1. Introduction
Tumors are inherently heterogeneous biological entities critically challenging the establishment of personalized treatment strategies in cancer therapy [1,2]. Precision oncology aims to design optimal therapeutic strategies based on the molecular and phenotypic characteristics of individual tumors [3,4]. Although advances in genomic profiling have enabled the identification of tumor specific molecular markers and facilitated the clinical implementation of various targeted therapies, their ability to accurately predict patient-specific treatment responses remains limited [5,6]. This limitation arises because of the biological complexity of tumors, which is shaped by intratumoral heterogeneity, clonal evolution, and interactions with the tumor microenvironment (TME), which cannot be sufficiently captured solely by genomic alteration data [5,7,8]. Consequently, patients at identical disease stages or with similar genetic alterations often exhibit markedly different responses to therapies. Such variability in treatment outcomes is driven by a combination of factors, including intratumoral heterogeneity, differences in the TME, discordance between genomic alterations and functional drug responses, emergence of adaptive drug resistance, and inter-individual variability in immune responses [1,7,8,9]. Together, these factors represent a major unresolved limitation in the predictive capacity of current precision medicine approaches [4,5].
Owing to the fundamental heterogeneity of cancer, conventional two-dimensional (2D) cancer cell line-based models, which have been widely used in preclinical research, exhibit substantial limitations in recapitulating the complex biological features of patient-specific tumors, particularly intratumoral heterogeneity and tumor–TME interactions [2,8,10]. In 2D culture systems, the spatial architecture of tumor cells, as well as cell–cell and cell–matrix interactions, is highly simplified, which may result in discrepancies between the experimentally observed drug responses and those observed in clinical settings [10,11]. Patient-derived xenograft (PDX) models, in contrast, preserve the histological and genetic characteristics of the parental tumors relatively well; however, their application is fundamentally constrained by longer establishment times, higher costs, limited scalability, and the requirement for immunodeficient host animals, which restricts their utility in immunotherapy research [1,12]. Owing to these limitations, the development of novel experimental platforms capable of rapidly and accurately reflecting patient-specific tumor biology and functional drug responses has emerged as a critical prerequisite for the clinical implementation of precision oncology [4,13].
The demand for experimental systems that precisely represent individual tumor characteristics has driven advancements in patient-derived organoid (PDO)–based research. Because PDOs are established directly from patient tumor tissues, they preserve not only the histological architecture and genetic features of the parental tumors, but also a substantial degree of intratumoral phenotypic heterogeneity [2,13,14]. Notably, organoid models significantly reduce the long establishment times associated with PDX models and enable the direct ex vivo evaluation of therapeutic responses to specific anti-cancer agents. In this regard, PDOs have considerable potential as functional drug response prediction platforms that complement genomics-centered precision oncology approaches [4,14].
Furthermore, recent efforts have focused on integrating tumor derived organoids with microfluidic chip technologies to reconstruct in vivo tumor conditions more faithfully. By co-culturing tumor cells with key components of the TME, including stromal elements and vascular structures, these advanced platforms aim to recapitulate the physiological tumor complexity with greater fidelity [15,16,17]. Through such technological advancements, organoid-based platforms are increasingly recognized not merely as experimental disease models, but as functional tools that support patient-specific treatment stratification and clinical decision-making in precision oncology [4,14].
In this review, we selected head and neck cancer (HNC), lung cancers, and hematological malignancies as representative disease groups to illustrate the diverse challenges faced by precision medicine. Although these malignancies differ substantially in terms of their tissue origin, genetic architecture, and clinical treatment strategies, there is a common unmet need for functional platforms capable of predicting patient-specific therapeutic responses. HNCs exemplify treatment resistance driven by pronounced intratumoral heterogeneity and variability in response to radiotherapy [18,19]. Lung cancer is a paradigm case of genotype-driven targeted therapy complicated by the rapid emergence of drug resistance [20]. In contrast, hematological malignancies underscore the critical role of a specialized TME and immune interactions in determining therapeutic outcomes (
Figure 1). By integrating organoid-based studies across these distinct disease contexts, this article aims to elucidate how organoid-based approaches are reshaping the paradigm of precision oncology, while highlighting the technical and translational challenges that must be addressed to facilitate broader clinical adoption [21].
2. Organoid Technologies in Cancer Research
Organoid technology has gained increasing recognition as a model well-suited for the development of patient-specific therapeutic strategies. In recent years, organoid platforms have moved beyond their original role as experimental disease models and have emerged as decision-support tools that functionally complement genomics-centered precision oncology for various cancer types, including lung, gastric, and ovarian cancers [4,22]. From a translational research perspective, organoids can recapitulate the key biological features of tumors with relatively high fidelity, thereby offering the potential to predict clinical treatment responses [23]. In particular, PDOs retain major driver mutations, copy number alterations, and transcriptomic characteristics of the parental tumors, allowing them to reflect intratumoral phenotypic heterogeneity to a meaningful extent [14]. Organoid platforms have been increasingly applied in drug response screening and biomarker discovery and are being explored as clinically relevant tools for optimizing personalized treatment strategies in precision medicine [22].
In this context, the present review aims to comparatively examine the functional value of organoid-based approaches across distinct clinical settings by focusing on three representative cancer types [2,4]. HNCs exhibit marked interpatient variability in treatment response, underscoring the need for functional response prediction beyond established standard-of-care regimens [18,19]. Lung cancer provides a paradigmatic context in which discordance between genomic information and actual therapeutic outcomes can be systematically evaluated [21]. In contrast, hematological malignancies offer a distinct context in which the applicability and limitations of organoid platforms can be critically assessed, owing to their specialized microenvironmental and immune dependencies [19]. By examining these heterogeneous disease contexts, this review aimed to delineate the role of organoid technologies as functional predictive tools that address shared unmet needs in precision oncology across diverse cancer types.
3. Organoid Models in Head and Neck Cancers
3.1. Overview of Head and Neck Cancers and Clinical Challenges
HNC is a malignancy of the oral cavity, pharynx, larynx, and other head and neck regions. Approximately 90% of these cases are head and neck squamous cell carcinomas (HNSCC) [18]. Although tobacco and alcohol consumption have long been established as major risk factors, human papillomavirus (HPV) infection has recently emerged as a particularly important etiological factor for oropharyngeal cancer [18,19,24]. Unlike HPV-negative HNC, which is typically associated with smoking and alcohol consumption, HPV-positive HNC tends to occur in younger patients and generally responds better to therapy, making these two subtypes clinically distinguishable [25].
Despite these differences, the overall prognosis for advanced-stage HNC remains poor, with high rates of recurrence and metastasis, even after aggressive standard treatment. In fact, 50–60% of patients with locoregionally advanced disease experience recurrence within 5 years of the completion of concurrent chemoradiotherapy or similar intensive regimens [26,27]. Moreover, multidisciplinary treatment (MDT) approaches, including surgery, radiotherapy, and chemotherapy, often result in severe side effects, and some patients are unable to complete the entire course of treatment [28].
Another critical challenge in the management of HNC is the molecular heterogeneity of the disease [18]. Each patient may exhibit a unique profile of genetic mutations and distinct TME, leading to widely varying responses to the same therapeutic regimen. However, reliable biomarkers that are capable of predicting individual treatment responses are lacking. As a result, the current clinical practice typically relies on a one-size-fits-all treatment strategy, regardless of the specific tumor characteristics of the individual patient [26,29]. This approach not only reduces therapeutic efficacy but also increases the risk of unnecessary toxicity. Therefore, there is an urgent need for novel model systems that faithfully represent patient-specific tumor biology and allow preclinical testing of therapeutic responses, ultimately paving the way for personalized treatment strategies [30].
Particularly, in HNSCC differences in radiosensitivity, the high rate of post-treatment recurrence, and the clinical and molecular heterogeneity between HPV-positive and HPV-negative subtypes collectively hinder the implementation of personalized therapeutic strategies [31,32]. Whereas HPV-positive tumors are highly immunogenic due to viral antigen expression and generally exhibit greater radiosensitivity and improved survival, HPV-negative tumors are characterized by a higher mutational burden, increased radioresistance, and a greater risk of recurrence [32]. This heterogeneity underscores the need for precision-medicine platforms capable of prospectively evaluating patient-specific radiosensitivity and treatment responsiveness. Recent studies have demonstrated that PDOs enable direct in vitro assessment of individual tumor responses to radiation and chemotherapeutic agents, while preserving HPV subtype–specific molecular features and therapeutic vulnerabilities [26,33,34]. Therefore, PDOs represent a promising platform for addressing precision-medicine needs in HNSCC, including the prediction of radioresistance, assessment of recurrence risk, and the development of HPV subtype–tailored therapeutic strategies.
3.2. Establishment of HNC Organoids
3.2.1. Methods for Obtaining Patient Samples and Success Rates
The establishment of HNC organoids typically begins with the immediate processing of tumor tissue obtained from surgical resection or biopsy [33]. Fresh tumor specimens are mechanically dissociated or enzymatically digested into single cells or small cell clusters, which are then embedded in an extracellular matrix (ECM) commonly based on Matrigel. Culturing is performed using customized media supplemented with specific growth factors such as EGF, R-spondin, and Noggin to promote tumor cell proliferation [26,30].
The reported success rate for establishing organoid cultures from patient-derived HNC samples ranges from approximately 35% to 77%, depending on factors such as tumor cell content, prior radiation treatment, and the time elapsed between tissue collection and processing [26]. Organoid establishment tends to be less successful in older patients or when pretreated tissue samples are used. For example, Fisch et al. reported a 77% success rate when using high-quality tissue samples containing ≥30% tumor cells. In contrast, delayed tissue processing and low tumor cell content have been identified as major causes of culture failure [34].
3.2.2. Modeling Differences Between HPV-Positive and HPV-Negative Tumors
HNSCC exhibits markedly different molecular characteristics and clinical behaviors depending on the status of the HPV. HPV-positive tumors predominantly arise in the oropharynx (e.g., tonsils and base of the tongue), are more common in younger patients, and generally have a more favorable prognosis. This is largely attributed to their higher sensitivity to radiation and chemoradiation therapies than HPV-negative tumors [32]. HPV-positive tumors are characterized by the degradation of p53 protein and inactivation of Rb protein via the viral oncoproteins E6 and E7. Notably, these cells usually retained wild-type TP53 and overexpressed p16 [35,36].
In contrast, HPV-negative tumors are typically associated with risk factors such as tobacco and alcohol use and tend to harbor TP53 mutations, EGFR pathway activation, and a higher overall mutational burden [31]. Although HPV-positive tumors exhibit a lower total mutational burden, they are often more immunogenic owing to the presence of viral antigens and are associated with higher levels of immune cell infiltration. HPV-negative tumors, in contrast, demonstrate greater genetic heterogeneity due to mutation accumulation [26,31]. Indeed, large-scale immunogenomic analyses have revealed that HPV-positive HNSCCs possess significantly higher levels of tumor infiltrating lymphocytes (TILs) and immune-related gene expression than HPV-negative tumors, although both subtypes exhibit immune microenvironments potentially responsive to immune checkpoint blockade [31,36,37].
PDO models are actively utilized to recapitulate and investigate the differential treatment responses and biological characteristics of HPV-positive and HPV-negative HNSCC in vitro. Recent studies have established biobanks for HNSCC organoids, including HPV16-positive and TP53 wild-type tumors. These organoids preserve the histological and molecular features of their parental tumors and effectively represent HPV-specific subtypes. For example, organoids derived from HPV-positive tumors exhibited p16 overexpression and TP53 wild-type status, and transcriptomic profiling confirmed the retention of HPV-associated gene expression signatures that distinctly separated HPV-positive and HPV-negative subgroups [26,33,38]. Functionally, HPV-positive organoids demonstrate increased radiosensitivity. Notably, the ectopic expression of HPV E6/E7 in normal cells or HPV-negative tumor organoids induces cell cycle arrest and enhances radiation sensitivity, suggesting that HPV oncogene-mediated cell cycle vulnerability may underlie the heightened therapeutic responsiveness of HPV-positive tumors [38,39].
Furthermore, a study investigating disease progression and prognostic factors in HPV-related HNSCC using patient-derived models found that organoid and PDX growth rates in HPV-positive patients were positively correlated with the tumor mutational burden. This implies that, within the HPV-positive subgroup, higher genetic heterogeneity may be associated with more aggressive tumor biology [10,40]. Conversely, in HPV-negative HNSCC, a recent study employed 3D collagen-based cultures of PDOs to analyze the invasive behavior and prognostic factors. Single-cell transcriptomic profiling has revealed that tumor cells with activated YAP signaling exhibit enhanced collective invasion, and this invasive gene signature is associated with poor patient survival [41]. These findings highlight the utility of organoid models in faithfully reproducing the molecular and phenotypic distinctions between HPV-positive and -negative HNSCC, serving as powerful preclinical platforms for exploring subtype-specific therapeutic responses and prognostic indicators.
3.3. Drug Screening and Biomarker Discovery
3.3.1. Target Evaluation
In HNSCC, the epidermal growth factor receptor (EGFR) and PI3K/AKT/mTOR signaling pathways represent major molecular targets. EGFR is overexpressed or activated in approximately 80% to 90% of patients with HNSCC and is known to contribute to tumor cell proliferation and survival. Cetuximab, a monoclonal antibody targeting EGFR, was the first agent to be approved for the treatment of HNSCC [26]. However, its efficacy varies among the patient subgroups. In particular, the additional benefits of cetuximab in patients with HPV-positive oropharyngeal cancer appear to be limited [42]. From a molecular epidemiological perspective, PIK3CA mutations are found in over 30% of HPV-positive HNSCC cases, which is significantly higher than in HPV-negative cases. This suggests that the PI3K/AKT/mTOR pathway may serve as a key therapeutic target for HPV-positive tumors [10].
In contrast, EGFR gene amplification or pathway activation is more frequently observed in HPV-negative tumors, with more than half of all HPV-negative HNSCC cases showing activation of either the EGFR or PI3K pathway [24,31,43]. These findings underscore the importance of selecting targeted therapies based on the patient’s molecular profile, and PDO drug testing is emerging as a valuable tool for implementing such precision medicine strategies.
Drug sensitivity studies using HNSCC organoids have provided evidence supporting the prediction of individual tumor responses to targeted therapies. Driehuis et al. established a panel of organoids derived from both HNSCC tumors and normal tissues and conducted a medium-scale drug screening study. They demonstrated that the sensitivity of tumor organoids to clinically relevant anti-cancer and targeted agents closely mirrored the actual treatment responses of the patient [33]. For instance, organoids that respond to EGFR inhibitors are likely to represent patients who would benefit from cetuximab, whereas organoids sensitive to PI3K inhibitors often harbor pathway alterations, providing a rational basis for clinical administration [26,43,44].
Consistent with this, clinical trial results have shown that the efficacy of PI3K inhibitors differs according to the molecular subtype. In the BERIL-1 trial evaluating buparlisib, improvement in progression-free survival (PFS) was limited in the overall HNSCC population but was significantly better in patients with PIK3CA mutations or PTEN loss, indicating alterations in the PI3K pathway [44]. These findings suggest that evaluating patient-specific drug responsiveness using organoid assays may help to identify optimal targeted therapies.
A recent study by Millen et al. assessed drug responses across 40 HNSCC organoid models and analyzed their correlation with clinical treatment outcomes [26]. In some patients, organoid sensitivity to chemoradiotherapy is closely associated with actual clinical outcomes, such as complete remission. This study also showed the potential of organoids to identify predictive biomarkers for patient prognosis [26,38,45]. Interestingly, in experiments treating organoids with radiation alone or in combination with chemotherapy, the addition of cetuximab reduced radiosensitivity. This suggests that combining cetuximab with radiotherapy might result in inferior outcomes compared with radiotherapy alone, mirroring clinical observations in HPV-positive patients, where such combination therapy was associated with decreased survival [26,42]. These findings highlight how organoid models can reflect not only treatment efficacy but also potential antagonistic interactions in combination therapies, thereby informing treatment strategies [26].
Organoid-based studies on PI3K-targeted therapies are ongoing. In several preclinical models, organoids and PDXs harboring PIK3CA mutations have shown selective sensitivity to PI3Kα inhibitors such as alpelisib, resulting in suppressed tumor growth [43]. Given the frequent activation of the PI3K pathway in HPV-negative HNSCC, PI3K inhibition has also been proposed as a potential radiosensitizing strategy in this subgroup, which often exhibits intrinsic radioresistance [24,31,43]. Collectively, organoid-based drug sensitivity analyses serve as a platform for predicting patient-specific responses to molecular targeting and identifying rational combination therapies in precision medicine settings [26,43].
3.3.2. Predicting Immunotherapy Response
The advent of immune checkpoint inhibitors (ICIs) has improved clinical outcomes in patients with recurrent or metastatic HNSCC. Notably, pembrolizumab, an anti-PD-1 antibody, has been approved as a first-line treatment for metastatic HNSCC and has demonstrated long-term survival benefits in a subset of patients. Similarly, nivolumab has shown significant survival advantages in patients who have failed prior therapies [45,46]. However, the objective response rate to ICIs remains limited to approximately 20% of patients, highlighting the urgent need for predictive biomarkers. Although tumor PD-L1 expression levels and tumor mutational burden are currently used as potential indicators of treatment response, their predictive accuracy is insufficient [47]. Therefore, there is a growing demand for models that can directly assess the immune responsiveness of individual patient tumors [48].
Organoid models naturally lack immune components, which poses a challenge in evaluating immune responses. Several recent technological advancements have addressed this issue. One approach involves co-culture systems, in which patient-derived peripheral blood mononuclear cells (PBMCs) or TILs are cultured together with tumor organoids to recreate tumor–immune interactions in vitro [49,50].
For example, Neal et al. (2018) established air–liquid interface (ALI) cultures from various human tumors, including lung cancer, which preserved endogenous immune cells within the organoid. Upon treatment with anti-PD-1 antibodies, they observed increased CD8+ T-cell infiltration and enhanced tumor killing activity, successfully recapitulating the immune checkpoint blockade responses ex vivo [49]. In other words, co-culturing patient-derived mini-tumors with autologous immune cells allows researchers to directly test whether immune cells can eliminate tumor cells in response to ICIs. Similarly, Dijkstra et al. developed a method for co-culturing colorectal cancer organoids with autologous blood-derived T cells, leading to the enrichment and identification of tumor specific T cell populations [50]. These techniques may apply to HNCs in the future and could potentially serve as a platform for personalized immunotherapy testing. Recent studies have reported efforts to predict ICI responses using organoid–immune cell co-cultures. Organoid-based models have been successfully used to predict patient responses to immunotherapy [47,48].
Another strategy is to modify the organoid culture environment to retain the endogenous immune components [49]. For example, ALI-based organoids can maintain not only tumor cells but also resident immune cells, such as T cells, natural killer (NK) cells, and macrophages, for a certain duration. These “immune cell-retaining” organoids offer a unique opportunity to study the TME and immune checkpoint pathways without the need for exogenous immune cell supplementation. A recent study on HNSCC using an ALI-based organoid platform demonstrated that tumor derived organoids preserved CD3+ TILs and that PD-1/PD-L1 blockade enhanced T-cell infiltration and interferon-gamma secretion, indicating that immune-responsive tumors can exhibit comparable immunologic changes in vitro [48,51,52]. However, these technologies are still in the research phase and further development is needed to ensure the long-term viability and proper function of immune cells in organoid cultures, particularly in HNCs.
Nevertheless, PDO platforms have already been used for personalized immune profiling. For instance, some models have been used to test the efficacy of CAR-T cells or tumor vaccines against individual tumors [53,54].
In summary, organoid models represent a promising tool for assessing responsiveness to immune therapy at the patient level, which is difficult to predict using conventional methods. They are expected to contribute to the stratification of patients for immunotherapy and discovery of novel immune targets in the future [48,49,50].
3.4. Clinical Translation and Limitations
Efforts to explore the clinical applications of PDOs are gradually emerging [34,55,56]. Recently, a preclinical study demonstrated the use of organoids derived from patients with HNSCC to design personalized treatment strategies. In this pilot study, tumor tissues from patients with locally advanced HNSCC were used to establish organoids, and their responses to various chemotherapeutic agents and radiation were evaluated to help inform clinical decision-making [34]. Although limited in scale, this study provides meaningful evidence that drug sensitivity results from PDOs may correlate with patient responses and aid in optimizing treatment strategies. Although organoid-based personalized treatment remains largely at the research stage, these early findings suggest its potential for integration into clinical practice in the near future [34,55,56].
Despite this promise, several key challenges must be addressed before the PDO technology can be widely adopted in clinical settings. First, organoid establishment and culture require a considerable amount of time (often several weeks), and the success rate of organoid generation is not uniform across all patient samples. In some cases, insufficient tumor tissue or intrinsic properties of the tumor may lead to failure of organoid formation [34,55]. Second, current organoid cultures generally lack critical components of the TME, such as immune cells and vascular structures, limiting their ability to model responses to immunotherapies or tumor TME interactions [49]. Third, the generation and testing of organoids require specialized techniques and costly equipment, restricting their use to large medical centers or advanced research institutes [26]. Fourth, prolonged in vitro culture may lead to the accumulation of genetic or phenotypic alterations in tumor cells, necessitating caution when interpreting results from long-term cultured organoids [57,58].
Various approaches have been explored for overcoming these limitations. These include co-culturing organoids with immune cells and mimicking vascular structures using 3D bioprinting to better replicate the TME [49]. Additionally, efforts are underway to automate organoid culture and drug-screening platforms to improve the efficiency and success rates of organoid establishment [59]. Despite current limitations, PDOs are emerging as promising tools for guiding precise treatment decisions and accelerating the development of new therapies for HNC. With continued advancements, integration into clinical practice is expected [26,34].
In summary, PDO models in HNSCC are well suited for [18] capturing interpatient heterogeneity in therapeutic responses, including variations in radiosensitivity, chemotherapy sensitivity, and risk of recurrence and preserving HPV subtype–specific molecular features relevant to precision treatment stratification [26,34,38]. Conversely, PDOs have a limited ability to [18] recapitulate the immune and stromal components required to model immunotherapy responses, and [26] provide sufficiently rapid turnaround times (TAT) compatible with real-time clinical decision-making for newly diagnosed patients. Continued advances in microenvironmental modeling, co-culture systems, and automated screening platforms may help bridge these gaps and expand the clinical utility of PDOs in HNSCC [49,53,54].
4. Organoid-Based Precision Medicine in Lung Cancer
4.1. Lung Cancer Landscape: Genetic Heterogeneity Non-Small Cell Lung Cancer and Small-Cell Lung Cancer
Lung cancer is one of the leading causes of cancer-related deaths globally [60], reflecting both its high incidence and persistent difficulty in achieving durable treatments. In clinical settings, lung cancer management is challenged by the significant genetic heterogeneity across both Non-Small Cell Lung Cancer (NSCLC) and Small-Cell Lung Cancer (SCLC). This heterogeneity exists not only between tumors (intertumoral), but also within different regions of the same tumor (intratumoral), leading to variations in mutations, gene expression patterns, and chromosomal abnormalities. Such heterogeneity complicates accurate treatment selection and prediction of drug response and is a critical driver of acquired resistance to cancer therapies [1,61], limiting long-term clinical benefits.
In this context, the increasing importance of driver mutation-based targeted therapies and the need for TAT platforms highlight the critical need for preclinical models that can faithfully capture patient-specific tumor dependencies in clinically actionable timeframes [62].
4.2. Development of Lung Cancer Organoids
Effective precision medicine for lung cancer requires preclinical models that can preserve tumor heterogeneity, retain clinically relevant driver mutations, and deliver rapid results useful clinically. Historically, traditional 2D cell lines, such as A549, have been instrumental in cancer research; however, their inability to reflect patient-specific heterogeneity and treatment responses has limited their translational relevance.
PDX models, wherein tumor tissues from patients are implanted into immunodeficient mice, offer much improved preservation of tumor heterogeneity compared to 2D cultures [63]; however, their low engraftment rates, high costs, and ethical concerns limit their utility for clinical decision-making [64,65,66].
Compared with PDX models, PDOs offer faster expansion, the ability to be cryopreserved, and are suitable for high-throughput drug screening (HTS) [57]. Unlike traditional cell lines, lung cancer PDOs retain the patient’s tumor complexity and accurately reproduce the pathological and genomic characteristics of the parental tumors, including driver gene mutations [67,68,69]. Thus, PDOs offer a helpful middle ground between traditional cell lines and in vivo models.
PDOs are typically derived from biopsies, surgical resections, or malignant pleural effusions and have shown moderate to high success rates in cell cultures [68,69,70,71]. The derivation process typically involves dissociating tumor tissues into single cells, embedding them in ECM hydrogels, and culturing them under conditions that allow 3D growth [70]. However, because biopsy-derived PDOs represent only a limited spatial fraction of the tumor, the overall tumor heterogeneity may be underestimated because of sampling bias [72]. Even closely related cells from the same tumor can show markedly different responses to anti-cancer drugs [73].
Conventional lung cancer organoids maintain only malignant cancer cells and fail to preserve stromal components, such as immune cells. The interactions of these components with the epithelium play vital roles in tumorigenesis, tumor growth, and the response of the tumor to therapies [58,74].
One strategy developed to address this limitation is the use of ALI culture techniques. In ALI systems, tumor organoids are embedded in a gel, wherein one surface is in contact with the culture medium and the other is exposed to air. These PDOs retain the tumor epithelium and its stromal microenvironment with fibroblasts and immune cells [75,76]. The use of ALI systems to culture lung cancer PDOs is promising for observing therapies. while maintaining the key features of the TME. However, despite these advantages, ALI systems are constrained by their limited culture duration and difficulties with standardization, which restrict their scalability and routine implementation [58].
4.3. Applications in Targeted and Immunotherapy
4.3.1. Evaluation of Therapies Targeting EGFR, ALK, and KRAS
Given the central role of actionable driver mutations in lung cancer treatment, PDOs offer a clinically relevant model for evaluating the efficacy of targeted therapies for lung cancer. Because they retain the critical molecular characteristics of the original tumor, they are ideal for modeling therapeutic response and resistance in precision interventions [68].
PDOs have demonstrated high fidelity in recapitulating the clinical responses to EGFR-targeted therapies for NSCLC. Yao et al. showed that lung cancer PDOs retain driver mutations found in their parental tumors, including EGFR exon 19 deletions and BRAF G464A mutations. These PDOs mirror patient responses to clinically approved tyrosine kinase inhibitors (TKIs), such as afatinib and the dabrafenib/trametinib combination, successfully predicting PFS and objective responses [71].
PDOs can also serve as valuable tools for modeling rare and underrepresented variants. In a 2025 study, Jeong et al. demonstrated that the PDOs YOU-076 and YOU-139, derived from patients with prior EGFR-TKI exposure, showed sensitivity to lazertinib-amivantamab. This combination enhanced the antitumor potency six-fold in vitro and was validated using a xenograft model [77]. Other case studies have found that PDOs were able to provide moderate prediction of drug sensitivity in NSCLC metastases, complex mutation fusions, and rare cases [71,78].
Recent advances have enabled rapid turnaround in drug sensitivity testing using PDOs. In a small-scale study, cancer organoid-based diagnosis reactivity prediction (CODRP)-based testing using malignant pleural effusion-derived PDOs accurately predicted EGFR-TKI response within 10 days. Lee et al. found that when drug sensitivity assays used the CODRP index method, cancer cells isolated from 14 patients outperformed conventional approaches [79]. Hu et al. reduced the time for drug sensitivity testing of lung cancer PDOs to 1 week on a moderate scale [62]. HTS systems built around PDO models have also allowed for the scalable testing of EGFR and HER2 inhibitors, expanding the translational pipeline [80].
PDOs also show promise as guiding therapies for anaplastic lymphoma kinase (ALK)-rearranged NSCLC. CODRP-based models outperformed conventional metrics in predicting sensitivity to ALK inhibitors (crizotinib, alectinib, and brigatinib) and accurately mirrored clinical outcomes [81].
A 2024 case study used lung cancer PDOs to evaluate a patient with ceritinib-resistant ALK-rearranged NSCLC, correctly predicting sensitivity to brigatinib and lorlatinib [82]. In a case report of a novel ALK fusion in a patient with stage IIIA lung adenocarcinoma, crizotinib was selected as adjuvant therapy using PDOs, and the patient achieved more than 3-year disease-free survival [81].
PDOs harboring KRAS mutations have also been used to evaluate therapeutic efficacy. Bian et al. utilized PDO models and found that aldehyde dehydrogenase 1 family member A1 (ALDH1A1) expression conferred resistance to KRAS inhibitors; however, combining ALDH1A1 inhibition with KRAS blockade restored the therapeutic efficacy in PDO models [83]. Additionally, PDOs harboring KRAS(G12C) mutations have been used to model the responses to sotorasib. The in vivo PDO metastasis model allows for the effective evaluation of anti-metastatic therapies [76]. These studies demonstrate the value of PDOs in the evaluation of therapies targeting mutations in EGFR, ALK, or KRAS.
4.3.2. The Role of PDOs in Immunotherapy
Another method used to address the absence of immune and stromal components in standard organoid models is the introduction of co-culture systems. Co-culture systems typically either retain and expand endogenous immune cells within tumor organoids or add immune cells to organoids, more accurately preserving tumor heterogeneity and allowing for more accurate dissection of the effects of various therapies [75].
Several organoid-based co-culture systems have demonstrated the potential to predict immune checkpoint inhibitor responsiveness. Co-culturing tumor organoids with cytotoxic T cells reintroduces essential elements of the TME and allows for the real-time evaluation of responses to ICIs that more closely mimic the in vivo response [84,85].
Organoid-based co-cultures are also being developed and screened for advanced immunotherapies. These include testing chimeric antigen receptor (CAR) T cells, generating tumor reactive lymphocytes for adoptive cell therapies, and reprogramming T lymphocytes to enhance tumor specific cytotoxicity [86,87]. As these technologies continue to mature, PDO-based co-culture platforms have promising implications for personalized immunotherapy landscapes.
4.4. Integration with Multi-Omics and Artificial Intelligence (AI)
The integration of PDOs with multi-omics profiling and AI presents a cutting-edge approach for advancing precision medicine for lung cancer. Multi-omics profiling can enhance the reliability of applying PDOs in discovering novel therapeutic targets using technologies like CRISPR/Cas9 and RNAi to validate therapeutic targets [88,89].
AI can be used to predict interactions between immune cells and tumor components and to compare multi-omics characteristics between treatment responders and non-responders [88]. Deep learning models are being developed to improve organoid analysis and culture processes, as well as to assess PDO responses to anti-cancer therapies [90,91].
Additionally, microfluidic systems mimicking physiological conditions (shear stress, fluid flow, and tissue interfaces) have been integrated with PDOs. Referred to as organoids-on-chips (OrgOCs), when integrated with multi-omic profiling, OrgOCs can identify how various omics layers respond under specific physiological conditions. In precision medicine, this allows for the real-time monitoring of PDOs, analysis using omics, and prediction of clinical responses [92].
Challenges remain in standardizing data pipelines and addressing technical concerns such as scalability and costs [93]. Ethical concerns, such as patient privacy and computational bias, must also be considered as AI models have become increasingly common [94]. Nonetheless, the convergence of PDOs, multi-omics, and AI holds immense promise for lung cancer treatment.
In lung cancer, PDOs excel at recapitulating patient-specific driver mutations and associated targeted therapy responses and show immense promise for enabling rapid functional drug sensitivity testing with clinically relevant turnaround times [80,81]. These strengths make lung cancer PDOs effective platforms for prioritizing targeted agents in genomically stratified NSCLC [71,77,95], particularly in settings that require accelerated treatment decisions. Despite this promise, lung cancer PDOs may still incompletely capture intratumoral heterogeneity due to sampling bias and remain relatively limited in their ability to stably reproduce immune-stromal interactions over extended culture periods [96]. Addressing these limitations is essential for broader clinical implementation of organoid-based precision medicine for lung cancer.
5. Organoids in Hematological Malignancies
5.1. Unique Challenges in Modeling Blood Cancers
Given that hematological malignancies are forecasted to reach 4.5 million cases by 2030 [60], the burden of developing stronger preclinical models for these diseases is increasing. Hematological malignancies present unique biological and modeling challenges owing to their non-solid tumor characteristics and dependence on specialized microenvironments [97]. Unlike solid tumors that can be surgically resected and modeled in structured 3D cultures, many blood cancers consist of circulating cells whose proliferation and therapeutic responses are dictated by dynamic, spatially organized niches, such as the bone marrow (BM) or lymphoid tissues [98,99]. Therefore, the inherent complexity of the blood cancer TME means that traditional modeling efforts are often forced to rely on simpler systems, predominantly 2D cell cultures [97].
The TME in blood cancers is exceptionally complex and heterogeneous, encompassing a remarkably diverse array of non-hematopoietic cells at various differentiation stages, including tumor associated macrophages, endothelial cells, various immune infiltrates, and other non-tumor cell types [97]. The nature of circulating cells implies a level of independence from the primary tumor reservoir, but their pathogenesis and drug response are still greatly influenced by their particular niches. However, these 2D culture systems fail to mimic the interactions between hematopoietic cells and their non-hematopoietic counterparts, limiting their ability to model disease biology and ex vivo responses to immunotherapy [97].
These limitations highlight the necessity for improved biomimetic 3D systems that can better replicate the spatial, mechanical, and biochemical complexities of the blood cancer microenvironment. These 3D systems aim to better simulate cell-cell and cell-matrix interactions, oxygen and nutrient gradients, and niche-specific signaling pathways [97]. Organoids offer a promising approach to model hematological malignancies in a manner that preserves microenvironmental dependence and supports ex vivo evaluation of immunotherapy in patient-relevant microenvironments [100].
5.2. Bone Marrow and Lymphoid Organoids
Although 2D cultures oversimplify the tumor ecosystem, organoids present a promising solution to these limitations [67,101]. Recent generations of bioengineering have enabled the development of BM and lymphoid organoids that can accurately simulate the cellular architecture of human cells by integrating mesenchymal stromal cells, endothelial networks, and hematopoietic progenitors [102,103].
However, because 2D cultures oversimplify in vivo BM function, BM organoids (BMOs) incorporate essential cellular and architectural features of the marrow [104]. Multiple studies have demonstrated the success of human induced pluripotent stem cell (iPSC)-derived BMOs in recapitulating normal and malignant hematopoiesis. The differentiation process involves the precise application of a series of cytokine cocktails and embedding of cells within a hydrogel matrix. These cells then organize into well-vascularized architectures that closely resemble native hematopoietic tissue [98,105]. These structures successfully supported the engraftment of patient-derived cells from multiple hematological malignancies, including acute myeloid leukemia (AML), CLL, and multiple myeloma (MM), while maintaining a disease-specific morphology [106].
This in vivo-like behavior makes BMOs invaluable tools for modeling the hematopoietic developmental processes. A 2024 study demonstrated the generation of BM-like organoids containing multipotent mesenchymal and vascular networks capable of autonomous hematopoiesis [107]. Similar findings regarding patient-derived hematopoietic stem and progenitor cells (hPSCs) from patients have been described for patients with myelodysplastic syndromes (MDS) [108]. This indicates that iPSC BMOs exhibit disease-specific lineage differentiation [109].
BMOs show great promise for application to specific types of hematological malignancies. In MM, cells actively modify the BM microenvironment, enhance disease progression, and promote drug resistance [110,111]. Organoids have shown promise as a platform to overcome drug resistance in MM by sensitizing tumor cells with small molecules that can correct dysregulated microRNAs (miRNAs), which are strongly associated with refractory disease in patients with MM [112].
AML is a particularly challenging blood cancer originating from the BM. Within the BM, AML cells establish a unique TME that shields the leukemic cells. The lack of long-term in vitro models that can be scaled for HTS poses a significant challenge in advancing AML research. Recent breakthroughs have introduced BMOs that support long-term culture and HTS [113].
Thus, the development of lymphoid organoids has gained traction [114]. In 2024, freshly resected lymphoma biopsy fragments were cultured ex vivo to develop lymphoid organoids that retained the histological and molecular features of the native tissue [115]. Thus, they were able to perform parallel drug screening that correlated with the clinical responses.
Both BM and lymphoid organoids have strong potential to bridge the translational gap between preclinical models and their application in the treatment of hematological malignancies [98]. Nonetheless, even state-of-the-art immune organoid models have limitations in fully recapitulating tissue structure and function [116]. Many lymphoid organoids rely on reaggregation techniques to generate morphologically unstable constructs, thereby reducing their downstream utility.
5.3. Personalized Drug Screening and Resistance Profiling
Tumor organoids have emerged as valuable in vitro models for drug testing because of their high fidelity to the native tumor [57,101]. However, the translation of organoid models into platforms for precision oncology relies heavily on their ability to sustain viable malignant hematopoietic cells long enough for functional drug testing [99]. The following studies demonstrate promise that the 3D culture of organoids may help overcome these limitations and enable patient-specific drug profiling.
For instance, in MM, the use of a 3D Matrigel-based BM niche significantly extended primary tumor cell viability by approximately 10 days. This was compared with the approximately 2 days in traditional 2D cultures. This viability allows for more comprehensive drug sensitivity testing and demonstrates the importance of stromal components [117]. In AML, recently introduced human BM perivascular organoids have demonstrated success in long-term cultures and HTS capabilities [98].
The aforementioned lymphomas, which preserve key histological and molecular features, also support HTS applications. These lymphomoids not only retain tumor architecture but also enable individualized testing of therapeutic agents [115].
Similarly, hematopoietic organoids formed via stepwise induction and gelatin-methacryloyl (GelMA) scaffolding have been used as platforms to evaluate therapeutic responses [118]. Platforms such as MagIC-PDX further advance this field by enabling the efficient creation of drug-resistant hematological malignancy organoids [110,117].
5.4. Toward Immunotherapy Testing Platforms
Immunotherapies, including CAR-T cells, bispecific antibodies, and NK cell-based approaches, can also be evaluated using organoid-based platforms that allow the analysis of patient-specific immune responses in a controlled ex vivo setting [49,119,120].
A 2025 study introduced patient-derived lymphoma organoids (PDLOs) constructed from follicular lymphoma samples that retained the original TILs and the TME for up to three weeks. When T cell-engaging bispecific antibodies were used to treat these PDLOs, marked B cell killing and T cell activation were observed, providing evidence that autologous immune-tumor interactions can be preserved and evaluated with organoids [119]. In another study, T cells expressing T-cell receptors (TEGs) were shown to infiltrate 3D organoids and eliminate tumor cells more effectively than that achieved in 2D cultures [117].
Beyond T cells, CAR-T therapies have demonstrated clinical success in hematological malignancies, and the integration of PDO systems offers a scalable patient-specific method to test these therapies in vitro [120]. The use of bispecific antibodies also holds promise for treating blood cancers, although the research is underdeveloped thus far. In colorectal cancer, T-cell bispecific antibodies have been used to aid in the dissection of immunotherapy resistance mechanisms [75].
Although standardization challenges remain [121], as these platforms mature and technological advances occur, organoid-based immunotherapy may enable greater preclinical evaluation of patient-specific immunotherapeutic strategies.
Across hematological malignancies, organoid-based models have shown strong promise for recapitulating disease-relevant microenvironmental niches that govern malignant cell survival and, therefore, enable ex vivo functional evaluation of immunotherapies within patient-matched stromal and immune contexts [98,110,114,115]. These capabilities are especially valuable in blood cancers, where the therapeutic response is tightly coupled with BM or lymphoid niche signaling [99].
However, hematological malignancy organoids remain technically complex and time-intensive to create and are still limited in their ability to fully reproduce long-term immune dynamics [98]. Additionally, it remains challenging to standardize and has variable scalability potential across disease subtypes [121,122,123,124]. Overcoming these challenges is critical for translating niche-informed organoid platforms into robust tools for precision medicine.
Organoid platforms are increasingly recognized as versatile functional systems for modeling both solid and hematological malignancies. While organoid models in solid tumors, including head and neck and lung cancers, primarily aim to recapitulate tumor architecture and epithelial heterogeneity, hematological organoids uniquely focus on reconstructing specialized microenvironmental niches, such as the bone marrow and lymphoid tissues, which are critical for sustaining malignant cell survival and therapeutic response [99,123]. Despite these structural and technical distinctions, organoid platforms consistently provide valuable functional insights for drug screening, resistance profiling, and immunotherapy evaluation across diverse disease contexts [122,123]. A comparative summary of organoid applications, strengths, and current limitations across head and neck cancer, lung cancer, and hematological malignancies is presented in
Table 1.
6. Toward Functional Precision Oncology
6.1. Shared Functional Gaps Across Distinct Cancer Types
Through HNCs, lung cancer, and hematological malignancies, this review reveals a shared albeit context-dependent limitations of the current genomics-centered precision oncology: the inability to functionally predict patient-specific therapeutic responses (
Figure 2). In HNCs, the pronounced heterogeneity in radiosensitivity and recurrence risk undermines uniform treatment strategies. In lung cancer, rapid resistance to genotype-matched targeted therapies highlights the persistent discordance between molecular alterations and clinical responses. In hematological malignancies, therapeutic outcomes are critically shaped by specialized microenvironmental niches that cannot be captured by conventional reductionist models.
Although these diseases differ substantially in terms of tissue origin, genetic architecture, and clinical management, they share a common unmet need for functional platforms capable of bridging the gap between molecular profiling and real-world treatment responses.
6.2. Redefining Organoids as Functional Stratification Platforms
By synthesizing disease-specific evidence across these distinct clinical contexts, this review reframes PDOs not merely as experimental disease models but as functional stratification platforms that enable patient-specific therapeutic decision-making. Unlike conventional preclinical systems, organoids directly investigate treatment sensitivity, resistance dynamics, and microenvironment-dependent vulnerability in patient-matched contexts. Through this functional lens, organoid platforms function as an intermediate layer between genomic information and clinical responses, complementing molecular profiling and real-time biological validation. This shift repositions organoids from passive disease representations to active decision-support tools in precision oncology workflows.
6.3. From Genotype-Guided to Response-Driven Precision Medicine
This review addressed the growing divide between the expanding molecular classification of cancer and the persistent unpredictability of treatment responses observed in clinical practice. Although genomic profiling has transformed oncology, genotyping alone is insufficient for capturing the functional complexity of tumor biology. By positioning organoid-based systems at the interface between molecular features and therapeutic outcomes, we propose a conceptual transition from genotype-guided precision medicine to response-driven precision medicine. This paradigm emphasizes functional evidence as a critical determinant of treatment selection and enables adaptive and patient-centered therapeutic strategies.
7. Future Directions and Clinical Translation
Several critical challenges must be addressed for organoid-guided precision oncology to transition from proof-of-concept to routine clinical implementation. Standardization of culture conditions, extracellular matrices, assay readouts, and quality control metrics is essential to ensure reproducibility across institutions. Equally important is the reduction in turnaround time through automation, miniaturization, and integration with microfluidic platforms to enable real-time clinical decision support.
Incorporating immune, stromal, and vascular components into organoid systems is necessary to comprehensively model therapeutic responses, particularly in immunotherapies and microenvironment-dependent cancers. The convergence of organoid platforms with multi-omics profiling and artificial intelligence-driven analytics will further enhance predictive accuracy by enabling data integration, pattern recognition, and patient stratification at scale.
Finally, regulatory harmonization, ethical governance, and equitable access must be considered for organoid-based technologies that move toward clinical adoption. Establishing clinically certified workflows and companion diagnostic frameworks is crucial for integrating functional testing into standard oncology practice. Together, these advances will enable a new generation of adaptive, response-driven, precision oncology strategies that place functional biology at the center of therapeutic decision-making.
Author Contributions
Conceptualization: Hyejoung Kim., SeulBee Lee., Alyssa Kim.; literature review and data curation: Hyejoung Kim., SeulBee Lee., Alyssa Kim.; Writing original draft preparation: Hyejoung Kim., SeulBee Lee., Alyssa Kim.; Writing review and editing: Hyejoung Kim., SeulBee Lee., Alyssa Kim.; Supervision: Hyejoung Kim. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Alchemist Project (Grant No. 1415180884; Project No. 20012378, Development of Meta Soft Organ Module Manufacturing Technology without Immunity Rejection and Module Assembly Robot System) and the Technology Innovation Program (Grant No. RS-2025-02314555), funded by the Ministry of Trade, Industry and Energy (MOTIE, Republic of Korea). This work was also supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (Grant Nos. RS-2025-00514943 and NRF-2022R1I1A1A01063972).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
No new data were created or analyzed in this study. Data sharing is not applicable.
Acknowledgments
The authors thank their collaborators and clinical partners for insightful discussions that contributed to the conceptual development of this review.
Conflicts of Interest
The authors declare no competing interests.
Abbreviations
The following abbreviations are used in this manuscript:
| ALI |
Air–liquid interface |
| ALK |
Anaplastic lymphoma kinase |
| AML |
Acute myeloid leukemia |
| BM |
Bone marrow |
| BMO |
Bone marrow organoid |
| CLL |
Chronic lymphocytic leukemia |
| GelMA |
Gelatin methacryloyl |
| HNC |
Head and neck cancer |
| HNSCC |
Head and neck squamous cell carcinoma |
| HPV |
Human papillomavirus |
| HPSC |
Hematopoietic stem and progenitor cell |
| HTS |
High-throughput screening |
| ICI |
Immune checkpoint inhibitor |
| iPSC |
Induced pluripotent stem cell |
| MDT |
Multidisciplinary treatment |
| MM |
Multiple myeloma |
| NK |
Natural killer |
| NSCLC |
Non-small cell lung cancer |
| PD-1 |
Programmed cell death protein 1 |
| TAT |
Turnaround time |
| TME |
Tumor microenvironment |
| TIL |
Tumor infiltrating lymphocyte |
| PDX |
Patient-derived xenograft |
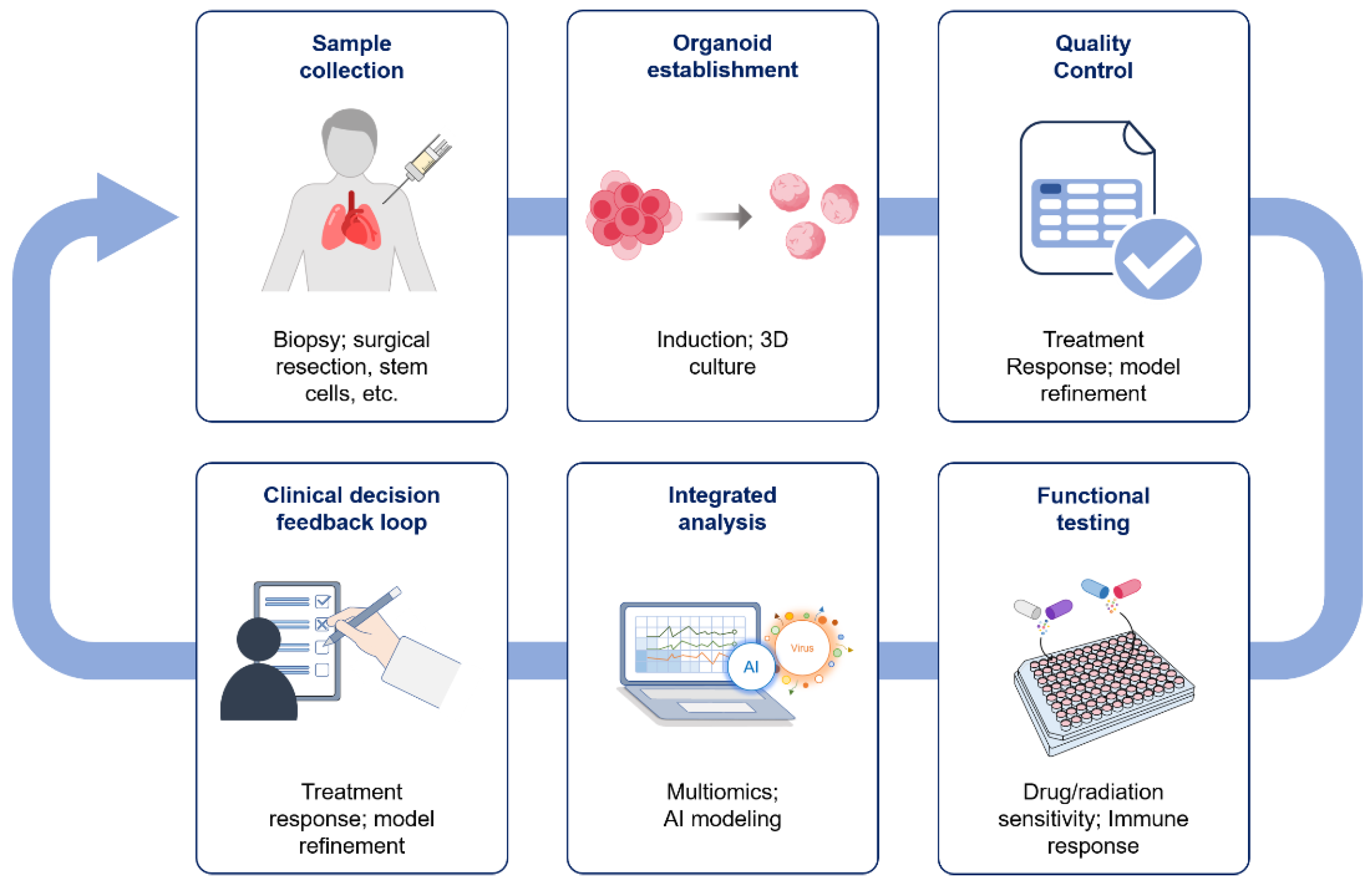
Figure 1. Workflow of organoid-based precision oncology.
Schematic illustration of the workflow for organoid-based precision oncology. Patient-derived samples obtained from biopsies or surgical resections are used to establish tumor organoids through induction and three-dimensional culture. Established organoids undergo quality control to validate genotypic and phenotypic fidelity to the parental tumors. Functional testing, including drug response, radiation sensitivity, and immune responsiveness, is performed, followed by integrated analysis using multi-omics approaches and computational modeling. The resulting functional data are fed back into clinical decision-making to refine treatment strategies and improve predictive accuracy.
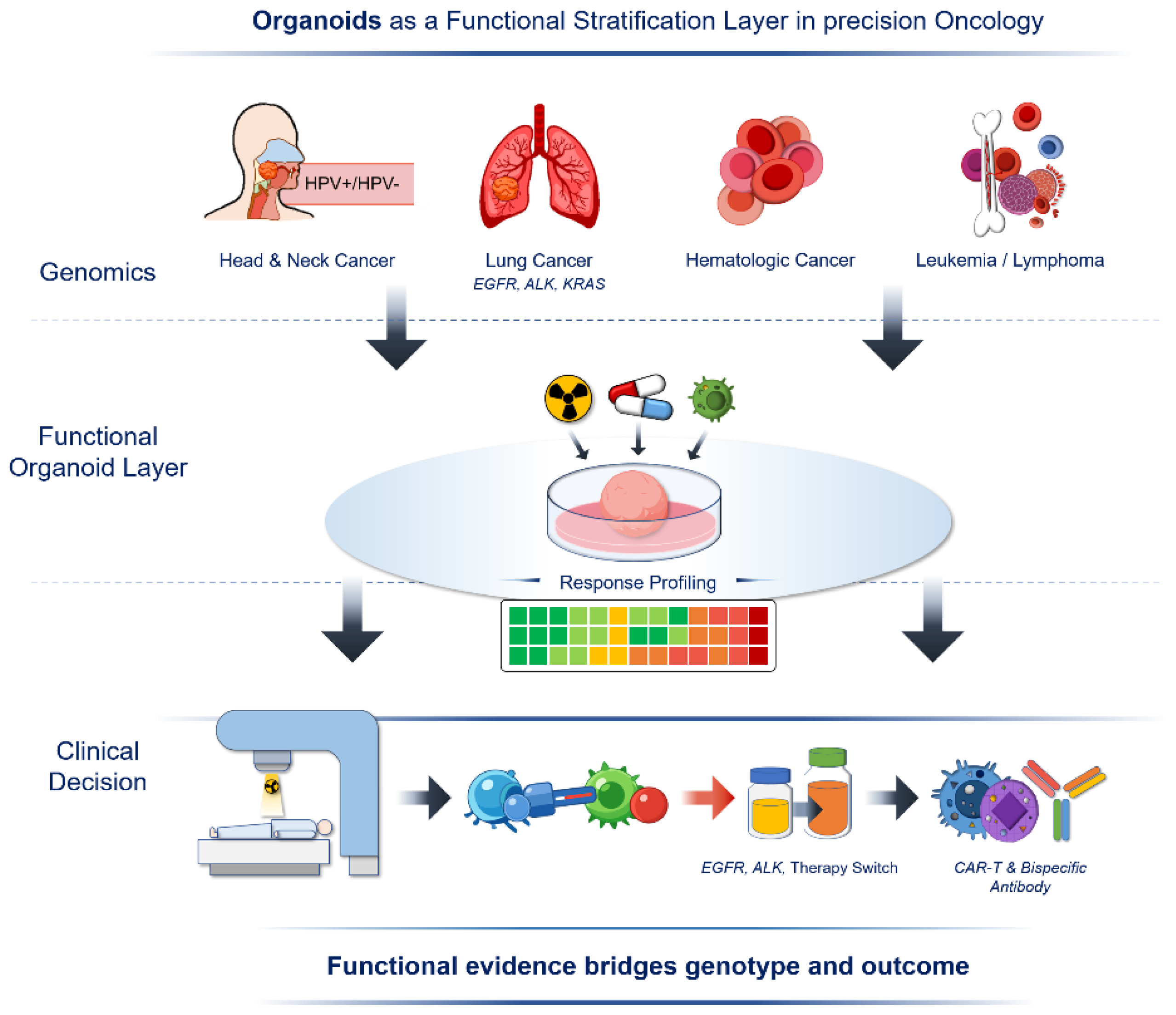
Figure 2. Organoids as a functional stratification layer in precision oncology.
Schematic illustration of organoids as a functional stratification layer bridging genomic information and clinical outcomes in precision oncology. Across multiple cancer types, including head and neck cancer, lung cancer, and hematological malignancies, genomic profiling alone is often insufficient to predict therapeutic response. Patient-derived organoids enable functional response profiling to targeted therapies, radiotherapy, and immunotherapies, generating quantitative sensitivity patterns. These functional readouts complement molecular profiling and support informed clinical decision-making, including therapy selection, treatment switching, and advanced immunotherapeutic strategies such as CAR-T cells and bispecific antibodies.
Table 1. Cross-disease comparison of organoids in precision medicine readiness.
Overview of organoid-based platforms across head and neck cancer, lung cancer, and hematological malignancies. The table summarizes key parameters, including sample sources, establishment success rates, turnaround times, functional readouts, feasibility of immune modeling, levels of clinical evidence, and major limitations. This comparison highlights disease-specific strengths and challenges of organoid models in supporting precision medicine applications.
References
- Dagogo-Jack, I., Shaw, A.T., 2018. Tumour heterogeneity and resistance to cancer therapies. Nat. Rev. Clin. Oncol. 15 (2), 81–94. [CrossRef]
- Letai, A., 2017. Functional precision cancer medicine—moving beyond pure genomics. Nat. Med. 23 (9), 1028–1035. [CrossRef]
- Collins, F.S., Varmus, H., 2015. A new initiative on precision medicine. N. Engl. J. Med. 372 (9), 793–795. [CrossRef]
- Tsimberidou, A.M., et al., 2020. Precision oncology: lessons learned and future directions. Nat. Rev. Clin. Oncol. 17 (11), 676–690. [CrossRef]
- Tannock, I.F., Hickman, J.A., 2016. Limits to personalized cancer medicine. N. Engl. J. Med. 375 (13), 1289–1294. [CrossRef]
- Hyman, D.M., Taylor, B.S., Baselga, J., 2017. Implementing genome-driven oncology. Cell 168 (4), 584–599. [CrossRef]
- McGranahan, N., Swanton, C., 2017. Clonal heterogeneity and tumor evolution. Cell 168 (4), 613–628. [CrossRef]
- Quail, D.F., Joyce, J.A., 2013. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19 (11), 1423–1437. [CrossRef]
- Ribas, A., Wolchok, J.D., 2018. Cancer immunotherapy using checkpoint blockade. Science 359 (6382), 1350–1355. [CrossRef]
- Ben-David, U., et al., 2018. Genetic and transcriptional evolution alters cancer cell line drug response. Nature 560 (7718), 325–330. [CrossRef]
- Breslin, S., O’Driscoll, L., 2013. Three-dimensional cell culture: the missing link in drug discovery. Drug Discov. Today 18 (5–6), 240–249. [CrossRef]
- Byrne, A.T., et al., 2017. Interrogating open issues in cancer precision medicine with patient-derived xenografts. Nat. Rev. Cancer 17 (4), 254–268. [CrossRef]
- Clevers, H., 2016. Modeling development and disease with organoids. Cell 165 (7), 1586–1597. [CrossRef]
- Vlachogiannis, G., et al., 2018. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science 359 (6378), 920–926. [CrossRef]
- An, L., Liu, Y., Liu, Y., 2025. Organ-on-a-chip applications in microfluidic platforms. Micromachines 16 (2), 201. [CrossRef]
- Kimura, H., Nishikawa, M., Kutsuzawa, N., et al., 2025. Advancements in microphysiological systems: exploring organoids and organ-on-a-chip technologies in drug development with a focus on pharmacokinetics-related organs. Drug Metab. Pharmacokinet. 60, 101046. [CrossRef]
- Wang, J., Tao, X., Zhu, J., et al., 2025. Tumor organoid-immune co-culture models: exploring a new perspective of tumor immunity. Cell Death Discov. 11 (1), 195. [CrossRef]
- Leemans, C.R., Snijders, P.J.F., Brakenhoff, R.H., 2018. The molecular landscape of head and neck cancer. Nat. Rev. Cancer 18 (5), 269–282. [CrossRef]
- Marur, S., Forastiere, A.A., 2016. Head and neck squamous cell carcinoma: update on epidemiology, diagnosis, and treatment. Mayo Clin. Proc. 91 (3), 386–396. [CrossRef]
- Yu, H.A., et al., 2013. Analysis of tumor specimens at the time of acquired resistance to EGFR-TKI therapy. Clin. Cancer Res. 19 (8), 2240–2247. [CrossRef]
- Morrison, S.J., Scadden, D.T., 2014. The bone marrow niche for haematopoietic stem cells. Nature 505 (7483), 327–334. [CrossRef]
- Tiriac, H., et al., 2018. Organoid profiling identifies common responders to chemotherapy in pancreatic cancer. Cancer Discov. 8 (9), 1112–1129. [CrossRef]
- Ooft, S.N., et al., 2019. Patient-derived organoids can predict response to chemotherapy in metastatic colorectal cancer. Sci. Transl. Med. 11 (513), eaay2574. [CrossRef]
- Johnson, D.E., et al., 2020. Head and neck squamous cell carcinoma. Nat. Rev. Dis. Primers 6 (1), 92. [CrossRef]
- Ang, K.K., Harris, J., Wheeler, R., Weber, R., Rosenthal, D.I., Nguyen-Tân, P.F., Westra, W.H., et al., 2010. Human papillomavirus and survival of patients with oropharyngeal cancer. N. Engl. J. Med. 363 (1), 24–35. [CrossRef]
- Millen, R., de Kort, W.W.B., Koomen, M., et al., 2023. Patient-derived head and neck cancer organoids allow treatment stratification and serve as a tool for biomarker validation and identification. Med 4 (5), 290–310.e12. [CrossRef]
- Pignon, J.P., le Maître, A., Maillard, E., Bourhis, J., et al., 2009. Meta-analysis of chemotherapy in head and neck cancer (MACH-NC): An update on 93 randomised trials and 17,346 patients. Radiother. Oncol. 92 (1), 4–14. [CrossRef]
- Vokes, E.E., Weichselbaum, R.R., Lippman, S.M., Hong, W.K., 1993. Head and neck cancer. N. Engl. J. Med. 328 (3), 184–194. [CrossRef]
- Yang, H.T., Shah, R.H., Tegay, D., Onel, K., 2019. Precision oncology: Lessons learned and challenges for the future. Cancer Manag. Res. 11, 7525–7536. [CrossRef]
-
Drost, J., Clevers, H., 2018. Organoids in cancer research. Nat. Rev. Cancer 18, 407–418. [CrossRef]
- Lechner, M., et al., 2022. HPV-associated oropharyngeal cancer: epidemiology, molecular biology and clinical management. Nat. Rev. Clin. Oncol. 19 (5), 306–327. [CrossRef]
- Mirghani, H., Blanchard, P., 2018. Treatment de-escalation for HPV-driven oropharyngeal cancer: where do we stand? Lancet Oncol. 19 (5), e195–e196. [CrossRef]
- Driehuis, E., Kolders, S., Spelier, S., et al., 2019. Oral mucosal organoids as a potential platform for personalized cancer therapy. Cancer Discov. 9 (7), 852–871. [CrossRef]
- Fisch, A.-S., Unger, S., Schulte, C., et al., 2024. Feasibility analysis of using patient-derived tumour organoids for treatment decision guidance in locally advanced head and neck squamous cell carcinoma. Eur. J. Cancer 213, 115100. [CrossRef]
- Moody, C.A., Laimins, L.A., 2010. Human papillomavirus oncoproteins: Pathways to transformation. Nat. Rev. Cancer 10 (8), 550–560. [CrossRef]
- The Cancer Genome Atlas Network., 2015. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517, 576–582. [CrossRef]
- Mandal, R., Şenbabaoğlu, Y., Desrichard, A., Havel, J.J., Dalin, M.G., Riaz, N., Lee, K.W., Ganly, I., Hakimi, A.A., Chan, T.A., Morris, L.G., 2016. The head and neck cancer immune landscape and its immunotherapeutic implications. JCI Insight 1 (17), e89829. [CrossRef]
- Issing, C., et al., 2025. Head and neck tumor organoid biobank for modeling individual responses to radiotherapy according to TP53/HPV status. J. Exp. Clin. Cancer Res. 44, 85. [CrossRef]
- Kimple, R.J., Smith, M.A., Blitzer, G.C., Torres, A.D., Martin, J.A., Yang, R.Z., Peet, C.R., Lorenz, L.D., Nickel, K.P., Klingelhutz, A.J., Lambert, P.F., Harari, P.M., 2013. Enhanced radiation sensitivity in HPV-positive head and neck cancer. Cancer Res. 73 (15), 4791–4800. [CrossRef]
- Facompré, N.D., et al., 2020. Identifying predictors of HPV-related head and neck cancer progression using patient-derived models. Int. J. Cancer 147 (11), 3236–3249. [CrossRef]
- Haughton, P.D., et al., 2024. Differential transcriptional invasion signatures from patient-derived organoids define a prognostic tool in head and neck cancer. Oncogene 43 (11), 2463–2474. [CrossRef]
- Mehanna, H., et al., 2019. Radiotherapy plus cisplatin or cetuximab in low-risk HPV-positive oropharyngeal cancer (De-ESCALaTE HPV). Lancet 393 (10166), 51–60. [CrossRef]
- Li, Q., et al., 2023. Targeted therapy for head and neck cancer: signaling pathways and clinical studies. Sig. Transduct. Target. Ther. 8 (1), 31. [CrossRef]
- Soulières, D., et al., 2017. Buparlisib plus weekly paclitaxel versus placebo plus weekly paclitaxel for recurrent/metastatic head and neck squamous cell carcinoma (BERIL-1): a randomized, double-blind phase II trial. Lancet Oncol. 18 (3), 321–330. [CrossRef]
- Harrington, K.J., Ferris, R.L., Blumenschein, G., Jr., Colevas, A.D., Fayette, J., Licitra, L., Kasper, S., Even, C., Vokes, E.E., Worden, F., Saba, N.F., Kiyota, N., Haddad, R., Tahara, M., Grünwald, V., Shaw, J.W., Monga, M., Lynch, M., Taylor, F., DeRosa, M., et al., 2017. Nivolumab versus standard, single-agent therapy of investigator’s choice in recurrent or metastatic squamous cell carcinoma of the head and neck (CheckMate 141): health-related quality-of-life results from a randomised, phase 3 trial. Lancet Oncol. 18 (8), 1104–1115. [CrossRef]
- Burtness, B., et al., 2019. Pembrolizumab versus cetuximab in recurrent or metastatic head-and-neck cancer (KEYNOTE-048). Lancet 394 (10212), 1915–1928. [CrossRef]
- Cristescu, R., et al., 2018. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science 362 (6411), eaar3593. [CrossRef]
- Voabil, P., et al., 2021. An ex vivo platform to dissect tumor response to PD-1 blockade in MSI colorectal cancer. Nat. Med. 27 (8), 1410–1416. [CrossRef]
- Neal, J.T., Li, X., Zhu, J., et al., 2018. Organoid modeling of the tumor immune microenvironment. Cell 175 (7), 1972–1988.e16. [CrossRef]
- Dijkstra, K.K., et al., 2018. Generation of tumor reactive T cells by co-culture of peripheral blood lymphocytes and tumor organoids. Cell 174 (6), 1586–1598. [CrossRef]
- Chalabi, M., et al., 2020. Neoadjuvant immunotherapy leads to pathological responses in MMR-deficient and MMR-proficient early-stage colon cancers (NICHE study). Nat. Med. 26 (4), 566–576. [CrossRef]
- Yuki, K., et al., 2020. Organoid models of tumor immunology. Trends Immunol. 41 (8), 652–664. [CrossRef]
- Meng, R., et al., 2025. Head and neck cancer organoids: a panoramic perspective from basic construction to clinical translation. Stem Cell Res. Ther. 16 (1), 534. [CrossRef]
- Wang, J., et al., 2022. Patient-derived tumor organoids: new progress and opportunities for cancer therapy. Front. Oncol. 12, 872531. [CrossRef]
- Ganesh, K., et al., 2019. A rectal cancer organoid platform to study individual responses to chemoradiation. Nat. Med. 25, 1607–1614. [CrossRef]
- Yao, Y., et al., 2020. Patient-derived organoids predict chemoradiation responses of locally advanced rectal cancer. Nat. Med. 26, 1275–1284. [CrossRef]
- Bleijs, M., et al., 2019. Xenograft and organoid model systems in cancer research. EMBO J. 38 (15), e101654. [CrossRef]
- Veninga, V., et al., 2021. Tumor organoids: opportunities and challenges to guide precision medicine. Cancer Cell 39 (9), 1190–1201. [CrossRef]
- Schutgens, F., Clevers, H., 2020. Human organoids: Tools for understanding biology and treating diseases. Cell 182 (5), 1036–1049. [CrossRef]
- Bray, F., Laversanne, M., Sung, H., Ferlay, J., Siegel, R.L., Soerjomataram, I., Jemal, A., 2024. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74 (3), 229–263. [CrossRef]
- Lim, Z.F., Ma, P.C., 2019. Emerging insights of tumor heterogeneity and drug resistance mechanisms in lung cancer targeted therapy. J. Hematol. Oncol. 12 (1), 134. [CrossRef]
- Hu, Y., Sui, X., Song, F., et al., 2021. Lung cancer organoids analyzed on microwell arrays predict drug responses of patients within a week. Nat. Commun. 12, 2581. [CrossRef]
- Wang, D., et al., 2017. Molecular heterogeneity of non-small cell lung carcinoma patient-derived xenografts closely reflect their primary tumors. Int. J. Cancer 140 (3), 662–673. [CrossRef]
- Abdolahi, S., et al., 2022. Patient-derived xenograft (PDX) models, applications and challenges in cancer research. J. Transl. Med. 20 (1), 206. [CrossRef]
- Yoshida, G.J., 2020. Applications of patient-derived tumor xenograft models and tumor organoids. J. Hematol. Oncol. 13 (1), 4. [CrossRef]
- Gao, H., et al., 2015. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat. Med. 21 (11), 1318–1325. [CrossRef]
- Wang, H.M., et al., 2023. Using patient-derived organoids to predict locally advanced or metastatic lung cancer tumor response: A real-world study. Cell Rep. Med. 4 (2), 100911. [CrossRef]
- Shi, R., Radulovich, N., et al., 2020. Organoid cultures as preclinical models of non-small cell lung cancer. Clin. Cancer Res. 26 (5), 1162–1174. [CrossRef]
- Kim, M., et al., 2019. Patient-derived lung cancer organoids as in vitro cancer models for therapeutic screening. Nat. Commun. 10 (1), 3991. [CrossRef]
- Dost, A.F.M., Moye, A.L., Vedaie, M., Tran, L.M., Fung, E., Heinze, D., et al., 2020. Organoids model transcriptional hallmarks of oncogenic KRAS activation in lung epithelial progenitor cells. Cell Stem Cell 27 (4), 663–678.e8. [CrossRef]
- Kim, S.Y., Kim, S.M., Lim, S., Lee, J.Y., Choi, S.J., Yang, S.D., Yun, M.R., Kim, C.G., Gu, S.R., Park, C., Park, A.Y., Lim, S.M., Heo, S.G., Kim, H., Cho, B.C., 2021. Modeling clinical responses to targeted therapies by patient-derived organoids of advanced lung adenocarcinoma. Clin. Cancer Res. 27 (15), 4397–4409. [CrossRef]
- Bar-Ephraim, Y., Kretzschmar, K., Clevers, H., 2020. Organoids in immunological research. Nat. Rev. Immunol. 20, 279–293. [CrossRef]
- Roerink, S.F., Sasaki, N., Lee-Six, H., et al., 2018. Intra-tumour diversification in colorectal cancer at the single-cell level. Nature 556, 457–462. [CrossRef]
- Meng, Y., et al., 2025. Lung cancer organoids: a new strategy for precision medicine research. Transl. Lung Cancer Res. 14 (2). [CrossRef]
- Grönholm, M., et al., 2021. Patient-derived organoids for precision cancer immunotherapy. Cancer Res. 81 (12), 3149–3155. [CrossRef]
- Liu, Y., Lankadasari, M., Rosiene, J., Johnson, K.E., Zhou, J., Bapat, S., Chow-Tsang, L.L., Tian, H., Mastrogiacomo, B., He, D., Connolly, J.G., Lengel, H.B., Caso, R., Dunne, E.G., Fick, C.N., Rocco, G., Sihag, S., Isbell, J.M., Bott, M.J., Li, B.T., et al., 2024. Modeling lung adenocarcinoma metastases using patient-derived organoids. Cell Rep. Med. 5 (10), 101777. [CrossRef]
- Oh, S.Y., Park, S., Lee, S., Lee, E.J., Kim, T.H., Choi, S.J., Park, S.Y., Kim, J.H., Lim, S.M., Lee, J.B., Cho, B.C., Hong, M.H., Yun, M.R., 2025. The potential of lazertinib and amivantamab combination therapy as a treatment strategy for uncommon EGFR-mutated NSCLC. Cell Rep. Med. 6 (2), 101929. [CrossRef]
- Kim, J., Minna, J.D., 2023. Moving toward precision medicine with lung cancer organoids. Cell Rep. Med. 4 (2), 100952. [CrossRef]
- Lee, S.H., Kim, K., Lee, E., Lee, K., Ahn, K.H., Park, H., Kim, Y., Shin, S., Jeon, S.Y., Hwang, Y., Ahn, D.H., Kwon, Y.J., Moon, S.W., Moon, M.H., Kim, K.S., Hyun, K., Kim, T.J., Sung, Y.E., Choi, J.Y., Park, C.K., Kim, S.J., 2024. Prediction of TKI response in EGFR-mutant lung cancer patients-derived organoids using malignant pleural effusion. NPJ Precis. Oncol. 8 (1), 111. [CrossRef]
- Takahashi, N., Hoshi, H., Higa, A., Hiyama, G., Tamura, H., Ogawa, M., Takagi, K., Goda, K., Okabe, N., Muto, S., Suzuki, H., Shimomura, K., Watanabe, S., Takagi, M., 2019. An in vitro system for evaluating molecular targeted drugs using lung patient-derived tumor organoids. Cells 8 (5), 481. [CrossRef]
- Lee, S.Y., Cho, H.J., Choi, J., Ku, B., Moon, S.W., Moon, M.H., Kim, K.S., Hyun, K., Kim, T.J., Sung, Y.E., Hwang, Y., Lee, E., Ahn, D.H., Choi, J.Y., Lim, J.U., Park, C.K., Kim, S.W., Kim, S.J., Koo, I.S., Jung, W.S., Lee, D.W., 2023. Cancer organoid-based diagnosis reactivity prediction (CODRP) index-based anti-cancer drug sensitivity test in ALK-rearrangement positive non-small cell lung cancer (NSCLC). J. Exp. Clin. Cancer Res. 42 (1), 309. [CrossRef]
- Xie, Y., Zhang, Y., Wu, Y., Xie, X., Lin, X., Tang, Q., Zhou, C., Xie, Z., 2024. Analysis of the resistance profile of real-world alectinib first-line therapy in ALK rearrangement-positive advanced non-small cell lung cancer using organoid technology: a case report. J. Thorac. Dis. 16 (6), 3854–3863. [CrossRef]
- Bian, Y., Shan, G., Bi, G., Liang, J., Hu, Z., Sui, Q., Shi, H., Zheng, Z., Yao, G., Wang, Q., Fan, H., Zhan, C., 2024. Targeting ALDH1A1 to enhance the efficacy of KRAS-targeted therapy through ferroptosis. Redox Biol. 77, 103361. [CrossRef]
- Jeong, S.R., Kang, M., 2023. Exploring tumor immune interactions in co-culture models of T cells and tumor organoids derived from patients. Int. J. Mol. Sci. 24 (19), 14609. [CrossRef]
- Schoenfeld, A.J., Hellmann, M.D., 2020. Acquired resistance to immune checkpoint inhibitors. Cancer Cell 37 (4), 443–455. [CrossRef]
- Magré, L., Verstegen, M.M.A., Buschow, S., van der Laan, L.J.W., Peppelenbosch, M., Desai, J., 2023. Emerging organoid-immune co-culture models for cancer research: from oncoimmunology to personalized immunotherapies. J. Immunother. Cancer 11 (5), e006290. [CrossRef]
- Chen, D., Xu, L., Xuan, M., Chu, Q., Xue, C., 2024. Unveiling the functional roles of patient-derived tumour organoids in assessing the tumour microenvironment and immunotherapy. Clin. Transl. Med. 14 (9), e1802. [CrossRef]
- Wang, Q., Yuan, F., Zuo, X., Li, M., 2025. Breakthroughs and challenges of organoid models for assessing cancer immunotherapy: a cutting-edge tool for advancing personalised treatments. Cell Death Discov. 11 (1), 222. [CrossRef]
- Zhu, Z., Shen, J., Ho, P.C., Hu, Y., Ma, Z., Wang, L., 2025. Transforming cancer treatment: Integrating patient-derived organoids and CRISPR screening for precision medicine. Front. Pharmacol. 16, 1563198. [CrossRef]
- Park, T., Kim, T.K., Han, Y.D., Kim, K.A., Kim, H., Kim, H.S., 2023. Development of a deep learning-based image processing tool for enhanced organoid analysis. Sci. Rep. 13 (1), 19841. [CrossRef]
- Bian, X., Li, G., Wang, C., Liu, W., Lin, X., Chen, Z., Cheung, M., Luo, X., 2021. A deep learning model for detection and tracking in high-throughput images of organoid. Comput. Biol. Med. 134, 104490. [CrossRef]
- Wang, H., Ning, X., Zhao, F., Zhao, H., Li, D., 2024. Human organoids-on-chips for biomedical research and applications. Theranostics 14 (2), 788–818. [CrossRef]
- Zhu, L., Zhang, J., Guo, Q., Kuang, J., Li, D., Wu, M., Mo, Y., Zhang, T., Gao, X., Tan, J., 2023. Advanced lung organoids and lung-on-a-chip for cancer research and drug evaluation: A review. Front. Bioeng. Biotechnol. 11, 1299033. [CrossRef]
- Zeng, X., Ma, Q., Li, X.K., You, L.T., Li, J., Fu, X., You, F.M., Ren, Y.F., 2023. Patient-derived organoids of lung cancer based on organoids-on-a-chip: Enhancing clinical and translational applications. Front. Bioeng. Biotechnol. 11, 1205157. [CrossRef]
- Tian, H., Ren, J., Mou, R., Jia, Y., 2023. Application of organoids in precision immunotherapy of lung cancer (Review). Oncol. Lett. 26 (5), 484. [CrossRef]
- Liu, C., Shi, C., Wang, S., Qi, R., Gu, W., Yu, F., Zhang, G., Qiu, F., 2025. Bridging the gap: How patient-derived lung cancer organoids are transforming personalized medicine. Front. Cell Dev. Biol. 13, 1554268. [CrossRef]
- Mattioda, C., Voena, C., Ciardelli, G., Mattu, C., 2025. In vitro 3D models of haematological malignancies: Current trends and the road ahead? Cells 14 (1), 38. [CrossRef]
- Méndez-Ferrer, S., Bonnet, D., Steensma, D.P., Hasserjian, R.P., Ghobrial, I.M., Gribben, J.G., Andreeff, M., Krause, D.S., 2020. Bone marrow niches in haematological malignancies. Nat. Rev. Cancer 20 (5), 285–298. [CrossRef]
- Kokkaliaris, K.D., Scadden, D.T., 2020. Cell interactions in the bone marrow microenvironment affecting myeloid malignancies. Blood Adv. 4 (15), 3795–3803. [CrossRef]
- Khan, A.O., et al., 2023. Human bone marrow organoids for disease modeling, discovery, and validation of therapeutic targets in hematologic malignancies. Cancer Discov. 13 (2), 364–385. [CrossRef]
- Kim, J., Koo, B.K., Knoblich, J.A., 2020. Human organoids: model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol. 21 (10), 571–584. [CrossRef]
- Torisawa, Y.-S., Spina, C.S., Mammoto, T., et al., 2014. Bone marrow–on–a–chip replicates hematopoietic niche physiology in vitro. Nat. Methods 11, 663–669. [CrossRef]
- Bessy, T., Itkin, T., Passaro, D., 2021. Bioengineering the bone marrow vascular niche. Front. Cell Dev. Biol. 9, 645496. [CrossRef]
- De Janon, A., et al., 2023. Three-dimensional human bone marrow organoids for the study and application of normal and abnormal hematoimmunopoiesis. J. Immunol. 210 (7), 895–904. [CrossRef]
- Olijnik, A., et al., 2024. Generating human bone marrow organoids for disease modeling and drug discovery. Nat. Protoc. 19, 2117–2146. [CrossRef]
- Frenz-Wiessner, S., et al., 2024. Generation of complex bone marrow organoids from human induced pluripotent stem cells. Nat. Methods 21 (5), 868–881. [CrossRef]
- Ren, K., et al., 2025. Development of iPSC-derived human bone marrow organoid for autonomous hematopoiesis and patient-derived HSPC engraftment. Blood Adv. 9 (1), 54–65. [CrossRef]
- Zheng, H., Chen, Y., Luo, Q., Zhang, J., Huang, M., Xu, Y., Huo, D., Shan, W., Tie, R., Zhang, M., Qian, P., Huang, H., 2023. Generating hematopoietic cells from human pluripotent stem cells: Approaches, progress and challenges. Cell Regen. 12 (1), 31. [CrossRef]
- Cerneckis, J., Cai, H., Shi, Y., 2024. Induced pluripotent stem cells (iPSCs): Molecular mechanisms of induction and applications. Sig. Transduct. Target. Ther. 9, 112. [CrossRef]
- Wang, L., Shi, M., Sung, A.Y., Yin, C.C., Bai, Y., Chen, M., 2025. Role of the bone marrow microenvironment in multiple myeloma: Impact of niches on drug resistance mechanisms. Semin. Diagn. Pathol. [CrossRef]
- Forster, S., Radpour, R., 2022. Molecular impact of the tumor microenvironment on multiple myeloma dissemination and extramedullary disease. Front. Oncol. 12, 941437. [CrossRef]
- Li, Q., Nikolova, M.T., Zhang, G., Cervenka, I., Valigi, F., Burri, D., Plantier, E., Mazzoleni, A., Lamouline, A., Schwaller, J., Treutlein, B., Martin, I., García-García, A., 2025. Macro-scale, scaffold-assisted model of the human bone marrow endosteal niche using hiPSC-vascularized osteoblastic organoids. Cell Stem Cell 32 (12), 1941–1958.e8. [CrossRef]
- Koenig, L., Juglair, L., Tao, T.P., et al., 2025. A microfluidic bone marrow chip for the safety profiling of biologics in pre-clinical drug development. Commun. Biol. 8, 754. [CrossRef]
- Purwada, A., Jaiswal, M.K., Ahn, H., Nojima, T., Kitamura, D., Gaharwar, A.K., Cerchietti, L., Singh, A., 2015. Ex vivo engineered immune organoids for controlled germinal center reactions. Biomaterials 63, 24–34. [CrossRef]
- Santamaria-Martínez, A., Epiney, J., Srivastava, D., et al., 2024. Development of patient-derived lymphomoids with preserved tumor architecture for lymphoma therapy screening. Nat. Commun. 15, 10650. [CrossRef]
- Suhito, I.R., Sunil, C., Tay, A., 2024. Engineering human immune organoids for translational immunology. Bioact. Mater. 44, 164–183. [CrossRef]
- Lourenço, D., 2022. Patient-derived multiple myeloma 3D models for personalized medicine—are we there yet? International Journal of Molecular Sciences 23 (21), 12888. [CrossRef]
- Chen, K., et al., 2024. Establishment of human hematopoietic organoids for evaluation of hematopoietic injury and regeneration effect. Stem Cell Res. Ther. 15, 133. [CrossRef]
- Kastenschmidt, J.M., et al., 2024. A human lymphoma organoid model for evaluating and targeting the follicular lymphoma tumor immune microenvironment. Cell Stem Cell 31 (3), 410–420.e4. [CrossRef]
- Park, J.H., et al., 2018. Long-term follow-up of CD19 CAR therapy in acute lymphoblastic leukemia. N. Engl. J. Med. 378 (5), 449–459. [CrossRef]
- Ybarra, M., Lee, J., Chen, Y.Y., et al., 2025. Human preclinical multiple myeloma in vitro models for disease modeling and therapy screening. J. Biol. Eng. 19, 98. [CrossRef]
- Turhan, A., Foudi, A., Hwang, J.W., Desterke, C., Griscelli, F., Bennaceur-Griscelli, A., 2019. Modeling malignancies using induced pluripotent stem cells: From chronic myeloid leukemia to hereditary cancers. Exp. Hematol. 71, 61–67. [CrossRef]
- Silvestri, G., Chatterjee, A., 2025. Rebuilding the marrow in vitro: Translational advances in the 3D modeling of blood cancers. Onco 5 (4), 51. [CrossRef]
- Andrews, M.G., Kriegstein, A.R., 2022. Challenges of organoid research. Annu. Rev. Neurosci. 45, 23–39. [CrossRef]
- Volpini, L., Monaco, F., Casalone, E., Herman, E.J., Filomena, R., Manzotti, S., Strogovets, O., Moroncini, G., Mozzicafreddo, M., Poloni, A., Marinelli Busilacchi, E., Gonnelli, F., Barbisan, F., Goteri, G., Zuccatosta, L., Bonifazi, M., Matullo, G., Neuzil, J., Santarelli, L., Rahman, N.M., Blyth, K., Tomasetti, M., Mei, F., 2025. Use of patient-derived organoids for pleural mesothelioma 3-D modeling. Am. J. Physiol. Lung Cell Mol. Physiol. 329 (6), L756–L769. [CrossRef]
- Ekanger, C.T., Ramnefjell, M.P., Guttormsen, M.S.F., Hekland, J., Dahl-Michelsen, K., Lotsberg, M.L., Lu, N., Stuhr, L.E.B., Hoareau, L., Salminen, P.R., Gärtner, F., Aanerud, M., Akslen, L.A., Lorens, J.B., Engelsen, A.S.T., 2025. An organoid model for translational cancer research recapitulates histoarchitecture and molecular hallmarks of non-small-cell lung cancer. Cancers (Basel) 17 (11), 1873. [CrossRef]
- Finnberg, N.K., Gokare, P., Lev, A., Grivennikov, S.I., MacFarlane, A.W. 4th, Campbell, K.S., Winters, R.M., Kaputa, K., Farma, J.M., Abbas, A.E., Grasso, L., Nicolaides, N.C., El-Deiry, W.S., 2017. Application of 3D tumoroid systems to define immune and cytotoxic therapeutic responses based on tumoroid and tissue slice culture molecular signatures. Oncotarget 8 (40), 66747–66757. [CrossRef]
- Bottaro, A., Nasso, M.E., Stagno, F., Fazio, M., Allegra, A., 2025. Modeling the bone marrow niche in multiple myeloma: From 2D cultures to 3D systems. Int. J. Mol. Sci. 26 (13), 6229. [CrossRef]
- Rodriguez, C., et al., 2019. High throughput 3D bioprinting of patient-derived multiple myeloma organoid models for niche recapitulation and chemosensitivity assessment. Clin. Lymphoma Myeloma Leuk. 19 (10). [CrossRef]
- de la Puente, P., Muz, B., Gilson, R.C., Azab, F., Luderer, M., King, J., Achilefu, S., Vij, R., Azab, A.K., 2015. 3D tissue-engineered bone marrow as a novel model to study pathophysiology and drug resistance in multiple myeloma. Biomaterials 73, 70–84. [CrossRef]
- Raje, N., et al., 2019. Anti-BCMA CAR T-cell therapy bb2121 in relapsed or refractory multiple myeloma. N. Engl. J. Med. 380 (18), 1726–1737. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).