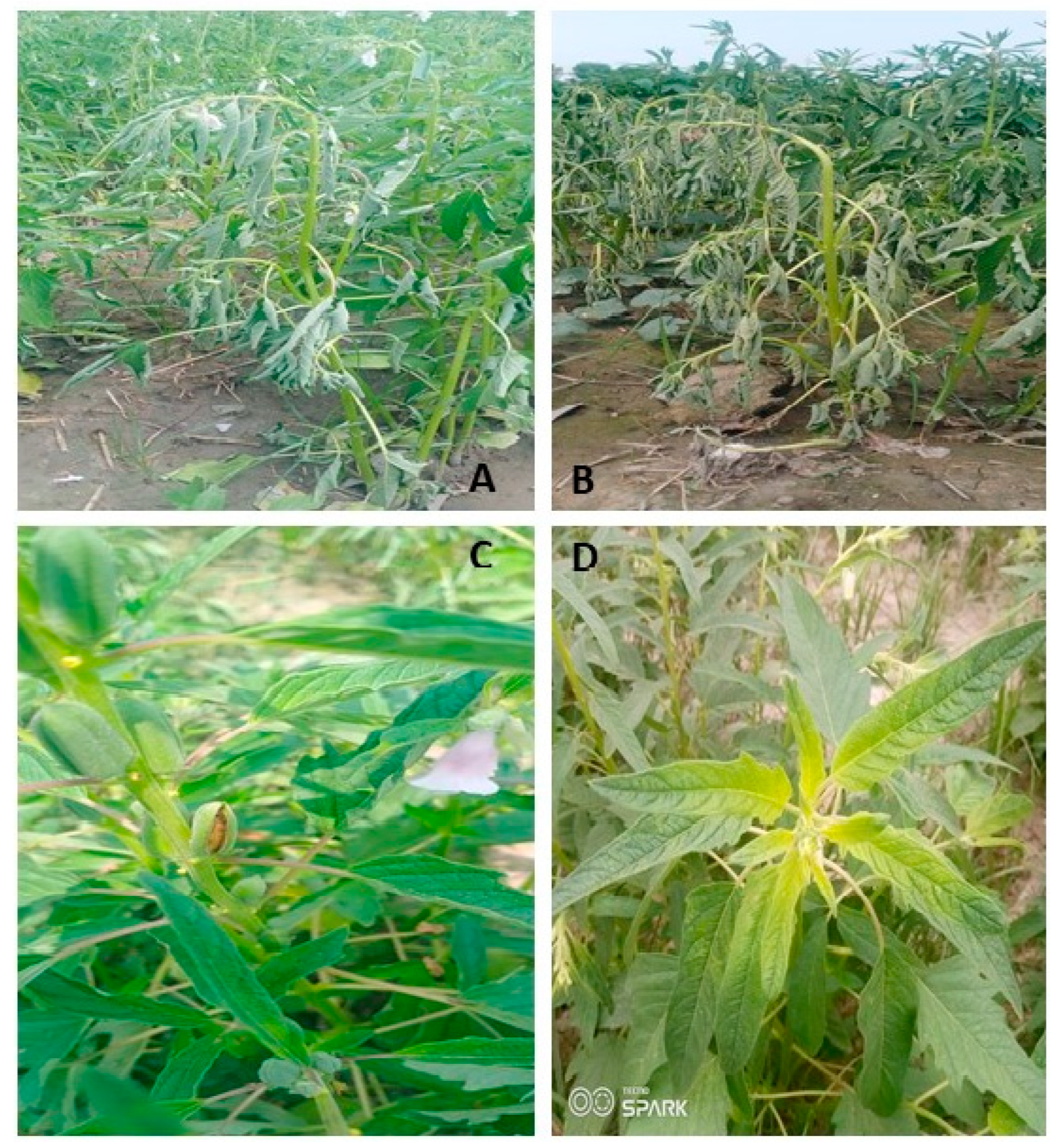
A significant increase in the browning of the stem in the wilted plants of sesame was observed after 5–6 days of rain (20 mm) in heavy soils of Shahkot, Pakistan indicative of charcoal rot infestation after wilting (
Figure 3). Resistance breeding may be required to cope with the sudden wilting in the sesame. Resistance breeding may require the development of varieties which may survive hypoxia conditions within roots after heavy rainfall or irrigation [
40]. Moreover, introgression of charcoal or root rot disease resistance may further reduce the chances of fungal infestation after the wiling. Combining ability analyses of the sesame breeding lines was done to determine the breeding value for charcoal rot resistance. Selectable variation for charcoal rot resistance was low to high and there were significant environmental and dominance effects associated with the charcoal rot resistance which indicate that recurrent selection and genotypic selection will be required to improve the charcoal rot resistance [
41]. The sesame genetic map was developed from 424 new polymorphic markers and 548 recombinant inbred lines [
42]. These markers detected 14 QTL which explained 3-14% of the total genetic variation contributing to the charcoal rot resistance (Wang et al. 2024).
qCRR12.
2 contributed the highest to charcoal rot resistance in all four environments and
qCRR8.
2 and
qCRR8.
3 were environmental specific. These QTL were used to predict the disease-resistance genes. Genome-wide association research may be required to tag genes related to the disease and breeding value for resistance breeding.
6. Abiotic Stresses Tolerance in Sesame
Sesame is a drought-tolerant crop with nitrogen recovery ability, making it suitable for arid climates with low rainfall and soil fertility [
43]. Climate change, characterized by increasing CO2 levels and temperature fluctuations, poses significant challenges for sesame production. While sesame thrives in high temperatures, it experiences reproductive failures when temperatures exceed 33ºC [
43]. Developing new cultivars with enhanced drought and heat tolerance, combined with high yield potential, is essential for sesame cultivation under changing climate conditions [
44].
Selection for high yield potential and stability has been effectively done through QTL mapping of economically important traits linked with yield, resistance against stresses, and stability (
Table 2). Molecular markers and QTL mapping have been used to select high-yielding and stress-resistant cultivars [
45]. Transcriptomic analysis of drought-tolerant sesame accessions has identified genes related to osmotic stress [
46]. The root profile of drought-tolerant sesame accessions showed differential expression of genes related to catalytic activity, ion binding, and transferase activity compared to susceptible accessions [
46]. The study, therefore, helps to identify the genes related to abiotic stresses such as osmotic stress.
Flowering time in sesame is influenced by day length and temperature, with the CONSTANS gene playing a key role [
53]. Understanding the genetic regulation of flowering and maturity can help develop cultivars suited to specific environments [
47]. Flowering in sesame is initiated in response to day length and temperature, and day length has profound effects on the flowering time in sesame [
53]. Sesame is a short-day plant, and flowering is initiated in response to day length. A gene called
CONSTANS (
CO) was identified, which dictates the vegetative growth period of sesame. A biometrical analysis of sesame flowering time across six generations revealed that two genes have additive, dominant, and epistatic effects [
53]. Days to flowering initiation were negatively related to traits such as main stem length, capsule number plant-1, and thousand seeds weight (TSW) [
54]. Accessions with later flowering had a small reproductive period.
Gene regulation of flowering initiation in sesame could help to understand the mechanism underlying flowering and maturity under a specific set of environments. The gene
CO has two homologues that have been found:
SiCOL1 and
SiCOL2.
SICOL2, lacking the B-box motif, delays flowering compared to genotypes with the
SiCOL1 sequence.
SiCOL1 is similar to the
CO gene, showing high expression in leaves before flowering and diurnal rhythmic expression in response to day length. It has 16 haplotypes within Asian genotypes, and the mutant type does not respond to day length, indicating it is non-functional or photoperiod-insensitive. Genotypes having a mutant allele of
SiCOL1 initiated early flowering under long days [
47]. Hence, genome sequencing and annotation may help identify differentially expressed genes and understand the adaptability of sesame genotypes to a particular environment.
7. Inducing New Variability within Sesame Germplasm
7.1. Mutation Breeding
Physical or chemical mutagens were used to expand genetic variability within the breeding population (
Table 3). Twenty-five mutants from 8 countries were released for general cultivation, out of those, five were released in China with improved agronomic and host plant resistance to pathogens [
55]. Exposure to sesame accessions with 0.5% or 1% ethyl methane sulfonate (EMS) induced novel genetic variability such as tetra or tri capsule per leaf axil, determinate growth habit, and seed colour [
56]. The optimum dose for inducing LD50 for EMS in sesame was between 0.34% and 0.44% [
57]. The optimum lethal dose for gamma radiation was 389–417 Gy [
58]. Advanced mutant lines (M5) were evaluated in replicated trials and selected as candidate lines (SM-06, SM-07, and SM-04), which yielded higher than control and novel traits such as earliness, and the highest capsule plant-1 was observed within the mutant population [
59]. Genetic divergence among the mutant lines showed that phenological traits such as the height of the first capsule and days to 50% flowering had the highest divergence [
60].
CRISPR/Cas9 has great potential to for improving crops through gene editing with the least detrimental effect on yield and components. With the availability of sequence information about key genes related to fatty acid and photoperiod insensitivity, CRISPR/Cas may be efficiently used to downregulate the gene related to linoleic acid to develop breeding lines with high oleic acid without yield drags. Moreover, genes related to shattering problems may also be downregulated, which helps induce shattering resistance in soybeans. Shattering resistance may also facilitate sesame for mechanized harvesting, apart from reducing yield losses. Sesame-specific adaptability may be improved by downregulating genes related to photoperiod, Photoperiod insensitivity helps to initiate flowering under long day lengths [
61].
7.2. Development of Semi-Dwarf Plants and Various Mutant Types
A semi-dwarf sesame genotype (
dw607) having reduced intermodal length and stem lodging resistance and a higher thousand seeds weight (TSW) was developed by Miao
et al., [
68]. The gene
Sidwf1 affects plant height, as determined by the molecular analysis of 824 germplasm accessions carrying 58 genomic variants.
Sidwf1-inducing semi-dwarf genotypes carried SNP mutation of nucleotide substitution from C to T, causing an amino acid change from proline to serine [
68]. An EMS-induced sesame mutant (
sc1) with few seed capsules and short capsules was due to a mutation of the
SICS1 allele [
7]. The
SICS1 contained a point mutation at intron 5 and exon 6 junctions, which caused a frame-shift of the reading frame, causing defective protein [
7]. Indeterminate growth habits in sesame were dominant over the determinate type [
69]. Crosses among the indeterminate and determinate types identified multiple recessive alleles of determinate growth, such as
dt1 and
dt2 [
69]. In a cross between long and dense capsules and short and sparse capsules, a F
1 showed dominance for a long and dense capsule. A segregating ratio of 12:3:1 was obtained, which showed that two genes had epistasis interactions [
70]. In another study, Liu
et al. [
71] developed a yellow-leaf dwarf (
yl1) mutant having lower carotenoid and chlorophyll contents than wild types. The yl1 mutant also had a longer growth duration with a slower photosynthesis rate, later flower initiation, and shorter plant height than the wild type [
71].
7.3. Indehiscence Capsule
Capsule shattering is a major impediment to the cultivation and mechanical harvesting of the sesame crop. Generally, the farmer harvests the crop with partial pod maturity and then threshes it to obtain the seeds. Crossing the indehiscent parent with the dehiscent parent results in an indehiscent F1 hybrid that is dominant to the dehiscence capsule [
72]. A segregation ratio of 3:1 was obtained, which shows a single recessive allele was controlling the indehiscent capsule [
72]. Genes controlling shattering or non-shattering type were mapped onto SNP marker S8_5062843 (78.9 cM) near the distal end of LG8 (chromosome 8) [
73]. The sequence of the capsule non-shattering allele carried a frameshift mutation, which caused a shift in the splicing and introduced SNP variation compared with the shattering alleles [
73]. To facilitate the selection of non-shattering types, CAPS markers were developed which differentiated both types in sesame [
53].
7.4. Male Sterility and Hybrid Vigour
The possibility of breeding hybrid cultivars of sesame has been documented. Many articles described heterosis in sesame for yield and components, indicating the possibility for commercial exploitation of hybrid vigour [74, 75]. Heterosis ranged between 9.5 and 32.7%, estimated from 1,636 crosses [
74]. However, commercial development of hybrid seed was only possible through exploitation of genetic or cytoplasmic male sterility. A list of various nuclear alleles identified or induced in sesame has been shown in
Table 4.
Cytoplasmic male sterility sources were identified through wide hybridization and species such as
S. malabaricum were identified as possible sources of cytoplasmic male sterility [
80]. Genetic male sterility was controlled by the single recessive nuclear male sterility alleles and generally maintained in heterozygous conditions. Spontaneous recessive male sterility was identified in the Chinese cultivar "Zhuzhi,” which was transferred and maintained through sib mating and crossing with male fertile plants [
81]. The developed male sterile line was named “D248A” [
81]. The cultivar “D248 A” had a small green anther which did not contain pollens. The absence of pollen occurred due to abnormality in the microspore mother cells and microsporocytes had less cytoplasm and anucleate [
78]. Moreover, microspore mother cells were not able to undergo normal meiotic division [
78]. Research showed that males had a segregating ratio of 1:3 in the F
2 generation. A dominant genetic male sterility allele was also identified in sesame [
77], which may be easier to maintain than recessive genetic male sterility within the inbred line [
78]. Genetic male sterility also requires visible markers to effectively select plants with genetic male sterility at the seedling stage [
82]. In sesame, genetic male sterility was linked with wrinkled leaves to differentiate male sterile plants from normal plants [
82].
Chemical restoration agents have been recommended for the maintenance of genetic male sterility in sesame [
83]. For example, the genotype CRA-02 showed the highest restoration of pollen fertility when sprayed at bud initiation stages and restored pollen fertility by 33% [
83]. Bulk segregant analyses followed by next-generation tools identified the putative regions of the 4 chromosomes related to pollen sterility in male sterile plants along with leaf shape [
82], SSRs and InDel markers further narrowed down the region associated with traits in the backcross population (BC1). The qPCR identified suppression of 4 genes in sterile pollen and 1 gene during leaf development [
82]. The cell wall invertase CWINV1 was involved in the supply of carbohydrates to the developing pollen and their suppression in anther caused male sterility [
84]. Gene
Sicwinv1 that induced male sterility in sesame was cloned to study the function of the allele which indicated its accumulation during the tetrad stage in the tapetum cells of the anther [
84].
8. Genetics of Economic Traits
Generally, yield and its components are affected by numerous QTL across the genome and their inheritance was further complicated by the interactions between loci, and environment. Yield components also show pleiotropic loci. Genome-wide association mapping resulted in the identification of major QTL related to yield and its components, such as thousand seed weight, seed size, and seed coat colour [
85]. They mapped major QTL for seed size and seed coat colour on LG04 and LG11 and these linkage groups contained 155 candidate genes for seed coat and size [
85]. Recent advancements in genomic technologies have significantly accelerated the identification of QTL in sesame. For instance, the genome-wide association study (GWAS) approach has enabled researchers to identify novel loci associated with complex traits. This approach has been instrumental in uncovering the genetic architecture of oil content and fatty acid composition in sesame seeds [
45].
Whole genome sequencing paved the way for high-density markers such as SNP or SSR which helped to map quantitative traits of sesame [85, 86]. For instance, a study in sesame identified 2,159 SNP markers on 13 linkage maps, the distance between the markers was about 0.99 cM [
85]. About 83,135 non-redundant SSR marker positions and motif sequences have been published and obtained from 151 published genomic sequences [
86]. Genetic regions containing clusters of genes in sesame were considered ‘hotspots’ for economic traits. In another study, a high-density linkage map of 19,309 markers was constructed which revealed 84 QTL associated with yield components and seed mineral components [
87]. Thirteen QTL on 7 LG and 17 QTL on 10 LG were identified for yield components identified through 1,230 markers in the recombinant inbred lines (RIL) population [
88]. Thus, developing high-throughput phenotyping technologies has complemented genomic advances, allowing for more precise and efficient evaluation of complex traits. For example, near-infrared spectroscopy (NIRS) enables rapid and non-destructive assessment of seed quality traits, facilitating the screening of large populations for breeding programs [
89].
A high-density map of specific locus amplified fragment (SLAF) comprised 3,528 markers with an average distance of 0.37 cM and the SLAF makers revealed 46 significant QTLs for 7 yield components across 4 environments [
90], where QTL were spread over 11 linkage groups and 23 stable QTL were detected in all environments and favourable QTL were found concentrated in the Chinese line Yuzhi4 [
90]. Furthermore, integrating multi-omics data, including genomics, transcriptomics, and metabolomics, has provided a more comprehensive understanding of sesame's molecular mechanisms underlying complex traits. This systems biology approach has revealed intricate gene regulatory networks involved in oil biosynthesis and stress responses [
20]. QTL for traits related to water stress condition i.e. root length and relative root length were identified on chromosome 12 of sesame [
91]. A density linkage map spanned over 12 chromosomes comprising 1,354 markers developed through a whole genome re-sequencing strategy [
91]. There was significant genetic variation for yield components such as capsule size, capsule number and seed size-related traits [
47]. They identified two novel candidate genes (
SiLPT3 and
SiACS8c) affecting capsule number and capsule length. A major QTL qLS15-1, stable across the environment located on LG15 was shown to affect leaf size and development [
92](Sheng et al., 2021). Sequencing of QTL and transcriptome analyses also revealed three candidate genes
SIN_1004875,
SIN_1004882, and
SIN_1004883 associated with leaf size [
92](Sheng et al., 2021). In another study, A high-density linkage map of 20,294 SNP markers also identified major QTL on LG2 for flowering date and yield components [
93](Sabag et al., 2021). Significant QTL identified a candidate gene
SIN_1019016 on chromosome 10 related to phytophthora blight resistance which explained up to 13.34% of phenotypic variation [
94]. The advent of genome editing technologies, particularly CRISPR-Cas9, has opened new avenues for targeted modification of sesame traits. While still in its early stages for sesame, this technology holds promise for precisely manipulating genes controlling key agronomic traits, potentially accelerating the breeding process [
95]. Hence, whole genome sequencing and genome-wide association studies paved the way for the identification of major QTL related to yield and its components in sesame.
9. Prospects for Sesame Improvement
Sesame is an oilseed crop with great potential to be ranked among the leading oilseed crop in the world. Prospects for sesame improvement lie in integrating conventional breeding with advanced molecular techniques such as marker-assisted selection, genomic selection, and gene editing. Recent advances in high-throughput phenotyping and genotyping technologies have significantly accelerated sesame improvement. For example, unmanned aerial vehicles (UAVs) equipped with multispectral sensors now allow for rapid assessment of large-scale field trials, enabling more efficient selection of superior genotypes that will expand the area of this crop for new germplasm with adaptability to diverse environmental conditions. On the other hand, sesame has low yield potential due to poor response to farm inputs. Sesame accessions having characteristics such as determinate growth, semi-dwarf stature, and lodging resistance are required to improve response to farm inputs and the same ideotype for mechanized harvesting. Moreover, accessions with indehiscence capsules should be bred to reduce yield losses during harvesting. Additionally, developing sesame varieties with synchronized maturity is another critical breeding objective, as it would significantly improve the efficiency of mechanical harvesting and reduce post-harvest losses. Some of the work has already been done and germplasm accessions with novel agronomic traits were developed through mutation breeding, selection or hybridization. The application of CRISPR-Cas9 technology holds promise for targeted gene editing to develop high-yielding, pest-resistant, and climate-resilient sesame varieties. For example, CRISPR-Cas9 could modify genes involved in lignin biosynthesis, potentially developing cultivars with improved oil content and easier oil extraction. Further, expanding transcriptomic and proteomic studies will enhance our understanding of the regulatory mechanisms controlling important agronomic traits. Hence, integrating omics approaches with metabolomics data can offer a comprehensive view of the molecular pathways underlying sesame's nutritional and medicinal properties, potentially opening new avenues for value-added product development. However, sesame cultivation is potentially grown in developing or resource-poor countries which may have seldom research priorities for advanced molecular techniques. Hence, international collaborative efforts among institutes, researchers, breeders, and policymakers are essential to address sesame cultivation's challenges and exploit its full potential as a sustainable oilseed crop. Climate-smart sesame cultivars must be developed to withstand extreme weather events and utilize resources more efficiently to ensure the crop's sustainability in the face of climate change. Thus, this requires a multidisciplinary approach that combines traditional breeding techniques, genomic tools, and advanced agronomic practices.
10. Conclusion
In conclusion, sesame provides valuable opportunities for oil production and medicinal uses. However, its cultivation is hindered by several factors, like low yields, poor input response, and susceptibility to pests and diseases. Recent advances in genomics and molecular breeding provide new ways to overcome these challenges and improve the productivity and profitability of sesame farming. Researchers and breeders can develop superior sesame varieties by utilizing the extensive genetic diversity found in sesame germplasm and employing advanced breeding techniques. These cultivars are designed to meet growers' and consumers' demands worldwide, enhancing the crop's utility and market potential. Thus, this comprehensive approach will promote the sustainable growth of the sesame industry, making it a more resilient and valuable crop for future generations.
Author Contributions
Conceptualization, S.R and T.B; Software: M.N, S.R and T.B; Data analysis: all coauthors; Writing–original draft: M.N, S.R, T.B, M.E, A.G., M.R. and R.O.; Writing – review and editing: all coauthors; funding acquisition: R.O. All authors have read and agreed to the published version of the manuscript.
Funding
A.G. and R.O. acknowledges grant funding CTS 23:2654 for the project “Fine mapping of plant height and seed coat color quantitative trait loci of sesame (Sesamum indicum L.) using single nucleotide polymorphisms”.
Data Availability Statement
Not applicable.
Acknowledgments
We would like to thank the Department of Plant Breeding and Genetics, University of Agriculture, Faisalabad, Pakistan and the Swedish University of Agricultural Sciences, for the time and covering APC of the first author of this article. from the Carl Trigger Foundation.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Onkware, A., et al., Genetic relationship between sesame (Sesamum indicum L.) and related wild species based on chromosome counts and isozyme markers. 2015.
- Elsafy, M., Status of sesame breeding. Introductory paper at the Faculty of Landscape Architecture, Horticulture and Crop Production Science, 2023 (2023: 2).
- Canton, H., Food and Agriculture Organization of the United Nations—FAO, in The Europa directory of international organizations 2021. 2021, Routledge. p. 297-305.
- FAO, Food and Agricultural Organization. Italy, Rome, 2022.
- Wacal, C., et al., Seed yield, crude protein and mineral nutrient contents of sesame during a two-year continuous cropping on upland field converted from a paddy. Field Crops Research, 2019. 240: p. 125-133. [CrossRef]
- Singh, A.K., et al., Plant Genetic Resources Management in India: A Historical Perspective.
- Wei, X., et al., SesameFG: an integrated database for the functional genomics of sesame. Scientific Reports, 2017. 7(1): p. 2342. [CrossRef]
- Bedigian, D., Slimy leaves and oily seeds: distribution and use of wild relatives of sesame in Africa. Economic Botany, 2004. 58(1): p. S3-S33. [CrossRef]
- Azeez, M.A. and J.A. Morakinyo, Genetic diversity of fatty acids in sesame and its relatives in Nigeria. European Journal of Lipid Science and Technology, 2011. 113(2): p. 238-244. [CrossRef]
- Bedigian, D., Characterization of sesame (Sesamum indicum L.) germplasm: a critique. Genetic Resources and Crop Evolution, 2010. 57: p. 641-647. [CrossRef]
- Yang MinMin, Y.M., et al., Production and identification of F1 interspecific hybrid between Sesamum indicum and wild relative S. indicatum. 2017.
- Stavridou, E., et al., Characterization of the genetic diversity present in a diverse sesame landrace collection based on phenotypic traits and EST-SSR markers coupled with an HRM analysis. Plants, 2021. 10(4): p. 656. [CrossRef]
- Wei, X., et al., Development of simple sequence repeat (SSR) markers of sesame (Sesamum indicum) from a genome survey. Molecules, 2014. 19(4): p. 5150-5162. [CrossRef]
- Nyongesa, B.O., et al., Genetic diversity in cultivated sesame (Sesamum indicum L.) and related wild species in East Africa. Journal of Crop Science and Biotechnology, 2013. 16: p. 9-15. [CrossRef]
- Wang, M., et al., Improved assembly and annotation of the sesame genome. DNA Research, 2022. 29(6): p. dsac041. [CrossRef]
- Purru, S., et al., GinMicrosatDb: a genome-wide microsatellite markers database for sesame (Sesamum indicum L.). Physiology and Molecular Biology of Plants, 2018. 24: p. 929-937. [CrossRef]
- Dutta, D., V.K. Awon, and G. Gangopadhyay, Transcriptomic dataset of cultivated (Sesamum indicum), wild (S. mulayanum), and interspecific hybrid sesame in response to induced Macrophomina phaseolina infection. Data in Brief, 2020. 33: p. 106448.
- Mondal, N., et al., Effects of domestication bottleneck and selection on fatty acid desaturases in Indian sesame germplasm. Plant Genetic Resources, 2016. 14(2): p. 81-90. [CrossRef]
- Kurt, C., Variation in oil content and fatty acid composition of sesame accessions from different origins. Grasas y aceites, 2018. 69(1): p. e241-e241. [CrossRef]
- Zhang, Y.-P., et al., Integration of miRNAs, degradome, and transcriptome omics uncovers a complex regulatory network and provides insights into lipid and fatty acid synthesis during sesame seed development. Frontiers in Plant Science, 2021. 12: p. 709197.
- Kancharla, P.K. and N. Arumugam, Variation of oil, sesamin, and sesamolin content in the germplasm of the ancient oilseed crop Sesamum indicum L. Journal of the American Oil Chemists' Society, 2020. 97(5): p. 475-483.
- Mohammed, F., et al., Chemical composition and mineralogical residence of sesame oil from plants grown in different Yemeni environments. Microchemical Journal, 2018. 140: p. 269-277. [CrossRef]
- Zeb, A., B. Muhammad, and F. Ullah, Characterization of sesame (Sesamum indicum L.) seed oil from Pakistan for phenolic composition, quality characteristics and potential beneficial properties. Journal of Food Measurement and Characterization, 2017. 11(3): p. 1362-1369. [CrossRef]
- Asghar, A. and M.N. Majeed, Chemical characterization and fatty acid profile of different sesame verities in Pakistan. Am J Sci Ind Res, 2013. 4(6): p. 540-545.
- Carvalho, R., et al., Extraction, fatty acid profile and antioxidant activity of sesame extract (Sesamum Indicum L.). Brazilian Journal of Chemical Engineering, 2012. 29: p. 409-420. [CrossRef]
- Hama, J.R., Comparison of fatty acid profile changes between unroasted and roasted brown sesame (Sesamum indicum L.) seeds oil. International Journal of Food Properties, 2017. 20(5): p. 957-967. [CrossRef]
- Uzun, B., Ç. Arslan, and Ş. Furat, Variation in fatty acid compositions, oil content and oil yield in a germplasm collection of sesame (Sesamum indicum L.). Journal of the American Oil Chemists' Society, 2008. 85(12): p. 1135-1142.
- Awasthi, C., et al., Biochemical composition and fatty acid profile of some promising sesame (Sesamum indicum L.) genotypes. Indian Journal of Agricultural Biochemistry, 2006. 19(2): p. 67-70.
- Santos, H.O., et al., Small dense low-density lipoprotein-cholesterol (sdLDL-C): analysis, effects on cardiovascular endpoints and dietary strategies. Progress in cardiovascular diseases, 2020. 63(4): p. 503-509.
- Rauf, S., et al., Progress in modification of sunflower oil to expand its industrial value. Journal of the Science of Food and Agriculture, 2017. 97(7): p. 1997-2006. [CrossRef]
- Huang, H., et al., Modifications of fatty acid profile through targeted mutation at BnaFAD2 gene with CRISPR/Cas9-mediated gene editing in Brassica napus. Theoretical and Applied Genetics, 2020. 133: p. 2401-2411. [CrossRef]
- Lee, K.-R., et al., Increasing monounsaturated fatty acid contents in hexaploid Camelina sativa seed oil by FAD2 gene knockout using CRISPR-Cas9. Frontiers in Plant Science, 2021. 12: p. 702930. [CrossRef]
- Salas, J.J., et al., High stearic sunflower oil: Latest advances and applications. OCL, 2021. 28: p. 35. [CrossRef]
- Dar, A.A., et al., The FAD2 gene in plants: occurrence, regulation, and role. Frontiers in plant science, 2017. 8: p. 1789. [CrossRef]
- Jin, U.-H., et al., Characterization and temporal expression of a ω-6 fatty acid desaturase cDNA from sesame (Sesamum indicum L.) seeds. Plant science, 2001. 161(5): p. 935-941. [CrossRef]
- Chen, Z., et al., Variation in seed fatty acid composition and sequence divergence in the FAD2 gene coding region between wild and cultivated sesame. Journal of Agricultural and Food Chemistry, 2014. 62(48): p. 11706-11710. [CrossRef]
- Kim, M.J., et al., Seed-specific expression of sesame microsomal oleic acid desaturase is controlled by combinatorial properties between negative cis-regulatory elements in the SeFAD2 promoter and enhancers in the 5′-UTR intron. Molecular Genetics and Genomics, 2006. 276: p. 351-368. [CrossRef]
- Kavak, H. and E. Boydak, Trends of sudden wilt syndrome in sesame plots irrigated with delayed intervals. Afr. J. Microbiol. Res, 2011. 5: p. 1837-1841.
- Kavak, H. and E. Boydak, Screening of the resistance levels of 26 sesame breeding lines to Fusarium wilt disease. Plant Pathology Journal, 2006.
- Wang, L., et al., Global gene expression responses to waterlogging in roots of sesame (Sesamum indicum L.). Acta physiologiae plantarum, 2012. 34: p. 2241-2249. [CrossRef]
- Mahmoud, S.A., E.S. El-Sharkawy, and M. Emam, Breeding sesame for resistance to charcoal rot caused by Macrophomina phaseolina. SVU-International Journal of Agricultural Sciences, 2024. 6(2): p. 18-35. [CrossRef]
- Wang, L., et al., Development of an SSR-based genetic map in sesame and identification of quantitative trait loci associated with charcoal rot resistance. Scientific Reports, 2017. 7(1): p. 8349. [CrossRef]
- Baath, G.S., et al., Quantifying and modeling the influence of temperature on growth and reproductive development of sesame. Journal of Plant Growth Regulation, 2022: p. 1-10. [CrossRef]
- Kim, M.-S., et al., Changes in cuticular waxes of developing leaves in sesame (Sesamum indicum L.). Journal of Crop Science and Biotechnology, 2009. 12: p. 161-167. [CrossRef]
- Zhou, R., et al., Genome-wide association studies of 39 seed yield-related traits in sesame (Sesamum indicum L.). International Journal of Molecular Sciences, 2018. 19(9): p. 2794. [CrossRef]
- Song, Q., et al., Comparative analysis of root transcriptome profiles of sesame (Sesamum indicum L.) in response to osmotic stress. Journal of Plant Growth Regulation, 2021. 40(4): p. 1787-1801. [CrossRef]
- Zhou, R., et al., Photoperiod response-related gene SiCOL1 contributes to flowering in sesame. BMC Plant Biology, 2018. 18: p. 1-16. [CrossRef]
- Li, D., et al., Genome-wide analysis of WRKY gene family in the sesame genome and identification of the WRKY genes involved in responses to abiotic stresses. BMC Plant Biology, 2017. 17: p. 1-19. [CrossRef]
- Wang, L., et al., High-resolution temporal transcriptome sequencing unravels ERF and WRKY as the master players in the regulatory networks underlying sesame responses to waterlogging and recovery. Genomics, 2021. 113(1): p. 276-290. [CrossRef]
- Wang, Y., et al., Identification and characterization of the bZIP transcription factor family and its expression in response to abiotic stresses in sesame. PLoS One, 2018. 13(7): p. e0200850. [CrossRef]
- Chowdhury, S., A. Basu, and S. Kundu, Overexpression of a new osmotin-like protein gene (SindOLP) confers tolerance against biotic and abiotic stresses in sesame. Frontiers in Plant Science, 2017. 8: p. 410. [CrossRef]
- Dossa, K., et al., Transcriptomic, biochemical and physio-anatomical investigations shed more light on responses to drought stress in two contrasting sesame genotypes. Scientific Reports, 2017. 7(1): p. 8755. [CrossRef]
- Zhang TiDe, Z.T., et al., Genetic analysis of flowering time with the mixed major gene plus polygene inheritance model in sesame. 2019.
- Shim, K.B., et al., Effect of temperature and daylength on flowering and growth characteristics. KOREAN JOURNAL OF CROP SCIENCE, 2020. 65(3): p. 241-247.
- YU, M., et al., Research advances on induced mutation breeding of sesame. Biotechnology Bulletin, 2017. 33(11): p. 8.
- Kouighat, M., et al., Novel genetic variability in sesame induced via ethyl methane sulfonate. Journal of Crop Improvement, 2021. 35(5): p. 654-665. [CrossRef]
- Sandhiya, V., et al., Determination of optimum dose of chemical mutagen for large scale seed treatment of white seeded sesame (Sesamum indicum L.) varieties. Electronic Journal of Plant Breeding, 2020. 11(1): p. 238-242.
- Parthasarathi, G., et al., Optimal lethal dose determination for gamma rays and EMS induced mutagenesis in TMV7 and SVPR1 Sesame (Sesamum indicum L.) varieties. Current Journal of Applied Science and Technology, 2020. 39(28): p. 136-144. [CrossRef]
- Bhuiyan, M.S.H., et al., Genetic variance and performance of sesame mutants for yield contributing characters. Malaysian Journal of Sustainable Agriculture, 2019. 3(2): p. 27-30.
- Patil, M.K., R. Lokesha, and J. Diwan, Genetic divergence of advanced mutant breeding lines in sesame (Sesamum indicum L.) assessed through D2 statistics. International Journal of Current Microbiology and Applied Science, 2018. 6(9): p. 3133-3139.
- Ashri, A., Sesamum Indicum, in Handbook of Flowering. 2019, CRC Press. p. 309-312.
- Weldemichael, M.Y., et al., Effect of sodium azide on quantitative and qualitative stem traits in the M2 generation of Ethiopian sesame (Sesamum indicum L.) genotypes. The Scientific World Journal, 2021. 2021(1): p. 6660711. [CrossRef]
- Jayaramachandran, M., et al., Genetic improvement of a neglected and underutilised oilseed crop: sesame (Sesamum indicum L.) through mutation breeding. The Nucleus, 2020. 63: p. 293-302.
- Aristya, V.E., T. Taryono, and R.A. Wulandari, Yield Components of Some Sesame Mutant Populations Induced by Gamma Irradiation. Bul. Tanam. Tembakau Serat Miny. Ind, 2019. 10(64): p. 2018.64-71. [CrossRef]
- Kumari, V., et al., Identification of Phytophthora blight resistant mutants through induced mutagenesis in sesame (Sesamum indicum L.). Indian Phytopathology, 2019. 72: p. 71-77. [CrossRef]
- Ravichandran, V. and S. Jayakumar, Effect of mutagens on quantitative characters in M2 and M3 generation of sesame (Sesamum indicum L.). International Letters of Natural Sciences, 2015(42).
- Gadri, Y., L.E. Williams, and Z. Peleg, Tradeoffs between yield components promote crop stability in sesame. Plant Science, 2020. 295: p. 110105. [CrossRef]
- Miao, H., et al., Identification of a Sidwf1 gene controlling short internode length trait in the sesame dwarf mutant dw607. Theoretical and Applied Genetics, 2020. 133: p. 73-86.
- Cheng, J.-Z., et al., Computer-aided diagnosis with deep learning architecture: applications to breast lesions in US images and pulmonary nodules in CT scans. Scientific Reports, 2016. 6(1): p. 24454. [CrossRef]
- Yol, E., Inheritance of long and dense capsule characteristics in sesame. Turkish Journal of Field Crops, 2017. 22(1): p. 8-13. [CrossRef]
- Liu HongYan, L.H., et al., Anatomical structure and photosynthetic characteristics of a yellow leaf mutant YL1 in sesame (Sesamum indicum L.). 2017.
- Yol, E. and B. Uzun, Inheritance of indehiscent capsule character, heritability and genetic advance analyses in the segregation generations of dehiscent x indehiscent capsules in sesame. Journal of Agricultural Sciences, 2019. 25(1): p. 79-85.
- Zhang, H., et al., Identification of a SiCL1 gene controlling leaf curling and capsule indehiscence in sesame via cross-population association mapping and genomic variants screening. BMC Plant Biology, 2018. 18: p. 1-12. [CrossRef]
- Duhoon, S. Exploitation of heterosis for raising productivity in sesame. in 4th International Crop Science Congress, Brisbane, Australia. 2004.
- Jeeva, G., K. Saravanan, and C. Sowmiya, Assessment of combining ability and standard heterosis through diallel analysis in sesame (Sesamum indicum L.). Electronic Journal of Plant Breeding, 2020. 11(02): p. 386-391.
- Zhao, Y., et al., Characterization and genetic mapping of a novel recessive genic male sterile gene in sesame (Sesamum indicum L.). Molecular breeding, 2013. 32: p. 901-908. [CrossRef]
- Liu, H., et al., Inheritance and molecular mapping of a novel dominant genic male-sterile gene in Sesamum indicum L. Molecular Breeding, 2015. 35: p. 1-14. [CrossRef]
- Liu, H., et al., Cytological characterization and molecular mapping of a novel recessive genic male sterility in sesame (Sesamum indicum L.). Plos One, 2018. 13(9): p. e0204034. [CrossRef]
- Wu, K., et al., Histological and transcriptional characterization of a novel recessive genic male sterility mutant in sesame (Sesamum indicum L.). Acta physiologiae plantarum, 2014. 36: p. 421-431. [CrossRef]
- Prabakaran, A., S.S. Rangasamy, and R. Ramalingam, Identification of cytoplasm-induced male sterility in sesame through wide hybridization. Current Science, 1995: p. 1044-1047.
- Liu, H., et al., Development, inheritance and breeding potential of a recessive genic male sterile line D248A in Sesame (Sesamum indicum L.). SpringerPlus, 2013. 2: p. 1-7. [CrossRef]
- Liu, H., et al., Fine mapping of a novel male-sterile mutant showing wrinkled-leaf in sesame by BSA-Seq technology. Industrial Crops and Products, 2020. 156: p. 112862. [CrossRef]
- Zheng YongZhan, Z.Y., et al., An analysis of chemical maintaining effect on genic male sterility in sesame (Sesamum indicum L.) I. The effect of CRA in restoring fertility for genic male sterility in sesame. 2000.
- Zhou, T., et al., Sicwinv1, a cell wall invertase from sesame, is involved in anther development. Journal of Plant Growth Regulation, 2019. 38: p. 1274-1286.
- Du, H., et al., A high-density genetic map constructed using specific length amplified fragment (SLAF) sequencing and QTL mapping of seed-related traits in sesame (Sesamum indicum L.). BMC Plant Biology, 2019. 19: p. 1-20. [CrossRef]
- Dossa, K., A physical map of important QTLs, functional markers and genes available for sesame breeding programs. Physiology and Molecular Biology of Plants, 2016. 22: p. 613-619. [CrossRef]
- Teboul, N., et al., Genetic architecture underpinning yield components and seed mineral–nutrients in sesame. Genes, 2020. 11(10): p. 1221. [CrossRef]
- Wu, K., et al., High-density genetic map construction and QTLs analysis of grain yield-related traits in Sesame (Sesamum indicum L.) based on RAD-Seq technology. BMC Plant Biology, 2014. 14: p. 1-14. [CrossRef]
- Rife, T.W., et al., Prospector: A mobile application for portable, high-throughput near-infrared spectroscopy phenotyping. The Plant Phenome Journal, 2021. 4(1): p. e20024.
- Mei, H., et al., QTL mapping of yield-related traits in sesame. Molecular Breeding, 2021. 41(7): p. 43. [CrossRef]
- Liang, J., et al., QTL mapping of PEG-induced drought tolerance at the early seedling stage in sesame using whole genome re-sequencing. PLoS One, 2021. 16(2): p. e0247681. [CrossRef]
- Sheng, C., et al., QTL-seq and transcriptome analysis disclose major QTL and candidate genes controlling leaf size in Sesame (Sesamum indicum L.). Frontiers in Plant Science, 2021. 12: p. 580846. [CrossRef]
- Sabag, I., G. Morota, and Z. Peleg, Genome-wide association uncovers the genetic architecture of tradeoff between flowering date and yield components in sesame. bioRxiv, 2021: p. 2021.04. 22.440889. [CrossRef]
- Asekova, S., et al., An integrated approach of QTL mapping and genome-wide association analysis identifies candidate genes for phytophthora blight resistance in sesame (Sesamum indicum L.). Frontiers in Plant Science, 2021. 12: p. 604709. [CrossRef]
- You, J., et al., CRISPR/Cas9-mediated efficient targeted mutagenesis in sesame (Sesamum indicum L.). Frontiers in Plant Science, 2022. 13: p. 935825. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).