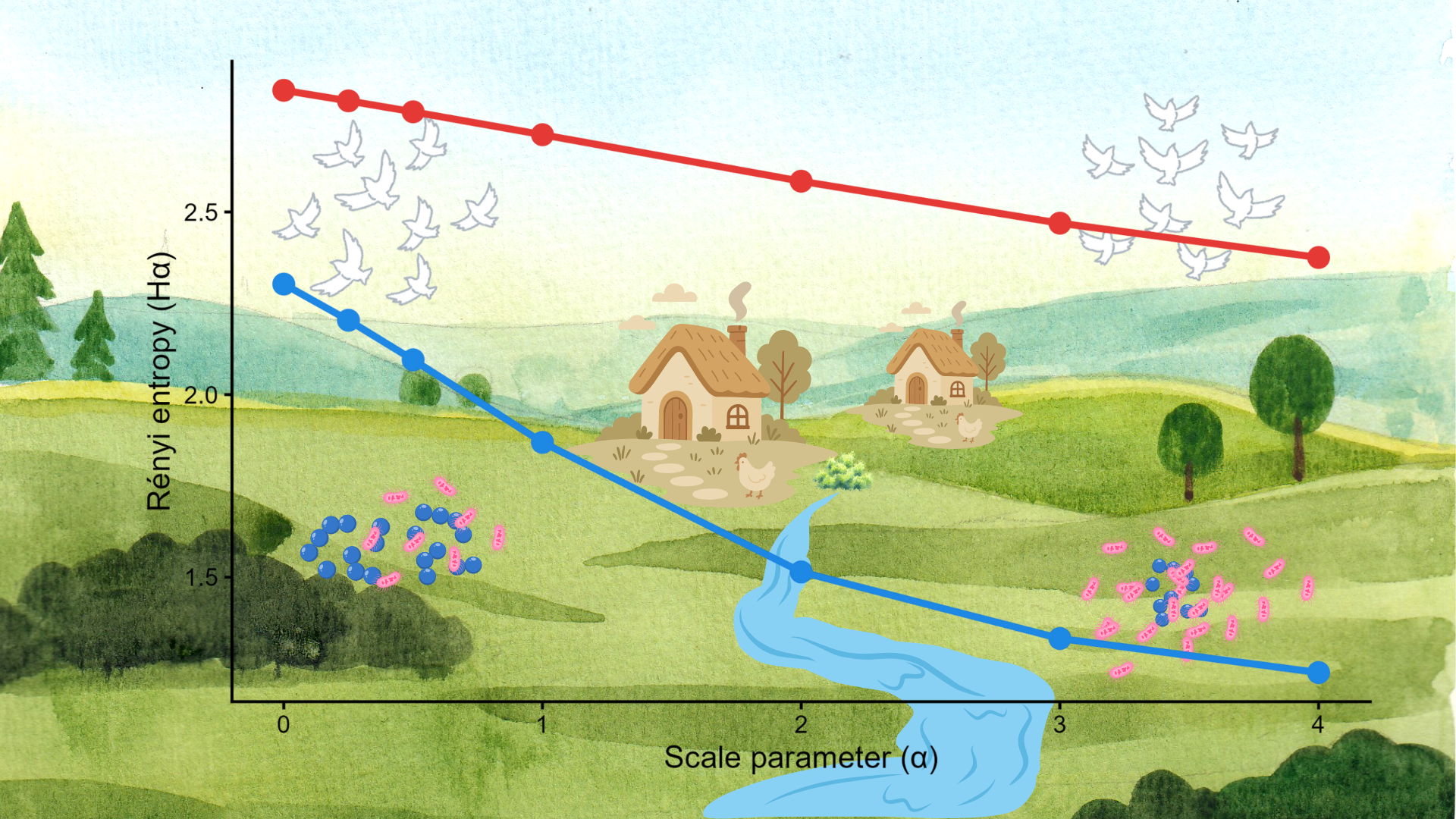
Antimicrobial resistance (AMR) is a silently escalating global crisis, presenting a specific challenge for the One Health approach. Landscapes can serve as reservoirs of AMR, while synurban wildlife may act as vectors of bidirectional exchange. However, these species can also be utilised as sentinels of landscape AMR load. Herbivorous avian bioindicators, such as the Common Wood Pigeon (Columba palumbus), continuously sample the landscape during foraging and drinking, providing unbiased data on the state of AMR. This study aimed to investigate the potential of this species for assessing the impact of landscape diversity on bacterial communities and their AMR patterns. Toward this objective, two landscape units of 4-km-diameter located at an upstream and a downstream section of a river, relative to a provincial town, were compared using 16 cloacal samples per site. Heterotrophic plate count techniques resulted in 60 isolates, of which 48 were identified, and 35 were tested for AMR using the VITEK 2 Compact system. Rényi diversity profiles of landscape compositions, bacterial communities, and AMR patterns revealed that higher landscape diversity was associated with lower bacterial but higher AMR pattern diversity. Additionally, the structure of more diverse bacterial communities shifted toward Gram-negative taxa. These findings support the hypothesis that culture-based methods using Common Wood Pigeons, complemented by Rényi diversity analysis and the determination of Gram-positive/Gram-negative ratios, provide valuable data on landscape health, even with small sample sizes.