Submitted:
01 March 2026
Posted:
03 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
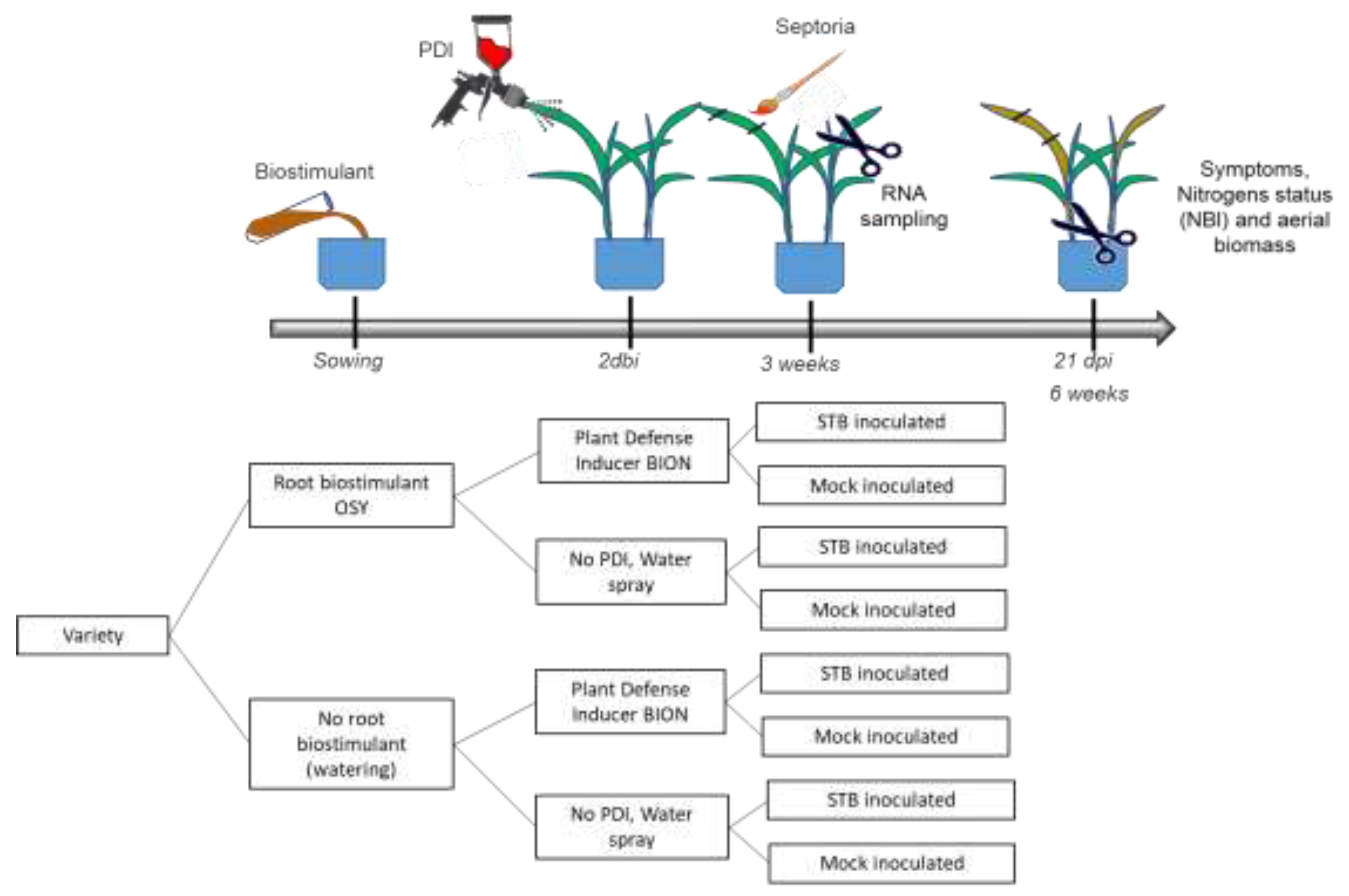
2.1. Experimental Design
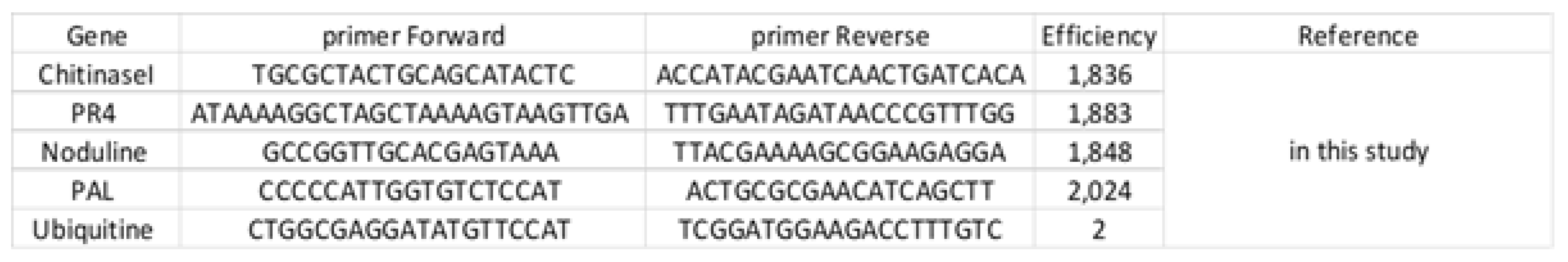
2.2. Molecular Experiment
2.3. Statistical Analysis
3. Results
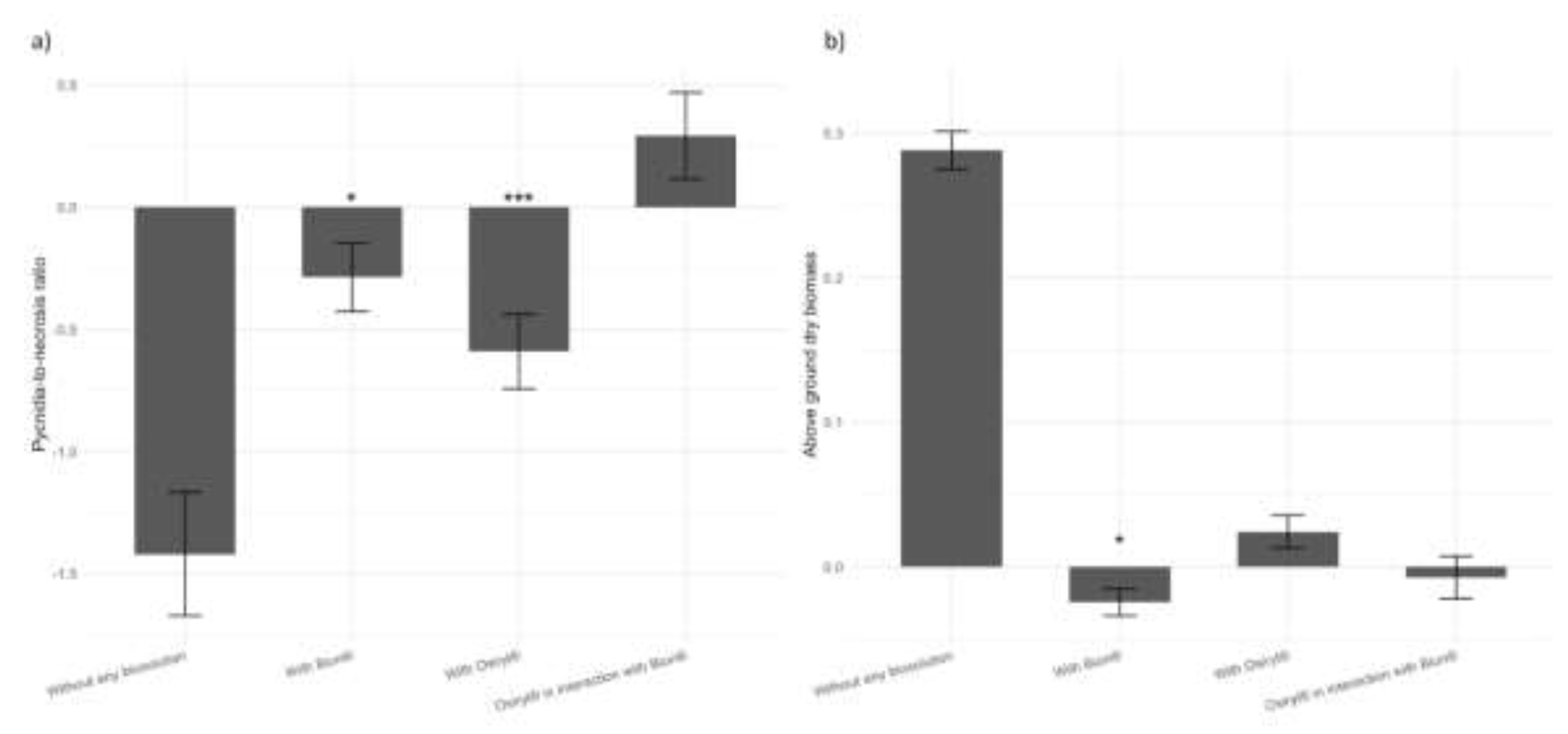
3.1. Salicylic Acid Analogue Enhances Resistance but Reduces Growth in Durum Wheat
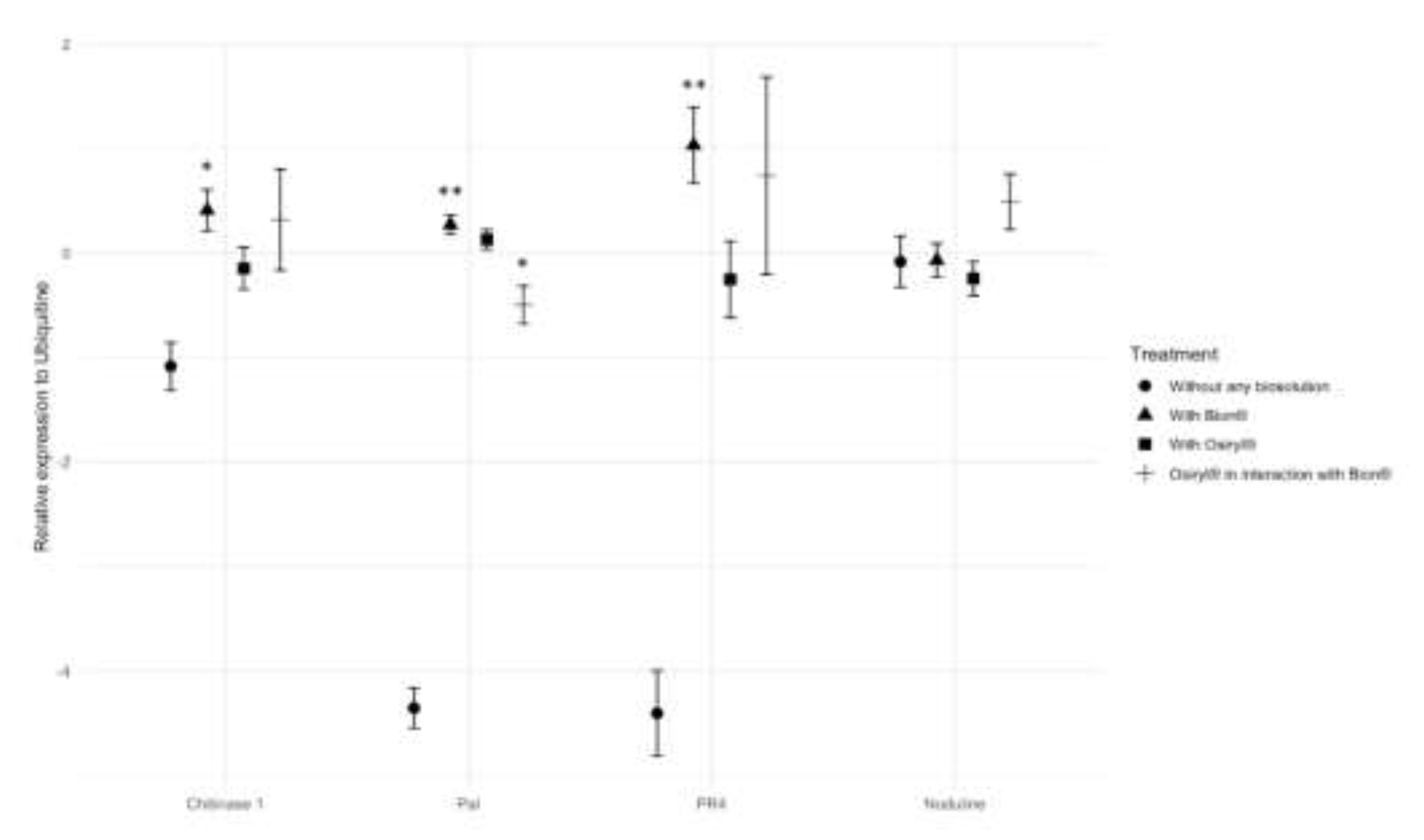
3.2. Four Defense-Related Genes Expression Induced by Salicylic Acid Analogue can be Modulated by the Biostimulant
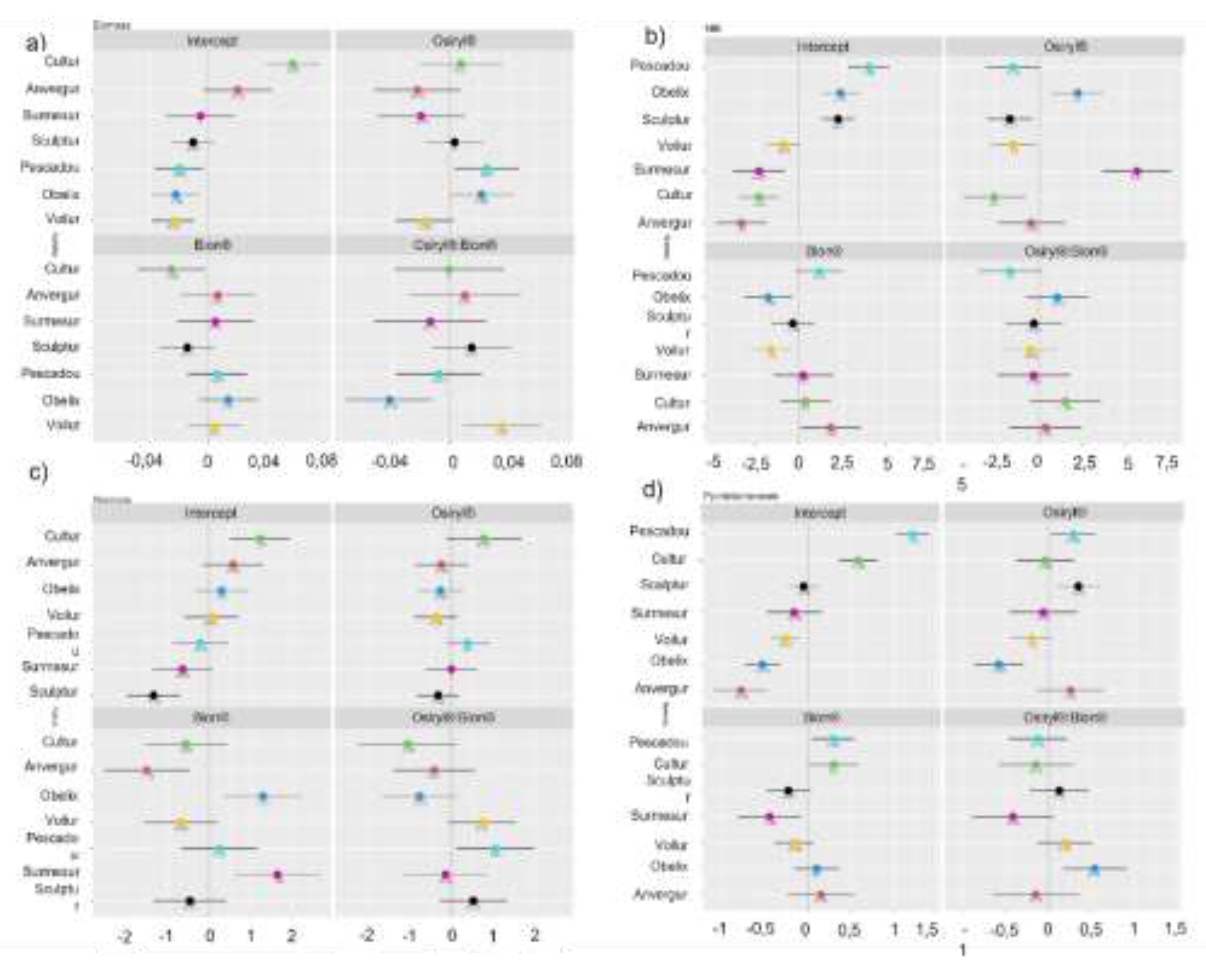
3.3. Variety Dictates Resistance, Defense and Growth Outcomes in Durum Wheat
3.4. The Interaction Between Factors Is Variable Among Varieties
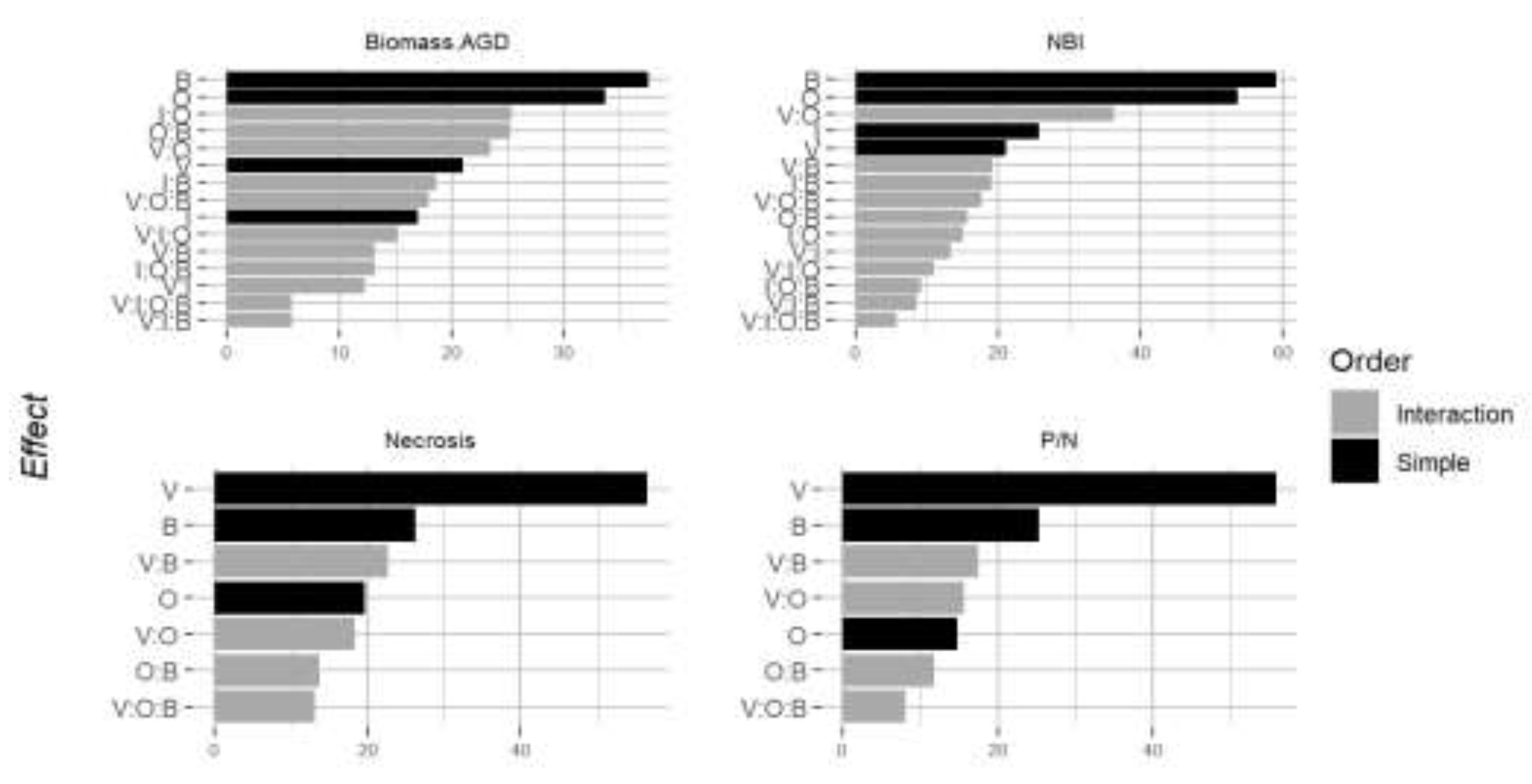
3.5. Beyond Main Effects: Interacting Factors Govern Phenotypic Variance
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| PDI | Plant Defense Inducer |
| STB | Septoria tritici blotch |
| NBI | Nitrogen balance Index |
Appendix A. Comprises Two Appendices Providing Additional Information on the Methodologies Used in This Article
Appendix A.1
Appendix A.2
| LD | Linear dichroism |
Appendix B
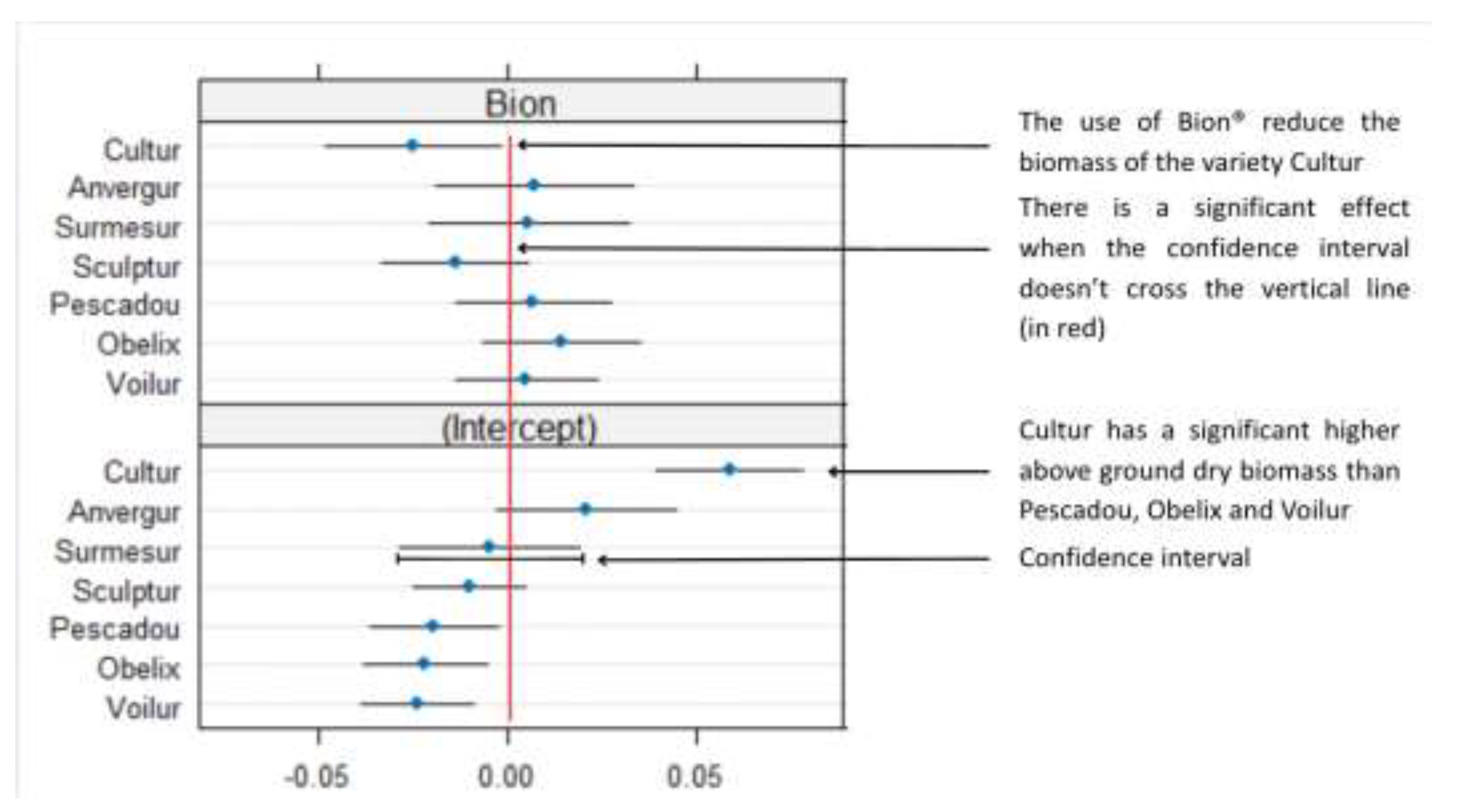
Appendix B.1
| Fixed effects | ||||
|---|---|---|---|---|
| Variables | Coefficients | Estimate | Std.error | p.value |
| Above ground dry biomass | Intercept | 0.288 | 0.0131 | 2.98E-07 |
| Osiryl® | 0.0244 | 0.0113 | 0.0724 | |
| Bion® | -0.0244 | 0.0094 | 0.0399 | |
| Osiryl®:Bion® | -0.0072 | 0.0147 | 6.38E-01 | |
| NBI | Intercept | 18.42 | 1.148 | 3.10E-06 |
| Osiryl® | 1.786 | 1.211 | 1.91E-01 | |
| Bion® | 0.145 | 0.692 | 8.40E-01 | |
| Osiryl®:Bion® | -1.56 | 0.765 | 8.61E-02 | |
| Necrosis | Intercept | 0.7758 | 0.3171 | 1.44E-02 |
| Osiryl® | 0.1375 | 0.2192 | 5.31E-01 | |
| Bion® | 0.0474 | 0.4312 | 9.12E-01 | |
| Osiryl®:Bion® | 0.2996 | 0.3636 | 4.10E-01 | |
|
Pycnidia-to- necrosis ratio |
Intercept | -1.42 | 0.252 | 1.80E-08 |
| Osiryl® | -0.591 | 0.152 | 0.000105 | |
| Bion® | -0.287 | 0.139 | 0.0387 | |
| Osiryl®:Bion® | 0.292 | 0.177 | 0.0985 | |
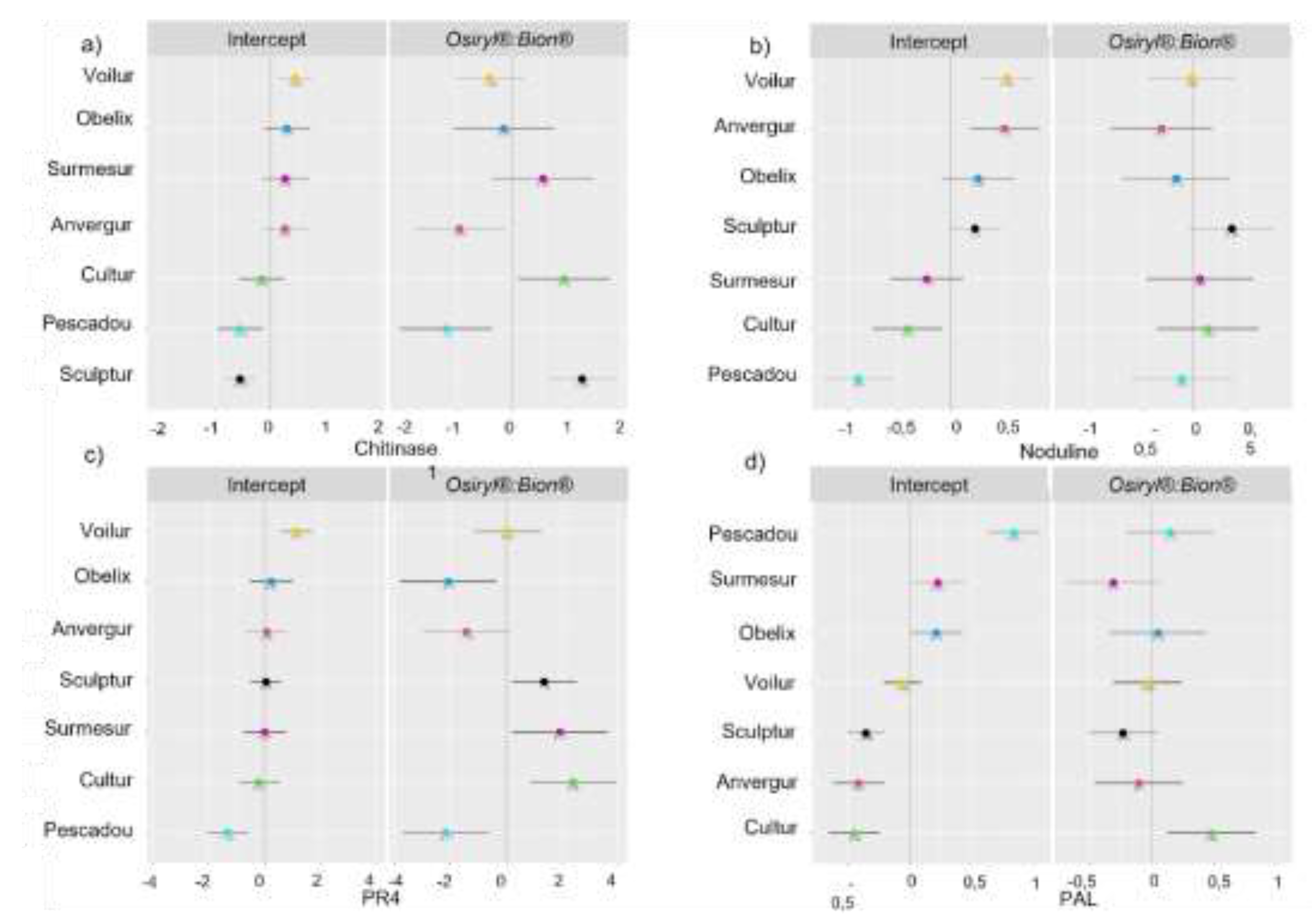
Appendix B.2
| Fixed effects | ||||
|---|---|---|---|---|
| Genes | Coefficients | Estimate | Std.error | p.value |
| Chitinase 1 | Intercept | -1.0847 | 0.228 | 0.00052 |
| Osiryl | -0.149 | 0.1991 | 0.4557 | |
| Bion | 0.408 | 0.1975 | 0.041 | |
| Osiryl:Bion | 0.3164 | 0.4833 | 0.52705 | |
| Noduline | Intercept | -0.08314 | 0.24333 | 0.741 |
| Osiryl | -0.24449 | 0.16348 | 0.1374 | |
| Bion | -0.07036 | 0.1622 | 0.6652 | |
| Osiryl:Bion | 0.49009 | 0.26633 | 0.0779 | |
| PR4 | Intercept | -4.4064 | 0.4068 | 2.16E-07 |
| Osiryl | -0.2528 | 0.367 | 0.49226 | |
| Bion | 1.0302 | 0.3641 | 0.00546 | |
| Osiryl:Bion | 0.7379 | 0.9469 | 0.45562 | |
| Pal | Intercept | -4.35642 | 0.19171 | 7.12E-08 |
| Osiryl | 0.13048 | 0.09312 | 0.1637 | |
| Bion | 0.26984 | 0.09238 | 0.00416 | |
| Osiryl:Bion | -0.49797 | 0.17991 | 0.0157 | |
Appendix B.3
>Appendix B4
References
- Li, J.; Van Gerrewey, T.; Geelen, D. A Meta-Analysis of Biostimulant Yield Effectiveness in Field Trials. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Rouphael, Y.; Colla, G. Synergistic Biostimulatory Action: Designing the Next Generation of Plant Biostimulants for Sustainable Agriculture. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef]
- Flors, V.; Kyndt, T.; Mauch-Mani, B.; Pozo, M.J.; Ryu, C.-M.; Ton, J. Enabling Sustainable Crop Protection with Induced Resistance in Plants. Front. Sci. 2024, 2. [Google Scholar] [CrossRef]
- Reglinski, T.; Havis, N.; Rees, H.J.; de Jong, H. The Practical Role of Induced Resistance for Crop Protection. Phytopathology® 2023, 113, 719–731. [Google Scholar] [CrossRef]
- Garcia-Brugger, A.; Lamotte, O.; Vandelle, E.; Bourque, S.; Lecourieux, D.; Poinssot, B.; Wendehenne, D.; Pugin, A. Early Signaling Events Induced by Elicitors of Plant Defenses. Mol. Plant-Microbe Interactions® 2006, 19, 711–724. [Google Scholar] [CrossRef]
- Llorens, E.; Agustí-Brisach, C. Biocontrol of Plant Diseases by Means of Antagonist Microorganisms, Biostimulants and Induced Resistance as Alternatives to Chemicals. Plants 2022, 11, 3521. [Google Scholar] [CrossRef] [PubMed]
- Ciriello, M.; Fusco, G.M.; Woodrow, P.; Carillo, P.; Rouphael, Y. Unravelling the Nexus of Plant Response to Non-Microbial Biostimulants under Stress Conditions. Plant Stress 2024, 11, 100421. [Google Scholar] [CrossRef]
- Raja, B.; Vidya, R. Application of Seaweed Extracts to Mitigate Biotic and Abiotic Stresses in Plants. Physiol. Mol. Biol. Plants 2023. [Google Scholar] [CrossRef]
- Song, W.; Shao, H.; Zheng, A.; Zhao, L.; Xu, Y.; Song, W.; Shao, H.; Zheng, A.; Zhao, L.; Xu, Y. Advances in Roles of Salicylic Acid in Plant Tolerance Responses to Biotic and Abiotic Stresses. Plants 2023, 12. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Ning, C.; Pan, T.; Cai, K.; Wang, L.; Ning, C.; Pan, T.; Cai, K. Role of Silica Nanoparticles in Abiotic and Biotic Stress Tolerance in Plants: A Review. Int. J. Mol. Sci. 2022, 23. [Google Scholar] [CrossRef] [PubMed]
- Yakhin, O.I.; Lubyanov, A.A.; Yakhin, I.A.; Brown, P.H. Biostimulants in Plant Science: A Global Perspective. Front. Plant Sci. 2017, 7. [Google Scholar] [CrossRef]
- Lee Díaz, A.S.; Macheda, D.; Saha, H.; Ploll, U.; Orine, D.; Biere, A. Tackling the Context-Dependency of Microbial-Induced Resistance. Agronomy 2021, 11, 1293. [Google Scholar] [CrossRef]
- Mendes, R.; de Barros, I.; D’Andréa, P.A.; D’Andréa-Kühl, M.S.C.; Rodrigues, G.S. A Multi-Attribute Approach to Evaluating the Impact of Biostimulants on Crop Performance. Front. Plant Sci. 2023, 14. [Google Scholar] [CrossRef]
- Stiling, P.; Cornelissen, T. What Makes a Successful Biocontrol Agent? A Meta-Analysis of Biological Control Agent Performance. Biol. Control 2005, 34, 236–246. [Google Scholar] [CrossRef]
- Pagani, A.P.S.; Dianese, A.C.; Café-Filho, A.C. Management of Wheat Blast with Synthetic Fungicides, Partial Resistance and Silicate and Phosphite Minerals. Phytoparasitica 2014, 42, 609–617. [Google Scholar] [CrossRef]
- Samain, E.; Aussenac, T.; Selim, S. The Effect of Plant Genotype, Growth Stage, and Mycosphaerella Graminicola Strains on the Efficiency and Durability of Wheat-Induced Resistance by Paenibacillus Sp. Strain B2. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Ji, H.; Wang, J.; Chen, F.; Fan, N.; Wang, X.; Xiao, Z.; Wang, Z. Meta-Analysis of Chitosan-Mediated Effects on Plant Defense against Oxidative Stress. Sci. Total Environ. 2022, 851, 158212. [Google Scholar] [CrossRef]
- Val-Torregrosa, B.; Bundó, M.; San Segundo, B. Crosstalk between Nutrient Signalling Pathways and Immune Responses in Rice. Agriculture 2021, 11, 747. [Google Scholar] [CrossRef]
- Choudhary, A.; Senthil-Kumar, M. Drought: A Context-Dependent Damper and Aggravator of Plant Diseases. Plant Cell Environ. 2024, 47, 2109–2126. [Google Scholar] [CrossRef]
- Dallacorte, L.V.; Bosse, M.A.; Capelin, D.; Paladini, M.V.; Cattani, F.; Remor, M.B.; de Lima, J.D.; Perboni, A.T.; Marchese, J.A. Economic versus Technical Efficiency in Using ASM Combined with Fungicides to Elicit Wheat Varieties with Different Disease Susceptibilities. Heliyon 2023, 9. [Google Scholar] [CrossRef]
- Jacquens, L.; Trouvelot, S.; Lemaitre-Guillier, C.; Krzyzaniak, Y.; Clément, G.; Citerne, S.; Mouille, G.; Moreau, E.; Héloir, M.-C.; Adrian, M. Biostimulation Can Prime Elicitor Induced Resistance of Grapevine Leaves to Downy Mildew. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef] [PubMed]
- Petrova, M.; Miladinova-Georgieva, K.; Geneva, M.; Petrova, M.; Miladinova-Georgieva, K.; Geneva, M. Influence of Abiotic and Biotic Elicitors on Organogenesis, Biomass Accumulation, and Production of Key Secondary Metabolites in Asteraceae Plants. Int. J. Mol. Sci. 2024, 25. [Google Scholar] [CrossRef] [PubMed]
- Azmat, A.; Yasmin, H.; Hassan, M.N.; Nosheen, A.; Naz, R.; Sajjad, M.; Ilyas, N.; Akhtar, M.N. Co-Application of Bio-Fertilizer and Salicylic Acid Improves Growth, Photosynthetic Pigments and Stress Tolerance in Wheat under Drought Stress. PeerJ 2020, 8, e9960. [Google Scholar] [CrossRef]
- Iula, G.; Miras-Moreno, B.; Rouphael, Y.; Lucini, L.; Trevisan, M. The Complex Metabolomics Crosstalk Triggered by Four Molecular Elicitors in Tomato. Plants 2022, 11, 678. [Google Scholar] [CrossRef]
- Jacquens, L.; Trouvelot, S.; Lemaitre-Guillier, C.; Krzyzaniak, Y.; Clément, G.; Citerne, S.; Mouille, G.; Moreau, E.; Héloir, M.-C.; Adrian, M. Biostimulation Can Prime Elicitor Induced Resistance of Grapevine Leaves to Downy Mildew. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Brzozowski, L.J.; Smith, K.P.; Ohm, J.-B.; Clark, A.J.; Palakurthi, R.; Sorrells, M.E.; Gore, M.A.; Jannink, J.-L. Response to Plant Defense Elicitation Is Genotype- and Context-Dependent in Diverse Oat (Avena Sativa L.) Germplasm. J. Exp. Bot. 2025, eraf313. [Google Scholar] [CrossRef]
- van Butselaar, T.; Ackerveken, G.V. den Salicylic Acid Steers the Growth–Immunity Tradeoff. Trends Plant Sci. 2020, 25, 566–576. [Google Scholar] [CrossRef] [PubMed]
- Karasov, T.L.; Chae, E.; Herman, J.J.; Bergelson, J. Mechanisms to Mitigate the Trade-Off between Growth and Defense. Plant Cell 2017, 29, 666–680. [Google Scholar] [CrossRef]
- Kliebenstein, D.J. False Idolatry of the Mythical Growth versus Immunity Tradeoff in Molecular Systems Plant Pathology. Physiol. Mol. Plant Pathol. 2016, 95, 55–59. [Google Scholar] [CrossRef]
- Proietti, S.; Caarls, L.; Coolen, S.; Van Pelt, J.A.; Van Wees, S.C.M.; Pieterse, C.M.J. Genome-Wide Association Study Reveals Novel Players in Defense Hormone Crosstalk in Arabidopsis. Plant Cell Environ. 2018, 41, 2342–2356. [Google Scholar] [CrossRef]
- Samanta, S.; Roychoudhury, A. Molecular Crosstalk of Jasmonate with Major Phytohormones and Plant Growth Regulators During Diverse Stress Responses. J. Plant Growth Regul. 2025, 44, 62–88. [Google Scholar] [CrossRef]
- Urban, L.; Lauri, F.; Ben Hdech, D.; Aarrouf, J. Prospects for Increasing the Efficacy of Plant Resistance Inducers Stimulating Salicylic Acid. Agronomy 2022, 12, 3151. [Google Scholar] [CrossRef]
- Chan, C. Progress in Salicylic Acid-Dependent Signaling for Growth–Defense Trade-Off. Cells 2022, 11, 2985. [Google Scholar] [CrossRef]
- Kitagawa, M. SA-Related Defense Signaling Supports the Trade-Off between Shoot Growth and Low-Ca Tolerance. Plant Cell Physiol. 2023, 64, 1576–1578. [Google Scholar] [CrossRef]
- Pokotylo, I.; Hodges, M.; Kravets, V.; Ruelland, E. A Ménage à Trois: Salicylic Acid, Growth Inhibition, and Immunity. Trends Plant Sci. 2022, 27, 460–471. [Google Scholar] [CrossRef]
- Matysiak, K.; Siatkowski, I.; Kierzek, R.; Kowalska, J.; Krawczyk, R. Effect of Foliar Applied Acetylsalicilic Acid on Wheat (Triticum Aestivum L.) under Field Conditions. Agronomy 2020, 10, 1918. [Google Scholar] [CrossRef]
- Perelló, A. e.; Dal Bello, G. m. Suppression of Tan Spot and Plant Growth Promotion of Wheat by Synthetic and Biological Inducers under Field Conditions. Ann. Appl. Biol. 2011, 158, 267–274. [Google Scholar] [CrossRef]
- Sissons, M. Durum Wheat Products—Recent Advances. Foods 2022, 11, 3660. [Google Scholar] [CrossRef] [PubMed]
- Fones, H.; Gurr, S. The Impact of Septoria Tritici Blotch Disease on Wheat: An EU Perspective. Fungal Genet. Biol. 2015, 79, 3–7. [Google Scholar] [CrossRef]
- Porras, R.; Miguel-Rojas, C.; Lorite, I.J.; Pérez-de-Luque, A.; Sillero, J.C. Characterization of Durum Wheat Resistance against Septoria Tritici Blotch under Climate Change Conditions of Increasing Temperature and CO2 Concentration. Agronomy 2023, 13, 2638. [Google Scholar] [CrossRef]
- Hekımhan, H.; Boyraz, N. The effects of acibenzolar-S-methyl application on root and foot rot disease (F. culmorum) on the some bread wheat varieties. TABAD Tarım Bilim. Araștırma Derg. 2013, 6, 56–61. [Google Scholar]
- Hafez, Y.; Soliman, N.K.; Mostafa, S.; Imbabi, I.A.; Abd-Elaziz, A.S. Induced Resistance against Puccinia Triticina, the Causal Agent of Wheat Leaf Rust by Chemical Inducers. Egypt. J. Biol. Pest Control 2014, 24, 173–181. [Google Scholar]
- Bellameche, F.; Pedrazzini, C.; Mauch-Mani, B.; Mascher, F. Efficiency of Biological and Chemical Inducers for Controlling Septoria Tritici Leaf Blotch (STB) on Wheat (Triticum Aestivum L.). Eur. J. Plant Pathol. 2020, 158, 99–109. [Google Scholar] [CrossRef]
- Mejri, S.; Siah, A.; Abuhaie, C.-M.; Halama, P.; Magnin-Robert, M.; Randoux, B.; Reignault, P.; Rigo, B.; Ghinet, A. New Salicylic Acid and Pyroglutamic Acid Conjugated Derivatives Confer Protection to Bread Wheat against Zymoseptoria Tritici. J. Sci. Food Agric. 2019, 99, 1780–1786. [Google Scholar] [CrossRef] [PubMed]
- Mathieu, L.; Ducasse, A.; Ballini, E.; Morel, J.-B. Plant–Plant Interactions in Wheat Mixtures Modulate Mean and Variance of Susceptibility to Septoria Tritici Blotch. J. Exp. Bot. 2025, 76, 6714–6722. [Google Scholar] [CrossRef] [PubMed]
- Ballini, E.; Tavaud, M.; Ducasse, A.; Sanchez, D.; Paux, E.; Kitt, J.; Charmet, G.; Audigeos, D.; Roumet, P.; David, J.; et al. Genome Wide Association Mapping for Resistance to Multiple Fungal Pathogens in a Panel Issued from a Broad Composite Cross-Population of Tetraploid Wheat Triticum Turgidum. Euphytica 2020, 216, 92. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, J.; Xiao, Y.; Shi, X.; Zeng, Y. Diversity Analysis of Chlorophyll, Flavonoid, Anthocyanin, and Nitrogen Balance Index of Tea Based on Dualex. Phyton-Int. J. Exp. Bot. 2021, 90, 1549–1558. [Google Scholar] [CrossRef]
- Duba, A.; Goriewa-Duba, K.; Wachowska, U. A Review of the Interactions between Wheat and Wheat Pathogens: Zymoseptoria Tritici, Fusarium Spp. and Parastagonospora Nodorum. Int. J. Mol. Sci. 2018, 19, 1138. [Google Scholar] [CrossRef]
- Delteil, A.; Blein, M.; Faivre-Rampant, O.; Guellim, A.; Estevan, J.; Hirsch, J.; Bevitori, R.; Michel, C.; Morel, J.-B. Building a Mutant Resource for the Study of Disease Resistance in Rice Reveals the Pivotal Role of Several Genes Involved in Defence. Mol. Plant Pathol. 2012, 13, 72–82. [Google Scholar] [CrossRef]
- Vergne, E.; Ballini, E.; Marques, S.; Sidi Mammar, B.; Droc, G.; Gaillard, S.; Bourot, S.; DeRose, R.; Tharreau, D.; Nottéghem, J.-L.; et al. Early and Specific Gene Expression Triggered by Rice Resistance Gene Pi33 in Response to Infection by ACE1 Avirulent Blast Fungus. New Phytol. 2007, 174, 159–171. [Google Scholar] [CrossRef]
- Bates, D.; Mächler, M.; Bolker, B.; Walker, S. Fitting Linear Mixed-Effects Models Using Lme4. J. Stat. Softw. 2015, 67, 1–48. [Google Scholar] [CrossRef]
- Leisner, C.P.; Potnis, N.; Sanz-Saez, A. Crosstalk and Trade-offs: Plant Responses to Climate Change-associated Abiotic and Biotic Stresses. Plant Cell Environ. 2023, 46, 2946–2963. [Google Scholar] [CrossRef]
- Tripathi, R.; Tewari, R.; Singh, K.P.; Keswani, C.; Minkina, T.; Srivastava, A.K.; De Corato, U.; Sansinenea, E. Plant Mineral Nutrition and Disease Resistance: A Significant Linkage for Sustainable Crop Protection. Front. Plant Sci. 2022, 13. [Google Scholar] [CrossRef]
- Figueroa-Macías, J.P.; García, Y.C.; Núñez, M.; Díaz, K.; Olea, A.F.; Espinoza, L. Plant Growth-Defense Trade-Offs: Molecular Processes Leading to Physiological Changes. Int. J. Mol. Sci. 2021, 22. [Google Scholar] [CrossRef]
- He, Z.; Webster, S.; He, S.Y. Growth–Defense Trade-Offs in Plants. Curr. Biol. 2022, 32, R634–R639. [Google Scholar] [CrossRef]
- Bénéjam, J.; Ravon, E.; Gaucher, M.; Brisset, M.-N.; Durel, C.-E.; Perchepied, L. Acibenzolar-S-Methyl and Resistance Quantitative Trait Loci Complement Each Other to Control Apple Scab and Fire Blight. Plant Dis. 2021, 105, 1702–1710. [Google Scholar] [CrossRef] [PubMed]
- Matros, A.; Schikora, A.; Ordon, F.; Wehner, G. QTL for Induced Resistance against Leaf Rust in Barley. Front. Plant Sci. 2023, 13. [Google Scholar] [CrossRef] [PubMed]
- Soleimani, B.; Lehnert, H.; Schikora, A.; Stahl, A.; Matros, A.; Wehner, G. Bacterial N-Acyl Homoserine Lactone Priming Enhances Leaf-Rust Resistance in Winter Wheat and Some Genomic Regions Are Associated with Priming Efficiency. Microorganisms 2024, 12, 1936. [Google Scholar] [CrossRef]
- Du, B.; Haensch, R.; Alfarraj, S.; Rennenberg, H. Strategies of Plants to Overcome Abiotic and Biotic Stresses. Biol. Rev. Camb. Philos. Soc. 2024, 99, 1524–1536. [Google Scholar] [CrossRef]
- Walters, D.R.; Ratsep, J.; Havis, N.D. Controlling Crop Diseases Using Induced Resistance: Challenges for the Future. J. Exp. Bot. 2013, 64, 1263–1280. [Google Scholar] [CrossRef] [PubMed]
- Lundberg, D.S.; Kersten, S.; Boz, E.M.; Ayutthaya, P.P.N.; Zhu, W.; Poersch, K.; Yuan, W.; Swartz, S.; Müller, D.; Bezrukov, I.; et al. A Major Trade-off between Growth and Defense in Arabidopsis Thaliana Can Vanish in Field Conditions. PLOS Biol. 2025, 23, e3003237. [Google Scholar] [CrossRef]
- Walters, D.R.; Havis, N.D.; Sablou, C.; Walsh, D.J. Possible Trade-off Associated with the Use of a Combination of Resistance Elicitors. Physiol. Mol. Plant Pathol. 2011, 75, 188–192. [Google Scholar] [CrossRef]
- Suarez-Fernandez, M.; De Francesco, A. Embracing Biological Control of Septoria Tritici Blotch for Sustainable Wheat Protection. J. Phytopathol. 2024, 172, e13395. [Google Scholar] [CrossRef]
- Napier, J.D.; Heckman, R.W.; Juenger, T.E. Gene-by-Environment Interactions in Plants: Molecular Mechanisms, Environmental Drivers, and Adaptive Plasticity. Plant Cell 2023, 35, 109–124. [Google Scholar] [CrossRef] [PubMed]
- Bolker, B. Linear and Generalized Linear Mixed Models; 2015. [Google Scholar] [CrossRef]
- Bolker, B. GLMM. Available online: https://bbolker.github.io/mixedmodels-misc/glmmFAQ.html.
- brooks, mollie E.; Kristensen, K.; Van Benthem, K.J.; Magnusson, A.; Berg, C.W.; Nielsen, A.; Skaug, H.J.; Machler, M.; Bolker, B. glmmTMB Balances Speed and Flexibility Among Packages for Zero-Inflate. R J 2017. [Google Scholar] [CrossRef]
| Above ground dry biomass | NBI | Necrosis | pycnidia/necrosis | Chitinase 1 | Noduline | PR4 | Pal |
| 0,2879 | 18,413 | 0,6848 | 0,2417 | -1,0847 | -0,0831 | -4,4064 | -4,3564 |
| Response variable | Type of data | Tested effects | Statistical model | Statistical law |
|---|---|---|---|---|
| Above ground dry biomass | Quantitative continue | O, B, O:B, Exp, V, O:B, B:V, O:B:V | Linear mixed with random effects | Normal (0, σ2) |
| NBI | Quantitative continue | O, B, O:B, Exp, V, O:B, B:V, O:B:V | Linear mixed with random effects | Normal (0, σ2) |
| Necrosis | Surface (%) | O, B, O:B, Exp, V, O:B, B:V, O:B:V | Generalized linear with random effects | Beta-binomial BB(n, α, β) |
| Pycnidia-to-necrosis ratio | Counting | O, B, O:B, Exp, V, O:B, B:V, O:B:V | Generalized linear with random effects | Negative Binomial BN(n,p) |
| Genes | Quantitative continue | O, B, O:B, Exp, V, O:B:V | Linear mixed with random effects | Normal (0, σ2) |
| Variety | Osiryl® : Variety | Bion® : Variety | Osiryl®:Bion® : Variety | Residual | |
|---|---|---|---|---|---|
| Above ground dry biomass | 0.0318 (***) | 0.0239 (**) | 0.0181 | 0.0299 (**) | 0.0894 |
| NBI | 2.939 (***) | 3.033 (***) | 1.54 (**) | 1.436 | 4.661 |
| Chitinase 1 | 0.4674 (***) | 1.0264 (***) | 0.8319 | ||
| Noduline | 0.5627 (***) | 0.3332 | 0.6831 | ||
| PR4 | 0.8146 (***) | 2.0719 (***) | 1.5333 | ||
| Pal | 0.4748 (***) | 0.3164 (*) | 0.389 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.





