Submitted:
14 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
Introduction
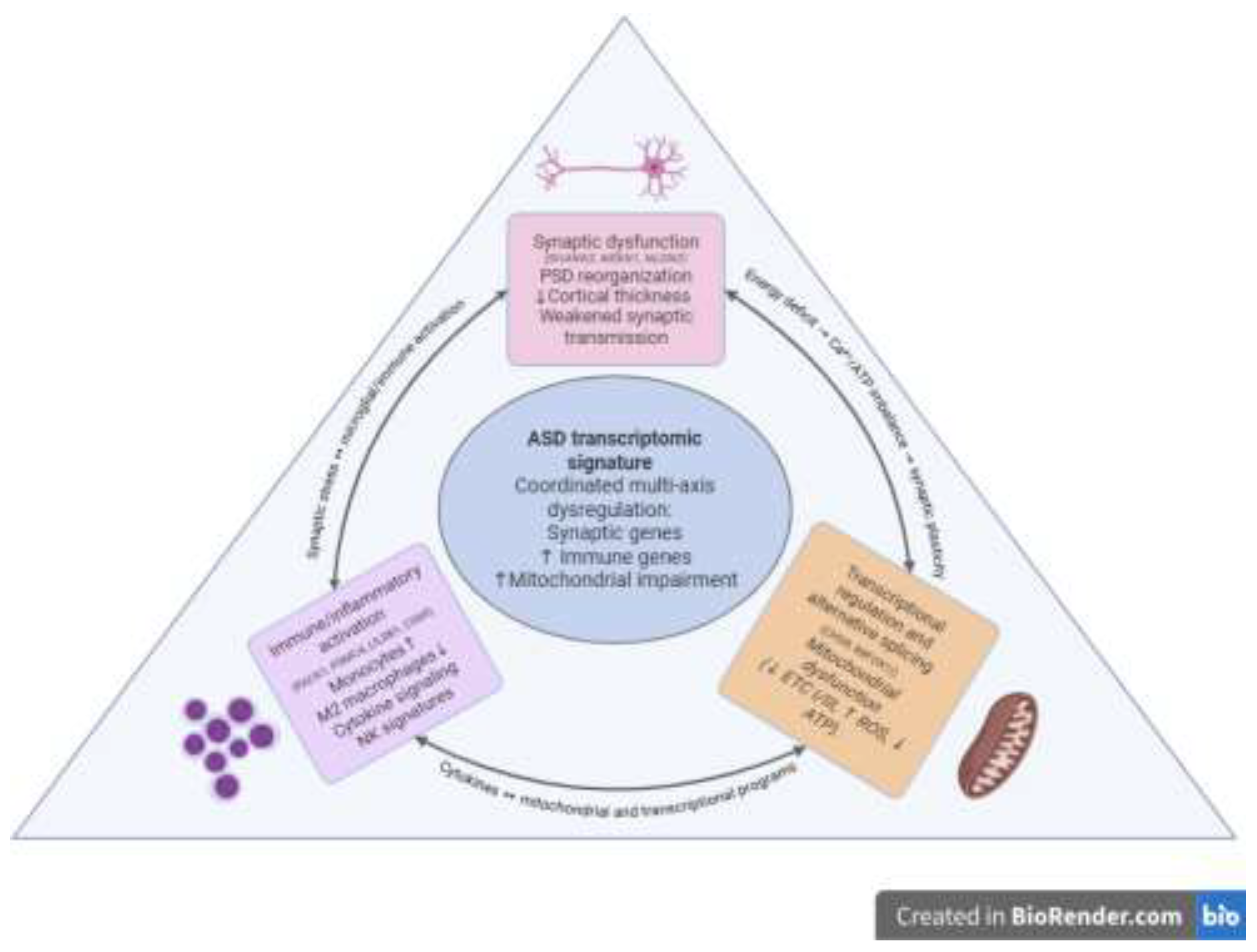
Pathogenetic Axes and Transcriptomics
Molecular Context: Synaptic Dysfunction (Transcriptomic Correlates)
Immune and Inflammatory Mechanisms: Transcriptomic Data of Peripheral Blood.
Transcriptional Control and Mitochondrial Disorders in ASD.
Transcriptomics.
Modern Behavioural Diagnostic Tools for ASD and Their Limitations.
Definition of Transcriptomics and RNA-Seq Technologies
The Use of RNA-Seq in ASD Research
Comparison of Transcriptomic Methods for Pediatric Diagnosis of ASD.
Review of Transcriptomic Markers of ASD
Synaptic Genes
Immune and Inflammatory Markers
Regulators of Transcription and Splicing
Behavioral and Neuromodulatory Axes (Preclinical Data)
Existing Panels for the Diagnosis of ASD
Early Works.
Salivary Small-RNA Panel/Test.
Peripheral Blood-Based Panels.
Ethics and Clinical Translation of Transcriptome Tests in ASD
Conclusion
Summary Assessment of Potential.
Balance of Strengths and Limitations.
Prospects and Prerequisites for Translation.
STARD 2015 Mapping for Transcriptomic Diagnostic Studies.
Funding
Ethical approval
Informed consent
Acknowledgments
Conflicts of Interest
Declaration: of generative AI and AI-assisted technologies in the manuscript preparation process
References
- American Psychiatric Association (2013) Diagnostic and statistical manual of mental disorders (5th ed.). Washington, DC: American Psychiatric Association. [CrossRef]
- Maenner, M.J., Shaw, K.A., Bakian, A.V., Bilder, D.A., Durkin, M.S., Esler, A., et al. (2023) ‘Prevalence and characteristics of autism spectrum disorder among children aged 8 years-Autism and Developmental Disabilities Monitoring Network, 11 sites, United States, 2020’, MMWR Surveillance Summaries, 72(2), pp. 1-14. [CrossRef]
- Zwaigenbaum, L., Bauman, M.L., Choueiri, R., Kasari, C., Carter, A., Granpeesheh, D., et al. (2015) ‘Early intervention for children with autism spectrum disorder under 3 years of age: recommendations for practice and research’, Pediatrics, 136(Suppl 1), pp. S60-S81. [CrossRef]
- Ji, S.-I., Park, H., Yoon, S.A. and Hong, S.-B. (2023) ‘A validation study of the CARS-2 compared with the ADOS-2 in the diagnosis of autism spectrum disorder: suggestion for cutoff scores’, Journal of the Korean Academy of Child and Adolescent Psychiatry, 34(1), pp. 45-50. [CrossRef]
- Pierce, K., Gazestani, V.H., Bacon, E., Barnes, C.C., Cha, D., Nalabolu, S., et al. (2019) ‘Evaluation of the diagnostic stability of the early autism spectrum disorder phenotype in the general population starting at 12 months’, JAMA Pediatrics, 173(6), pp. 578-587. [CrossRef]
- Lockwood Estrin, G., Milner, V., Spain, D., Happé, F. and Colvert, E. (2021) ‘Barriers to autism spectrum disorder diagnosis for young women and girls: a systematic review’, Review Journal of Autism and Developmental Disorders, 8(4), pp. 454-470. [CrossRef]
- Gandal, M.J., Haney, J.R., Wamsley, B., Yap, C.X., Parhami, S., Eilbott, J., et al. (2022) ‘Broad transcriptomic dysregulation occurs across the cerebral cortex in ASD’, Nature, 611(7936), pp. 532-539. [CrossRef]
- Gandal, M.J., Zhang, P., Hadjimichael, E., Walker, R.L., Chen, C., Liu, S. PsychENCODE Consortium, (2018) ‘Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder’, Science, 362(6420), eaat8127. [CrossRef]
- Wang, Z., Gerstein, M. and Snyder, M. (2009) ‘RNA-Seq: a revolutionary tool for transcriptomics’, Nature Reviews Genetics, 10(1), pp. 57-63. [CrossRef]
- Voinsky, I., Fridland, O.Y., Aran, A., Frye, R.E. and Gurwitz, D. (2023) ‘Machine learning-based blood RNA signature for diagnosis of autism spectrum disorder’, International Journal of Molecular Sciences, 24(3), 2082. [CrossRef]
- Voinsky, I., Zoabi, Y., Shomron, N., Harel, M., Cassuto, H., Tam, J., Rose, S., Scheck, A.C., Karim, M.A., Frye, R.E., Aran, A. and Gurwitz, D. (2022) ‘Blood RNA sequencing indicates upregulated BATF2 and LY6E and downregulated ISG15 and MT2A expression in children with autism spectrum disorder’, International Journal of Molecular Sciences, 23(17), 9843. [CrossRef]
- He, Y., Zhou, Y., Ma, W., et al. (2019) ‘An integrated transcriptomic analysis of autism spectrum disorder’, Scientific Reports, 9, 11818. [CrossRef]
- Tomaiuolo, P., Piras, I.S., Sain, S.B., et al. (2023) ‘RNA sequencing of blood from sex- and age-matched discordant siblings supports immune and transcriptional dysregulation in autism spectrum disorder’, Scientific Reports, 13, 807. [CrossRef]
- Filosi, M., Kam-Thong, T., Essioux, L., et al., on behalf of the Italian Autism Network (Domenici, E.) (2020) ‘Transcriptome signatures from discordant sibling pairs reveal changes in peripheral blood immune cell composition in autism spectrum disorder’, Translational Psychiatry, 10, 106. [CrossRef]
- Bourgeron, T. (2015) ‘From the genetic architecture to synaptic plasticity in autism spectrum disorder’, Nature Reviews Neuroscience, 16(9), 551-563. [CrossRef]
- Parikshak, N.N., et al. (2016) ‘Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism’, Nature, 540, 423-427. [CrossRef]
- Voineagu, I., et al. (2011) ‘Transcriptomic analysis of autistic brain reveals convergent molecular pathology’, Nature, 474, 380-384. [CrossRef]
- Romero-Garcia, R., et al. (2019) ‘Synaptic and transcriptionally downregulated genes are associated with cortical thickness differences in autism’, Molecular Psychiatry, 24(7), 1053-1064. [CrossRef]
- Li, X. (2024) ‘Unravelling the role of SHANK3 mutations in targeted therapies for autism spectrum disorders’, Discover Psychology, 4, 110. [CrossRef]
- Schwede, M., Nagpal, S., Gandal, M.J., Parikshak, N.N., Mirnics, K., Geschwind, D.H. and Morrow, E.M. (2018) ‘Strong correlation of downregulated genes related to synaptic transmission and mitochondria in post-mortem autism cerebral cortex’, Journal of Neurodevelopmental Disorders, 10(1), 19. [CrossRef]
- Li, H., Xu, Y., Li, W., Zhang, L., Zhang, X., Li, B., Chen, Y., Wang, X. and Zhu, C. (2023) ‘Novel insights into the immune cell landscape and gene signatures in autism spectrum disorder by bioinformatics and clinical analysis’, Frontiers in Immunology, 13, 1082950. [CrossRef]
- Horiuchi, F., Yoshino, Y., Kumon, H., Hosokawa, R., Nakachi, K., Kawabe, K., Iga, J.-I. & Ueno, S.-I. (2021). Identification of aberrant innate and adaptive immunity based on changes in global gene expression in the blood of adults with autism spectrum disorder. Journal of Neuroinflammation, 18, 102. [CrossRef]
- Wang, X., Xu, Q., Bey, A.L., Lee, Y. and Jiang, Y.-H. (2014) ‘Transcriptional and functional complexity of Shank3 provides a molecular framework to understand the phenotypic heterogeneity of SHANK3 causing autism and Shank3 mutant mice’, Molecular Autism, 5, 30. [CrossRef]
- Anitha, A., Nakamura, K., Thanseem, I., Matsuzaki, H., Miyachi, T., Yamada, K., (2012). Brain region-specific altered expression and association of mitochondria-related genes in autism. Molecular Autism, 3, 12. [CrossRef]
- Rossignol, D.A. & Frye, R.E. (2014). Evidence linking oxidative stress, mitochondrial dysfunction and inflammation in the brain of individuals with autism. Frontiers in Physiology, 5, 150. [CrossRef]
- Lebersfeld, J.B., Swanson, M., Clesi, C.D. & O’Kelley, S.E. (2021). Systematic review and meta-analysis of the clinical utility of the ADOS-2 and the ADI-R in diagnosing autism spectrum disorders in children. Journal of Autism and Developmental Disorders, 51(11), 4101-4114. [CrossRef]
- Yu, Y., Ozonoff, S. & Miller, M. (2023). Assessment of autism spectrum disorder. Assessment, 31(1), 24-41. [CrossRef]
- Wieckowski, A.T., Williams, L.N., Rando, J., Lyall, K. & Robins, D.L. (2023). Sensitivity and specificity of the Modified Checklist for Autism in Toddlers: A systematic review and meta-analysis. JAMA Pediatrics, 177(4), 373-383. [CrossRef]
- Robins, D.L., Casagrande, K., Barton, M., Chen, C.M.A., Dumont-Mathieu, T. & Fein, D. (2014). Validation of the Modified Checklist for Autism in Toddlers, Revised with Follow-up (M-CHAT-R/F). Pediatrics, 133(1), 37-45. [CrossRef]
- Moon, S.J., Hwang, J.S., Shin, A.L., Kim, J.Y., Bae, S.M., Sheehy-Knight, J. & Kim, J.W. (2019). Accuracy of the Childhood Autism Rating Scale: A systematic review and meta-analysis. Developmental Medicine & Child Neurology, 61(9), 1030-1038. [CrossRef]
- Santos, C.L.D., Barreto, I.I., Floriano, I., Tristão, L.S., Silvinato, A. & Bernardo, W.M. (2024). Screening and diagnostic tools for autism spectrum disorder: Systematic review and meta-analysis. Clinics, 79, 100323. [CrossRef]
- Yuen, T., Penner, M., Carter, M.T., Szatmari, P. & Ungar, W.J. (2018). Assessing the accuracy of the Modified Checklist for Autism in Toddlers: A systematic review and meta-analysis. Developmental Medicine & Child Neurology, 60(11), 1093-1100. [CrossRef]
- Maddox, B.B., Brodkin, E.S., Calkins, M.E., Shea, K., Mullan, K., Hostager, J., Mandell, D.S. & Miller, J.S. (2017). The accuracy of the ADOS-2 in identifying autism among adults with complex psychiatric conditions. Journal of Autism and Developmental Disorders, 47(9), 2703-2709. [CrossRef]
- Conesa, A., Madrigal, P., Tarazona, S., Gómez-Cabrero, D., Cervera, A., McPherson, A., (2016) ‘A survey of best practices for RNA-seq data analysis’, Genome Biology, 17, 13. [CrossRef]
- Rea, H.M., Øien, R.A., Shic, F., Webb, S.J. and Ratto, A.B. (2023) ‘Sex differences on the ADOS-2’, Journal of Autism and Developmental Disorders, 53(7), 2878-2890. [CrossRef]
- Hull, L., Petrides, K.V. and Mandy, W. (2020) ‘The female autism phenotype and camouflaging: a narrative review’, Review Journal of Autism and Developmental Disorders, 7, 306-321. [CrossRef]
- Pearson, A. and Rose, K. (2021) ‘A conceptual analysis of autistic masking: understanding the narrative of stigma and the illusion of choice’, Autism in Adulthood, 3(1), 21-29. [CrossRef]
- Stevanovic, D., et al. (2021) ‘Measurement invariance of the Childhood Autism Rating Scale across six countries: India, Jamaica, Mexico, Spain, Turkey and USA’, Autism Research. [CrossRef]
- Velmeshev, D., et al. (2019) ‘Single-nucleus RNA-seq identifies molecular changes in autism’, Science, 364(6436), 685-689. [CrossRef]
- Pardo-Palacios, F.J., Wang, D., Reese, F., Diekhans, M., Carbonell-Sala, S., Williams, B., et al. (2024) ‘Systematic assessment of long-read RNA-seq methods for transcript identification and quantification’, Nature Methods, 21(7), 1349-1363. [CrossRef]
- Lim, E.T., Chan, Y., Dawes, P., Guo, X., Erdin, S., Tai, D.J.C., et al. (2022) ‘Orgo-Seq integrates single-cell and bulk transcriptomic data to identify cell type-specific driver genes associated with autism spectrum disorder’, Nature Communications, 13, 3243. [CrossRef]
- Lu, X., Ni, P., Suarez-Meade, P., Ma, Y., Forrest, E.N., Wang, G., Wang, Y., Quiñones-Hinojosa, A., Gerstein, M. & Jiang, Y.-H. (2024) Transcriptional determinism and stochasticity contribute to the complexity of autism-associated SHANK family genes. Cell Reports, 43(7), 114376. [CrossRef]
- Erbescu, A., Papuc, S.M., Budisteanu, M., Arghir, A. & Neagu, M. (2022) Re-emerging concepts of immune dysregulation in autism spectrum disorders. Frontiers in Psychiatry, 13, 1006612. [CrossRef]
- Hamada, N., Ito, H., Nishijo, T., Iwamoto, I., Morishita, R., Tabata, H., Momiyama, T. & Nagata, K.-I. (2016) Essential role of the nuclear isoform of RBFOX1, a candidate gene for autism spectrum disorders, in brain development. Scientific Reports, 6, 30805. [CrossRef]
- Huang, K., Wu, Y., Shin, J., Zheng, Y., Siahpirani, A.F., Lin, Y., Ni, Z., Chen, J., You, J., Keleş, S., Wang, D., Roy, S. & Lu, Q. (2021) Transcriptome-wide transmission disequilibrium analysis identifies novel risk genes for autism spectrum disorder. PLOS Genetics, 17(2), e1009309. [CrossRef]
- Gora, C., Dudas, A., Vaugrente, O., Drobecq, L., Pecnard, E., Lefort, G. & Pellissier, L.P. (2024) Deciphering autism heterogeneity: A molecular stratification approach in four mouse models. Translational Psychiatry, 14, 416. [CrossRef]
- Hicks, S.D., Rajan, A.T., Wagner, K.E., Barns, S., Carpenter, R.L. & Middleton, F.A. (2018) Validation of a salivary RNA test for childhood autism spectrum disorder. Frontiers in Genetics, 9, 534. [CrossRef]
- Glatt, S.J., Tsuang, M.T., Winn, M., Chandler, S.D., Collins, M., Lopez, L., Weinfeld, M., Carter, C., Schork, N., Pierce, K. & Courchesne, E. (2012) Blood-based gene expression signatures of infants and toddlers with autism. Journal of the American Academy of Child & Adolescent Psychiatry, 51(9), 934-944.e2. [CrossRef]
- Varma, S. & Simon, R. (2006) Bias in error estimation when using cross-validation for model selection. BMC Bioinformatics, 7, 91. [CrossRef]
- Cohen, J.F., Korevaar, D.A., Altman, D.G., Bruns, D.E., Gatsonis, C.A., Hooft, L., et al. (2016) STARD 2015 guidelines for reporting diagnostic accuracy studies: Explanation and elaboration. BMJ Open, 6(11), e012799. [CrossRef]
- Sprang, M., Andrade-Navarro, M.A. & Fontaine, J.-F. (2022) Batch effect detection and correction in RNA-seq data using machine-learning-based automated assessment of quality. BMC Bioinformatics, 23(Suppl 6), 279. [CrossRef]
- Sosina, O.A., Tran, M.N., Maynard, K.R., Tao, R., Taub, M.A., Martinowich, K., Semick, S.A., et al. (2021) Strategies for cellular deconvolution in human brain RNA sequencing data [version 1, peer review: 1 approved, 1 approved with reservations]. F1000Research, 10, 750. [CrossRef]
- European Union. (2016) Regulation (EU) 2016/679 (General Data Protection Regulation). Official Journal of the European Union, L119, 1-88 (04 May 2016). Available at: https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:32016R0679 (esp. Art. 5, Art. 89).
- McPartland, J.C., Bernier, R.A., Jeste, S.S., Dawson, G., Nelson, C.A., Chawarska, K., … ABC-CT Consortium. (2020) The Autism Biomarkers Consortium for Clinical Trials (ABC-CT): Scientific context, study design, and progress toward biomarker qualification. Frontiers in Integrative Neuroscience, 14, 16. [CrossRef]
- Frye, R.E., Vassall, S., Kaur, G., Lewis, C., Karim, M. & Rossignol, D. (2019) Emerging biomarkers in autism spectrum disorder: A systematic review. Annals of Translational Medicine, 7(23), 792. [CrossRef]
- Weymann, D., Buckell, J., Fahr, P., Loewen, R., Ehman, M., Pollard, S., et al. (2024) Health care costs after genome-wide sequencing for children with rare diseases in England and Canada. JAMA Network Open, 7(7), e2420842. [CrossRef]
- Beck, K.B., MacKenzie, K.T., Kirby, A.V., McDonald, K., Moura, I., Breitenfeldt, K., et al., WIRE Consortium. (2025) Guidelines for the creation of accessible consent materials and procedures: Lessons from research with autistic people and people with intellectual disability. Autism in Adulthood (advance online publication). [CrossRef]
- Ament, I.H., DeBruyne, N., Wang, F. & Lin, L. (2025) Long-read RNA sequencing: A transformative technology for exploring transcriptome complexity in human diseases. Molecular Therapy, 33(3), 883-894. [CrossRef]
- Bossuyt PMM, Reitsma JB, Bruns DE, Gatsonis CA, Glasziou PP, Irwig L, et al. (2015) STARD 2015: An updated list of essential items for reporting diagnostic accuracy studies. BMJ 351: h5527. [CrossRef]
| Instrument | Sample / design | Sensitivity | Specificity | Source |
| ADOS-2 | Systematic review and HSROC meta-analysis, 22 studies | 0.89-0.92 | 0.81-0.85 | Lebersfeld et al., 2021 |
| ADI-R | Systematic review and meta-analysis | ~0.75 | ~0.82 | Lebersfeld et al., 2021 |
| ADOS-2 + ADI-R (combined) | Clinical studies (ranges reported from clinical series, not a meta-analysis) | 0.70-0.98 | 0.80-0.96 | Lebersfeld et al., 2021 |
| M-CHAT-R/F | Systematic review and meta-analysis: 50 studies (51 samples) | 0.83 (95% CI: 0.77-0.88) | 0.94 (95% CI: 0.89-0.97) | Wieckowski et al., 2023 |
| M-CHAT-R/F (two-step protocol) | Prospective study, >16,000 children (16-30 months) | 0.833 (95% CI: 0.73-0.93) | 0.992 (95% CI: 0.98-0.99) | Robins et al., 2014 |
| M-CHAT-R/F (independent meta-analysis) | Independent systematic review | 0.78 (95% CI: 57-91%) | 0.98 (95% CI: 88-100%) | Santos et al., 2024 |
| CARS / CARS-2 | Systematic review (n = 4,433) | 0.71-0.86 | 0.75-0.79 | Moon et al., 2019 |
| CARS-2ST (validation) | 237 children, 2-12 years, ROC analysis relative to ADOS-2 | Optimal cut-offs: 30 (autism), 28.5 (ASD) | Sensitivity: 98.9% (Autism, 30.25), 94.9% (Autism + ASD, 28.25) Specificity: 86.1% (Autism, 30.25), 100% (Autism + ASD, 28.25). |
Ji et al., 2023 |
| Method | Material | Cell res. | Cost | Complexity | Pediatric suitability | Primary use case | Notes / Key sources |
|---|---|---|---|---|---|---|---|
| Bulk RNA-seq | B/T | - | ↑ | ↑ | ✓ | DE genes, pathway enrichment | Requires cell-composition adjustment*, classic overview: Wang, Gerstein & Snyder (2009). |
| sc/snRNA-seq | T/Org | ++ | ↑↑ | ↑↑ | △ | Cell-type attribution, signatures | Sensitive to tissue/nuclei quality, Velmeshev et al. (2019) (ASD cortex). Integration with bulk**. |
| Long-read RNA-seq (PacBio/ONT) | B/T | + | ↑↑ | ↑ | △ | Isoforms, complex splicing | Most suitable as targeted post hoc after short-read***, Pardo-Palacios et al. (2024), Ament et al. (2025). |
| Saliva small RNAs | S | - | - | - | ✓ | Add-on after behavioral screening | 32-feature small-RNA panel, AUC ≈ 0.88 in children, Hicks et al. (2018). See M-CHAT-R/F: Robins et al. (2014). |
| Axis | Marker / panel | Context / level | Method | Key effect | Metric | Source |
|---|---|---|---|---|---|---|
| Transcriptomics / splicing (RBFOX1/A2BP1) | GRIN1 (NR1) | Postmortem cortex (frontal/temporal), H | RNA-seq, A2BP1 motif/positional-rule search, RT-PCR validation | Among top A2BP1-dependent DS events, annotated as synaptogenesis-related protein | A2BP1 motifs: p = 1.09×10⁻⁷, RT-PCR: ≈85% of targets confirmed (incl. GRIN1) | Voineagu et al., 2011 |
| Transcriptomics / splicing (RBFOX1/A2BP1) | CAMK2G (CaMKIIγ) | Postmortem cortex, H | RNA-seq, A2BP1 motif analysis, RT-PCR | Among top predicted A2BP1-dependent DS events, maps to neural module M12 | A2BP1 motifs: p = 1.09×10⁻⁷, RT-PCR: ≈85% of targets confirmed | Voineagu et al., 2011 |
| Transcriptomics / splicing (RBFOX1/A2BP1) | NRCAM | Postmortem cortex, H | RNA-seq, A2BP1 motif analysis, RT-PCR | Among top predicted A2BP1-dependent DS events, synaptogenesis-related protein | A2BP1 motifs: p = 1.09×10⁻⁷, RT-PCR: ≈85% of targets confirmed | Voineagu et al., 2011 |
| Transcriptomics / splicing (DTU) | ANK2 (isoform ANK2-013) | Postnatal frontal/temporal cortex, H (PsychENCODE) | Bulk RNA-seq, DTU (isoform level), co-expression & disease-specific PPI | ANK2-013 ↑ in ASD (DTU FDR < 0.05), co-expression network links with NRCAM, SCN4B, TAF9 | DTU FDR < 0.05 | Gandal et al., 2018 |
| Synapse | SHANK3 | Postmortem cortex, H | cDNA-capture + long-read RNA-seq | Region-specific differences of SHANK3 transcripts in ASD cortex | NR | Lu et al., 2024 |
| Immune-glial | IL-6, TNF-α | Plasma, postmortem cortex, H | Evidence synthesis (ELISA/IHC/expression studies) | Neuroinflammatory activation | NR | Erbescu et al., 2022 |
| Immune-glial / peripheral immunity | NMUR1, HMGB3, PTPRN2 | Peripheral blood, children, M | RNA-seq + xCell/CIBERSORT, WGCNA | ↓ NK signature, markers persist after adjusting for cell composition, some signals at trend level | FDR < 0.25 | Filosi et al., 2020 |
| Transcriptomics / splicing | RBFOX1 (nuclear isoform, iso1) | Preclinical-mouse, P | In utero shRNA knockdown, time-lapse migration, morphometry | Defects in radial migration and terminal translocation, ↓ axonal growth and dendritic arborization | NR | Hamada et al., 2016 |
| Transcriptional regulation | POU3F2 | Brain-integrative, H | TWAS/TITANS + fetal Hi-C + spatiotemporal expression + TFBS-LDSC | “Master regulator” targets enriched for ASD genes and LoF-DNMs, TFBS explain part of ASD heritability | TFBS h² = 11.7% (5.3×, p = 0.054), ASD-target enrichment 2.1-2.68× (p ≤ 0.012) | Huang et al., 2021 |
| Neuromodulation / behavior | Egr1, Foxp1, Homer1a, Oxt, Oxtr | Preclinical-mouse (4 ASD models), P | RT-qPCR, modular stratification, correlation with behavioral metrics | Expression panel separating models along socio-behavioral and neuroendocrine (OXT-system) axes | NR | Gora et al., 2024 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





