Introduction
High-concentrate diets, rich in nutrients and energy, rapidly replenish animals’ energy and protein requirements, promoting growth and enhancing production performance.Therefore, it is primarily used for fattening. However, with the development of intensive farming, the proportion of high-concentrate diets in sheep rations has increased, readily resulting in subacute ruminal acidosis (SARA). This leads to impaired ruminal fermentation and gastrointestinal dysfunction [
1]. During SARA, volatile fatty acids (VFAs) and lactic acid accumulate in rumen contents, pH levels decrease significantly, and microbial diversity decreases, leading to metabolic disorders and weakening the absorptive barrier function of the rumen epithelium [
2]. Large amounts of lactic acid and endotoxins (primarily lipopolysaccharides [LPS] from gram-negative bacteria) migrate from the rumen to peripheral tissues, triggering systemic inflammatory responses [
3]. This subsequently leads to an imbalance in the host-microbiota equilibrium of the hindgut. Studies indicate that under SARA conditions, elevated levels of LPS and the pro-inflammatory cytokine tumor necrosis factor-α (TNF-α) are detected in sheep blood and tissues. Significant activation of TLR4 receptors and the NF-κB signaling pathway occurs in tissues, accompanied by abnormal expression of tight junction proteins, leading to an impaired mucosal barrier [
3]. Disruption of the intestinal barrier allows pathogenic bacteria and their toxins to readily penetrate the intestinal mucosa, triggering excessive mucosal immune activation and chronic inflammation, thereby exacerbating metabolic stress in the body [
4]. Therefore, under a high-concentrate diet feeding regimen, sheep intestines are prone to a vicious cycle of acidosis, barrier damage, and microbial community disruption, severely impacting animal health and production performance.
Current research predominantly focuses on the effects of high-concentrate diets on the rumen and small intestine, with limited investigation into the hindgut. The colon exhibits highly complex and diverse structural, functional, and internal environmental characteristics, posing significant challenges for disease diagnosis and treatment. Within the colon, approximately 17% of digestible fiber undergoes microbial fermentation, producing substantial amounts of volatile fatty acids (VFAs) such as acetate, propionate, and butyrate, which provide a vital supplementary energy source for the host [
5]. Moreover, an intact colonic mucosal barrier helps prevent the transmural transfer of harmful microbial metabolites (e.g., endotoxins), thereby safeguarding intestinal and systemic health [
6]. Over 70% of the body’s immune cells reside within the sheep colon and primarily functioning to maintain tolerance toward commensal microorganisms while preventing pathogen invasion. This critical equilibrium is disrupted during dietary acidosis. High-concentrate diets can damage the hindgut barrier in sheep, triggering mucosal inflammation. In a trial involving Hu sheep, high-concentrate diets reduced expression of the goblet cell mucoprotein MUC2 and tight junction proteins, impairing colonic barrier function. This was accompanied by marked inflammation and tissue thickening in the colonic mucosa [
7]. These findings demonstrate that under normal conditions, the colonic barrier provides a physiological advantage against the leakage of pro-inflammatory substances into the systemic circulation. Its impairment leads to inflammatory responses in the sheep colon and compromised health.
Yeast culture (YC), a microbial fermentation product, is widely used in ruminant diets to alleviate gastrointestinal disorders caused by dietary imbalances [
8]. Yeast culture is typically obtained through the complete fermentation of yeast on specific media, comprising extracellular metabolites of yeast cells, transformed fermentation medium, and a small amount of inactivated yeast cells [
9]. Research indicates that adding yeast culture stabilizes rumen fermentation, enhances nutrient digestibility, and modulates immune responses to some extent [
10]. Concurrently, YC indirectly reduces hindgut acid load and barrier damage risk by stabilizing ruminal fermentation and pH levels [
11].
Research on the effects of high-concentrate diets on sheep colonic microbiota and barrier function remains limited, and the mechanism of yeast culture intervention is unclear. Therefore, this study selected Duhan crossbred sheep as experimental subjects to investigate the potential impacts of high-concentrate diets on sheep and the intervention mechanism of yeast culture, aiming to achieve synergistic optimization of efficient feeding and animal health in sheep.
Materials and Methods
Feeding Trial and Sampling
This study employed a randomized design using individual animals as experimental units. Forty-five healthy, similar-condition 3-month-old Duhan crossbred rams were randomly assigned to 45 separate pens and fed according to their respective groups. The treatment groups were designed as follows: (1) Control group (CON); (2) High-concentrate diet group (HC); (3) Yeast Culture Treatment Group (Xi’an Xinhanbao Biotechnology Co., Ltd.) fed (HCY). The feed formulation is shown in
Table 1 (inactive yeast). Water was provided ad libitum. After 60 days of feeding, five healthy sheep were randomly selected from each group and transported to a local slaughterhouse for slaughter upon trial completion. Five milliliters of the colonic contents were collected into cryogenic tubes, rapidly frozen in liquid nitrogen, and stored at -80 °C for microbiome analysis and VFA determination. The colon walls were then gently rinsed with sterile saline to remove residual contents. Two tissue samples were obtained: one fixed was in 4% paraformaldehyde solution at 4 °C for histological sectioning; and the other was stored in an RNA preservative (RNAlater) at -80℃ for RNA extraction.
Hematoxylin and Eosin (H&E) Staining
Fixed colon samples were retrieved, subjected to routine dehydration, cleared, paraffin embedded, and sectioned into 4 μm-thick slices. After xylene dewaxing and gradient ethanol hydration, the sections were stained with hematoxylin for 5–8 minutes. Following differentiation with 1% hydrochloric acid ethanol and counterstaining with water, the sections were counterstained with 0.5% eosin solution for 1–3 minutes. The sections were then dehydrated with graded ethanol, cleared with xylene, and finally sealed with neutral resin. Histological structures were examined under a microscope to assess intestinal barrier integrity.
16. S rRNA Amplification and Sequencing
Total DNA was extracted from each sample using magnetic bead technology (100–200 mg fecal sample was mixed with lysis buffer, Proteinase K, and 1 mm grinding beads, homogenized by mechanical disruption, and lysed at 70 °C). After centrifuging the lysis buffer and collecting the supernatant, RNase A and binding buffer were sequentially added. Following centrifugation to remove impurities, the supernatant was transferred to a DNA binding column for binding. Subsequently, the sample was washed sequentially with wash buffers to remove residual impurities and salts. Finally, elution with TE buffer or nuclease-free water yielded fecal genomic DNA, with DNA concentration, purity, and integrity subsequently assessed. Barcoded primers were synthesized targeting the 341F (CCTACGGGNGGCWGCAG) and 806R (GGACTACHVGGGTWTCTAAT) regions for PCR amplification. Amplified products were purified and sequenced using paired-end (PE250) read lengths on an Illumina MiSeq platform. Raw sequencing reads underwent quality control using Trimmomatic to remove adapter sequences, low-quality bases, and N-containing reads. Subsequently, paired-end reads were assembled using FLASH or PEAR based on read overlap, with a minimum overlap length of 10 bp and a maximum mismatch rate of 0.1. Chimeric sequences were removed from Fasta files using the Uchime method against known databases. For unknown databases, chimeras were eliminated via self-alignment (Denovo) alongside short sequences failing quality thresholds. Clean Tags were then subjected to bioinformatics analysis using the Uparse clustering method to generate OTUs at 97% similarity. OTU-based species annotation (using Silva128 as reference database), diversity analysis, and differential analysis were subsequently performed. Functional prediction of microbial communities was performed using PICRUSt2 software based on 16S rRNA sequencing data. Significantly altered metabolic pathways were identified via LEfSe (LDA > 2.0, P < 0.05), providing insights into potential metabolic functional impacts of microbial community shifts. Due to limitations in functional prediction, results serve only as supplementary interpretation.
Transcriptome Sequencing to Observe the Molecular Effects of the Microbiota on the Gut
Total RNA was extracted from 15 colon samples via an appropriate volume of TRIzol reagent. The RNA integrity and concentration were assessed via a bioanalyzer (Agilent Bioanalyzer 2100 system). Following cDNA library construction, high-throughput transcriptome sequencing was performed via the MGI sequencing platform to obtain raw data. High-quality data (clean reads) were filtered and aligned to the sheep genome ARS-UI_Ramb_v3.0 via HISAT2 alignment software. Transcriptome assembly was then performed via stringtie. New transcripts were identified by comparing the assembly with existing annotations via cuffcompare. CPC was subsequently employed to predict the protein-coding potential of these new transcripts. TransDecoder was then used to predict coding regions for transcripts with protein-coding potential, yielding new genes with coding capacity. These new genes were then integrated into the reference gene set. Finally, quantitative analysis of the integrated gene set was performed via RSEM. Differential gene expression analysis was conducted with the DESeq2 package, setting thresholds at |log2 Fold Change| > 0.58 and FDR-adjusted P < 0.05. The DEGs were subjected to Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis to screen for genes associated with the immune response and barrier function pathways, and differences in gene expression were observed.
GC Targeted Detection of VFAs
Detection was performed via an Agilent 8890 GC gas chromatograph. A 5 mL filtrate was centrifuged at 4,000 r/min for 15 min. One milliliter of the supernatant was then mixed with 25% metaphosphoric acid solution at a 4:1 ratio. After mixing for 30 min in an ice–water bath, the mixture was centrifuged at 4,000 r/min for 15 min in a 4 °C centrifuge. The filtrate was transferred into dedicated vials. The operating conditions for the gas chromatograph were as follows: FFAP column, 30 m × 0.32 mm × 0.25 μm. The forward injection port with an injector heater set to 300 °C and a split ratio of 10:1 was used. The column temperature was as follows: initial temperature of 80 °C held for 1 min; ramped at 10 °C/min to 170 °C, held for 1 min; and continued at 20 °C/min to 250 °C, held for 5 min, with a maximum column oven temperature of 300 °C. Hydrogen flame ionization detection (FID) was employed with nitrogen as the carrier gas; the detector heater temperature was 300 °C, the injection volume was 1 μL, and the run time was 20 min. 2-Ethylbutyric acid served as the internal standard.
Determination of Ammonia Nitrogen
Take 10 mL of colonic contents and centrifuge at 3500–4000 rpm for 10 minutes. Measure 2 mL of supernatant (if ammonia nitrogen content is high, take 1 mL supernatant and add 1 mL distilled water), place in a 15 mL test tube, then add 8 mL of 0.2 mol/L hydrochloric acid to a final volume of 10 mL and mix thoroughly. Prepare standard solution, stock solution (dissolve 0.382 g ammonium chloride in 0.2 mol/L hydrochloric acid and dilute to 100 mL), and working solution (take 10 mL stock solution and dilute with distilled water to 100 mL) sequentially. Sequentially measure 0, 1, 2, 4, and 6 mL of the working solution into five labeled 50 mL volumetric flasks. Add distilled water to each to a final volume of 10 mL, then add Solution A (0.08 g Na2 [Fe(CN)5NO]·2H₂O dissolved in 100 mL of 14% NaC6H4(OH)CO2 solution) and Solution B (2 mL NaClO mixed into 100 mL of 0.3 M NaOH solution), then add 0.2 mol/L hydrochloric acid to the mark. Measure the absorbance using a 721 spectrophotometer. Measure at 700 nm using a 0.5 cm cuvette. Use nitrogen-free blank solution (No. 0 tube) as the blank control. Record the absorbance values for each sample. Calculate the ammonia nitrogen content using the standard curve equation..
qPCR Validation
To validate the reliability of RNA-Seq results, we selected key differentially expressed genes and performed qPCR validation using the same colon RNA samples (n=5 per group) as those used for transcriptome sequencing. Colonic RNA was extracted from each group, reverse transcribed into cDNA, and amplified using specific primers targeting the genes of interest. Expression levels were compared to the housekeeping gene (
GAPDH) and calculated as relative expression (using the 2^
-ΔΔCt method). qPCR results generally aligned with sequencing trends, confirming the accuracy and reliability of the transcriptomic analysis. Primer sequences are listed below, synthesized by Wuhan Savier Biotechnology Co., Ltd.:, are listed below
Table 2.
Western Blot Validation of Relevant Protein Expression Levels
All operations were performed on ice. Approximately 20 mg of colon tissue was lysed thoroughly with RIPA lysis buffer. The lysate was centrifuged at 4 °C and 12,000 r/min for 20 min. The supernatant was collected, and the protein concentration was standardized via the BCA assay. A 1:1 volume of loading buffer was added. The mixture was heated at 100 °C for 10 min, cooled, and stored at –80 °C. SDS‒PAGE electrophoresis (120 V) was performed with 20 µg of protein. The gel (200 mA, 60 min) was transferred after the marker reached the bottom of the separating gel. The samples were blocked with 5% nonfat milk powder at room temperature for 1 hour. The samples were incubated overnight at 4 °C with primary antibodies against CLDN1 (Hangzhou Hua’an Biotechnology Co., Ltd., HA721999, 1:2000) and GAPDH (Wuhan Sanying Biotechnology Co., Ltd., 60004-1-Ig, 1:10000) at 4 °C. After the membranes were washed with TBST, they were incubated with secondary antibodies for 1 hour. Following another wash, the samples were visualized with an enhanced chemiluminescence (ECL) reagent. The blot bands were detected via a chemiluminescent imaging system. The target protein molecular weights were determined relative to those of the prestained marker, and quantitative analysis was performed via ImageJ (1.53e) software.
Data Statistics and Analysis
ANOVA was performed on mucosal phenotypic characteristics using SPSS Statistics 27 software. Multiple comparisons were followed by LSD post hoc tests, with results expressed as mean±SEM. Spearman correlation analysis was performed between differential microbes, differential genes, and VFAs. Results only indicate correlations without assessing significance, with FDR controlled using Benjamini–Hochberg correction. VFAs underwent one-way ANOVA using GraphPad Prism 10.1.2, with p < 0.05 as the significance threshold.
Discussion
Dietary composition differences significantly influence the sheep gut microbiota. Across dietary treatments, the gut microbial community structure, VFA profiles, and host gene expression patterns presented overall similarity, yet specific bacterial abundances, VFA levels, and related gene expression levels presented variations. This study investigated how different diets affect the colonic microbiota, VFAs, and host genes and the interrelationships among these three components.
We observed that the high-concentrate diet disrupted the integrity of the colonic mucosal barrier, whereas the YC intervention had protective and regulatory effects on the barrier. Consistent with previous studies [
12], the high-concentrate diet allowed undigested starch and rapidly fermentable substrates to enter the hindgut, causing a decrease in pH, transient VFA accumulation, and osmotic pressure changes. This leads to disrupted epithelial tight junctions, goblet cell depletion, and increased local inflammation, ultimately manifesting as histological mucosal epithelial damage [
13].
This study characterized the microbiota via 16S rRNA sequencing combined with functional prediction, revealing substantial microbial shifts. The
[Eubacterium]-xylanophilum group, Lachnospiraceae and
Ruminococcaceae have been repeatedly identified as important butyrate producers, positively correlated with barrier homeostasis [
14]. Our functional predictions yielded similar results: the CON group was enriched in bile acid metabolism and butyrate metabolism, whereas the HCY group was enriched in DNA repair and protein secretion-related functions. Following injury, the intestinal epithelium requires the DNA damage response, repair, and re-epithelialization to restore barrier integrity. Furthermore, bacterial secreted proteins and exosomes interact with host receptors to regulate tight junctions and immune signaling, forming a close link with HCY group enrichment in protein secretion-related functions [
15].
Microbial alterations indicate corresponding metabolic shifts, prompting us to analyze their metabolites: VFAs and ammonia nitrogen. VFAs serve as the primary energy source for ruminants, whose composition is largely influenced by feed composition [
16]. Changes in ammonia nitrogen levels reflect variations in feed protein degradation efficiency and microbial activity [
17]. In this study, the HC group presented the lowest VFA concentration in the colonic contents and the highest ammonia nitrogen (NH₃-N) content. This finding indicates that during hindgut fermentation, the intake of a high-concentrate diet promotes the proliferation of lactic acid bacteria, creating an acidic environment that inhibits microorganisms from utilizing lactic acid to produce butyric acid and propionic acid [
8]. Additionally, protein fermentation generates ammonia and amines [
18]. The HCY group presented significantly elevated acetate, propionate, butyrate, and valerate (
P < 0.05), which is consistent with the metabolic regulatory function of YC [
8]. Studies indicate that yeast culture enhances microbial nitrogen assimilation, increasing VFAs while reducing ruminal ammonia [
19]. Overall, feeding a high-concentrate diet alone may reduce colonic fermentation efficiency, whereas adding yeast culture can rebalance microbial activity to increase VFAs and suppress excessive ammonia production, a trend broadly consistent with prior research [
19,
20]. Severe or prolonged high-concentrate feeding disrupts fiber fermentation, leading to reduced VFAs and increased proteolytic end products [
21].
Our analysis of the DuHan F1 colon transcriptome revealed substantial changes in DEGs In the HCY group, indicating extensive transcriptional reprogramming. This finding aligns with previous reports in ruminant high-concentrate diet models [
13]. When performing WB and qPCR for the barrier protein CLDN1, we observed inconsistent expression patterns between transcript and protein abundance. We hypothesize that this discrepancy may stem from endoplasmic reticulum stress and the inflammatory environment induced by the HC diet, which accelerates the turnover and degradation of tight junction proteins via the endocytosis-lysosomal pathway [
22,
23]. Concurrently, the mucosal epithelium may transcritionally
upregulate CLDN1 as a compensatory response, leading to a discordant pattern of increased mRNA but decreased protein levels. This process may be further promoted by E3 ubiquitin ligases (e.g., LNX1), which mediate endocytosis and subsequent lysosomal degradation of claudins—a mechanism validated across diverse epithelial cell types [
23,
24,
25]. Moreover, under inflammatory conditions or upon stimulation by acidic loads or toxic metabolites, the UPR/ISR (PERK–eIF2α) pathway is readily activated, leading to a global reduction in protein translation. This results in increased transcription but restricted protein synthesis, with translation being suppressed [
26,
27]. In the HC group,
IL-8,
IL-23A, and
Ovar-DRB1 were downregulated, whereas
ADIPOQ was upregulated. A literature review indicated that decreased
IL-8 aligns with subsequent YC-mediated suppression, which is consistent with probiotic yeast classic NF-κB inhibition [
28].
Given sample size constraints and multiple corrections, correlations are presented as exploratory findings. Overall, in the CON group, VFAs were negatively correlated with proinflammatory factors (e.g
., CXCL8), which is consistent with the anti-inflammatory regulation by SCFAs, particularly butyrate [
29,
30]. Concurrently, tissue damage and reduced immune cell counts induced by HC also decrease the overall expression of these immune genes [
31]. Tao et al. similarly reported that a high-concentrate diet significantly downregulated the expression of proinflammatory factors such as
IL-1β [
32]. VFAs can also limit
IL-23 production by dendritic cells to regulate immunity [
33]; conversely, 65% grain intake downregulates tight junction proteins, causing epithelial damage accompanied by local inflammation [
34]. Under high-concentrate diet conditions
, these patterns exhibited diet-dependent rearrangements with partial directional reversals (e.g., barrier-related genes and ammonia nitrogen showed parallel trends), suggesting potential limitations in VFA absorption or utilization during inflammation. YC supplementation stabilized the overall pattern: the negative coupling between VFAs and inflammation weakened, whereas the positive coupling between barrier-related indicators and key metabolites strengthened.
The microbiota strongly influences the diet and is closely associated with VFAs [
35,
36]. Correlations are presented as exploratory patterns. The CON group maintains a healthy fermentation network in which dominant symbionts efficiently convert fibers into short-chain fatty acids [
14,
37]. However, the consumption of a high-concentrate diet disrupts this equilibrium. Microorganisms associated with fiber fermentation show weakened or absent links to VFAs, whereas novel abnormal associations emerge. This finding aligns with the finding that HC inhibits normal fiber fermentation pathways and reduces fiber-degrading bacteria in the hindgut [
38]. Certain rumenobacteria within the family Ruminococcaceae shift their metabolic function toward producing or tolerating valeric acid under high-concentrate diet conditions. When the abundance of fiber-degrading bacteria decreases, ammonia-producing proteolytic bacteria may proliferate substantially. The addition of YC partially restored the stability of microbial‒host metabolic interactions. The abundance of the
[Eubacterium]-xylanophilum-group was strongly negatively correlated with propionic acid (ρ = -1.00) and butyric acid (ρ = -0.90) contents. This likely reflects functional redistribution and suppression of specific acid-producing pathways following improved colonic digestion and controlled nutrient overflow [
39].
Under normal conditions, host mucosal gene expression exhibits is well-coordinated regulation with the microbial community. However, diet-dependent rearrangements emerged following high-concentrate diet feeding. In the HC group, the abundance
of Ruminococcaceae;
uncultured bacteria was negatively correlated with
CLDN1, ADIPOQ, and TNFSF13, whereas these associations were either absent or extremely weak in the CON group. This pattern aligns with reported HC-associated barrier stress/inflammatory stress [
40] and is consistent with transcriptional compensation and metabolic environment alterations under stress. In the HCY group,
[Eubacterium]-xylanophilum-group abundance was positively correlated with
CLDN1 expression (ρ=0.90), but exhibited enhanced inverse coupling with inflammatory factors (e.
g., IL23A). This aligns with the anti-inflammatory and barrier-supporting effects of the SCFA–GPR41 and GPR43 axes documented in the literature [
14].
We propose that YC primarily functions as a microbial regulator, fostering a community structure more conducive to glycolytic fermentation. This is evidenced by YC-induced inhibition of certain bacteria and promotion of a metabolic environment characterized by increased VFA production and reduced ammonia nitrogen content. The re-regulated microbial-metabolite interactions subsequently create a favorable environment for the host mucosa. This hypothesis is supported by our observed correlation analysis: the rebound of VFAs following YC supplementation correlates with the normalization of immune gene expression (e.g., downregulation of CXCL8 and ADIPOQ) and the restoration of CLDN1 protein abundance. However, this study has certain limitations. Particularly in metabolite analysis, it focused solely on changes in VFAs (acetic, propionic, butyric, and valeric acids), failing to comprehensively assess whether other metabolites were affected by the high-concentrate diet and YC intervention. It lacked broader metabolomics and causal validation, and the small sample size precludes causal discussions. Therefore, future studies should employ larger sample sizes under different dietary interventions and utilize metabolomics or metagenomics to further investigate the interactions among colonic metabolites, microbial communities, and host health in sheep.
Figure 1.
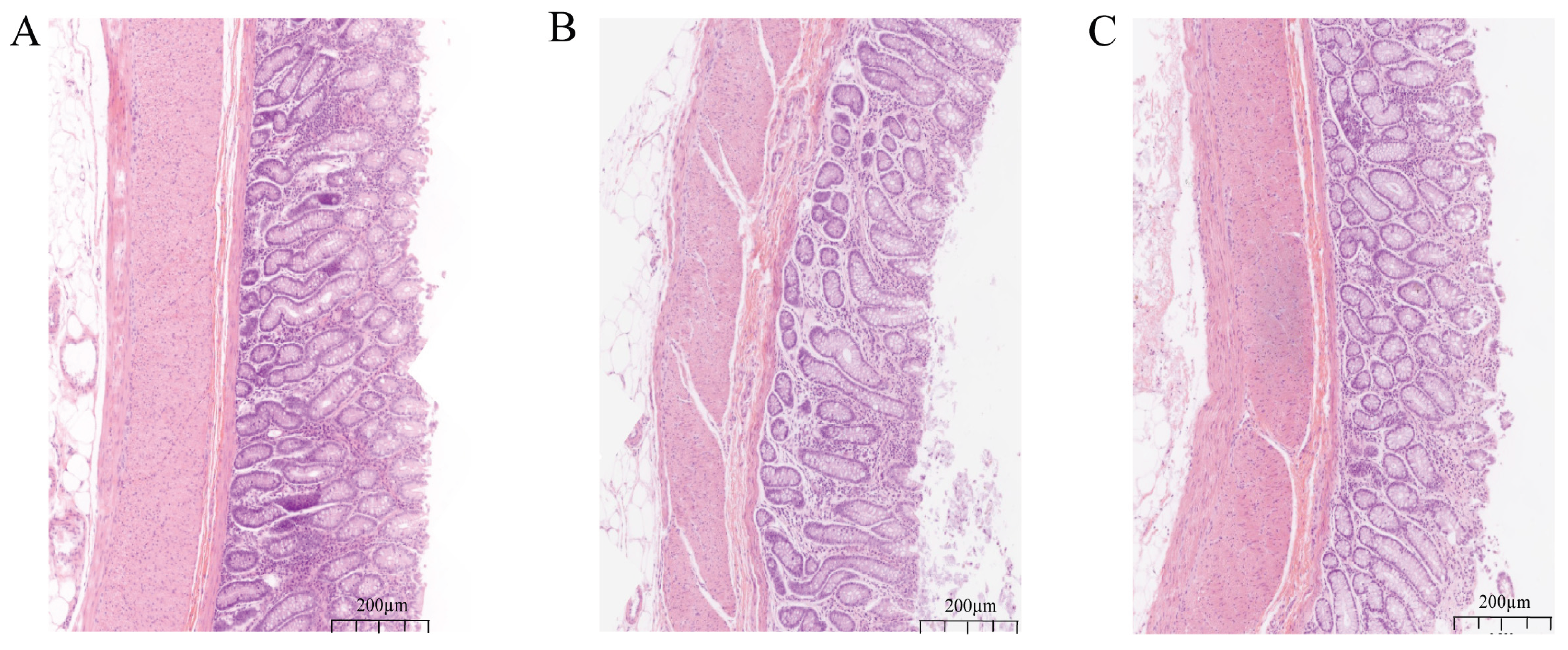
Effects of different diets on colonic tissue morphology in sheep. (A) CON: basal diet control. (B) HC: high-concentrate feeding. (C) HCY: high-concentrate diet + yeast culture feeding.
Figure 1.
Effects of different diets on colonic tissue morphology in sheep. (A) CON: basal diet control. (B) HC: high-concentrate feeding. (C) HCY: high-concentrate diet + yeast culture feeding.
Figure 2.
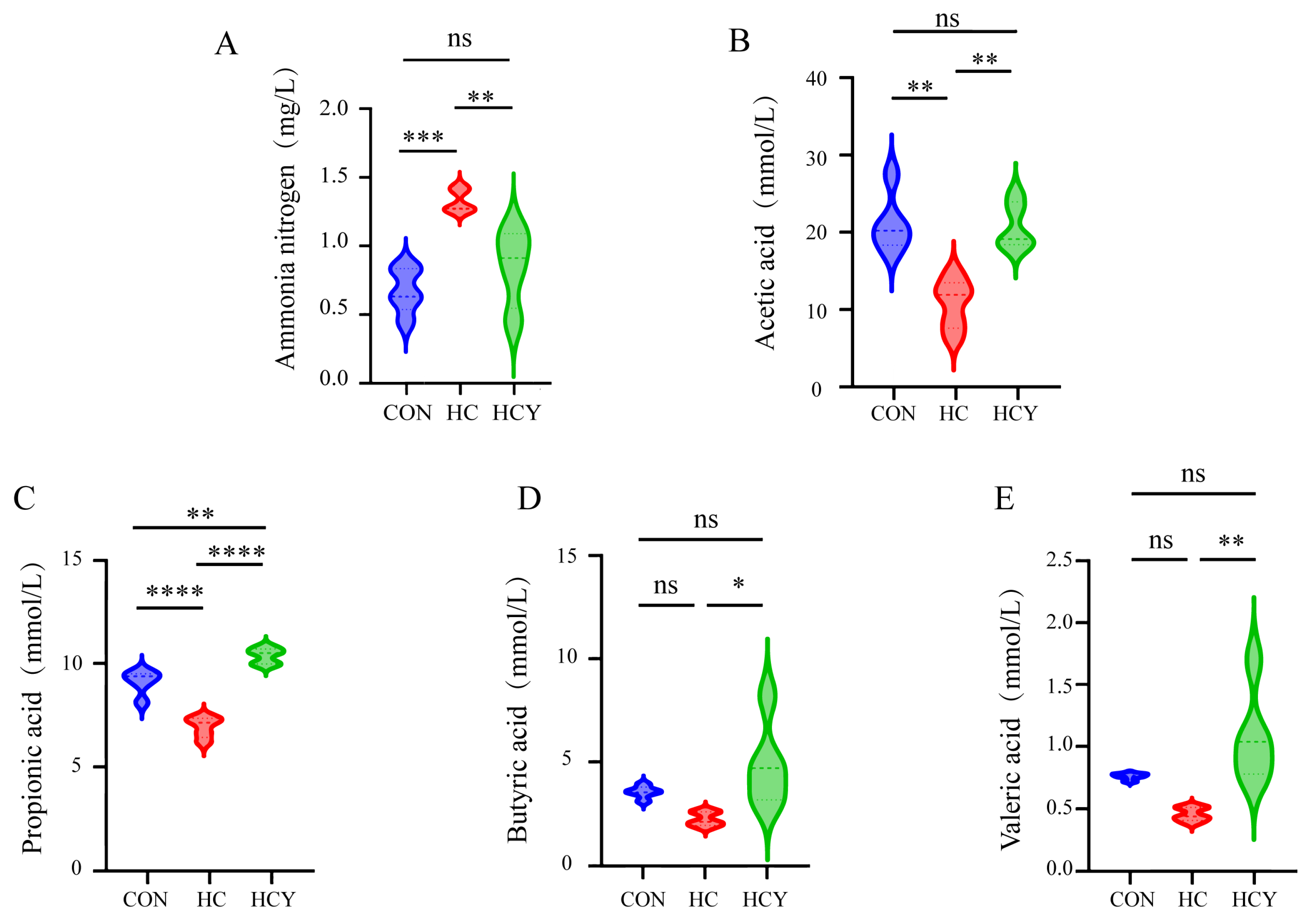
VFA content under different feeding conditions. (A) Ammonia nitrogen content in the colonic contents. (B) Acetic acid content in the colonic contents. (C) Propionic acid content in the colonic contents. (D) Butyric acid content in the colonic contents. (E) Valeric acid content in the colonic contents. Among these, no significant difference was observed (ns, P > 0.05); a significant difference was present (*, P < 0.05); and an extremely significant difference was noted (**, P < 0.01).
Figure 2.
VFA content under different feeding conditions. (A) Ammonia nitrogen content in the colonic contents. (B) Acetic acid content in the colonic contents. (C) Propionic acid content in the colonic contents. (D) Butyric acid content in the colonic contents. (E) Valeric acid content in the colonic contents. Among these, no significant difference was observed (ns, P > 0.05); a significant difference was present (*, P < 0.05); and an extremely significant difference was noted (**, P < 0.01).
Figure 3.
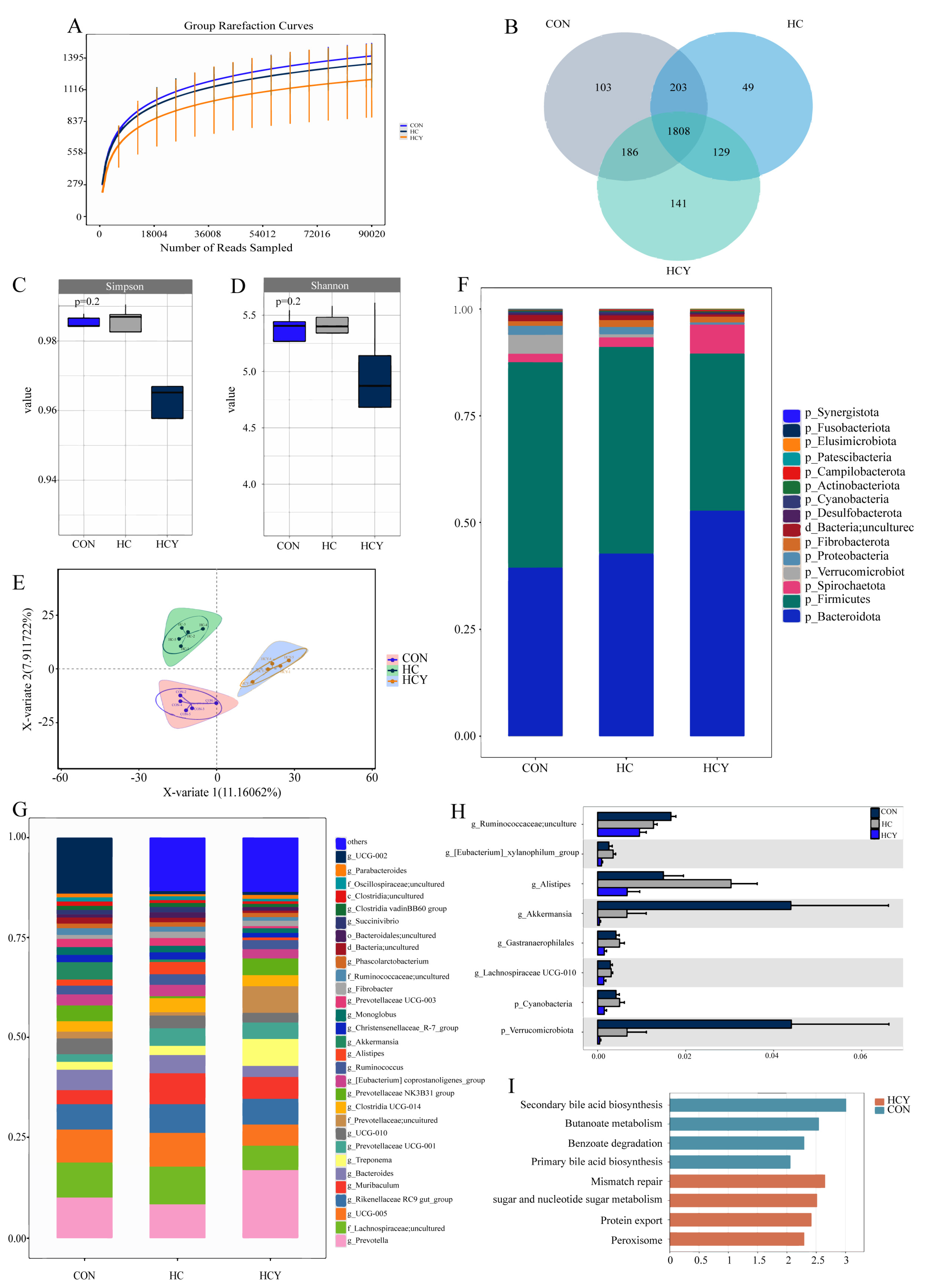
Effects of different feeding conditions on the microbial community of sheep. (A) Rarefaction curves showing the cumulative number of operational taxonomic units (OTUs) in each treatment group at different sequencing depths. (B) Venn diagram of OTU sharing and specificity among the three groups. (C) Box plot of the Simpson index. (D) Box plot of the Shannon index. (E) Two-dimensional scatter plot of partial least squares discriminant analysis (PLS-DA). (F) Stacked bar chart of the relative abundance of the top 15 dominant taxa at the phylum level. (G) Stacked bar chart of the relative abundance of the top 30 dominant taxa at the genus level. (H) Kruskal‒Wallis method for differential species analysis at the phylum and genus levels. (I) LEfSe microbial KEGG functional prediction.
Figure 3.
Effects of different feeding conditions on the microbial community of sheep. (A) Rarefaction curves showing the cumulative number of operational taxonomic units (OTUs) in each treatment group at different sequencing depths. (B) Venn diagram of OTU sharing and specificity among the three groups. (C) Box plot of the Simpson index. (D) Box plot of the Shannon index. (E) Two-dimensional scatter plot of partial least squares discriminant analysis (PLS-DA). (F) Stacked bar chart of the relative abundance of the top 15 dominant taxa at the phylum level. (G) Stacked bar chart of the relative abundance of the top 30 dominant taxa at the genus level. (H) Kruskal‒Wallis method for differential species analysis at the phylum and genus levels. (I) LEfSe microbial KEGG functional prediction.
Figure 4.
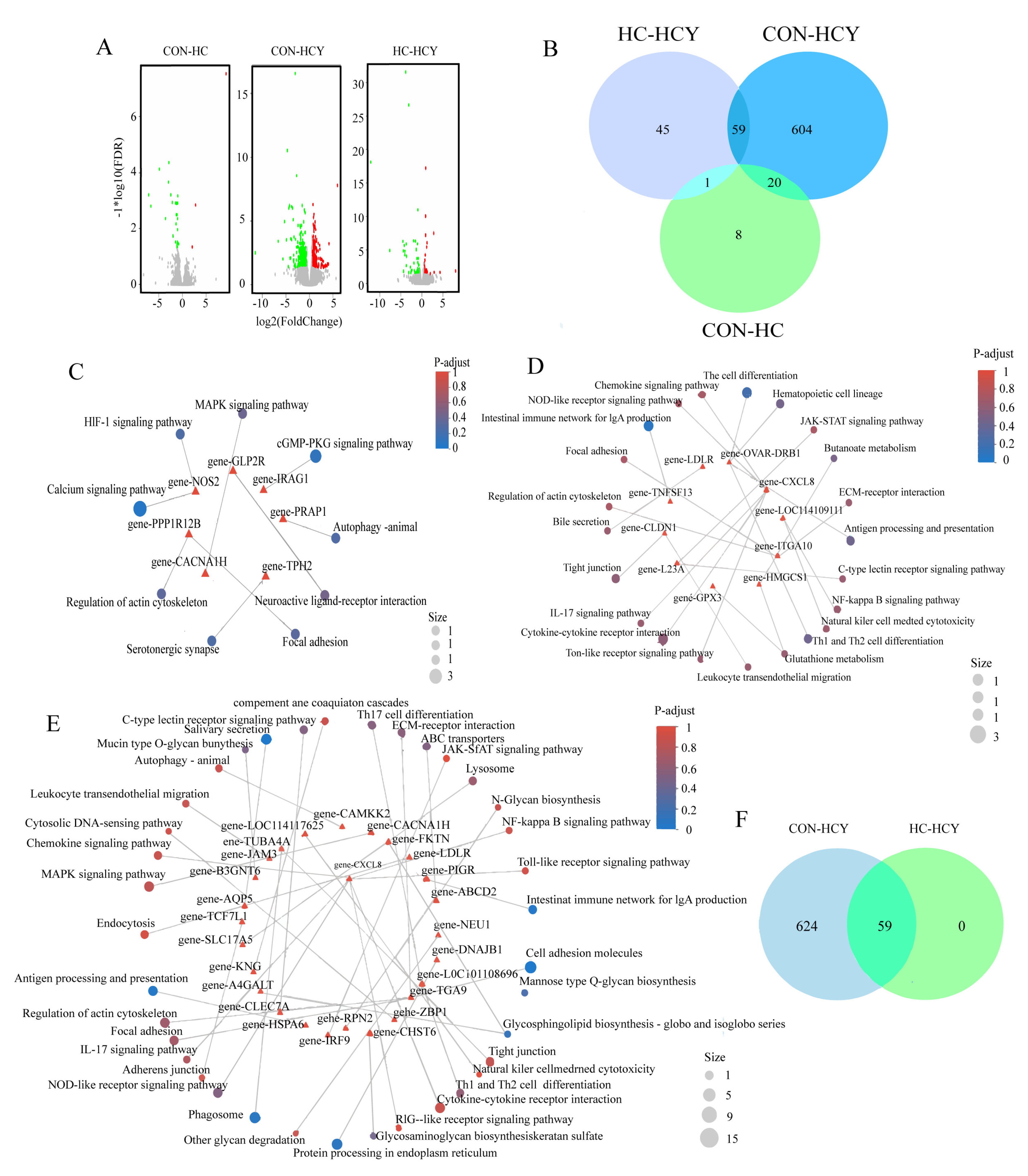
The influence of different feeding conditions on the transcriptome of sheep. (A) Volcano map of genes that were differentially expressed between groups. (B) Venn diagrams of differentially expressed genes in different groups. (C) CON-HC KEGG functional enrichment circle map of differentially expressed genes related to immune barriers. (D) HC-HCY KEGG functional enrichment circle map of differentially expressed genes related to immune barriers. (E) CON-HCY KEGG functional enrichment circle map of differentially expressed genes related to immune barriers. (F) Venn diagram of immune barrier-related DEGs between CON-HCY and HC-HCY.
Figure 4.
The influence of different feeding conditions on the transcriptome of sheep. (A) Volcano map of genes that were differentially expressed between groups. (B) Venn diagrams of differentially expressed genes in different groups. (C) CON-HC KEGG functional enrichment circle map of differentially expressed genes related to immune barriers. (D) HC-HCY KEGG functional enrichment circle map of differentially expressed genes related to immune barriers. (E) CON-HCY KEGG functional enrichment circle map of differentially expressed genes related to immune barriers. (F) Venn diagram of immune barrier-related DEGs between CON-HCY and HC-HCY.
Figure 5.
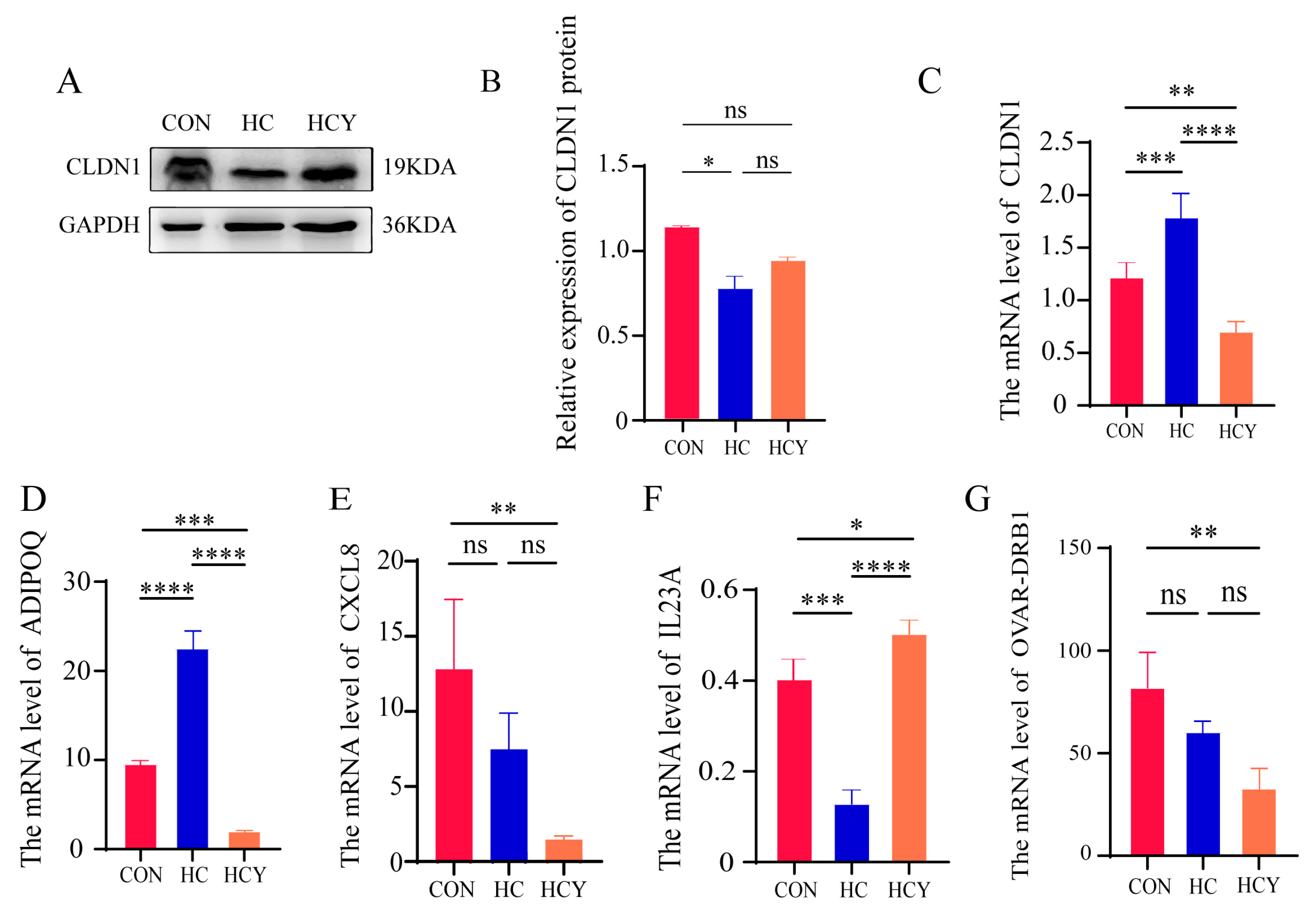
Effects of different diets on immune barrier genes in the colon tissue of sheep. (A) CLDN1 protein abundance levels in sheep colon tissue, including representative bands from Western blotting analysis and quantified volumes of specific bands. Protein expression levels were normalized against the corresponding GAPDH protein level. (B) CLDN1 protein expression levels in sheep colon tissue. (C), (D), (E), (F), and (G) Relative mRNA expression levels of CLDN1, ADIPOQ, CXCL8, IL23A, and Ovar-DRB1 in sheep colon tissue, respectively. GAPDH served as the reference gene for normalization. The difference was not significant (ns, P > 0.05); the difference was significant (*, P < 0.05); the difference was extremely significant (**, P < 0.01).
Figure 5.
Effects of different diets on immune barrier genes in the colon tissue of sheep. (A) CLDN1 protein abundance levels in sheep colon tissue, including representative bands from Western blotting analysis and quantified volumes of specific bands. Protein expression levels were normalized against the corresponding GAPDH protein level. (B) CLDN1 protein expression levels in sheep colon tissue. (C), (D), (E), (F), and (G) Relative mRNA expression levels of CLDN1, ADIPOQ, CXCL8, IL23A, and Ovar-DRB1 in sheep colon tissue, respectively. GAPDH served as the reference gene for normalization. The difference was not significant (ns, P > 0.05); the difference was significant (*, P < 0.05); the difference was extremely significant (**, P < 0.01).
Figure 6.
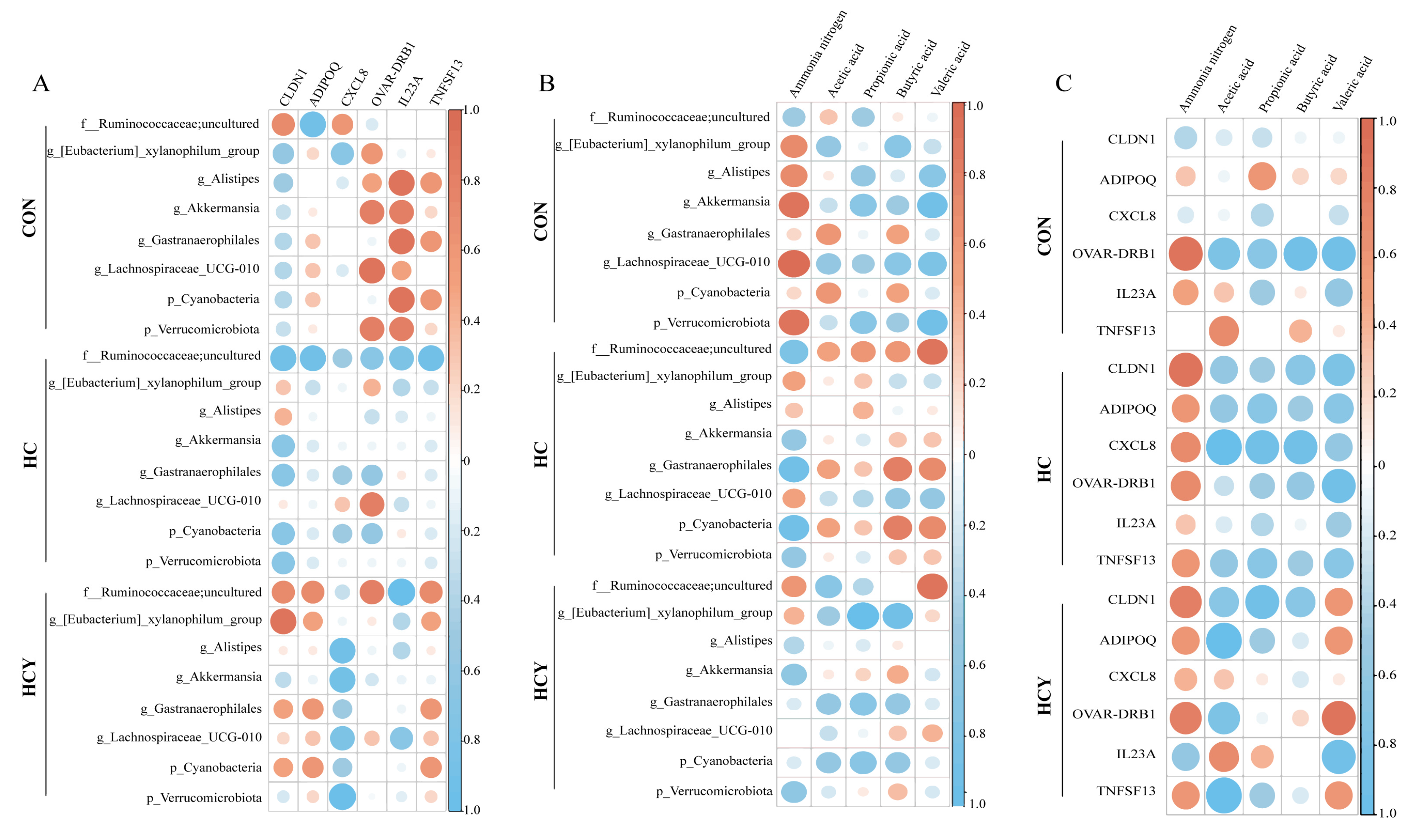
Heatmaps displaying Spearman’s ρ (effect sizes) only. Following FDR control (BH, 10%), no pairwise tests met the significance thresholds; thus, no significant markers are shown. (A) Spearman correlation analysis heatmap of differentially expressed genes related to immune barriers and differentially abundant bacteria. (B) Spearman correlation analysis heatmap among bacteria with differences in VFAs. (C) Spearman correlation analysis heatmap between VFAs and DEGs.
Figure 6.
Heatmaps displaying Spearman’s ρ (effect sizes) only. Following FDR control (BH, 10%), no pairwise tests met the significance thresholds; thus, no significant markers are shown. (A) Spearman correlation analysis heatmap of differentially expressed genes related to immune barriers and differentially abundant bacteria. (B) Spearman correlation analysis heatmap among bacteria with differences in VFAs. (C) Spearman correlation analysis heatmap between VFAs and DEGs.
Table 1.
Compositions and nutrient levels of the diets (DM basis).
Table 1.
Compositions and nutrient levels of the diets (DM basis).
| Items (%) |
Groups |
| CON |
HC |
HCY |
| Corn |
20.96 |
36.29 |
36.29 |
| Soybean hull |
18.7 |
15.8 |
15.5 |
| Spray-coated corn husk |
15.5 |
15 |
15 |
| Corn germ meal |
13.5 |
7.9 |
7.9 |
| Rice hull powder |
7.6 |
5 |
5 |
| Flour |
7 |
10.5 |
10.5 |
| Bran |
4.2 |
|
|
| Stone powder |
2.64 |
2.51 |
2.51 |
| Soybean meal |
|
2.5 |
2.5 |
| Rice bran |
2.3 |
|
|
| Pumpkin seed hull |
2.1 |
|
|
| Sugarcane molasses |
2 |
2 |
2 |
| Bentonite |
1 |
|
|
| Expanded urea1
|
1 |
1 |
1 |
| Sodium chloride |
0.5 |
0.5 |
0.5 |
| Yeast culture |
|
|
0.3 |
| 1% premix1
|
1. |
1 |
1 |
|
Chemical composition, % of DM
|
|
|
|
| DM |
89.58 |
88.72 |
88.72 |
| CP |
14.90 |
14.91 |
14.92 |
| EE |
3.25 |
2.52 |
2.52 |
| Starch-Ew |
25.99 |
34.96 |
34.98 |
| NDF |
31.83 |
25.07 |
25.07 |
| ADF |
11.77 |
12.58 |
12.58 |
| Ca |
1.20 |
1.10 |
1.10 |
| P-total |
0.40 |
0.35 |
0.35 |
Table 2.
List of primer sequences for quantitative real-time PCR.
Table 2.
List of primer sequences for quantitative real-time PCR.
| Gene |
GeneBank accession no. |
Direction |
Primer sequence (5′–3′) |
Length (bp) |
| CXCL8 |
NM_001009401.2 |
Forward |
AACACATTCCACACCTTTCCACC |
173 |
| Reverse |
CTCTCTTCAAAAATGCCTGCACA |
| IL23A |
NM_001185122.2 |
Forward |
GGGAACAGAGTTGTGATGCT |
164 |
| Reverse |
TCCACATGTCCCATTGGTAGG |
| Ovar-DRB1 |
NM_001280698.2 |
Forward |
CTGTATTTCTCCAGAGGCTCCC |
124 |
| Reverse |
GACACTCGCTCTTAGCATACTCC |
| CLDN1 |
NM_001185016.1 |
Forward |
AAGACGACGAGGCACAGAAGA |
168 |
| Reverse |
GAGCCTGACCAAATTCATACCTG |
| ADIPOQ |
NM_001308565.1 |
Forward |
TTCCACACCTGAGGGACTC |
136 |
| Reverse |
AATGCCTGCCATCCAACCTG |
| GAPDH |
NM_001190390.1 |
Forward |
GTCAAGGCAGAGAACGGGAA |
106 |
| Reverse |
CGTACTCAGCACCAGCATCA |
Table 3.
Effects of Different Diets on Sheep Colon Mucosa.
Table 3.
Effects of Different Diets on Sheep Colon Mucosa.
| Indicators |
CON |
HC |
HCY |
P |
| Crypt depth (mm) |
381.0896±7.1817A |
218.4259±8.3947BC |
322.4586±9.2712AB |
<0.01 |
| Crypt width (mm) |
40.626±1.9262 |
36.7474±1.5182 |
36.8027±1.0481 |
0.144 |
| Muscle layer thickness (mm) |
313.4273±8.2009A |
196.2983±2.9708AB |
175.1622±3.6256B |
<0.01 |
| Submucosal thickness (mm) |
75.5931±1.8456A |
38.9403±1.0286BC |
46.7882±2.7101AB |
<0.01 |
| Serosal thickness (mm) |
55.838±1.9128 |
52.4145±1.8280 |
52.8953±1.4258 |
0.334 |