Submitted:
09 May 2023
Posted:
10 May 2023
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
- Molecular weight ≤ 500
- Indicator of lipophilicity, LogP ≤ 5
- Number of hydrogen bond donors ≤ 10
- Number of hydrogen bond acceptors ≤ 5
- Molecular weight > 400
- Indicator of lipophilicity, LogP > 4
- Number of cyclic structures > 4
- Number of hydrogen bond acceptors > 4
2. Materials and Methods
2.1. Molecular generation model
2.2. Scoring function
2.3. Computational experiments
3. Results
3.1. Inducing exploration through reinforcement learning
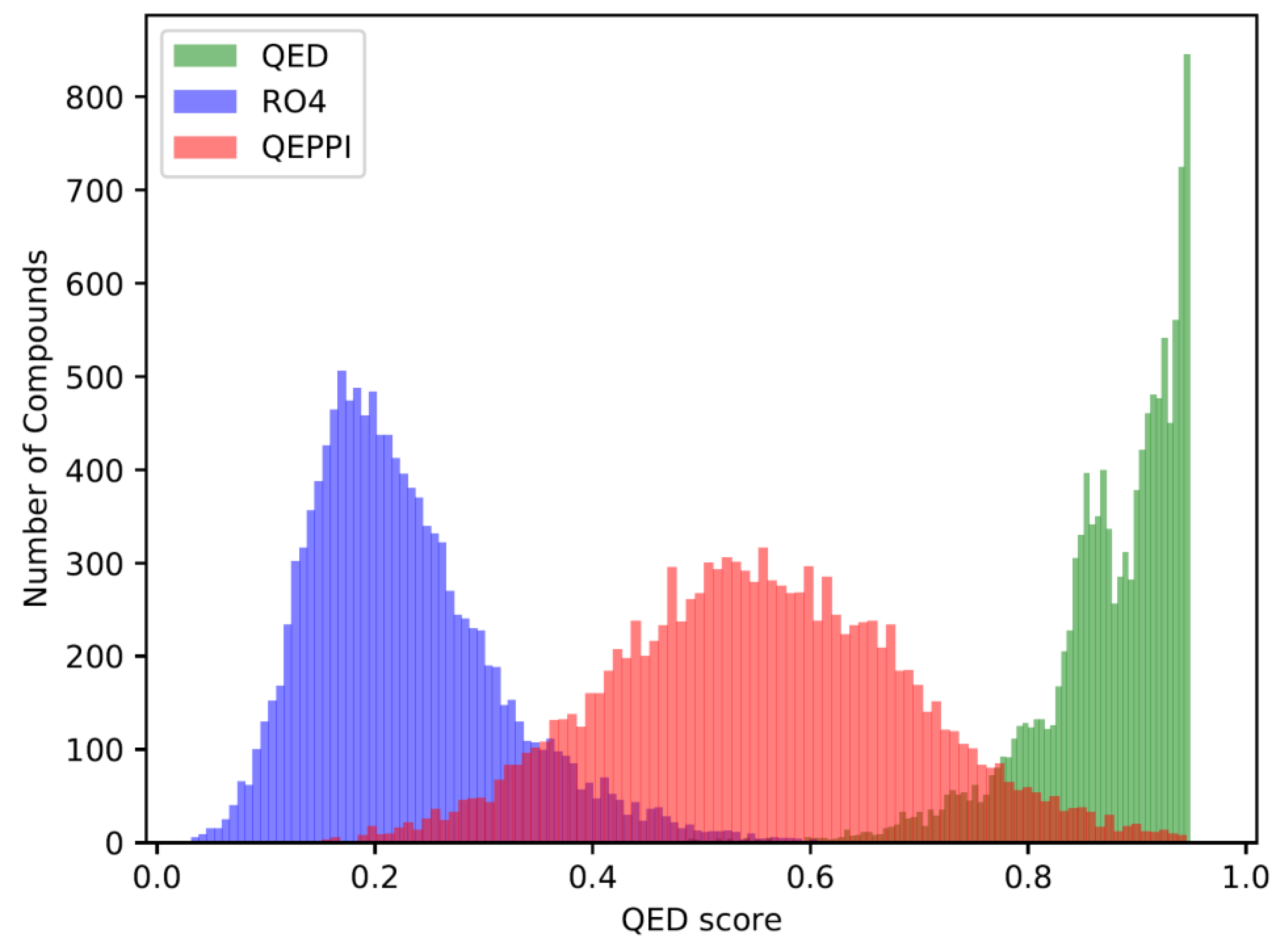
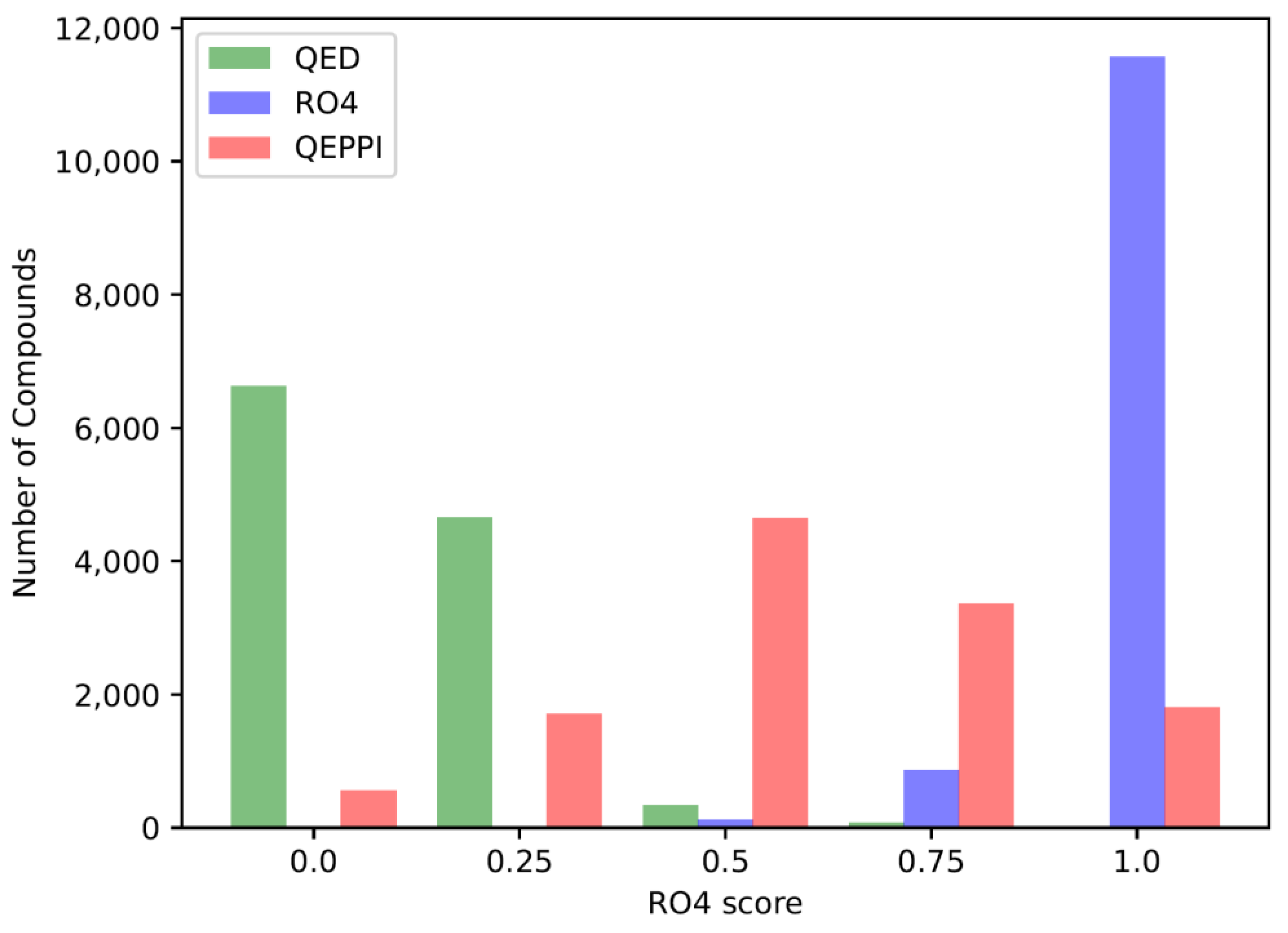
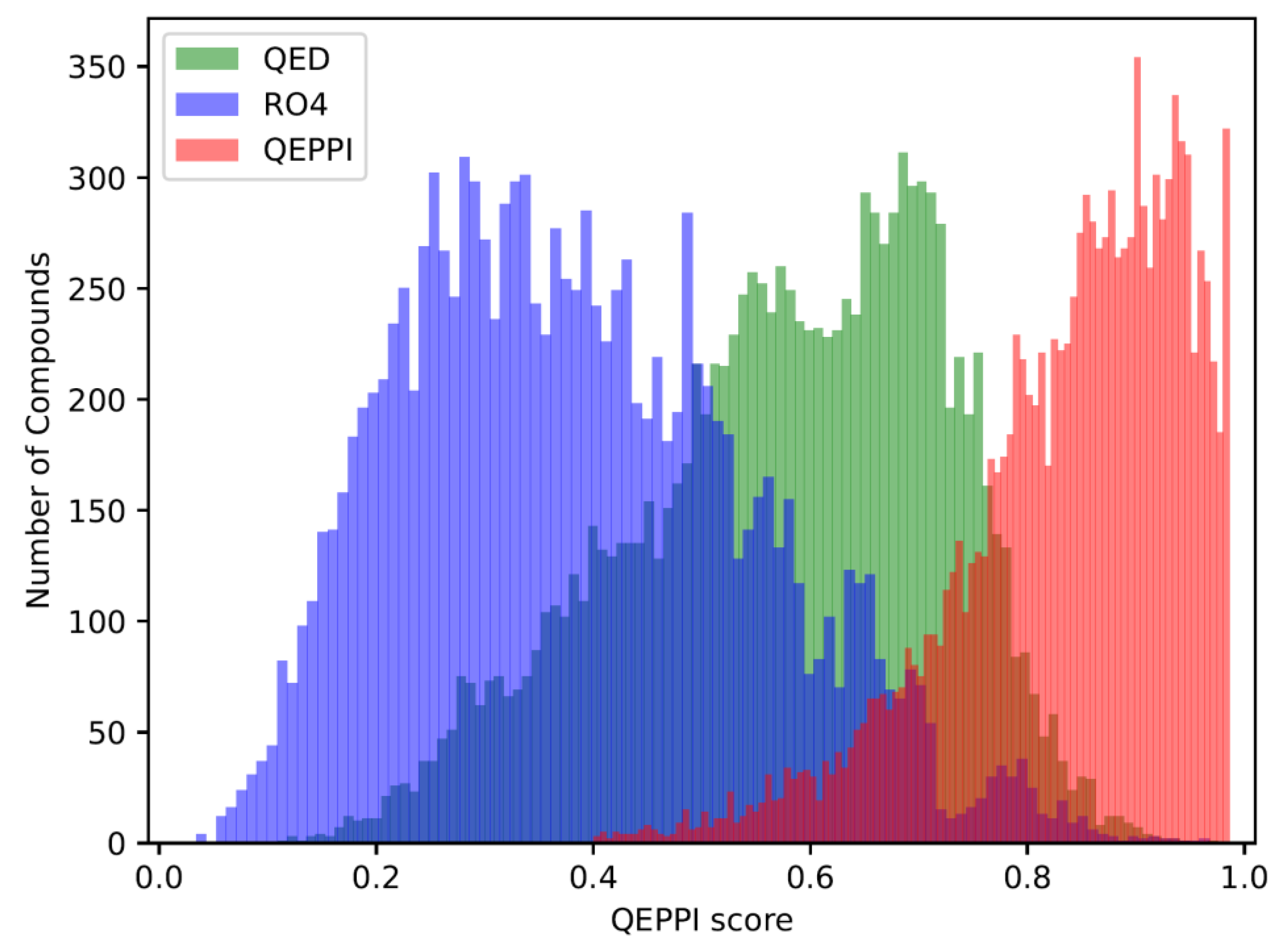
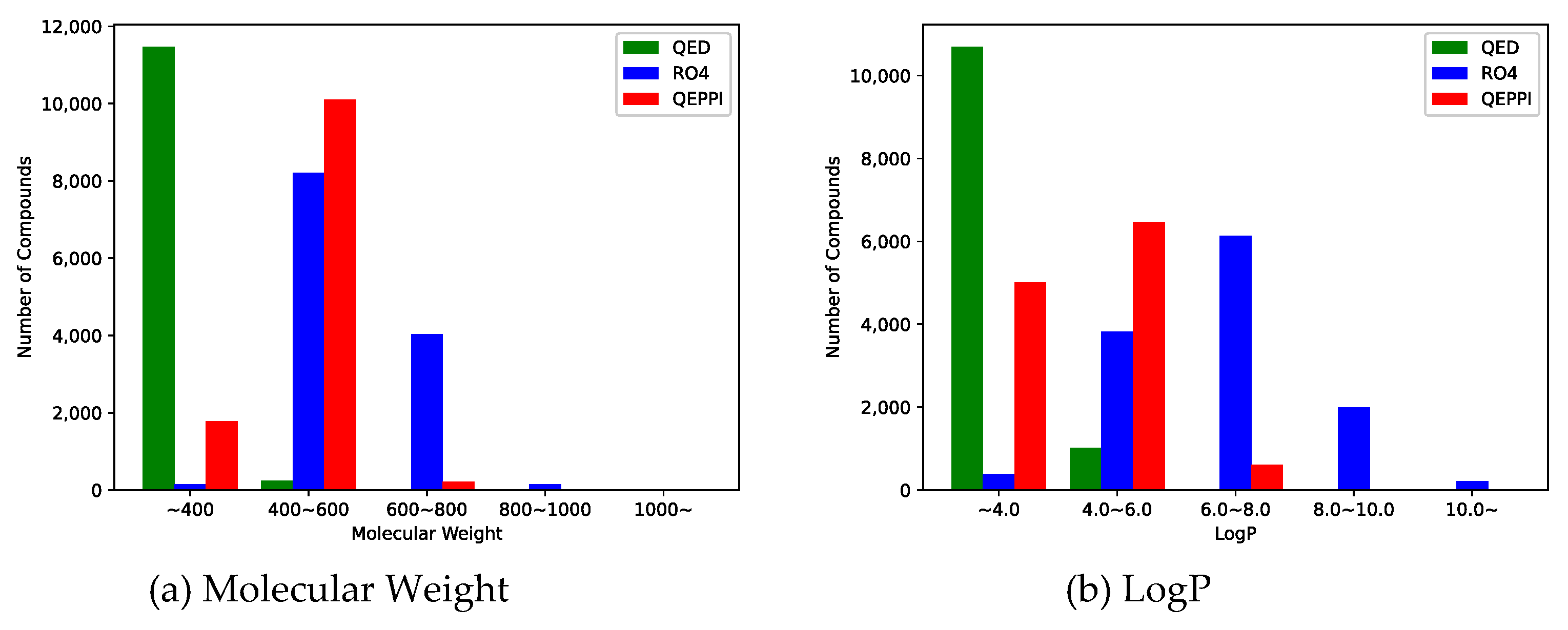
3.2. Distribution of compounds generated by REINVENT
3.3. Indicators for oral bioavailability
- Number of rotatable bonds, Rbond ≤ 10
- Topological polar surface area TPSA ≤ 140
3.4. Constructing virtual libraries of PPI-target compounds
4. Discussion
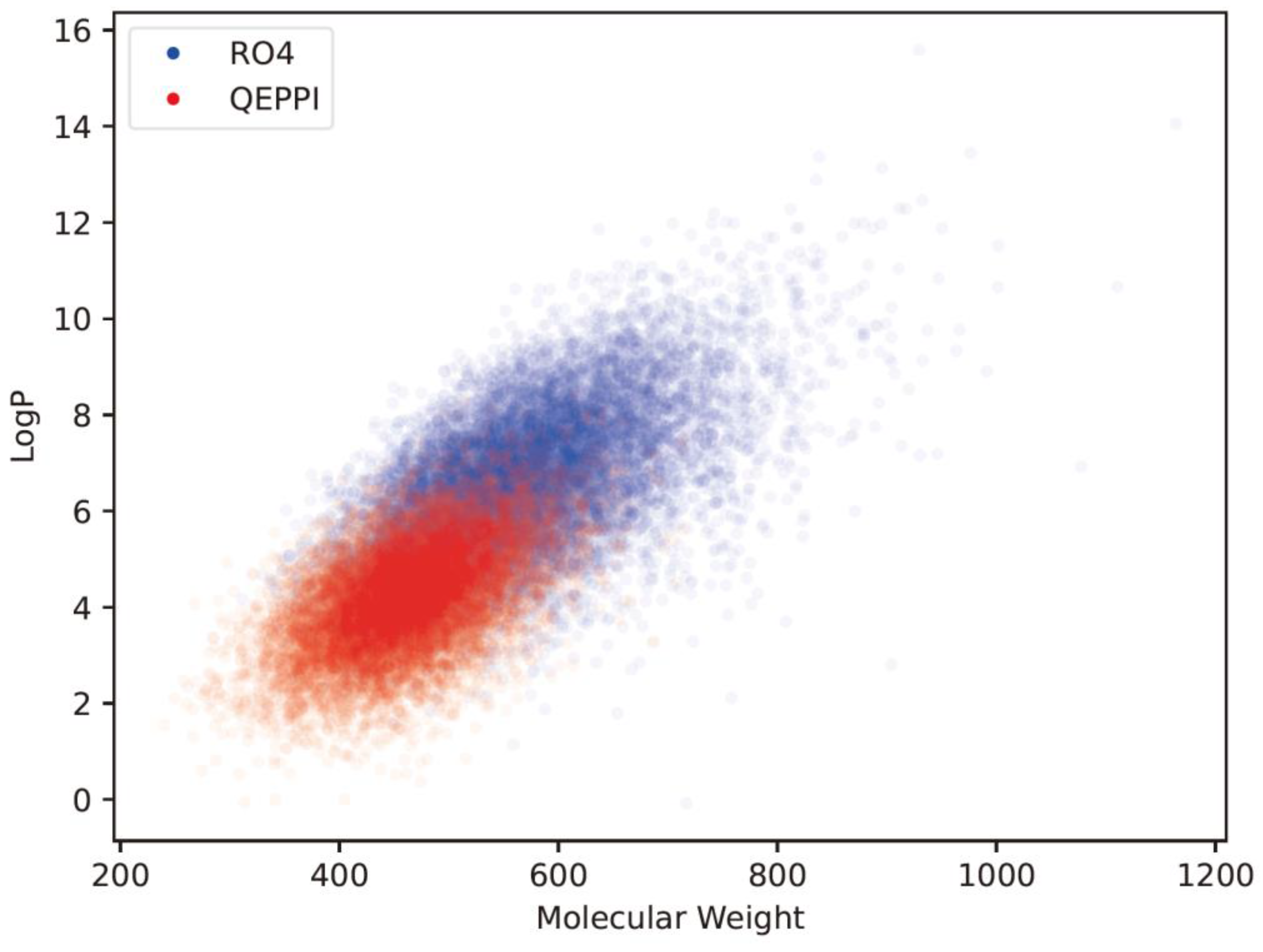
4.1. Chemical space of generated compounds
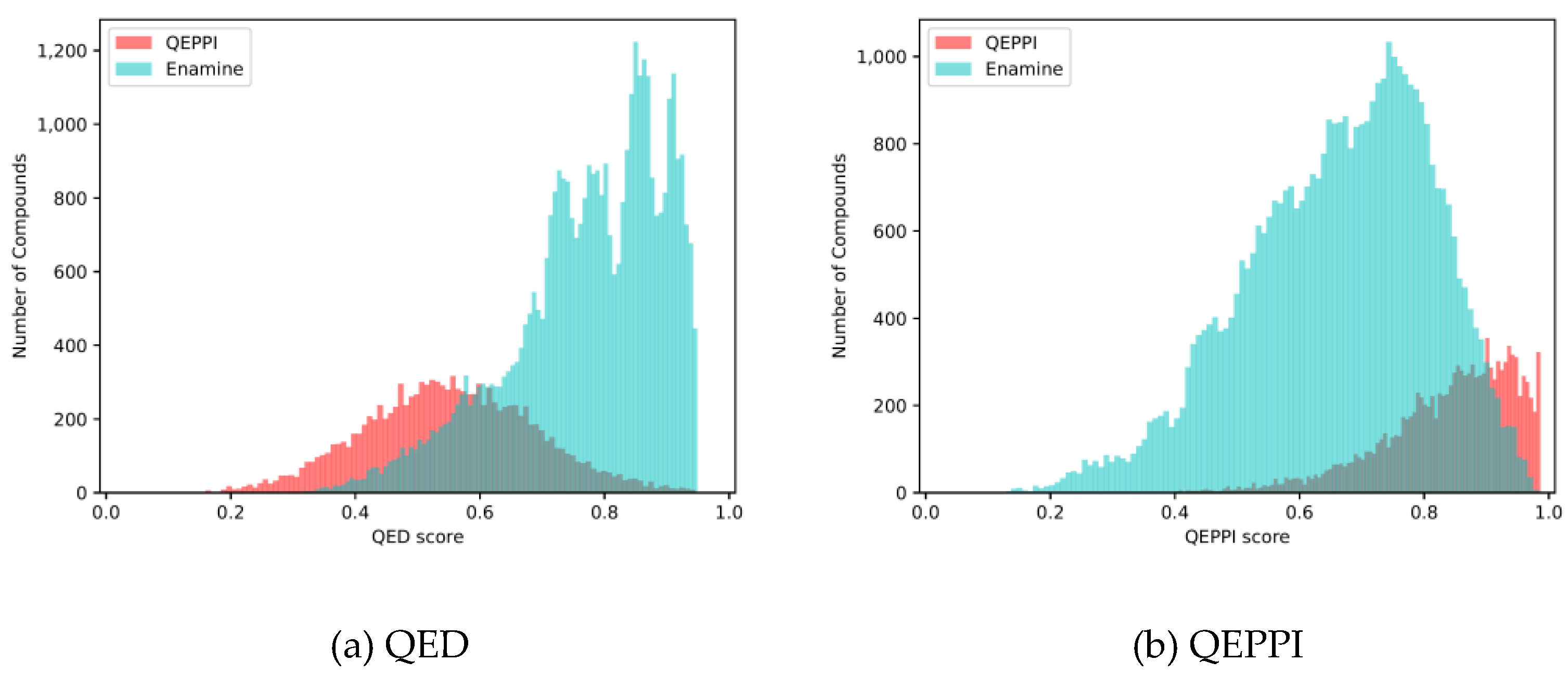
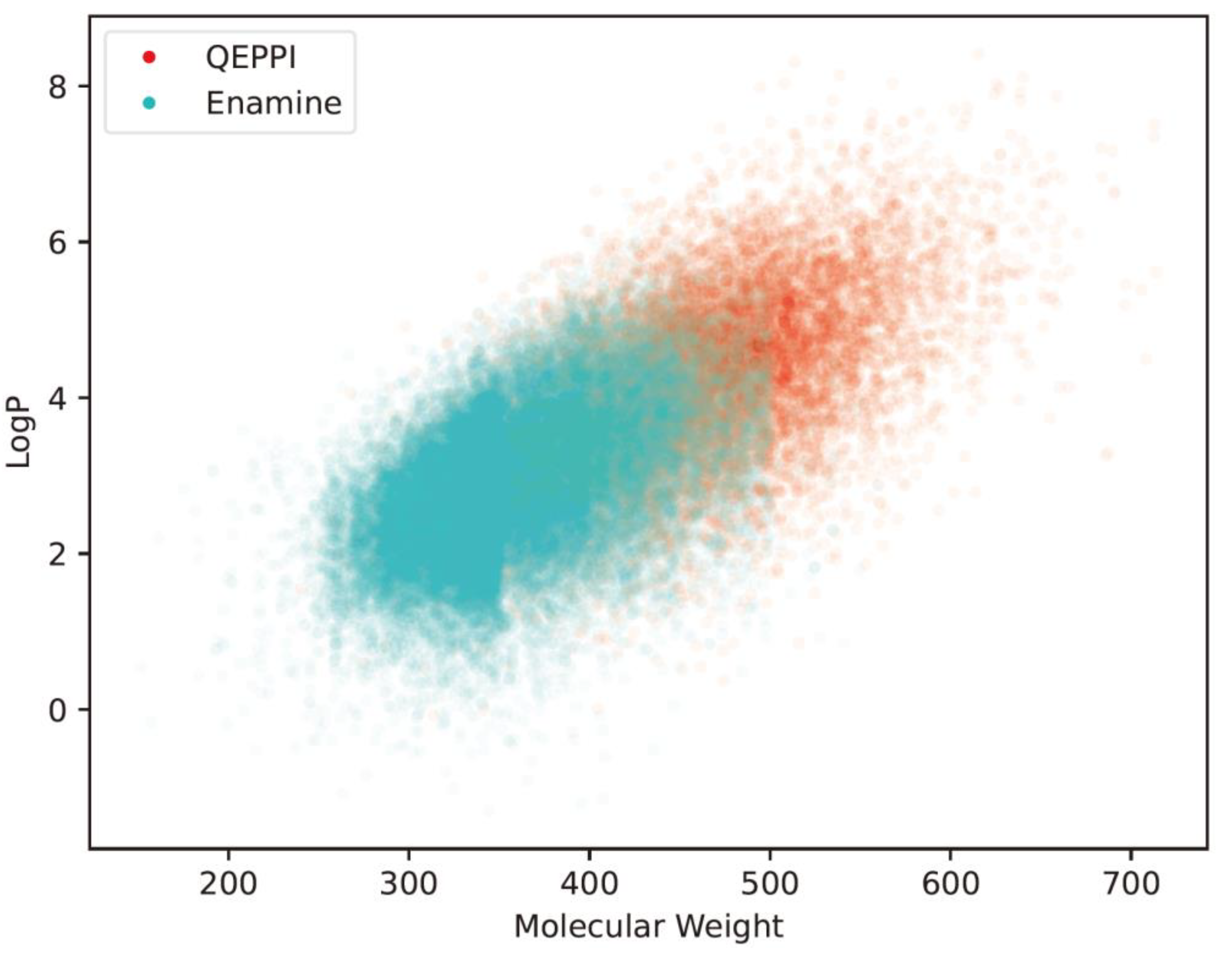
4.2. Comparison with existing PPI libraries
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Scannell, J.W.; Blanckley, A.; Boldon, H.; Warrington, B. Diagnosing the decline in pharmaceutical R&D efficiency. Nat Rev Drug Discov. 2012, 11, 191–200. [Google Scholar] [CrossRef] [PubMed]
- Toogood, P.L. Inhibition of protein-protein association by small molecules: Approaches and progress. J Med Chem. 2002, 45, 1543–1558. [Google Scholar] [CrossRef] [PubMed]
- Arkin, M.R.; Wells, J.A. Small-molecule inhibitors of protein-protein interactions: Progressing towards the dream. Nat Rev Drug Discov. 2004, 3, 301–317. [Google Scholar] [CrossRef] [PubMed]
- Dev, K.K. Making protein interactions druggable: Targeting PDZ domains. Nat Rev Drug Discov. 2004, 3, 1047–1056. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Wang, W.; Fang, G. Targeting protein-protein interaction by small molecules. Annu Rev Pharmacol Toxicol. 2014, 54, 435–456. [Google Scholar] [CrossRef]
- Ivanov, A.A.; Khuri, F.R.; Fu, H. Targeting protein–protein interactions as an anticancer strategy. Trends Pharmacol Sci. 2013, 34, 393–400. [Google Scholar] [CrossRef]
- Mao, Y.; Fisher, D.W.; Yang, S.; Keszycki, R.M.; Dong, H. Protein-protein interactions underlying the behavioral and psychological symptoms of dementia (BPSD) and Alzheimer’s disease. PLOS ONE. 2020, 15, e0226021. [Google Scholar] [CrossRef]
- Shin, W.H.; Kumazawa, K.; Imai, K.; Hirokawa, T.; Kihara, D. Current challenges and opportunities in designing protein-protein interaction targeted drugs. Adv Appl Bioinform Chem. 2020, 13, 11–25. [Google Scholar] [CrossRef]
- Shin, W.H.; Christoffer, C.W.; Kihara, D. In silico structure-based approaches to discover protein-protein interaction-targeting drugs. Methods. 2017, 131, 22–32. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev. 1997, 23, 3–26. [Google Scholar] [CrossRef]
- Lipinski, C.A. Lead- and drug-like compounds: The rule-of-five revolution. Drug Discov Today Technol. 2004, 1, 337–341. [Google Scholar] [CrossRef] [PubMed]
- Morelli, X.; Bourgeas, R.; Roche, P. Chemical and structural lessons from recent successes in protein-protein interaction inhibition (2P2I). Curr Opin Chem Biol. 2011, 15, 475–481. [Google Scholar] [CrossRef] [PubMed]
- Basse, M.J.; Betzi, S.; Morelli, X.; Roche, P. 2P2Idb v2: Update of a structural database dedicated to orthosteric modulation of protein-protein interactions. Database (Oxford). 2016, 2016, No. baw007. [Google Scholar] [CrossRef]
- Jnoff, E.; Albrecht, C.; Barker, J.J.; Barker, O.; Beaumont, E.; Bromidge, S.; Brookfield, F.; Brooks, M.; Bubert, C.; Ceska, T.; et al. Binding mode and structure-activity relationships around direct inhibitors of the Nrf2-Keap1 complex. ChemMedChem. 2014, 9, 699–705. [Google Scholar] [CrossRef] [PubMed]
- Bosc, N.; Muller, C.; Hoffer, L.; Lagorce, D.; Bourg, S.; Derviaux, C.; Gourdel, M.E.; Rain, J.C.; Miller, T.W.; Villoutreix, B.O.; et al. Fr-PPIChem: An academic compound library dedicated to protein-protein interactions. ACS Chem Biol. 2020, 15, 1566–1574. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, Y.; Yonezawa, T.; Sakamoto, J.; Furuya, T.; Osawa, M.; Ikeda, K. Identification of novel inhibitors of Keap1/Nrf2 by a promising method combining protein–protein interaction-oriented library and machine learning. Sci Rep. 2021, 11, 7420. [Google Scholar] [CrossRef]
- Blaschke, T.; Aŕus-Pous, J.; Chen, H.; Margreitter, C.; Tyrchan, C.; Engkvist, O.; Papadopoulos, K.; Patronov, A. REINVENT 2.0: An AI tool for de novo drug design. J Chem Inf Model. 2020, 60, 5918–5922. [Google Scholar] [CrossRef]
- Gaulton, A.; Hersey, A.; Nowotka, M.; Bento, A.P.; Chambers, J.; Mendez, D.; Mutowo, P.; Atkinson, F.; Bellis, L.J.; Cibrían-Uhalte, E.; Davies, M.; Dedman, N.; Karlsson, A.; Magariños, M.P.; Overington, J.P.; Papadatos, G.; Smit, I.; Leach, A.R. The ChEMBL database in 2017. Nucleic Acids Res. 2017, 45, D945–D954. [Google Scholar] [CrossRef]
- MolecularAI. ReinventCommunity. Available online: https://github.com/MolecularAI/ReinventCommunity.
- Bickerton, G.R.; Paolini, G.V.; Besnard, J.; Muresan, S.; Hopkins, A.L. Quantifying the chemical beauty of drugs. Nat Chem. 2012, 4, 90–98. [Google Scholar] [CrossRef]
- Kosugi, T.; Ohue, M. Quantitative estimate index for early-stage screening of compounds targeting protein-protein interactions. Int J Mol Sci. 2021, 22, 10925. [Google Scholar] [CrossRef]
- Wang, J.; Chu, Y.; Mao, J.; Jeon, H.N.; Jin, H.; Zeb, A.; Jang, Y.; Cho, K.H.; Song, T. De novo molecular design with deep molecular generative models for PPI inhibitors. Brief Bioinform. 2022, 23, bbac285. [Google Scholar] [CrossRef] [PubMed]
- Veber, D.F.; Johnson, S.R.; Cheng, H.Y.; Smith, B.R.; Ward, K.W.; Kopple, K.D. Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem. 2002, 45, 2615–2623. [Google Scholar] [CrossRef] [PubMed]
- Baell, J.B.; Holloway, G.A. New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. J Med Chem. 2010, 53, 2719–2740. [Google Scholar] [CrossRef] [PubMed]
- Enamine; PPI Library. Available online: https://enamine.net/compound-libraries/targeted-libraries/ppi-library.
- Ertl, P.; Schuffenhauer, A. Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J Cheminform. 2009, 1, 8. [Google Scholar] [CrossRef]
- Genheden, S.; Thakkar, A.; Chadimová, V.; Reymond, J.L.; Engkvist, O.; Bjerrum, E. AiZynthFinder: A fast, robust and flexible open-source software for retrosynthetic planning. J Cheminform. 2020, 12, 70. [Google Scholar] [CrossRef]
- Kengkanna, A.; Ohue, M. Enhancing Model Learning and Interpretation Using Multiple Molecular Graph Representations for Compound Property and Activity Prediction. arXiv, 2023; arXiv:2304.06253. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).



