Submitted:
09 March 2026
Posted:
12 March 2026
You are already at the latest version
Abstract
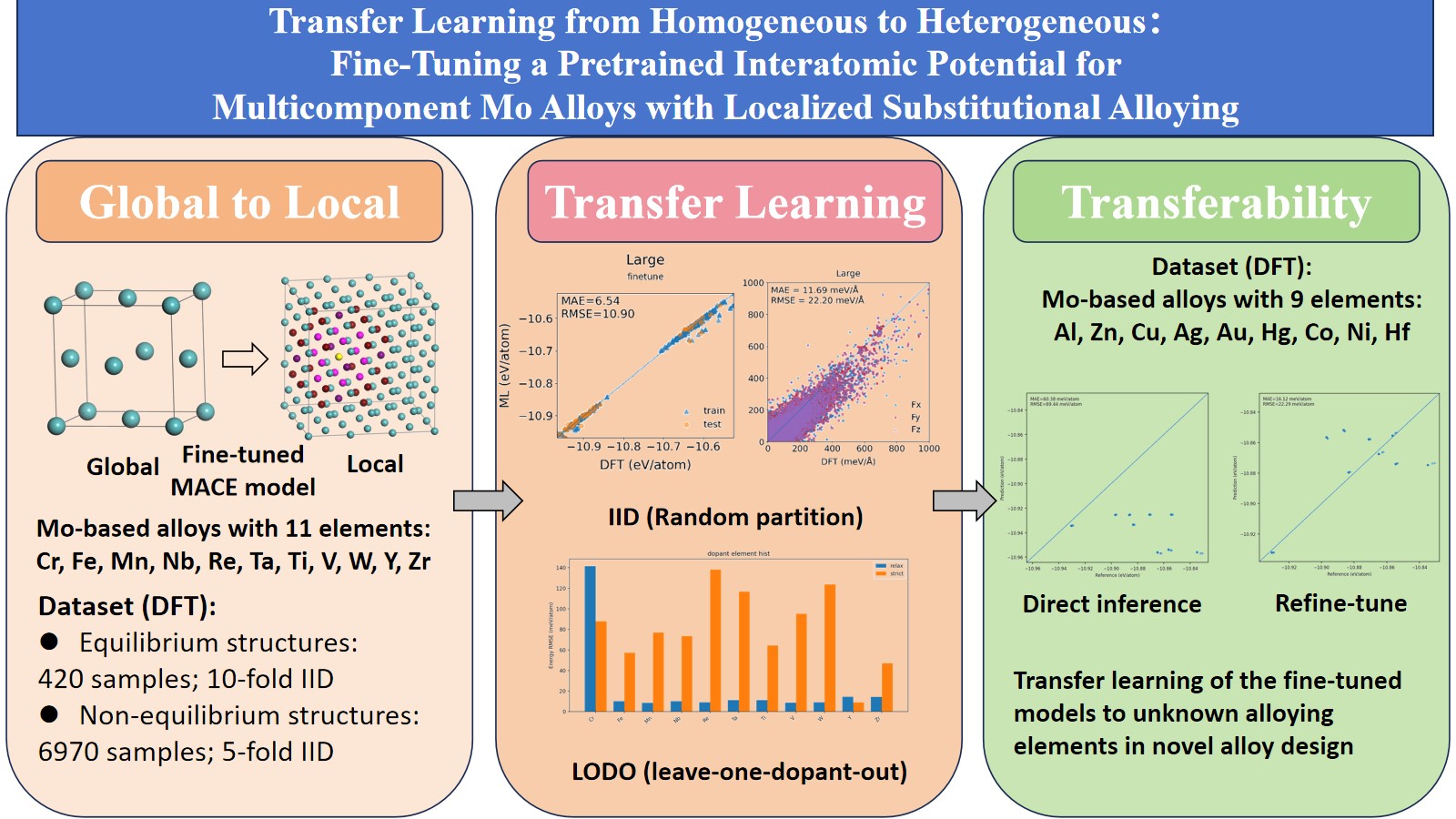
Keywords:
1. Introduction
2. Methods and Datasets
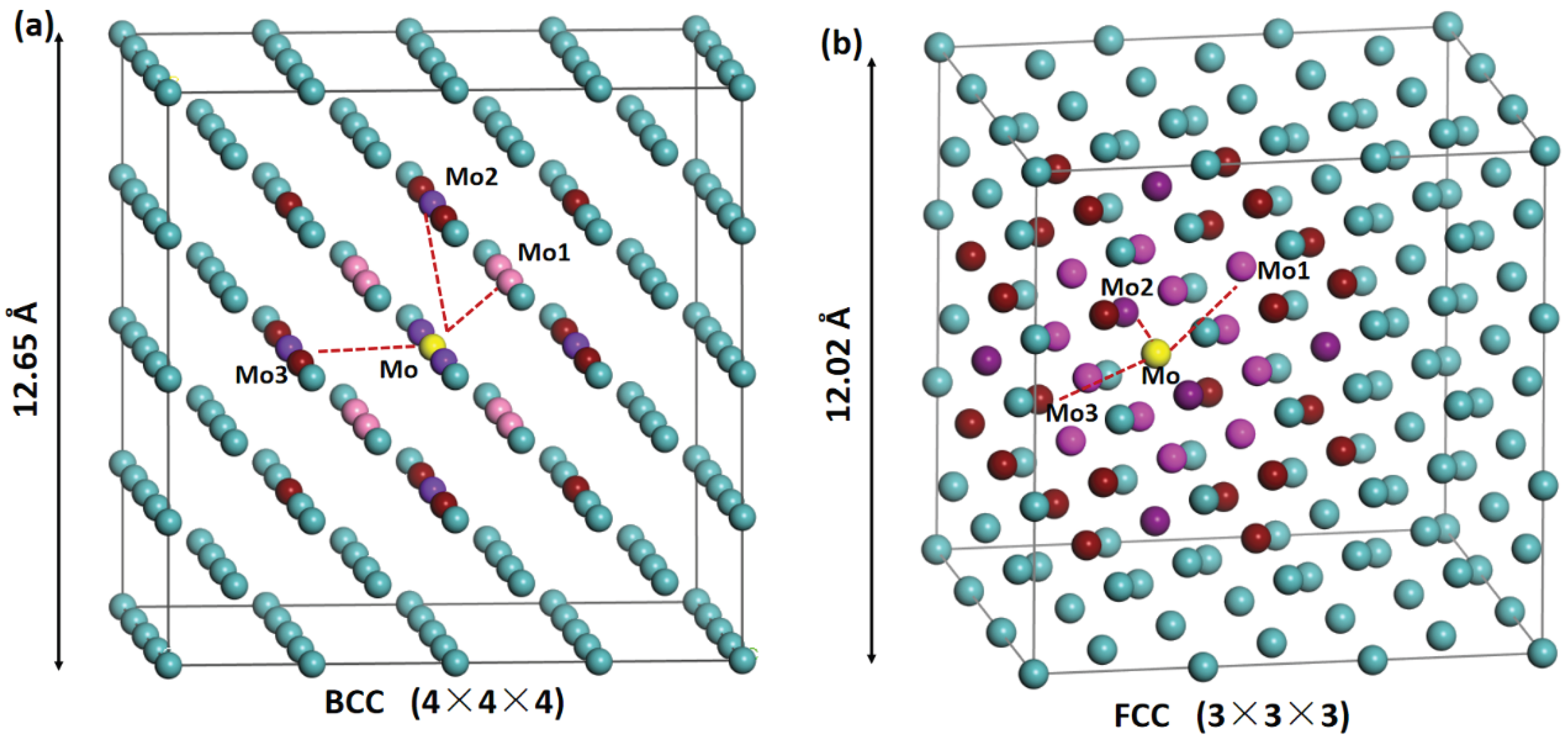
2.1. Fine-Tuning Dataset of Doped Mo Alloys
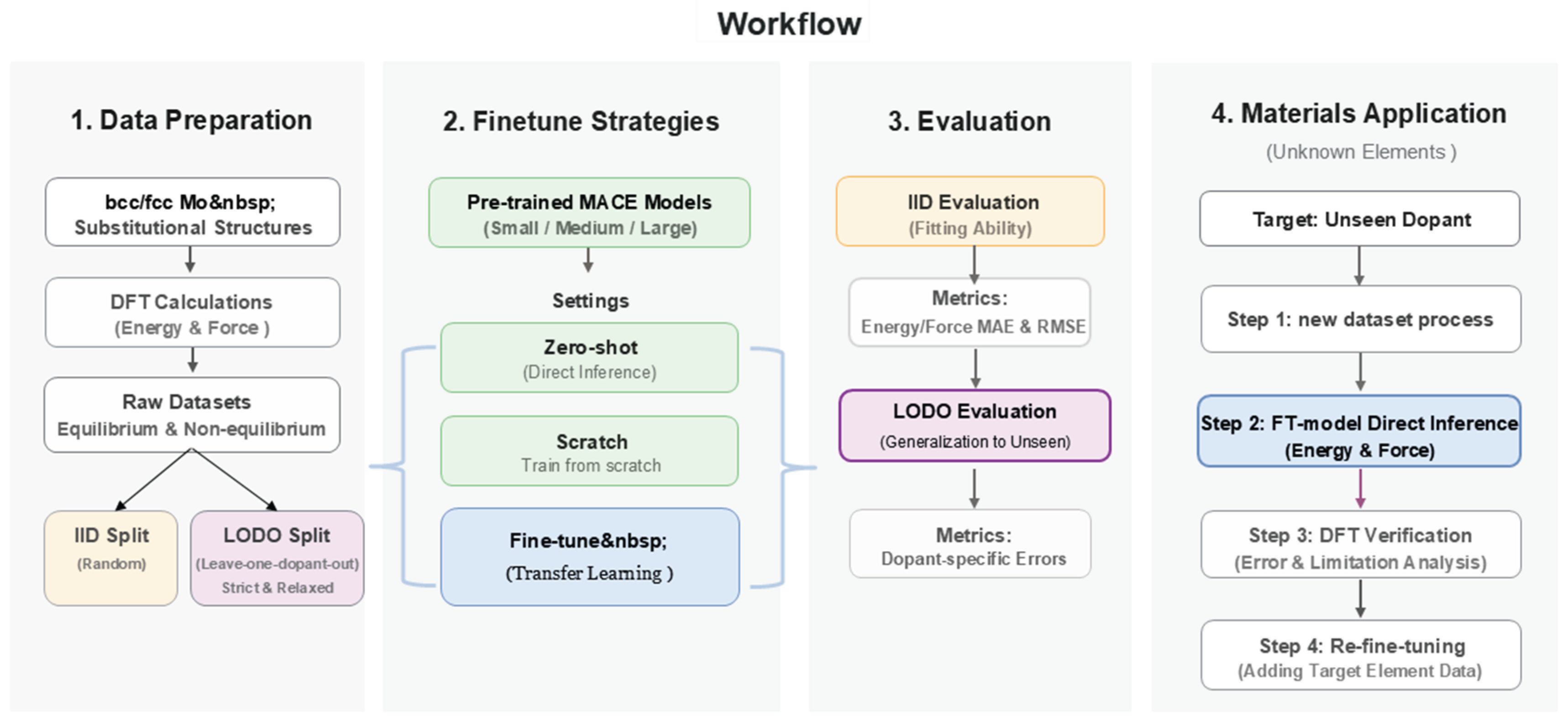
2.2. Model Training Protocols and Evaluation Metrics
3. Results and Discussion
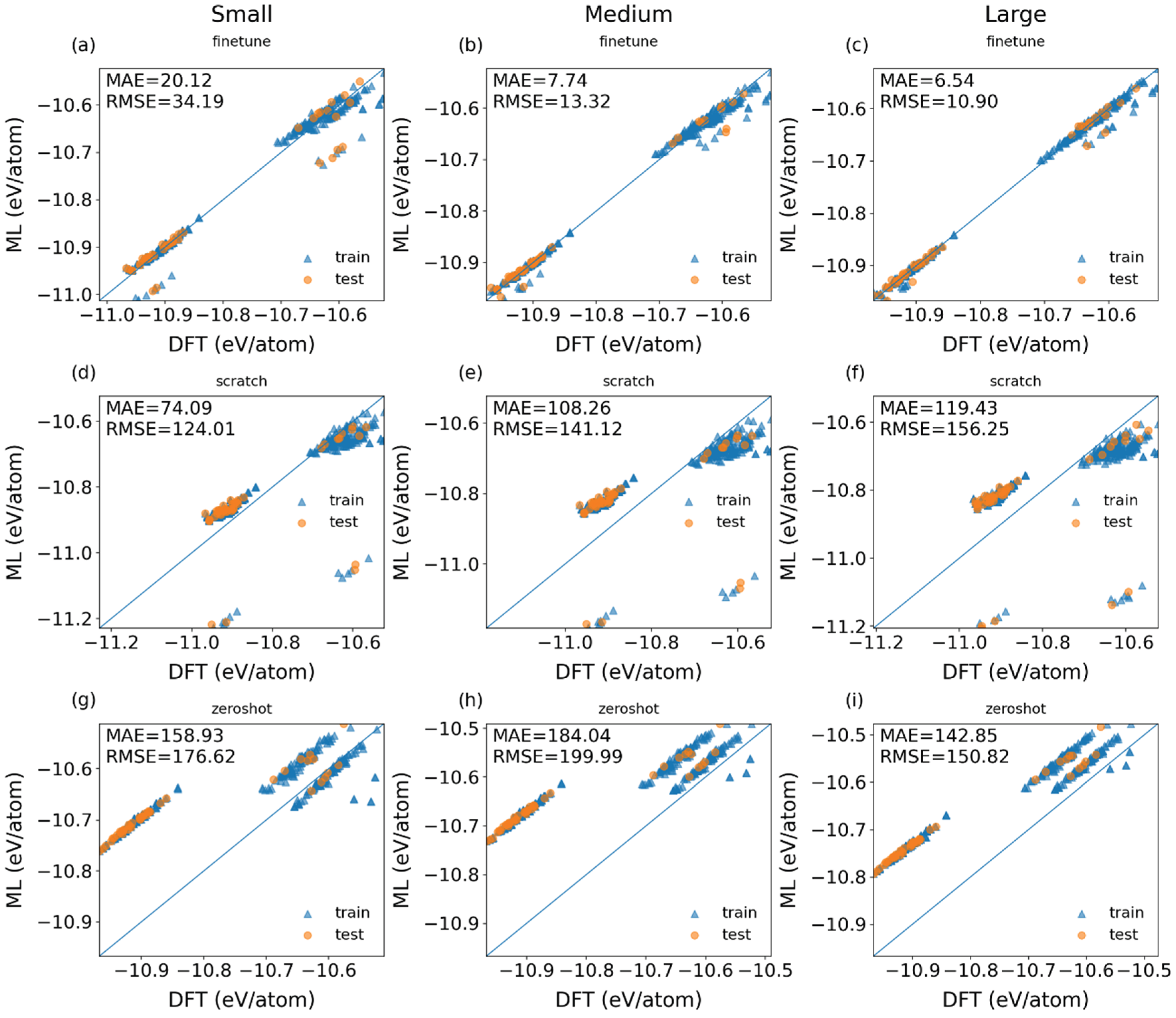
3.1. Construction of Fine-Tuned Models with Randomly Partitioned Datasets
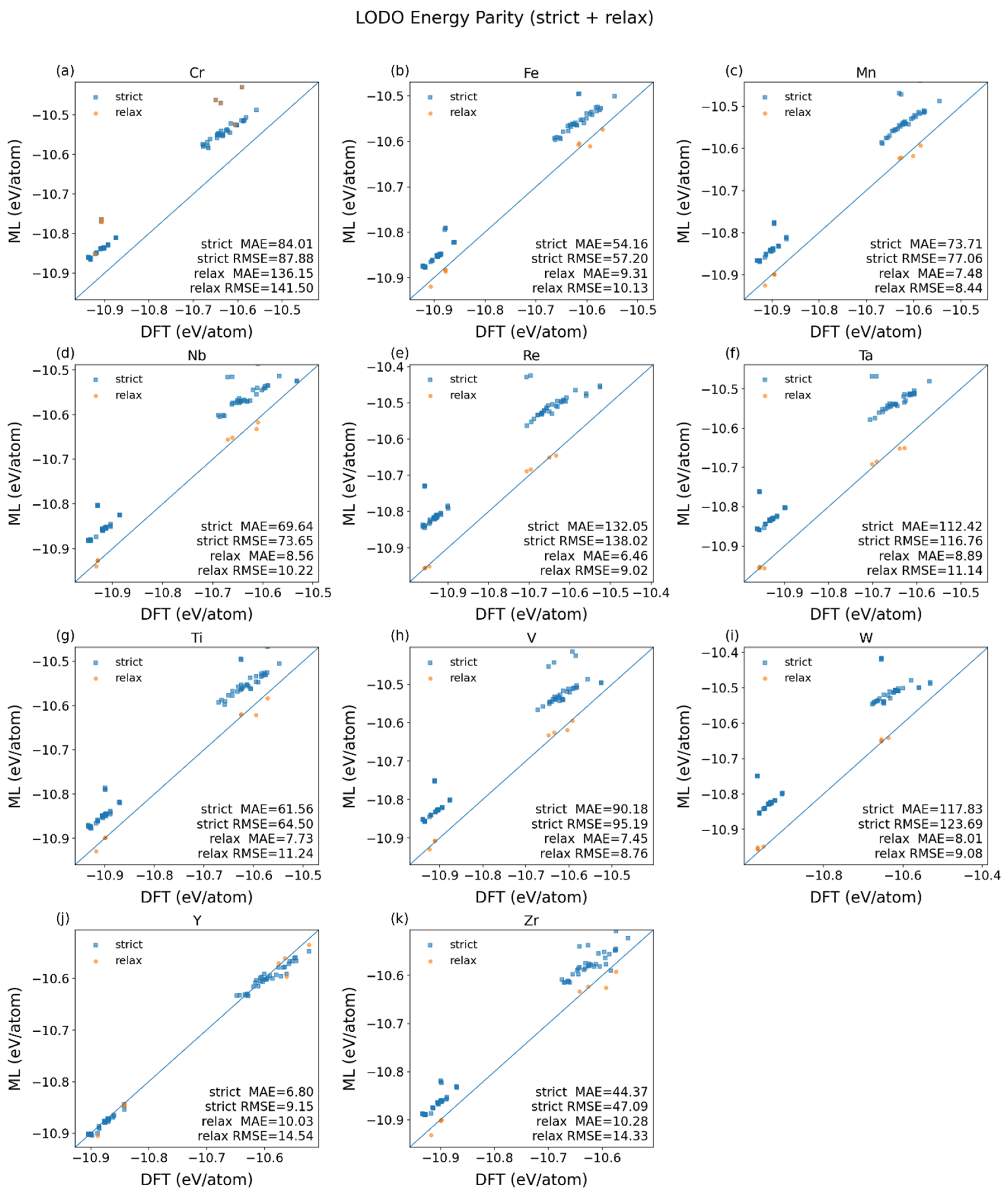
3.2. Verification of Fine-Tuned Models with Leave-One-Dopant-Out Partitioned Datasets
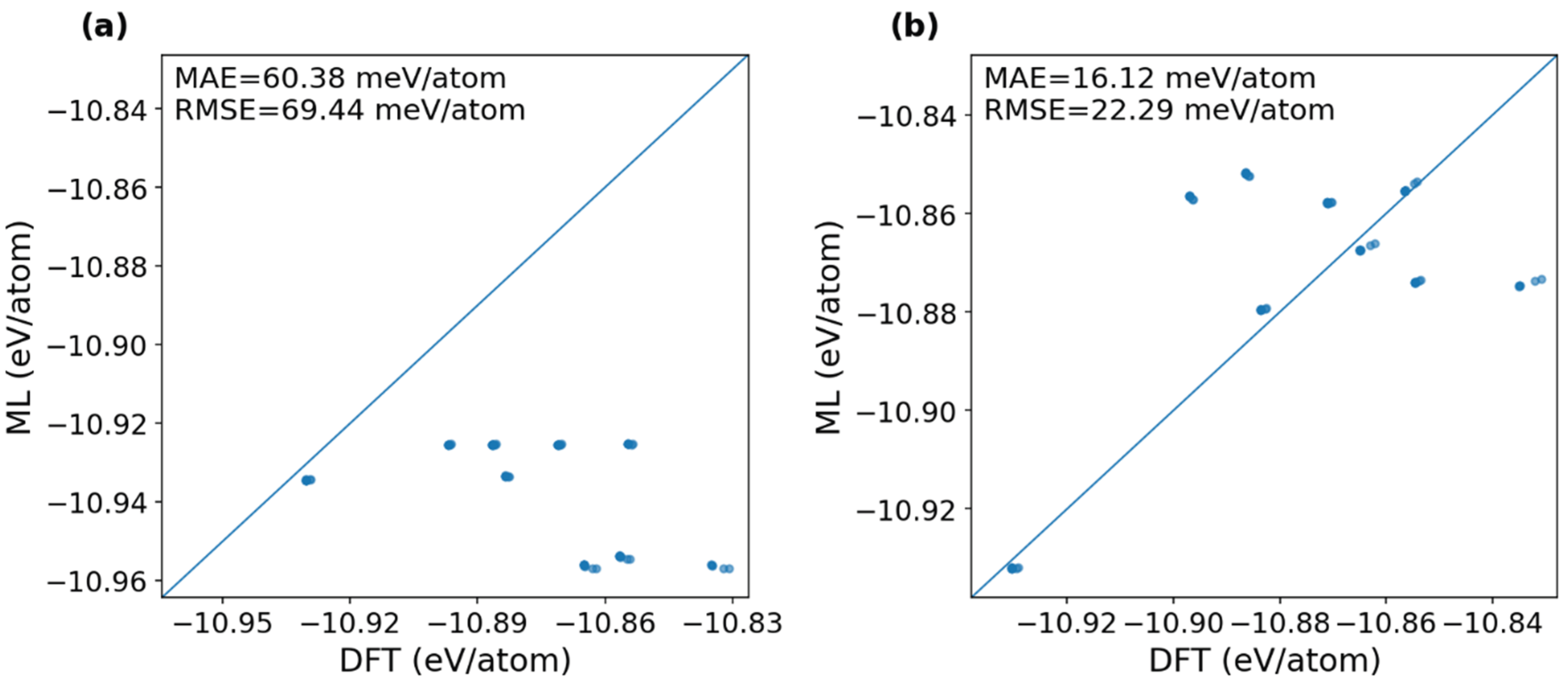
3.3. Transferability of Fine-Tuned Models to Unknown Alloying Elements
4. Conclusions
Supplementary Materials
CRediT authorship contribution statement
Acknowledgments
Declaration of Competing Interest
Code Availability
References
- Xu, H.; Cui, T.; Tang, C.; Ma, J.; Zhou, D.; Li, Y.; Gao, X.; Gong, X.; Ouyang, W.; Zhang, S. Evidential Deep Learning for Interatomic Potentials. Nature Communications 2025. [Google Scholar] [CrossRef]
- Yang, Z.; Wang, X.; Li, Y.; Lv, Q.; Chen, C. Y.-C.; Shen, L. Efficient Equivariant Model for Machine Learning Interatomic Potentials. NPJ Computational Materials 2025, 11(1), 49. [Google Scholar] [CrossRef]
- Ran, N.; Yin, L.; Qiu, W.; Liu, J. Recent Advances in Machine Learning Interatomic Potentials for Cross-Scale Computational Simulation of Materials. Sci. China Mater. 2024, 67(4), 1082–1100. [Google Scholar] [CrossRef]
- Behler, J.; Parrinello, M. Generalized Neural-Network Representation of High-Dimensional Potential-Energy Surfaces. Phys. Rev. Lett. 2007, 98(14), 146401. [Google Scholar] [CrossRef]
- Bartók, A. P.; Payne, M. C.; Kondor, R.; Csányi, G. Gaussian Approximation Potentials: The Accuracy of Quantum Mechanics, without the Electrons. Phys. Rev. Lett. 2010, 104(13), 136403. [Google Scholar] [CrossRef]
- Bartók, A. P.; Kondor, R.; Csányi, G. On Representing Chemical Environments. Phys. Rev. B 2013, 87(18), 184115. [Google Scholar] [CrossRef]
- Thompson, A. P.; Swiler, L. P.; Trott, C. R.; Foiles, S. M.; Tucker, G. J. Spectral Neighbor Analysis Method for Automated Generation of Quantum-Accurate Interatomic Potentials. Journal of Computational Physics 2015, 285, 316–330. [Google Scholar] [CrossRef]
- Shapeev, A. V. Moment Tensor Potentials: A Class of Systematically Improvable Interatomic Potentials. Multiscale Model. Simul. 2016, 14(3), 1153–1173. [Google Scholar] [CrossRef]
- Drautz, R. Atomic Cluster Expansion for Accurate and Transferable Interatomic Potentials. Phys. Rev. B 2019, 99(1), 014104. [Google Scholar] [CrossRef]
- Schütt, K.; Kindermans, P.-J.; Sauceda Felix, H. E.; Chmiela, S.; Tkatchenko, A.; Müller, K.-R. Schnet: A Continuous-Filter Convolutional Neural Network for Modeling Quantum Interactions. Advances in neural information processing systems 2017, 30. [Google Scholar]
- Gasteiger, J.; Groß, J.; Günnemann, S. Directional Message Passing for Molecular Graphs. arXiv 2022. [Google Scholar] [CrossRef]
- Gasteiger, J.; Becker, F.; Günnemann, S. Gemnet: Universal Directional Graph Neural Networks for Molecules. Advances in neural information processing systems 2021, 34, 6790–6802. [Google Scholar]
- Schütt, K.; Unke, O.; Gastegger, M. Equivariant Message Passing for the Prediction of Tensorial Properties and Molecular Spectra. In International conference on machine learning; 2021, PMLR; pp. 9377–9388.
- Batzner, S.; Musaelian, A.; Sun, L.; Geiger, M.; Mailoa, J. P.; Kornbluth, M.; Molinari, N.; Smidt, T. E.; Kozinsky, B. E (3)-Equivariant Graph Neural Networks for Data-Efficient and Accurate Interatomic Potentials. Nature communications 2022, 13(1), 2453. [Google Scholar] [CrossRef]
- Batatia, I.; Kovacs, D. P.; Simm, G.; Ortner, C.; Csányi, G. MACE: Higher Order Equivariant Message Passing Neural Networks for Fast and Accurate Force Fields. Advances in neural information processing systems 2022, 35, 11423–11436. [Google Scholar]
- Gasteiger, J.; Shuaibi, M.; Sriram, A.; Günnemann, S.; Ulissi, Z.; Zitnick, C. L.; Das, A. GemNet-OC: Developing Graph Neural Networks for Large and Diverse Molecular Simulation Datasets. arXiv 2022. [Google Scholar] [CrossRef]
- Chen, C.; Ong, S. P. A Universal Graph Deep Learning Interatomic Potential for the Periodic Table. Nature Computational Science 2022, 2(11), 718–728. [Google Scholar] [CrossRef] [PubMed]
- Deng, B.; Zhong, P.; Jun, K.; Riebesell, J.; Han, K.; Bartel, C. J.; Ceder, G. CHGNet as a Pretrained Universal Neural Network Potential for Charge-Informed Atomistic Modelling. Nature Machine Intelligence 2023, 5(9), 1031–1041. [Google Scholar] [CrossRef]
- Batatia, I.; Benner, P.; Chiang, Y.; Elena, A. M.; Kovács, D. P.; Riebesell, J.; Advincula, X. R.; Asta, M.; Avaylon, M.; Baldwin, W. J. A Foundation Model for Atomistic Materials Chemistry. The Journal of chemical physics 2025, 163(18). [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Hu, C.; Zhou, Y.; Liu, X.; Shi, Y.; Li, J.; Li, G.; Chen, Z.; Chen, S.; Zeni, C. A Deep Learning Atomistic Model Across Elements, Temperatures and Pressures. arXiv 2024, arXiv:2405.04967. [Google Scholar] [CrossRef]
- Ma, J.; Fu, X.; Xie, W.; Hu, P. From Pretrained to Precision: Fine-Tuning Universal Interatomic Potentials for Accurate Catalytic Reaction Simulations. J. Chem. Theory Comput. 2026, 22(4), 1920–1930. [Google Scholar] [CrossRef]
- Zhang, L.; Han, J.; Wang, H.; Car, R.; E, W. Deep Potential Molecular Dynamics: A Scalable Model with the Accuracy of Quantum Mechanics. Phys. Rev. Lett. 2018, 120(14), 143001. [Google Scholar] [CrossRef]
- Musaelian, A.; Batzner, S.; Johansson, A.; Sun, L.; Owen, C. J.; Kornbluth, M.; Kozinsky, B. Learning Local Equivariant Representations for Large-Scale Atomistic Dynamics. Nature Communications 2023, 14(1), 579. [Google Scholar] [CrossRef]
- Chanussot, L.; Das, A.; Goyal, S.; Lavril, T.; Shuaibi, M.; Riviere, M.; Tran, K.; Heras-Domingo, J.; Ho, C.; Hu, W.; Palizhati, A.; Sriram, A.; Wood, B.; Yoon, J.; Parikh, D.; Zitnick, C. L.; Ulissi, Z. Open Catalyst 2020 (OC20) Dataset and Community Challenges. ACS Catal. 2021, 11(10), 6059–6072. [Google Scholar] [CrossRef]
- Tran, R.; Lan, J.; Shuaibi, M.; Wood, B. M.; Goyal, S.; Das, A.; Heras-Domingo, J.; Kolluru, A.; Rizvi, A.; Shoghi, N.; Sriram, A.; Therrien, F.; Abed, J.; Voznyy, O.; Sargent, E. H.; Ulissi, Z.; Zitnick, C. L. The Open Catalyst 2022 (OC22) Dataset and Challenges for Oxide Electrocatalysts. ACS Catal. 2023, 13(5), 3066–3084. [Google Scholar] [CrossRef]
- Kirklin, S.; Saal, J. E.; Meredig, B.; Thompson, A.; Doak, J. W.; Aykol, M.; Rühl, S.; Wolverton, C. The Open Quantum Materials Database (OQMD): Assessing the Accuracy of DFT Formation Energies. npj Computational Materials 2015, 1(1), 1–15. [Google Scholar] [CrossRef]
- Radova, M.; Stark, W. G.; Allen, C. S.; Maurer, R. J.; Bartók, A. P. Fine-Tuning Foundation Models of Materials Interatomic Potentials with Frozen Transfer Learning. npj Computational Materials 2025, 11(1), 237. [Google Scholar] [CrossRef] [PubMed]
- Smith, J. S.; Nebgen, B. T.; Zubatyuk, R.; Lubbers, N.; Devereux, C.; Barros, K.; Tretiak, S.; Isayev, O.; Roitberg, A. E. Approaching Coupled Cluster Accuracy with a General-Purpose Neural Network Potential through Transfer Learning. Nature communications 2019, 10(1), 2903. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Zhou, L.; Hou, P.; Liu, Y.; Wang, R.; Guo, T.; Liu, J.-C. A Machine Learning Interatomic Potential Data Set and Model for Catalysis with Local Fine-Tuning to Chemical Accuracy. JACS Au 2025, 5(12), 6151–6161. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xiao, B.; Tang, Y.; Liu, F.; Wang, X.; Yan, F.; Liu, Y. Center-Environment Feature Model for Machine Learning Study of Spinel Oxides Based on First-Principles Computations. J. Phys. Chem. C 2020, 124(52), 28458–28468. [Google Scholar] [CrossRef]
- Tang, Y.; Xiao, B.; Chen, S.; Qian, Q.; Liu, Y. Predefined Attention-Focused Mechanism Using Center-Environment Features: A Machine Learning Study of Alloying Effects on the Stability of Nb 5 Si 3 Alloys. Digital Discovery 2025, 4(7), 1870–1883. [Google Scholar] [CrossRef]
- Bin, X.; Fu, L. I. U.; Wan, D. U.; Yi-heng, S.; Xue, F. A. N.; Quan, Q.; Yi, L. I. U. Machine Learning with Center-Environment Attention Mechanism for Multi-Component Nb Alloys. Transactions of Nonferrous Metals Society of China 2025, 35(11), 3813–3823. [Google Scholar]
- Ramakrishnan, R.; Dral, P. O.; Rupp, M.; Von Lilienfeld, O. A. Quantum Chemistry Structures and Properties of 134 Kilo Molecules. Scientific data 2014, 1(1), 1–7. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.; Ong, S. P.; Hautier, G.; Chen, W.; Richards, W. D.; Dacek, S.; Cholia, S.; Gunter, D.; Skinner, D.; Ceder, G. Commentary: The Materials Project: A Materials Genome Approach to Accelerating Materials Innovation. APL materials 2013, 1(1). [Google Scholar] [CrossRef]
- Saal, J. E.; Kirklin, S.; Aykol, M.; Meredig, B.; Wolverton, C. Materials Design and Discovery with High-Throughput Density Functional Theory: The Open Quantum Materials Database (OQMD). JOM 2013, 65(11), 1501–1509. [Google Scholar] [CrossRef]
- Tang, Y.; Xiao, B.; Chen, J.; Liu, F.; Du, W.; Guo, J.; Liu, Y.; Liu, Y. Multi-Component Alloying Effects on the Stability and Mechanical Properties of Nb and Nb–Si Alloys: A First-Principles Study. Metall Mater Trans A 2023, 54(2), 450–472. [Google Scholar] [CrossRef]
- Guo, J.; Xiao, B.; Li, Y.; Zhai, D.; Tang, Y.; Du, W.; Liu, Y. Machine Learning Aided First-Principles Studies of Structure Stability of Co3 (Al, X) Doped with Transition Metal Elements. Computational Materials Science 2021, 200, 110787. [Google Scholar] [CrossRef]
- Horton, M. K.; Huck, P.; Yang, R. X.; Munro, J. M.; Dwaraknath, S.; Ganose, A. M.; Kingsbury, R. S.; Wen, M.; Shen, J. X.; Mathis, T. S.; Kaplan, A. D.; Berket, K.; Riebesell, J.; George, J.; Rosen, A. S.; Spotte-Smith, E. W. C.; McDermott, M. J.; Cohen, O. A.; Dunn, A.; Kuner, M. C.; Rignanese, G.-M.; Petretto, G.; Waroquiers, D.; Griffin, S. M.; Neaton, J. B.; Chrzan, D. C.; Asta, M.; Hautier, G.; Cholia, S.; Ceder, G.; Ong, S. P.; Jain, A.; Persson, K. A. Accelerated Data-Driven Materials Science with the Materials Project. Nat. Mater. 2025, 24(10), 1522–1532. [Google Scholar] [CrossRef]
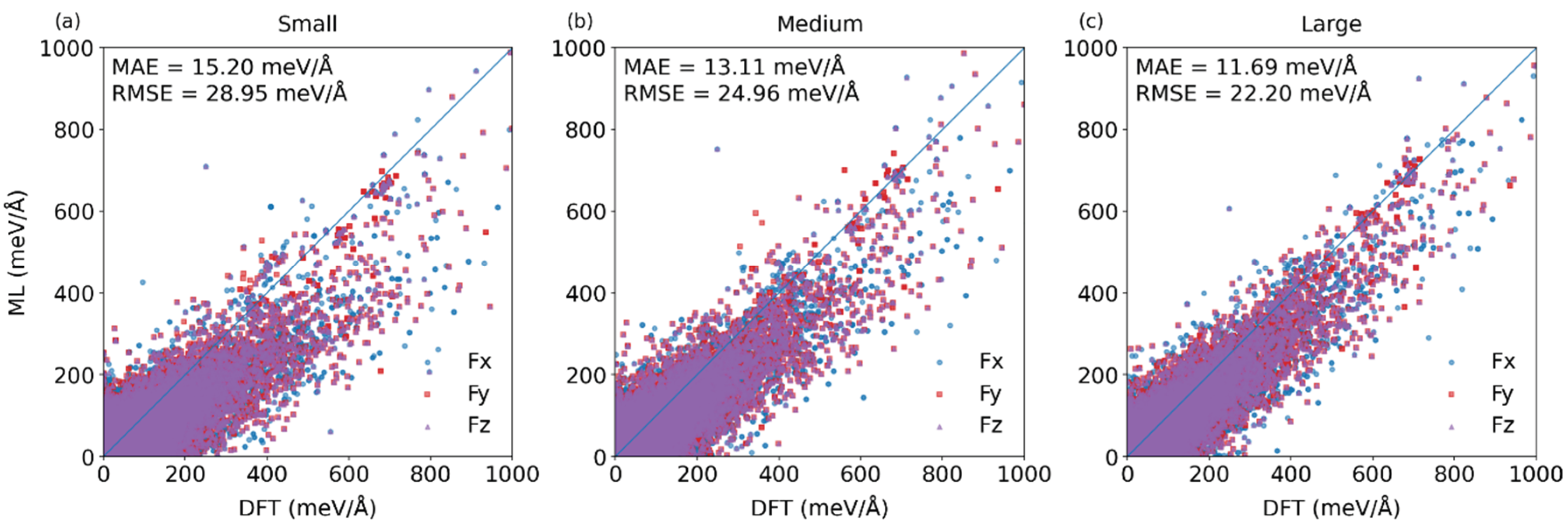

| ML models | Energy <MAE> (meV/atom) |
Energy <RMSE> (meV/atom) |
Force <MAE> (meV/Å) |
Force <RMSE> (meV/Å) |
| FT-Eq (Small) FT-Eq (Medium) FT-Eq (Large) |
17.07 ± 4.74 8.10 ± 1.40 7.34 ± 1.71 |
30.31 ± 11.40 12.45 ± 2.79 13.20 ± 4.02 |
11.00 ± 0.75 11.11 ± 0.72 12.98 ± 1.21 |
22.82 ± 1.69 22.25 ± 1.66 26.32 ± 2.64 |
| FT-NonEq (Small) FT-NonEq (Medium) FT-NonEq (Large) |
5.55 ± 0.14 3.51 ± 0.13 2.27 ± 0.07 |
7.47 ± 0.49 4.95 ± 0.80 3.79 ± 0.89 |
18.28 ± 0.27 15.69 ± 0.18 13.83 ± 0.18 |
31.58 ± 0.75 27.33 ± 0.62 24.26 ± 0.48 |
| Scratch-Eq (Small) Scratch-Eq (Medium) Scratch-Eq (Large) |
75.04 ± 22.02 111.96±11.91 121.53 ± 8.58 |
124.60±28.00 144.37±20.92 153.38±18.81 |
17.08 ± 3.04 11.26 ± 2.46 3.57 ± 1.08 |
31.17 ± 5.58 21.03 ± 4.57 5.91 ± 1.94 |
| Zero-shot-Eq (Small) Zero-shot-Eq(Medium) Zero-shot-Eq (Large) |
159.20 ± 6.90 184.43 ± 7.04 143.93 ± 4.39 |
176.53 ± 4.62 200.03 ± 4.99 151.50 ± 3.35 |
69.44 ± 5.50 50.26 ± 3.46 44.55 ± 3.33 |
146.13 ± 7.86 100.56 ± 4.79 88.57 ± 4.55 |
| Element | Strict-LODO | Relaxed-LODO | ||||||
|---|---|---|---|---|---|---|---|---|
| Energy MAE |
Energy RMSE |
Force MAE |
Force RMSE |
Energy MAE |
Energy RMSE |
Force MAE |
Force RMSE |
|
| Fe | 54.16 | 57.20 | 50.3 | 108.8 | 9.31 | 10.13 | 17.8 | 41.9 |
| Mn | 73.71 | 77.06 | 45.5 | 93.4 | 7.48 | 8.44 | 15.3 | 29.6 |
| Nb | 69.64 | 73.65 | 36.2 | 74.4 | 8.56 | 10.22 | 9.0 | 14.8 |
| Re | 132.05 | 138.02 | 40.9 | 69.3 | 6.46 | 9.02 | 12.1 | 23.8 |
| Ta | 112.42 | 116.76 | 28.0 | 60.8 | 8.89 | 11.14 | 11.0 | 19.1 |
| Ti | 61.56 | 64.50 | 31.4 | 69.6 | 7.73 | 11.24 | 10.2 | 19.3 |
| V | 90.18 | 95.19 | 34.6 | 71.8 | 7.45 | 8.76 | 11.8 | 23.6 |
| W | 117.84 | 123.69 | 49.4 | 118.9 | 8.01 | 9.09 | 9.9 | 18.8 |
| Y | 6.80 | 9.15 | 43.7 | 74.8 | 10.03 | 14.54 | 18.5 | 32.4 |
| Zr | 44.37 | 47.09 | 37.1 | 71.2 | 10.28 | 14.33 | 13.9 | 24.3 |
| Cr | 84.01 | 87.88 | 54.1 | 126.8 | 136.15 | 141.50 | 77.4 | 177.2 |
| MeanNC | 76.27 | 80.23 | 39.71 | 81.30 | 8.42 | 10.69 | 12.95 | 24.76 |
| MeanAll Std. | 76.98 ±36.05 | 80.93 ±37.18 | 41.0 ±8.3 | 85.4 ±22.8 | 20.03 ±38.53 | 22.58 ±39.49 | 18.8 ±19.7 | 38.6 ±46.6 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




