1. Introduction
Chili pepper (
Capsicum spp.), a member of the Solanaceae family, is a highly valued cash crop. The global chili pepper production in 2022 was 59.0 million tonnes from an area of 44.7 million hectares. China, Mexico, Indonesia, Türkiye and India are primary producers along with Thailand [
1]. The productivity of chili peppers has been declining due to climate change [
2] and the incidence of insects and diseases [
3,
4]. The major pepper diseases prevalent in tropical and subtropical regions are anthracnose, fruit rot, bacterial wilt, and Pepper leaf curl virus disease.
Pepper leaf curl virus (PepLCV) belongs to the Begomovirus genus. Two species including
Pepper yellow leaf curl Kanchanaburi virus (PepYLCKaV) and
Pepper yellow leaf curl Thailand virus (PepYLCTHV) were diagnosed and characterized from chili pepper growing areas of Thailand [
5]. The virus-causing leaf curl disease is transmitted by whitefly (
Bemissai tabaci) and was first identified in Indonesia in the early 2000s [
6]. It was initially observed in Thailand in 1995. Chiemsombat et al. [
5] reported a significant increase in PepYLCV cases in 2014 and rapidly expanded to
Capsicum chinense and
C. frutescens species in Kanchanaburi province, Thailand. Currently, most of the commercial cultivars of
C. annuum L. widely grown in Thailand are susceptible to PepYLCV. The disease is difficult to control as the vector of the virus (whiteflies) has a wide host range like okra, tomato, eggplant, soybean, angled luffa, cucumber, bottle gourd and pumpkin [
5,
7]. Developing PepYLCV-resistant cultivars in chili peppers has been challenging due to the disease's nature, the vector and the complex infection strategies of viruses [
8,
9]. Begomovirus-induced studies on various sources of resistance have revealed qualitative and quantitative gene actions for resistance, depending on the virus strain, chili pepper genotype, and their interaction [
10].
The resistant sources to PepYLCTHV, PepGMV and PHV diseases were reported in C.
annuum include 9853-123, PSP11-4 [
11,
12], DLS-Sel-10, WBC-Sel-5 and PBC142 [
13]. However, no chili pepper commercial have been reported resistant to the disease and also none of these genotypes was purely resistant as inoculated plants of each entry segregated into resistant and susceptible plants. Due to the resistant traits were controlled by various genetic mechanisms [
14] such as single recessive and dominant genes with inherited by additive, non-additive and epitasis gene action [
15]. Thus, the purification of resistant plants is imperative.
Mass selection has been an efficient method for populations controlled by polygenic genes. Chili peppers are self-pollinating crops and classical methods such as pure line and mass selection have been utilized for germplasm purification, depending on the heritability trait [
16,
17]. However, there has been no reports on the selection of individual plants resistant to PepYLCV. In this communication we report a comprehensive approach to select resistant genotypes and improve the resistance levels of individual plants against PepYLCTHV using the viruliferous whitefly inoculations.
2. Materials and Methods
2.1. Plant Materials
Plant materials for this investigation consisted of four screening cycles. In the 1
st cycle, we screened 31 chili pepper genotypes comprising of 24 breeding lines sourced from Khon Kaen University (KKU), Thailand, known for their resistance to virus diseases under field conditions. The remaining lines were selected from the World Vegetable Center, Taiwan (improved lines). Pong Charian and 9853-123 were used as susceptible and resistant checks, respectively (
Table 1). A randomized complete block design was used with three replications and 10 plants per replication. The experiment was performed during January to April 2020. For the 2
nd cycle, two resistant lines (PSP11-4 and 9853-123) from the first cycle selection based on low disease severity means, and the highly resistant plants (score 0). The seeds of the selected plants were screened for resistance to PepYLCV from June to October 2021 (
Figure 1). Based on phenotypic screening and PCR results, the resistant line PSP11-4 (coded as PEP6) was selected from a highly resistant genotype (R-), while line 9853-123 (code PEP12) was selected from a highly resistant genotype (R+). The seeds from the selected plants of the second cycle were continuously challenged and screened for the third and the fourth cycles from January to May 2022, October 2022 to April 2023, respectively. Screening, selection, and generation advancement from C
1 (30 plants/line), C
2 (60 plants/line) and C
3 (200 plants/line) generations were performed at Kasetsart University, Kamphaengsaen Campus, Nakhon Pathom, Thailand. Subsequently, screening and selection of C
4 (100 plants/line) were performed at the School of Agricultural of Technology, King’s Mongkut Institute of Technology Ladkrabang, Thailand by artificial whitefly transmission. The number of plant screenings for each cycle depends on the extent of seed settings in resistant plants.
2.2. Viruliferus Whitefly, Inoculation and Disease Evaluation
Nonviruliferous whiteflies (Bemisia tabaci Genn) were maintained and multiplied on cotton plants (Gossypium hirsutum) in cages. Twenty-one days old adult whiteflies were. The non-viruliferous whiteflies were released on PepYLCTHV-infected chili pepper plants (Pong Charian) for 24 hours acquisition access period. Chili pepper leaf was sampled for the confirmation of PepYLCTHV using universal primers specific for pepper yellow leaf curl, PepYLCTHV-F: 5’ ATGGCGAAGCGTCCCGCAGAT 3’, and PepYLCTHV-R: 5’CTGCAGTTAATTTTGAACCGAACC3’.
Resistance tests were conducted in insect-free greenhouses to keep seedlings free from viruses. Forty-five days after sowing, the seedling was transferred individually to insect-proof net cages 60 mesh size 100 x 100 x 100 cm. 10 to 15 sterile whitefly imagoes were introduced to each plant for 48 hours (during feeding time, the plants were shanked). After transmission, the whiteflies were killed by spraying insecticide (acetamiprid 20% SP).
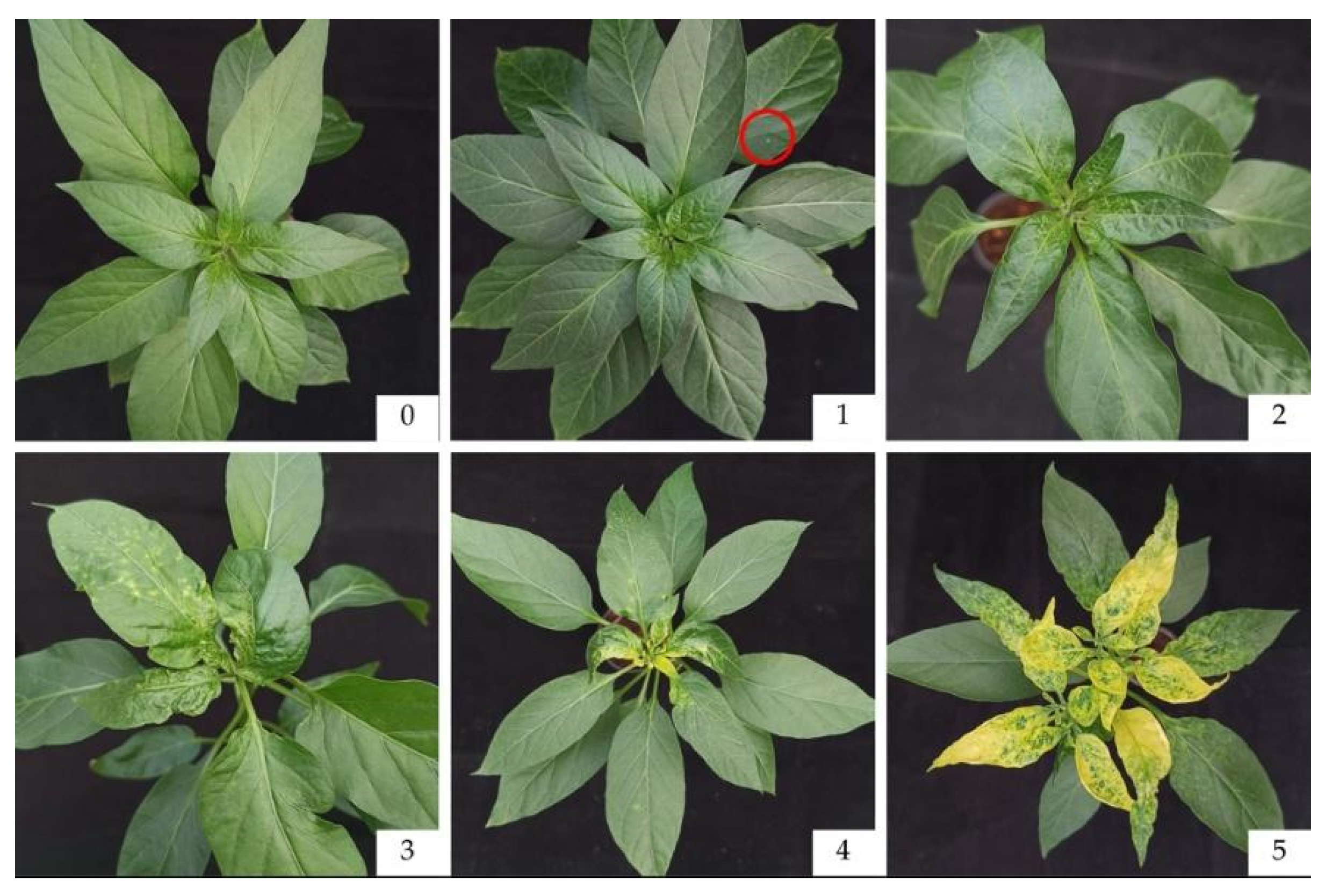
The disease scores were recorded at 0, 7, 14, 21, and 28 days after inoculation, using 6 score levels (0, 1, 2, 3, 4, and 5) following Suwor et al [
12] (
Figure 2,
Table 2).
The 6 score levels were calculated to disease incidence (DI%) by using the following formula:
With i: 0-5, ni = number of symptomatic plants to value of a particular score, vi = value symptom score, N = the total number of plants, observed, and V = the highest score value.
Disease severity and percentage disease index were analyzed using variance analysis of variance using the Statistix 10 version software.
2.3. DNA Extraction and Verification of Viral Genomes
At 28 DAI, the no-symptom plants of all genotypes were screened for the present or absence of PepYLCTHV genome by using specific PepYLCTHV primers. Total DNA was extracted from individual plants at 28 DAI. Three young leaves were homogenized in 750 µL of 60°C extraction buffer, which contained 2% CTAB, 10% 1M Tris HCl (pH 8.0), 28% 5M NaCl, 4% 0.5M EDTA, and 0.2% mercaptone. The plant sap was then incubated at 60°C for 30 minutes. Next, 750 μl of chloroform:isoamyl alcohol (24:1) was added and mixed. The samples were precipitated by centrifugation at 6000 rpm for 10 minutes. Afterward, 450 µL of the supernatant was transferred to a 1.5 µL tube. Subsequently, 450 µL of 95% ethanol was added and the mixture was stored at -20°C for 30 minutes. The DNA was precipitated by centrifuging at 13,000 rpm for 10 minutes. The supernatant was poured off, and then 450 µL of 70% ethanol was added and centrifuged for 5 minutes. The liquid was removed, and the dry DNA was resuspended in 100 µL of dH2O.
The viral genome of PepYLCTHV was successfully detected using the polymerase chain reaction (PCR) with specific PepYLCTHV primers. The PCR reaction mixture consisted of 1 μl of DNA template, 1 μl of reverse and forward primers, 1 μl of 50 mM MgC
l2, 2.5 μl of 10x PCR buffer, 0.1 μl of Taq DNA polymerase, and 18.4 μl of dH
2O. The PCR cycle included an initial denaturation at 94 °C for 1 min, followed by 30 cycles of amplification, with each cycle consisting of 94 °C for 1 s, 55 °C for 2 s, and 72 °C for 2 min. Finally, there was a final extension period at 72 °C for 10 min [
18]. The PCR products were then electrophoresed on a 0.8% agarose gel and stained with ethidium bromide for visualization.
3. Results
3.1. Resistance of Chili Germplasms to PepYLCTHV
A total of 930 individual plants of 31 chili pepper genotypes were screened against Pepper yellow leaf curl Thailand virus (PepYLCTHV). Upon free-choice whitefly inoculation, all genotypes exhibited disease symptoms ranged from 0 to 5 scores. Disease severity progressed during 14, 21, and 28 days after inoculation (DAI), with the first symptom appearing at 14 DAI. At 21 DAI, the susceptible check (PEP31) and genotypes PEP15, PEP23, and PEP25 displayed the highest average disease score of 5, while others ranged from 2.90 to 4.96. On final scoring date, among advanced breeding lines (PEP1, PEP2, PEP3, PEP4, PEP5, PEP6, PEP7, PEP8, PEP9, PEP10, and PEP11, PEP26, PEP27, PEP28, PEP29, and PEP30, disease scores ranged from 3.46 to 5.0 and disease indices ranged from 69.33% to 100%. Pure-line chili pepper genotypes sourced from the World Vegetable Center (WorldVeg.) (PEP12, PEP13, PEP14, PEP15, PEP16, PEP17, PEP18, and PEP19) had disease scores ranging from 3.83 to 5.0, and disease indices from 76.67% to 100%. Thailand's commercial chili genotypes (PEP20, PEP21, PEP22, PEP23, PEP24, and PEP25) showed the highest disease scores ranging from 4.96 to 5.0 with disease indices ranging from 99.33% to 100%. Notably, two chili genotypes, PEP6 (advance breeding line) and PEP12 (WorldVeg.), exhibited the lowest average disease scores of 3.46 and 3.83, respectively, with disease indices of 69.33% and 76.67% (
Table 3).
2.3.2. Number of Resistance Plants and Virus Detection
Out of the 31 chili genotypes studied at the first screening cycle, their response to PepYLCTHV was assessed across six score levels. Among them, 13 genotypes displayed symptoms (score 1-5) and no symptoms (score 0). Based on PCR analysis, all inoculated plants were infected with the virus. Plants with a disease score of 0 were further categorized into two groups: those with virus detection (0+) and those without (0-), while scores 1-5 were identified as susceptible to virus detection (
Table 4). Among the genotypes, PEP6 and PEP12 exhibited more resistant plants (no symptoms) than others. Specifically, PEP6 had 9 plants classified as 0+. Conversely, 5 plants of PEP12 were classified as 0+ and 2 as 0-. Additionally, various plants from different genotypes, such as PEP2, PEP3, PEP5, PEP7, PEP8, PEP9, PEP10, PEP11, PEP12, PEP13, PEP14, and PEP19, displayed varying levels of 0+ (ranging from 1 to 4 plants). The genotypes PEP1, PEP4, PEP16, PEP18, PEP20, PEP21, PEP22, PEP23, PEP24, PEP25, PEP26, PEP27, PEP28, and PEP31 were found to be entirely susceptible to PepYLCTHV.
5.3.3. Effective Selection of Resistance to PepYLCTHV
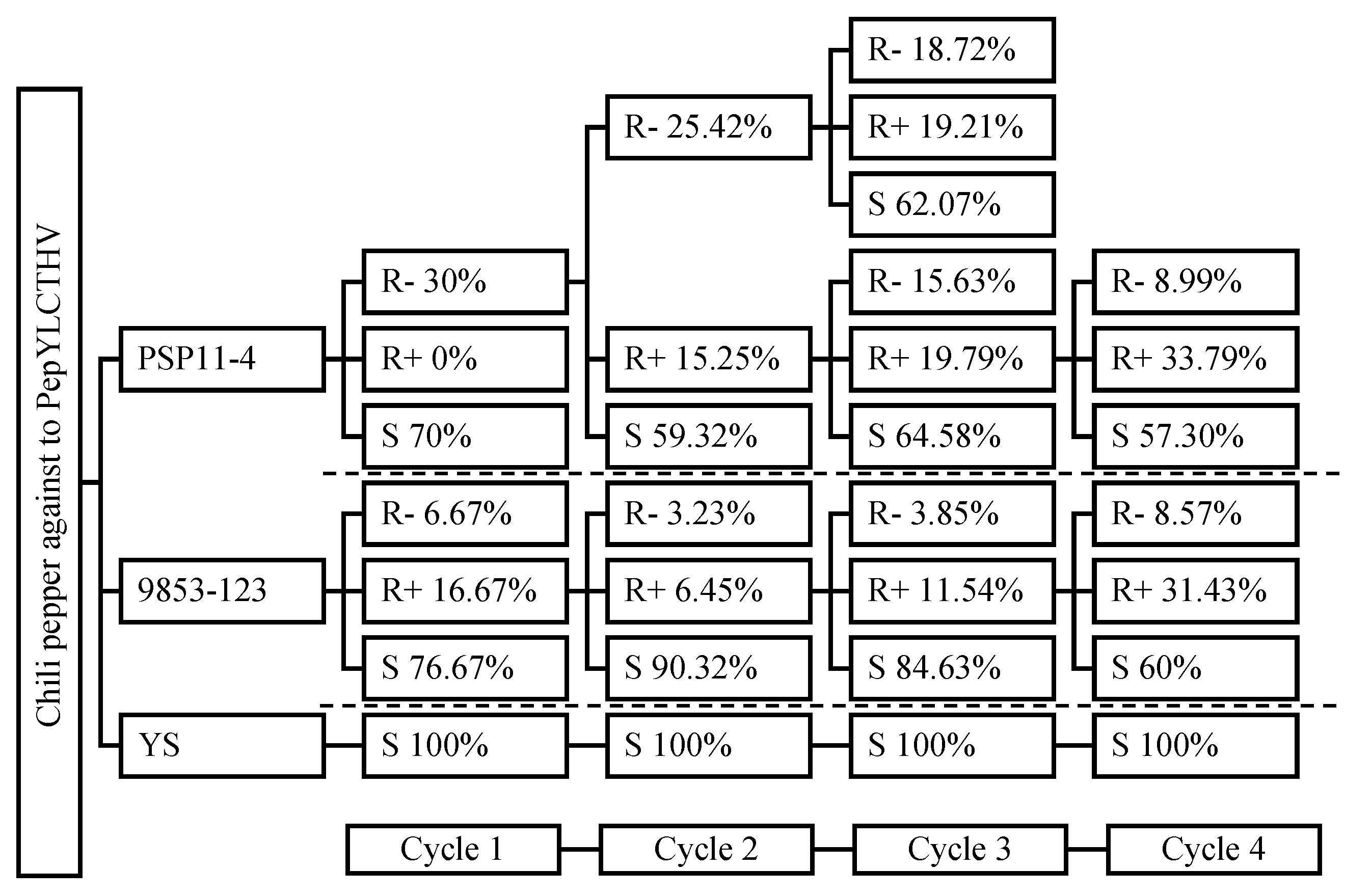
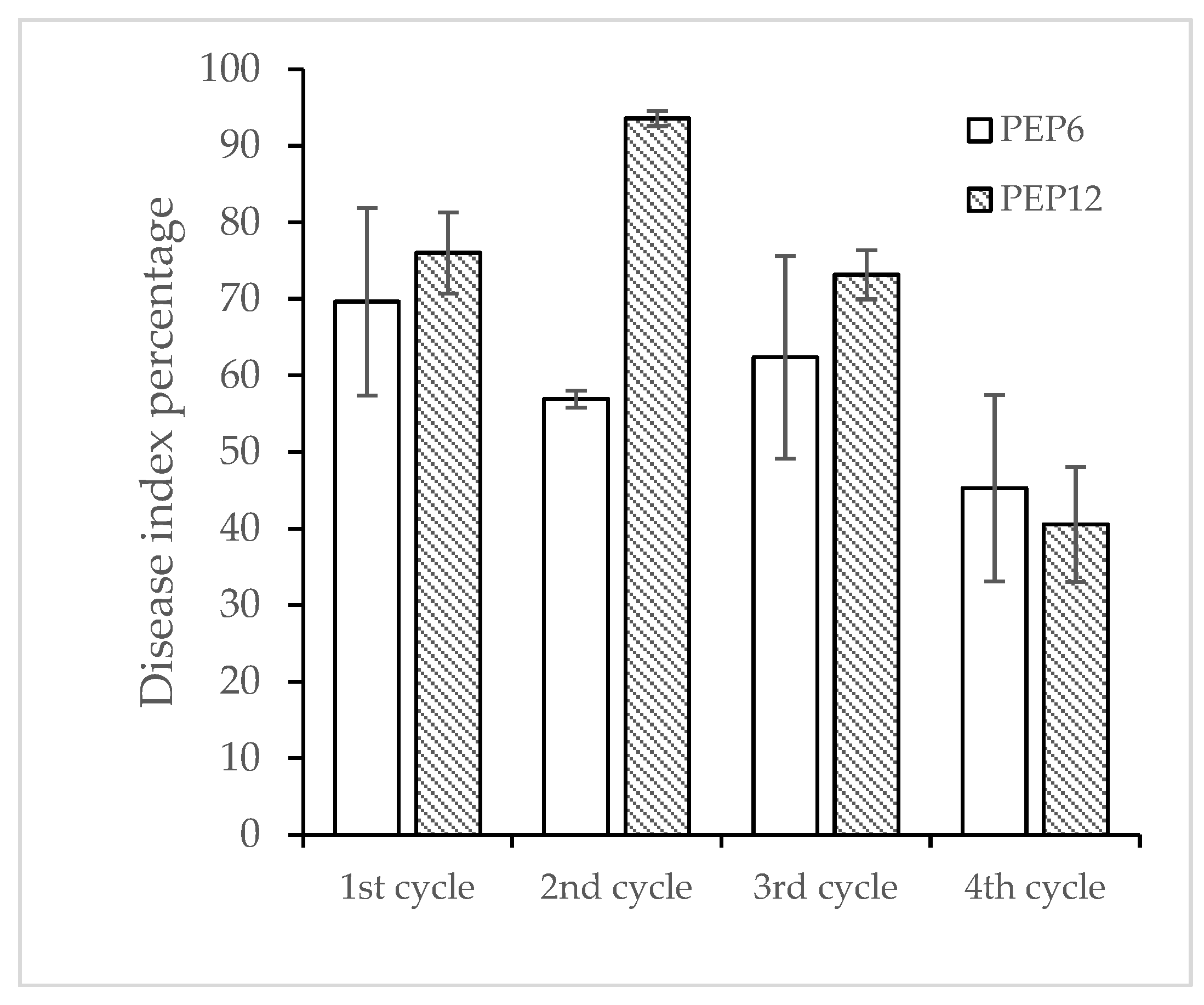
The response to selection for resistance to PepYLCTHV in PEP6 and PEP12 through four advanced cycles is described in
Figure 3. The percentage of resistance to the disease in both genotypes showed a slight increase from the first to the fourth cycle (
Figure 4). Resistance in PEP6 increased from 30% to 42.78%, and in PEP12 from 23.34% to 40%, while susceptibility remained at 100% for the susceptible plants. In the initial screening cycle, PEP6 exhibited a disease response score of 0- refer to R-, whereas PEP12 displayed 0+ refer to R+, and R-. Plants with the resistant phenotypes R- from PEP6 and R+ from PEP12 were subsequently selfed to assess the disease response within the R- and R+ groups. The resistance of PEP6 (R-) and PEP12 (R+) in the 3
rd and 4
th exhibited similar segregated responses across three groups: R-, R+, and S (
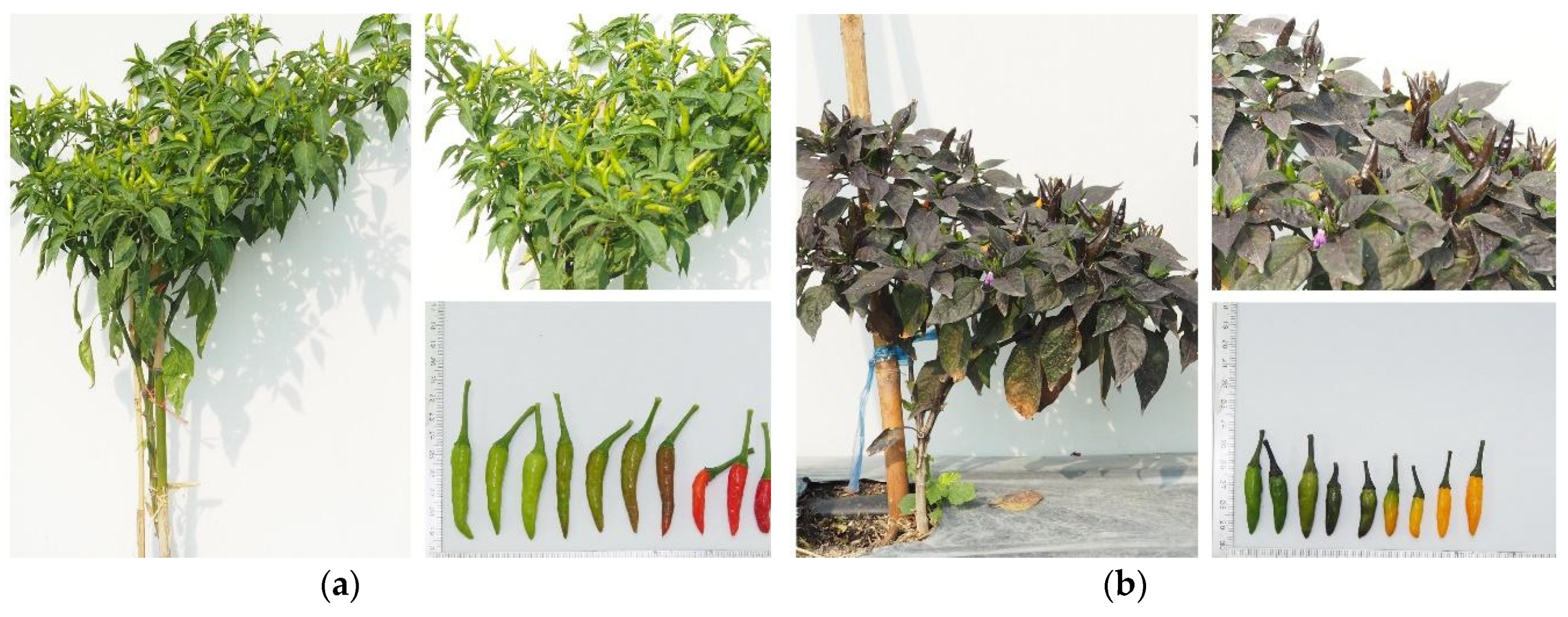
Figure 3). In addition, the morphological characterization of selected chili pepper lines resistant to PepYLCTHV PEP6 and PEP12 was described (
Figure 5). They were
C. annuum L. and showed differences in plant growth, leaf shape, leaf color, flower color, fruit shape, and fruit color. PEP6 was observed to have a green color in plant stem, leaf, and immature fruit, while PEP12 showed a purple color.
4. Discussion
We discovered that two out of thirty-one lines exhibited the highest resistance to PepYLCTHV when subjected to viruliferous whitefly inoculation. One of these resistant lines, PEP6, originated from The World Vegetable Center, Taiwan, was found to be highly resistant under field screenings at Khon Kaen University (KKU) and moderately resistant after graft screenings [
12]. This accession has been found to be highly resistance for the first time after whitefly artificial inoculation. Additionally, we identified PEP12 (an advanced generation derived from the 9853-123) as resistant to PepYLCTHV as Both absence (immune) and presence (tolerant) of the virus were observed in symptomless plants. PEP12 also demonstrated PepYLCV resistance in field net-house screenings by viruliferous whiteflies at 120 days after inoculation (DAI) and was highly resistant in graft screenings [
12]. We observed differences in the responses of individual resistant plants, categorized as R- (virus absent or immune) and R+ (virus present or tolerant). The R- resistance could be attributed to mechanisms such as non-preference by whiteflies, including pre-existing mechanisms like wax, hair, leaf color, and trichomes which initially protect the plant [
19,
20]. For example, chilli genotypes with a high density of glandular trichomes were associated with the whitefly non-preference [
21].The R- resistance may involve the elimination of pathogens from plant cells, known as hypersensitivity. This mechanism entails the production of thickened cell walls, increased cuticle layers, higher trichome density, and the accumulation of secondary products in plants, which collectively deter pests or inhibit pathogen growth, potentially leading to unsuccessful infection expansion [
22,
23]. In the case of R+ responses, plants may exhibit a local reaction involving the release of molecules triggered by the virus (elicitor), or they may interact with specific proteins produced by both the virus and the plants, such as the nuclear shuttle protein (NSP) and movement protein (MP) These responses can be suppressed by specific proteins or secondary metabolites from resistant host plants, resulting in the restriction or delay of infection [
24].
We demonstrated mass selection is efficient in enhancing resistance to PepYLCTHV in two chili inbred lines. By the fourth cycle of selection, the resistance level increased to a maximum ratio of 1:1 of resistance to susceptibility, while the agronomic traits and phenotypic appearance were uniform. This evidence suggests that only resistance to PepYLCTHV segregates, possibly indicating that the resistant gene is controlled by more than two genes [
25] with one position showing resistance in heterozygous form, likely due to the presence of a lethal gene. The evidence presented elucidated the role of the chlorophyll formation gene in rice leaf pigments, where distinct green (heterozygous) and yellow (homozygous) colors were segregated at a 1:1 ratio among self-progenies from S2 to S6 [
26]. In this study, PEP6 and PEP12 were identified as potential improve sources of PepYLCTHV, which could be used to develop commercial cultivars resistant to PepYLCTHV disease. Future research is underway to understand the genetic mechanisms underlying resistance to PepYLCTHV and enhance our understanding.
Author Contributions
Conceptualization, N.K., S.W., O.C. and S.T.; methodology, N.K., S.W., T.T, W.T. and N.J.; software, N.K. and S.W.; validation, N.K., S.W. and S.Ku.; formal analysis, N.K. and S.W.; writing—original draft preparation, N.K. and S.W.; writing—review and editing, N.K., S.W., S.Kr. and S.Ku.; visualization, N.K. and S.W.; supervision, M.T., O.C. and S.T. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by King’s Mongkut Institute of Technology Ladkrabang (KMITL) and the National Science and Technology Development Agency (NSTDA) (FDA-CO-2562-9606-TH).
Acknowledgments
This work was supported by King’s Mongkut Institute of Technology Ladkrabang. We wish to thank Kasetsart University Kamphaeng Saen Campus for experimental facilities support and the Faculty of Agriculture, Khon Kaen University and the World Vegetable Center, Taiwan for providing germplasm.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Food and agriculture organization of the United Nations. 2022.
- Bhutia, K.V. Khanna & Ngasepam, Tombisana & Dolma, Nangsol. Effects Of Climate Change On Growth And Development Of Chilli. Agrotechnology 2018, 7. [CrossRef]
- Prabaningrum, L.M., Tonny & Hasyim, A. & Setiawati, Wiwin & Murtiningsih, Rini & Udiarto, B.K. & Sulastrini, Ineu & Korlina, Eli & Gunaeni, Neni & Wulandari, A.W. & Gunadi, N. & Priani, R.A. & Lukman, L. & Mejaya, Made. Diversity of insect pests and their natural enemies in hot pepper (Capsicum annuum L.) ecosystem of Indonesia. Applied Ecology and Environmental Research 2022, 20, 3367–3377. [CrossRef]
- Nalla, M.S., Roland & Pappu, H. & Barchenger, Derek. Current status, breeding strategies and future prospects for managing chilli leaf curl virus disease and associated begomoviruses in Chilli (Capsicum spp.). Frontiers in Plant Science 2023, 14. [CrossRef] [PubMed]
- Chiemsombat, P.Y., Sopana & Srikamphung, Bunnada. Begomoviruses Associated to Pepper Yellow Leaf Curl Disease in Thailand. Begomoviruses Associated to Pepper Yellow Leaf Curl Disease in Thailand. J Agri Res 2018, 3, 000183. [CrossRef]
- Hidayat, S.H.; Chatchawankanpanich, O.; Rusli, E.; Aidawati, N. Begomovirus Associated with Pepper Yellow Leaf Curl Disease in West Jva, Indonesia. Jurnal Mikrobiologi Indonesia 2006, 11, 87–90. [Google Scholar]
- Kenyon, L.; Kumar, S.; Tsai, W.-S.; Jacqueline d, A.H. Virus Diseases of Peppers (Capsicum spp.) and Their Control. Adv Virus Res 2014, 90, 297–354. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.S.; SK, R.; Prasad, V.; Singh, V. Recent Research Findings Related to Management Strategies of Begomoviruses. Plant Pathology & Microbiology 2015, 6. [Google Scholar] [CrossRef]
- Thakur, H.; Jindal, S.K.; Sharma, A.; Dhaliwal, M.S. A monogenic dominant resistance for leaf curl virus disease in chilli pepper (Capsicum annuum L.). Crop Protection 2019, 116, 115–120. [Google Scholar] [CrossRef]
- Pandey, V.; Srivastava, A.; Shahmohammadi, N.; Nehra, C.; Nehra, R.; Guar, R.K.; Golnaraghi, A. Begomovirus Exploiting the Host Machinery for Their Survival. Journal of Modern Agriculture and Biotechnology 2022, 2. [Google Scholar] [CrossRef]
- Barchenger, D.Y., Sopana & Jeeatid, Nakarin & Lin, Susan & Wang, Yen-Wei & Lin, Tsung-han & Chan, Yuan-Li & Kenyon, Lawrence. A Novel Source of Resistance to Pepper yellow leaf curl Thailand virus (PepYLCThV) (Begomovirus) in Chile Pepper. Hortscience 2019, 54, 2146–2149. [CrossRef]
- Suwor, P.; Masirayanan, T.; Khingkumoungk, H.; Tsai, W.S.; Saetiew, K.; Techawongstien, S.; Kumar, S.; Kramchote, S. Chili (Capsicum annuum L.) genotypes resistant to Pepper yellow leaf curl Thailand virus (PepYLCTHV). Journal of Plant Protection Research 2021, 61, 377–383. [Google Scholar] [CrossRef]
- Srivastava, A.; Mangal, M.; Saritha, R.K.; Kalia, P. Screening of chilli pepper (Capsicum spp.) lines for resistance to the begomoviruses causing chilli leaf curl disease in India. Crop Protection 2017, 100, 177–185. [Google Scholar] [CrossRef]
- Koeda, S.; Mika Onouchi; Namiko Mori; Nadya Syafira Pohan; Atsushi J. Nagano; Elly Kesumawati. A recessive gene pepy-1 encoding Pelota confers resistance to begomovirus isolates of PepYLCIV and PepYLCAV in Capsicum annuum. Theoretical and applied genetic 2021, 134, 2497–2964. [CrossRef] [PubMed]
- Dwivedi, N.; Mishra, M.; Sharma, S.S.; Singh, R.K. Genetic analysis and QTLs identification for resistance to the Begomovirus causing pepper leaf curl virus (PepLCV) disease. Journal of Plant Biochemistry and Biotechnology 2023. [Google Scholar] [CrossRef]
- Acquaah, G. Principle of plant genetics and breeding, 2nd ed.; Jonh Willey & Sons, Ltd: West susex. UK, 2012; p. 740. [Google Scholar]
- Padilha, H.K.M.; Barbieri, R.L. Plant breeding of chili peppers (Capsicum, Solanaceae). Australian Journal of Basic and Applied Science 2016, 10, 148–154. [Google Scholar]
- Tsai, W.S.; Shih, S.L.; Kenyon, L.; Green, S.K.; Jan, F.-J. Temporal distribution and pathogenicity of the predominant tomato-infecting begomoviruses in Taiwan. Plant Pathology 2011, 60, 787–799. [Google Scholar] [CrossRef]
- Jeevanandham, N.M., Murugan & Natesan, Senthil & Gandhi, Karthikeyan & Appachi, Sathiyamurthy. Plant resistance in chillies Capsicum spp against whitefly, Bemisia tabaci under field and greenhouse condition. Journal of Entomology and Zoology Studies 2018, 6, 1904–1914.
- Sandra, Y.M.A.; Awang Maharijaya; Sobir. Screening of resistance to geminivirus and whitefly in pepper. Euphytica 2022, 218. [CrossRef]
- Yadav, R.K.J., Pagadala Damodaram & Kumar, Manish & Parepely, Saravan & Rao, Vala & Kambham, Madhavi. Screening chilli genotypes for whitefly (Bemisia tabaci Genn.) resistance: A vector for chilli leaf curl virus. International Journal of Chemical Studies 2020, 8, 971–979. [CrossRef]
- Slater, A.; Fowler, M.R.; Scott, N.W. Plant biotechnology: the genetic manipulation of plant; Oxford University Press: 2003; p. 346.
- Hasanuzzaman, A.T.M.; Islam, M.N.; Zhang, Y.; Zhang, C.-Y.; Liu, T.-X. Leaf Morphological Characters Can Be a Factor for Intra-Varietal Preference of Whitefly Bemisia tabaci (Hemiptera: Aleyrodidae) among Eggplant Varieties. PLOS ONE 2016, 11. [Google Scholar] [CrossRef]
- Hanley-Bowdoin, L.; Bejarno, E.R.; Robertson, D.; Mansoor, S. Geminiviruses: masters at redirecting and reprogramming plant processes. Nat Rev Microbiol 2013, 11, 777–788. [Google Scholar] [CrossRef] [PubMed]
- Siddique, M.I.; Lee, J.-H.; Ahn, J.-H.; Kusumawardhani, M.K.; Safitri, R.; Harpenas, A.; Kwon, J.-K.; Kang, B.-C. Genotyping-by-sequencing-based QTL mapping reveals novel loci for Pepper yellow leaf curl virus (PepYLCV) resistance in Capsicum annuum. PLOS ONE 2022, 17, e0264026. [Google Scholar] [CrossRef] [PubMed]
- Katayama, Y.; Shida, S. Bahavior of balance lethal gene occurring in progeny of X-rayed rice. The Japanese Journal of Genetics 1968, 43, 129–136. [Google Scholar] [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).