Introduction
The folding pathways problem originates from Levinthal (1968), who posed a paradox (Levinthal, 1969) bearing his name: how can a protein attain its natural state within a biologically feasible timeframe while randomly exploring the whole conformational space? A proper answer to this query is of foremost importance since protein folds from milliseconds to seconds rather than (~1052) years—as foreseen by an exhaustive enumeration of all possible conformations for a 100-residue protein (Karplus, 2011). As the reader must be aware, several possible solutions to this apparent contradiction—also known as Levinthal’s paradox—exist in the literature (Zwanzig et al., 1992; Karplus, 1997; Finkelstein & Badretdinov, 1977; Dill & Chan, 1997; Bogatyreva & Finkelstein, 2001; Rooman et al., 2002; Garbuzynskiy et al., 2012; Ben-Naim, 2012; Finkelstein & Garbuzynskiy, 2013; Martinez, 2014; Ivankov & Finkelstein, 2020; Vila, 2023a). Although we will not revisit each possible solution here, it is worth noting that solving this paradox is relevant because it could enable us to determine clear answers to the following key questions: Why and how can proteins reach their native state in a biologically reasonable time? Regarding the ‘why,’ we demonstrated—from a thermodynamic viewpoint—that the range of ‘slowest’ folding times for two-state monomeric proteins (τ) spans from ~milliseconds to ~seconds (Vila, 2023a), which closely aligns with the observed data (Garbuzynskiy et al., 2012; Ivankov & Finkelstein, 2020). These time scales for protein folding kinetics arise from the fulfillment of the thermodynamic hypothesis (Anfinsen, 1973), also known as the Anfinsen dogma, which allowed us to demonstrate—through a statistical-thermodynamics analysis (Vila, 2019)—the existence of an upper limit to the marginal stability of globular proteins (ΔG) of approximately 7.4 kcal/mol (Vila, 2022), beyond which proteins either unfold or lose their functionality (Vila, 2021). This threshold applies to any fold class, sequence, or protein size, and it is significant because it arises from a quasi-equilibrium of forces that occurs at the lowest accessible protein's free-energy minimum (Martin & Vila, 2020). As to the ‘how’—which mainly focuses on clarifying the mechanisms by which proteins achieve their native states, including the pathways and trajectories of folding—it is a query that remains largely unresolved, despite the extensive literature available on the subject (Anfinsen & Scheraga, 1975; Creighton, 1985; Šali et al., 1994; Thirumalai, 1995; Baldwin, 1994; Baldwin, 1995; Wolynes et al., 1995; Lazaridis & Karplus, 1997; Dill & Chan, 1997; Pande et al., 1998; Dobson et al., 1998; Honig, 1999; Bakk et al., 2000; Englander et al., 2007; Karplus, 2007; Karplus, 2011; Englander & Mayne, 2014; Wolynes, 2015; Neupane et al., 2016; Lapidus, 2021; Kikuchi, 2022; Zhao et al., 2023; Fakhoury et al., 2023; Zhao et al., 2024; Chang & Perez, 2025). For a more profound understanding of the complexity of this problem, we should keep in mind that the protein folding process cannot be represented in a single dimension; rather, it results from interactions across multiple dimensions, creating complex multidimensional pathways and trajectories. This feature highlights the importance of understanding the interplay of complex forces at work, such as van der Waals interactions, hydrogen bonds, solvent effects, and electrostatic forces, all of which influence obtaining an accurate solution to the protein folding problem in a manner that demands solving it as an ‘analytical whole’ rather than a sum of parts (Vila, 2023b).
It is essential to acknowledge two key points prior to moving forward. Firstly, the terms ‘pathway’ and ‘trajectory’ represent distinct concepts. The distinction between pathways and trajectories is that pathways consist of discrete sequences of events—whether parallel or linear—to go from point A (terminus a quo) to point B (terminus ad quem). In contrast, trajectories encompass the entire duration of the process (Vila, 2025). Secondly, it is well-documented experimentally that proteins can reliably attain the folded state at the same rate in a proper environment (Harrington & Schellman, 1956; Gromiha, 2005; Huang et al., 2008; Finkelstein & Garbuzynskiy, 2013), thereby suggesting that all folding pathways—if multiple exist—may be characterized by an effective trajectory. To gain a better grasp of this conjecture, we will analyze it from the standpoint of the “least action principle,” a fundamental physical principle (Fee, 1941) and a powerful model for describing how systems behave under the constraints mentioned above (Feynman et al., 1963; Hanc & Taylor, 2004). This principle's significance encompasses all domains of physics, rendering it the nearest approximation to a theory of everything, as it pertains to classical mechanics (Landau & Lifshitz, 1982), relativity (Landau & Lifshitz, 1987), quantum mechanics (Feynman, 2005), thermodynamics (Kaila & Annila, 2008), and biophysics (Simmons & Weiner, 2013). This approach will allow us to further explore the more probable pathways that proteins may follow to achieve their folded or unfolded states within a biologically reasonable timeframe. This endeavor also aims to tackle the Levinthal paradox from a fresh perspective, distinguishing the current study from a previous resolution (Vila, 2023a). Furthermore, we seek to uncover the basic physical principles that dictate how proteins fold in response to alterations in their environment or sequence.
Outlining the System Being Studied
The existence of metamorphic proteins (Murzin, 2008) indicates that the role of environmental factors cannot be overstated (Vila, 2020), as even slight changes in conditions can lead to dramatically different folding outcomes (Dishman & Volkman, 2018). This viewpoint emphasizes that the dynamic nature of proteins means their behavior is not solely determined by intrinsic properties—encoded in their amino acid sequence (Anfinsen, 1973)—but also heavily influenced by external stimuli. This complexity highlights the necessity of studying protein behavior under varied conditions to understand the principles of folding and stability fully. A single-point change in the amino acid sequence—such as due to mutations—only exacerbates this problem by giving rise, among other things, to epistatic effects (Vila, 2024a).
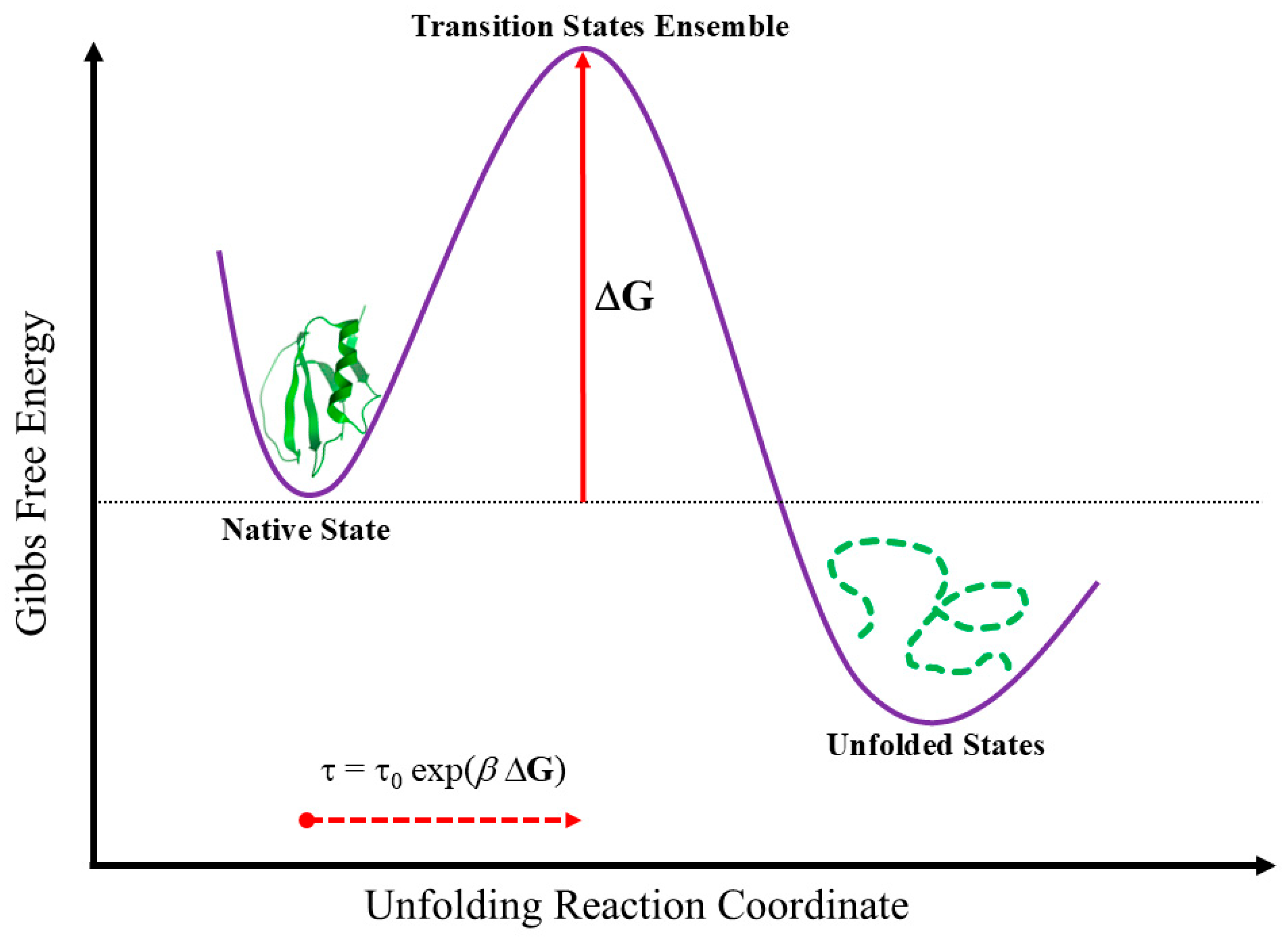
In summary, based on the considerations mentioned above, we will now focus on analyzing the folding of a single two-state protein in a fixed environmental setting. The analysis of two-state proteins (Jackson, 1998) is chosen for a specific reason: the native state and unfolded states are separated by an energetic barrier that exceeds the energy of thermal fluctuations (Akmal & Muñoz, 2004; Kuwajima, 2020). In other words, the folded and unfolded states are separated by a collection of high-energy native-like structures known as the Transition State Ensemble (TSE), which represents (as shown in
Figure 1) the energetic barrier (ΔG) for the process (Matouschek
et al., 1989; Itzhaki
et al., 1995; Englander, 2000; Ding
et al., 2002; Vendruscolo
et al., 2003; Akmal & Muñoz, 2004; Shakhnovich, 2006; Englander & Mayne, 2014; Lange
et al., 2008; Kuwajima, 2020; Stiller
et al., 2022; Li & Gong, 2022). In this simple folding model, there are no stable intermediate states necessary to complete the process. Therefore, the treatment of protein folding and unfolding can be viewed as interchangeable processes (Lazaridis & Karplus, 1997; Jackson, 1998; Ivankov & Finkelstein, 2020)—as long as the Gibbs free-energy barrier (ΔG) remains the same for both cases. The folding and unfolding data from 108 proteins, of which 70 display two-state kinetics (Glyakina & Galzitskaya, 2020), support these claims, as the logarithm of the folding and unfolding rates demonstrates a strong correlation (
R ~0.8). Henceforth, these terms shall be considered synonymous from this point forward.
The Least Action Principle and the Effective-Trajectory Conjecture
For a comprehensive and detailed exploration of the application of the principle of least action in classical mechanics, thermodynamics, relativity, and quantum mechanics, we recommend consulting the existing literature (Landau & Lifshitz, 1982; Landau & Lifshitz, 1987; Feynman, 2005; Kaila & Annila, 2008). We will skip over all those specific uses and instead provide, firstly, a layman's definition of the concept of least action and, later, a more concise definition of the principle from a physics-mathematics perspective.
The principle of least action in physics states that a system going from a starting to an ending point—such as the folding process of proteins—will naturally follow a path that minimizes the ‘action’ (a functional of the trajectory) among all feasible pathways connecting these two points. The implication here is that nature operates with optimal efficiency, rather than randomly searching among all feasible pathways.
Let us now focus on a brief description of the principle of least action within the framework of Lagrangian mechanics—an approach that enables us to frame the system using a function encompassing both kinetic and potential energy contributions (Feynman
et al., 1963; Hanc & Taylor, 2004). Let us begin by characterizing the folding of a protein as a process that occurs in a high-dimensional dihedral-angle space, which could be represented by a smooth, continuous function:
f)
—where {
} denotes a set of internal dihedral angles of the protein at time
t. This function determines how the protein conformation evolves along a given folding pathway under a given
all-atom force field—which, in simple terms, refers to
all the lines of force around and between atoms—such as the empirical conformational energy program for peptides (Arnautova
et al., 2006). Additionally, we can define the time derivative of this function [
], which identifies the folding velocity along a specified path that must be traversed within a well-defined range of folding rates (τ). Specifically, it must satisfy the following boundary constraint: ~milliseconds < τ < ~seconds (Vila, 2023a)—as per an analysis grounded in transition state theory (Ivankov & Finkelstein, 2020)—with the folding rate calculated as τ = τ
0 exp (
βΔG). Here, ΔG represents the protein's marginal stability (Vila, 2022), namely, the Gibbs free energy difference between the native state and the highest-energy native-like structures—of the transition state ensemble (TSE)—coexistent with it (see
Figure 1); τ
0 is a pre-exponential factor that indicates the folding speed limit of two-state proteins—also known as the barrier-less limit (Zana, 1975; McCammon, 1996; Mayor
et al., 2000; Muñoz & Cerminara, 2016; Eaton, 2021)—and
β = 1/
RT, with
R representing the gas constant and
T being the absolute temperature. After providing some details of the folding process, the action (
S), a functional of the trajectory, can be defined as the integral of the Lagrangian (
along any folding pathway, namely as:
The path integral spans the entire duration of the folding process, subject to two key constraints. First, the upper limit of the integration (t) must be less than a few seconds (Vila, 2023a). Second, the lower limit of the integration (t0) should fall between ten nanoseconds and ten microseconds, as this range represents the folding speed limit for two-state proteins (Zana, 1975; McCammon, 1996; Mayor et al., 2000; Muñoz & Cerminara, 2016; Eaton, 2021). It is important to highlight, on one hand, that the cap on the upper-bound limit of the integration (t)—applicable to any two-state protein, irrespective of its fold class, length, or sequence—has been demonstrated to stem from the validity of the thermodynamic hypothesis, also known as Anfinsen's dogma (Vila, 2023a). On the other hand, this constraint on the upper bound also provides explanations for why protein folding consistently occurs within a biologically reasonable time frame. In summary, the validity of the Anfinsen dogma delineates an integration limit that constrains viable folding pathways, as their trajectories are ruled by temporal constraints.
The task of solving equation (1) presents an enormous mathematical challenge that exceeds the current analysis's limits, as it requires solving—among other things—the Euler-Lagrange equations [
] that represent the solution to virtual variations on the functional
S (Coopersmith, 2017); instead, we will focus on highlighting its main features, aiming to clarify the potential relationship between the effective-trajectory conjecture and the least action principle. To achieve this, it is important to note that the Hamilton principle (of stationary action) states that the variations (δ) that make the integral zero (δ
S = 0), by infinitesimal virtual variation on
f, correspond to a stationary point of
S—where the functional must be at a minimum or a saddle point, but not a maximum. All things considered, a pathway is considered a stationary point of the action (
S) if—and only if—the Euler-Lagrange equations are satisfied (Coopersmith, 2017). Hence, protein folds by following a pathway that renders the action (
S) stationary. However, how can we guarantee that the action has a unique path that makes it stationary? The latter represents an important question, as the number of folding pathways—specifically, whether there is one, a few, or a countless number—has been, and continues to be, a topic of debate (Baldwin, 1994; Baldwin, 1995; Lazaridis & Karplus, 1997; Dill & Chan, 1997; Bakk
et al., 2000; Rooman
et al., 2002; Englander & Mayne, 2014; Eaton & Wolynes, 2017; Englander & Mayne, 2017). Then, what if numerous pathways exist, each represented by trajectories that share the same folding rate (τ) as their sole common feature? This viewpoint associates the uniqueness of the solution to Eq. (1) with the time necessary for the protein to navigate the most efficient pathways—those that render the action (
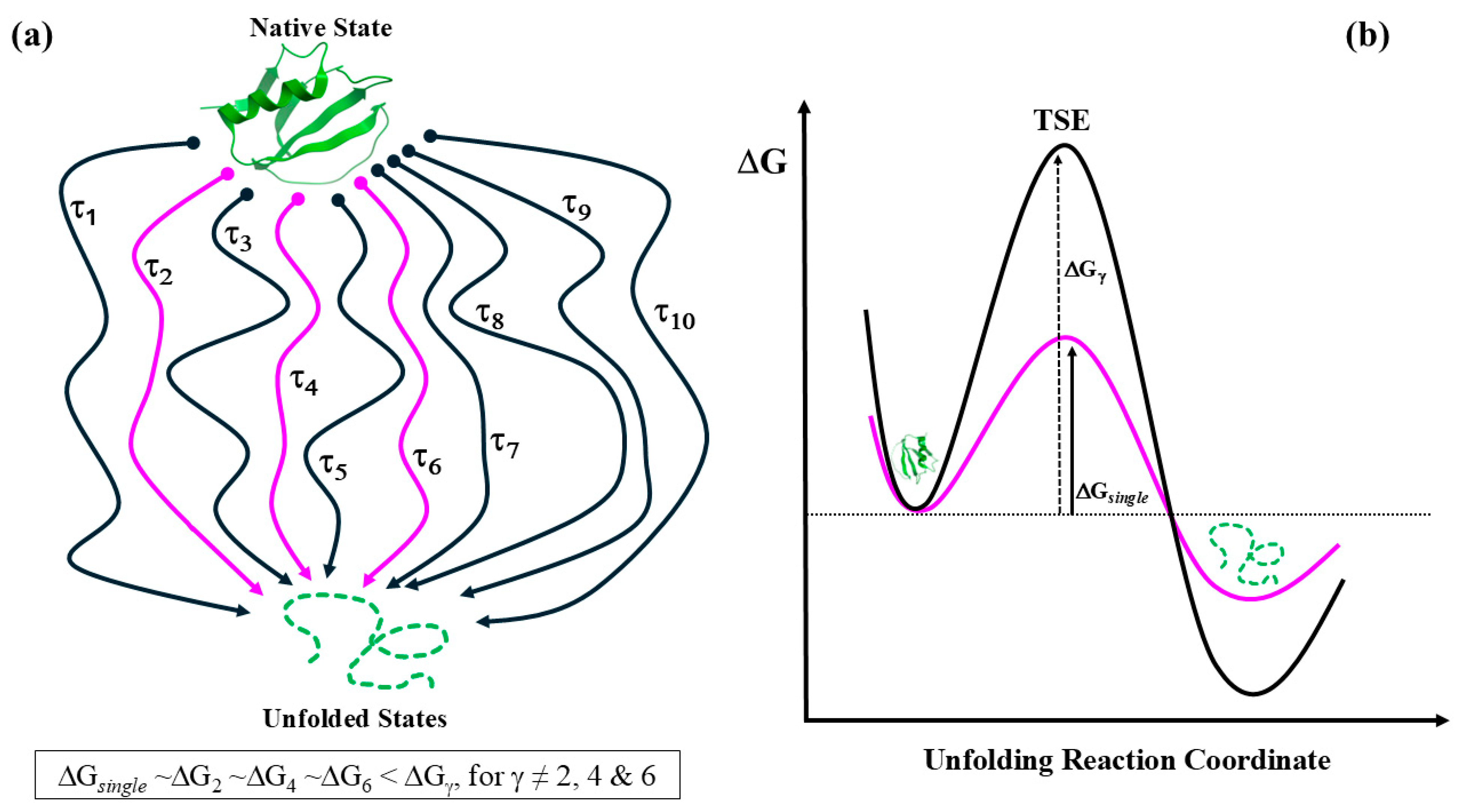
S) stationary—rather than suggesting the existence of a specific path (refer to
Figure 2). This leads to the conjecture of an ‘effective trajectory,’ which encompasses
all folding pathways characterized by a common single folding rate (τ
single), depending on the specific conditions described below. Firstly, the folding process must take place in a proper and controlled environment, which includes factors such as pH, temperature, and ionic strength, among others. Secondly, the single-folding rate must satisfy the following constraint: τ
single < ~seconds, as explained earlier.
The proposed effective-trajectory conjecture is based on a crucial observation: proteins unfold at a single-folding rate determined by DG (Vila, 2023a), which defines the protein's marginal stability while also serving as a threshold beyond which a protein may unfold or lose its functionality (refer to
Figure 1). This scenario offers a simple and credible explanation for how multiple pathways may converge on the protein native state at the same rate (see
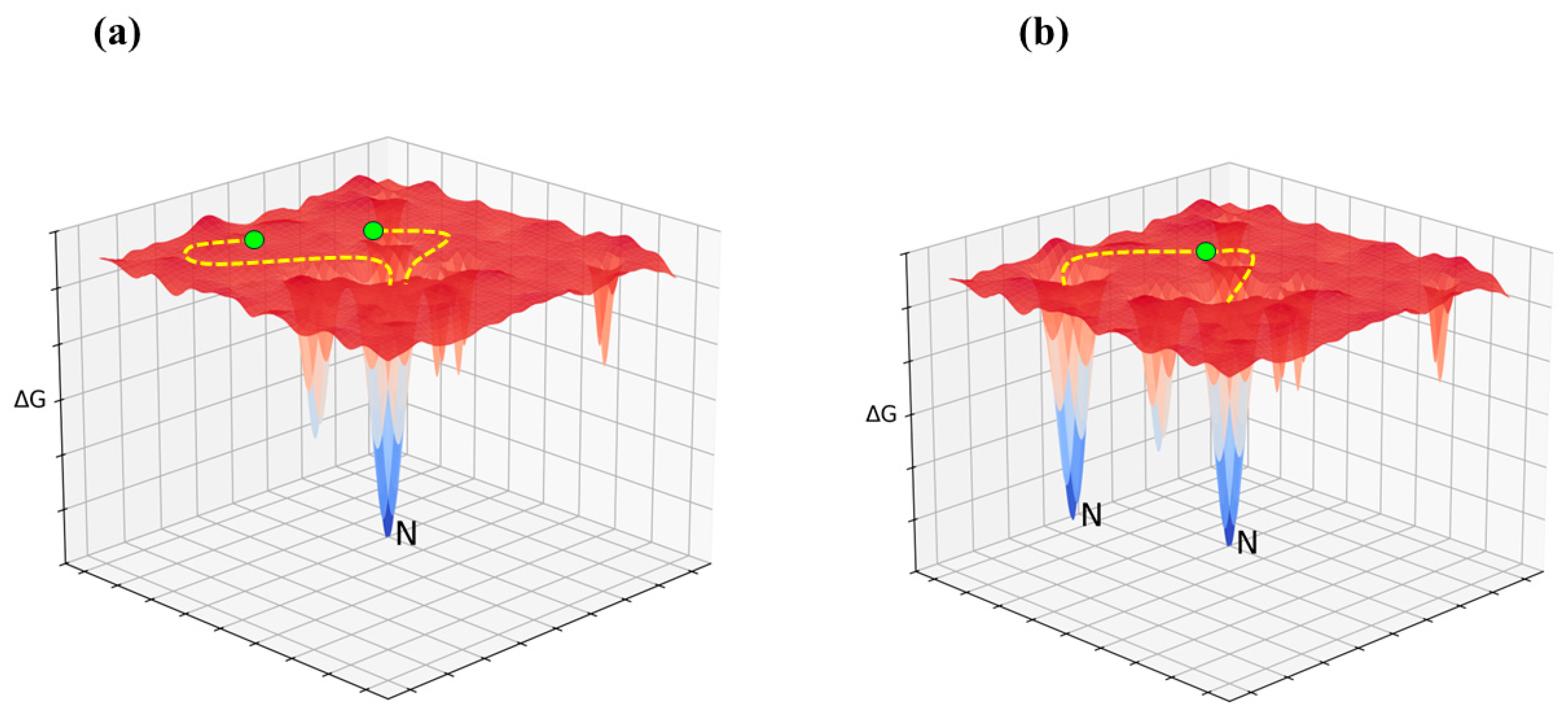
Figure 2)—even in the presence of a degenerate Gibbs free energy minimum (see
Figure 3), a possibility that cannot be ruled out (Vila
et al., 2003; Martin
et al., 2019). It is important to recognize that different proteins—whether they have varying sequences, differing numbers of residues, or both—can exhibit identical folding rates. Proteins with Protein Data Bank identifiers 1PBA, 1AYE, and 2VIK illustrate this phenomenon (Huang
et al., 2008). At first glance, it may appear odd that such proteins can exhibit identical folding rates while possessing distinct folding pathways. However, upon recognizing two pivotal facts, this phenomenon becomes readily comprehensible. Firstly, the folding rates are limited—as previously mentioned—by the protein's marginal stability—regardless of fold class, sequence, or protein size (Vila, 2023a). This proposal is consistent with evidence suggesting that folding rates are insensitive to a protein's sequence details (Plaxco
et al., 2000). Paths and trajectories are not synonymous, as previously noted. A path refers to a sequence of parallel or linear events linking the starting and ending points of a process, such as protein unfolding. On the other hand, a trajectory accounts for the time required for the protein to traverse that path, thereby defining the folding rate. The latter clarification is of paramount importance, as the thermodynamic hypothesis (1973), proposes that the amino acid sequence determines, in a proper environment, a protein's unique three-dimensional structure—a statement that remains valid even for metamorphic proteins (Vila, 2020)—regardless of how long it takes for the protein to fold or unfold. In other words, the amino acid sequence and its surrounding environment encode its folding pathways and consequently the protein's structure.
All in all, for more than half a century, chemists, biologists, mathematicians, and physicists have been unable to solve the protein folding problem analytically (Vila, 2023b). Consequently, as an alternative, we have employed a heuristic argument to both demonstrate the physical plausibility of the effective-trajectory conjecture and to illustrate the factors controlling the feasible folding pathways in a more intuitive manner, i.e., without relying on exhaustive mathematical rigor. The upcoming section will examine the validity range of the proposed solution, its relation to the Levinthal paradox, and its implications on critical issues in structural and evolutionary biology.
Avenues for Future Research
Knowledge of the single folding rate (τ
single) that characterizes the effective-trajectory solution of Eq. (1)—for a given environment—is crucial in practical applications. For example, it allows researchers to more precisely forecast changes in protein stability upon varying initial conditions (Vila, 2024b), such as those arising from single-point mutations (Vila, 2022). This analysis does not undermine the significance of understanding the pathway a protein could take, as this knowledge offers vital information regarding the key factors affecting protein evolvability (Vila, 2025). Unfortunately, as already mentioned, resolving this issue requires an analytical solution to Eq. (1), which—as previously indicated (Vila, 2023b)—presents a major unsolved challenge. This nuanced comprehension highlights the complexity of protein folding dynamics, suggesting that while multiple pathways might exist, all of them ultimately converge at a common destination: the native state, which must be attained at a given single folding rate (τ
single)—as determined by ΔG
single (as shown in
Figure 2). This convergence strengthens the idea that the folding process can be simplified—for practical applications—into an ‘effective trajectory.’ The latter aligns with the thorough analysis of the unfolding pathways of
chymotrypsin inhibitor 2 conducted by Lazaridis and Karplus (1997). Their simulations indicate that a “…
statistically preferred unfolding pathway…” is feasible, providing evidence that supports the proposed single effective-trajectory conjecture.
The whole analysis enables us, on the one hand, to bridge the effective-trajectory conjecture—originating from the validity of the least action principle—with the phenomenon of multiple pathways. This is a critical point, as it does not contradict the ‘new view’ of protein folding (Baldwin, 1994; Lazaridis & Karplus, 1997), which posits that there are multiple paths to the native state. On the other hand, it solves the Levinthal paradox (regarding the folding search timescale), as only those pathways exhibiting a common single folding rate (τsingle)—as provided by the least action principle—are allowed during the conformational search. This hypothesis dismantles the idea that the conformational search for the native state could be random. From this perspective, the paradox vanishes because each feasible protein folding pathway represents a solution.
We have thus far examined how proteins attain their native states regarding pathways and trajectories, but we have not addressed the mechanisms underlying that conformational search—such as the ‘funnel model’ (Dill & Chan, 1997) or the ‘foldon model’ (Bai et al., 1995)—as this falls outside the primary focus of our research. It is critical, however, to recognize these mechanisms' significance for comprehending a wide range of biological processes and illnesses. Thus, studying how these mechanisms affect protein pathways and trajectories may improve our understanding of protein folding and misfolding, which should be considered in future research.
Overall, future research in structural and evolutionary biology should prioritize solving Eq. (1). By solving this equation, researchers will gain crucial information that deepens our understanding of the main factors influencing protein folding and misfolding, as well as the feasible folding pathways. This knowledge will enable us to precisely and accurately determine how the amino acid sequence encodes its folding and, hence, which mechanism better represents the protein folding process.
Conclusions
This work yields three principal conclusions. Firstly, we have been able to provide a reasonable answer to a critical question in evolutionary biology: how a protein always attains—through a single or multiple folding pathways—its native state within a biologically acceptable timeframe. We have used the ‘least action principle’ as a physical framework to rationalize this process, i.e., to disclose why proteins must fold by following specific pathways—those that make the ‘action’ stationary. All these specific pathways have in common a unique folding rate determined mainly by the protein's marginal stability (ΔG)—which ultimately is set by the equilibrium of native and native-like conformations populating the transition state ensemble. This indicates that if multiple pathways are present, they must ‘all’ be characterized by the same ΔG to ensure proteins fold at the most efficient rate, regardless of which path nature chooses to reach the native state. This proposal implies that proteins will achieve the native state within a biologically acceptable timeframe. The latter is assured by the validity of the Anfinsen dogma which determines a constraint for the slowest single-folding rate (τsingle < ~seconds). In general, the principle of least action should be recognized as a fundamental physical law that allows us to understand how proteins fold in accordance with rigorous physical-mathematical rules and specific boundary conditions—such as limiting the folding pathways to those that render the ‘action’ stationary and whose slowest folding rate is capped at a few seconds. Secondly, we concluded, after analyzing how proteins fold from the perspective of the principle of least action, that the Anfinsen dogma—“…that is, that the native conformation is determined by the totality of interatomic interactions and hence by the amino acid sequence, in a given environment (Anfinsen, 1973)”—should be rewritten as follows: “The amino acid sequence determines—in a proper environment—their folding pathways and, hence, their native conformation.” This rewriting of the Anfinsen dogma highlights the link that exists between the amino acid sequence and the native structure. The link specifically refers to the folding pathways, which are determined by all interatomic interactions, in accordance with the principle of least action (see Eq. 1). Additionally, it emphasizes that the amino acid sequence dictates the protein's native state rather than the latter being determined by the former. At first glance, the difference may seem subtle, but it is significant: the amino acid sequence alone is sufficient to determine—in a proper environment—the lowest accessible free-energy conformation in solution (also known as the protein native state). The analysis also shows us that all feasible folding trajectories share a common feature: the time required to traverse any feasible folding path—represented by the effective folding trajectory—must align with the protein-folding rate. This folding rate, by definition, may be the same for proteins with different sequences, lengths, or both, as it is primarily determined by the protein's marginal stability. Finally, our analysis, grounded in the principle of least action, indicates that neither the question of whether ‘there are pathways for protein folding’ (Levinthal, 1968) should have been viewed as a challenge nor the timescale for conformational search (Levinthal, 1969) as a paradox. It is important to note that the principle of least action, introduced by Maupertuis (Fee, 1941), predates Levinthal's query by more than 200 years. Therefore, had researchers considered this principle from the outset, it might have prompted them to propose—despite their limited understanding of the forces at play in the folding process at that time—that nature tends to pursue the most efficient pathways for folding rather than exploring them randomly, as suggested by some early attempts to tackle the protein folding problem. In conclusion, it is crucial to underscore the significance of the principle of least action for understanding both the genesis of the Levinthal paradox and its solution. At this point, it is worth noting that the real challenge is understanding the mechanisms that drive proteins to fold into their native states, and this question remains unanswered after more than 50 years of research in structural biology. The reason for these difficulties is that the central question of the protein folding problem remains unresolved: specifically, “how a sequence of amino acids—in a proper environment—encodes its folding pathways.” However, advancements in the state of the art of computational modeling, simulations, and experimental techniques are gradually shedding light on what these pathways could be, providing optimism for potential breakthroughs in our understanding of such essential biological processes.
Founding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Ethical approval
Not applicable.
Acknowledgments
The author acknowledges support from the Institute of Applied Mathematics San Luis (IMASL), the National University of San Luis (UNSL), and the National Research Council of Argentina (CONICET). I want to extend my gratitude to Pablo Garay for his assistance in preparing Figure 3.
Conflicts of Interest
The author declares no competing interest.
References
- Akmal, A; Muñoz, V. The nature of the free energy barriers to two-state folding. Proteins 2004, 57, 142–52. [Google Scholar] [CrossRef]
- Anfinsen, CB; Scheraga, HA. Experimental and theoretical aspects of protein folding. Adv Protein Chem 1975, 29, 205–300. [Google Scholar]
- Anfinsen, CB. Principles that govern the folding of protein chains. Science 1973, 181, 223–230. [Google Scholar] [CrossRef]
- Arnautova, YA; Jagielska, A; Scheraga, HA. A new force field (ECEPP-05) for peptides, proteins, and organic molecules. J Phys Chem B 2006, 110(10), 5025–5044. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y; Sosnick, TR; Mayne, L; Englander, SW. Protein folding intermediates: native-state hydrogen exchange. Science 1995, 269(5221), 192–197. [Google Scholar] [CrossRef]
- Bakk, A; Høye, JS; Hansen, A; Sneppen, K; Jensen, MH. Pathways in two-state protein folding. Biophys J 2000, 79(5), 2722–2727. [Google Scholar] [CrossRef]
- Baldwin, RL. Protein folding. Matching speed and stability. Nature 1994, 369(6477), 183–184. [Google Scholar] [CrossRef]
- Baldwin, RL. The nature of protein folding pathways: the classical versus the new view. J Biomol NMR 1995, 5(2), 103–109. [Google Scholar] [CrossRef] [PubMed]
- Ben-Naim, A. Levinthal’s Paradox Revisited and Dismissed. Open Journal of Biophysics 2012, 2, 23–32. [Google Scholar] [CrossRef]
- Bogatyreva, NS; Finkelstein, AV. Cunning simplicity of protein folding landscapes. Protein Eng 2001, 14(8), 521–523. [Google Scholar] [CrossRef]
- Chang, L; Perez, A. Rapid estimation of protein folding pathways from sequence alone using AlphaFold2. Nat Commun 2025. [Google Scholar] [CrossRef]
- Coopersmith, J. The Lazy Universe: An Introduction to the Principle of Least Action; Oxford University Press, 2017. [Google Scholar] [CrossRef]
- Creighton, TE. The problem of how and why proteins adopt folded conformations. The Journal of Physical Chemistry 1985, 89(12), 2452–2459. [Google Scholar] [CrossRef]
- Dill, KA; Chan, HS. From Levinthal to pathways to funnels. Nat Struct Biol 1997, 4(1), 10–19. [Google Scholar] [CrossRef] [PubMed]
- Ding, F; Dokholyan, NV; Buldyrev, SV; Stanley, HE; Shakhnovich, EI. Direct molecular dynamics observation of protein folding transition state ensemble. Biophys J 2002, 83(6), 3525–3532. [Google Scholar] [CrossRef]
- Dishman, AF; Volkman, BF. Unfolding the Mysteries of Protein Metamorphosis. ACS Chemical Biology 2018, 13(6), 1438–1446. [Google Scholar] [CrossRef]
- Dobson, CM; Šali, A; Karplus, M. Protein Folding: A Perspective from Theory and Experiment. Angew Chem Int Ed Engl 1998, 37(7), 868–893. [Google Scholar] [CrossRef]
- Eaton, WA; Wolynes, PG. Theory, simulations, and experiments show that proteins fold by multiple pathways. Proc Natl Acad Sci USA 2017, 114(46), E9759–E9760. [Google Scholar] [CrossRef]
- Eaton, WA. Modern Kinetics and Mechanism of Protein Folding: A Retrospective. J Phys Chem B 2021, 19 125, 3452–3467. [Google Scholar] [CrossRef]
- Englander, SW; Mayne, L; Krishna, MM. Protein folding and misfolding: mechanism and principles. Q Rev Biophys 2007, 40(4), 287–326. [Google Scholar] [CrossRef]
- Englander, SW; Mayne, L. Reply to Eaton and Wolynes: How do proteins fold? Proc Natl Acad Sci USA 2017, 114(46), E9761–E9762. [Google Scholar] [CrossRef] [PubMed]
- Englander, SW; Mayne, L. The nature of protein folding pathways. Proc Natl Acad Sci USA 2014, 111(45), 15873–15880. [Google Scholar] [CrossRef] [PubMed]
- Englander, SW. Protein folding intermediates and pathways studied by hydrogen exchange. Annu Rev Biophys Biomol Struct 2000, 29, 213–38. [Google Scholar] [CrossRef]
- Fakhoury, Z; Sosso, GC; Habershon, S. Generating Protein Folding Trajectories Using Contact-Map-Driven Directed Walks. J Chem Inf Model 2023, 63(7), 2181–2195. [Google Scholar] [CrossRef]
- Fee, J. Maupertuis, and the Principle of Least Action. The Scientific Monthly, JSTOR, 1941; 52, pp. 496–503. Available online: http://www.jstor.org/stable/17253.
- Feynman, RP; Leighton, RB; Sands, M. The Feynman Lectures on Physics; Addison-Wesley: Reading, MA, 1963; Vol. II, p. Chap. 19. [Google Scholar]
- Feynman, RP. The Principle of Least Action in Quantum Mechanics. Feynman’s Thesis—A New Approach to Quantum Theory, World Scientific, 2005; pp. pp 1–69. [Google Scholar]
- Finkelstein, AV; Badretdinov, AYa. Rate of protein folding near the point of thermodynamic equilibrium between the coil and the most stable chain fold. Fold Des 1997, 2(2), 115–121. [Google Scholar] [CrossRef]
- Finkelstein, AV; Garbuzynskiy, SO. Levinthal’s question answered again? J Biomol Struct Dyn. 2013, 31(9), 1013–1015. [Google Scholar] [CrossRef]
- Garbuzynskiy, SO; Ivankov, DN; Bogatyreva, NS; Finkelstein, AV. Golden triangle for folding rates of globular proteins. Proc Natl Acad Sci USA 2013, 110(1), 147–150. [Google Scholar] [CrossRef]
- Glyakina, AV; Galzitskaya, OV. How Quickly Do Proteins Fold and Unfold, and What Structural Parameters Correlate with These Values? Biomolecules 2020, 10, 197. [Google Scholar] [CrossRef] [PubMed]
- Gromiha, MM. A statistical model for predicting protein folding rates from amino acid sequence with structural class information. J Chem Inf Model 2005, 45(2), 494–501. [Google Scholar] [CrossRef] [PubMed]
- Hanc, J; Taylor, EF. From conservation of energy to the principle of least action: a story line. Am J Phys 2004, 72, 514–521. [Google Scholar] [CrossRef]
- Harrington, WF; Schellman, JA. Evidence for the instability of hydrogen-bonded peptide structures in water, based on studies of ribonuclease and oxidized ribonuclease. C R Trav Lab Carlsberg Chim 1956, 30(3), 21–43. [Google Scholar]
- Honig, B. Protein folding: from the Levinthal paradox to structure prediction. J Mol Biol 1999, 293(2), 283–293. [Google Scholar] [CrossRef]
- Huang, LT; Gromiha, MM. Analysis and prediction of protein folding rates using quadratic response surface models. J Comput Chem 2008, 29(10), 1675–1683. [Google Scholar] [CrossRef]
- Itzhaki, LS; Otzen, DE; Fersht, AR. The structure of the transition state for folding of chymotrypsin inhibitor 2 analyzed by protein engineering methods: evidence for a nucleation-condensation mechanism for protein folding. J Mol Biol 1995, 254, 260–288. [Google Scholar] [CrossRef]
- Ivankov, DN; Finkelstein, AV. Solution of Levinthal’s Paradox and a Physical Theory of Protein Folding Times. Biomolecules 2020, 10(2), 250. [Google Scholar] [CrossRef]
- Jackson, SE. How do small single-domain proteins fold? Fold Des 1998, 3(4), R81–R91. [Google Scholar] [CrossRef]
- Kaila, VRI; Annila, A. Natural selection for least action. Proc R Soc A 2008, 464, 3055–3070. [Google Scholar] [CrossRef]
- Karplus, M. Behind the folding funnel diagram. Nat Chem Biol 2011, 7(7), 401–404. [Google Scholar] [CrossRef] [PubMed]
- Karplus, M. The Levinthal paradox: yesterday and today. Fold Des 1997, 2(4), S69–S75. [Google Scholar] [CrossRef]
- Kikuchi, T. Decoding an Amino Acid Sequence to Extract Information on Protein Folding. Molecules 2022, 27(9), 3020. [Google Scholar] [CrossRef]
- Kuwajima, K. The Molten Globule, and Two-State vs. Non-Two-State Folding of Globular Proteins. Biomolecules 2020, 10, 407. [Google Scholar] [CrossRef] [PubMed]
- Landau, LD; Lifshitz, EM. Course of Theoretical Physics. In Mechanics, 3rd ed.; Butterworth-Heinemann: Oxford, 1982; Volume 1. [Google Scholar]
- Landau, LD; Lifshitz, EM. Course of Theoretical Physics. In The Classical Theory of Fields, 4th ed.; Butterworth-Heinemann: Oxford, 1987; Volume 2. [Google Scholar]
- Lange, OF; Lakomek, N-A; Farès, C; Schröder, GF; Walter, KFA; Becker, S; Meiler, J; Grubmüller, H; Griesinger, C; de Groot, BL. Recognition Dynamics Up to Microseconds Revealed from an RDC-Derived Ubiquitin Ensemble in Solution. Science 2008, 13, 1471–1475. [Google Scholar] [CrossRef]
- Lapidus, LJ. The road less traveled in protein folding: evidence for multiple pathways. Curr Opin Struct Biol 2021, 66, 83–88. [Google Scholar] [CrossRef]
- Lazaridis, T; Karplus, M. ‘New view’ of protein folding reconciled with the old through multiple unfolding simulations. Science 1997, 278(5345), 1928–1931. [Google Scholar] [CrossRef]
- Levinthal, C. Are There Pathways for Protein Folding? Journal de Chimie Physique 1968, 65, 44–45. [Google Scholar] [CrossRef]
- Levinthal, C. How to fold graciously. In Mossbauer Spectroscopy in Biological Systems, Proceedings of a Meeting held at Allerton House, Monticello, Illinois; Debrunner, P, Tsibris, JCM, Münck, E, Eds.; University of Illinois Press: Urbana, 1969; p. 22. [Google Scholar]
- Li, Y; Gong, H. Identifying a Feasible Transition Pathway between Two Conformational States for a Protein. J Chem Theory Comput 2022, 18, 4529–4543. [Google Scholar] [CrossRef]
- Martin, AO; Vila, JA. The Marginal Stability of Proteins: How the Jiggling and Wiggling of Atoms is Connected to Neutral Evolution. Journal of Molecular Evolution 2020, 88, 424–426. [Google Scholar] [CrossRef]
- Martin, OA; Vorobjev, Y; Scheraga, HA; Vila, JA. Outline of an experimental design aimed to detect a protein A mirror image in solution. PeerJ Phys Chem 2019, 1, e2. [Google Scholar] [CrossRef] [PubMed]
- Martinez, L. Introducing the Levinthal’s Protein Folding Paradox and Its Solution. J Chem Educ 2014, 91(11), 1918–1923. [Google Scholar] [CrossRef]
- Matouschek, A; Kellis, JT, Jr.; Serrano, L; Fersht, AR. Mapping the transition state and pathway of protein folding by protein engineering. Nature 1989, 340, 122–126. [Google Scholar] [CrossRef] [PubMed]
- Mayor, U; Johnson, CM; Daggett, V; Fersht, AR. Protein folding and unfolding in microseconds to nanoseconds by experiment and simulation. Proc Natl Acad Sci USA 2000, 97, 13518–13522. [Google Scholar] [CrossRef]
- McCammon, JA; Gelin, BR; Karplus, M. Dynamics of folded proteins. Nature 1977, 267, 585–590. [Google Scholar] [CrossRef] [PubMed]
- McCammon, JA. A speed limit for protein folding. Proc Natl Acad Sci USA 1996, 93, 11426–11427. [Google Scholar] [CrossRef] [PubMed]
- Muñoz, V; Cerminara, M. When fast is better: protein folding fundamentals and mechanisms from ultrafast approaches. Biochem J 2016, 473, 2545–2559. [Google Scholar] [CrossRef]
- Murzin, A.G. Biochemistry. Metamorphic proteins. Science 2008, 320, 1725–1726. [Google Scholar] [CrossRef] [PubMed]
- Neupane, K.; Manuel, A.; Woodside, M. Protein folding trajectories can be described quantitatively by one-dimensional diffusion over measured energy landscapes. Nature Phys 2016, 12, 700–703. [Google Scholar] [CrossRef]
- Pande, VS; Grosberg, AY; Tanaka, T; Rokhsar, DS. Pathways for protein folding: is a new view needed? Curr Opin Struct Biol 1998, 8(1), 68–79. [Google Scholar] [CrossRef]
- Plaxco, KW; Simons, KT; Ruczinski, I; Baker, D. Topology, stability, sequence, and length: defining the determinants of two-state protein folding kinetics. Biochemistry 2000, 37, 11177–11183. [Google Scholar] [CrossRef]
- Rooman, M; Dehouck, Y; Kwasigroch, JM; Biot, C; Gilis, D. What is paradoxical about Levinthal paradox? J Biomol Struct Dyn 2002, 20(3), 327–329. [Google Scholar] [CrossRef]
- Šali, A; Shakhnovich, E; Karplus, M. How does a protein fold? Nature 1994, 369(6477), 248–251. [Google Scholar] [CrossRef]
- Shakhnovich, E. Protein folding thermodynamics and dynamics: where physics, chemistry, and biology meet. Chem Rev 2006, 106, 1559–1588. [Google Scholar] [CrossRef]
- Simmons, W; Weiner, JL. The principle of stationary action in biophysics: stability in protein folding. J Biophys 2013, 2013, 697529. [Google Scholar] [CrossRef] [PubMed]
- Stiller, JB; Otten, R; Häussinger, D; Rieder, PS; Theobald, DL; Kern, D. Structure determination of high-energy states in a dynamic protein ensemble. Nature 2022, 603(7901), 528–535. [Google Scholar] [CrossRef]
- Thirumalai, D. From Minimal Models to Real Proteins: Time Scales for Protein Folding Kinetics. Journal de Physique I 1995, 5(11), 1457–1467. [Google Scholar] [CrossRef]
- Tsuboyama, K; Dauparas, J; Chen, J; Laine, E; Mohseni Behbahani, Y; Weinstein, JJ; Mangan, NM; Ovchinnikov, S; Rocklin, GJ. Mega-scale experimental analysis of protein folding stability in biology and design. Nature 2023, 620(7973), 434–444. [Google Scholar] [CrossRef] [PubMed]
- Vendruscolo, M; Paci, E; Dobson, CM; Karplus, M. Rare Fluctuations of Native Proteins Sampled by Equilibrium Hydrogen Exchange. J Am Chem Soc 2003, 125(51), 15686–15687. [Google Scholar] [CrossRef]
- Vila, JA; Ripoll, DR; Scheraga, HA. Atomically detailed folding simulation of the B domain of staphylococcal protein A from random structures. Proc Natl Acad Sci USA 2003, 100(25), 14812–14816. [Google Scholar] [CrossRef] [PubMed]
- Vila, JA. Analysis of proteins in the light of mutations. Eur Biophys J 2024b, 53(5-6), 255–265. [Google Scholar] [CrossRef]
- Vila, JA. Factors controlling protein evolvability—at the molecular scale. Eur Biophys J 2025, 55, 101–109. [Google Scholar] [CrossRef]
- Vila, JA. Forecasting the upper bound free energy difference between protein native-like structures. Physica A 2019, 533, 122053. [Google Scholar] [CrossRef]
- Vila, JA. Metamorphic Proteins in Light of Anfinsen's Dogma. J Phys Chem Lett 2020, 11(13), 4998–4999. [Google Scholar] [CrossRef]
- Vila, JA. Protein Evolution upon Point Mutations. ACS Omega 2022, 7, 14371–14376. [Google Scholar] [CrossRef]
- Vila, JA. Protein folding rate evolution upon mutations. Biophys Rev 2023a, 15, 661–669. [Google Scholar] [CrossRef]
- Vila, JA. Rethinking the protein folding problem from a new perspective. Eur Biophys J 2023b, 52(3), 189–193. [Google Scholar] [CrossRef]
- Vila, JA. The origin of mutational epistasis. Eur Biophys J 2024a, 53, 473–480. [Google Scholar] [CrossRef]
- Vila, JA. Thoughts on the Protein’s Native State. J Phys Chem Lett 2021, 12, 5963–5966. [Google Scholar] [CrossRef] [PubMed]
- Wolynes, PG; Onuchic, JN; Thirumalai, D. Navigating the folding routes. Science 1995, 267(5204), 1619–1620. [Google Scholar] [CrossRef] [PubMed]
- Wolynes, PG. Evolution, energy landscapes, and the paradoxes of protein folding. Biochimie 2015, 119, 218–230. [Google Scholar] [CrossRef] [PubMed]
- Zana, R. On the rate determining step for helix propagation in the helix-coil transition of polypeptides in solution. Biopolymers 1975, 14, 2425–2428. [Google Scholar] [CrossRef]
- Zhao, K; Xia, Y; Zhang, F; Zhou, X; Li, SZ; Zhang, G. Protein structure and folding pathway prediction based on remote homologs recognition using PAthreader. Commun Biol 2023, 6(1), 243. [Google Scholar] [CrossRef]
- Zhao, K; Zhao, P; Wang, S; Xia, Y; Zhang, G. FoldPAthreader: predicting protein folding pathway using a novel folding force field model derived from the known protein universe. Genome Biol 2024, 25(1), 152. [Google Scholar] [CrossRef]
- Zwanzig, R; Szabo, A; Bagchi, B. Levinthal’s paradox. Proc Natl Acad Sci USA 1992, 89, 20–22. [Google Scholar] [CrossRef] [PubMed]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).