1. Introduction
Stripe rust, caused by
Puccinia striiformis f. sp.
tritici (Pst), is one of the most destructive diseases of wheat worldwide [
1,
2]. The first stripe rust incursion in Eastern Australia occurred in 1979 [
3,
4] and in Western Australia in 2002 [
5]. Subsequently it caused several epidemics, including the most recent in 2022, with devastating impacts on the agricultural industry with approximate yield reduction of 84% [
6]. Now stripe rust pathogen has a history of 44 years in Australia and it still remains a major constraint for wheat production with an estimated average annual loss of A
$ 127 m [
7].
Although more than 80 loci for stripe rust resistance have been characterized and formally named [
https://wheat.pw.usda.gov/GG3/wgc; visited on July 21, 2023], most of these belong to the race-specific category. Resistance genes that condition ASR often succumb to the acquisition of virulence in pathogen populations. Wellings and McIntosh[
8] summarised the evolution of virulence in Pst population and concluded the stepwise attainment in virulence for stripe rust resistance genes
YrA,
Yr6 and
Yr7 individually and in different combinations. The other commercially important event included the detection of virulence for
Yr17 in 1999. All derivatives of the 1979 introduction (104 E137A-) were virulent on genotypes carrying
Yr3 and
Yr4. In contrast, the 2002 introduction (134 E16A+) and all its current derivatives are avirulent on
Yr3 and
Yr4. In addition, this group carried virulence for resistance genes
Yr8 and
Yr9 that have not been deployed widely in Australian wheat cultivars. Stripe rust resistance genes
Yr6,
Yr7,
Yr9,
Yr17 and
Yr27 have been postulated in spring wheat cultivars grown worldwide. Pathotypes carrying virulence for these have been reported in many countries including Australia.
All stage resistance (ASR) and adult plant resistance (APR) are two main categories of resistance to rust diseases in wheat. ASR genes protect plants throughout all growth stages, while APR genes provide post-seedling resistance. To achieve long-lasting control, it's essential to combine ASR and APR genes in a single cultivar which is now possible through advancement in molecular technologies. It has enabled efficient mapping of resistance genes and identification of closely linked markers using whole genome scanning [
9] using different high-throughput platforms such as Diversity Arrays Technology (DArT) [
10], genotyping-by-sequencing (GBS) [
11], selective genotyping (SG) [
12], and SNP-genotyping arrays [
13]. The development of ordered draft sequence of the 17-gigabase with more than 75,000 genes positioned along 21 chromosomes of wheat [
14] adds further impetus to mapping and marker development research. Bariana et al. [
15] identified genotypes carrying four to six QTL for stripe rust resistance and a genotype combining closely linked genes
Yr15 and
Yr24 were identified by Zakeri et al. [
16].
To achieve durable resistance to stripe rust, discovery and characterisation of diverse sources of resistance is essential. Bariana and co-workers screened the Watkins Collection of common wheat landraces [
17] and identified genotypes for detailed genetic analysis. Daetwyler et al. [
18] predicted the involvement of several chromosomes in conditioning stripe rust resistance among this collection of landraces. Two landraces AUS27507 and AUS27894 exhibited seedling resistance against a range of Pst pathotypes in Australia. This study covers genomic location of resistance carried by these genotypes and identification of closely linked markers with the underlying gene(s).
2. Materials and Methods
2.1. Plant Material
AUS27507 and AUS27894 from France and Spain, respectively, were crossed with a susceptible landrace AUS27229. F3 populations comprising 106 and 100 families were developed from AUS27507/AUS27229 and AUS27894/AUS27229 crosses. Only AUS27507/AUS27229 F3 was advanced to generate an F6 recombinant inbred line (RIL) population for further analysis. AUS27894 and AUS27507 were crossed to determine the allelic relationship of the gene(s) carried by these genotypes. Twenty-one Australian wheat cultivars were used to validate markers closely linked with the resistance gene identified in this study.
2.2. Greenhouse Tests
Twenty seeds of each AUS27507/AUS27229 F3 line were sown in 9 cm plastic pots filled with mixture of pine bark and river sand (2:1 ratio). Parental genotypes and the susceptible control Morocco were sown in each experiment. AUS27894/AUS27507 F3 families and AUS27507/AUS27229 F6 RIL population were evaluated against Pst pathotype 134 E16A+17+27+ (Plant Breeding Institute, Cobbitty culture no. 617).
2.3. Isolation of Genomic DNA
Leaf tissue was collected from 12 day-old seedlings in 2 ml Eppendorf tubes and was dried on silica gel for 72 hours. Genomic DNA was extracted using the modified CTAB method [
19] and quantified using Nano Drop spectrophotometer (NanoDrop® ND1000).
2.4. Bulked Segregant Analysis
Bulked segregant analysis (BSA) was performed on resistant and susceptible bulks prepared by pooling equal amount of DNA from 20 homozygous resistant (HR) and 20 homozygous susceptible (HS) F
3 lines, respectively. The F
1 artificial bulk was prepared combining an equal amount of DNA from rest of the population. DNA of bulks and F
1 were sent to Diversity Arrays Technology Canberra for BSA using high density DArT array Wheat
PstI (
TaqI) 3 (
http://www.diversityarrays.com).
2.5. STS and SSR Genotyping
Sequence of resistance linked DArT clones (kindly provided Dr A. Kilian) was used to develop STS markers using the Primer3 program [
22] (
http://primer3.sourceforge.net/). In addition, 52 SSR markers previously mapped on chromosome 2BL (
http://wheat.pw.usda.gov) were also genotyped to enrich the genetic map [
19,
20,
21]. PCRs were carried out in 10 µl reaction volume containing 2 µl of 30 ng/µl DNA, 1 µl of 1X PCR buffer containing MgCl
2, 0.75 µl of dNTPs, 0.4 µl of forward (1.25 mM) and reverse (5.0 mM) M13 tagged primers, 0.1 µl (0.5mM) of M13 fluorescent tag and 0.04 µl of Taq DNA polymerase. PCR amplification conditions described in Randhawa et al. [
22] were used. Markers polymorphic on parents and showing strong linkage among HR and HS lines were evaluated on the entire AUS27507/AUS27229 RIL population. Electrophoresis of PCR products was carried out on 2.5-3% agarose (Amresco) gel stained with GelRed™ (Biotium). PCR products were visualized using UV gel documentation system. Markers which could not be differentiated on agarose gel were separated on polyacrylamide gel using Analyzer Gene ReadIR 4300, Li-COR sequencing system (Li-COR Bio-sciences, USA) after denaturing PCR product at 95 °C for five minutes.
2.6. SNP Genotyping
Twenty-seven SNPs showed strong linkage with the stripe rust resistance locus in the AUS27507 AUS27894. Closely linked SNPs (
IWA5694,
IWA5839,
IWA6130,
IWA6334,
IWA6417,
IWA7265,
IWB10417,
IWB10455,
IWB12294,
IWB20875,
IWB21638,
IWB23209,
IWB24984,
IWB25015,
IWB28191,
IWB31823,
IWB33668,
IWB43166,
IWB45530,
IWB45652,
IWB49793,
IWB49793,
IWB62498,
IWB62757,
IWB62762,
IWB69000 and
IWB79078) were used to design kompetitive allele specific PCR assays (KASP) with two allele-specific and one common primer. These markers were genotyped on the parents and the CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad) was used for comparing KASP amplifications among different genotypes following the protocol given at LGC website (
http://www.ksre.ksu.edu/igenomics/doc1363.ashx).
2.7. Deletion Bin Mapping of Linked Markers
The DNA samples of chromosome 2BL deletion stocks [del2BL-9, del2BL-3, del2BL-7, del2BL-5 and del2BL-6 [
23]; kindly provided by Dr Evans Lagudah were used to confirm the location of resistance gene. Chinese Spring (CS) was used as control.
2.8. Data Analysis
Goodness of fit of observed segregation data to the expected genetic ratios was tested through Chi-squared analysis. Linkage map was constructed using Mapmaker version 3.0 [
24] and recombination fractions were transformed to centi Morgans (cM) using Kosambi mapping function [
25]. Final genetic linkage map was constructed using MapChart [
26].
3. Results
3.1. Inheritance Studies
Seedling stripe rust responses of AUS27507 and AUS27894 varied from 2C to 3C (
Figure 1), when evaluated against Pst pathotype 134 E16A+17+27+. Stripe rust screening results of F
3 populations derived from crosses AUS27507/AUS27229 and AUS27894/AUS27229 are presented in
Table 1. Both populations showed monogenic inheritance of stripe rust resistance. Both parents and homozygous resistant F
3 lines produced similar stripe rust responses suggesting the presence same gene. AUS27507 and AUS27894 were crossed and F
3 families were generated. All 80 F
3 families produced responses similar to parents confirming presence of the same gene in both genotypes. The resistance locus segregating in both populations was temporarily named
YrAW4. The AUS27507/AUS27229 derived population was advanced to generate F
6 RIL population. Segregation for
YrAW4 among the RIL population was confirmed (
Table 1).
3.2. Molecular Mapping
Twenty homozygous resistant and homozygous susceptible lines each from AUS27507/AUS27229 F
3 population were used to construct contrasting bulks. These bulks were subjected to BSA using DArT markers. DArT markers
wPt-2397, wPt-665550, wPt-7161, wPt-8916, wPt-1722, wPt-4197, wPt-6542, wPt-9104.2, wPt1650.2, wPt-2185, wPt-0510, wPt-0948, wPt-3632, wPt-7350, wPt-9104.1, wPt-9190, and wPt-9336 showed linkage with
YrAW4. All these markers are located in chromosome 2BL. Linked DArT clones were converted to STS markers and tested on the AUS27507/AUS27229 RIL population. The STS primers were named as “sun” (Sydney University). Four STS markers
sun483 (
wPt-2397),
sun481 (
wPt-665550),
sun482 (
wPt-7161) and
sun484 (
wPt-8916) showed polymorphism between parents and were genotyped on the entire RIL population. Markers
sun481 and
sun482 flanked
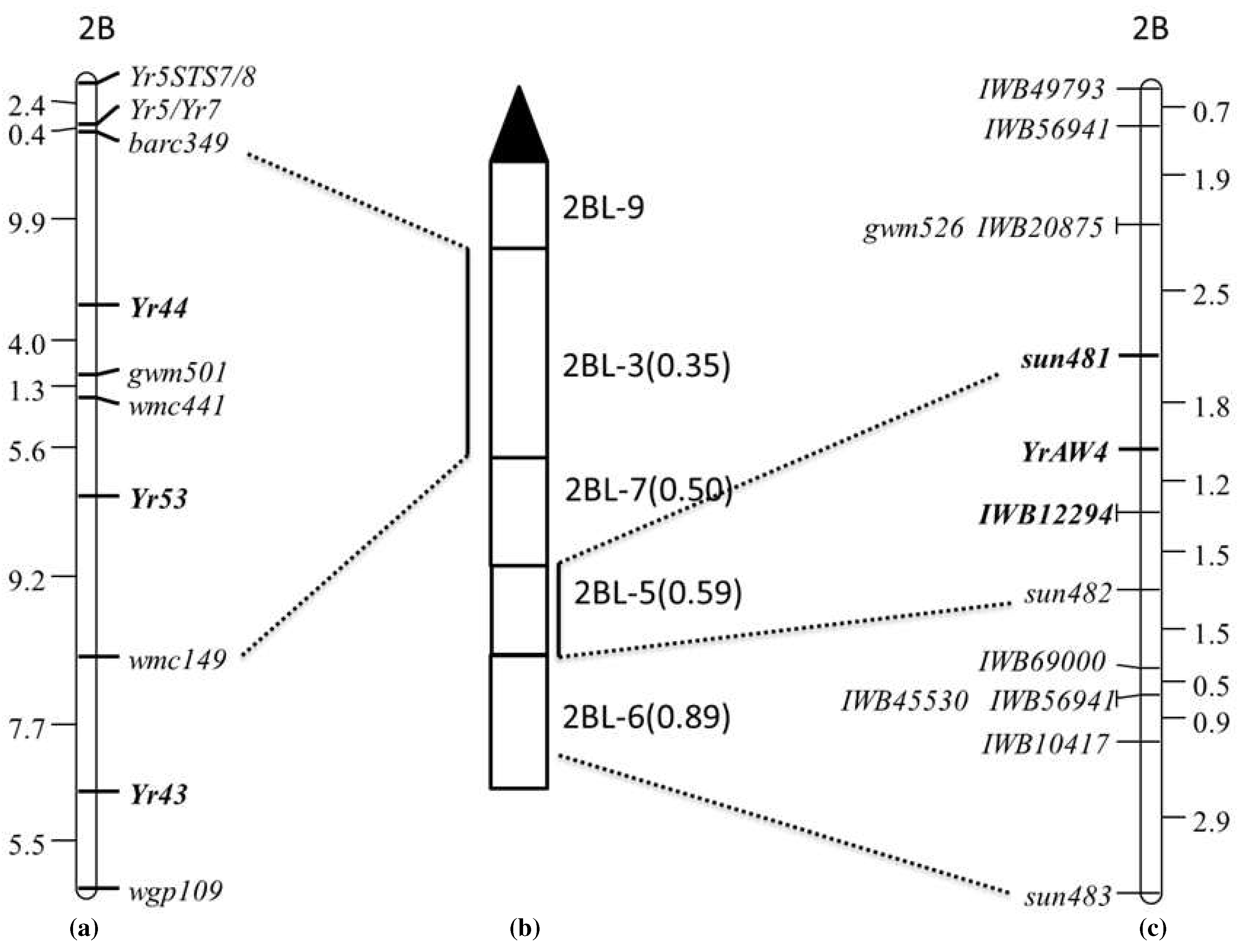
YrAW4 at genetic distances of 1.8 cM (proximally) and 2.7 cM (distally), respectively (
Figure 2).
3.3. Deletion Mapping of Flanking Markers
YrAW4-linked STS markers
sun481 and
sun482 were genotyped on DNA of the 2BL deletion stocks: del2BL-9, del2BL-3, del2BL-7, del2BL-5 and del2BL-6 to determine their precise genomic locations. Chinese Spring, AUS27507, AUS27894 (resistant parents) and AUS27229 (susceptible parent) were used as controls. The amplicons produced by deletion stocks were compared with Chinese Spring. Marker
sun482 amplified the expected product in del 2BL-5, del2BL-6 and CS, whereas
sun481 amplified the target band only in del 2BL-5 and CS. These results placed the closest marker
sun481 in the deletion bin 2BL-5 (
Figure 2b,c).
3.4. Saturation of Chromosome 2BL using SNP Markers
AUS27507/AUS27229 RIL population was genotyped using 90K Infinium SNP wheat array. Twenty-one SNPs markers on chromosome 2BL mapped close to
YrAW4 and spanned 759 to 784 Mbp region in physical map of Chinese Spring (IWGSC RefSeq_V2.0). These were converted into KASP assays, and those showing parental polymorphisms were genotyped on the entire population (
Figure 2). Eight KASP markers were incorporated into the AUS27507/AUS27229 chromossome 2BL map (
Table 3,
Figure 2). The final map included 1 SSR, 3 STS and 8 KASP markers (
Figure 2c). The SNP marker
IWB12294 mapped 1.2 cM distal to
YrAW4 and 1.5 proximal to
sun482 (
Figure 2). Sequences of the primers for markers included in the final map are given in
Table 2 and
Table 3.
3.5. Validation of YrAW4 Linked Markers
A set of Australian common wheat and durum wheat cultivars was tested with flanking markers
IWB12294 and
sun481.
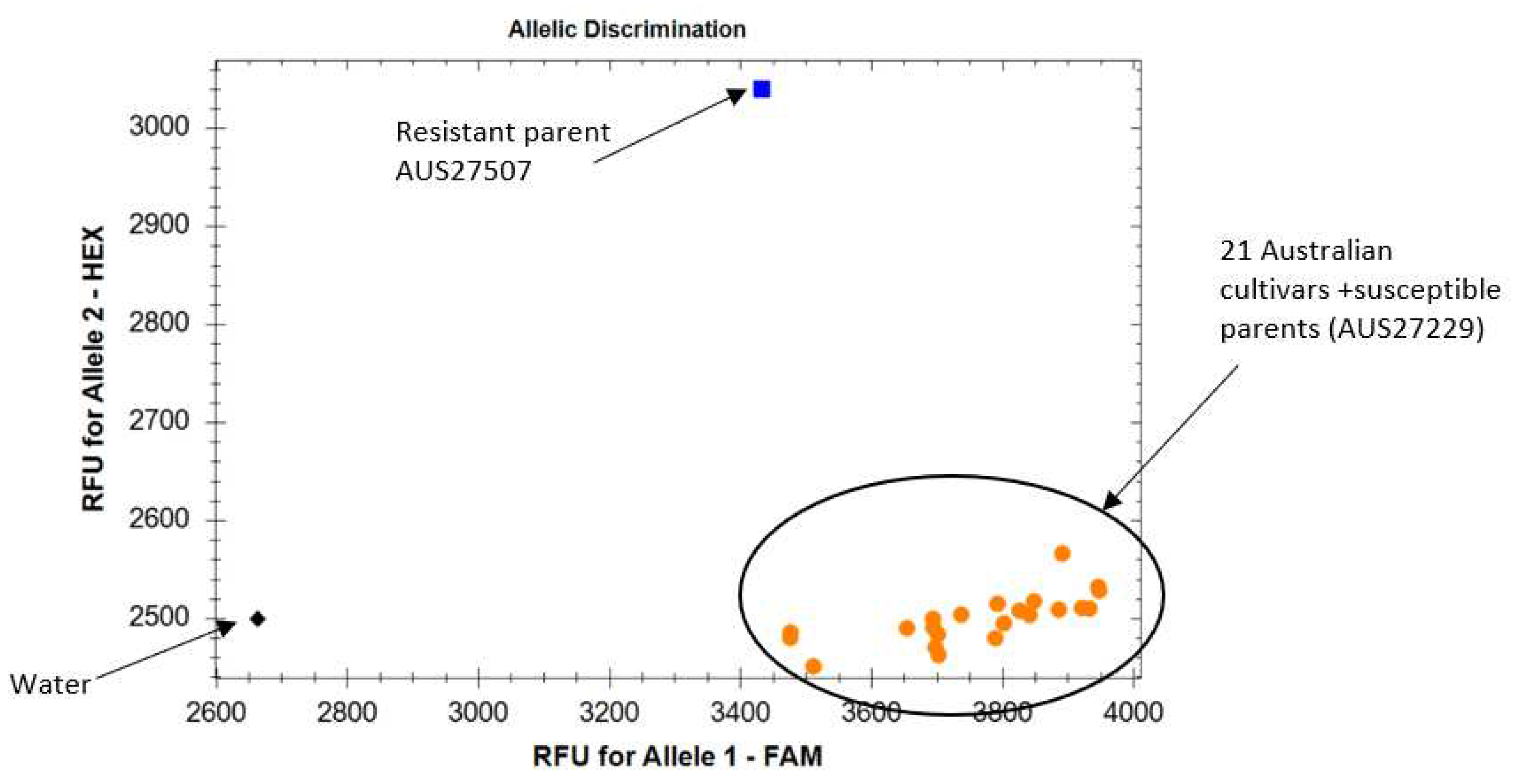
IWB12294 produced SNP allele ‘G’ in the resistant parent AUS27507 and allele ‘T’ in susceptible parent AUS27229 (
Table 4 and
Figure 3). SSR marker amplified
YrAW4 linked
sun481240bp and
sun481100bp allele in resistant and susceptible parents, respectively (
Table 4). Amplification of the ‘T’ allele and the
sun481100bp allele in all test cultivars confirmed the usefulness of these markers in marker assisted selection of
YrAW4 in breeding programs.
4. Discussion
This study identified a new stripe rust resistance locus in landraces AUS27507 and AUS27894. Different genomic resources were used to determine the genomic location and to identify closely linked markers for marker assisted selection of this gene. Absence of segregation among AUS27894/AUS27507 F3 population demonstrated presence of the same gene in both genotypes. The resistance locus was temporarily named
YrAW4. DArT based BSA indicated the location of
YrAW4 in chromosome 2BL and deletion mapping of flanking DArT markers refined the location of
YrAW4 in the deletion bin 2BL-5. Markers
sun481 and
sun482 mapped 1.8 cM proximal and 2.7 cM distal to
YrAW4, respectively. The target genomic region was enriched by converting
YrAW4-linked SNP markers to KASP assays. Marker
IWB12294 mapped 1.2 cM distal to
YrAW4 and 1.5 cM proximal to
sun482 (
Figure 2c).
Seedling stripe rust resistance genes
Yr5, Yr7, YrSp, Yr43, Yr44 and
Yr53 were previously reported in the long arm of chromosome 2B (2BL) [
https://wheat.pw.usda.gov/GG3/wgc].
Yr5 and
Yr7 were mapped 21cM away from centromere.
Yr7 was proved as an allele of
Yr5 [
27].
Yr5 produce IT from 0; to; against Australian Pst pathotype 134 E16A+ and its derivatives and
Yr7 is not effective against this group of pathotypes and hence
YrAW4 cannot be either of these genes. Moreover,
Yr5-linked marker,
Yr5STS7/8 [
28], Digenic segregation among
YrAW4/
YrSp F2 population (H.S. Bariana unpublished results) also excluded the possibility of
YrAW4 to be
YrSp.
Yr43 [
29]
, Yr44 [
30] and
Yr53 [
30] produce IT 2 on a 0 to 9 scale, which is much lower than IT 2C-3C (on a 0 to 4 scale) produced by
YrAW4. Xu et al. [
30] placed flanking markers for
Yr43,
Yr44 and
Yr53 in the chromosomal deletion bin 2BL-3, which is next to the centromeric deletion bin 2BL-9 (
Figure 2a, 2b). The
YrAW4-linked marker
sun481 was mapped in the deletion bin 2BL-5. Based on infection type and deletion bin location, we believe that
YrAW4 is a unique locus and therefore,
YrAW4 was formally named
Yr72.
The usefulness of markers closely linked with the resistance gene requires validation across potential backgrounds in which the target gene has to be transferred. Presence of the resistance-linked allele in genotypes carrying the target gene is referred to as ‘positive validation’ and the absence/amplification of an alternate allele in cultivars lacking the target locus is called ‘negative validation’. Since
YrAW4 has not yet been deployed in modern cultivars, ‘negative validation’ was performed. Markers
sun481 and
IWB12294 were negatively validated among a set of Australian wheat cultivars including two durum wheat genotypes (
Table 4 and
Figure 3). These robust markers can be used (
Table 2 and
Table 3) for marker assisted selection of
Yr72 in these backgrounds.
Markers linked with genes
Yr15 [
31],
Yr51 [
22] and
Yr57 [
32] that are effective against currently predominant Pst pathotypes and race non-specific APR genes
Yr18 [
33],
Yr36 [
34] and
Yr46 [
35] are available to deploy different combinations of these genes in future wheat cultivars. SNP markers linked with economic traits are preferred by breeding companies for their amenability for high-throughput testing.
YrAW4 linked SNP marker
IWB12294 offers this opportunity (
Table 3) for marker assisted selection and marker-assisted pyramiding of this gene with other ASR and APR genes.
YrAW4 is currently being transferred to leading Australian cultivars through marker assisted selection.
Author Contributions
This study was planned by H.B. and U.B. and H.B., U.B. and H.M. created the mapping population. H.B and M.C. performed phenotyping in the greenhouse. M.H. and D.W. carried out bulk segregant analysis. U.B. and M.C. developed STS and KASP markers and mapped those on the RIL population. The manuscript was ddrafted by M.C., U.B and H.B and was read and approved by all authors.
Funding
Financial support from the Australian Government (scholarship) via the Australia Awards program, University of Sydney and the GRDC Australia is gratefully acknowledged.
Institutional Review Board Statement
Not applicable involving wheat.
Informed Consent Statement
Not applicable as this study did not involve humans.
Data Availability Statement
All data included in this publication.
Acknowledgments
The first author acknowledges the Royal Government of Bhutan, Ministry of Agriculture and Forestry (MoAF), for granting study leave and the Australian Government for the Australian Leadership Award to pursue PhD studies at the University of Sydney. We are thankful to Dr. Evans Lagudah for providing the deletion stocks.
Conflicts of Interest
The authors declare that they have no conflict of interest.
References
- Wellings, C.R. Global status of stripe rust: a review of historical and current threats. Euphytica 2011, 179, 129-141. [CrossRef]
- Stubbs, R.W. Stripe rust. In The Cereal Rusts Vol Il, Roelfs, A.P., Bushnell, W.R., Eds.; Academic Press: Orlando, Florida, USA, 1985; pp. 61-101.
- O'Brien, L.; Brown, J.S.; Young, R.M.; Pascoe, I. Occurrence and Distribution of Wheat Stripe Rust in Victoria and Susceptibility of Commercial Wheat Cultivars. Australasian Plant Pathology 1980, 9, 14. [CrossRef]
- O'Brien, L.; Brown, J.S.; Young, R.M.; Pascoe, I. Occurrence and distribution of wheat stripe rust in Victoria and susceptibility of commercial wheat cultivars. Aust Plant Pathol 1980, 9, 14. [CrossRef]
- Wellings, C.R.; Wright, D.G.; Keiper, F.; Loughman, R. First detection of wheat stripe rust in Western Australia: evidence for a foreign incursion. Australasian Plant Pathology 2003, 32, 321-322. [CrossRef]
- Murray, G.M.; Ellison, P.J.; Watson, A. Effects of stripe rust on the wheat plant. Australasian Plant Pathology 1995, 24, 261-270, doi:Doi 10.1071/App9950261.
- Murray, G.M.; Brennan, J.P. Estimating disease losses to the Australian wheat industry. Australasian Plant Pathology 2009, 38, 558-570, doi:Doi 10.1071/App9950261.8.
- Wellings, C.R.; McIntosh, R.A. Puccinia striiformis f.sp. tritici in Australasia: pathogenic changes during the first 10 years. Plant Pathol 1990, 39, 316-325. [CrossRef]
- Edae, E.A.; Olivera, P.D.; Jin, Y.; Poland, J.A.; Rouse, M.N. Genotype-by-sequencing facilitates genetic mapping of a stem rust resistance locus in Aegilops umbellulata, a wild relative of cultivated wheat. BMC genomics 2016, 17, 1039. [CrossRef]
- Akbari, M.; Wenzl, P.; Caig, V.; Carling, J.; Xia, L.; Yang, S.; Uszynski, G.; Mohler, V.; Lehmensiek, A.; Kuchel, H.; et al. Diversity arrays technology (DArT) for high-throughput profiling of the hexaploid wheat genome. Theor Appl Genet 2006, 113, 1409-1420. [CrossRef]
- Jaccoud, D.; Peng, K.; Feinstein, D.; Kilian, A. Diversity arrays: a solid state technology for sequence information independent genotyping. Nucleic Acids Res 2001, 29, E25. [CrossRef]
- Xue, S.; Zhang, Z.; Lin, F.; Kong, Z.; Cao, Y.; Li, C.; Yi, H.; Mei, M.; Zhu, H.; Wu, J.; et al. A high-density intervarietal map of the wheat genome enriched with markers derived from expressed sequence tags. Theor Appl Genet 2008, 117, 181-189. [CrossRef]
- Sun, C.; Dong, Z.; Zhao, L.; Ren, Y.; Zhang, N.; Chen, F. The Wheat 660K SNP array demonstrates.
- great potential for marker-assisted selection in polyploid wheat. Plant Biotechnol J 18:1354-1360.
- Eversole, K.; Feuillet, C.; Mayer, K.F.; Rogers, J. Slicing the wheat genome. Introduction. Science 2014, 345, 285-287. [CrossRef]
- Bariana, H.S.; Bansal, U.K.; Schmidt, A.; Lehmensiek, A.; Kaur, J.; Miah, H.; Gill, M.B.; Howes, N.; McIntyre, C.L. Molecular mapping of adult plant stripe rust resistance in wheat and identification of pyramided QTL genotypes. 2010 Euphytica 176:251-260. Euphytica 2010, 176, 251-260.
- Zakeri , A.; McIntosh, R.A.; Hovmoller, M.S.; Wellings, C.R.; Shariflou, M.R.; Hayden, M.; Bariana, H.S. Recombination of Yr15 and Yr24 in chromosome 1BS. In Proceedings of the Proceedings of 10th International Wheat Genetics Symposium, Paestum, Rome, Italy, 1-5 September 2003, 2003; pp. 417- 420.
- Miller, T.; Reader, S.; Ambrose, M. The Watkins wheat collection. Annu Wheat Newsl 2000, 46, 172.
- Daetwyler, H.D.; Bansal, U.K.; Bariana, H.S.; Hayden, M.J.; Hayes, B.J. Genomic prediction for rust resistance in diverse wheat landraces. Theor Appl Genet 2014, 127, 1795-1803. [CrossRef]
- Bansal, U.K.; Kazi, A.G.; Singh, B.; Hare, R.A.; Bariana, H.S. Mapping of durable stripe rust resistance in a durum wheat cultivar Wollaroi. Mol Breeding 2014, 33, 51-59. [CrossRef]
- Rozen, S.; Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. In Bioinformatics Methods and Protocols, Misener, S., Krawetz, S., Eds.; Methods in Molecular Biology™; Humana Press: Totowa, NJ, 1999; Volume 132, pp. 365-386.
- Somers, D.J.; Isaac, P.; Edwards, K. A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 2004, 109, 1105-1114. [CrossRef]
- Randhawa, M.; Bansal, U.; Valarik, M.; Klocova, B.; Dolezel, J.; Bariana, H. Molecular mapping of stripe rust resistance gene Yr51 in chromosome 4AL of wheat. Theor Appl Genet 2014, 127, 317-324. [CrossRef]
- Endo, T.R.; Gill, B.S. The deletion stocks of common wheat. Journal of Heredity 1996, 87, 295-307, doi:DOI 10.1093/oxfordjournals.jhered.a023003.
- Manly, K.F.; Cudmore, R.H., Jr.; Meer, J.M. Map Manager QTX, cross-platform software for genetic mapping. Mammalian genome : official journal of the International Mammalian Genome Society 2001, 12, 930-932. [CrossRef]
- Kosambi, D.D. The estimation of map distances from recombination values. Ann Eugenic 1944, 12, 172-175.
- Voorrips, R.E. MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 2002, 93, 77-78. [CrossRef]
- Zhang, P.; McIntosh, R.A.; Hoxha, S.; Dong, C. Wheat stripe rust resistance genes Yr5 and Yr7 are allelic. Theor Appl Genet 2009, 120, 25-29. [CrossRef]
- Chen, X.M.; Soria, M.A.; Yan, G.P.; Sun, J.; Dubcovsky, J. Development of sequence tagged site and cleaved amplified polymorphic sequence markers for wheat stripe rust resistance gene Yr5. Crop Sci 2003, 43, 2058-2064, doi:DOI 10.2135/cropsci2003.2058.
- Cheng, P.; Chen, X.M. Molecular mapping of a gene for stripe rust resistance in spring wheat cultivar IDO377s. Theor Appl Genet 121, 195–204 (2010). [CrossRef]
- Xu, L.S.; Wang, M.N.; Cheng, P.; Kang, Z.S.; Hulbert, S.H.; Chen, X.M. Molecular mapping of Yr53, a new gene for stripe rust resistance in durum wheat accession PI 480148 and its transfer to common wheat. Theor Appl Genet 2013, 126, 523-533. [CrossRef]
- Yaniv, E.; Raats, D.; Ronin, Y.; Korol, A.; Grama, A.; Bariana, H.; Dubcovsky, J.; Schulman, A.; Fahima, T. Evaluation of marker-assisted selection for the stripe rust resistance gene Yr15, introgressed from wild emmer wheat. Mol Breeding 2015, 35, 1-12. [CrossRef]
- Randhawa, M.S.; Bariana, H.S.; Mago, R.; Bansal, U.K. Mapping of a new stripe rust resistance locus Yr57 on chromosome 3BS of wheat. Mol Breeding 2015, 35, 1-8, doi:ARTN 65 10.1007/s11032-015-0270-0.
- Lagudah, E.S.; Krattinger, S.G.; Herrera-Foessel, S.; Singh, R.P.; Huerta-Espino, J.; Spielmeyer, W.; Brown-Guedira, G.; Selter, L.L.; Keller, B. Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor Appl Genet 2009, 119, 889-898. [CrossRef]
- Uauy, C.; Brevis, J.C.; Chen, X.; Khan, I.; Jackson, L.; Chicaiza, O.; Distelfeld, A.; Fahima, T.; Dubcovsky, J. High-temperature adult-plant (HTAP) stripe rust resistance gene Yr36 from Triticum turgidum ssp. dicoccoides is closely linked to the grain protein content locus Gpc-B1. Theor Appl Genet 2005, 112, 97-105. [CrossRef]
- Herrera-Foessel, S.A.; Singh, R.P.; Lillemo, M.; Huerta-Espino, J.; Bhavani, S.; Singh, S.; Lan, C.; Calvo-Salazar, V.; Lagudah, E.S. Lr67/Yr46 confers adult plant resistance to stem rust and powdery mildew in wheat. Theor Appl Genet 2014, 127, 781-789. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).