1. Introduction
Antimicrobial resistance (AMR) is one of the main challenges for public health because it compromises the effectiveness of antibiotic treatments [
1], particularly due to the increasing prevalence of extended-spectrum β-lactamase (ESBL)-producing Enterobacteriaceae, which inactivate β-lactam antibiotics such as penicillins and cephalosporins. These β-lactamase enzymes are encoded by acquired genes or variants such as:
bla_CTX-M, bla_TEM and
bla_SHV, which confer resistance to β-lactam antibiotics [
2], thereby limiting therapeutic options and increasing morbidity and mortality associated with infections. The World Health Organization (WHO) classifies ESBL-producing bacteria as priority pathogens for the development of new treatments, and reports indicate that 60% of nosocomial infections in Latin America are caused by β-lactam-resistant Enterobacteriaceae [
3].
Escherichia coli is an opportunistic bacterium associated with cases of cystitis, sepsis, and pneumonia due to the acquisition of resistance genes through mobile genetic elements (plasmids) [
4,
5].
Klebsiella pneumoniae also acquires ESBL production and is a nosocomial pathogen associated with approximately 50% of bacteremia cases, as well as a high capacity for resistance to multiple antibiotics [
6].
Medicinal plants have been valued for their therapeutic properties, playing a fundamental role in traditional medicine. This practice has been integrated into modern medicine through the isolation and characterization of phytochemicals or bioactive compounds such as terpenoids, phenolic compounds, glycosides, and alkaloids, which exhibit anti-inflammatory, antimicrobial, and antifungal effects [
7,
8].
Among the methods used for the extraction of bioactive compounds are non-conventional techniques such as ultrasound-assisted extraction and supercritical fluid extraction, as well as conventional methods including maceration, percolation, decoction, solvent extraction, and steam distillation [
9], the latter is particularly useful because it yields essential oils containing bioactive components with therapeutic potential [
10].
The essential oil of
Origanum vulgare (oregano), belonging to the Lamiaceae family, is rich in phenolic monoterpenes and phenylpropanoids, with carvacrol and thymol being the main compounds responsible for its antimicrobial activity [
11]. Studies by Nurzyńska demonstrate that essential oils exert inhibitory effects on the growth of both Gram-positive and Gram-negative bacteria, with particular emphasis on
Escherichia coli and
Klebsiella pneumoniae, due to the presence of metabolites in oregano essential oil that act through multiple mechanisms, including destabilization of the outer membrane, suppression of resistance gene expression, and inhibition of protein synthesis. In light of this emergency, oregano essential oil has been considered a natural adjuvant [
12].
Carvacrol has demonstrated synergistic effects with β-lactam antibiotics, reversing resistance to cephalosporins in resistant
E. coli strains [
13]. Si et al. reported minimum inhibitory concentrations (MIC) of
Origanum vulgare essential oil of 0.5 µg/mL and 0.8 µg/mL against ESBL-producing
E. coli strains, in addition to a synergistic effect with fluoroquinolones or doxycycline, which reduced the standard dose of these antibiotics by up to eightfold [
14].
2. Materials and Methods
2.1. Bacterial Strains
Four bacterial strains were used, provided by the National Institute of Public Health Research (INSPI) due to their epidemiological relevance regarding resistance mechanisms: E. coli ATCC 25922 and K. pneumoniae ATCC 700603, employed as quality control strains, and two ESBL-producing bacterial isolates: E. coli (ID 323) and K. pneumoniae (ID 132). The strains were preserved at −20 °C in cryovials with glycerol and reactivated in BHI. For each assay, fresh cultures (≤ 24 h) were used to ensure viability.
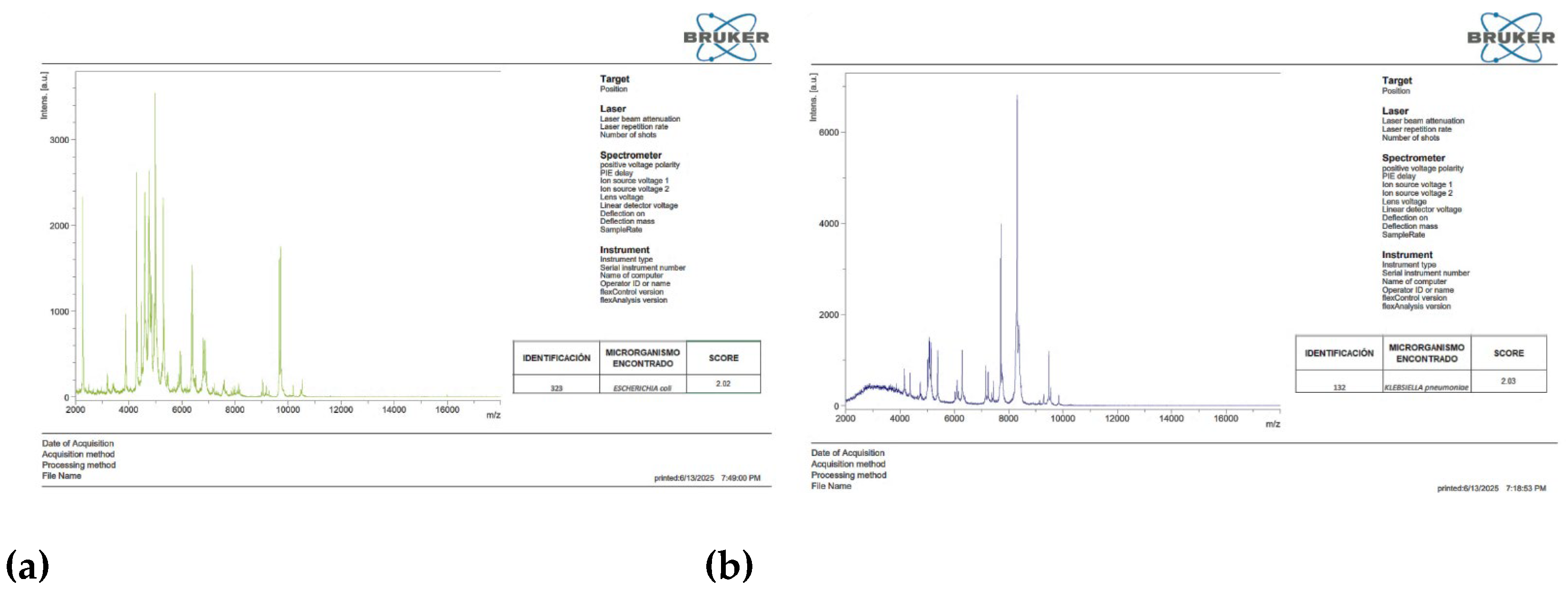
2.2. Identification by MALDI-TOF MS
Taxonomic identification of the study strains was performed by MALDI-TOF mass spectrometry at the MEDICULT laboratory (Cuenca, Ecuador). Fresh colonies (18 h) were taken from blood agar plates and placed onto a polished steel target plate for identification using the Microflex LT system (Bruker Daltonics, Bremen, Germany). The generated protein spectrum was compared with a reference database, providing score values, where scores ≥ 2.0 indicated reliable identification at the genus and species levels.
2.3. Susceptibility Analysis for Phenotypic Detection
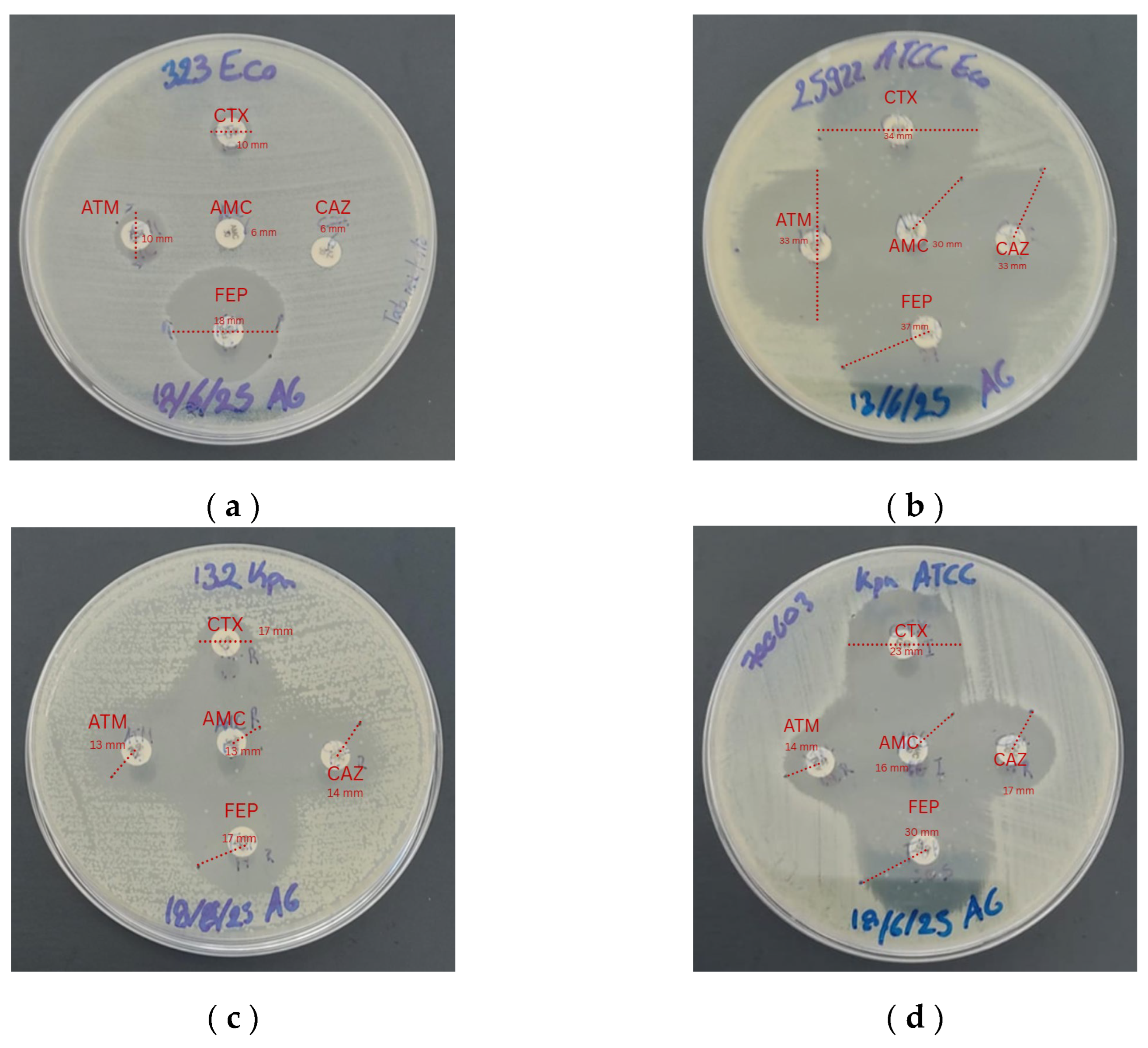
Phenotypic detection of ESBL production was carried out using the double-disk synergy test (Jarlier et al., 1988), employing the antibiotics amoxicillin/clavulanic acid (20/10 µg), aztreonam (30 µg), cefotaxime (30 µg), ceftazidime (30 µg), and cefepime (30 µg).
Mueller–Hinton agar plates were inoculated with a bacterial suspension adjusted to the 0.5 McFarland standard. The disks were placed at a distance of 25 mm and incubated at 37 °C for 18–24 h. The appearance of deformation or enlargement of the inhibition zone toward the amoxicillin/clavulanic acid disk was considered indicative of ESBL production. Inhibition zone diameters were measured and interpreted according to CLSI breakpoints (2025).
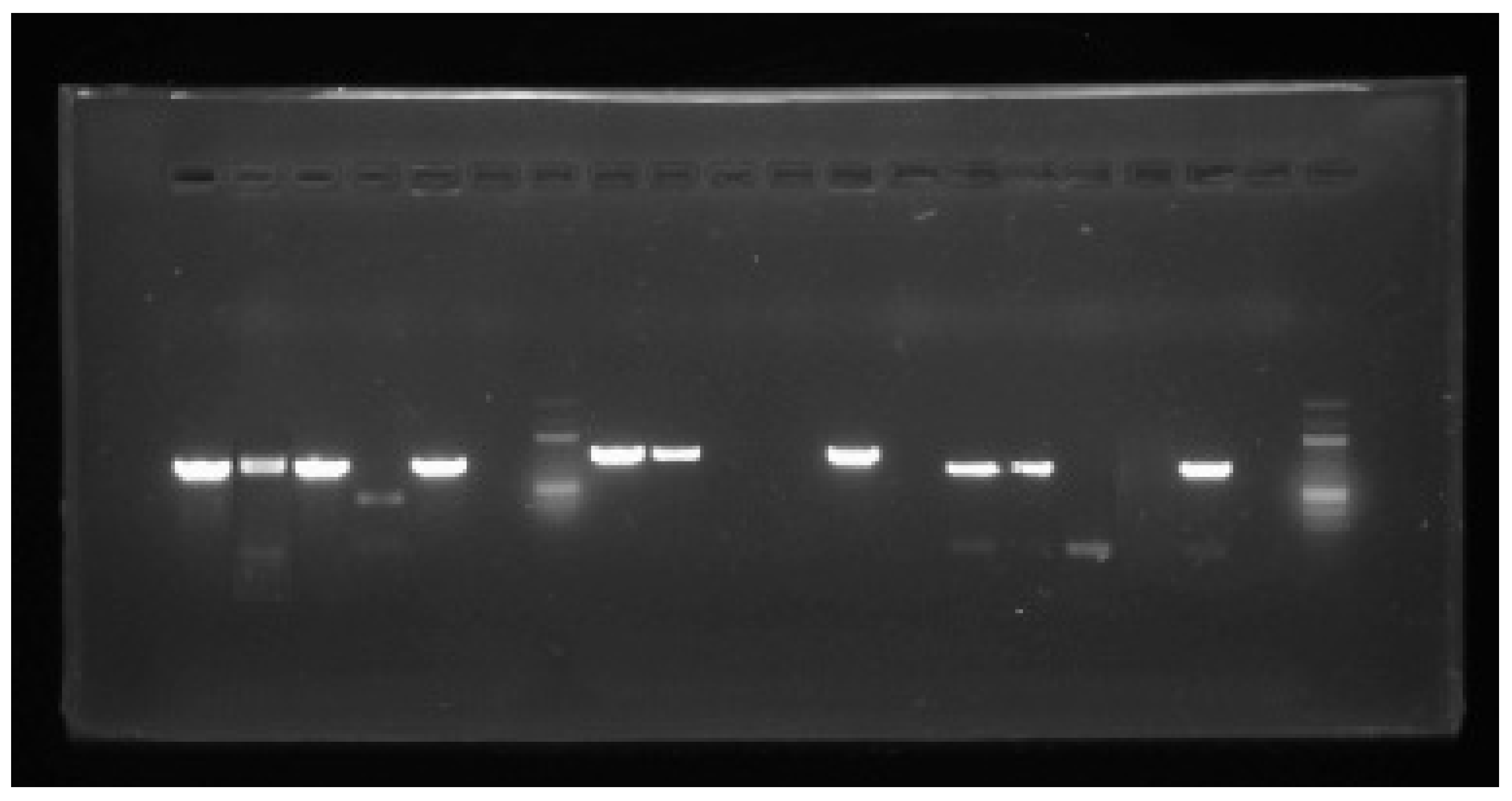
2.4. Molecular Identification by PCR
Five to six fresh colonies from each strain were collected and placed in 1 mL of TAE buffer, then incubated in a Thermomixer (Senacyt) at 95 °C with constant agitation for 10 minutes, followed by centrifugation to mechanically extract the genetic material. PCR reactions were prepared in a final volume of 24 µL containing 10 µL of molecular-grade water, 12 µL of GoTaq® Master Mix, 0.5 µL of forward primer, 0.5 µL of reverse primer, and 1 µL of DNA. Amplification was performed for two hours over 36 cycles in a Bio-Rad thermocycler. The samples were loaded onto a 1% agarose gel prepared with 1% TAE buffer and SafeView Classic® as the staining agent, and electrophoresis was carried out at 80 V for 40 minutes. After electrophoresis, the gel was visualized using a Bio-Rad gel documentation system for gene detection.
Table 1.
PCR conditions for DNA amplification.
Table 1.
PCR conditions for DNA amplification.
| Gene |
Primer sequence |
Size |
Denaturation |
Hybridization |
Extension |
Final extension |
| bla_SHV |
SHV-F |
ATGCGTTATATTCGCCTGTG |
449 |
94° for 30 seconds |
52.8° for 40 seconds |
72° for 50 seconds |
72° for 5 minutes |
| SHV-R |
TGCTTTGTTATTCGGGCCAA |
| bla_TEM |
TEM-F |
AAACGCTGGTGAAAGTA |
516 |
94° for 30 seconds |
52.8° for 40 seconds |
72° for 50 seconds |
72° for 5 minutes |
| TEM-R |
AGCGATCTGTCTAT |
| bla_CTX-M |
CTX-M-F |
TTTGCGATGTGCAGTACCAGTAA |
172 |
94° for 30 seconds |
52.8° for 40 seconds |
72° for 50 seconds |
72° for 5 minutes |
| CTXM-R |
CGATATCGTTGGTGGTGCCATA |
2.5. Origanum vulgare Essential Oil
The essential oil was acquired from the company KHIWA NATURAL (Quito, Ecuador), obtained by steam distillation, and accompanied by a certificate reporting the purity of the essential oil, density (0.935 g/mL), and the quantified content of secondary metabolites detailed in
Table 2.
Due to its hydrophobic nature, the essential oil was solubilized using 1% DMSO at a 1:1 ratio.
2.6. Determination of the Minimum Inhibitory Concentration (MIC)
The MIC of oregano essential oil and the antibiotic cefepime was determined using the broth microdilution method in 96-well plates, in accordance with CLSI guidelines (2025).
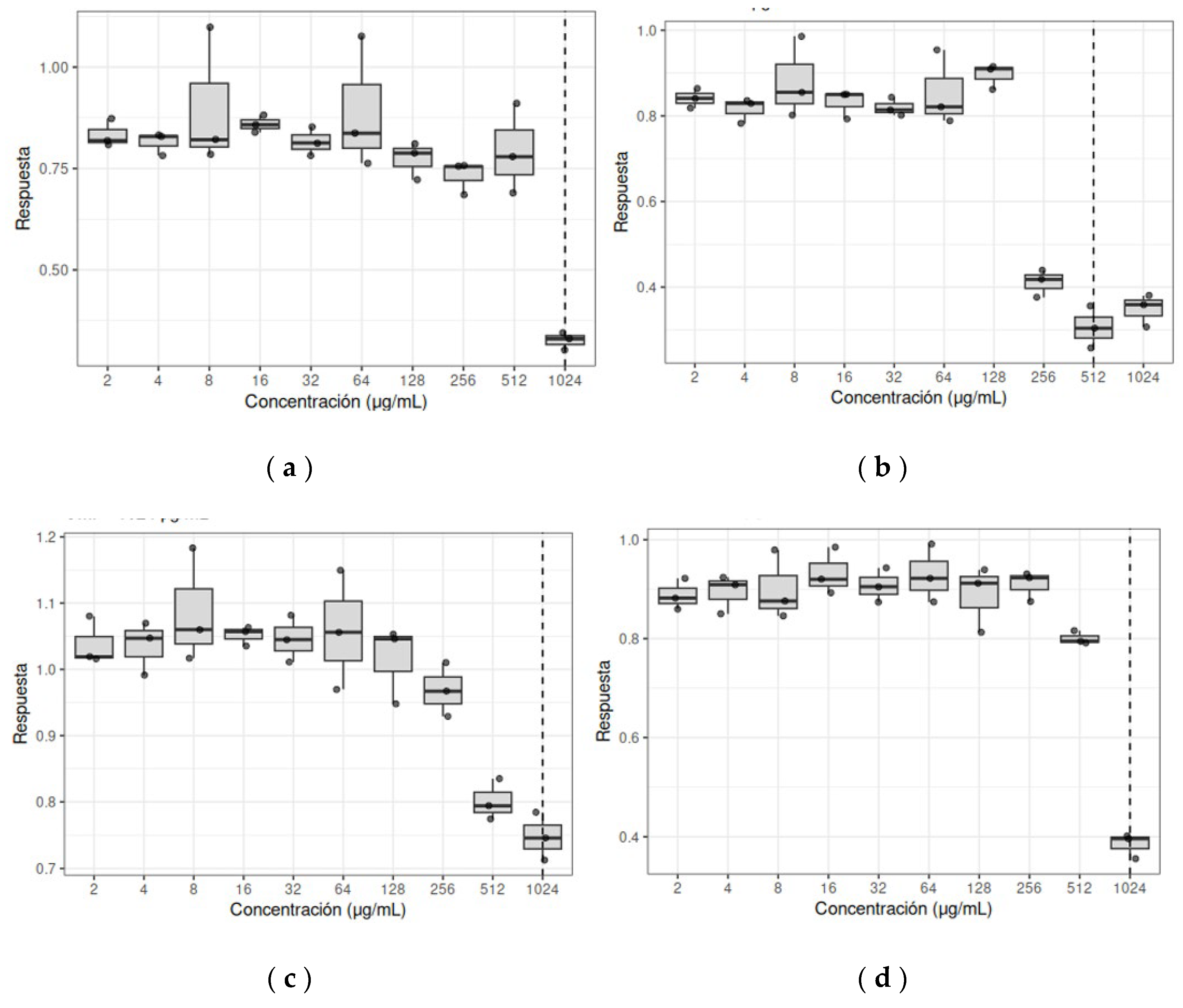
Ten serial dilutions of the essential oil were prepared with a bacterial inoculum of 5 × 10⁵ CFU/mL in Mueller–Hinton broth. Plates were incubated at 37 °C for 24 h, and absorbance was measured using an mcr® microplate reader at an OD of 630 nm. The MIC was defined as the lowest concentration with no visible growth. Each assay was performed in triplicate to ensure data consistency, including a DMSO control with no significant differences. For statistical analysis, data normality was initially assessed using the Shapiro–Wilk test. Since the data did not follow a normal distribution and corresponded to ordered concentrations in a dose–response design with a small sample size (n = 3), the non-parametric Jonckheere–Terpstra test was applied to evaluate the monotonic trend of the inhibitory effect of the essential oil as a function of concentration. Given the discrete dilution design and limited number of replicates, regression-based approaches were not applied.
The MIC was reported in an exploratory manner as the concentration associated with maximum inhibition (minimum median response), considering this criterion conservative and consistent with the microbiological interpretation of the assay, based on experimentally observed inhibition patterns rather than exclusively on statistical criteria. Statistical analyses were performed using RStudio software.
2.7. Determination of the Fractional Inhibitory Concentration
A 6 × 6 well matrix was prepared, in which the antibiotic was placed in serial horizontal dilutions across columns, and the essential oil was placed in serial vertical dilutions across rows. For this method, it is necessary to start from the MIC observed for each strain. Interpretation was performed according to the following criteria: FICI ≤ 0.5, synergy; 0.5 < FICI ≤ 1.0, additive effect; 1.0 < FICI ≤ 4.0, indifference; FICI > 4.0, antagonism. The interaction between the essential oil and cefepime was evaluated by calculating the FICI using the following formula:
Equation 1. Calculation of the FICI
4. Discussion
Phenotypic resistance profile
The results obtained from the disk diffusion test (Kirby–Bauer) for the
E. coli study strain showed resistance to AMC (6 mm), ATM (10 mm), FEP (18 mm), CTX (10 mm), and CAZ (6 mm), similar to the study by Kumar et al. (2014), in which 65% of clinical isolates exhibited complete resistance to cephalosporins (ceftazidime, cefotaxime, and cefepime). Likewise, Rehman et al. reported resistance rates of up to 100% to cefepime and cefotaxime, with inhibition zones < 16 mm, confirming the multidrug-resistant profile of the strain evaluated in the present study [
15], In contrast, the control strain
E. coli ATCC 25922 showed susceptibility to all evaluated antibiotics, consistent with the findings reported by Hudzicki, who indicated susceptibility to monobactams with inhibition zones equal to or greater than 30 mm in diameter [
16], ATCC strains are essential as reference controls to confirm the reliability of the assay [
17].
For the
K. pneumoniae study strain, reduced inhibition zones were observed: AMC (13 mm), ATM (13 mm), FEP (17 mm), CTX (17 mm), and CAZ (14 mm), in agreement with Smith et al., who established that inhibition zones below 18 mm for FEP, AMC, and ATM indicate the presence of ESBL production [
18] Similarly, Husna described that resistance to monobactams and combinations such as AMC/ATM in ESBL-producing
K. pneumoniae strains may exceed 90% of cases, which is consistent with the results obtained in this study [
19]. Regarding the control strain
K. pneumoniae ATCC 700603, the inhibition zones were within the quality control ranges established by EUCAST/CLSI, demonstrating a typical ESBL phenotype and validating the disk diffusion method. This strain is used as an ESBL control because it carries natural resistance associated with the
bla_SHV-18 gene [
20].
Antimicrobial activity of the essential oil (MIC)
The MIC of oregano essential oil for the
E. coli study strain was 1024 µg/mL. This value is consistent with, although slightly higher than, that reported by Burt, who obtained an average MIC of 500 µg/mL for oregano essential oil against ESBL-producing
E. coli strains [
21], This variability is attributed to differences in the chemical composition of the essential oil or to the study strains used, which may present distinct resistance profiles, resulting in greater natural or acquired susceptibility to natural compounds. For
E. coli ATCC 25922, the obtained concentration (256 µg/mL) is consistent with the study by Khan et al., who reported an MIC of 225 µg/mL for the compound carvacrol [
22]. Similarly, Stamova et al. reported a concentration of 250 µg/mL [
23], supporting the validity of the results obtained in this study, as the essential oil used contains 86.74% carvacrol, providing high efficacy and confirming its antimicrobial activity through damage to the cytoplasmic membrane and alteration of cellular metabolism [
21].
For the
K. pneumoniae study strain, the MIC was 2048 µg/mL. Several studies have also evaluated oregano essential oil against clinical
K. pneumoniae strains, including those with ESBL mechanisms, and have reported MIC values ranging from 128 to 256 µg/mL [
24]. Another study conducted in Bulgaria using clinical isolates reported a very broad MIC range of 390 to 12,500 µg/mL; therefore, the strain used in the present study falls within the range observed for resistant strains. Finally, for the control strain
K. pneumoniae ATCC 700603, the observed concentration of 1024 µg/mL represented significant inhibition due to its resistant nature. In comparison, a study that also evaluated the antimicrobial activity of oregano essential oil reported an MIC of 512 µg/mL [
25], while Bouhaddouda evaluated the same strain and reported MIC values of 1560 µg/mL, indicating a variable range [
26].
It should be noted that, although the statistical trend analysis supported a concentration-dependent inhibitory effect starting at 1024 µg/mL, the MIC for the K. pneumoniae study strain was conservatively reported as 2048 µg/mL. This decision was based on the microbiological criterion of complete and consistent inhibition observed across replicates, rather than solely on statistical significance. Given the exploratory nature of the study and the intrinsic resistance of K. pneumoniae, this conservative approach was adopted to ensure coherence with experimental inhibition patterns.
Antimicrobial activity of cefepime (MIC)
The
E. coli study strain exhibited a cefepime MIC of 64 µg/mL. This value is consistent with the study by Kumar et al. (2014), in which more than 60% of clinical isolates presented a cefepime MIC of 64 µg/mL, providing clear evidence of the epidemiology of cefepime resistance in ESBL-producing strains. In addition, CLSI and EUCAST [
27] indicate that MIC values above 16 µg/mL are associated with CTX-M–type enzymes and porin loss mechanisms, indicating a high resistance profile. In contrast, the cefepime MIC for
E. coli ATCC 25922 was 0.12 µg/mL, which falls within the quality control (QC) range established by the CLSI 2025 standard, defined as 0.016 to 0.12 µg/mL; therefore, the obtained values do not exceed the established threshold.
For the
K. pneumoniae study strain, the results showed a cefepime MIC of 64 µg/mL, confirming its resistant condition. Liang et al. evaluated 110 clinical ESBL-producing
K. pneumoniae isolates and reported that 47% of these strains exhibited resistance greater than 16 µg/mL, suggesting that such resistance levels are common in strains producing CTX-M or SHV-5 β-lactamases [
27]. Patel reported that ESBL-producing
K. pneumoniae with MIC values ranging from 2 to 8 µg/mL are associated with poor clinical response to cefepime [
28]. In the case of the control strain
K. pneumoniae ATCC 700603, the observed MIC was 1 µg/mL, which is within the QC ranges established by CLSI 2025 (0.25–2 µg/mL).
Antimicrobial interaction (FICI) between Origanum vulgare essential oil and cefepime
The combination of oregano essential oil with cefepime exhibited a synergistic effect against
E. coli, indicating that carvacrol and thymol enhance the action of the antibiotic by increasing outer membrane permeability and facilitating the entry of the β-lactam, a mechanism previously described in the literature [
13]. In addition, recent studies have reported that essential oils rich in carvacrol can reduce the expression of β-lactamases through gene suppression and protein inhibition, thereby decreasing the MIC of antibiotics in resistant Enterobacteriaceae, which is consistent with the marked MIC reduction observed in this study [
12,
29].
In
K. pneumoniae the observed effect was additive, which is consistent with investigations indicating that this species exhibits lower susceptibility to hydrophobic compounds due to its polysaccharide capsule and reduced membrane permeability
[30]. Despite this, the partial reduction of the antibiotic MIC in combination suggests an adjuvant effect of the essential oil. These results, together with findings reported in recent reviews, reinforce the potential use of essential oils as enhancers of antimicrobial therapies against multidrug-resistant bacteria.
5. Conclusions
This study demonstrates that the evaluated clinical strains of E. coli and K. pneumoniae exhibit a high resistance phenotype to β-lactam antibiotics, consistent with the expected profiles of ESBL-producing bacteria. The use of ATCC reference strains allowed validation of the applied methodologies and reinforced the reliability of the experimental results.
Origanum vulgare essential oil exhibited antimicrobial activity against all evaluated strains, although differences in susceptibility between species were observed. E. coli showed greater sensitivity to the essential oil, whereas K. pneumoniae required higher concentrations to achieve inhibition, reflecting its greater intrinsic resistance. Statistical analysis supported the observed trend, indicating a concentration-dependent inhibitory trend.
The evaluation of the fractional inhibitory concentration index (FICI) revealed synergistic interactions between the essential oil and cefepime in E. coli, as well as additive effects in K. pneumoniae. These findings suggest that the essential oil, particularly due to its high content of carvacrol and thymol, may act as an adjuvant in combination approaches against resistant Enterobacteriaceae.
Overall, the findings support the antimicrobial potential of O. vulgare essential oil and its ability to enhance the activity of β-lactam antibiotics against ESBL-producing bacteria. However, as this was a pilot study, further investigations involving a larger number of strains, cytotoxicity assessments, and evaluation in in vivo models are required to determine safety, efficacy, and potential clinical relevance.
Author Contributions
Conceptualization, A.G.; methodology, A.G. and R.A.; software, A.G.; validation, A.G. and M.E.; formal analysis, A.G., M.E., and R.A.; investigation, A.G.; resources, A.G. and M.E.; data curation, A.G. and M.E.; writing—original draft preparation, A.G. and M.E.; writing—review and editing, A.G., M.E., I.M., and R.A.; visualization, M.E. and R.A.; supervision, M.E. and R.A.; project administration, I.M. and M.E.; funding acquisition, I.M. and R.A. All authors have read and agreed to the published version of the manuscript.