Introduction
Living organisms persist because they are maintained “far-from-equilibrium” through metabolism, or the conversion of materials from the environment into new cells, tissues, biomass, etc. or into energy to form and maintain such organized material (England 2013). Natural selection is often hypothesized to favor individuals and their associated genotypes that can more efficiently or rapidly convert materials into growth or reproduction, (Odum 1993, Jorgensen et al. 1995, Brown et al. 2004, Jorgensen and Svirezhev 2007). However, selection might instead favor organisms that can persist longer or maintain stronger gradients of potential energy over longer times at the expense of maximizing rates. These may map onto trait trade-offs such as life history or resource economic strategies (Hirshfield and Tinkle 1975, Charnov 1997, Fridley 2017), where organisms may face trade-offs between maximizing rates and minimizing degradation (minimizing increases in entropy) in order to increase lifespan. Ultimately, the Second Law of Thermodynamics requires all organisms to increase the entropy of the surrounding environment, and selection may favor metabolic activity that best increases the rate at which entropy increases in surroundings in order to avoid degrading metabolic machinery, a concept known as maximum entropy production.
Another alternative is that metabolism reflects the outcomes of hundreds to thousands of reactions, and that the pattern of overall gas exchange rate with temperature may reflect the frequency distribution of reactions that can occur in the greatest number of possible ways, given limits on the supply of substrates and the rates at which molecules can move in and out of reaction sites (Harte 2011). Formally, this outcome can be determined by maximizing the information entropy of different reactions that produce a product (such as CO2) (Niven 2009) for a given substrate supply (such as various sugars and O2). In this case, metabolic rate may not track the temperature response of any single reaction, but rather emerge from the sum of different reactions and their relative contribution to the conversion of substrates to products. An analysis by Robert Niven (Niven 2009) suggests that this outcome will also produce maximum entropy production if reactions are near steady state.
These alternative “objectives” for the function of metabolic systems can be formulated into different hypotheses about what limits or drives metabolic and other vital rates and consequent associated times and states. Here, I will review and explore a framework for metabolic rate and its temperature dependence. I employ a thermodynamic framework that can make predictions from “objectives” grounded in both the First Law of Thermodynamics (maximum rates or power) and the Second Law (maximum entropy production). Many biologists consider “the matter of temperature” to be solved (West 2011) by the established paradigm that declining physiological rates above a Topt are driven by thermally induced degradation of enzymes (Sharpe and DeMichele 1977, Schoolfield et al. 1981, DeLong et al. 2017). However, evidence is lacking for physical degradation of enzymes at temperatures above that where metabolic rates begin to decline, Topt (20-40 oC) as most enzymes do not lose their catalytic capability until T> 55 oC (Daniel et al. 1996, Daniel and Danson 2010). Moreover, physiologists and ecologists frequently are faced with evidence that metabolic-temperature relationships are influenced by environmental resources, metabolic function (growth and reproduction versus locomotion and respiration), thermal expectations (acclimation), and body size (DeLong et al. 2018). These influences are not predicted by the current “denaturation” paradigm, so clearly the physical and chemical foundations of temperature effects on metabolism need further exploration.
Thermodynamics explicitly considers different forms of energy, including heat and chemical potential energy, and their contribution to metabolism and order, as measured by chemical reaction rates and changes in entropy in organisms and in surroundings. Thermodynamics necessarily requires that a “system” be defined so as to clearly identify the surroundings “outside” the system, and conditions “inside” the system. We begin by considering the most fundamental of metabolic processes – biochemical reactions – and the different factors that influence the course and rate of such reactions. Metabolism of course depends on thousands of different reactions that collectively occur at many different reaction sites within cells. However, to understand metabolism as a thermodynamic process, we first focus on a single hypothetical reversible, enzyme-catalyzed reaction.
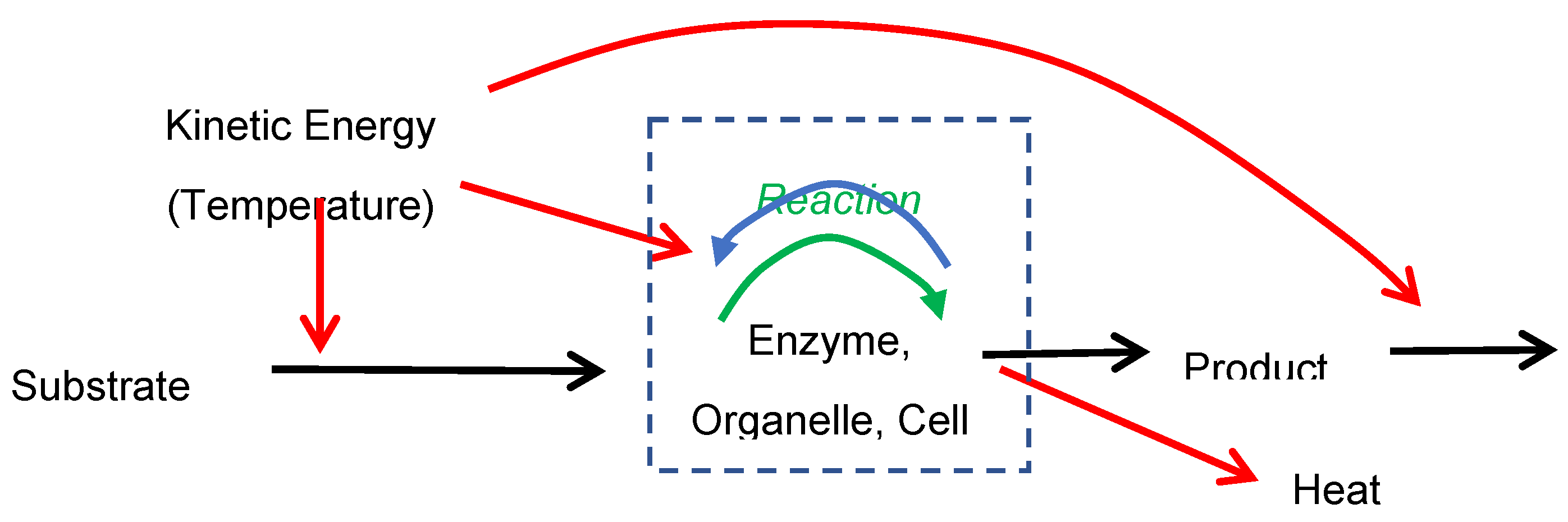
The operationally defined “system” in question will be the neighborhoods with substrate and product molecules close enough to catalytic enzymes for reactions to proceed, which I will call “reaction sites.” Materials, called substrates, move from the surrounding environment (outside the reaction site) to inside reaction sites (near the catalytic enzyme) and heat (if generated) and reaction products move from reaction sites into the surroundings outside (far from catalytic enzyme molecules). Reactions can be described as a “reaction-displacement” process, where substrates and products move into and out of reaction sites through various transport mechanisms such as diffusion, active transport, bulk flow, etc.
Figure 1.
Description of a reaction-displacement system, with substrate displaced into and product displaced (black arrows) out of the reaction site (dashed box) where conversion occurs through a forward (green arrow) and reverse reaction (blue arrow). The temperature of the surroundings is assumed to reflect kinetic energy of both displacement and reactions, and heat (if produced in an exothermic reaction) is dissipated to the surroundings outside the reaction site. Reaction sites can be operationally defined at a variety of levels of organizations, such as enzyme, organelle, etc.
Figure 1.
Description of a reaction-displacement system, with substrate displaced into and product displaced (black arrows) out of the reaction site (dashed box) where conversion occurs through a forward (green arrow) and reverse reaction (blue arrow). The temperature of the surroundings is assumed to reflect kinetic energy of both displacement and reactions, and heat (if produced in an exothermic reaction) is dissipated to the surroundings outside the reaction site. Reaction sites can be operationally defined at a variety of levels of organizations, such as enzyme, organelle, etc.
The reaction component of metabolism, and particularly exothermic reactions that generate heat (energy) or that drive the formation of energy carriers such as ATP or NADH, are reasonably well-understood (Haynie 2008). However, thermodynamics focuses on additional questions such as whether such reactions generate net negative Gibbs energy, are likely to run in reverse, and the degree to which they result in degradation of enzymes and other metabolic machinery or an increase in undissipated heat. All of these outcomes are associated with changes in entropy and the potential for the reaction to generate “useful” energy.
We begin with an evaluation of the thermodynamic potential of a reaction or the amount of useful or “free” energy generated from an exothermic (heat-producing) reaction. This potential depends on the ratio of the concentrations of substrate inside the reaction site, Ai, to product inside the reaction site, Pi and whether this exceeds the ratio at which the forward reaction and reverse reaction balance, also known as the equilibrium constant, Keq. Reactions with high Keq are described as “favorable” in that the forward reaction proceeds faster than the reverse even when product is at much higher concentration than substrate. Conversely, reactions with low Keq, even Keq << 1, are described as “unfavorable” because forward reactions are slower than reverse reactions even when substrate occurs at vastly higher concentrations than product. The ratio of actual concentrations of product : substrate relative to Keq, or chemical activity a, where a = Pi/(AiKeq), determines the magnitude of Gibbs energy generated.
From standard chemical thermodynamics, the actual free (or useful) energy produced by converting a unit of substrate (typically expressed in moles) into product is defined as Gibb’s energy Δ
Gr (kJ/mol) and is expressed as a negative value.
where ΔH is defined as the change in enthalpy (kJ/mol), the heat released by converting substrate into product, and ΔS
r is the change in entropy (kJ.K
-1mol
-1) within the system related to the work done (change in frequency, energy, and location of different molecules) in the overall transition from substrate to product. The frequency, energy, and spatial location of molecules in the vicinity of catalytic enzymes can be abstracted as “microstates,” and entropy change for a reaction is simply a change in the number of microstates over the course of the reaction. The subscript
r refers to these changes in the vicinity of the catalytic enzymes, or the reaction site.
Both change in enthalpy
∆Hr and entropy ∆
Sr near the catalytic enzymes are measured as kJ
oK
-1 mol
-1. Gibbs energy can also be expressed as a function of the chemical activity based on the ratio of product to substrate concentrations at the reaction site, as discussed above. Because ΔG
r is by definition negative, to make our subsequent measures of Gibb’s energy positive, we estimate
Immediately we see that the Gibbs energy produced per unit substrate will depend on the details of the reaction, and thus on reaction characteristics and displacement of molecules that determine the value of a.
A simple example is provided by a reaction-displacement model with an enzyme catalyzed reaction,
r = f(
k,Ai Z)
, a function with reaction constant
k, enzyme concentration
Z and reaction site substrate concentration
Ai. Diffusion or transport of substrate is driven by the concentration gradient between environmental (outside the reaction site) concentrations of substrate,
Ao, and
Ai. Likewise, the reaction yields product which accumulates to a concentration
Pi, and this product then diffuses or is transported away from the reaction site according to a concentration gradient
Pi to
Po, the product concentration outside the reaction site. Further, displacement is described as a diffusion or transport coefficient, multiplied by the concentration gradients for substrate and product.
This system of equations yields the steady state solutions (d
Ai/d
t = 0, d
Pi/d
t = 0) that derive the steady-state value of
a or
a*. A general result for a variety of reaction descriptions (Michaelis Menten, first order, etc.) suggests (Ritchie 2018):
This means that, as would be expected, greater Gibbs energy production will be associated with a greater ratio of substrate to product in the surroundings (Ao/Po), greater transport coefficient D, and greater reaction favorability, Keq. Counterintuitively, greater ΔGr is associated with less efficient (k) and less concentrated (Z) catalytic enzymes. More efficient and/or concentrated enzymes are likely to produce product faster reactions for the same capacity to displace products, which results in a greater steady- state product concentration Pi at the reaction site (inside the system), a faster reverse reaction and thus lower overall reaction and ΔGr. This outcome means that thermodynamic potential may be decoupled from and contrast with reaction rate in its response to reaction characteristics. Such a contrast can create trade-offs that ultimately influence the optimal temperature and other aspects of the response of organisms to temperature.
Gibbs energy and temperature dependence
To further explore how different metabolic “objectives” might produce different metabolic temperature dependence, we can now consider the temperature relationships embedded in the reaction-displacement model in equation (3). Over the range of
T below an optimal
Topt we can approximate the temperature dependence of the two coefficients
D for transport and
k for the reaction
where
R is the gas constant and
D0 and
k0 are normalization constants that incorporate specific information about the reaction and transport. The parameter
ED is the temperature sensitivity of diffusion or transport and Δ
G‡ is the Gibbs energy of
activation (not of the overall reaction as was introduced earlier). Activation energy is the energy required to form transition states with higher total bond energy (change in enthalpy), such as the enzyme-substrate complexes that form during an enzyme-catalyzed reaction, and the accompanying changes in frequencies of molecule types, their energy states, and positions (change in entropy) (see Fig. 2). Experimentally these are estimated as slopes of Arrhenius relationships (assumed to be linear) for the logarithm of rate versus 1/T (Fujita et al. 1960, Dell et al. 2011, Knapp and Huang 2022). However, it should be noted that “activation energies” estimated from empirical relationships between rates and temperature are first approximations of Δ
G‡ over certain ranges of
T, as increases in Δ
S‡ with temperature mean that Δ
G‡ is
not constant with temperature.
A recent meta-analysis (Ritchie 2018) shows that diffusion and active transport processes exhibit about 50% lower temperature sensitivity such that
ED ≈ Δ
G‡/2. This difference has significant ramifications because it means that, as temperature increases, the rate of displacement of molecules increases more slowly than the rate at which products are formed. If we assume these differences in temperature sensitivity in equation (5), we obtain:
which shows that chemical activity likely declines with increasing temperature (Δ
G‡ -
ED > 0) but that no temperature dependence would result if Δ
G‡ =
ED, which is what is usually assumed (Haynie 2008, Niven 2009).
The effect on Gibbs energy
of the overall reaction (converting substrate into product, not forming an intermediate or transition state) is obtained by substituting equation (6) into equation (3)
Gibbs energy for the overall reaction therefore increases with temperature and the factors discussed above that increase or decrease chemical activity. Note especially the positive dependence of ΔGr on the diffusion or transport coefficient, which shows that limits to the displacement of molecules to and from the reaction site can strongly affect the energy obtained from the reaction.
Figure 2.
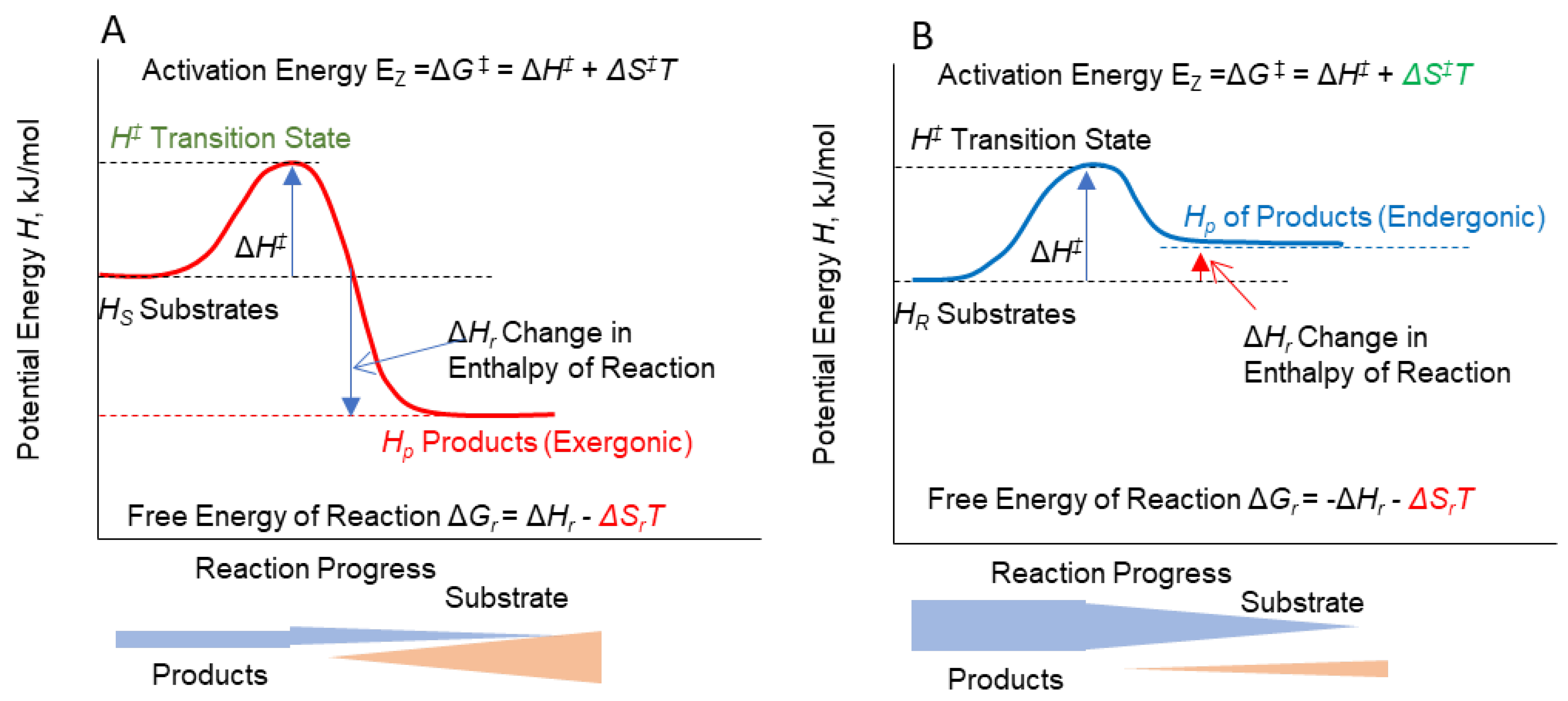
Conceptualization of energy of reaction activation and the Gibbs energy of the overall reaction for a conversion of substrate to product for two reaction types, both of which require input of an activation energy ΔG‡ composed of an increase in bond energy, ΔH‡ , from substrate to a transition state (enzyme-substrate or enzyme-product complex) and an accompanying increase in entropy ΔS‡ due to an increase in the number of molecule types, their energy state, and positions at the reaction site. A. exergonic reaction that produces heat and yields products with a lower total bond energy HP than the substrate HS, resulting in a release of Gibbs energy from the reaction as a whole. B. Endergonic reaction that requires heat for activation but products retain heat in the form of higher bond energy than the substrate.
Figure 2.
Conceptualization of energy of reaction activation and the Gibbs energy of the overall reaction for a conversion of substrate to product for two reaction types, both of which require input of an activation energy ΔG‡ composed of an increase in bond energy, ΔH‡ , from substrate to a transition state (enzyme-substrate or enzyme-product complex) and an accompanying increase in entropy ΔS‡ due to an increase in the number of molecule types, their energy state, and positions at the reaction site. A. exergonic reaction that produces heat and yields products with a lower total bond energy HP than the substrate HS, resulting in a release of Gibbs energy from the reaction as a whole. B. Endergonic reaction that requires heat for activation but products retain heat in the form of higher bond energy than the substrate.
Entropy and Steady-State
A hypothesis can be put forward that performance will cease when entropy production for the overall reaction ΔSr is less than entropy change inside the reaction site, ΔS‡ in order to satisfy both the Second Law of Thermodynamics and the requirement of persistence far-from-equilibrium ΔSr > ΔS‡. This inequality would correspond to the situation where entropy increases inside the reaction sites, due to formation of enzyme transition states and products in addition to substrates and enzymes, but is not balanced by an increase in entropy outside reaction sites, thus violating the Second Law of Thermodynamics. In practice this might correspond with damage to enzymes or reaction sites that would cause the reactions to stop and performance to cease. In any case, if entropy changes inside the reaction site exceed those outside, it should not be possible to achieve steady state.
Expanding from equation (10) in Ritchie (2018)
From equation (9), under the assumption that entropy of activation changes are driven by differences in heat capacity among transition states, we obtain
Re-evaluating inequality (1) and substituting equations (2) and (3), we obtain the condition for T
max as
This equation defines the conditions that satisfy the Second Law of Thermodynamics for the reaction at steady-state and can be solved numerically for
T as an estimate of
Tmax , and such solutions are useful because most of the component parameters are strongly bounded or averages are well known (
Table 1). Essentially, the reaction cannot maintain steady-state if this inequality is not satisfied and must either change the reaction characteristics, inactivate the enzymes, or otherwise cease the reaction.
Box 1 Entropy and Entropy Production
A key metric in thermodynamics is entropy, usually referred to with the variable
S. Technically it is defined as the number of “microstates” in a system. For example, the number of molecules for multiple chemical species, their position in space, energy states and physical conformation, all reflect relevant microstates. Such a direct measure of entropy in the context of an enzyme-catalyzed biochemical reaction could use the informatic metric (Kurzynski 2006)
where
pi,j,k might represent the proportion of all molecules of that belong to chemical species i, have energy states and conformations
j, and positions
k. While it is difficult to know all this information, researchers, however, can rather easily measure
changes in entropy, Δ
S, from changes in numbers of molecules of different chemical species associated with a specific chemical reaction. Without going into too many details (Haynie 2008), a change in Gibbs energy (equation (1)), includes energy incorporated in the changes in microstates that occur during activation (∆
SŦ) of the reaction, or from the overall conversion of substrate to product (∆
Sr). For example, a commonly cited source of change in entropy is changes in heat capacity that result from conformational changes in catalysts and transitions states during activation that may be measured from changes in the capacity of these molecules to absorb heat (Wynne-Jones and Eyring 1935, Hobbs et al. 2013, Arcus et al. 2016).
Figure 3.
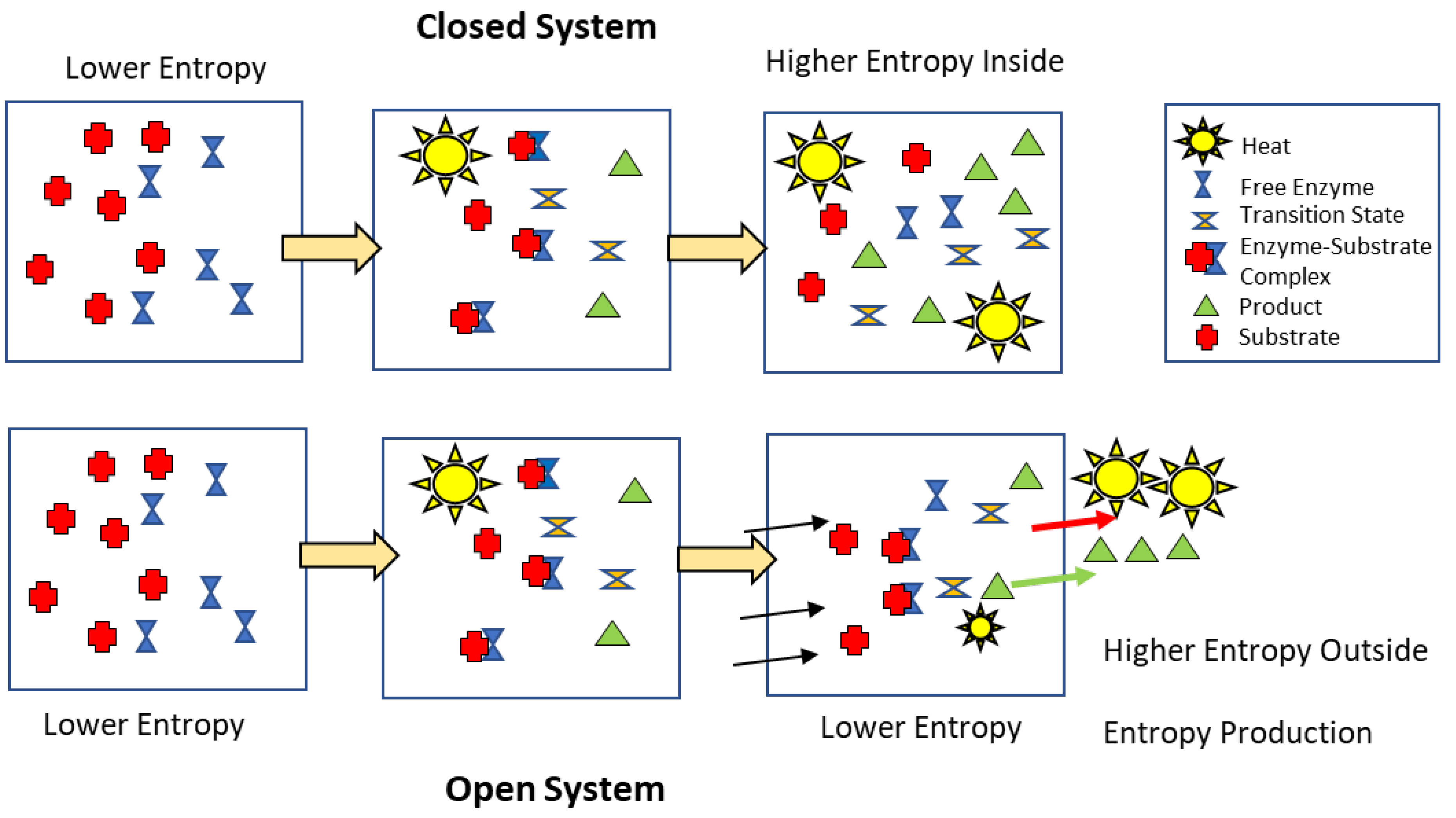
Following the example in the diagram, the hydrolysis of ATP in a hypothetical closed system will lead to an increase in entropy due to decline in the potential energy from the substrates ATP + H2O to the products ADP + PO42- + H2O and changes in the number of types of molecules, their energy states and positions and an increase in temperature of the system from the heat released by the reaction. In an open system where the reaction products can be displaced away from catalysts and heat dissipated, the entropy within the system (reaction site) increases much less because the flux of heat and molecules increases the entropy of the surroundings at a rate defined as entropy production.
Figure 3.
Following the example in the diagram, the hydrolysis of ATP in a hypothetical closed system will lead to an increase in entropy due to decline in the potential energy from the substrates ATP + H2O to the products ADP + PO42- + H2O and changes in the number of types of molecules, their energy states and positions and an increase in temperature of the system from the heat released by the reaction. In an open system where the reaction products can be displaced away from catalysts and heat dissipated, the entropy within the system (reaction site) increases much less because the flux of heat and molecules increases the entropy of the surroundings at a rate defined as entropy production.
Metabolic Goals and Testable Predictions
At this point we are ready to consider different metabolic “goals” that might be favored by natural selection. While many might be proposed, and some have been considered in previous studies (Hall and McWhirter 2023) , here I consider just three because they illustrate different predictions for how metabolism might change with temperature.
Maximum reaction rate – this hypothesis assumes that organisms have greater fitness if they can perform their reactions and thus replicate as fast as possible (Dean and Thornton 2007)
Maximum power – this hypothesis presumes that biological work derives from “useful” Gibbs energy, and that organisms will have greater fitness if they can generate greater Gibbs energy per unit time (Lotka 1922, Odum 1993, Hall 1995).
Maximum entropy production – this hypothesis presumes that fitness is enhanced by being able to perform reactions while maximizing the likelihood of persistence, that is, increase in entropy in the surroundings to minimize entropy increases at reaction sites (Hill and Simmons 1976, Dewar 2003, Jorgensen and Svirezhev 2004, Niven 2009, Martyushev 2013).
Superficially these may sound like very similar objectives, but as we will see, they predict very different responses to temperature. In a thermodynamic framework that assumes the reaction site is an open system, we will build the theoretical predictions for each of these hypotheses using the reaction-displacement system with the supply of substrates and formation of products in balance.
Figure 4.
Comparison of metabolic goal rates as a function of temperature A. Simulations with the parameters in
Table 1 of the response of steady-state entropy production (red), reaction rate (blue), and power (black), expressed as a % of their maximum rate at their respective T
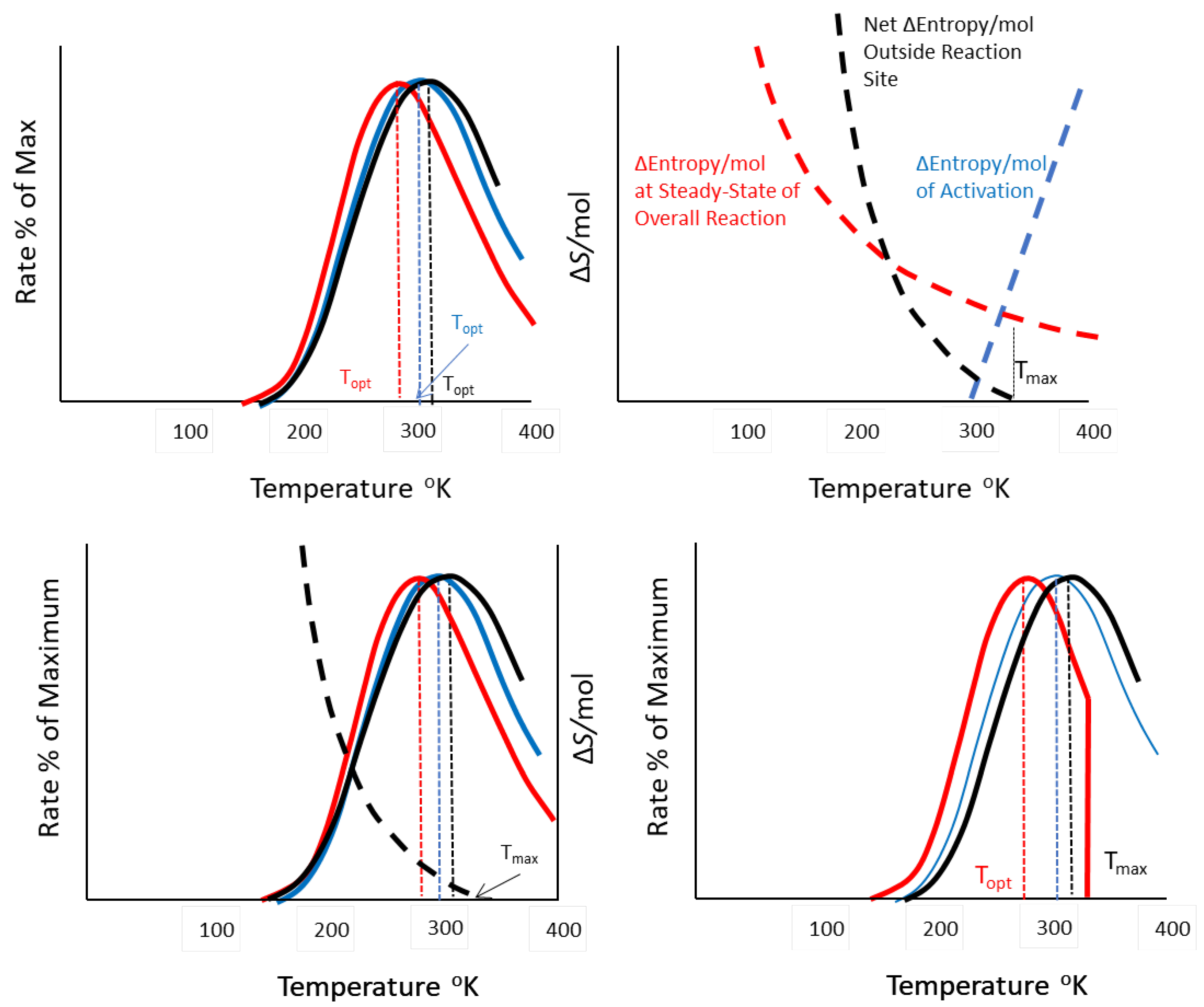
opt as a function of temperature. B. Changes in entropy produced per unit substrate outside (red dashed) and inside (blue dashed) the reaction site as a function of temperature, showing the maximum temperature, above which entropy increase outside the reaction system is less than inside, violating the conditions for steady-state. C. A double y-axis graph showing how the relationships in A and B overlap. D. Limits to entropy production at a maximum temperature Tmax producing a (red) temperature performance curve (TPC) that is sharply truncated above T
opt .
Figure 4.
Comparison of metabolic goal rates as a function of temperature A. Simulations with the parameters in
Table 1 of the response of steady-state entropy production (red), reaction rate (blue), and power (black), expressed as a % of their maximum rate at their respective T
opt as a function of temperature. B. Changes in entropy produced per unit substrate outside (red dashed) and inside (blue dashed) the reaction site as a function of temperature, showing the maximum temperature, above which entropy increase outside the reaction system is less than inside, violating the conditions for steady-state. C. A double y-axis graph showing how the relationships in A and B overlap. D. Limits to entropy production at a maximum temperature Tmax producing a (red) temperature performance curve (TPC) that is sharply truncated above T
opt .
Simulations of the three objective rates and the entropy change per unit substrate for activation and the reaction as a whole are shown as illustrations of generally how the temperature responses for each quantity compare (Fig. 4). The relevant steady-state rates for all three objectives show the expected temperature optimum Topt (Fig. 4). However, the objectives differ in their Topt although their values are surprisingly close and, after exploration of a variety of parameter inputs, generally differ by less than 10 oK. Interestingly, entropy production is maximized at the lowest temperature, followed by reaction rate, and then power, suggesting that persistence in more resource-poor habitats might be favored in cooler habitats for ectotherms or for endotherm species with lower core body temperatures. In contrast, power was maximized at the warmest temperatures, an outcome consistent with arguments for the evolution of endotherms, which trade-off efficiency and possibly longevity (for the same body size) for the ability to engage in high power activities at all times. These varying outcomes suggest testable hypotheses for discriminating among metabolic “objectives” and the response of metabolism to temperature.
In addition, the prediction of Tmax is above all the Topt predictions, but by only a few degrees, which is consistent with the pattern observed for in vivo measured physiological rates. This outcome suggests that warmer temperatures may force cells and organisms to “shut down” reactions prior to destruction or degradation of enzymes in order to maintain steady state. In any case these predictions of Tmax form a nascent beginning of a biochemical theory for the breadth of individuals’ and species’ thermal niches.
Finally, embedded in the chemical activity term in the calculation of Gibbs energy are a host of reaction characteristics and environmental influences on
Topt (Ritchie 2018) (Table 2). These dependences make many unique predictions about how performance at different temperatures depends on organism function and environmental resource conditions (Table 2). For example, organisms engaged primarily in synthesis (growth) or catabolic reactions will feature thermodynamically unfavorable synthesis reactions with a low ln(
Keq) and should exhibit lower
Topt and
Tmax (Ritchie 2018) (Fig. 5 A, B). In contrast, organisms with lower concentrations of enzymes are expected to have higher optimal temperatures (Ritchie 2018), and this pattern is very strong for reactions conducted with varying concentrations of
E. coli enzymes (Fig. 5 C).
| |
|
Effect on Optimal Temperature |
| Parameter |
Variable |
Transition State |
Entropy Production |
| Activation Energy |
ΔHŧ
|
+ |
+ |
| Change in Heat Capacity |
ΔCP
|
- |
NA |
| Reaction Constant |
k |
NA |
- |
| Enzyme Concentration |
Z |
NA |
- |
| Reaction Favorability |
Keq
|
NA |
+ |
| Environmental Substrate |
Ao
|
NA |
+ |
| Environmental Product |
Po
|
NA |
- |
| Diffusivity |
D |
NA |
+ |
These biochemical relationships extend to the field. A key prediction is that organisms with high concentrations of enzymes or other metabolic machinery will maximize entropy production and power at cooler temperatures. Fig. 5D shows an example for herbaceous plants, in which greater chlorophyll and RuBisco, the enzyme that catalyzes the carboxylation step of photosynthesis, account for the majority of plant nitrogen content (Chapin et al. 1986, Tateno and Chapin 1997). In these unpublished data, greater leaf nitrogen (%) in grasses and herbaceous dicots (forbs) was strongly correlated with lower leaf temperatures under the same radiative environment in both temperate and tropical grasslands. In contrast, greater substrate (resource) supply increases the thermodynamic gradient from substrate to product, and theoretically increases both Topt and Tmax. This prediction is consistent with a number of recent studies with aquatic organisms that demonstrate increased Topt and/or Tmax at greater substrate supply (Tang and Riley 2024) Conversely, this prediction also suggests that increasing resource limitation limits the ability of organisms to increase Topt and Tmax when environmental temperatures increase (Alster et al 2023, Tang and Riley 2024). A great example is the 4-5 oC lower Tmax (measured as CTmax) observed in oxygen-starved lizards as compared to those in normal atmospheric O2 (Claunch et al. 2021).
The predictions from the thermodynamic framework even have implications for the relationship between body size and temperature. Body size is expected to influence delivery rates of substrates and removal of reaction products to and from reaction sites. If so, key thermal niche parameters, such as Topt and Tmax may exhibit predictable declines with increasing body size. For example, delivery rates of substrate to cells and removal rates of products may determine the transport coefficient D in the chemical activity term 1/a that influences Topt and Tmax for entropy production and power. If mass-specific delivery rates to the reaction sites in the “invariant units” scale as expected (West et al 2005) as mass M-1/4, then the ln(1/a) term predicts that thermoregulated target body temperatures and maximum critical temperatures should scale as –(1/4) ln(M), implying a -0.25 oC decline in Tmax for every 2.76- fold increase in body mass. In addition, the upper and lower bounds on Tmax should range from 36 – 52 oC for the smallest (ants) to 32.6 – 48.5 for the largest ectotherms measured (~ 50 g lizard). A recent review of upper critical temperature CTmax, a laboratory measure that estimates Tmax, as a function of body mass for ectotherms (Leiva et al. 2019) shows clearly that such expected bounds encompass the vast majority of observations. Despite this variation Leiva et al (2019) found a significant decline in CTmax with body mass of various ectotherms and the observed slope of -0.502 (+ 0.22 SE) is not significantly different from the expected -0.25 (Fig. 5 E).
Figure 5.
A. – C. Patterns in temperature optima T
opt for in vitro enzyme -catalyzed reaction rates as a function of different rection characteristics (from data in Ritchie 2018). A. Significant (P < 0.001, N=79) increase in T
opt with increasing reaction favorability, measured as K
eq the ratio of product: substrate required for a reaction to proceed, expressed here as ln(K
eq). B. Significantly lower T
opt (P < 0.001) for anabolic (synthesis) reactions, which typically have low favorability, as compared to catabolic reactions, which typically have high favorability. C. Significant (P < 0.001) negative relationship between T
opt and the logarithm of enzyme concentration for reactions with E. coli hydrolysis enzymes. D. Unpublished mean leaf temperatures (N = 10 for each of 17 species from temperate Cedar Creek Ecosystem Science Reserve, Minnesota, USA and 10 species from tropical Serengeti National Park, Tanzania) for different herbaceous plant species (grasses and herbs) versus leaf nitrogen content (%). Regressions are significant (P < 0.003). E. Relationship for upper critical temperature (CT
max) versus body mass for various ectotherms, reviewed by Leiva et al (2019). Black line is the least squares regression fit to data. Red line is the predicted upper bound for Tmax as a function of body size using the numerical solution to equation (15) with the parameters in
Table 1 and assuming metabolic function is focused on locomotion and respiration with high favorability. Blue line is the predicted lower bound assuming metabolic function is focused on growth and/or reproduction with low favorability (biochemical synthesis).
Figure 5.
A. – C. Patterns in temperature optima T
opt for in vitro enzyme -catalyzed reaction rates as a function of different rection characteristics (from data in Ritchie 2018). A. Significant (P < 0.001, N=79) increase in T
opt with increasing reaction favorability, measured as K
eq the ratio of product: substrate required for a reaction to proceed, expressed here as ln(K
eq). B. Significantly lower T
opt (P < 0.001) for anabolic (synthesis) reactions, which typically have low favorability, as compared to catabolic reactions, which typically have high favorability. C. Significant (P < 0.001) negative relationship between T
opt and the logarithm of enzyme concentration for reactions with E. coli hydrolysis enzymes. D. Unpublished mean leaf temperatures (N = 10 for each of 17 species from temperate Cedar Creek Ecosystem Science Reserve, Minnesota, USA and 10 species from tropical Serengeti National Park, Tanzania) for different herbaceous plant species (grasses and herbs) versus leaf nitrogen content (%). Regressions are significant (P < 0.003). E. Relationship for upper critical temperature (CT
max) versus body mass for various ectotherms, reviewed by Leiva et al (2019). Black line is the least squares regression fit to data. Red line is the predicted upper bound for Tmax as a function of body size using the numerical solution to equation (15) with the parameters in
Table 1 and assuming metabolic function is focused on locomotion and respiration with high favorability. Blue line is the predicted lower bound assuming metabolic function is focused on growth and/or reproduction with low favorability (biochemical synthesis).

Scaling up to biological performance.
Biological performance can be operationally defined at many scales of organization, from individual molecular machines, such as ribosomes, to membranes with multiple catalytic enzymes, to organelles such as mitochondria, endoplasmic reticula, to entire cells, to entire multicellular organisms. While the vast work on the physics and chemistry of enzyme catalyzed reactions provides a basis for hypotheses about how metabolic rate responds to temperature, metabolism is a phenomenon of organisms. As such, it is the result of dozens to hundreds of different types of reactions that may vary spatially within cells (among organelles) and among cells (specializations and organs). Because entropy and or Gibbs energy are extensive quantities (they add across time and space), framing metabolism in these currencies makes it easier to scale up from the biochemical to higher levels of organization.
The thermodynamic framework presented here, and particularly the quantification of metabolism through entropy changes over time, has the potential to explicitly scale up understanding of biochemical reactions to whole-organism, population, community, and ecosystem metabolism. The alignment of data with entropy-driven and -constrained reaction rates suggests that an important feature of entropy can allow results to be scaled up across multiple reaction types and replications of reaction sites across multicellular organisms: entropy is an extensive physical quantity, which means it adds across multiple localities, entities and across space and time. Moreover, techniques for assessing frequency distributions of the contribution of different reaction types, such as MaxEnt methods (Harte 2011) can help coalesce the vast amount of biochemical detail that normally daunts efforts to scale up.