1. Introduction
Head and neck squamous cell carcinoma (HNSCC) represents a significant global health burden, with tongue squamous cell carcinoma (TSCC) emerging as one of its most aggressive and clinically challenging subtypes [
1]. TSCC is characterized by a constellation of distinct biological features: early local invasion into the intricate musculature of the tongue, a high propensity for bilateral cervical lymph node metastasis (particularly to levels I–III) [
2], and an increasing incidence among younger patients, frequently independent of HPV infection [
3]. These anatomical and molecular attributes collectively contribute to poor clinical outcomes [
4]. Moreover, TSCC tumors frequently exhibit intrinsic resistance to conventional radiotherapy and chemotherapy. For patients with recurrent or metastatic disease, the prognosis remains dismal, with limited therapeutic options beyond first-line regimens [
5]. Consequently, there is an urgent need to elucidate the molecular drivers underlying TSCC pathogenesis and to identify novel targets for more effective therapeutic strategies.
To identify key oncogenic drivers in HNSCC, we interrogated the TCGA database for prognosis-associated genes and identified Caveolin-2 (CAV2) as a top candidate. CAV2 is a core structural component of caveolae—plasma membrane invaginations abundantly expressed in epithelial cells [
6,
7]. While elevated CAV2 expression has been linked to poor prognosis in malignancies such as lung and pancreatic cancer [
8,
9,
10], its functional role in HNSCC remained unexplored, prompting our investigation. Subsequent immunoprecipitation-mass spectrometry (IP-MS) and proteomic analyses revealed that CAV2 interacts with the double-stranded RNA-dependent protein kinase PKR, a key mediator of cellular stress responses frequently dysregulated in cancer [
11,
12]. Notably, PKR expression was significantly upregulated upon CAV2 knockdown. Given that PACT serves as the primary activator of PKR under stress conditions [
12,
13], we further demonstrated that CAV2 interacts with and modulates PACT expression, positioning CAV2 upstream of the PACT–PKR stress signaling axis.
The clinical relevance of these findings is particularly significant in HNSCC, where over 90% of patients exhibit elevated EGFR expression yet demonstrate limited and heterogeneous responses to the EGFR-targeted antibody cetuximab [
14,
15]. Resistance to cetuximab—whether intrinsic or acquired—represents a major therapeutic challenge. In this context, we discovered that CAV2 depletion markedly enhances the sensitivity of HNSCC cells to cetuximab, suggesting a mechanistic link between CAV2-driven signaling and therapeutic resistance.
Here, we elucidate the functional role of CAV2 in HNSCC proliferation and delineate its molecular mechanism of action. Our findings reveal a previously unrecognized oncogenic pathway in which CAV2 drives tumor growth by regulating the PACT–PKR axis via the ubiquitin–proteasome system. Furthermore, we establish that targeting CAV2 sensitizes HNSCC cells to cetuximab, positioning CAV2 as a promising therapeutic target to overcome treatment resistance in this disease.
2. Materials and Methods
Human Head and Neck Squamous Cell Carcinoma Tissue Samples
In the study, samples were gathered from patients undergoing surgical removal of HNSCC at the Department of Maxillofacial and Otorhinolaryngology Oncology and Department of Head and Neck Oncology, Cancer Hospital of Tianjin Medical University from 2024.10-2024.12. The tumor samples from all patients were pathologically identified as HNSCC. Cancer tissues and their adjacent tissues were collected separately and frozen using liquid nitrogen rapidly. The preparation of protein extracts involved homogenizing tissues in a cold environment, followed by centrifugation at 4 °C. The supernatant was then collected for Western blotting analysis.
Cell Culture, Chemicals, and Reagents
In this study, the human HNSCC cell lines SCC15 and SCC25 were acquired from the American Type Culture Collection (ATCC) and supplied by the Department of Tumor Cell Biology at the Cancer Research Institute, Tianjin Cancer Hospital. The cell lines were grown in DMEM/F-12 medium (Corning, USA) with an addition of 10% fetal bovine serum (FBS; PAN-Biotech) and 1% penicillin/streptomycin (Hyclone). The HEK293T cell line, derived from human embryonic kidney cells and sourced from the American Type Culture Collection, was cultured in DMEM (Corning, USA) with the addition of 10% FBS and 1% penicillin/streptomycin. A humidified atmosphere with 5% CO₂ was used to incubate all cell lines at 37 °C.
CAV2 Expression Variability and Prognostic Analysis
Stable Transfected Cell LINES establishment
Lentiviral vectors encoding non-targeting control (Ctrl), shPKR, and shPACT were obtained from GeneChem (Shanghai, China). To create lentiviral particles, HEK293T cells were co-transfected with the respective transfer plasmid (shCtrl, shCAV2, shPKR, or shPACT) and the packaging plasmids VSVG and δR using polyethylenimine (PEI; Polysciences). Viral supernatants were obtained 48h following transfection. Target cells were infected using polybrene (Solarbio) and selected with 1μg/mL puromycin (Gibco) for a duration of 5 days to establish stable knockdown pools. The efficiency of the knockdown was verified using Western blotting and qRT-PCR.
Western Blotting and Antibodies
Proteins were isolated using SDS-PAGE at 80–140V and then transferred onto Immobilon-P membranes from Millipore. The membranes were blocked with 5% non-fat milk or 3% BSA in TBST for an hour at room temperature before being incubated with primary antibodies overnight at 4℃. The day after, membranes were incubated with HRP-conjugated secondary antibodies for one hour at room temperature, and signals were detected using an ECL kit by Pierce.
The following antibodies were employed: CAV2 (1:1000, Novus, NBP1-31116), PKR (1:1000, CST, 12297S), PACT (1:1000, CST, 13490S), Ki67 (1:1000, CST, 62548SF), Ubiquitin (1:1000, CST, 3936S), PARP (1:1000, CST, 9532S), HRP-conjugated Goat Anti-Rabbit IgG (1:4000, Proteintech, SA00001-2), and HRP-conjugated Goat Anti-Mouse IgG (1:4000, Proteintech, SA00001-1).
EdU Assay
The BeyoClick™ EdU Cell Proliferation Kit with Alexa Fluor 594 (Beyotime, C00788L) was utilized to assess cell proliferation. Cells were seeded in 48-well plates at a density of 1×10⁴ cells per well and cultured for 24 hours. The medium was then supplemented with 10μM EdU, followed by incubation for 2h at 37 °C under 5% CO₂.
Cells were washed with PBS, fixed with 4% paraformaldehyde for 30 minutes at room temperature, and permeabilized with 0.3% Triton X-100 for 15 minutes after the EdU-containing medium was removed. To visualize incorporated EdU, the Click reaction was carried out following the manufacturer’s guidelines. DAPI was used to counterstain the nuclei for 10 minutes, and images were captured with an inverted fluorescence microscope equipped with a digital camera. Three distinct replicates were carried out for each condition in the experiment.
Cell Viability Assay
Cells were placed in 96-well plates at a density of 1,000 cells per well in 100μL of culture medium. At each time of measurement, 10μL of CCK-8 solution (Dojindo Laboratories, Japan) was added to each well, followed by incubation at 37 °C for 2-4h. A microplate reader was used to measure the optical density at 450 nm. Cell proliferation curves were generated from measurements taken continuously over a period of 4-5 days. Each experiment was conducted with a minimum of three independent replicates. The CCK-8 absorbance at 450 nm was adjusted by removing the blank control and then normalized to Day 0, where the fold change is set to 1. Cell growth was determined by dividing the corrected optical density on any given day by the corrected optical density on Day 0.
RNA Extraction and qRT-PCR
Trizol reagent (Ambion, Austin, TX, USA) was utilized to extract total RNA from adherent cells following the manufacturer’s guidelines, and RNA quantification was achieved using a NanoDrop spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA). PrimeScript RT Master Mix (TaKaRa, Japan) was utilized for cDNA synthesis. qRT-PCR was performed using AceQ SYBR qPCR Master Mix (Vazyme, Q111) on a BioRad CFX96 system, as instructed. The primers used are listed in the following.
IHC
The Tianjin Medical University Cancer Institute and Hospital provided tissue sections embedded in paraffin. These sections were deparaffinized with xylene and rehydrated through a graded series of ethanol. Citrate buffer (pH=6.0) was used for antigen retrieval at 95 °C for 15 minutes, followed by blocking endogenous peroxidase activity with 3% hydrogen peroxide at room temperature for 15 minutes. The sections were incubated overnight at 4 °C with primary antibodies targeting CAV2 (1:100, Novus, NBP1-31116) or Ki67 (1:200, CST, 62548SF) diluted in an antibody diluent. The sections were incubated with an HRP-conjugated secondary antibody (PV-6001 kit, Zhongshan Biotechnology) for one hour at 37 °C after washing. For signal detection, a DAB substrate kit (Zhongshan Biotechnology) was used, followed by counterstaining with hematoxylin. Images were acquired using a brightfield microscope (Olympus BX61) with different objective lenses.
TUNEL Assay
Cells were placed on sterile coverslips in 12-well plates a day before the experiment. After incubation, the cells were treated with 4% paraformaldehyde at room temperature for 30 minutes and then permeabilized using 0.3% Triton X-100 for 10 minutes at room temperature. Following the manufacturer’s instructions, apoptotic cells were detected using a TUNEL assay kit by applying 50μL of TUNEL reaction mixture to each sample and incubating at 37 °C for one hour in the dark. The nuclei were counterstained with DAPI for 10 minutes after PBS washing. Coverslips were mounted with an anti-fade medium and examined under a fluorescence microscope.
Co-Immunoprecipitation (Co-IP)
A commercial kit (Proteintech, PK10008) was used to perform Co-IP assays according to the manufacturer’s instructions, with minor adjustments. The adherent cells were washed three times with pre-chilled PBS and lysed in IP lysis buffer. The lysates were spun at 12,000g for 10 minutes at 4 °C to clear them. Equal protein lysate amounts were incubated overnight at 4 °C with either the target antibody or a species-matched IgG control. Protein A/G beads were subsequently added and incubated for 2h at 4 °C with gentle rotation. The beads were washed 4–5 times with 800μL of wash buffer, and the proteins that were bound were eluted using an elution buffer. The samples were then neutralized, combined with 5× SDS loading buffer, and heated at 95 °C for 5 minutes.
Flow Cytometry Analysis
Cell apoptosis was examined using the PE Annexin V Apoptosis Detection Kit (BD Pharmingen™, 559763). After washing with chilled PBS, cells were resuspended in 1× Binding Buffer at a concentration of 1×10⁶ cells/mL. 100μL cell suspension was placed into a flow cytometry tube and mixed with 5μL of PE Annexin V and 5μL of 7-AAD, then left to incubate for 15 minutes at room temperature in the dark. After incubation, 400μL of 1× Binding Buffer was added to each tube. Samples were subjected to flow cytometry using a BD FACS instrument within an hour. The analysis distinguished between viable cells (Annexin V⁻/7-AAD⁻), early apoptotic cells (Annexin V⁺/7-AAD⁻), late apoptotic cells (Annexin V⁺/7-AAD⁺), and necrotic cells (Annexin V⁻/7-AAD⁺).
Apoptosis Enzyme-Linked Immunosorbent Assay (ELISA)
Using the Cellular DNA Fragmentation ELISA kit (Roche, 11585045001), apoptosis was quantitatively assessed according to the manufacturer’s instructions. Cells were seeded in 96-well plates at a density of 1×10⁴ cells per well and pre-labeled with BrdU for 24 hours. Cells were treated with the apoptosis inducer CCCP (10μM) and incubated for 1-6 hours at 37 °C. After incubation, the cells were collected by centrifuging at 300 ×g for 10 minutes, and the supernatant was gently removed. Cell pellets were lysed with the incubation buffer supplied for 30 minutes at room temperature, then centrifuged at 800 ×g for 10 minutes to collect the supernatant with fragmented DNA. The supernatant underwent the usual ELISA process to measure DNA fragmentation. Absorbance was recorded at 450 nm with a microplate reader, and the results were compared to untreated controls. Three independent experiments were performed in triplicate.
Proximity Ligation Assay
The Duolink® PLA was conducted on paraffin-embedded HNSCC tissue sections from the Department of Maxillofacial, Ear, Nose and Throat Oncology at Tianjin Medical University Cancer Hospital to identify protein-protein interactions in situ. Sections were deparaffinized and rehydrated, then antigen retrieval was performed using a sodium citrate buffer with a pH of 6.0. The Duolink® InSitu Red Starter Kit Mouse/Rabbit (Sigma-Aldrich) was utilized as per the manufacturer’s guidelines. The primary antibodies used were CAV2 (Rabbit, Novus, 1:100), PKR (Rabbit, Proteintech, 1:100 or Mouse, Santa Cruz, 1:100), and PACT (Mouse, Santa Cruz, 1:100), and they were incubated overnight at 4 °C in a humidified chamber. After washing, the steps of ligation and amplification were performed as directed. Nuclei were counterstained with a mounting medium containing DAPI from the kit. Images were captured using a fluorescence microscope. hree independent experiments were performed with appropriate controls.
Statistical Analysis
Differences between the two groups were assessed using Student’s t-test, and the Wilcoxon rank sum test was utilized for analyzing expression differences. The prognostic outcomes were analyzed using the univariate Kaplan–Meier method and the multivariate Cox proportional hazards model, with data evaluation performed using R version 4.2.1. Differences were deemed statistically significant if the p-value was below 0.05.
4. Discussion
Globally, head and neck squamous cell carcinoma (HNSCC) is a significant health issue, ranking among the top common cancers[
25]. In China alone, the disease incidence and associated mortality remain alarmingly high, with tens of thousands of new cases and deaths projected annually[
26]. Although the majority of patients have locally advanced disease, which is linked to worse survival rates, treatment options are still limited[
27]. In particular, resistance, whether inherent or acquired, often diminishes the efficacy of EGFR-targeted treatments like cetuximab, underscoring the urgent need to find new targets and combination strategies to improve outcomes.
Caveolin-2 (CAV2), a core structural component of caveolae, has emerged from our TCGA-based screening as a strong prognostic indicator in HNSCC[
28]. CAV2 is the most strongly associated molecule with patient prognosis in head and neck squamous carcinoma that we mined from the TCGA database. Although CAV1, its well-characterized paralog, has been extensively studied in cancer biology[
29,
30], the functional role of CAV2 remains comparatively undefined, particularly in HNSCC.
In this study, we systematically defined the oncogenic role of CAV2 in HNSCC. Analysis of public databases and our clinical specimens consistently revealed that CAV2 is highly expressed in HNSCC tissues, and its elevated expression correlates with poor patient prognosis. Analysis of public databases along with our clinical specimens consistently demonstrated that CAV2 is prominently expressed in HNSCC tissues, and its elevated levels are connected to poor patient outcomes.
Proteomic profiling identified PKR as a key molecule associated with CAV2. Although PKR was initially recognized for its role in viral infection, recent studies have implicated it in diverse biological processes, including endoplasmic reticulum stress, oxidative stress, and tumorigenesis [
31,
32,
33]. In our models, CAV2 silencing led to upregulation of PKR, which subsequently promoted apoptosis. Given that PACT is a well-established upstream activator of PKR and facilitates its function through direct binding, we initially hypothesized that CAV2, PKR, and PACT form a ternary complex, with CAV2 acting as a scaffold to modulate the PKR–PACT interaction. However, co-immunoprecipitation assays refuted this assumption, indicating that the three proteins do not form a stable complex but instead engage in pairwise interactions independent of the third partner. Since CAV2 knockdown did not alter the mRNA levels of PKR or PACT, we postulated that CAV2 regulates their expression post-transcriptionally. Using cycloheximide (CHX) chase assays, we found that the stability of both PKR and PACT was enhanced in CAV2-knockdown cells. Furthermore, among several protease pathway inhibitors, only the proteasome inhibitor MG132 caused significant accumulation of PKR and PACT. Subsequent ubiquitination immunoprecipitation confirmed that CAV2 depletion promotes polyubiquitination and proteasomal degradation of both proteins, indicating that CAV2 stabilizes PKR and PACT by inhibiting their ubiquitin-mediated degradation.
PKR has been observed to have context-dependent functions in different cancers, where it acts as an oncogene in breast cancer, melanoma, and colon cancer, yet it seems to serve as a tumor suppressor in HNSCC[
9,
34], consistent with our findings. In rescue experiments, concurrent knockdown of PACT or PKR partially reversed the anti-proliferative effect caused by CAV2 deletion, supporting the notion that the PACT–PKR axis operates downstream of CAV2[
35]. However, PKR was identified as a tumor suppressor in head and neck squamous cell carcinoma cells[
36,
37], which is consistent with our experimental results. The RESCUE experiments found that PACT-PKR knockdown could partially rescue the decline in cell malignancy caused by CAV2 deletion.
PKR suppresses tumor growth primarily by phosphorylating eIF2α, which reduces overall protein synthesis during stress[
34]. Phosphorylation of the eIF2α is a core mechanism for the downregulation of protein synthesis under various stress conditions [
38,
39]. PKR can phosphorylate eIF2α to shut down the most common translation initiation using the eIF2 complex[
35]
. Phosphorylated eIF2α expression is upregulated after CAV2 silence (Supplementary 2A), suggesting that CAV2 deletion can shut down the core pathway of protein translation, explaining the mechanism by which CAV2 can promote tumor progression from another perspective. eIF2α is central in endoplasmic reticulum stress[
40]. We observed upregulation of phosphorylated eIF2α following CAV2 silencing, suggesting that CAV2 deletion attenuates protein synthesis via the PKR–eIF2α pathway. We further activated endoplasmic reticulum (ER) stress using tunicamycin in PKR- or PACT-deficient cells. In these cells, tunicamycin failed to significantly suppress proliferation, underscoring the essential role of the PACT–PKR axis in mediating ER stress-induced growth inhibition in HNSCC (Supplementary 2B, C). In contrast to the prevailing model[
31,
41], we observed that PKR and its upstream activator, PACT, were concordantly upregulated following CAV2 knockdown.
The majority of HNSCC tumors have high levels of EGFR expression and activate the EGFR/PI3K/AKT pathway[
14], although cetuximab, an EGFR monoclonal antibody, is FDA-approved for HNSCC treatment, its effectiveness is limited due to acquired resistance. In an orthotopic xenograft model, CAV2-deficient tumors showed reduced sensitivity to cetuximab compared with controls. This finding positions CAV2 as a potential target for combination therapy, which may help overcome cetuximab resistance in HNSCC patients.
Our study systematically elucidates the oncogenic role of CAV2 in HNSCC and its underlying molecular mechanisms. We demonstrated that CAV2 is significantly overexpressed in HNSCC tissues and cell lines, and the expression level is closely linked to a poor prognosis in patients. Functional tests showed that reducing CAV2 enhances the breakdown of the PACT–PKR axis through the ubiquitin–proteasome pathway, thereby inhibiting tumor proliferation and inducing apoptosis. Rescue experiments further confirmed that PACT–PKR acts as a critical downstream effector through which CAV2 exerts its tumor-promoting functions. Notably, CAV2 does not serve as a scaffold protein facilitating direct interaction between PACT and PKR; rather, it regulates their protein stability to modulate the activity of this signaling axis.
Our findings are consistent with and extend recent studies on HNSCC progression and drug resistance. MRPL21 has been found to promote cisplatin resistance in HNSCC by activating the PI3K/AKT/mTOR pathway and blocking autophagy [
42]. Similarly, mutations in the FAT1 gene have been found to promote resistance to immune checkpoint inhibitors by shaping an immunosuppressive tumor microenvironment [
43]. These studies collectively suggest that the malignant progression and therapeutic resistance of HNSCC involve the coordinated dysregulation of multiple signaling pathways. Our work positions CAV2 and its regulated PACT–PKR axis within this complex molecular network, providing a novel perspective for understanding HNSCC pathogenesis.
Figure 1.
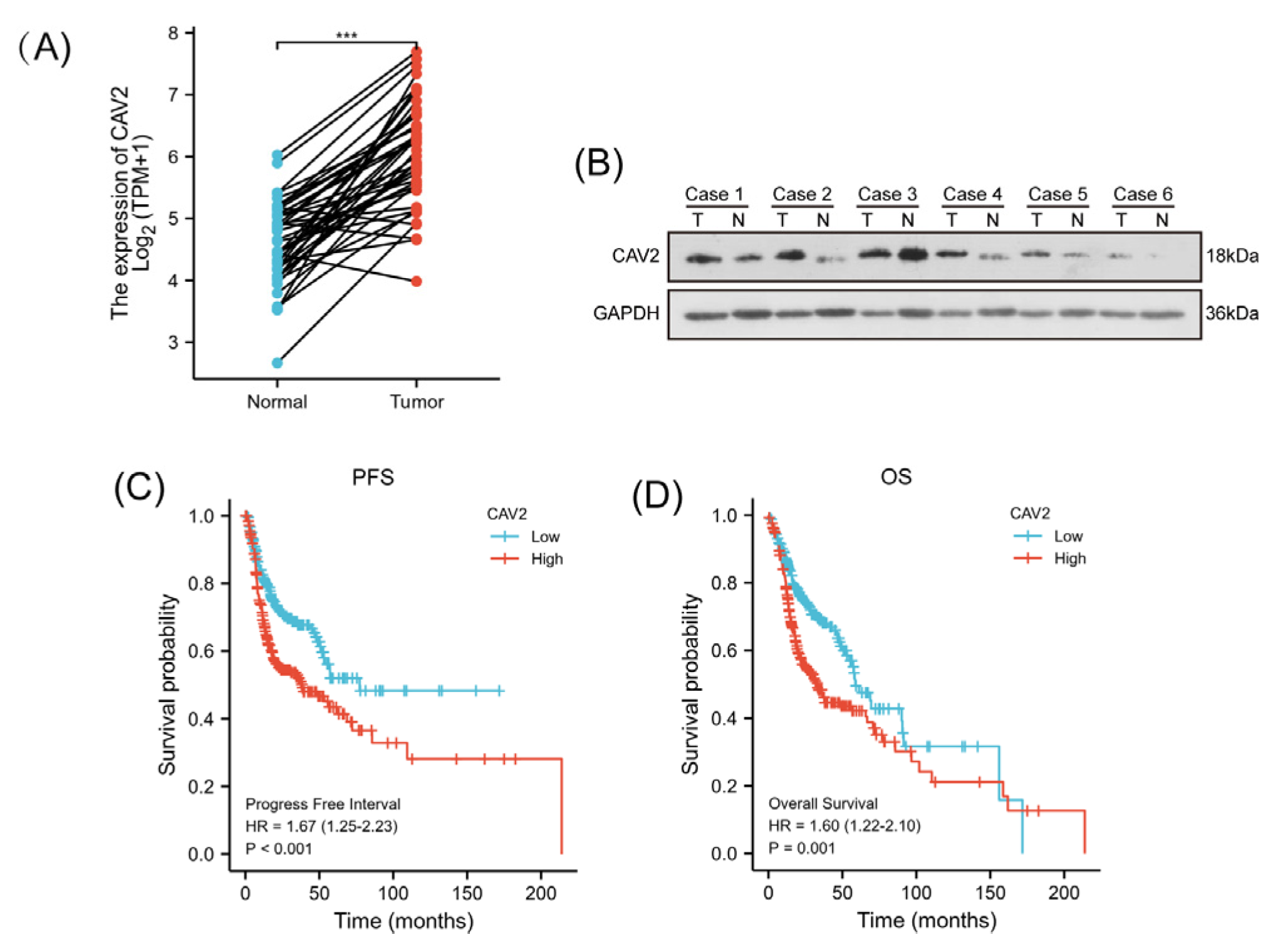
CAV2 is overexpressed in HNSCC and correlates with poor patient prognosis. (A). The TCGA dataset analysis reveals CAV2 mRNA expression in tumor and corresponding normal tissues across 43 HNSCC (T) pairs and adjacent non-tumor tissues (N). (B). Western blot was used to analyze CAV2 in five pairs of human HNSCC tumors and nearby non-cancerous tissues, with GAPDH serving as a loading control (C, D). Kaplan–Meier survival curves based on TCGA dataset showing progress-free survival (C) and overall survival (D) of HNSCC patients stratified by high vs. low CAV2 expression. Log-rank test was used for statistical comparison.
Figure 1.
CAV2 is overexpressed in HNSCC and correlates with poor patient prognosis. (A). The TCGA dataset analysis reveals CAV2 mRNA expression in tumor and corresponding normal tissues across 43 HNSCC (T) pairs and adjacent non-tumor tissues (N). (B). Western blot was used to analyze CAV2 in five pairs of human HNSCC tumors and nearby non-cancerous tissues, with GAPDH serving as a loading control (C, D). Kaplan–Meier survival curves based on TCGA dataset showing progress-free survival (C) and overall survival (D) of HNSCC patients stratified by high vs. low CAV2 expression. Log-rank test was used for statistical comparison.
Figure 2.
CAV2 promotes HNSCC cell proliferation in vitro and in vivo.(A). The infection of SCC15 and SCC25 cells with lentivirus carrying shRNA against CAV2 was performed. Knockdown efficiency was confirmed by Western blotting for proteins and qRT-PCR for mRNA, with GAPDH as the loading control. (B) Colony formation assay and (C-D) EdU staining were performed to assess proliferative capacity in control and CAV2-knockdown SCC15(C) and SCC25(D) cell lines. Scale bar, 50μm. The bar diagram represents the fraction of EdU-positive cells. (E-F). Cell growth curves of SCC15(E) and SCC25(F) cells following CAV2 knockdown. The Y-axis shows the fold change in cell count compared to Day 0. The data are expressed as mean ± SEM from three separate experiments. (G). Images representing subcutaneous xenograft tumors from control and CAV2-knockdown cells. (H). Immunohistochemical staining of CAV2 and the proliferation marker Ki67 in tumor sections. Scale bar, 100μm. (I). Tumor size was tracked from day 6 following inoculation, and the tumor’s weight was documented at the endpoint (J). Data are presented as mean ± SEM; n = 13 mice per group. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; ns, not significant.
Figure 2.
CAV2 promotes HNSCC cell proliferation in vitro and in vivo.(A). The infection of SCC15 and SCC25 cells with lentivirus carrying shRNA against CAV2 was performed. Knockdown efficiency was confirmed by Western blotting for proteins and qRT-PCR for mRNA, with GAPDH as the loading control. (B) Colony formation assay and (C-D) EdU staining were performed to assess proliferative capacity in control and CAV2-knockdown SCC15(C) and SCC25(D) cell lines. Scale bar, 50μm. The bar diagram represents the fraction of EdU-positive cells. (E-F). Cell growth curves of SCC15(E) and SCC25(F) cells following CAV2 knockdown. The Y-axis shows the fold change in cell count compared to Day 0. The data are expressed as mean ± SEM from three separate experiments. (G). Images representing subcutaneous xenograft tumors from control and CAV2-knockdown cells. (H). Immunohistochemical staining of CAV2 and the proliferation marker Ki67 in tumor sections. Scale bar, 100μm. (I). Tumor size was tracked from day 6 following inoculation, and the tumor’s weight was documented at the endpoint (J). Data are presented as mean ± SEM; n = 13 mice per group. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; ns, not significant.
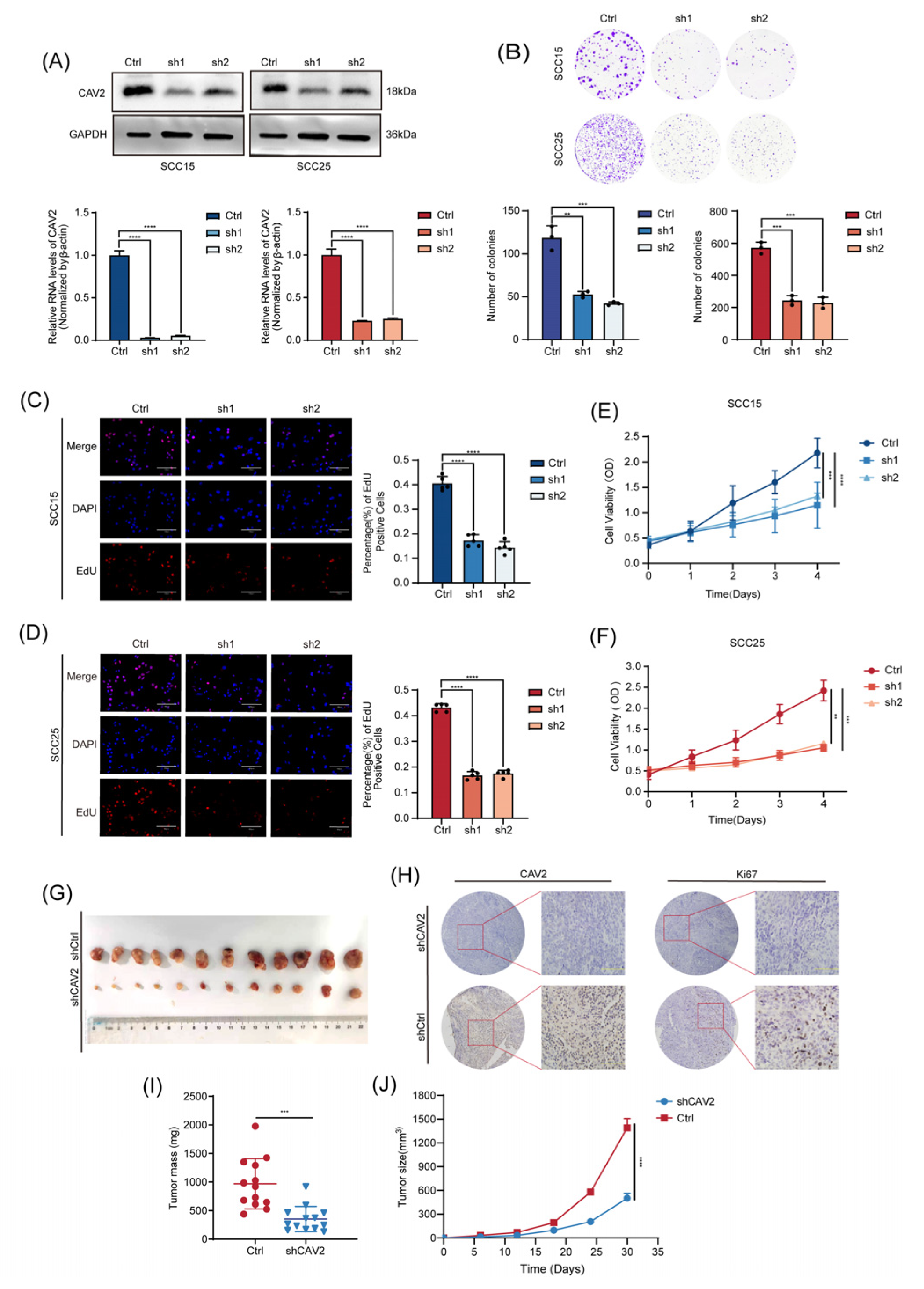
Figure 3.
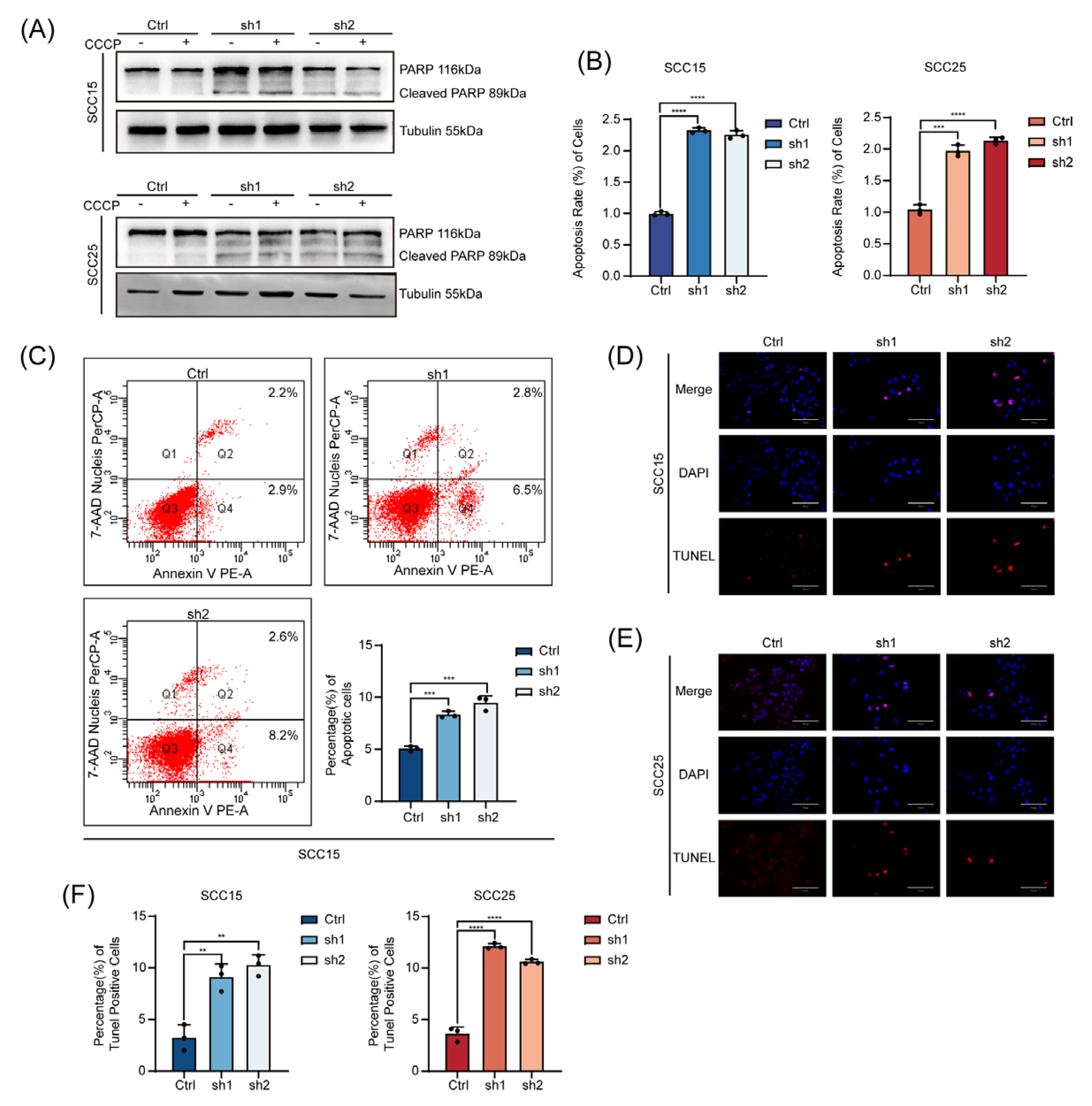
CAV2 suppresses apoptosis in HNSCC cells. (A). Western blot was used to analyze cleaved PARP protein levels in SCC15 and SCC25 cells with CAV2 knockdown and control, both with and without CCCP treatment. GAPDH served as the loading control. (B-C). The apoptotic rates in SCC15 and SCC25 cells were determined using Annexin V/7-AAD double staining post-CAV2 knockdown(B). Representative images of Flow cytometry in control and CAV2-knockdown SCC15(C). (D-E). Representative images of TUNEL staining in control and CAV2-knockdown SCC15 (D) and SCC25 (E) cell lines. Scale bar, 50μm. (F). Quantification of TUNEL-positive cells from multiple fields. Data are presented as mean ± SEM from three independent experiments. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; ns, not significant.
Figure 3.
CAV2 suppresses apoptosis in HNSCC cells. (A). Western blot was used to analyze cleaved PARP protein levels in SCC15 and SCC25 cells with CAV2 knockdown and control, both with and without CCCP treatment. GAPDH served as the loading control. (B-C). The apoptotic rates in SCC15 and SCC25 cells were determined using Annexin V/7-AAD double staining post-CAV2 knockdown(B). Representative images of Flow cytometry in control and CAV2-knockdown SCC15(C). (D-E). Representative images of TUNEL staining in control and CAV2-knockdown SCC15 (D) and SCC25 (E) cell lines. Scale bar, 50μm. (F). Quantification of TUNEL-positive cells from multiple fields. Data are presented as mean ± SEM from three independent experiments. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; ns, not significant.
Figure 4.
CAV2 interacts with the PACT-PKR axis but does not function as a scaffold. (A). Workflow of quantitative proteomic analysis performed in control versus CAV2-knockdown SCC15 cells. (B). Volcano plot displaying differentially expressed proteins following CAV2 silencing. Significantly upregulated (Red) and downregulated (Blue) proteins are highlighted (Fold change > 1.5, p < 0.05). (C-D). Co-immunoprecipitation (Co-IP) assays in SCC15(C) and SCC25(D) cells using antibodies against CAV2, PKR, and PACT, followed by immunoblotting with the indicated antibodies. Total cell lysates (Input) and IgG controls are shown. (E). Duo-link PLA was performed on human HNSCC tissue sections to detect protein-protein interactions between CAV2, PKR, and PACT. Red fluorescent dots represent specific protein interactions. Scale bar, 50μm. (F). Co-IP of endogenous PKR and PACT in control and CAV2-knockdown SCC15 cells. (G). Co-IP of CAV2 and PACT in control and PKR-knockdown SCC15 cells. (H). Co-IP of CAV2 and PKR in control and PACT-knockdown SCC15 cells. All Co-IPs were performed with the indicated antibodies and analyzed by western blotting.
Figure 4.
CAV2 interacts with the PACT-PKR axis but does not function as a scaffold. (A). Workflow of quantitative proteomic analysis performed in control versus CAV2-knockdown SCC15 cells. (B). Volcano plot displaying differentially expressed proteins following CAV2 silencing. Significantly upregulated (Red) and downregulated (Blue) proteins are highlighted (Fold change > 1.5, p < 0.05). (C-D). Co-immunoprecipitation (Co-IP) assays in SCC15(C) and SCC25(D) cells using antibodies against CAV2, PKR, and PACT, followed by immunoblotting with the indicated antibodies. Total cell lysates (Input) and IgG controls are shown. (E). Duo-link PLA was performed on human HNSCC tissue sections to detect protein-protein interactions between CAV2, PKR, and PACT. Red fluorescent dots represent specific protein interactions. Scale bar, 50μm. (F). Co-IP of endogenous PKR and PACT in control and CAV2-knockdown SCC15 cells. (G). Co-IP of CAV2 and PACT in control and PKR-knockdown SCC15 cells. (H). Co-IP of CAV2 and PKR in control and PACT-knockdown SCC15 cells. All Co-IPs were performed with the indicated antibodies and analyzed by western blotting.
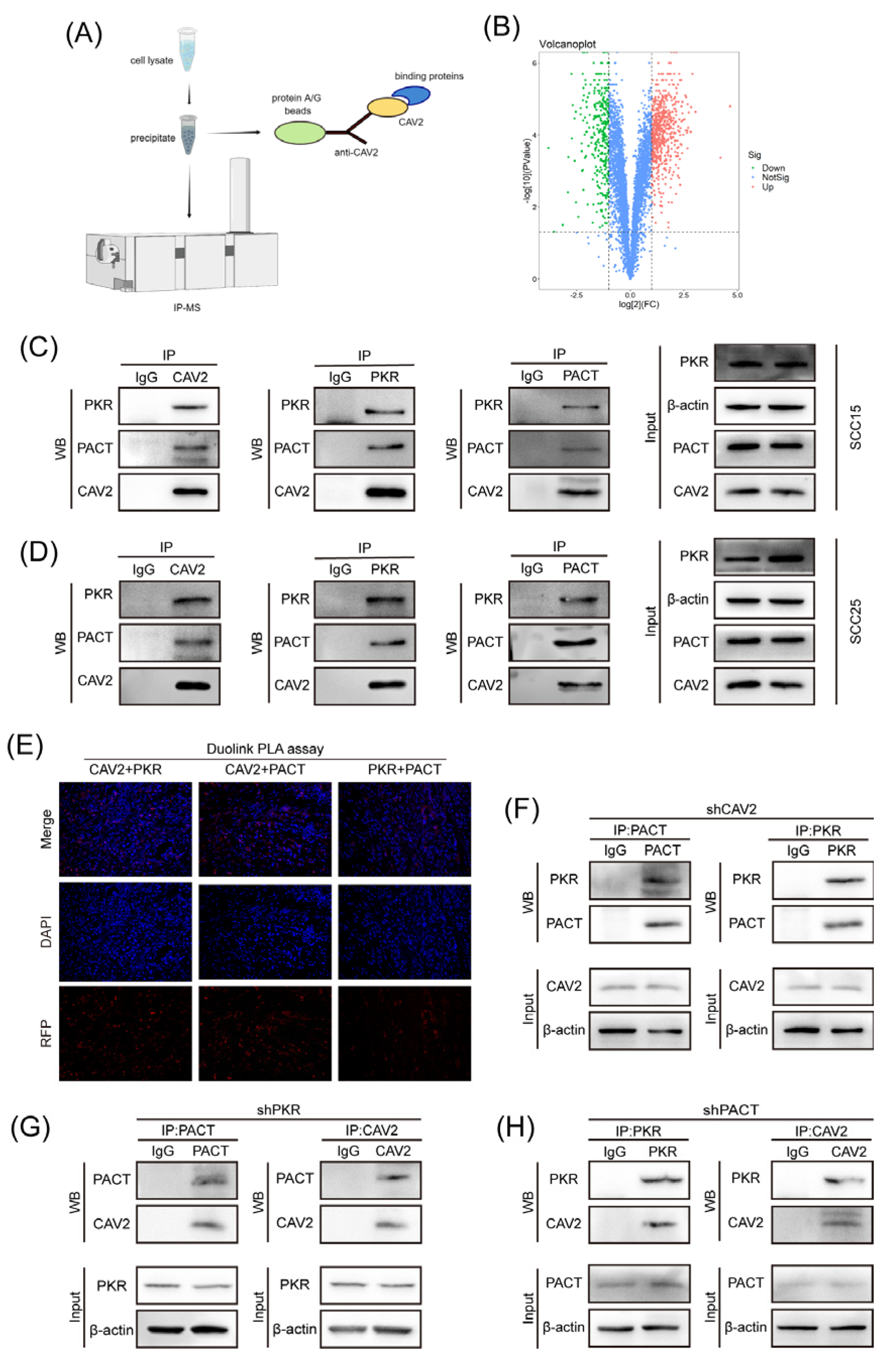
Figure 5.
CAV2 promotes the ubiquitin-mediated proteasomal degradation of PKR and PACT.(A). Western blot analysis of PKR and PACT protein levels in control and CAV2-knockdown SCC15 and SCC25 cells. GAPDH served as a loading control. (B). Immunoblot analysis of PKR and PACT expression in control and CAV2-knockdown cells treated with various protease inhibitors: MG-132 (proteasome inhibitor), chloroquine (lysosome inhibitor), calpeptin (calpain inhibitor), and Z-VAD-FMK (caspase inhibitor). Cells were treated with 10μM of each inhibitor for 24h. (C-F). (C-D) Western blot analysis and (E-F) quantification of PKR and PACT protein stability in control and CAV2-knockdown cells treated with 100μg/mL cycloheximide (CHX) for the indicated time periods. Protein bands were quantified using ImageJ software and normalized to GAPDH. Data are presented as mean ± SEM from three independent experiments. (G-H). Immunoblot analysis of polyubiquitinated PKR and PACT in control and CAV2-knockdown SCC15 and SCC25 cells. Cells were treated with 10μM MG-132 for 24 hours prior to immunoprecipitation with anti-PKR or anti-PACT antibodies, followed by western blotting with anti-ubiquitin antibody.
Figure 5.
CAV2 promotes the ubiquitin-mediated proteasomal degradation of PKR and PACT.(A). Western blot analysis of PKR and PACT protein levels in control and CAV2-knockdown SCC15 and SCC25 cells. GAPDH served as a loading control. (B). Immunoblot analysis of PKR and PACT expression in control and CAV2-knockdown cells treated with various protease inhibitors: MG-132 (proteasome inhibitor), chloroquine (lysosome inhibitor), calpeptin (calpain inhibitor), and Z-VAD-FMK (caspase inhibitor). Cells were treated with 10μM of each inhibitor for 24h. (C-F). (C-D) Western blot analysis and (E-F) quantification of PKR and PACT protein stability in control and CAV2-knockdown cells treated with 100μg/mL cycloheximide (CHX) for the indicated time periods. Protein bands were quantified using ImageJ software and normalized to GAPDH. Data are presented as mean ± SEM from three independent experiments. (G-H). Immunoblot analysis of polyubiquitinated PKR and PACT in control and CAV2-knockdown SCC15 and SCC25 cells. Cells were treated with 10μM MG-132 for 24 hours prior to immunoprecipitation with anti-PKR or anti-PACT antibodies, followed by western blotting with anti-ubiquitin antibody.
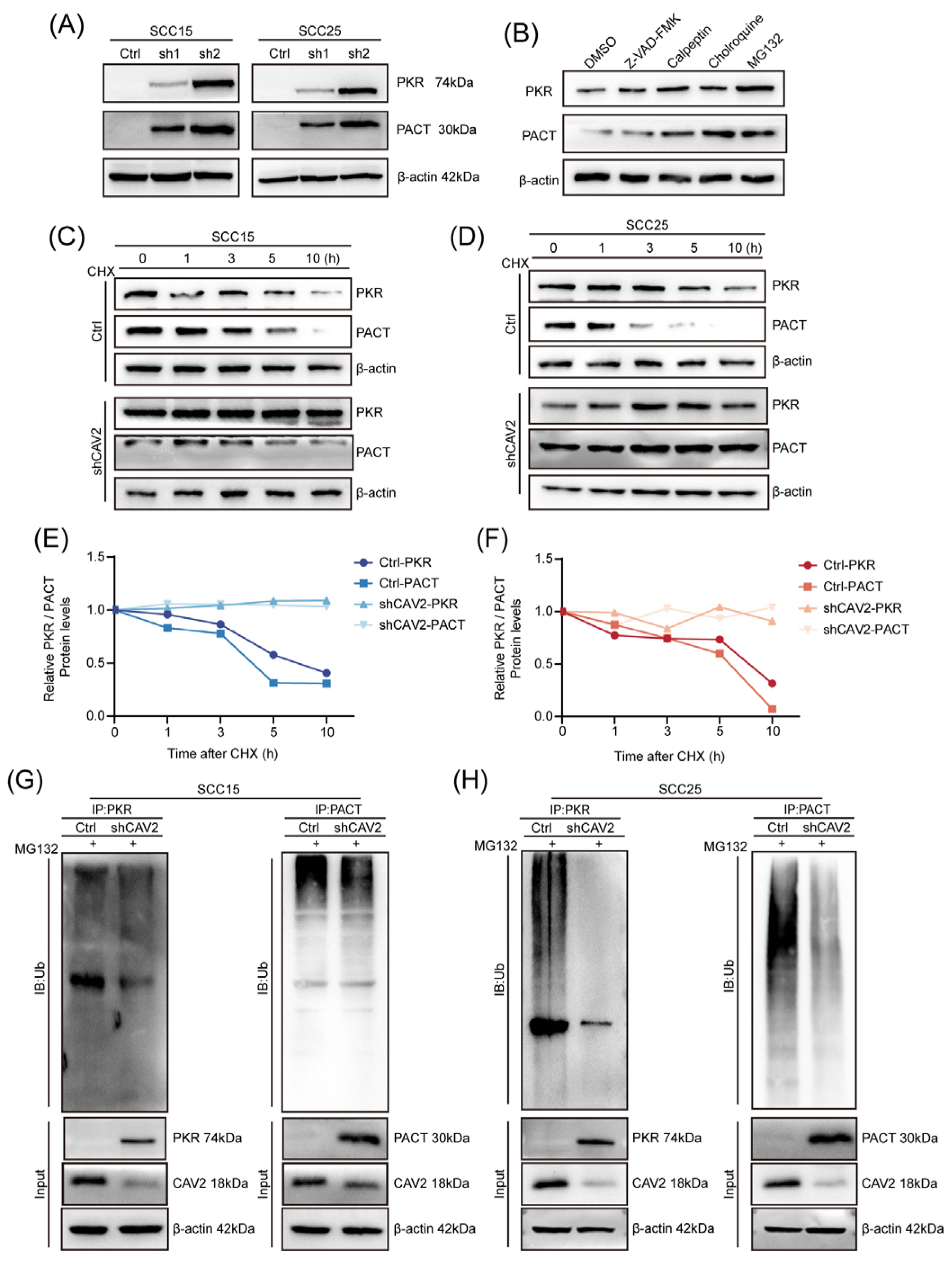
Figure 6.
PKR and PACT mediate the pro-oncogenic effects of CAV2 in HNSCC.(A). Western blot analysis confirming PKR protein levels in control and PKR-knockdown SCC15 and SCC25 cells. GAPDH served as a loading control. (B). (Left) Representative images of EdU staining in control and PKR-knockdown cells. Scale bar, 50μm. (Right) Quantification of EdU-positive cells. Data are presented as mean SEM from three independent experiments. (C). Western blot analysis confirming PACT protein levels in control and PACT-knockdown SCC15 and SCC25 cells. GAPDH served as a loading control. (D). (Left panel) Representative images of EdU staining in ±control and PACT-knockdown cells. Scale bar, 50μm (Right panel). Quantification of EdU-positive cells. Data are presented as mean ± SEM from three independent experiments. (E). EdU assay demonstrating that the anti-proliferative effect of CAV2 knockdown was partially reversed by co-depletion of PKR or PACT. (F). Quantification of (E) EdU-positive cells. Data are presented as mean ± SEM from three independent experiments. *p < 0.05, **p < 0.01, ***p < 0.001; ns, not significant.
Figure 6.
PKR and PACT mediate the pro-oncogenic effects of CAV2 in HNSCC.(A). Western blot analysis confirming PKR protein levels in control and PKR-knockdown SCC15 and SCC25 cells. GAPDH served as a loading control. (B). (Left) Representative images of EdU staining in control and PKR-knockdown cells. Scale bar, 50μm. (Right) Quantification of EdU-positive cells. Data are presented as mean SEM from three independent experiments. (C). Western blot analysis confirming PACT protein levels in control and PACT-knockdown SCC15 and SCC25 cells. GAPDH served as a loading control. (D). (Left panel) Representative images of EdU staining in ±control and PACT-knockdown cells. Scale bar, 50μm (Right panel). Quantification of EdU-positive cells. Data are presented as mean ± SEM from three independent experiments. (E). EdU assay demonstrating that the anti-proliferative effect of CAV2 knockdown was partially reversed by co-depletion of PKR or PACT. (F). Quantification of (E) EdU-positive cells. Data are presented as mean ± SEM from three independent experiments. *p < 0.05, **p < 0.01, ***p < 0.001; ns, not significant.
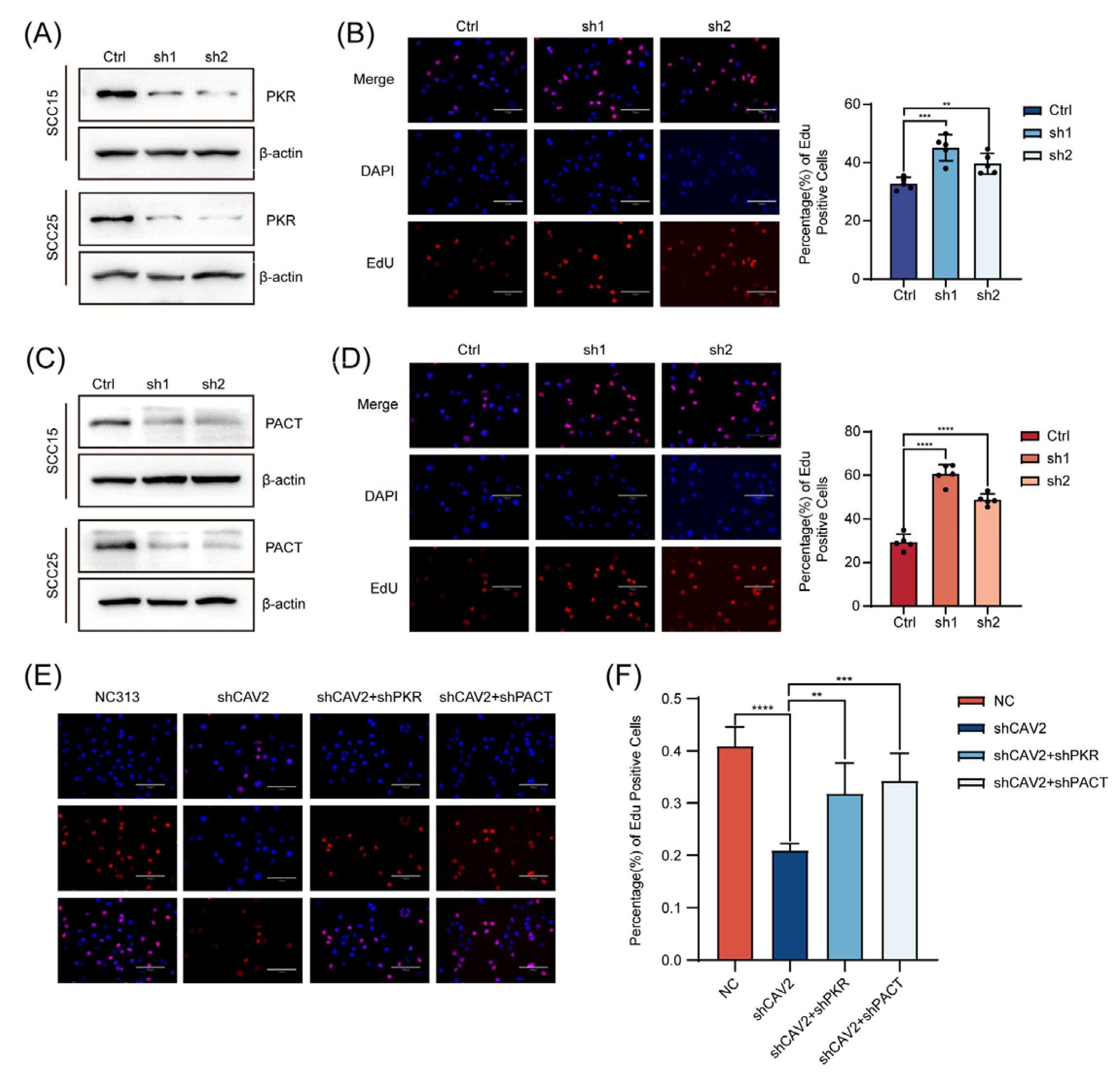
Figure 7.
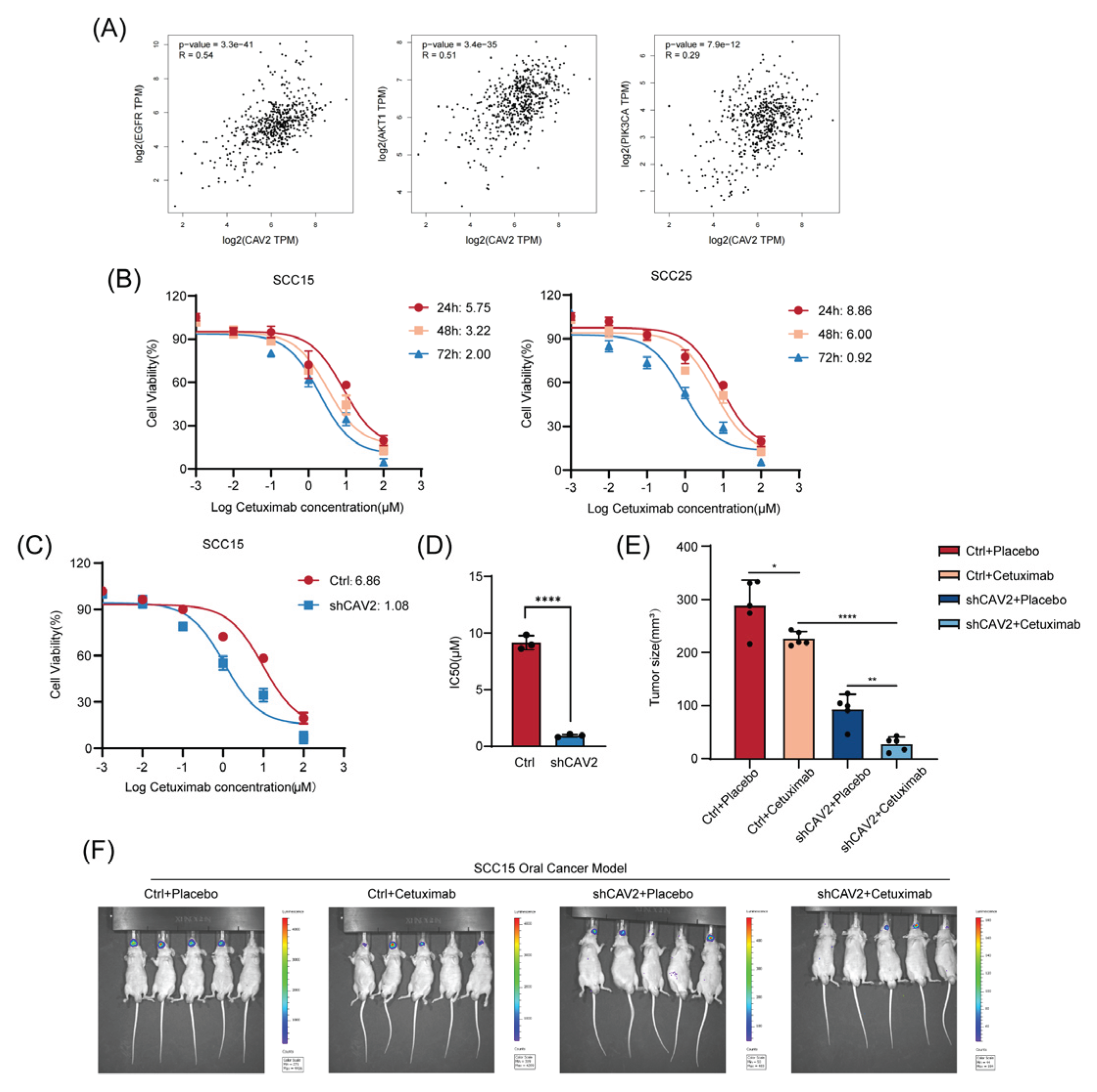
Targeting CAV2 Overcomes Cetuximab Resistance and Suppresses Tumor Growth.(A). Scatter plots showing the expression correlation of CAV2 with representative molecules of the EGFR signalling pathway from a public transcriptomic database GEPIA (Gene Expression Profiling Interactive Analysis). Pearson correlation coefficient (r) and p-value are indicated. (B). IC₅₀ values of cetuximab were determined at 24h, 48h, and 72h in (Left panel) SCC15 and (Right panel) SCC25 cell lines using CCK-8 assays. (C-D). Dose-response curves of Cetuximab after 48h treatment in SCC15-shCAV2 and SCC15-shCtrl cells (C). Bar graph comparing the IC₅₀ values between the two groups (D). Data are presented as mean ± SD (n = 3). ****p<0.0001 by unpaired Student’s t-test. (E-F). Representative bioluminescence images of mice bearing SCC15-Ctrl-luci or SCC15-shCAV2-luci tumors at the experimental endpoint after treatment with either PBS (control) or cetuximab (F). Mice were randomly assigned to treatment groups (n = 5 per group). Data are presented as mean ± SD from three independent experiments. Bar graph showed the final tumor volumes measured from the four experimental groups described in (E). Data are presented as mean ± SEM (n = 5). *p < 0.05, **p < 0.01, ****p<0.0001 by one-way ANOVA with Tukey’s post-hoc test.
Figure 7.
Targeting CAV2 Overcomes Cetuximab Resistance and Suppresses Tumor Growth.(A). Scatter plots showing the expression correlation of CAV2 with representative molecules of the EGFR signalling pathway from a public transcriptomic database GEPIA (Gene Expression Profiling Interactive Analysis). Pearson correlation coefficient (r) and p-value are indicated. (B). IC₅₀ values of cetuximab were determined at 24h, 48h, and 72h in (Left panel) SCC15 and (Right panel) SCC25 cell lines using CCK-8 assays. (C-D). Dose-response curves of Cetuximab after 48h treatment in SCC15-shCAV2 and SCC15-shCtrl cells (C). Bar graph comparing the IC₅₀ values between the two groups (D). Data are presented as mean ± SD (n = 3). ****p<0.0001 by unpaired Student’s t-test. (E-F). Representative bioluminescence images of mice bearing SCC15-Ctrl-luci or SCC15-shCAV2-luci tumors at the experimental endpoint after treatment with either PBS (control) or cetuximab (F). Mice were randomly assigned to treatment groups (n = 5 per group). Data are presented as mean ± SD from three independent experiments. Bar graph showed the final tumor volumes measured from the four experimental groups described in (E). Data are presented as mean ± SEM (n = 5). *p < 0.05, **p < 0.01, ****p<0.0001 by one-way ANOVA with Tukey’s post-hoc test.