1. Introduction
Foodborne diseases, as defined by the World Health Organization (WHO), are illnesses caused by consuming food contaminated with harmful microorganisms and their toxins [
1]. These diseases can lead to a range of severe symptoms, including organ failure, digestive issues, and immune system and neurological complications [
2]. Contaminated food poses a significant threat to both food safety and human health. Pathogens such as
Listeria (L.) monocytogenes,
Salmonella spp.,
Staphylococcus (S.) aureus and
Clostridium (C.) botulinum are frequently associated with outbreaks linked to cheese consumption, even after the cheese has been ripened [
3]. Monitoring of milk and cheese production is crucial to detect and prevent the presence of these pathogens. For instance, although high incidence of
S. aureus has been found in sheep (39.4%) and goat (35.2%) raw milk, it appears as more prevalent in the final formulation of cheeses made from goat milk (9.83%) than in those from sheep milk (2.94%) [
4].
Fermented milk products, such as cheese, have a long history of preservation and culinary significance. In 2022, the European Union produced 160 million tons (m.t.) of raw milk, predominantly sourced from bovine species (154.3 m.t.) but also including ovine (3 m.t.), caprine (2.5 m.t.) and bubaline (0.3 m.t.) origins [
5]. A portion of this raw milk was converted into cheese (10.4 m.t. of total production), with artisanal cheese production thriving particularly in the Mediterranean regions [
6,
7]. However, artisanal cheese-making operations face challenges related to microbial contamination. To address this issue, there is a growing trend toward employing natural lactic acid bacteria (LAB) cultures to mitigate contamination risks and ensure the microbial safety and quality of cheese [
7,
8].
LAB constitute a specific taxonomic group of microorganisms characterized by their Gram-positive nature, lack of catalase enzyme, and characterized by their limited mobility [
9]. Autochthonous LAB in milk have been commonly sought, owing to their technological and probiotic potential, since traditional cheeses made from raw milk provide rich flavor and diversity of LAB strains [
10]. Artisanal cheeses, particularly those made from raw milk, are rich sources of microbial diversity, offering opportunities for the discovery of new LAB strains. However, ensuring the safety of raw milk cheese relies on various factors, including the type of cheese, initial microbial contamination, and the presence of harmful pathogens [
11].
During fermentation, LAB produce substances that stop harmful bacteria from growing in food. The main mechanism to achieve this is by producing lactic acid, which lowers the pH and negatively influences bacteria survival [
12]. LAB also release other compounds such as hydrogen peroxide, carbon dioxide, diacetyl, and bacteriocins, which further prevent bacterial growth [
10]. Traditional cheeses made from raw milk, without added starter cultures, harbor indigenous LAB strains with protective properties against pathogenic bacteria, which makes them better suited for cheese-making environments than commercial starters [
13]. Moreover, using LAB for natural food preservation meets the demand for healthier food with fewer chemical additives and contributes to expanding the range of functional dairy products [
14].
Additionally, given the association of milk and its derivatives with various pathogenic microorganisms, improving the microbiological quality of cheeses is imperative, warranting intensified efforts to ensure the safety of these products. Research on bacteriocins produced by dairy strains holds promise for enhancing food safety and expanding the types of healthy dairy products [
15].
This study aimed to (1) perform a systematic review and meta-analysis of the in vitro susceptibility of foodborne pathogens, namely, L. monocytogenes, Salmonella spp., and S. aureus, to autochthonous LAB species isolated from dairy products; and (2) to assess the relative importance of different moderators in the LAB’s supernatant antimicrobial activity. To achieve this, separate multilevel meta-analyses were conducted using up-to-date information extracted from relevant literature sources.
2. Materials and Methods
2.1. Articles Search
Electronic searches were conducted in PubMed, Scopus and Web of Science databases to gather articles summarizing results from in vitro assays carried out with autochthonous dairy LAB and their antimicrobial effects against the foodborne pathogens: L. monocytogenes, S. enterica, and S. aureus. The search was carried out by two reviewers on November 12th, 2022. The abstract bibliographic search was defined by terms encompassing cheese, starter culture and antimicrobial properties against specific pathogens as following:
(((cheese OR Milk OR Dairy) AND (lactic acid bacteria)) OR (starter culture) OR biocontrol) OR (lactobacillus)) AND (antimicrobial**)) OR (functional)) OR (preserv*)) OR (biopreserv*)) OR (antagoni*)) AND (Salmonella)) OR (Listeria monocytogenes)) OR (Staphylococcus aureus)) OR (pathogen*).
The searches were carried out in the specific syntax of PubMed, Scopus and Web of Science.
2.2. Data Extraction
Duplicate references were identified with the Rayyan software [
16]; reviewers manually verified and removed duplicates. Subsequently, individual reviewers examined the titles and abstracts of the citations retrieved. The title and abstract evaluation adhered to specific inclusion criteria: (i) the article must be written in English or Portuguese; (ii) the article must be a primary study, this is, must present original results; (iii) studies should present results of inhibition diameter of LAB against at least one of the selected pathogens
; (iv) articles must specify the origin of LAB, and it should be of dairy origin; and (v) studies must have LAB identified at species level
In the second step, the full text of selected articles were assessed to confirm their inclusion, considering any supplementary material if available. If the full text could not be retrieved, the corresponding author was contacted to request access [
17]. Additional inclusion criteria for the full-text screening phase included the reporting of inhibition diameter or radius in continuous numeric values. Furthermore, the article had to provide the sample size or number of repetitions (N).
Next, data extraction was performed from the selected articles using Excel [
18]. The extracted information encompassed details regarding the origin of LAB (milk source [cow, goat, sheep, NA], food type [milk, cheese, yogurt]), identification of LAB and pathogens (genus, species and strain), parameters from the susceptibility test method [agar spot, disc diffusion, well diffusion], duration of incubation, temperature of incubation, agar medium [MH, MRS, Nutrient, TSA, BHI], mode of LAB application (bacteriocin or supernatant) and variations in protocol: concentration of pathogen, concentration of LAB, aliquot volume, pH of LAB (bacteriocin or supernatant).
2.3. Meta-Regression Modeling
The meta-analysis considered the LAB evaluated for their antimicrobial activity as the study population. The primary measured outcome was the mean ID in millimeters, which refers to the total halo formed around the colony including the measurement of the colony, well or disk in the final ID calculation [
19]. If the articles presented the results in radius, the values were converted to diameter.
Two types of multilevel random-effects meta-analysis regressions were fitted to data in order to answer to different questions: the overarching meta-regression and the pathogen-specific meta-regressions.
2.3.1. Overarching Meta-Regression Models
The meta-regression model of the type
was adjusted to the full data, where
β0 is an intercept, and
β1,
β2 and
β3 are sets of fixed effects of the
p types of pathogens, the
g types of LAB genus, and the
a types of antagonism assay methods. The term
β4 is the effect of a one log increase in pathogen concentration (log CFU/ml);
β5 the effect of the incubation time (h) and
β6 the effect of the aliquot volume (μL). The error term
εpgaj represents the residuals and accounts for the variability between studies
j. The remaining unexplained variability was extracted by placing random effects
uj due to study
j in the intercept
β0.
The purpose of this model was to compare the extent of susceptibility to LAB supernatant among the three pathogens. Next, removing the pathogen effect β1 from Equation (1), the same model was fitted to data subsets partitioned by pathogen. The purpose of adjusting these three other models was to obtain meta-analytical estimates of ID (pooled ID) by LAB genus and susceptibility method, solved for a pathogen’s concentration of 7.0 log CFU/ml and an incubation time of 24 h. In this way, the pooled ID are aligned to the same conditions, and comparisons between pooled estimates can now be made for being more accurate.
2.3.2. Pathogen-Specific Meta-Regression Models
Three separate meta-regression models were adjusted by pathogen. Data were then partitioned by
L. monocytogenes,
Salmonella spp., and
S. aureus and the following models were adjusted, respectively,
Where the coefficients can be interpreted as exposed beforehand; and, in addition, in Equation (3),
β2 is the set of fixed effects for the types of agar. It can be observed that for each pathogen, a different meta-regression solution was achieved. This occurs because the partitions have different data structure, which is very usual in meta-analytical data sets; and, moreover, models are data-driven. For the three pathogen-specific models, only the significant terms were kept. Non-significant terms at α=0.10 were dropped. The purpose of the pathogen-specific meta-regressions was to assess the effect of moderators, and therefore deduce which moderators are more determinant in observing changes in the measured ID. To further understand the effect of certain quantitative moderators, bubble plots were generated. Meta-analysis models and graphs were built in R studio [
20] using the metafor package [
21].
2.4. Assessment of Heterogeneity and Publication Bias
Assessment of heterogeneity and publication bias were undertaken for each of the three pathogen-specific meta-regressions. First of all, from the null model (intercept only), the within-study variability (s²), and between-study variability (τ²) were obtained. The intra-class correlation (I
2) of the null model was then calculated as,
The I
2 statistic which measures the percentage of variation due to between-study heterogeneity, with values below 25% indicating low heterogeneity, between 25 and 50% moderate, and above 50% high heterogeneity. As rule of thumb, if I
2 > 25%, there should be an attempt to explain the variability between studies by incorporating significant moderators (also known as study characteristics). After fitting the full (final) model, the residual between-study variability (τ²
res) is obtained. Next, the between-study variability explained by significant moderators (R
2) can be estimated as,
Publication bias was ascertained using two methods: (1) by adding the sample size N as a moderator to each of the pathogen-specific models (Equations 2, 3 and 4), and applying the decision rule that publication bias is likely if p < 0.05; (2) by constructing funnel plots to visually identify potential publication bias and further evaluate the heterogeneity within a meta-regression, based on the spread of standard errors among individual outcomes [
24].
3. Results and Discussion
A total of 1665 articles were initially retrieved from the databases. Following the removal of duplicates and abstract screening, two-hundred and eighty-four articles remained. Subsequently, seventy-seven articles were included after the full-text screening phase. Of these, thirty-nine primary studies met the quality inclusion criteria, presenting sufficient data in extractable form. This resulted in 510 observations to be analyzed. The meta-analyses for each subset such as
L. monocytogenes were based on 24 primary studies with 220 observations [
25,
26,
27,
28,
29,
30,
31,
32,
33,
34,
35,
36,
37,
38,
39,
40,
41,
42,
43,
44,
45,
46,
47,
48],
Salmonella spp. based on 14 primary studies with 100 observations [
27,
31,
33,
34,
35,
36,
38,
49,
50,
51,
52,
53,
54,
55] and
S. aureus based on 25 studies with 190 observations [
25,
26,
27,
29,
30,
31,
33,
35,
36,
38,
39,
44,
49,
50,
51,
52,
53,
56,
57,
58,
59,
60,
61,
62].
The primary LAB found in starter cultures for producing fermented dairy products primarily comprise
Streptococcus, Lactobacillus, Bifidobacterium, Lactococcus and
Leuconostoc species [
63]. In this study, the main genus of LAB obtained was
Lactobacillus (n=310); although genus such as
Pediococcus (n=30),
Enterococcus (n=104),
Lactococcus (n=34),
Lacticaseibacillus (n = 19) and
Leuconostoc (n=13) were found. In terms of the types of data analyzed, there were no significant differences in ID observed between supernatant (n=476) and bacteriocin (n=37), leading to the decision to combine the data for analysis (results not shown).
In
Table 1, the outcomes of the overarching meta-regression models are compiled. First of all, these results demonstrate that, overall,
L. monocytogenes present greater susceptibility to LAB supernatant (i.e., higher inhibition values) compared to
Salmonella spp., and
S. aureus, though there were no differences between the latter two.
Table 1 separates the pooled outcome of the LAB genus by pathogen. There, it can be observed that
Enterococcus strains (15.90 ± 2.138 mm) were in general linked to higher inhibition values against
L. monocytogenes, followed by
Lacticaseibacillus (13.95 ± 2.313 mm),
Lactococcus (13.12 ± 2.129 mm),
Lactobacillus (11.96 ± 1.886 mm),
Leuconostoc (10.25 ± 2.712 mm) and
Pediococcus (8.735 ± 4.663 mm). Regarding
Salmonella spp.,
Enterococcus (12.00 ± 1.195),
Lactobacillus (12.36 ± 1.148 mm) and
Lactococcus (12.76 ± 1.194 mm) had no significant differences between their inhibition values, although
Lactococcus had numerically higher inhibition diameters than the other two.
With regards to S. aureus, the LAB genus which displayed higher inhibition were Enterococcus (11.01 ± 2.105 mm), Lacticaseibacillus (11.89 ± 0.573 mm), Lactobacillus (11.35 ± 1.096 mm), and Lactococcus (11.33 ± 9.578 mm), followed by Leuconostoc (7.173 ± 0.287 mm) and Pediococcus (7.173 ± 0.225 mm). The overall higher inhibition diameter against this pathogen was obtained by Lacticaseibacillus.
The pooled IDs were also estimated by susceptibility test method from the overarching model; and it was encountered that, as a whole, the well diffusion test method produced higher inhibition halos, than those produced by the spot and disk diffusion tests (
Table 1). There were a few exceptions, specifically, in the cases of
Lactobacillus against
L. monocytogenes, and
Lactobacillus against
S. aureus., for which spot, and disk tests were found to be more closely associated with higher inhibition levels. This could be attributed to the diffusion of the supernatant/bacteriocin in the well test being superior when compared to disk, for the mechanical perforation of the agar compared to solely diffusion [
64].
In the agar spot method, a small volume of the antimicrobial agent (such as a supernatant or bacteriocin) is directly placed onto the surface of an agar plate inoculated with the target microorganism. The antimicrobial agent diffuses outward from the spot, creating a zone of inhibition around it where bacterial growth is inhibited. The size of the inhibition zone is measured to assess the antimicrobial activity. Meanwhile, in disk diffusion, paper disks containing a standardized concentration of an antimicrobial substance are placed onto the surface of an agar plate seeded with the test pathogen. As the antimicrobial agent diffuses out of the disk into the agar, it creates a zone of inhibition around the disk where bacterial growth is inhibited. The size of the inhibition zone is measured to evaluate the susceptibility of the microorganism to the antimicrobial agent [
65].
In the well diffusion technique, wells or holes are made in the agar medium using a sterile cork-borer or similar instrument. Antimicrobial agents are then added to the wells in known concentrations. As the antimicrobial agent diffuses outward from the well into the agar, it creates a zone of inhibition around the well. Similar to the other methods, the size of the inhibition zone is measured to assess the effectiveness of the antimicrobial agent. In summary, while all three methods assess antimicrobial activity by measuring zones of inhibition, they differ in how the antimicrobial agent is applied to the agar medium: directly (agar spot), via a disk (disk diffusion), or through a well (well diffusion). Each method has its advantages and limitations and may be preferred based on factors such as the specific antimicrobial agent being tested or laboratory protocols [
66].
3.1. Listeria monocytogenes
The best-fit regression model comprised LAB genus, pathogen concentration and incubation time as the most important moderators of ID (
Table 2). It could be observed that
Enterococcus supernatant provided overall the higher inhibition of the pathogen; meanwhile
Leuconostoc and
Lactobacillus supernatants were significantly found to provide less inhibition, as can be deduced from the higher intercept for inhibition diameter. Enterococci LAB is commonly found in the gastrointestinal tracts of humans and animals, including dairy animals like cows, goats, and sheep. In milk and dairy products, enterococci can play various roles, both beneficial and potentially harmful, they can ferment lactose to produce lactic acid. This fermentation process contributes to the characteristic flavor and texture of fermented dairy products like yogurt, cheese, and kefir [
68].
Some species of enterococci are used as starter cultures or adjunct cultures in cheese production. They contribute to flavor development and texture characteristics during the ripening process of certain types of cheese. They may also help in maintaining gut health by promoting a balanced intestinal microbiota and supporting immune function. However, the safety and efficacy of enterococcal probiotics are subject to ongoing research and debate [
69].
While enterococci are generally considered beneficial in dairy fermentation processes, some species and strains can cause spoilage in milk and dairy products under certain conditions. Enterococci may also produce off-flavors and contribute to the deterioration of product quality if they grow unchecked during processing or storage. Furthermore, there are certain strains of enterococci, such as
Enterococcus faecalis and
Enterococcus faecium, which can harbor antibiotic resistance genes and may pose a risk if present in high numbers or under conditions that allow for the transmission of antibiotic resistance to other bacteria [
70].
The documentation of
L. monocytogenes is mainly carried out due to its numerous outbreaks related to dairy products including pasteurized milk and cheese. Moreover, nisin was found to be an effective biocontrol agent in dairy products against
L. monocytogenes with a decrease of up to 1-log
10 cycle when inoculated in cottage cheese at a pH ranging between 4.6 and 4.7 when stored at 20°C for 7 days. The addition of nisin to ricotta-type cheese was effective in controlling the growth of
L. monocytogenes when incubated at a temperature of 6–8°C for a period of 70 days [
67].
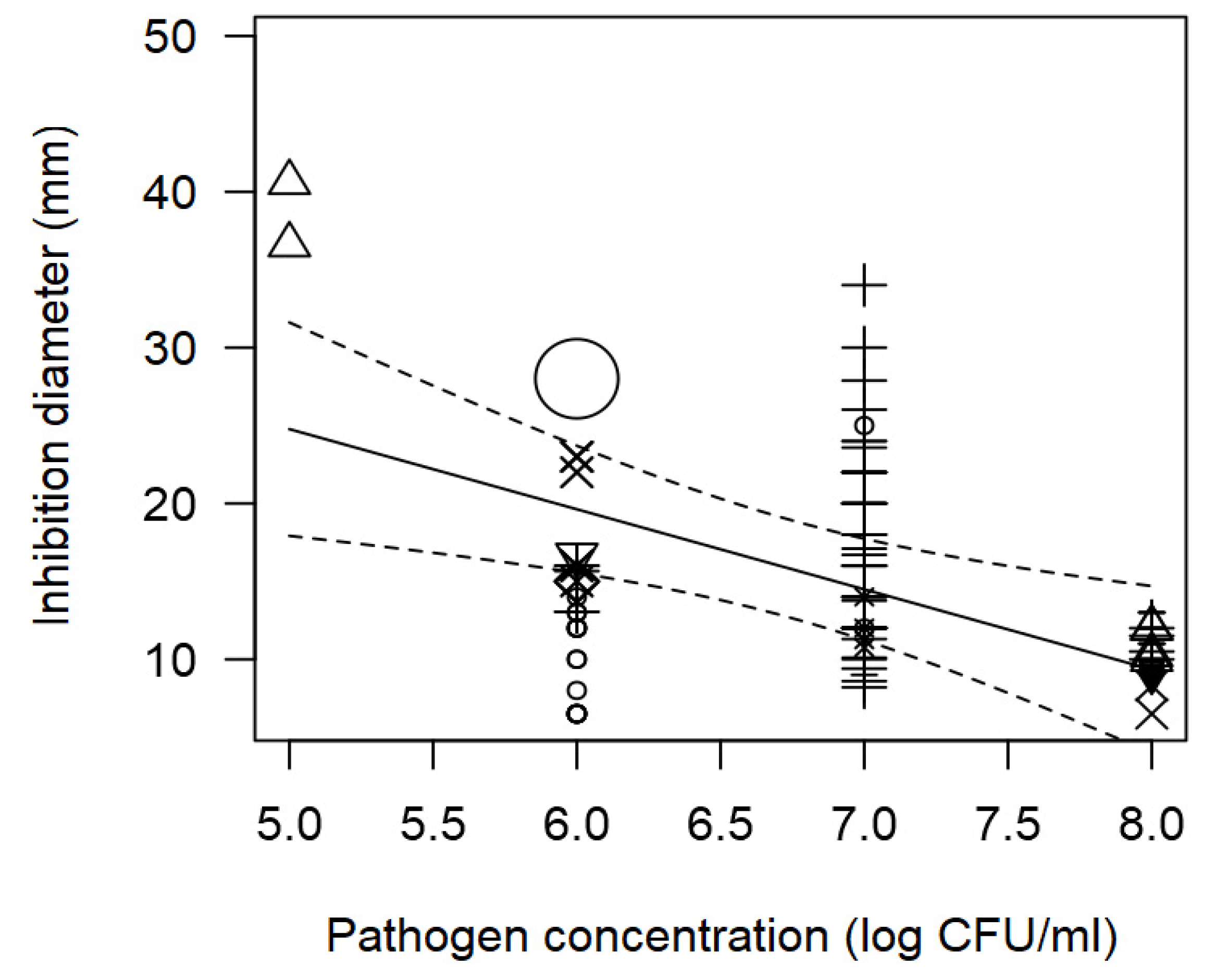
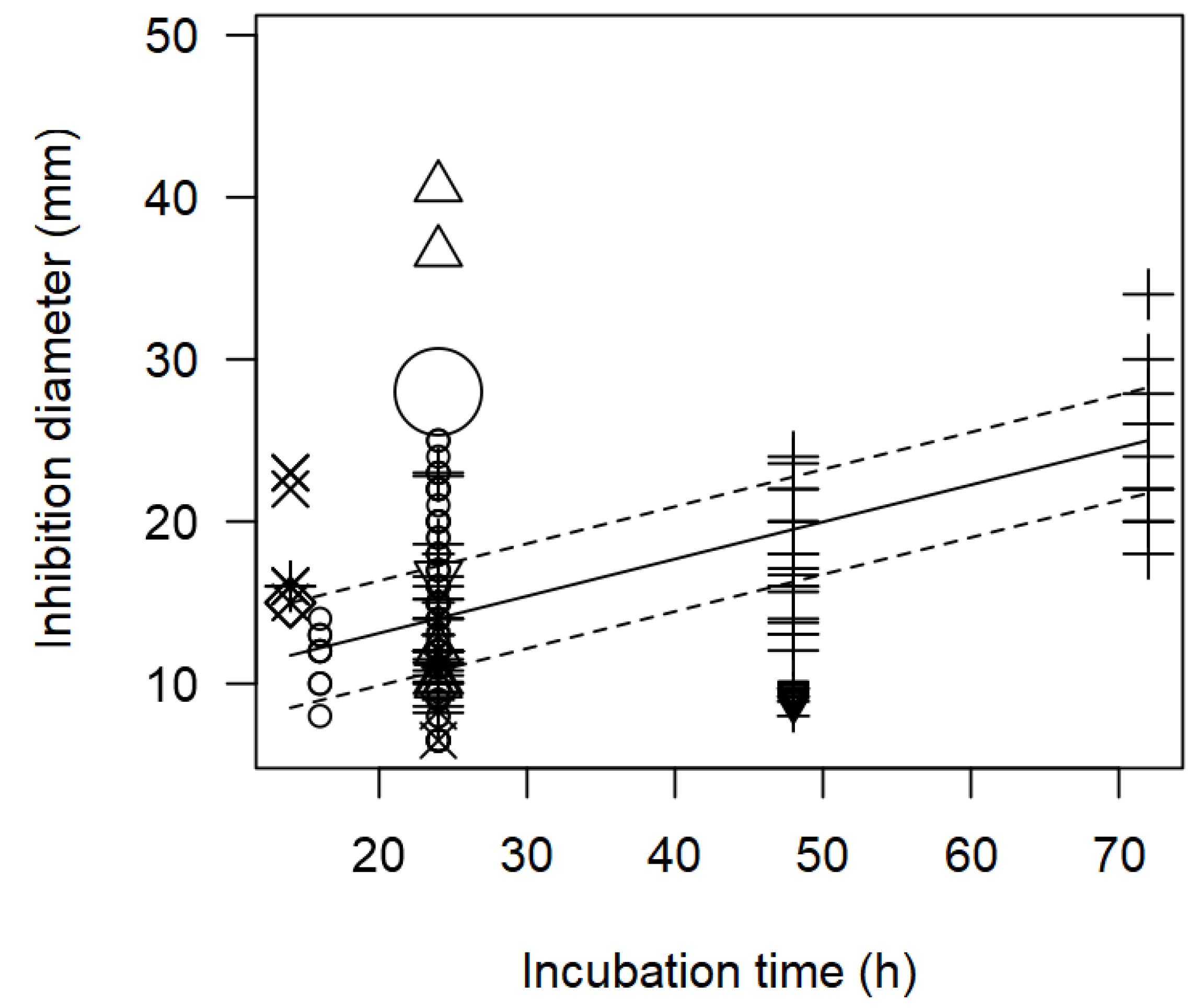
The meta-regression model also demonstrated the significant inverse association between the concentration of pathogen inoculated in the agar and the measured ID; and the direct association between incubation time and observed ID (
Table 2). Such tendencies are elucidated in
Figure 1 and
Figure 2. At a pathogen concentration of 8 log CFU/ml, ID values of ~12 mm were measured (
Figure 1); whereas, at longer incubation times, for instance 72 h, the inhibition values measured tended to be high.
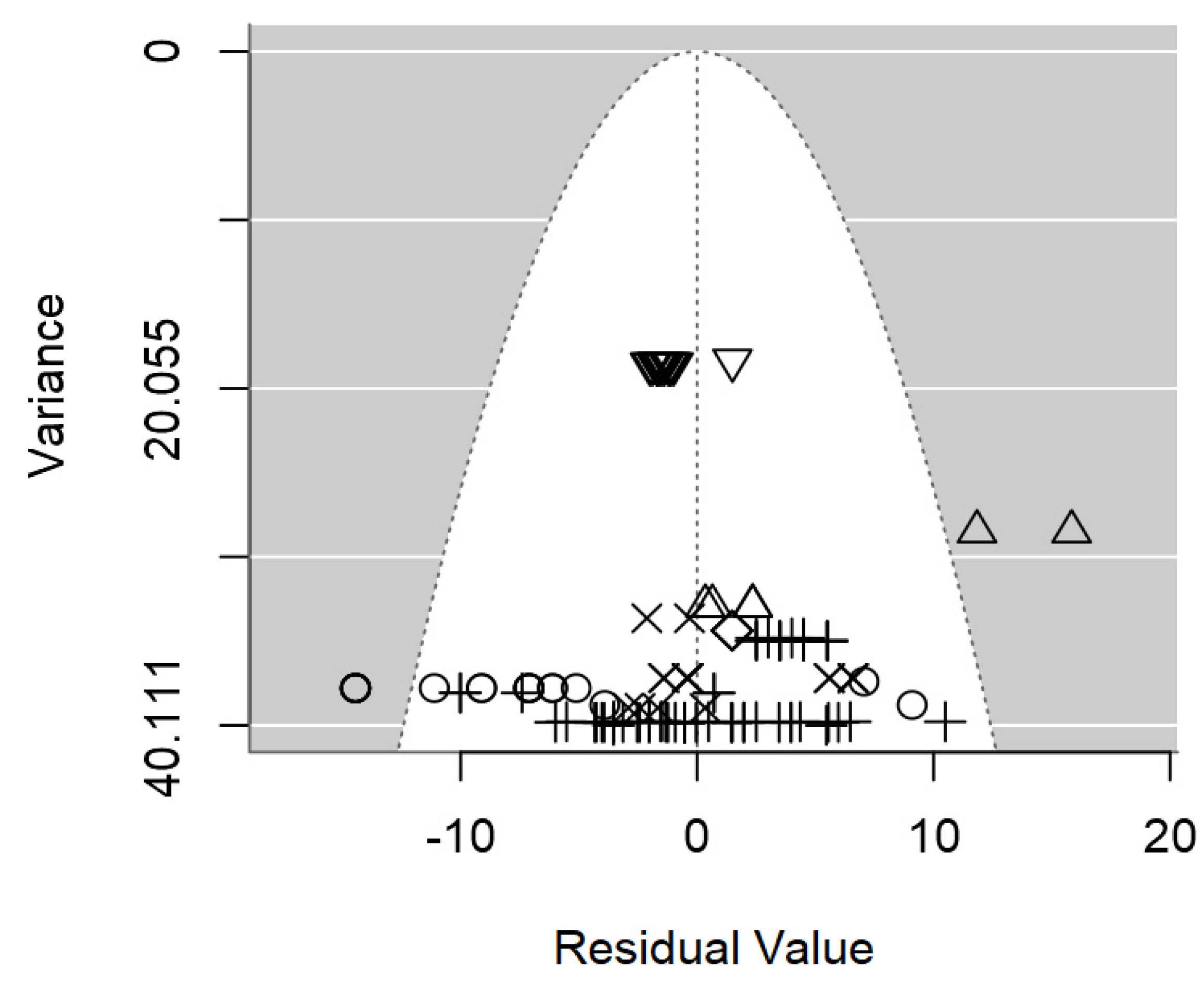
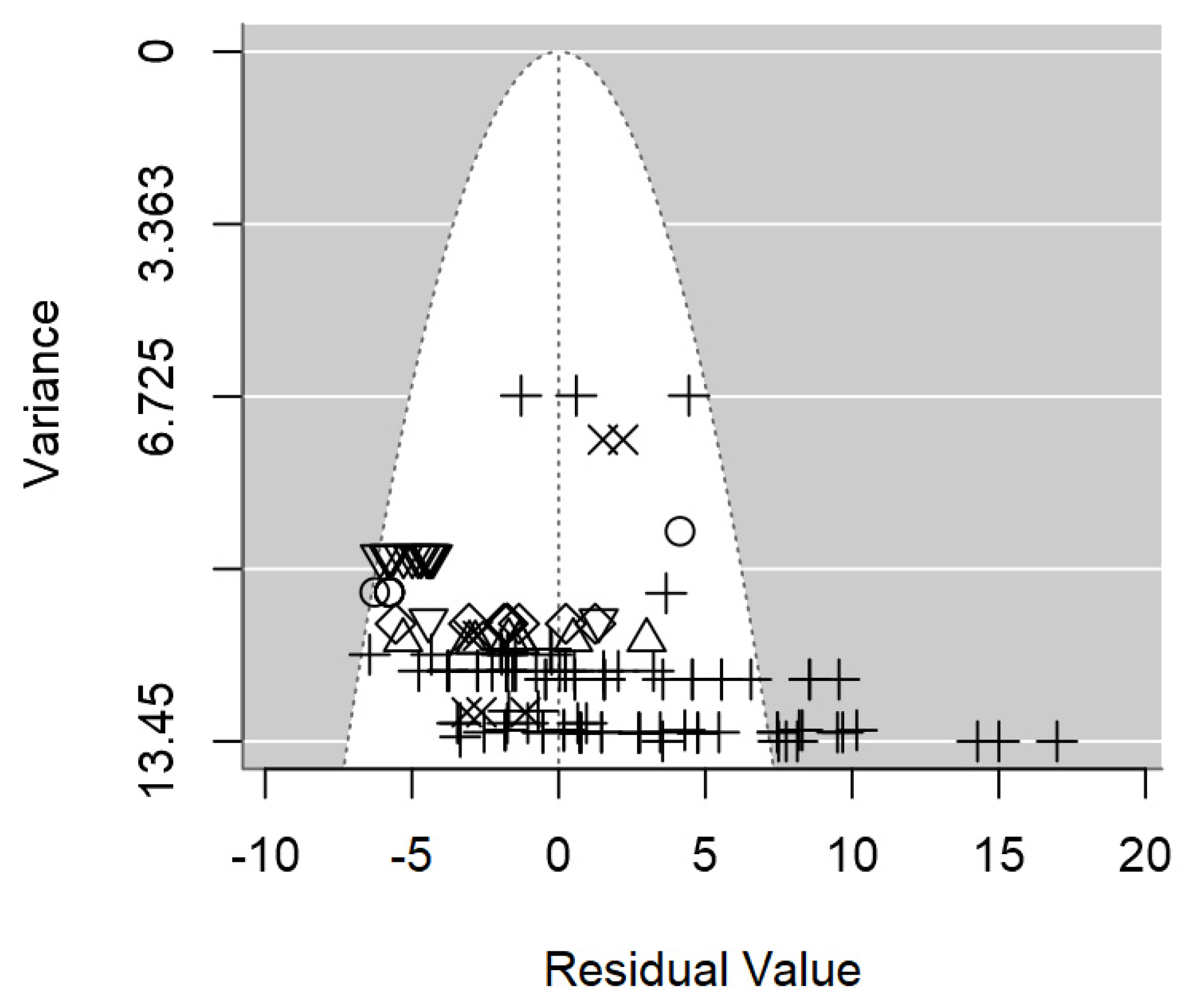
In terms of publication bias, a symmetrical distribution of data points could be appreciated within the funnel (
Figure 3), which provides no discernible evidence of publication bias detected. Likewise, the p-value testing the presence of publication bias was non-significant (p = 0.977).
Heterogeneity analysis pointed out a moderate intraclass correlation (I2=55.1%), from which 33.8% could be explained jointly by the three significant moderators. It is likely that the high unbalancedness of the data hindering the inclusion of other moderators was the cause of the relatively low R2.
3.2. Salmonella spp.
Salmonella contamination in milk and cheese can pose serious health risks to consumers.
Salmonella can contaminate milk and cheese during various stages of production, including milking, processing, and packaging. If the udders of dairy animals are contaminated with
Salmonella, the bacteria can be transferred to the milk during milking. Additionally, contamination can occur from equipment, surfaces, or human handlers during processing and packaging [
71,
72].
Although pasteurization is an effective method for eliminating most pathogens in milk, including Salmonella, if milk is not adequately pasteurized or if contamination occurs post-pasteurization, Salmonella can survive and remain viable in the final product. In cheese production, if raw milk or curd is contaminated with Salmonella, the bacteria can survive the fermentation and ripening processes, leading to contaminated final products.
The meta-regression model fitted on the ID outcomes for
Salmonella spp. allowed the assessment of the effects of susceptibility method, type of agar and incubation time (
Table 3). Other moderators were not significant. The well diffusion method and the Brain Heart Infusion (BHI) agar resulted in the highest inhibition diameters measured. Inversely, the disk diffusion method and the de Man Rogosa and Sharpe (MRS) agar were significantly correlated with lower values of inhibition diameter (mm).
As found with the meta-regression for
L. monocytogenes, the model of the inhibition against
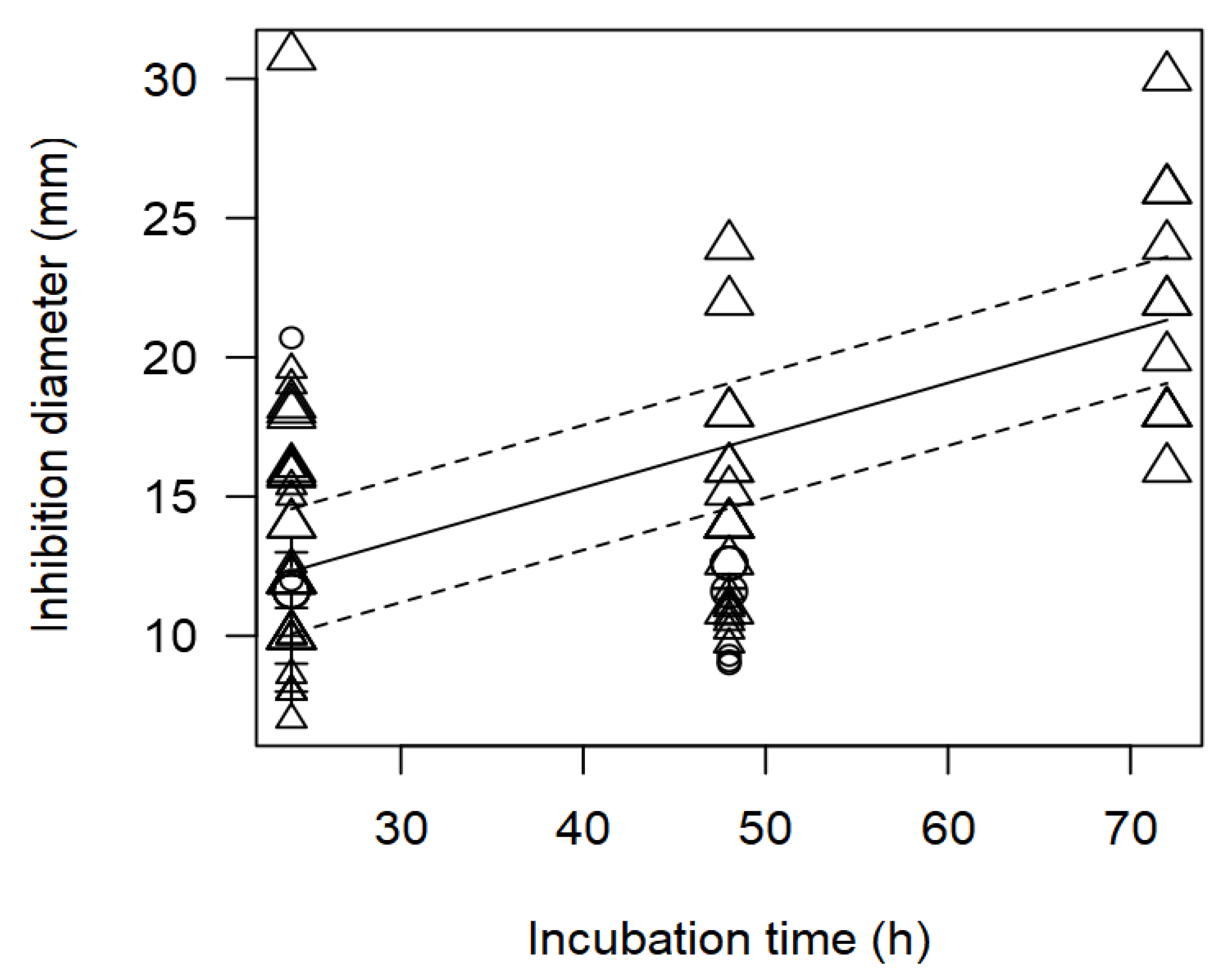
Salmonella spp. evidenced that the inverse relationship between incubation time and measured ID values (p<0.001).
Figure 4 shows the data scattered at three incubation times, 24, 48 and 72 h, and it is mainly
Lactobacillus strains that were associated with higher inhibition diameters at longer incubation times.
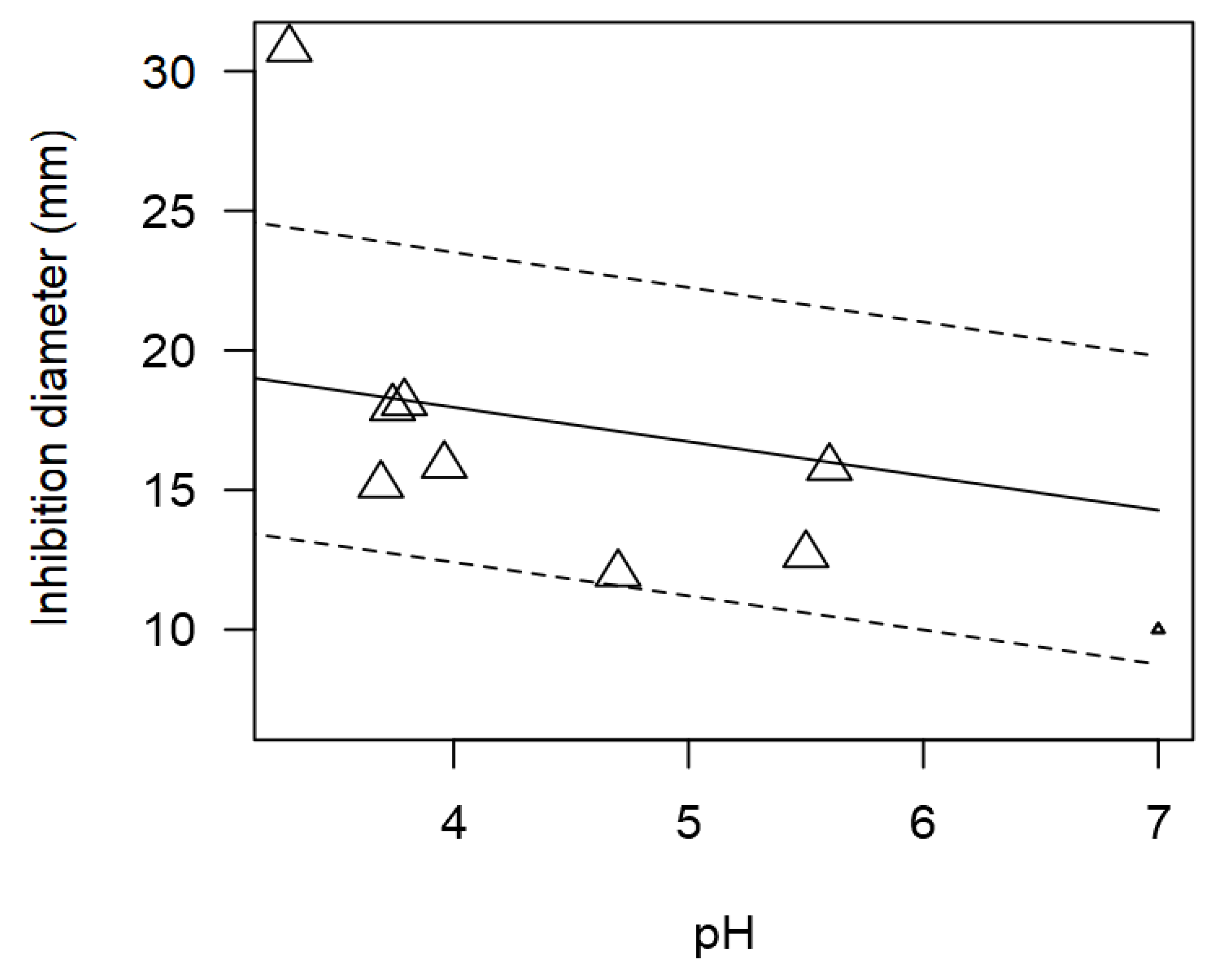
Although pH could not be included as a moderator in the meta-regression model due its very few incidences, still the relationship between pH and ID values was explored. The tendency of pH of the supernatant from LAB and inhibition diameter values were significantly correlated (p<0.001).
Figure 5 shows that
Lactobacillus displayed higher inhibitory activity at more acidic pH compared to neutral pH. The most relevant enzymes produced by LAB are proteases, lipases, bacteriocins, glucoamylases, peptidoglycan hydrolases, ureases, and phenol oxidases, among others. Antimicrobial peptides have emerged as a family of substances with huge potential as a means of microbiological control due to their broad spectrum of activity and low capacity to develop resistance.
Among the antimicrobial peptides used to achieve pathogenic bacteria inhibition, those produced by LAB have significance for applications in food, health, and agriculture. Antimicrobial peptides from bacteria are named bacteriocins and differ from antibiotics in a variety of ways: (1) bacteriocins are ribosomally synthesized, while antibiotics are synthesized by enzymes; (2) bacteriocins have activity in nano- and micromolar concentrations, while antibiotics are used in higher concentrations (μM and mM); and (3) bacteriocins can be used in the clinical and food sector, while antibiotics can only be used clinically.
The publication bias p-value of the meta-regression on ID values for
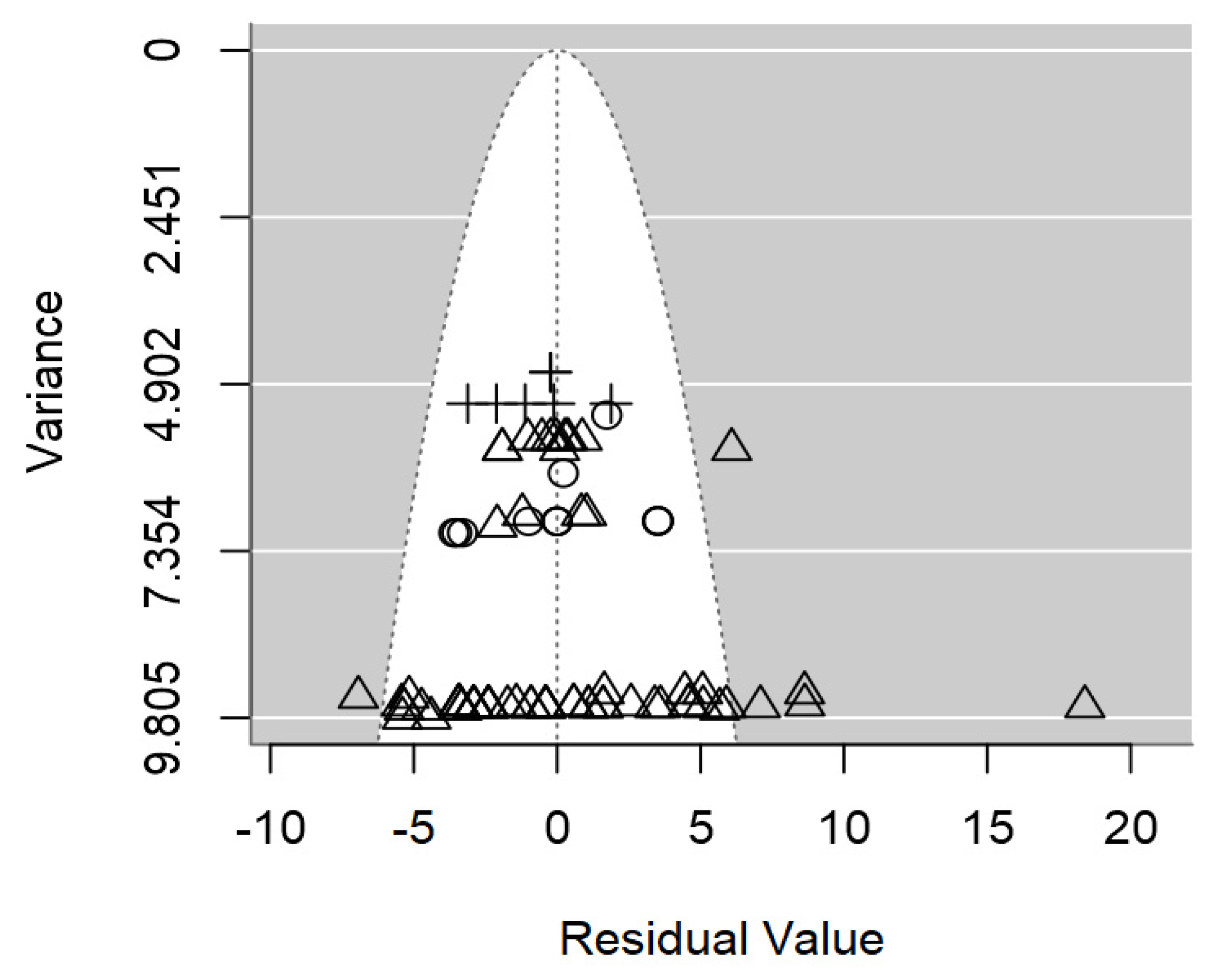
Salmonella spp. was non-significant (p=0.217;
Table 3); which was also supported by the symmetrical dispersion of the data points of the funnel plot shown in
Figure 6. The intraclass correlation of this meta-regression was fairly low (I
2=33.6%), and the three moderators were able to explain 52.3% of the between-study variability (
Table 3).
Lactobacillus inhibition of
Salmonella refers to the ability of certain strains of
Lactobacillus bacteria to inhibit the growth and activity of
Salmonella bacteria. This inhibition can occur through various mechanisms, including competition for nutrients and attachment sites, production of antimicrobial substances (such as organic acids, hydrogen peroxide, and bacteriocins), and modulation of the host's immune response.
Lactobacillus bacteria can outcompete
Salmonella for essential nutrients and attachment sites in the gastrointestinal tract. By colonizing and establishing themselves in the gut,
Lactobacillus strains can prevent
Salmonella from adhering to intestinal epithelial cells and forming biofilms, thus inhibiting its colonization and invasion [
73].
Lactobacillus species are known to produce antimicrobial compounds that can inhibit the growth and survival of pathogenic bacteria like
Salmonella. These antimicrobial substances include organic acids (such as lactic acid and acetic acid) that lower the pH of the intestinal environment, making it less hospitable for
Salmonella growth. Additionally,
Lactobacillus strains can produce hydrogen peroxide, which has antimicrobial properties against
Salmonella. Some
Lactobacillus strains also produce bacteriocins, which are proteinaceous antimicrobial peptides that specifically target and kill
Salmonella cells [
74].
Lactobacillus bacteria can interact with the host's immune system to enhance the body's defense mechanisms against
Salmonella infection. They can stimulate the production of antimicrobial peptides and cytokines, promote the development of a balanced immune response, and strengthen the intestinal barrier function, thereby reducing the likelihood of
Salmonella invasion and systemic infection [
75].
3.3. Staphylococcus aureus
Staphylococcus contamination in milk and dairy products poses a significant concern for food safety. They can be introduced into milk during the milking process. Contamination may occur due to poor udder hygiene, improper milking procedures, or contact with contaminated milking equipment. If milk comes into contact with surfaces, equipment, or personnel contaminated with
Staphylococcus bacteria during processing, it can lead to contamination of dairy products.
Staphylococcus aureus, in particular, is a common source of contamination in dairy processing facilities due to its halophilic nature [
76].
Insufficient cleaning and sanitation practices in dairy processing plants can result in the persistence and spread of
Staphylococcus contamination. Biofilms formed by Sta
phylococcus species on processing equipment and surfaces can serve as reservoirs for bacterial growth and contribute to ongoing contamination issues.
Staphylococcus bacteria are commonly found on human skin and in the nasal passages. If personnel handling milk and dairy products have poor personal hygiene practices or are carriers of
Staphylococcus, they may inadvertently introduce the bacteria into the dairy processing environment, leading to contamination [
77]. Improper handling and storage of dairy products can also result in
Staphylococcus contamination. If dairy products are stored at temperatures conducive to bacterial growth or are exposed to contaminated surfaces or environments after processing,
Staphylococcus bacteria can multiply and cause foodborne illness in consumers [
78].
With regards to the data-driven model fitted for the pathogen
S. aureus, the significant moderators were in this case four: LAB genus, pathogen concentration, incubation time and assay method. Results compiled in
Table 4 show that higher measurements of inhibition diameter were associated with the well diffusion method, as well as the
Enterococcus strain. By contrary, the disk diffusion method led to significantly lower inhibition values, as well as LAB strains belonging to the genus
Leuconostoc and
Pediococcus. In this model, the intraclass correlation was relatively low (I
2=38.8%), and the four moderators could explain only 19.8% of the total between-study variability (
Table 4). Since the variance associated with the different studies is not high, the unexplained part of the variance could be attributed to , more likely, “false” non-significant moderators arising from data unbalancedness, or, less likely, to traits not measured in this study.
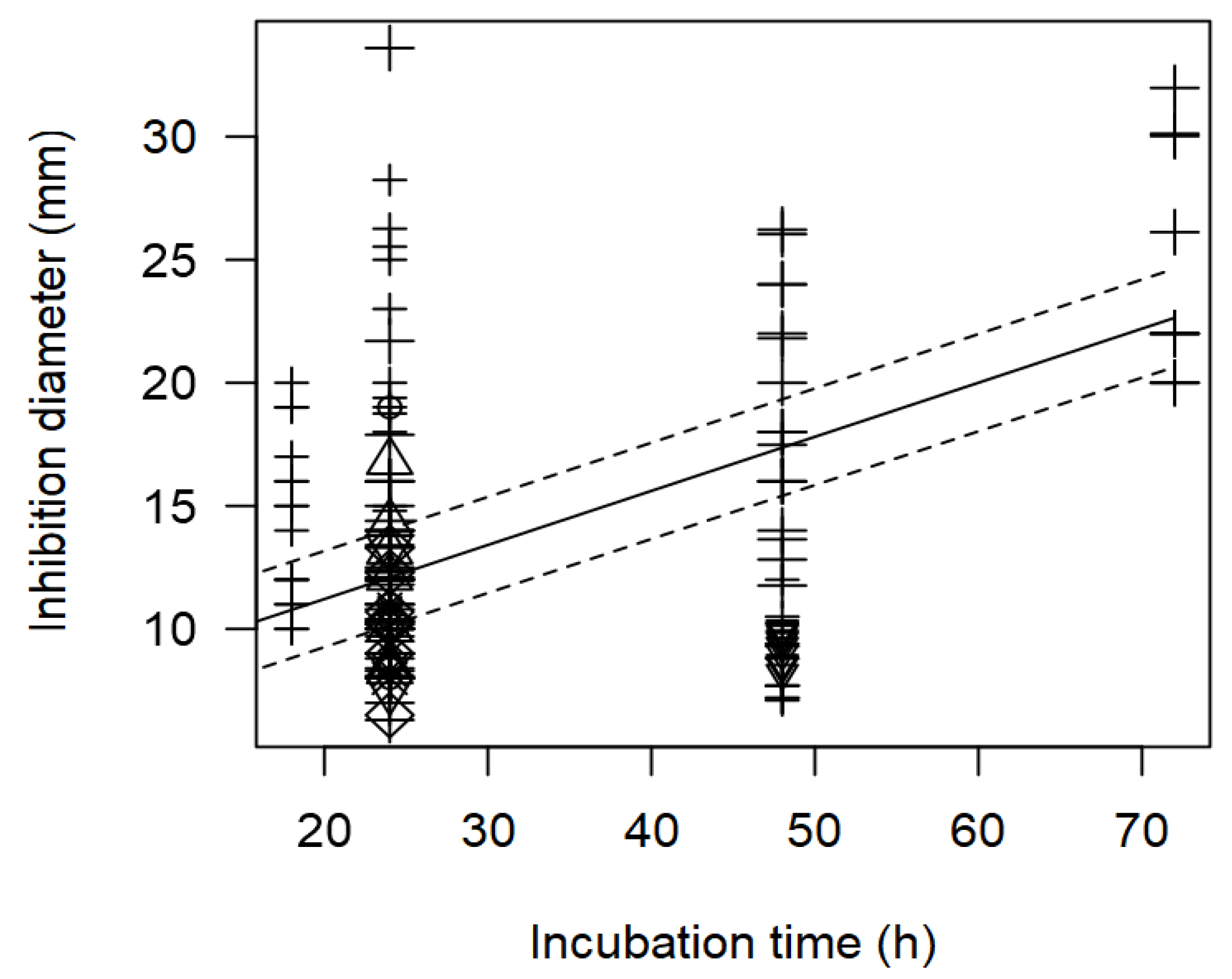
In addition, the meta-regression model showed a significant effect of incubation time (h) on the inhibition diameter of the indicator strains. The longer the incubation period of the indicator strain with the tested bacteriocin/supernatant, the higher the inhibition diameter. In
Figure 7, the significant effect of the incubation time (p<0.001) on the inhibition of
S. aureus is clearly shown. In this case,
Lactobacillus were associated with higher inhibition diameters (>20 mm) when plates were incubated for longer time (72 h)
LAB convert lactose in milk into lactic acid, and protein is degraded into polypeptides or amino acids. Such a process produces organic acids, alcohols, esters, ketones, and other flavor components, endowing fermented milk with specific flavors and rich nutrients. The fermentation process of naturally fermented milk is a dynamic ecological succession of the complex community of milk-associated microbiota that transforms the raw materials. This is mainly driven by the LAB community under specific fermentation conditions. Factors that control the fermentation process include the inoculum size, starter culture interaction with the raw milk and the endogenous microbiota, time of fermentation, oxygen, pH, temperature, and so on. By adjusting these parameters, the micro ecological succession in the milk fermentation process can be finely tuned to yield a variety of fermented milk products including kefir, cheese, cultured butter, matzoon, cultured buttermilk, yogurt, milk curd, and others. Fermentation not only extends food shelf life, but also improves the nutritional value of food and confers health benefits [
79].
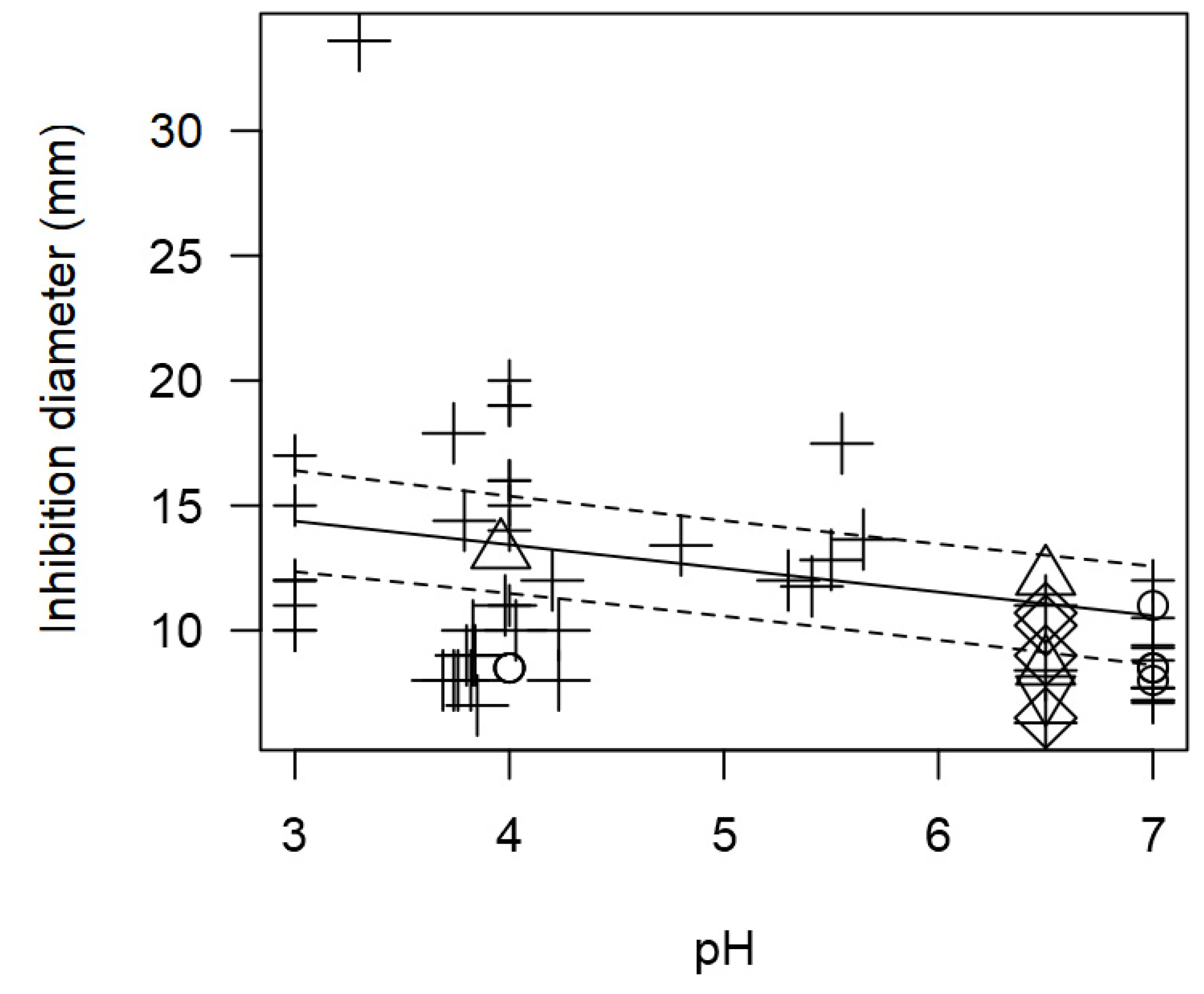
Once again, due to the outcomes with information on pH, this variable could not be assessed in the full meta-regression model. Nonetheless, the association between pH and inhibition diameter was still assessed, to find out again a significant inversion correlation (p<0.001).
Figure 8 shows that more acidic pH values (3 to 5) would tend to increase the measured inhibition diameters resulting in higher values (approximately higher than 12 mm). On the other hand, a more neutral (6 to 7) pH results in less inhibition with lower diameter values (
<10 mm). LAB has an arsenal of strategies to fight against other bacteria that occupy a given ecological niche, among them are the small ribosomal peptides known as bacteriocins. Due to their chemical nature, they have generally been considered as safe for consumption since they are degraded by gastric and pancreatic enzymes during food digestion, although some of them might present toxicity at high concentrations. Currently, the use of bacteriocins as food additives in dairy production is still limited in Europe to nisin (E 234) as approved by the European Food Safety Authority [
2]. This bacteriocin, which is synthesized by strains of
Lactococcus lactis, has antimicrobial activity against a large number of Gram-positive bacteria such as
Listeria,
Staphylococcus,
Bacillus or
Clostridium among others. Nisin is mainly used in cheeses, but it has proved its effectiveness against foodborne pathogens in other dairy products. In the case of milk and cheeses, the main foodborne pathogens are
Staphylococcus aureus and
Listeria monocytogenes, the incidence of
Salmonella enterica and
Escherichia coli being much lower [
80].
In
Figure 9, the symmetrical distribution of data in the plots indicates an absence of significant publication bias. Moreover, these plots imply minimal residual heterogeneity among studies (τ²
res), with no discernible evidence of publication bias detected (p = 0.492).
4. Conclusion
In conclusion, the present meta-analysis has shown that lactic acid bacteria strains naturally occurring in dairy products present variable in vitro inhibition activity against L. monocytogenes, Salmonella spp. and S. aureus, being L. monocytogenes, as a whole, more susceptible to lactic acid bacteria. Strains of Enterococcus, Lacticaseibacillus and Lactobacillus yielded supernatants with the greatest antimicrobial activity against L. monocytogenes and S. aureus, whereas against Salmonella spp. no difference in inhibition capacity was found among LAB genus. In addition, moderators that drive the inhibition diameter measurements are the pathogen’s concentration, incubation time, pH of LAB supernatant, susceptibility method and agar type. Thus, in order to obtain accurate and repeatable inhibition diameter measurements, it is important to optimize the essay protocol, or alternatively, to use well established protocols. Whereas opting for low pathogen concentrations can lead to “promising” susceptibility results, the adjustment to high pH can lead to needlessly discouraging results.
LAB strains, particularly those indigenous to raw milk, offer protective properties against harmful pathogens through the production of lactic acid and other inhibitory compounds during fermentation. Furthermore, employing LAB for natural food preservation aligns with consumer preferences for healthier, minimally processed foods. Continued research into the microbiological quality of cheeses, including the exploration of bacteriocins produced by dairy strains, is essential for enhancing food safety and expanding the range of functional dairy products. By understanding the dynamics of autochthonous LAB species and their antimicrobial activity against foodborne pathogens, stakeholders can implement targeted interventions to improve the safety and quality of dairy products, ultimately safeguarding public health.
Author Contributions
Conceptualization, V.C. and U.G.-B.; methodology, N.F., Y.L., V.C., U.G.-B.; software, N.F., U.G.-B.; validation, V.C. and U.G.B.; formal analysis, N.F., U.G.-B.; investigation, N.F., Y.L., V.C., U.G.-B.; resources, V.C. and U.G.B.; data curation, N.F., Y.L.; writing—original draft preparation, N.F., U.G.-B; writing—review and editing, N.F., V.C. and U.G.-B.; visualization, N.F., Y.L., V.C. and U.G.-B.; supervision, V.C. and U.G.-B.; project administration, V.C. and U.G.-B.; funding acquisition, V.C. and U.G.-B. All authors have read and agreed to the published version of the manuscript.
Funding
The research was funded by the Foundation for Science and Technology (FCT, Portugal) financial support through national funds FCT/MCTES (PIDDAC) to CIMO (UIDB/00690/2020 and UIDP/00690/2020) and SusTEC (LA/P/0007/2021) and through funding of the PAS-AGRO-PAS (PRIMA/0014/2022).
Data Availability Statement
Summary data will be available upon request.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Havelaar, A.H.; Kirk, M.D.; Torgerson, P.R.; Gibb, H.J.; Hald, T.; Lake, R.J.; Praet, N.; Bellinger, D.C.; Silva, N.R. de; Gargouri, N.; Speybroeck, N.; Cawthorne, A.; Mathers, C.; Stein, C.; Angulo, F.J.; Devleesschauwer, B. ; Group, on behalf of W. H. O. F. D. B. E. R. World Health Organization Global Estimates and Regional Comparisons of the Burden of Foodborne Disease in 2010. PLOS Medicine 2015, 12, e1001923. [Google Scholar] [CrossRef]
- Fung, F.; Wang, H.-S.; Menon, S. Food Safety in the 21st Century. Biomed J 2018, 41, 88–95. [Google Scholar] [CrossRef] [PubMed]
- Delgado, J.; Álvarez, M.; Cebrián, E.; Martín, I.; Roncero, E.; Rodríguez, M. Biocontrol of Pathogen Microorganisms in Ripened Foods of Animal Origin. Microorganisms 2023, 11, 1578. [Google Scholar] [CrossRef]
- Gonzales-Barron, U.; Gonçalves-Tenório, A.; Rodrigues, V.; Cadavez, V. Foodborne Pathogens in Raw Milk and Cheese of Sheep and Goat Origin: A Meta-Analysis Approach. Current Opinion in Food Science 2017, 18, 7–13. [Google Scholar] [CrossRef]
- EUROSTAT (2022) Milk and milk product statistics, Eurostat Statistics Explained. Available at: https://ec.europa.eu/eurostat/statistics-explained/index.php?title=Milk_and_milk_product_statistics (Accessed: 25 March 2024).
- Cardinali, F.; Osimani, A.; Taccari, M.; Milanović, V.; Garofalo, C.; Clementi, F.; Polverigiani, S.; Zitti, S.; Raffaelli, N.; Mozzon, M.; Foligni, R.; Franciosi, E.; Tuohy, K.; Aquilanti, L. Impact of Thistle Rennet from Carlina Acanthifolia All. Subsp. Acanthifolia on Bacterial Diversity and Dynamics of a Specialty Italian Raw Ewes’ Milk Cheese. International Journal of Food Microbiology 2017, 255, 7–16. [Google Scholar] [CrossRef] [PubMed]
- Cardinali, F.; Ferrocino, I.; Milanović, V.; Belleggia, L.; Corvaglia, M.R.; Garofalo, C.; Foligni, R.; Mannozzi, C.; Mozzon, M.; Cocolin, L.; Osimani, A.; Aquilanti, L. Microbial Communities and Volatile Profile of Queijo de Azeitão PDO Cheese, a Traditional Mediterranean Thistle-Curdled Cheese from Portugal. Food Research International 2021, 147, 110537. [Google Scholar] [CrossRef] [PubMed]
- Silva, B.N.; Fernandes, N.; Carvalho, L.; Faria, A.S.; Teixeira, J.A.; Rodrigues, C.; Gonzales-Barron, U.; Cadavez, V. Lactic Acid Bacteria from Artisanal Raw Goat Milk Cheeses: Technological Properties and Antimicrobial Potential. Italian Journal of Food Safety 2023, 12. [Google Scholar] [CrossRef]
- Nero, L.A.; Andretta, M.; Almeida, T.T.; Ferreira, L.R.; Camargo, A.C.; Yamatogi, R.S.; Carvalho, A.F.; Call, D.R. Lactic Microbiota of the Minas Artisanal Cheese Produced in the Serro Region, Minas Gerais, Brazil. LWT 2021, 148, 111698. [Google Scholar] [CrossRef]
- Alvarez-Sieiro, P.; Montalbán-López, M.; Mu, D.; Kuipers, O.P. Bacteriocins of Lactic Acid Bacteria: Extending the Family. Appl Microbiol Biotechnol 2016, 100, 2939–2951. [Google Scholar] [CrossRef]
- Favaro, L.; Basaglia, M.; Casella, S.; Hue, I.; Dousset, X.; Dora Gombossy de Melo Franco, B.; Todorov, S.D. Bacteriocinogenic Potential and Safety Evaluation of Non-Starter Enterococcus Faecium Strains Isolated from Home Made White Brine Cheese. Food Microbiology 2014, 38, 228–239. [Google Scholar] [CrossRef]
- Pineda, A.P.A.; Campos, G.Z.; Pimentel-Filho, N.J.; Franco, B.D.G. de M.; Pinto, U.M. Brazilian Artisanal Cheeses: Diversity, Microbiological Safety, and Challenges for the Sector. Front Microbiol 2021, 12, 666922. [Google Scholar] [CrossRef] [PubMed]
- Vandera, E.; Kakouri, A.; Koukkou, A.-I.; Samelis, J. Major Ecological Shifts within the Dominant Nonstarter Lactic Acid Bacteria in Mature Greek Graviera Cheese as Affected by the Starter Culture Type. Int J Food Microbiol 2019, 290, 15–26. [Google Scholar] [CrossRef] [PubMed]
- Lahiri, D.; Nag, M.; Dutta, B.; Sarkar, T.; Pati, S.; Basu, D.; Abdul Kari, Z.; Wei, L.S.; Smaoui, S.; Wen Goh, K.; Ray, R.R. Bacteriocin: A Natural Approach for Food Safety and Food Security. Front Bioeng Biotechnol 2022, 10, 1005918. [Google Scholar] [CrossRef] [PubMed]
- Silva, C.C.G.; Silva, S.P.M.; Ribeiro, S.C. Application of Bacteriocins and Protective Cultures in Dairy Food Preservation. Front Microbiol 2018, 9, 594. [Google Scholar] [CrossRef]
- Ouzzani, M.; Hammady, H.; Fedorowicz, Z.; Elmagarmid, A. Rayyan—a Web and Mobile App for Systematic Reviews. Systematic Reviews 2016, 5, 210. [Google Scholar] [CrossRef]
- Godolphin, P.J.; Bath, P.M.; Montgomery, A.A. Short Email with Attachment versus Long Email without Attachment When Contacting Authors to Request Unpublished Data for a Systematic Review: A Nested Randomised Trial. BMJ Open 2019, 9, e025273. [Google Scholar] [CrossRef]
- Microsoft Corporation. Microsoft Excel Online, Spreadsheet Software. www.microsoft.com. https://office.microsoft.com/excel. (Accessed: 15 January 2024).
- Priya, B.K.; Reddy, D.A.; Rani, A.D.; Kalahasthi, N.; Soliman, W.G.; Reddy, D.V.R.K. Automatic Inhibition Zone Diameter Measurement for Disc Diffusion Test Using Image Segmentation. IETE Journal of Research 2023, 69, 5708–5725. [Google Scholar] [CrossRef]
- R Core Team (2021). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/. (Accessed: 20 January 2024).
- Viechtbauer, W. Conducting Meta-Analyses in R with the Metafor Package. Journal of Statistical Software 2010, 36. [Google Scholar] [CrossRef]
- Pinheiro J, Bates D, R Core Team (2023). nlme: Linear and Nonlinear Mixed Effects Models. R package version 3.1-164. https://CRAN.R-project.org/package=nlme.
- Quintero, A.; Lesaffre, E. Multilevel Covariance Regression with Correlated Random Effects in the Mean and Variance Structure. Biom J 2017, 59, 1047–1066. [Google Scholar] [CrossRef]
- Doleman, B.; Freeman, S.C.; Lund, J.N.; Williams, J.P.; Sutton, A.J. Funnel Plots May Show Asymmetry in the Absence of Publication Bias with Continuous Outcomes Dependent on Baseline Risk: Presentation of a New Publication Bias Test. Res Synth Methods 2020, 11, 522–534. [Google Scholar] [CrossRef]
- Alvarado, C.; García-Almendárez, B.E.; Martin, S.E.; Regalado, C. Anti-Listeria Monocytogenes Bacteriocin-like Inhibitory Substances from Enterococcus Faecium UQ31 Isolated from Artisan Mexican-Style Cheese. Current Microbiology 2005, 51, 110–115. [Google Scholar] [CrossRef] [PubMed]
- Azizi, F.; Habibi Najafi, M.B.; Edalatian Dovom, M.R. The Biodiversity of Lactobacillus Spp. from Iranian Raw Milk Motal Cheese and Antibacterial Evaluation Based on Bacteriocin-Encoding Genes. AMB Express 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Bian, X.; Evivie, S.E.; Muhammad, Z.; Luo, G.-W.; Liang, H.-Z.; Wang, N.-N.; Huo, G.-C. In Vitro Assessment of the Antimicrobial Potentials of Lactobacillus Helveticus Strains Isolated from Traditional Cheese in Sinkiang China against Food-Borne Pathogens. Food Funct 2016, 7, 789–797. [Google Scholar] [CrossRef]
- Câmara, S.P.; Dapkevicius, A.; Riquelme, C.; Elias, R.B.; Silva, C.C.G.; Malcata, F.X.; Dapkevicius, M.L.N.E. Potential of Lactic Acid Bacteria from Pico Cheese for Starter Culture Development. Food Science and Technology International 2019, 25, 303–317. [Google Scholar] [CrossRef]
- Ferrari, I.D.S.; de Souza, J.V.; Ramos, C.L.; da Costa, M.M.; Schwan, R.F.; Dias, F.S. Selection of Autochthonous Lactic Acid Bacteria from Goat Dairies and Their Addition to Evaluate the Inhibition of Salmonella Typhi in Artisanal Cheese. Food Microbiology 2016, 60, 29–38. [Google Scholar] [CrossRef]
- Hejazi, M.A.; Ghafouri-Fard, S.; Eslami, S.; Afshar, D.; Barzegari, A.; Khorshidian, N. Polyphasic Characterization of Enterococcus Strains Isolated from Traditional Moghan Cheese in Iran. Journal of Food Safety 2019, 39. [Google Scholar] [CrossRef]
- Heredia-Castro, P.Y.; Méndez-Romero, J.I.; Hernández-Mendoza, A.; Acedo-Félix, E.; González-Córdova, A.F.; Vallejo-Cordoba, B. Antimicrobial Activity and Partial Characterization of Bacteriocin-like Inhibitory Substances Produced by Lactobacillus Spp. Isolated from Artisanal Mexican Cheese. Journal of Dairy Science 2015, 98, 8285–8293. [Google Scholar] [CrossRef]
- Jutinico-Shubach, A.; Gutiérrez-Cortés, C.; Suarez, H. Antilisterial Activity of Chitosan-Based Edible Coating Incorporating Cell-Free Supernatant from Pediococcus Pentosaceus 147 on the Preservation of Fresh Cheese. Journal of Food Processing and Preservation 2020, 44. [Google Scholar] [CrossRef]
- Kačániová, M.; Borotová, P.; Terentjeva, M.; Kunová, S.; Felšöciová, S.; Haščík, P.; Lopašovský, L.; Štefániková, J. Bryndza Cheese of Slovak Origin as Potential Resources of Probiotic Bacteria. Potravinarstvo Slovak Journal of Food Sciences 2020, 14, 641–646. [Google Scholar] [CrossRef]
- Kanak, E.K.; Yilmaz, S.Ö. Maldi-Tof Mass Spectrometry for the Identification and Detection of Antimicrobial Activity of Lactic Acid Bacteria Isolated from Local Cheeses. Food Science and Technology (Brazil) 2019, 39, 462–469. [Google Scholar] [CrossRef]
- Kanak, E.K.; Yilmaz, S.Ö. Identification, Antibacterial and Antifungal Effects, Antibiotic Resistance of Some Lactic Acid Bacteria. Food Science and Technology (Brazil) 2021, 41, 174–182. [Google Scholar] [CrossRef]
- Leboš Pavunc, A.; Kos, B.; Beganović, J.; Uroić, K.; Bučan, D.; Šušković, J. Antibiotic Susceptibility and Antimicrobial Activity of Autochthonous Starter Cultures as Safety Parameters for Fresh Cheese Production. Mljekarstvo 2013, 63, 185–194. [Google Scholar]
- Loessner, M.; Guenther, S.; Steffan, S.; Scherer, S. A Pediocin-Producing Lactobacillus Plantarum Strain Inhibits Listeria Monocytogenes in a Multispecies Cheese Surface Microbial Ripening Consortium. Applied and Environmental Microbiology 2003, 69, 1854–1857. [Google Scholar] [CrossRef] [PubMed]
- Muhammad, Z.; Ramzan, R.; Abdelazez, A.; Amjad, A.; Afzaal, M.; Zhang, S.; Pan, S. Assessment of the Antimicrobial Potentiality and Functionality of Lactobacillus Plantarum Strains Isolated from the Conventional Inner Mongolian Fermented Cheese Against Foodborne Pathogens. Pathogens 2019, 8, 71. [Google Scholar] [CrossRef] [PubMed]
- Nespolo, C.R.; Brandelli, A. Production of Bacteriocin-like Substances by Lactic Acid Bacteria Isolated from Regional Ovine Cheese. Brazilian Journal of Microbiology 2010, 41, 1009–1018. [Google Scholar] [CrossRef] [PubMed]
- Ołdak, A.; Zielińska, D.; Kołożyn-Krajewska, D. Comparison of antagonistic activity of lactic acid bacteria isolated from various types of traditional food. Zywnosc. Nauka. Technologia. Jakosc/Food. Science Technology. Quality 2019, 26, 60–72. [Google Scholar] [CrossRef]
- Partovi, R.; Gandomi, H.; Akhondzadeh Basti, A. Safety Aspects of Lactobacillus Plantarum Strains Isolated from Siahmazgi Cheese. Ankara Universitesi Veteriner Fakultesi Dergisi 2019, 66, 337–342. [Google Scholar] [CrossRef]
- Settanni, L.; Franciosi, E.; Cavazza, A.; Cocconcelli, P.S.; Poznanski, E. Extension of Tosèla Cheese Shelf-Life Using Non-Starter Lactic Acid Bacteria. Food Microbiology 2011, 28, 883–890. [Google Scholar] [CrossRef]
- Sip, A.; Więckowicz, M.; Olejnik-Schmidt, A.; Grajek, W. Anti-Listeria Activity of Lactic Acid Bacteria Isolated from Golka, a Regional Cheese Produced in Poland. Food Control 2012, 26, 117–124. [Google Scholar] [CrossRef]
- Toğay, S.Ö.; Ay, M.; Güneşer, O.; Yüceer, Y.K. Investigation of Antimicrobial Activity and entA and entB Genes in Enterococcus Faecium and Enterococcus Faecalis Strains Isolated from Naturally Fermented Turkish White Cheeses. Food Science and Biotechnology 2016, 25, 1633–1637. [Google Scholar] [CrossRef]
- Tulini, F.L.; Winkelströter, L.K.; De Martinis, E.C.P. Identification and Evaluation of the Probiotic Potential of Lactobacillus Paraplantarum FT259, a Bacteriocinogenic Strain Isolated from Brazilian Semi-Hard Artisanal Cheese. Anaerobe 2013, 22, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Vandera, E.; Tsirka, G.; Kakouri, A.; Koukkou, A.-I.; Samelis, J. Approaches for Enhancing in Situ Detection of Enterocin Genes in Thermized Milk, and Selective Isolation of Enterocin-Producing Enterococcus Faecium from Baird-Parker Agar. International Journal of Food Microbiology 2018, 281, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Vataščinová, T.; Pipová, M.; Fraqueza, M.J.R.; Maľa, P.; Dudriková, E.; Drážovská, M.; Lauková, A. Short Communication: Antimicrobial Potential of Lactobacillus Plantarum Strains Isolated from Slovak Raw Sheep Milk Cheeses. Journal of Dairy Science 2020, 103, 6900–6903. [Google Scholar] [CrossRef]
- Vera Pingitore, E.; Todorov, S.D.; Sesma, F.; Gombossy de Melo Franco, B.D. Application of Bacteriocinogenic Enterococcus Mundtii CRL35 and Enterococcus Faecium ST88Ch in the Control of Listeria Monocytogenes in Fresh Minas Cheese. Food Microbiology 2012, 32, 38–47. [Google Scholar] [CrossRef]
- Champiri, I.D.; Bamzadeh, Z.; Rahimi, E.; Rouhi, L. Isolation and Identification of Lactobacillus Brevis from Local Cheese of Bazoft and Evaluation of Antimicrobial Activity Against Some Pathogenic Microorganisms. Iranian Journal of Medical Microbiology 2022, 16, 17–34. [Google Scholar] [CrossRef]
- Erdoǧrul, Ö.; Erbilir, F. Isolation and Characterization of Lactobacillus Bulgaricus and Lactobacillus Casei from Various Foods. Turkish Journal of Biology 2006, 30, 39–44. [Google Scholar]
- Heredia-Castro, P.Y.; Reyes-Díaz, R.; Rendón-Rosales, M.Á.; Beltrán-Barrientos, L.M.; Torres-Llanez, M.J.; Estrada-Montoya, M.C.; Hernández-Mendoza, A.; González-Córdova, A.F.; Vallejo-Cordoba, B. Novel Bacteriocins Produced by Lactobacillus Fermentum Strains with Bacteriostatic Effects in Milk against Selected Indicator Microorganisms. Journal of Dairy Science 2021, 104, 4033–4043. [Google Scholar] [CrossRef] [PubMed]
- Hojjati, M.; Behabahani, B.A.; Falah, F. Aggregation, Adherence, Anti-Adhesion and Antagonistic Activity Properties Relating to Surface Charge of Probiotic Lactobacillus Brevis Gp104 against Staphylococcus Aureus. Microbial Pathogenesis 2020, 147. [Google Scholar] [CrossRef]
- Huang, L.; Goda, H.A.; Abdel-Hamid, M.; Renye, J.A., Jr.; Yang, P.; Huang, Z.; Zeng, Q.-K.; Li, L. Partial Characterization of Probiotic Lactic Acid Bacteria Isolated from Chinese Dairy Products. International Journal of Food Properties 2021, 24, 446–456. [Google Scholar] [CrossRef]
- Potočnjak, M.; Pušić, P.; Frece, J.; Abram, M.; Jankovic, T.; Gobin, I. Three New Lactobacillus Plantarum Strains in the Probiotic Toolbox against Gut Pathogen Salmonella Enterica Serotype Typhimurium. Food Technology and Biotechnology 2017, 55, 48–54. [Google Scholar] [CrossRef]
- Slyvka, I.; Tsisaryk, O.; Musii, L.; Kushnir, I.; Koziorowski, M.; Koziorowska, A. Identification and Investigation of Properties of Strains Enterococcus Spp. Isolated from Artisanal Carpathian Cheese. Biocatalysis and Agricultural Biotechnology 2022, 39. [Google Scholar] [CrossRef]
- Ait Chait, Y.; Gunenc, A.; Hosseinian, F.; Bendali, F. Antipathogenic and Probiotic Potential of Lactobacillus Brevis Strains Newly Isolated from Algerian Artisanal Cheeses. Folia Microbiologica 2021, 66, 429–440. [Google Scholar] [CrossRef] [PubMed]
- Jabbari, V.; Khiabani, M.S.; Mokarram, R.R.; Hassanzadeh, A.M.; Ahmadi, E.; Gharenaghadeh, S.; Karimi, N.; Kafil, H.S. Lactobacillus Plantarum as a Probiotic Potential from Kouzeh Cheese (Traditional Iranian Cheese) and Its Antimicrobial Activity. Probiotics and Antimicrobial Proteins 2017, 9, 189–193. [Google Scholar] [CrossRef] [PubMed]
- Karami, S.; Roayaei, M.; Hamzavi, H.; Bahmani, M.; Hassanzad-Azar, H.; Leila, M.; Rafieian-Kopaei, M. Isolation and Identification of Probiotic Lactobacillus from Local Dairy and Evaluating Their Antagonistic Effect on Pathogens. Int J Pharm Investig 2017, 7, 137–141. [Google Scholar] [CrossRef] [PubMed]
- Khalil, N.; Kheadr, E.; El-Ziney, M.; Dabour, N. Lactobacillus Plantarum Protective Cultures to Improve Safety and Quality of Wheyless Domiati-like Cheese. Journal of Food Processing and Preservation 2022, 46. [Google Scholar] [CrossRef]
- Madi, N.; Boushaba, R. Identification of Potential Biopreservative Lactic Acid Bacteria Strains Isolated from Algerian Cow’s Milk and Demonstration of Antagonism against S. Aureus in Cheese. Food Science and Technology Research 2017, 23, 679–688. [Google Scholar] [CrossRef]
- Ngamsomchat, A.; Kaewkod, T.; Konkit, M.; Tragoolpua, Y.; Bovonsombut, S.; Chitov, T. Characterisation of Lactobacillus Plantarum of Dairy-Product Origin for Probiotic Chèvre Cheese Production. Foods 2022, 11, 934. [Google Scholar] [CrossRef] [PubMed]
- Vahabzadeh, S.; Özpinar, H. Investigation of Some Biochemical Properties, Antimicrobial Activity and Antibiotic Resistances of Kefir Supernatants and Lactococcus Lactis Ssp. Lactis Strains Isolated from Raw Cow Milk and Cheese Samples. Kafkas Universitesi Veteriner Fakultesi Dergisi 2018, 24, 443–450. [Google Scholar] [CrossRef]
- Markusková, B.; Lichvariková, A.; Szemes, T.; Koreňová, J.; Kuchta, T.; Drahovská, H. Genome Analysis of Lactic Acid Bacterial Strains Selected as Potential Starters for Traditional Slovakian Bryndza Cheese. FEMS Microbiology Letters 2018, 365, fny257. [Google Scholar] [CrossRef]
- Braga, R.M.; Dourado, M.N.; Araújo, W.L. Microbial Interactions: Ecology in a Molecular Perspective. Braz J Microbiol 2016, 47, 86–98. [Google Scholar] [CrossRef]
- Flanagan, J.N.; Steck, T.R. The Relationship Between Agar Thickness and Antimicrobial Susceptibility Testing. Indian J Microbiol 2017, 57, 503–506. [Google Scholar] [CrossRef] [PubMed]
- Balouiri, M.; Sadiki, M.; Ibnsouda, S.K. Methods for in Vitro Evaluating Antimicrobial Activity: A Review. J Pharm Anal 2016, 6, 71–79. [Google Scholar] [CrossRef] [PubMed]
- Henderson, L.O.; Erazo Flores, B.J.; Skeens, J.; Kent, D.; Murphy, S.I.; Wiedmann, M.; Guariglia-Oropeza, V. Nevertheless, She Resisted - Role of the Environment on Listeria Monocytogenes Sensitivity to Nisin Treatment in a Laboratory Cheese Model. Front Microbiol 2020, 11, 635. [Google Scholar] [CrossRef]
- Taye, Y.; Degu, T.; Fesseha, H.; Mathewos, M. Isolation and Identification of Lactic Acid Bacteria from Cow Milk and Milk Products. ScientificWorldJournal 2021, 2021, 4697445. [Google Scholar] [CrossRef]
- Coelho, M.C.; Malcata, F.X.; Silva, C.C.G. Lactic Acid Bacteria in Raw-Milk Cheeses: From Starter Cultures to Probiotic Functions. Foods 2022, 11, 2276. [Google Scholar] [CrossRef]
- Gołaś-Prądzyńska, M.; Łuszczyńska, M.; Rola, J.G. Dairy Products: A Potential Source of Multidrug-Resistant Enterococcus Faecalis and Enterococcus Faecium Strains. Foods 2022, 11, 4116. [Google Scholar] [CrossRef]
- Freitas Ribeiro, L.; Akira Sato, R.; de Souza Pollo, A.; Marques Rossi, G.A.; do Amaral, L.A. Occurrence of Methicillin-Resistant Staphylococcus Spp. on Brazilian Dairy Farms That Produce Unpasteurized Cheese. Toxins (Basel) 2020, 12, 779. [Google Scholar] [CrossRef] [PubMed]
- Bonilla-Luque, O.M.; Possas, A.; Cabo, M.L.; Rodríguez-López, P.; Valero, A. Tracking Microbial Quality, Safety and Environmental Contamination Sources in Artisanal Goat Cheesemaking Factories. Food Microbiol 2023, 114, 104301. [Google Scholar] [CrossRef]
- Lam, L.H.; Monack, D.M. Intraspecies Competition for Niches in the Distal Gut Dictate Transmission during Persistent Salmonella Infection. PLoS Pathog 2014, 10, e1004527. [Google Scholar] [CrossRef]
- Anjana, null; Tiwari, S. K. Bacteriocin-Producing Probiotic Lactic Acid Bacteria in Controlling Dysbiosis of the Gut Microbiota. Front Cell Infect Microbiol 2022, 12, 851140. [Google Scholar] [CrossRef]
- Gou, H.-Z.; Zhang, Y.-L.; Ren, L.-F.; Li, Z.-J.; Zhang, L. How Do Intestinal Probiotics Restore the Intestinal Barrier? Front Microbiol 2022, 13, 929346. [Google Scholar] [CrossRef] [PubMed]
- Shafique, B.; Ranjha, M.M.A.N.; Murtaza, M.A.; Walayat, N.; Nawaz, A.; Khalid, W.; Mahmood, S.; Nadeem, M.; Manzoor, M.F.; Ameer, K.; Aadil, R.M.; Ibrahim, S.A. Recent Trends and Applications of Nanoencapsulated Bacteriocins against Microbes in Food Quality and Safety. Microorganisms 2022, 11, 85. [Google Scholar] [CrossRef] [PubMed]
- Latorre, A.A.; Oliva, R.; Pugin, J.; Estay, A.; Nualart, F.; Salazar, K.; Garrido, N.; Muñoz, M.A. Biofilms in Hoses Utilized to Divert Colostrum and Milk on Dairy Farms: A Report Exploring Their Potential Role in Herd Health, Milk Quality, and Public Health. Front Vet Sci 2022, 9, 969455. [Google Scholar] [CrossRef]
- Duvenage, S.; Korsten, L. Effect of Temperature and Nutrient Concentration on Survival of Foodborne Pathogens in Deciduous Fruit Processing Environments for Effective Hygiene Management. J Food Prot 2016, 79, 1959–1964. [Google Scholar] [CrossRef]
- Sun, L.; D’Amico, D.J. Composition, Succession, and Source Tracking of Microbial Communities throughout the Traditional Production of a Farmstead Cheese. mSystems 2021, 6, e0083021. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Mu, D.; Qiao, W.; Zhu, D.; Wang, X.; Liu, F.; Xu, H.; Saris, P.; Kuipers, O.P.; Qiao, M. Co-Expression of Nisin Z and Leucocin C as a Basis for Effective Protection Against Listeria Monocytogenes in Pasteurized Milk. Front Microbiol 2018, 9, 547. [Google Scholar] [CrossRef] [PubMed]
Figure 1.
Scatter plot depicting the effect (p=0.006) of L. monocytogenes concentration on the inhibition diameters. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus; and marker size is proportional to study size.
Figure 1.
Scatter plot depicting the effect (p=0.006) of L. monocytogenes concentration on the inhibition diameters. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus; and marker size is proportional to study size.
Figure 2.
Scatter plot depicting the effect (p<0.001) of incubation time on the inhibition diameters. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus; and marker size is proportional to study size.
Figure 2.
Scatter plot depicting the effect (p<0.001) of incubation time on the inhibition diameters. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus; and marker size is proportional to study size.
Figure 3.
Funnel plot of the meta-regression model on the inhibition diameter produced by lactic acid bacteria supernatant against L. monocytogenes. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus.
Figure 3.
Funnel plot of the meta-regression model on the inhibition diameter produced by lactic acid bacteria supernatant against L. monocytogenes. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus.
Figure 4.
Scatter plot depicting the effect (p<0.001) of incubation time on the inhibition diameters of Salmonella spp. Markers symbolize bacterium: ○= Enterococcus, ∆= Lactobacillus, += Lactococcus; and marker size is proportional to study size.
Figure 4.
Scatter plot depicting the effect (p<0.001) of incubation time on the inhibition diameters of Salmonella spp. Markers symbolize bacterium: ○= Enterococcus, ∆= Lactobacillus, += Lactococcus; and marker size is proportional to study size.
Figure 5.
Scatter plot depicting the effect (p<0.001) of pH on the inhibition diameters for Lactobacillus against Salmonella spp. Marker size is proportional to study size. The pH was not added in the final model as moderator for having too few incidences.
Figure 5.
Scatter plot depicting the effect (p<0.001) of pH on the inhibition diameters for Lactobacillus against Salmonella spp. Marker size is proportional to study size. The pH was not added in the final model as moderator for having too few incidences.
Figure 6.
Funnel plot of the meta-regression model on the inhibition diameter produced by lactic acid bacteria supernatant against Salmonella spp. Markers symbolize bacterium: ○= Enterococcus, ∆= Lactobacillus, += Lactococcus.
Figure 6.
Funnel plot of the meta-regression model on the inhibition diameter produced by lactic acid bacteria supernatant against Salmonella spp. Markers symbolize bacterium: ○= Enterococcus, ∆= Lactobacillus, += Lactococcus.
Figure 7.
Scatter plot depicting the effect (p<0.001) of incubation time on the inhibition diameters. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus; and marker size is proportional to study size.
Figure 7.
Scatter plot depicting the effect (p<0.001) of incubation time on the inhibition diameters. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus; and marker size is proportional to study size.
Figure 8.
Scatter plot depicting the effect (p<0.001) of pH on the inhibition diameters. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, + = Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus; and marker size is proportional to study size. pH was not added in the final model as moderator for having too few incidences.
Figure 8.
Scatter plot depicting the effect (p<0.001) of pH on the inhibition diameters. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, + = Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus; and marker size is proportional to study size. pH was not added in the final model as moderator for having too few incidences.
Figure 9.
Funnel plot of the meta-regression model on the inhibition diameter produced by lactic acid bacteria supernatant against S. aureus. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus.
Figure 9.
Funnel plot of the meta-regression model on the inhibition diameter produced by lactic acid bacteria supernatant against S. aureus. Markers symbolize bacterium: ○= Enterococcus, ∆= Lacticaseibacillus, += Lactobacillus, ×= Lactococcus, ◊= Leuconostoc, =Pediococcus.
Table 1.
Pooled inhibition diameters (mean and standard error in mm) produced by lactic acid bacteria (LAB) supernatant using pathogen’s concentration of 7.0 log CFU/ml and an incubation time of 24 h, as estimated by meta-regression models adjusted by foodborne pathogen. Number of observations (n), and number of primary studies (N) are shown.
Table 1.
Pooled inhibition diameters (mean and standard error in mm) produced by lactic acid bacteria (LAB) supernatant using pathogen’s concentration of 7.0 log CFU/ml and an incubation time of 24 h, as estimated by meta-regression models adjusted by foodborne pathogen. Number of observations (n), and number of primary studies (N) are shown.
| Pathogen 1.
|
LAB genus |
Method |
Pooled inhibition
Diameter2,3 (SE) [mm] |
n |
N |
|
Listeria monocytogenesA
|
Enterococcus |
Overall |
15.90A (2.138) |
|
38 |
|
| |
|
Well |
|
19.47a (2.409) |
28 |
|
| |
|
Disk |
|
10.98c (1.962) |
10 |
|
| |
Lacticaseibacillus |
Overall |
13.95B (2.313) |
|
8 |
|
| |
|
Well |
|
28.57a (5.058) |
4 |
|
| |
|
Disk |
|
11.73c (1.835) |
4 |
|
| |
Lactobacillus |
Overall |
11.96C (1.886) |
|
92 |
24 |
| |
|
Spot |
|
14.87b (2.883) |
14 |
|
| |
|
Well |
|
12.02c (2.541) |
18 |
|
| |
|
Disk |
|
10.88c (1.598) |
60 |
|
| |
Lactococcus |
Overall |
13.12B (2.129) |
|
20 |
|
| |
|
Spot |
|
17.75a (2.076) |
14 |
|
| |
|
Disk |
|
10.23c (1.792) |
6 |
|
| |
Leuconostoc |
Overall |
10.25C (2.712) |
|
5 |
|
| |
|
Spot |
|
14.04b (3.045) |
4 |
|
| |
Pediococcus |
Overall |
8.735C (4.663) |
|
14 |
|
| |
|
Well |
|
8.800c (3.025) |
14 |
|
|
Salmonella spp.B
|
Enterococcus |
Overall |
12.00A (1.195) |
|
13 |
|
| |
|
Well |
|
13.39a (1.408) |
12 |
|
| |
Lactobacillus |
Overall |
12.36A (1.148) |
|
69 |
|
| |
|
Well |
|
13.58a (1.362) |
18 |
14 |
| |
|
Disk |
|
11.13b (1.361) |
51 |
|
| |
Lactococcus |
Overall |
12.76A (1.194) |
|
7 |
|
| |
|
Disk |
|
10.82b (1.433) |
5 |
|
|
Staphylococcus aureus B
|
Enterococcus |
Overall |
11.01A (2.105) |
|
10 |
|
| |
|
Well |
|
14.16a (0.716) |
4 |
|
| |
|
Disk |
|
5.28d (0.366) |
6 |
|
| |
Lacticaseibacillus |
Overall |
11.89A (0.573) |
|
9 |
|
| |
|
Well |
|
12.36b (1.501) |
5 |
|
| |
|
Disk |
|
12.00b (1.197) |
4 |
|
| |
Lactobacillus |
Overall |
11.35A (1.096) |
|
133 |
25 |
| |
|
Well |
|
10.00c (1.772) |
67 |
|
| |
|
Disk |
|
12.03b (1.394) |
66 |
|
| |
Lactococcus |
Overall |
11.33A (9.578) |
|
7 |
|
| |
|
Disk |
|
11.24b (0.529) |
5 |
|
| |
Leuconostoc |
Overall |
7.173B (0.287) |
|
8 |
|
| |
|
Well |
|
7.198 (0.304) |
8 |
|
| |
Pediococcus |
Overall |
7.173B (0.225) |
|
15 |
|
| |
|
Well |
|
7.161 (0.232) |
15 |
|
Table 2.
Final meta-regression model on inhibition diameter produced by lactic acid bacteria (LAB) supernatant against L. monocytogenes, as a function of LAB genus, pathogen concentration (log CFU/ml) and incubation time (h). Number of observations (n) per LAB genus, pathogen concentration (log CFU/ml) and incubation time (h). Number of observations (n) per LAB genus, heterogeneity analysis and p-value of the publication bias test are shown.
Table 2.
Final meta-regression model on inhibition diameter produced by lactic acid bacteria (LAB) supernatant against L. monocytogenes, as a function of LAB genus, pathogen concentration (log CFU/ml) and incubation time (h). Number of observations (n) per LAB genus, pathogen concentration (log CFU/ml) and incubation time (h). Number of observations (n) per LAB genus, heterogeneity analysis and p-value of the publication bias test are shown.
| Parameter |
Estimate1
|
Standard error |
p-value |
n |
Heterogeneity analysis2
|
| Intercept |
45.67 |
12.41 |
0.002 |
|
s² = 53.0 |
| LAB genus |
|
|
|
|
τ² = 65.2 |
|
Lacticaseibacillus
|
-1.217 |
2.112 |
0.565 |
8 |
τ²res = 43.1 |
|
Lactobacillus
|
-3.375 |
1.580 |
0.033 |
92 |
R2 = 33.8% |
|
Lactococcus
|
-2.248 |
1.787 |
0.208 |
20 |
τ² = 65.2 |
|
Leuconostoc
|
-5.140 |
2.528 |
0.042 |
5 |
|
|
Enterococcus
|
- |
- |
- |
81 |
|
| Pathogen concentration |
-5.037 |
1.840 |
0.006 |
|
Publication bias |
| Incubation time |
0.229 |
0.229 |
<.0001 |
|
p = 0.977 |
Table 3.
Final meta-regression model on inhibition diameter produced by lactic acid bacteria (LAB) supernatant against Salmonella spp., as a function of susceptibility test method, agar and incubation time (h). Heterogeneity analysis and p-value of the publication bias test are shown.
Table 3.
Final meta-regression model on inhibition diameter produced by lactic acid bacteria (LAB) supernatant against Salmonella spp., as a function of susceptibility test method, agar and incubation time (h). Heterogeneity analysis and p-value of the publication bias test are shown.
| Parameter |
Estimate1
|
Standard error |
p-value |
n |
Heterogeneity analysis2
|
| Intercept |
10.41 |
2.250 |
<.0001 |
|
|
| Susceptibility method |
|
|
|
|
|
| Disk diffusion |
-2.529 |
0.979 |
0.009 |
|
s² = 25.1 |
| Agar |
|
|
|
|
τ² = 12.8 |
| MH |
-1.540 |
2.920 |
0.589 |
11 |
I2 = 33.6% |
| MRS |
-8.114 |
2.273 |
0.001 |
20 |
τ²res = 6.1 |
| Nutrient |
3.575 |
2.553 |
0.161 |
5 |
R2 = 52.3% |
| TSA |
-2.111 |
2.037 |
0.300 |
10 |
|
| BHI |
- |
- |
- |
43 |
Publication bias |
| Incubation time |
0.188 |
0.005 |
<.0001 |
|
p = 0.217 |
Table 4.
Final meta-regression model on inhibition diameter produced by lactic acid bacteria (LAB) supernatant against S. aureus, as a function of LAB genus, pathogen concentration (log CFU/ml), incubation time (h) and susceptibility test method. Heterogeneity analysis and p-value of the publication bias test are shown.
Table 4.
Final meta-regression model on inhibition diameter produced by lactic acid bacteria (LAB) supernatant against S. aureus, as a function of LAB genus, pathogen concentration (log CFU/ml), incubation time (h) and susceptibility test method. Heterogeneity analysis and p-value of the publication bias test are shown.
| Parameter |
Estimate1
|
Standard error |
p-value |
n |
Heterogeneity analysis2
|
| Intercept |
22.04 |
2.545 |
<.0001 |
|
|
| LAB genus |
|
|
|
|
s² = 31.8 |
|
Lacticaseibacillus
|
-0.923 |
1.481 |
0.533 |
9 |
τ² = 20.2 |
|
Lactobacillus
|
-1.295 |
1.208 |
0.283 |
133 |
I2 = 38.8% |
|
Lactococcus
|
-1.758 |
1.842 |
0.340 |
7 |
τ²res = 16.2 |
|
Leuconostoc
|
-5.666 |
1.835 |
0.002 |
8 |
R2 = 19.8% |
|
Pediococcus
|
-7.335 |
2.154 |
0.001 |
15 |
|
|
Enterococcus
|
- |
- |
- |
10 |
| Pathogen concentration |
-1.888 |
0.297 |
<.0001 |
|
Publication bias |
| Incubation time |
0.220 |
0.005 |
<.0001 |
|
p = 0.492 |
| Susceptibility method |
|
|
|
|
|
| Disk diffusion |
-2.373 |
0.483 |
<.0001 |
|
|
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).