Submitted:
14 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Materials and Reagents
2.3. Instruments and Equipment
2.4. Experimental Methods
2.4.1. Sample Processing
2.4.2. Fatty Acid Value
2.4.3. Sample Genome DNA Extraction and PCR Amplification
2.4.4. PE Library Construction and Metagenomic Sequencing
2.4.5. GC-MS Detection
2.5. Data Analysis
3. Results
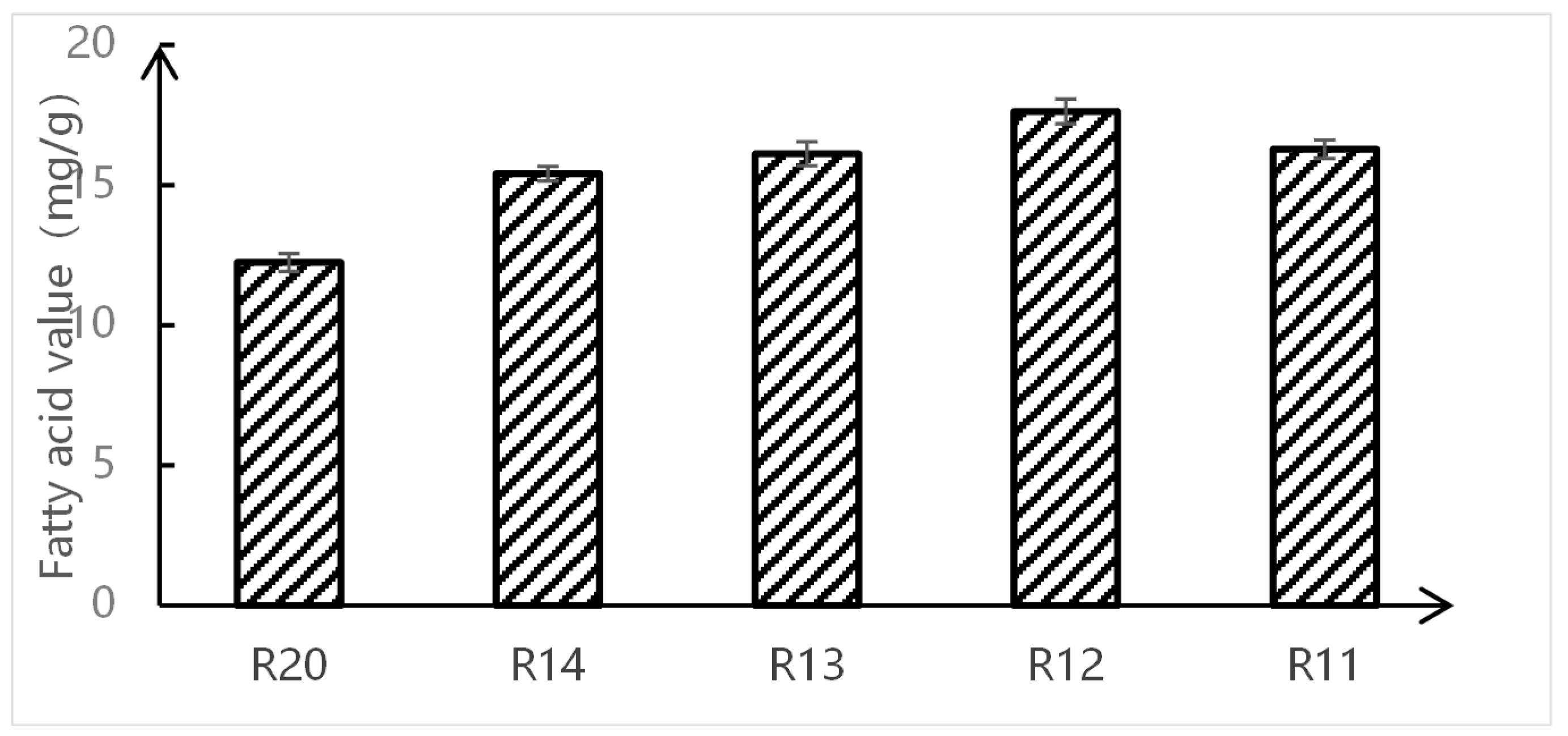
3.1. Changes in Rice Fatty Acid Content
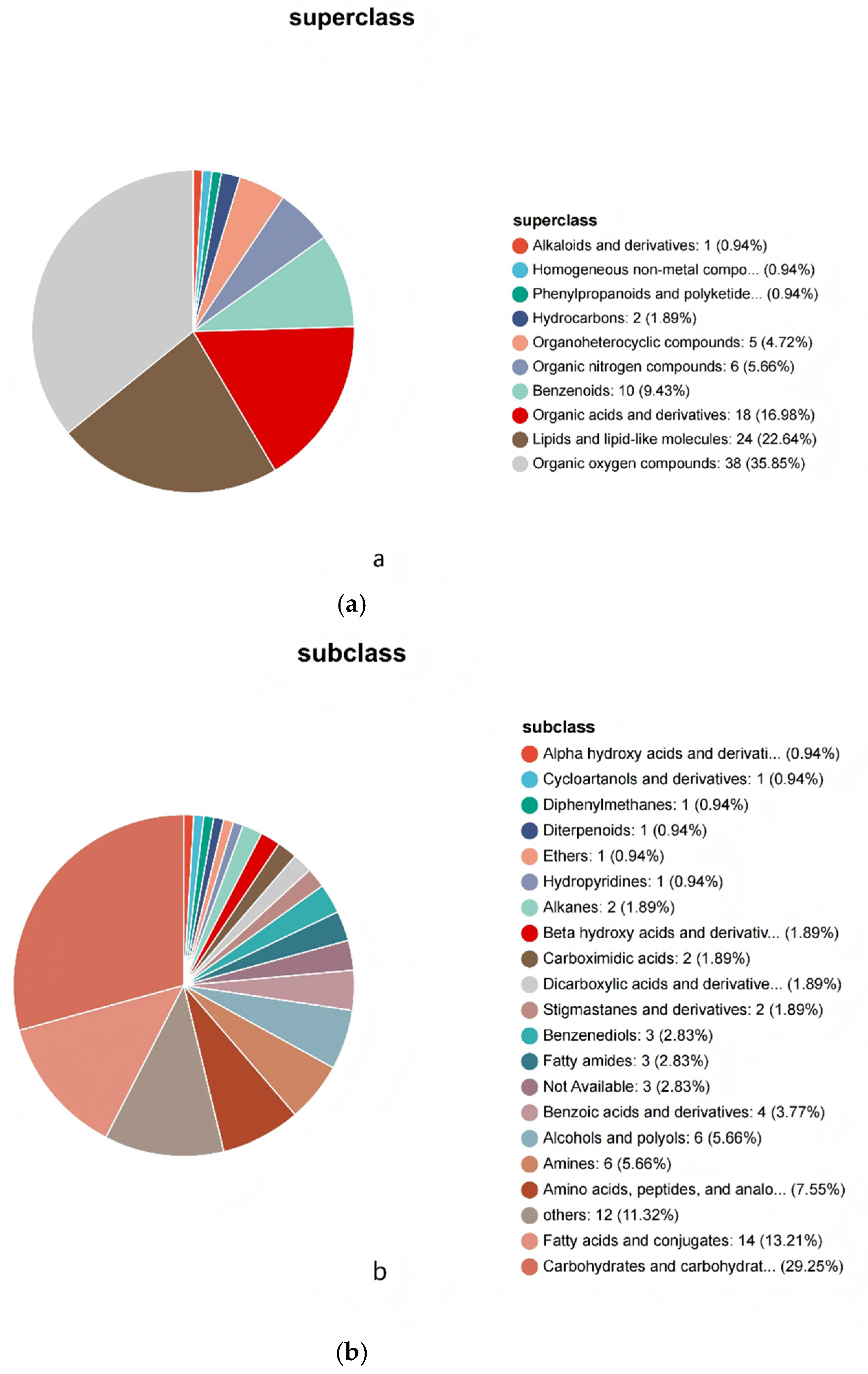
3.2. Analysis of Rice Metabolite Components
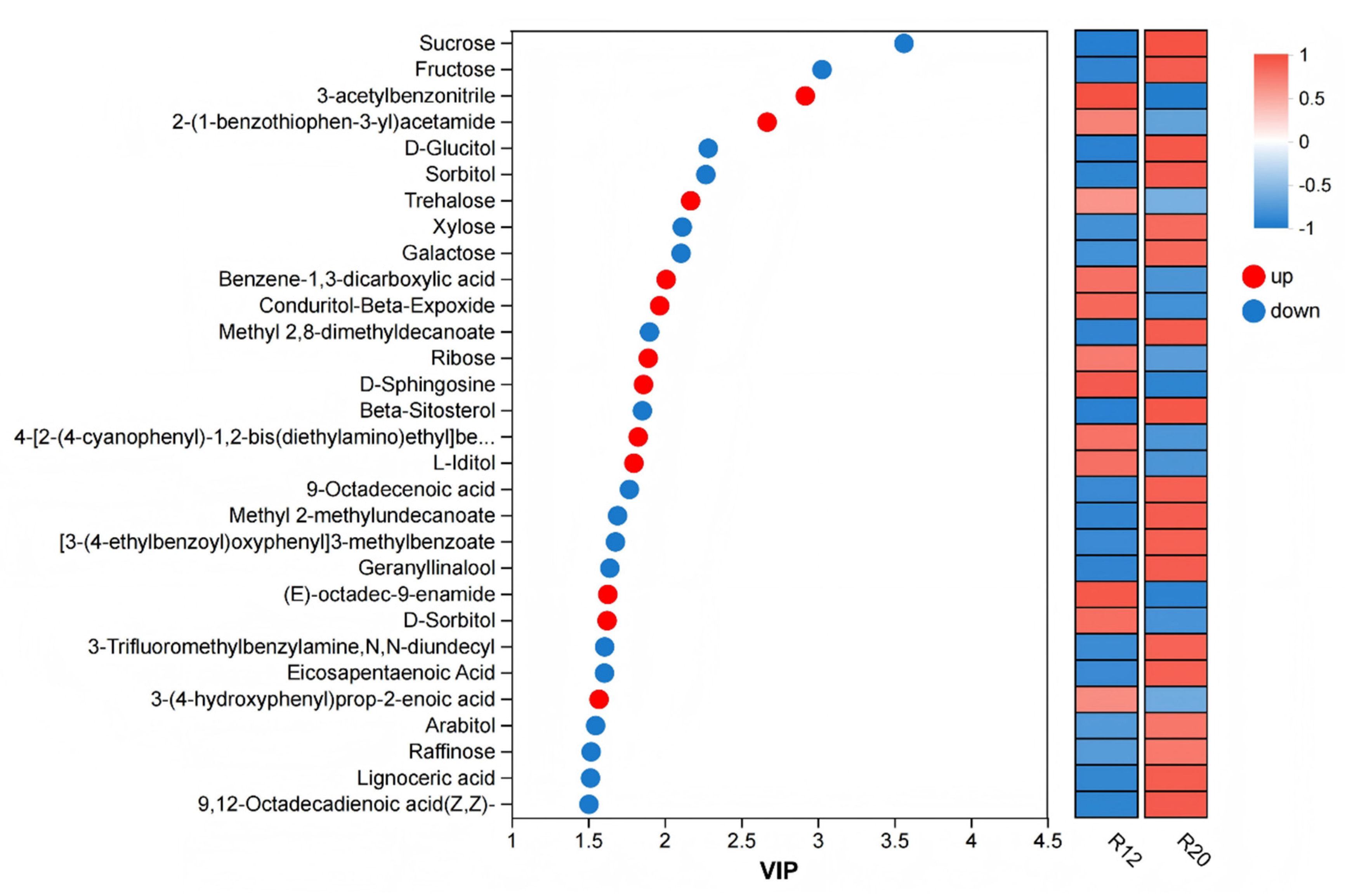
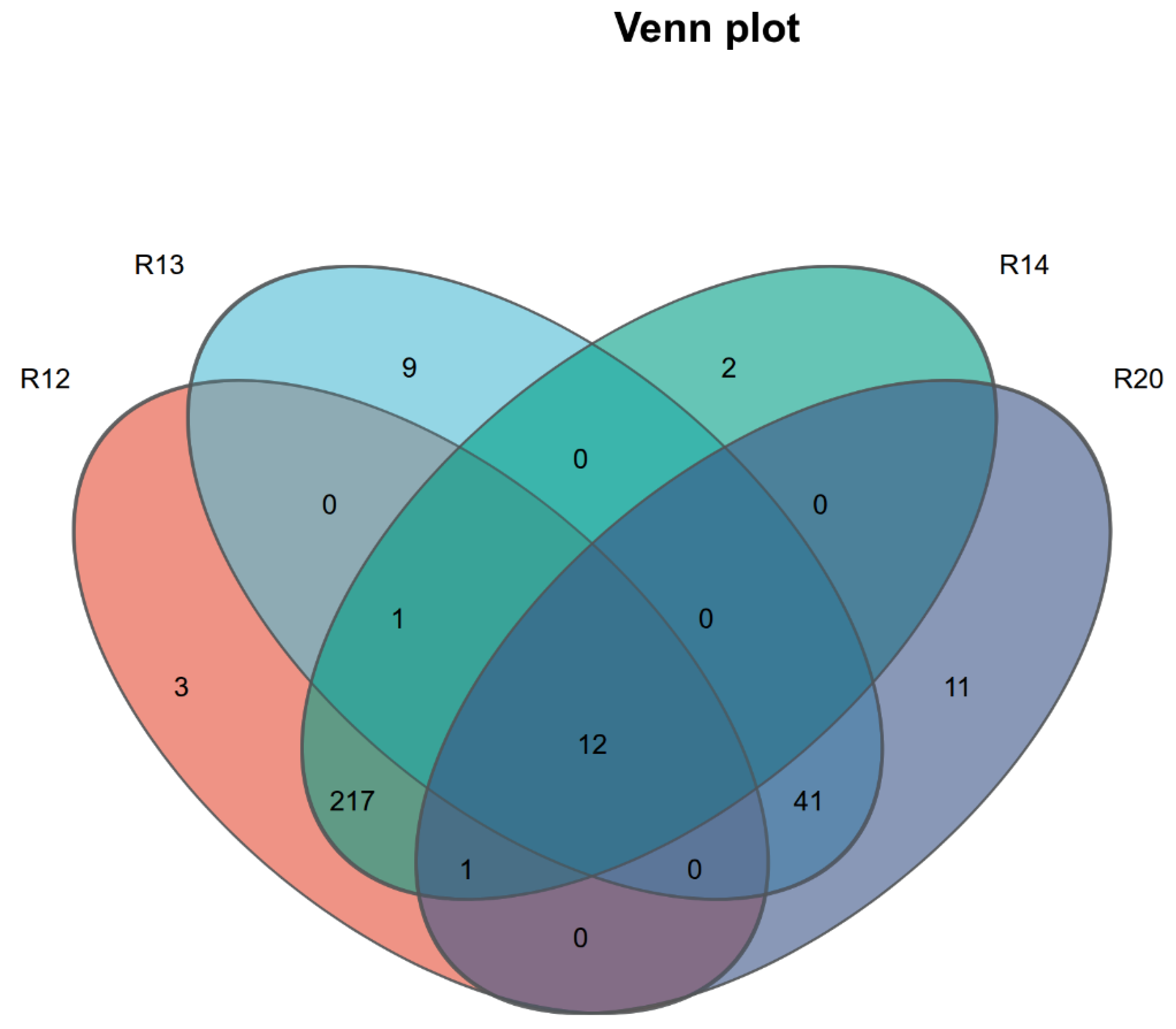
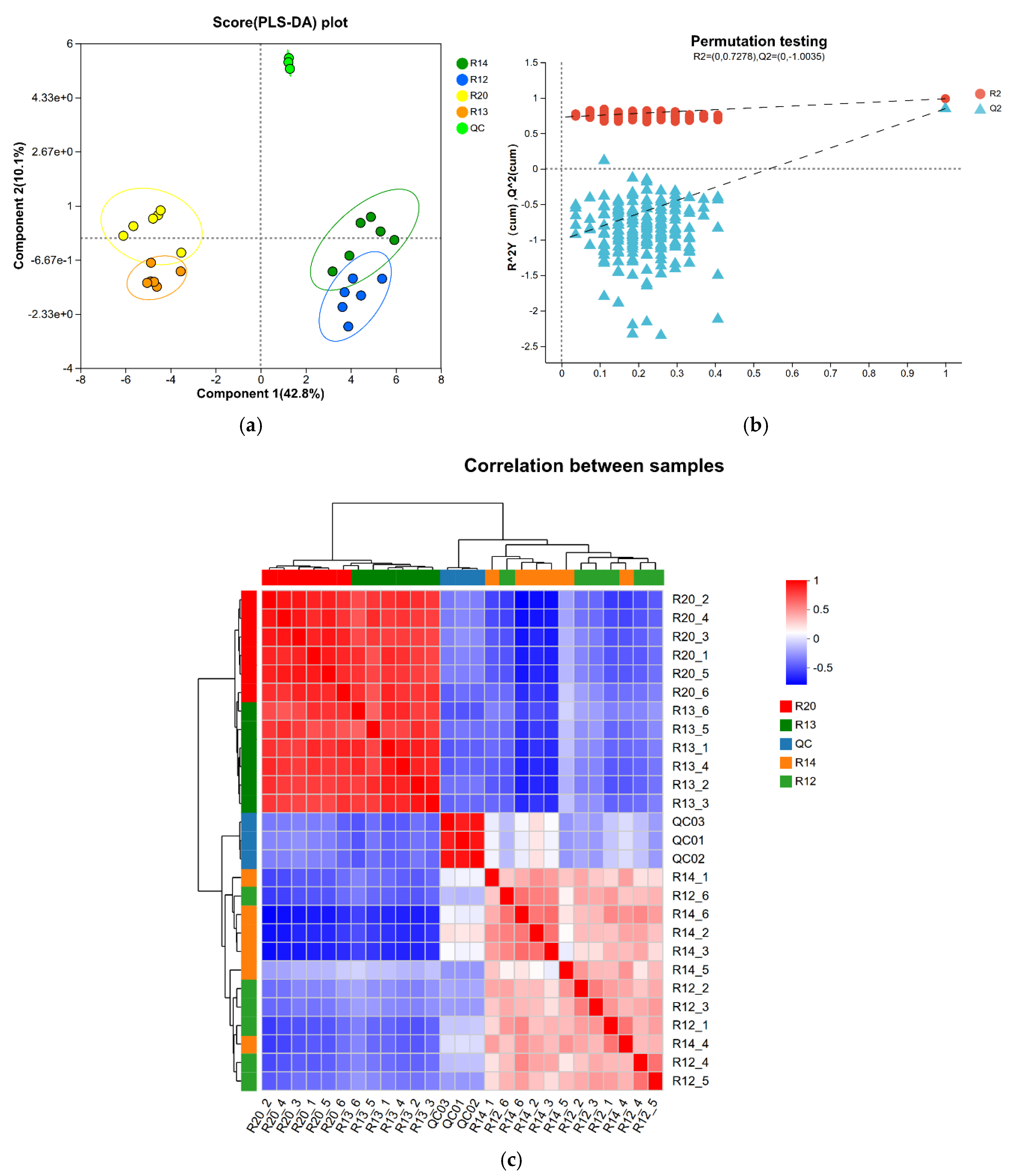
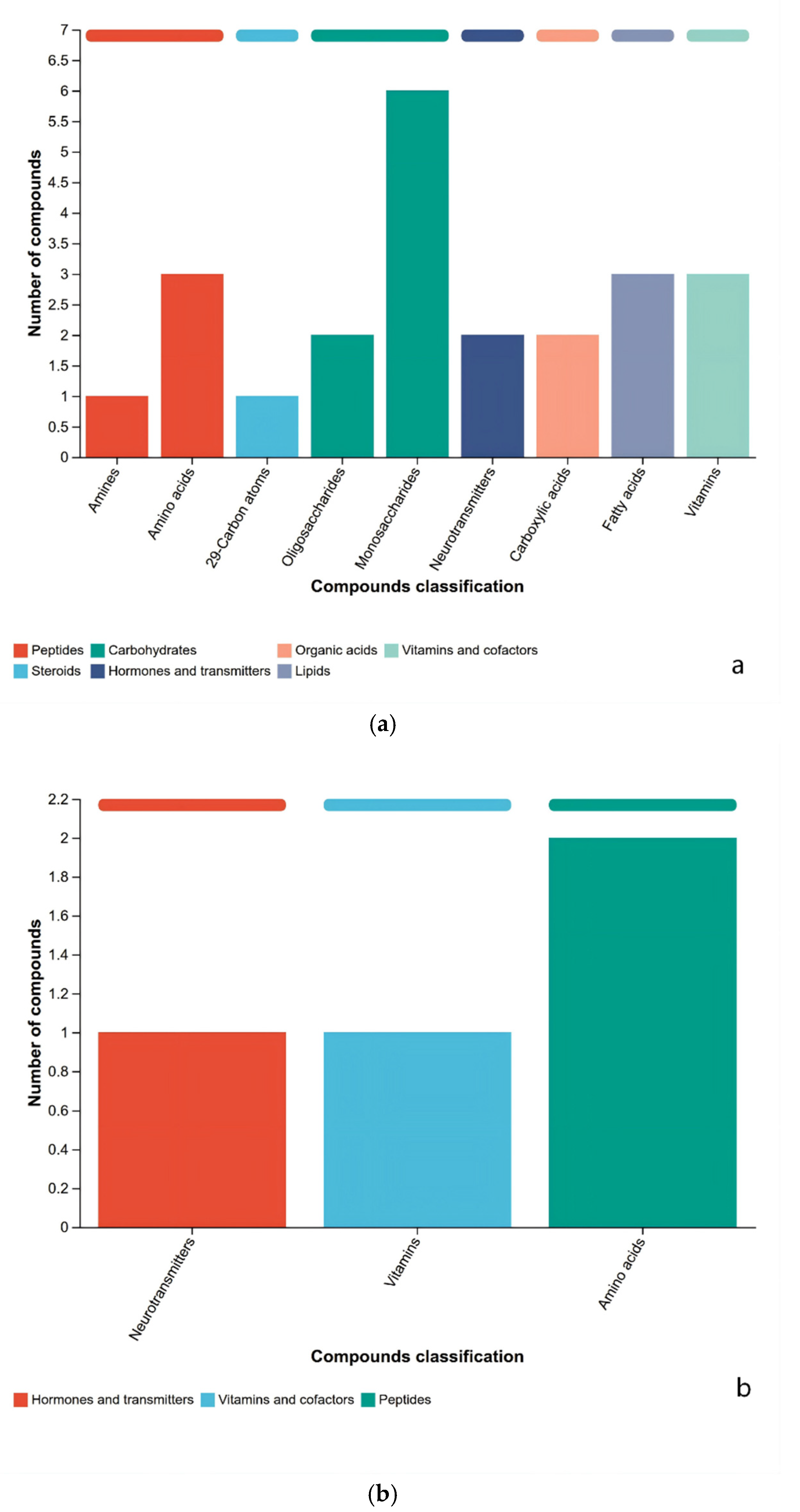
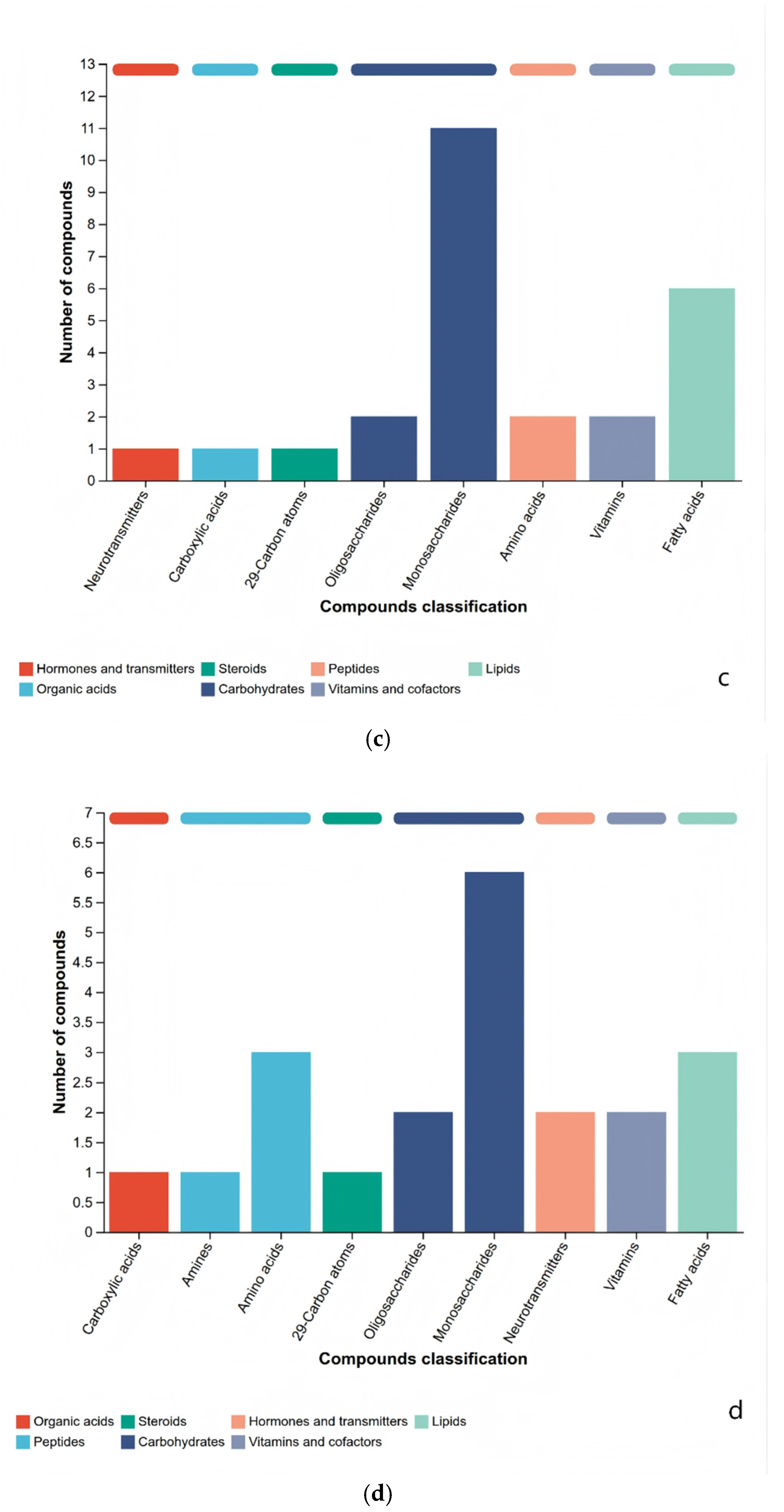
3.3. Analysis of Metabolite Composition Differences Among Rice Sample Groups
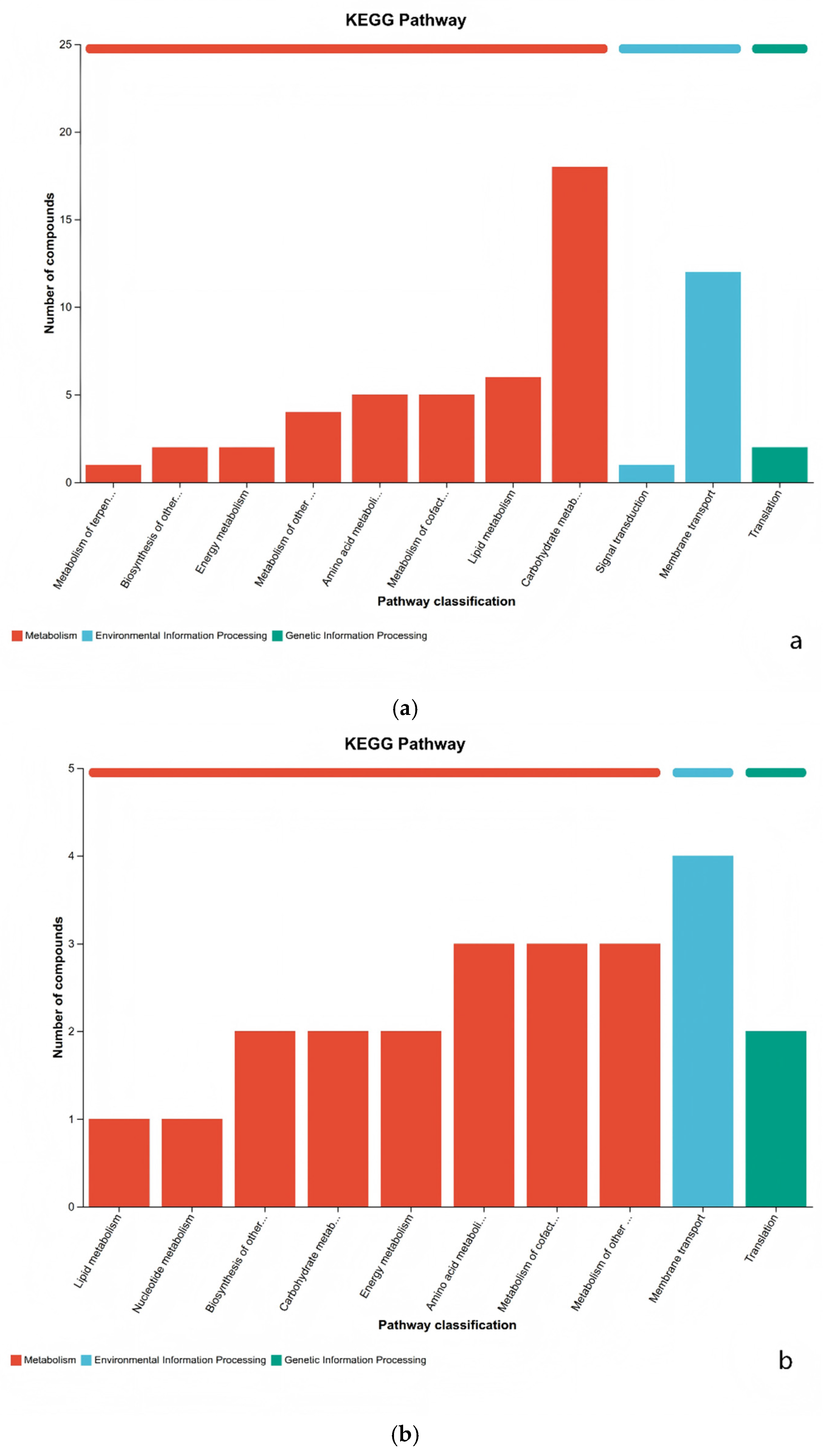
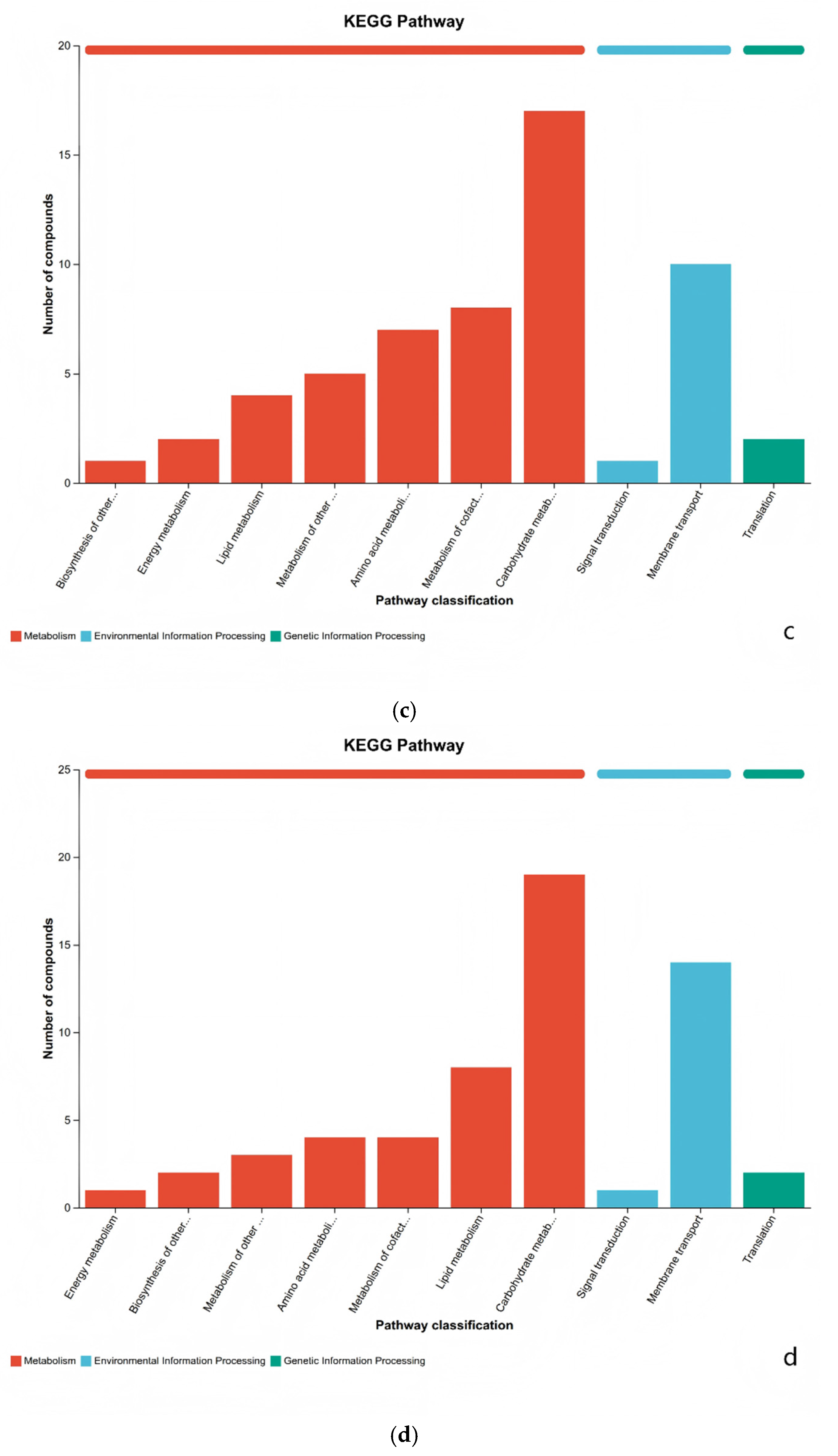
3.4. KEGG Differentially Metabolites Analysis
3.5. Metabolic Pathway Analysis of Differential Metabolites
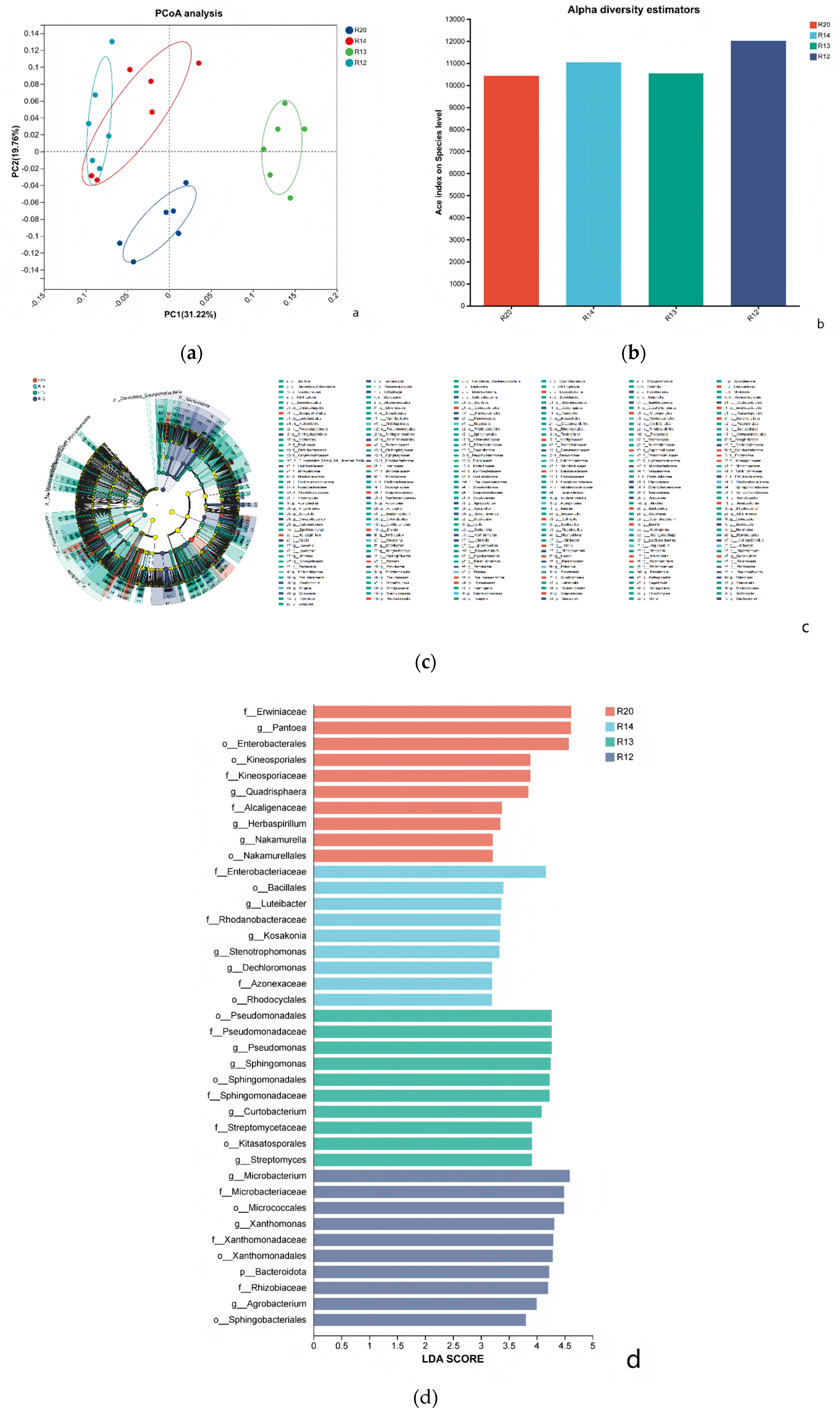
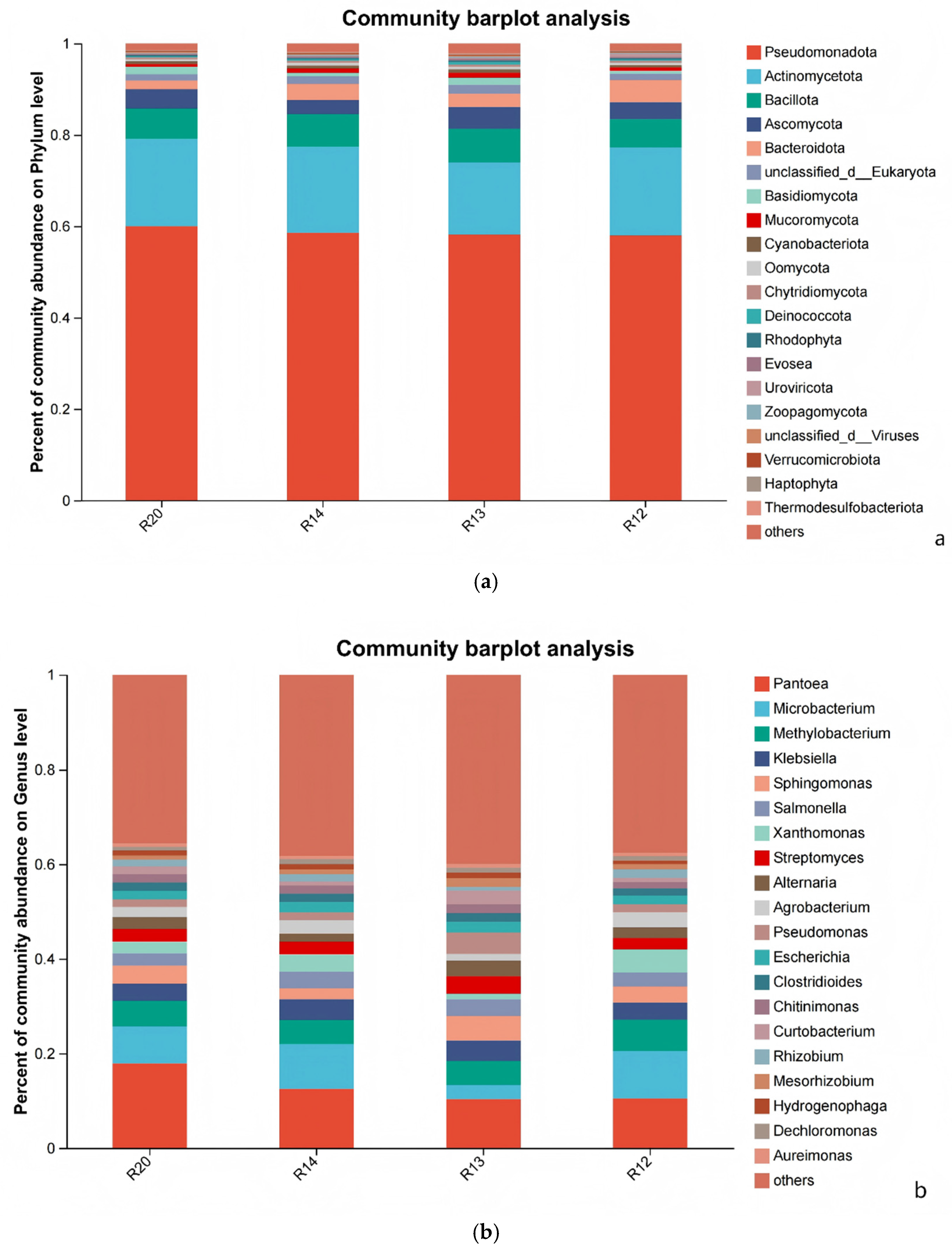
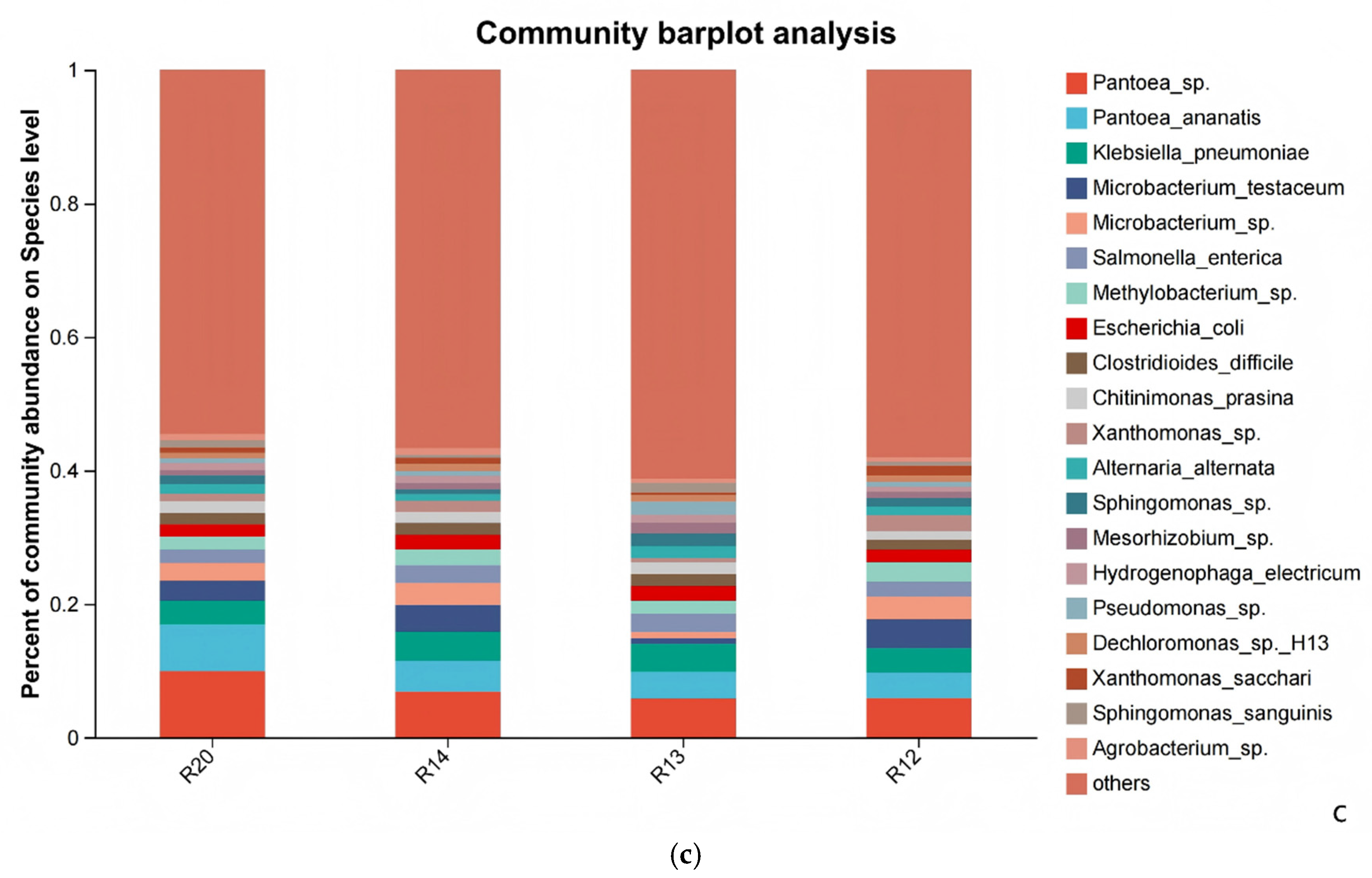
3.6. Analysis of Differences in Fungal Community Structure Composition of Rice at Different Storage Periods
3.7. Multi-Omics Association Analysis
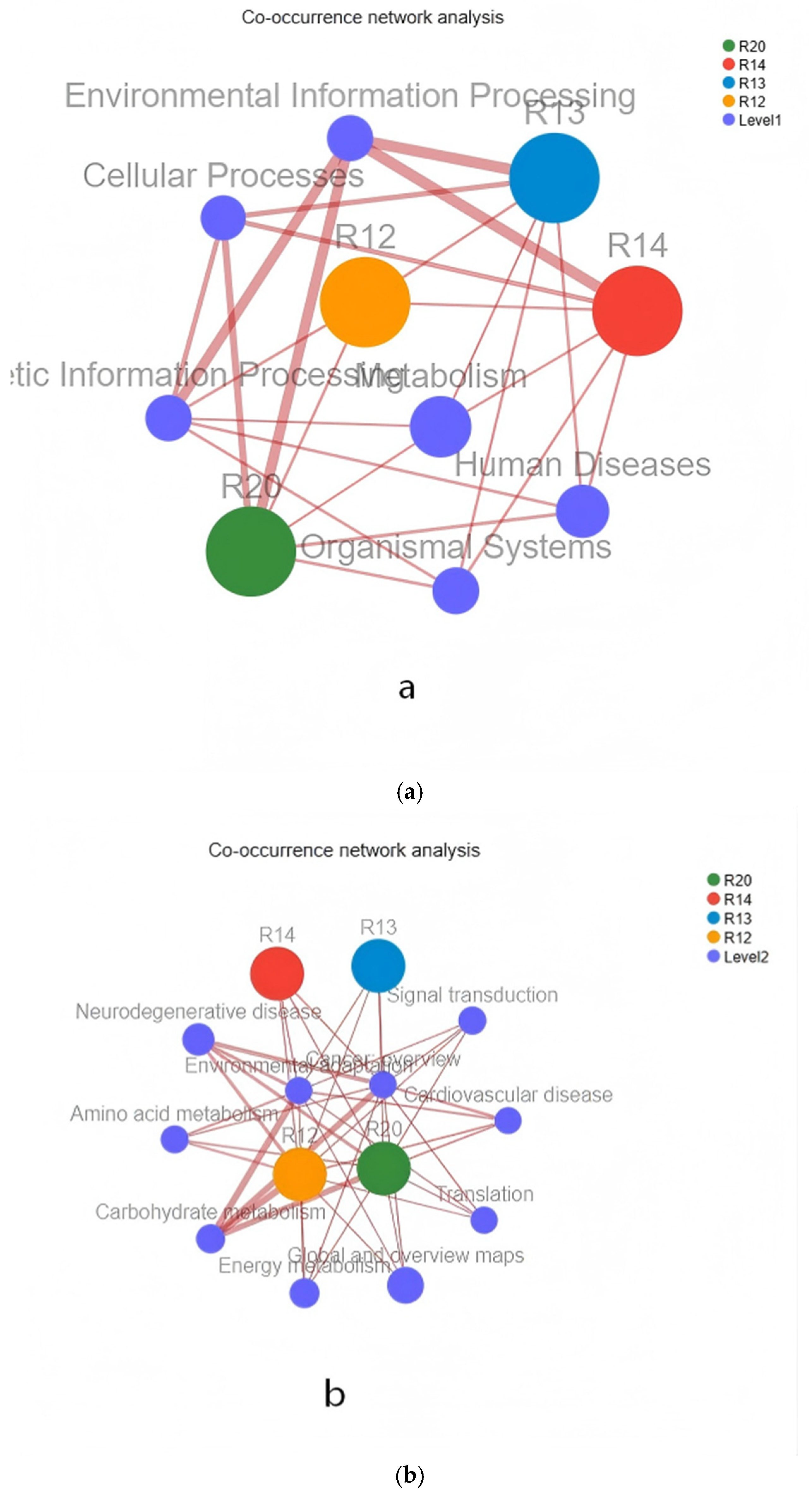
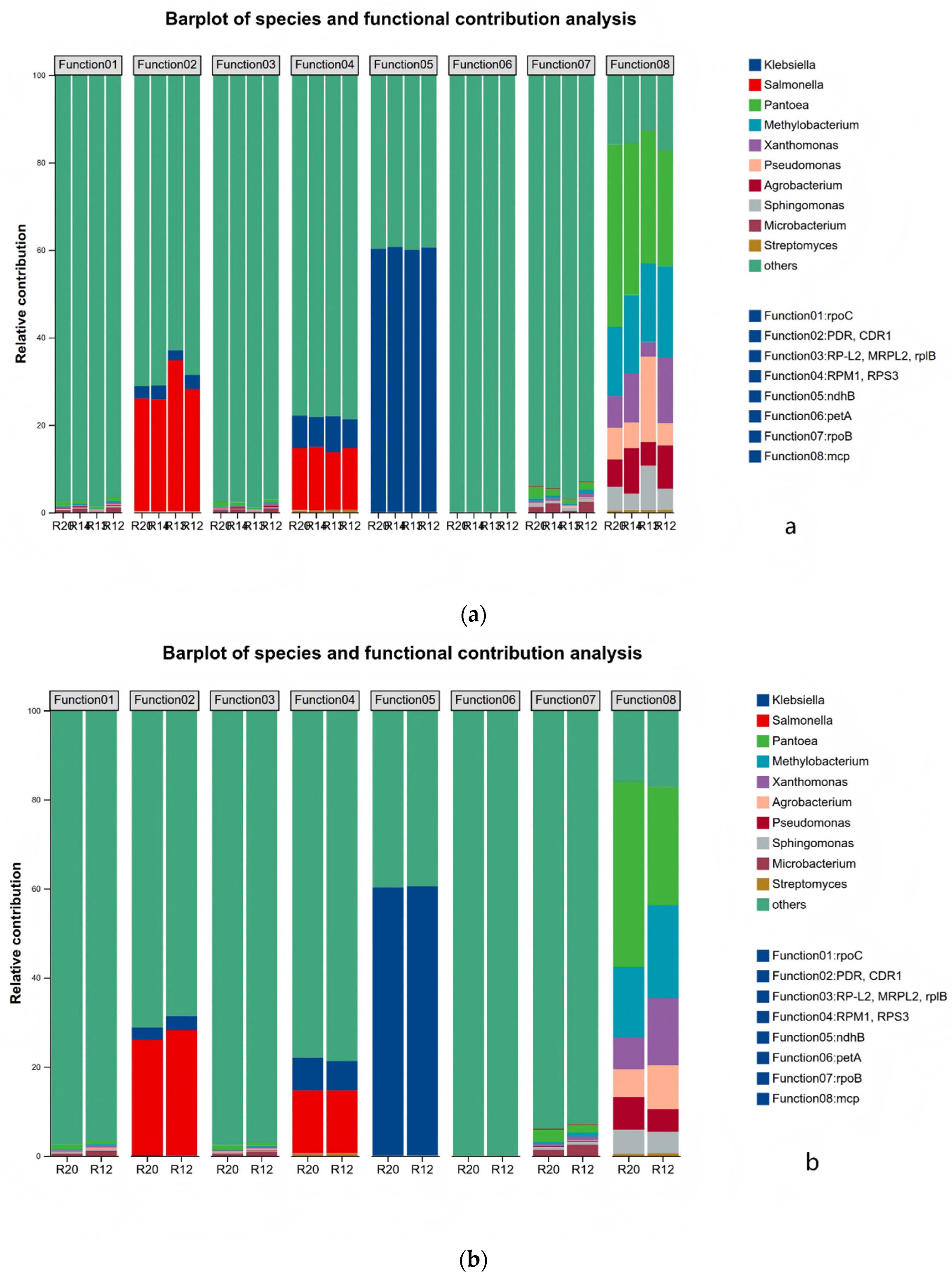
3.7.1. Functional Distribution Network Analysis of Rice Fungal Species
3.7.2. Key Functional Bacterial Genus in KEGG Metabolic Pathways
3.7.3. Correlation Analysis Between Dominant Bacterial Communities and Major Volatile Compounds
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Chen M, Zhang B, Gao Y, Liu R, Wang Y. Flavoromics-guided analysis of key odorants in fragrant japonica rice and their changes during storage. Food Chem. 2023;405:134876. [CrossRef]
- Fitzgerald MA, Bergman CJ, Resurreccion AP, Calingacion MN. The importance of eating quality in rice breeding: consumer preferences and sensory evaluation. Rice. 2021;14(1):89. [CrossRef]
- Liu Q, Wang H, Li J, Zhang G, Tian S. Postharvest storage-induced quality deterioration in rice: mechanisms, monitoring, and mitigation strategies. Compr Rev Food Sci Food Saf. 2024;23(1):321–345. [CrossRef]
- Zhou X, Zheng X, Zhu L, Liu X. Lipid oxidation and volatile compound formation in stored milled rice: effects of temperature and moisture content. J Agric Food Chem. 2022;70(15):4732–4741. [CrossRef]
- Zhao Y, Qu L, Xu X, Liu D. Starch retrogradation and texture deterioration in high-moisture brown rice during accelerated storage. Carbohydr Polym. 2023;301:120378. [CrossRef]
- Li Z, Wu C, Liu S, Zhang Y. Proteolysis and free amino acid accumulation in stored rice grains: correlation with sensory quality loss. LWT - Food Sci Technol. 2021;152:112345. [CrossRef]
- Xiong C, Liu Y, Wang H, Sun B, Shen Q. Integrated metagenomics and metabolomics reveal microbial drivers of metabolite shifts during postharvest rice storage. Microbiome. 2023;11(1):221. [CrossRef]
- Li J, Wang Y, Zhang G, Tian S. Fatty acid value as an indicator of rice storage quality: limitations and complementary approaches. J Cereal Sci. 2022;107:103521. [CrossRef]
- Tian S, Liu D, Ma Y, Zhang G. Correlation between lipid oxidation markers and sensory attributes in stored aromatic rice. Food Res Int. 2021;149:110678. [CrossRef]
- Chen H, Li Q, Guo W, Yu H. Dynamic profiling of volatile organic compounds in fragrant rice during storage using HS-SPME-GC-MS. Food Chem. 2023;404:134623. [CrossRef]
- Qu L, Zhao Y, Xu X, Liu D. Untargeted lipidomics reveal quality changes in high-moisture japonica brown rice at different storage temperatures. Foods. 2023;12(3):150. [CrossRef]
- Yu H, Li Q, Guo W, Chen H. Integrated metabolomic and transcriptomic analysis reveals key metabolic pathways involved in quality deterioration of stored rice. Int J Mol Sci. 2022;23(15):8521. [CrossRef]
- Mathew S, Lim LT. Biosynthesis and emission of volatile terpenoids in stored cereal grains: a review. Trends Food Sci Technol. 2022;128:1–12. [CrossRef]
- Wang Y, Li J, Zhang G, Tian S. Microbial contribution to ester formation in stored rice: evidence from co-culture experiments. Front Microbiol. 2024;15:1345678. [CrossRef]
- Callier MD, De Vos RCH, Hall RD, Bovy AG. Metabolomics reveals differences in volatile organic compounds and secondary metabolites between geographically indicated rice varieties. Metabolomics. 2021;17(5):67. [CrossRef]
- Capuano E, Oliviero T, van Boekel MAJS. Lipids as markers of oxidative stability in foods: A review. Trends Food Sci Technol. 2020;98:1–12. [CrossRef]
- Steinhaus M, Schieberle P. Characterization of the most odor-active compounds in an American Bourbon whisky by application of the aroma extract dilution analysis. J Agric Food Chem. 2000;48(2):417–421. [CrossRef]
- Zhang J, Li X, Liu Y, et al. Nitrogen metabolism and its regulation in plants. Plant Physiol. 2021;186(1):20–35. [CrossRef]
- Wang Y, Chen H, Guo W, et al. Oligosaccharides and sugar alcohols contribute to the viscoelasticity and textural refinement of stored rice. Food Chem. 2022;374:131655. [CrossRef]
- Zhang Q, Xu Y, Chen L, et al. Fatty amides inhibit lipoxygenase activity and delay the formation of hexanal during rice storage. LWT - Food Sci Technol. 2021;146:111455. [CrossRef]
- Langos D, Piironen V. Metabolic profiling reveals stabilization of metabolite types during later stages of rice storage. J Cereal Sci. 2022;105:103402. [CrossRef]
- Barker M, Rayens W. Partial least squares for discrimination. J Chemom. 2003;17(3):166–173. [CrossRef]
- Xia J, Wishart DS. Web-based inference of biological patterns, functions and pathways from metabolomic data using MetaboAnalyst. Nat Protoc. 2011;6(6):743–760. [CrossRef]
- Wang Y, Li J, Zhang G, Tian S. Dynamic changes in the rice metabolome during storage: insights into quality deterioration mechanisms. Food Chem. 2022;374:131655. [CrossRef]
- Liu H, Yang Y, Chen L, et al. Rice metabolomics reveals key metabolic changes during grain storage. Metabolites. 2021;11(11):767. [CrossRef]
- Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559. [CrossRef]
- Karp NA, Mason J, Beaudet AL, et al. Applying the ARRIVE guidelines to an in vivo database. PLOS Biol. 2017;15(9):e2002763. [CrossRef]
- Shi S, Wang H, Sun B, Shen Q. Lipidomics reveals fatty acid oxidation as a key driver of quality decline in stored grains. Metabolites. 2021;11(11):767. [CrossRef]
- Wang C, Zhang Y, Li X, et al. Metabolomics reveals core nutrient metabolic pathways dominate quality evolution during cereal storage. Food Chem. 2022;374:131655. [CrossRef]
- Coton M, Desmonts MH, Brillet B, et al. Metabolomics reveals carbohydrate metabolism as a key driver of rice quality evolution during storage. Food Chem. 2022;374:131655. [CrossRef]
- Johnson TS, Smith DL, Zhang Y, et al. Protein degradation during rice storage negatively correlates with taste quality. J Agric Food Chem. 2021;69(48):14756–14763. [CrossRef]
- Lee S, Kim Y, Park S, et al. Phenolic substance content is positively correlated with grain storage stability. Foods. 2023;12(3):150. [CrossRef]
- Zhang Z, Wang H, Li J, et al. Fungal community dynamics and mycotoxin contamination during rice storage under different temperature and humidity conditions. Food Microbiol. 2022;103:103948. [CrossRef]
- Segata N, Izard J, Waldron L, et al. Metagenomic biomarker discovery and explanation. Genome Biol. 2011;12(6):R60. [CrossRef]
- Faust K, Raes J. Microbial interactions: from networks to models. Nat Rev Microbiol. 2012;10(8):538–550. [CrossRef]
- Zhang Y, Figeys D, Wen B, Li J. Integration of multi-omics data: from networks to deep learning. Chem Rev. 2020;120(9):4563–4589. [CrossRef]
- Kanehisa M, Furumichi M, Sato Y, Ishiguro-Watanabe M, Tanabe M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 2021;49(D1):D545–D551. [CrossRef]
- Zhou X, Zheng X, Zhu L, Liu X. Volatile organic compounds in rice: identification and their roles in flavor. J Agric Food Chem. 2020;68(45):12687–12697. [CrossRef]
- Chen Y, Zhang Y, Li J, et al. Identification and quantification of volatile compounds in fragrant rice varieties using headspace solid-phase microextraction coupled with gas chromatography-mass spectrometry. Food Chem. 2022;374:131655. [CrossRef]
- Li Z, Wu C, Liu S, Zhang Y. Changes in volatile compounds during rice storage: A review. Trends Food Sci Technol. 2021;116:228–238. [CrossRef]
- Liu Q, Wang H, Li J, Zhang G, Tian S. Effects of phytosterols on rice quality during storage. Food Chem. 2022;374:131655. [CrossRef]
- Zhang Y, Wang H, Li J, et al. Role of phytosterols in maintaining rice quality during storage. J Agric Food Chem. 2021;69(48):14756–14763. [CrossRef]
- Chen Y, Zhang Y, Li J, et al. Impact of linalool on rice aroma and its retention during storage. J Agric Food Chem. 2022;70(15):4732–4741. [CrossRef]
- Zhang Y, Wang H, Li J, et al. Effect of sucrose on rice quality during storage: A review. J Stored Prod Res. 2021;94:101819. [CrossRef]
- Zhou Y, Li X, Wang J, et al. Multi-omics integration reveals the dynamic interplay between fungal communities and metabolites during rice storage. Food Res Int. 2023;174:113567. [CrossRef]
- Wang L, Chen S, Zhang H, et al. Network analysis of microbial co-occurrence patterns in stored grains uncovers keystone taxa driving quality deterioration. Microbiol Spectr. 2022;10(4):e01234-22. [CrossRef]
- Liu Y, Zhao Q, Sun C, et al. Topological features of metabolic networks predict rice quality changes during postharvest storage. J Agric Food Chem. 2021;69(30):8521–8530. [CrossRef]
- Huang R, Xu Z, Li M, et al. Fungal functional redundancy and its impact on metabolic pathway stability in stored rice ecosystems. ISME J. 2023;17(5):789–801. [CrossRef]
- Gao F, Zhang T, Wu D, et al. Temporal dynamics of KEGG pathway enrichment in rice microbiome during controlled atmosphere storage. Postharvest Biol Technol. 2022;193:112045. [CrossRef]
- Yang K, Li J, Wang P, et al. Klebsiella pneumoniae as a key driver of carbohydrate metabolism in deteriorating stored rice. Int J Food Microbiol. 2023;385:109987. [CrossRef]
- Zhang W, Liu X, Chen Y, et al. Salmonella enterica activates multidrug resistance pathways in response to oxidative stress during cereal storage. Appl Environ Microbiol. 2022;88(15):e00876-22. [CrossRef]
- Chen L, Wang H, Li Z, et al. Synergistic interactions among Pantoea, Pseudomonas, and Methylobacterium enhance chemotaxis and nutrient scavenging in stored rice. Front Microbiol. 2021;12:723456. [CrossRef]
- Zhao H, Liu S, Wang Y, et al. Metabolomic profiling links sucrose depletion to textural hardening in stored japonica rice. LWT - Food Sci Technol. 2023;174:114432. [CrossRef]
- Li X, Wang Q, Zhang R, et al. Galactose and xylose as critical modulators of perceived sweetness and stickiness in cooked rice. Food Chem. 2022;370:131025. [CrossRef]
- Xu M, Chen J, Liu B, et al. Raffinose family oligosaccharides contribute to the soft texture and mouthfeel of high-quality rice. Carbohydr Polym. 2021;272:118456. [CrossRef]
- Wang Z, Li Y, Zhang L, et al. Volatile ester profiles as fingerprints for authenticity and quality assessment of fragrant rice varieties. Food Control. 2023;145:109456. [CrossRef]
- Liu J, Zhang Y, Wang X, et al. Loss of linalool is a primary indicator of floral aroma degradation in stored aromatic rice. J Sci Food Agric. 2022;102(8):3210–3218. [CrossRef]
- Sun Y, Zhao X, Li H, et al. Phytosterols act as natural antioxidants to preserve lipid integrity and flavor in stored grains. Eur J Lipid Sci Technol. 2021;123(10):2100123. [CrossRef]
- Kumar V, Singh BP, Meena MC, et al. Multi-omics data fusion strategies for uncovering hidden biological insights in plant-microbe interactions. Brief Bioinform. 2023;24(1):bbac567. [CrossRef]
- Deng Y, Jiang YT, Yang YH, He ZG. Molecular ecological network analyses of microbial communities in extreme environments. Environ Microbiol. 2022;24(1):102–115. [CrossRef]
- Hu B, Jin J, Mao X, et al. The genetic architecture of sucrose metabolism and its impact on rice eating and cooking quality. Plant Biotechnol J. 2021;19(5):987–999. [CrossRef]
- Zhu L, Zhou X, Zheng X, et al. Comprehensive characterization of volatile compounds in rice using GC×GC-TOFMS and their contribution to flavor. Food Chem. 2023;403:134456. [CrossRef]
- Zhang N, Liu D, Tian S, et al. Integrated analysis of mycobiota and volatilome reveals fungal drivers of aroma loss in stored rice. Food Microbiol. 2024;117:104321. [CrossRef]
- Wang C, Li J, Zhang G, et al. Dynamic changes in free amino acid profiles and their correlation with taste quality in stored rice. J Cereal Sci. 2022;106:103489. [CrossRef]
- Liu Q, Ma Y, Wang H, et al. Correlation between lipid oxidation markers (LOX activity, MDA content) and sensory attributes in stored aromatic rice. LWT - Food Sci Technol. 2021;152:112389. [CrossRef]
- Chen H, Yu H, Guo W, et al. Real-time monitoring of volatile organic compound emissions from rice during storage using PTR-MS. Postharvest Biol Technol. 2023;195:112123. [CrossRef]
- Li Z, Wu C, Zhang Y, et al. Proteomic and metabolomic integration reveals the mechanism of protein degradation leading to quality loss in stored rice. Food Chem. 2022;387:132876. [CrossRef]
- Qu L, Xu X, Zhao Y, et al. Untargeted metabolomics coupled with chemometrics for discrimination of rice samples under different storage conditions. Metabolites. 2022;12(8):721. [CrossRef]
- Xiong C, Shen Q, Sun B, et al. Metagenomic and metatranscriptomic insights into the functional potential of the rice mycobiome during storage. Microbiome. 2024;12(1):45. [CrossRef]
- Johnson TS, Smith DL, Zhang Y, et al. Correlation between protein degradation products and off-flavor development in stored rice. J Agric Food Chem. 2022;70(10):3125–3133. [CrossRef]
- Lee S, Park S, Kim Y, et al. Phenolic compounds as natural preservatives to enhance the storage stability of cereal grains. Antioxidants. 2022;11(5):876. [CrossRef]
- Zhang Z, Li J, Wang H, et al. Mycotoxin risk assessment and fungal community succession in rice stored under simulated tropical conditions. World Mycotoxin J. 2023;16(2):145–158. [CrossRef]
- Kanehisa M, Sato Y, Morishima K. BlastKOALA and GhostKOALA: KEGG Tools for Functional Annotation of Genome and Metagenome Sequences. Nucleic Acids Res. 2016;44(W1):W50–W55. [CrossRef]
- Buchfink B, Reuter K, Drost HG. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat Methods. 2021;18(4):366–368. [CrossRef]

| order number | Metabolite name | retention time /min |
| 1 | (2,3,4,5,6-pentahydroxycyclohexyl)dihydrogen phosphate | 27.097 |
| 2 | (2-hydroxy-4-methoxyphenyl)-(2-hydroxyphenyl)methanone | 14.368 |
| 3 | (3S,8S,9S,10R,13R,14S,17R)-17-[(E,2R,5R)-5,6-dimethylhept-3-en-2-yl]-10,13-dimethyl-2,3,8,9,11,12,14,15,16,17-decahydro-1H-cyclopenta[a]phenanthren-3-ol | 33.285 |
| 4 | (4Z)-4-(furan-2-ylmethylidene)-2-phenyl-1,3-thiazol-5-one | 31.537 |
| 5 | (5S,8R,9S,10R,13S,14S)-4,4,10,13-tetramethyl-N-(2-phenylethyl)-1,5,6,7,8,9,11,12,14,15,16,17-dodecahydrocyclopenta[a]phenanthren-17-amine | 25.084 |
| 6 | (5Z)-5-methoxyiminohexane-1,2,3,4,6-pentol | 18.04 |
| 7 | (6Z)-6-methoxyiminoheptane-1,2,3,4,5,7-hexol | 23.859 |
| 8 | (9E,12E)-octadeca-9,12-dienoic acid | 25.089 |
| 9 | (E)-3-(4-hydroxy-3-methoxyphenyl)prop-2-enoic acid | 23.742 |
| 10 | (E)-but-2-ene-1,4-diol | 30.335 |
| 11 | (E)-hexadec-3-enenitrile | 23.807 |
| 12 | (E)-octadec-9-enamide | 27.207 |
| 13 | (Z)-octadec-9-enamide | 26.735 |
| 14 | (Z)-octadec-9-enenitrile | 23.703 |
| 15 | [1-[3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]sulfanylbut-3-enylideneamino]sulfate | 29.093 |
| 16 | [3-(4-ethylbenzoyl)oxyphenyl]3-methylbenzoate | 33.15 |
| 17 | [3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]dihydrogen phosphate | 24.982 |
| 18 | 1-(1-benzofuran-2-yl)ethanone | 6.873 |
| 19 | 1-(2-(3-Cyclohexenyl)Ethyl)Silatrane | 17.856 |
| 20 | 1-(2-methylazetidin-1-yl)ethanone | 6.19 |
| 21 | 1-(5-acetyl-4-methyl-1-phenyl-4H-pyridin-3-yl)ethanone | 14.007 |
| 22 | 1,3,5-Benzetriol | 16.92 |
| 23 | 10-bromodec-1-ene | 23.748 |
| 24 | 12A-Hydroxy-9-Demethylmunduserone-8-Carboxylic Acid | 7.412 |
| 25 | 17-(5,6-dimethylheptan-2-yl)-10,13-dimethyl-2,3,4,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-3-ol | 35.254 |
| 26 | 17-[(E)-5-ethyl-6-methylhept-3-en-2-yl]-10,13-dimethyl-2,3,4,7,8,9,11,12,14,15,16,17-dodecahydro-1H-cyclopenta[a]phenanthren-3-ol | 35.559 |
| 27 | 18-Deoxysagittarol | 33.57 |
| 28 | 1-iododecane | 21.629 |
| 29 | 1-Kestose | 29.44 |
| 30 | 1-methoxy-4-[2-(4-methoxyphenyl)propan-2-yl]-2-methylbenzene | 15.368 |
| 31 | 1-nitro-4-(4-nitrophenyl)benzene | 8.951 |
| 32 | 1-octyl-2,8,9-trioxa-5-aza-1-silabicyclo[3.3.3]undecane | 14.457 |
| 33 | 1-Stearoyl-Rac-Glycerol | 30.563 |
| 34 | 2-(1-benzothiophen-3-yl)acetamide | 6.149 |
| 35 | 2-(2-hydroxyethylamino)ethanol | 15.532 |
| 36 | 2-(3,4-dihydroxyphenyl)-2-hydroxyacetic acid | 27.683 |
| 37 | 2-(4-chlorophenyl)-8-methylquinoline-4-carboxylic acid | 31.792 |
| 38 | 2-(carboxyamino)pentanoic acid | 11.686 |
| 39 | 2-(carboxyamino)propanoic acid | 10.997 |
| 40 | 2-(ethylamino)ethanol | 12.507 |
| 41 | 2-(hydroxymethyl)-6-[3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxyoxane-3,4,5-triol | 36.738 |
| 42 | 2,2,4,4,6,6,8,8,10,10,12,12,14,14,16,16,18,18,20,20,22,22,24,24-tetracosamethyl-1,3,5,7,9,11,13,15,17,19,21,23-dodecaoxa-2,4,6,8,10,12,14,16,18,20,22,24-dodecasilacyclotetracosane | 28.95 |
| 43 | 2,2,4,4,6,6,8,8,10,10,12,12,14,14,16,16-hexadecamethyl-1,3,5,7,9,11,13,15-octaoxa-2,4,6,8,10,12,14,16-octasilacyclohexadecane | 17.741 |
| 44 | 2,2,4,4,6,6,8,8,10,10,12,12-dodecamethyl-1,3,5,7,9,11-hexaoxa-2,4,6,8,10,12-hexasilacyclododecane | 12.588 |
| 45 | 2,3,4,5-tetrahydroxypentanoic acid | 19.759 |
| 46 | 2,3,4,6-tetrahydroxyhexanoic acid | 21.138 |
| 47 | 2,3-dihydro-1-benzofuran-5-yl(morpholin-4-yl)methanone | 11.483 |
| 48 | 2,4-dimethylpentan-3-yloxy-dodecoxy-ethenyl-methylsilane | 32.385 |
| 49 | 2,5,7-trimethyl-3-phenylpyrazolo[1,5-a]pyrimidine | 33.85 |
| 50 | 2,5-bis(hydroxymethyl)-2-methoxyoxolane-3,4-diol | 19.704 |
| 51 | 2,6-dihydroxybenzoic acid | 22.617 |
| 52 | 2,6-ditert-butylphenol | 16.496 |
| 53 | 2-[[2-[[2-[carboxy(methyl)amino]acetyl]-methylamino]acetyl]-methylamino]acetic acid | 18.377 |
| 54 | 2-[1-(4-hydroxy-1,5,5-trimethyl-2-oxopyrrol-3-yl)ethylideneamino]acetic acid | 8.933 |
| 55 | 2-[3,4-dihydroxy-2,5-bis(hydroxymethyl)oxolan-2-yl]oxy-6-(hydroxymethyl)oxane-3,4,5-triol | 29.204 |
| 56 | 2-[4-(cyanomethyl)phenyl]acetonitrile | 9.758 |
| 57 | 2-[carboxy(methyl)amino]acetic acid | 15.914 |
| 58 | 2-amino-6,7-bis(3-phenylpropyl)-3H-pteridin-4-one | 6.98 |
| 59 | 2-aminoacetic acid | 7.865 |
| 60 | 2-aminobutanedioic acid | 16.113 |
| 61 | 2-aminoethanol | 12.026 |
| 62 | 2-Ethoxyethanol | 6.386 |
| 63 | 2-fluorobenzoic acid | 18.437 |
| 64 | 2-Hydroxypyridine | 8.032 |
| 65 | 2-methoxy-N,N-bis(2-methoxyethyl)ethanamine | 6.455 |
| 66 | 2-Methyl-2-Butanol | 26.784 |
| 67 | 2-methylnaphthalene | 13.3 |
| 68 | 2-methylpropan-2-ol | 24.52 |
| 69 | 2-nitrothiophene | 35.249 |
| 70 | 2-oxo-2-(6-phenylimidazo[1,2-a]pyridin-2-yl)acetic acid | 12.255 |
| 71 | 3-(4-hydroxyphenyl)prop-2-enoic acid | 20.018 |
| 72 | 3,4,5-trihydroxy-6-methoxyoxane-2-carboxylic acid | 30.038 |
| 73 | 3,4-Dihydroxybenzyl alcohol | 8.69 |
| 74 | 3,4-Dihydroxymandelic Acid | 24.936 |
| 75 | 3,5,5-trimethylimidazolidine-2,4-dione | 6.804 |
| 76 | 3-acetylbenzonitrile | 11.898 |
| 77 | 3-hydroxypropanoic acid | 9.854 |
| 78 | 3-Hydroxypyridine | 9.836 |
| 79 | 3-methyl-1-phenyl-3H-indol-2-imine | 12.988 |
| 80 | 3-methyl-N-prop-2-enylbut-2-enamide | 6.25 |
| 81 | 3-phenylmethoxybutane-1,2,4-triol | 32.261 |
| 82 | 3-Trifluoromethylbenzylamine,N,N-diundecyl | 16.92 |
| 83 | 4-(2-aminoethyl)-3-methoxyphenol | 11.801 |
| 84 | 4-(2-phenylpropan-2-yl)phenol | 21.466 |
| 85 | 4,6-ditert-butylbenzene-1,3-diol | 13.607 |
| 86 | 4-[[4-(methylamino)phenyl]methylideneamino]benzonitrile | 8.381 |
| 87 | 4-[2-(4-cyanophenyl)-1,2-bis(diethylamino)ethyl]benzonitrile | 6.409 |
| 88 | 4-[2-(4-hydroxyphenyl)propan-2-yl]phenol | 25.188 |
| 89 | 4-[4-(1,2,3,4,5,6,7,8-octahydroacridin-4-yl)phenyl]-1,2,3,4,5,6,7,8-octahydroacridine | 16.344 |
| 90 | 4-Coumaric acid | 20.92 |
| 91 | 4-cyanobenzoic acid | 6.367 |
| 92 | 4-Hydroxy-3-Methoxybenzoic Acid | 19.665 |
| 93 | 4-Hydroxyanthraquinone-2-carboxylic acid | 29.49 |
| 94 | 4-Hydroxybenzoate | 17.796 |
| 95 | 4-hydroxybenzonitrile | 6.7 |
| 96 | 4-methyl-15-thiatetracyclo[10.2.1.05,14.08,13]pentadeca-1(14),2,4,6,8(13),9,11-heptaene | 8.454 |
| 97 | 4-methyl-2-(2-nitro-5-piperidin-1-ylphenyl)phthalazin-1-one | 7.964 |
| 98 | 4-phenylquinolin-3-ol | 8.057 |
| 99 | 5-(Hydroxymethylfurfural) | 12.931 |
| 100 | 5,6-Dihydrouracil | 9.792 |
| 101 | 5,6-dimethyl-2-[(E)-2-phenylethenyl]benzotriazole | 7.73 |
| 102 | 5-Aminovaleric Acid | 15.957 |
| 103 | 5-Hydroxy Tryptamine | 11.049 |
| 104 | 5-Hydroxy Tryptophan | 14.572 |
| 105 | 5-hydroxypiperidine-2-carboxylic acid | 17.312 |
| 106 | 5-methylsulfanyl-2-phenyl-10H-pyrrolo[3,4-a]carbazole-1,3-dione | 27.711 |
| 107 | 6-ethyl-7-methoxy-2-methyl-3-phenylchromen-4-one | 22.657 |
| 108 | 6-Kestose | 27.46 |
| 109 | 7-benzyl-6-methyl-2-phenylindolizine | 31.49 |
| 110 | 8-acetyl-2,4,6-trimethyl-7-phenylpurino[7,8-a]imidazole-1,3-dione | 15.079 |
| 111 | 9,12-Octadecadienoic acid(Z,Z)- | 24.09 |
| 112 | 9-Octadecenoic acid | 24.15 |
| 113 | Acetylsalicylic Acid | 16.024 |
| 114 | Allantoin | 21.178 |
| 115 | Alpha-Tocopherol | 32.97 |
| 116 | Arabitol | 17.91 |
| 117 | Benzene-1,3-dicarboxylic acid | 14.95 |
| 118 | Benzene-1,4-diol | 14.333 |
| 119 | Beta-Gentiobiose | 27.725 |
| 120 | Beta-Lactose | 26.95 |
| 121 | Beta-Sitosterol | 35.42 |
| 122 | Bis[[hydroxy(dimethyl)silyl]oxy]-dimethylsilane | 10.215 |
| 123 | Butan-1-amine | 9.04 |
| 124 | Butan-2-ol | 29.225 |
| 125 | Butoxy-diethyl-pentoxysilane | 18.194 |
| 126 | Cadaverine | 14.618 |
| 127 | Carbamic acid | 8.433 |
| 128 | Carbamic Acid Ethyl Ester | 10.407 |
| 129 | Cellotetraose | 36.31 |
| 130 | Citric Acid | 20.296 |
| 131 | Clionasterol acetate | 32.74 |
| 132 | Conduritol-Beta-Expoxide | 22.876 |
| 133 | Cycloartenol | 36.45 |
| 134 | D-(+)-Altrose | 21.526 |
| 135 | D-(+)-Trehalose | 30.184 |
| 136 | D-Allose | 21.194 |
| 137 | D-beta-Galactopyranosyl-1,3-arabinose | 30.38 |
| 138 | Decane | 25.527 |
| 139 | Decoxy-diethyl-octoxysilane | 26.947 |
| 140 | D-Glucitol | 20.7 |
| 141 | D-Glucose | 21.274 |
| 142 | Diethylcarbamic acid | 8.316 |
| 143 | Diethyl-hexadecoxy-[(2,3,4,5,6-pentafluorophenyl)methoxy]silane | 36.207 |
| 144 | Dihydroxy(dimethyl)silane | 7.042 |
| 145 | Dimethoxymethane | 6.637 |
| 146 | Dimethyl-octadecoxy-(4-phenylphenoxy)silane | 35.207 |
| 147 | Diphenylphosphane | 9.373 |
| 148 | D-Malic Acid | 15.606 |
| 149 | D-Mannitol | 20.7 |
| 150 | D-manno-2-Heptulose | 20.17 |
| 151 | D-Mannose | 20.11 |
| 152 | Dodecanamide | 26.05 |
| 153 | D-Sorbitol | 21.772 |
| 154 | D-Sphingosine | 32.524 |
| 155 | D-Threitol | 15.869 |
| 156 | Eicosanoic acid | 26.49 |
| 157 | Eicosapentaenoic Acid | 35.01 |
| 158 | Elaidic Acid | 25.15 |
| 159 | Enolpyruvate | 8.847 |
| 160 | Erythritol | 14.86 |
| 161 | Ethanamine | 6.656 |
| 162 | Ethane-1,2-diol | 7.1 |
| 163 | Ethene-1,1,2-triol | 16.437 |
| 164 | Ethyl cyanate | 6.22 |
| 165 | Fructose | 19.93 |
| 166 | Galactinol | 33.48 |
| 167 | Galactitol | 26.562 |
| 168 | Galactose | 20.26 |
| 169 | Galactose oxime | 23.12 |
| 170 | Gamma-Aminobutyric Acid | 16.311 |
| 171 | Geranyllinalool | 33.57 |
| 172 | Gluconic Acid | 22.714 |
| 173 | Glucose | 21.26 |
| 174 | Glucose-1-Phosphate | 19.615 |
| 175 | Glycerol | 11.05 |
| 176 | Glycerol 1-Phosphate | 19.523 |
| 177 | Glycolic Acid | 8.633 |
| 178 | Glycyl-Proline | 17.266 |
| 179 | Harmaline | 10.812 |
| 180 | Heptadecanoic Acid | 24.345 |
| 181 | Heptanamide | 25.91 |
| 182 | Hexadecanamide | 24.784 |
| 183 | Hexadecanoic acid | 28.816 |
| 184 | Hexanamide | 26.84 |
| 185 | Hydroxy-[hydroxy(dimethyl)silyl]oxy-dimethylsilane | 7.895 |
| 186 | Hydroxy-methyl-trihydroxysilyloxysilicon | 19.838 |
| 187 | Icosanoic acid | 27.455 |
| 188 | Inositol | 23.639 |
| 189 | Ketohexose | 18.35 |
| 190 | Lactose | 30.45 |
| 191 | L-Asparagine | 18.312 |
| 192 | L-Glutamic Acid | 16.195 |
| 193 | L-Glycine | 12.707 |
| 194 | L-Iditol | 29.269 |
| 195 | Lignoceric acid | 30.23 |
| 196 | L-Serine | 13.558 |
| 197 | L-Tyrosine | 21.877 |
| 198 | Malonic Acid | 10.904 |
| 199 | Maltotriose | 37.187 |
| 200 | Melezitose | 28.16 |
| 201 | Methanediimine | 6.737 |
| 202 | Methanolphosphate | 10.429 |
| 203 | Methyl 2,8-dimethyldecanoate | 25.59 |
| 204 | Methyl 2-methylundecanoate | 25.59 |
| 205 | Methyl-(2-methyl-1,3-dioxoisoindol-4-yl)carbamic acid | 20.626 |
| 206 | Methylbeta-D-fructofuranoside | 18.69 |
| 207 | Methyl-Beta-D-Galactopyranoside | 27.841 |
| 208 | Methylimino(sulfanylidene)methane | 6.73 |
| 209 | Myo-Inositol | 22.65 |
| 210 | Myristic acid | 19.76 |
| 211 | N-(1,1-dioxo-2,3-dihydrothiophen-3-yl)-2-(4-methoxyphenoxy)acetamide | 28.538 |
| 212 | N-(4-aminobutyl)acetamide | 15.217 |
| 213 | N-[[3,6-dichloro-2,7-bis[2-(diethylamino)ethoxy]fluoren-9-ylidene]amino]acetamide | 6.375 |
| 214 | N-Acetylornithine | 10.104 |
| 215 | Nonacyclo[18.14.2.02,19.04,17.05,10.011,16.024,36.025,30.031,35]hexatriaconta-1(34),2(19),3,5,7,9,11,13,15,17,20,22,24(36),25,27,29,31(35),32-octadecaene | 18.147 |
| 216 | Nonadecanenitrile | 26.211 |
| 217 | Nonanamide | 26.06 |
| 218 | Norleucine | 12.089 |
| 219 | N-propan-2-ylacetamide | 6.487 |
| 220 | Octadecan-1-ol | 24.46 |
| 221 | Octadecanamide | 27.466 |
| 222 | Octamethylcyclotetrasiloxane | 6.994 |
| 223 | Oleamide | 27.336 |
| 224 | Oleic Acid | 24.15 |
| 225 | Palmitic Acid | 22.18 |
| 226 | Pantothenic Acid | 22.495 |
| 227 | P-Coummaric Acid | 21.951 |
| 228 | Pentane-1,5-diamine | 14.774 |
| 229 | Phosphoric Acid | 12.068 |
| 230 | P-Hydroxybenzaldehyde | 15.845 |
| 231 | Piperidin-2-one | 7.493 |
| 232 | P-Quinquephenyl | 31.9 |
| 233 | Propan-1-amine | 9.668 |
| 234 | Propane-1,2,3-triol | 33.99 |
| 235 | Propyl Lactate | 10.661 |
| 236 | Propylamine | 7.778 |
| 237 | Psicose | 20.945 |
| 238 | Pyridine-3-carboxylic acid | 12.602 |
| 239 | Pyrrolidine-2-carboxylic acid | 12.581 |
| 240 | Raffinose | 35.86 |
| 241 | Ribitol | 18.01 |
| 242 | Ribose | 17.11 |
| 243 | Serotonin | 15.008 |
| 244 | Sorbitol | 20.69 |
| 245 | Squalene | 30.07 |
| 246 | Stearic Acid | 24.43 |
| 247 | Stigmasterol | 34.62 |
| 248 | Succinic Acid | 12.832 |
| 249 | Sucrose | 27.18 |
| 250 | Tagatose | 21.063 |
| 251 | Terephthalic acid | 8.506 |
| 252 | Tetracosanoic acid | 31.1 |
| 253 | Tetrahydroxysilane | 11.203 |
| 254 | Threonic Acid | 16.656 |
| 255 | Trehalose | 29.32 |
| 256 | Tricosanoic acid | 32.204 |
| 257 | Tridec-2-yne | 9.881 |
| 258 | Trihydroxy(methyl)silane | 8.207 |
| 259 | Undecane | 11.772 |
| 260 | Urea | 11.584 |
| 261 | Vanillin | 18.076 |
| 262 | Xylitol,5TMS derivative | 19.015 |
| 263 | Xylose | 19.38 |
| Metabolite | Regulate | KEGG Pathway Description | Key Pathways |
| 3,4-Dihydroxymandelic Acid | up | Metabolic pathways;Tyrosine metabolism | Amino acid metabolism |
| 4-Coumaric acid | down | Metabolic pathways;Phenylpropanoid biosynthesis;Biosynthesis of secondary metabolites | Flavonoids |
| 9-Octadecenoic acid | down | Cutin, suberine and wax biosynthesis;Biosynthesis of unsaturated fatty acids;Fatty acid biosynthesis | Metabolism of short-chain fatty acids |
| Citric Acid | up | Metabolic pathways;Glyoxylate and dicarboxylate metabolism;Biosynthesis of secondary metabolites;Biosynthesis of cofactors;2-Oxocarboxylic acid metabolism;Carbon metabolism;Biosynthesis of amino acids;Alanine, aspartate and glutamate metabolism;Citrate cycle (TCA cycle) | Amino acid metabolism Energy metabolism |
| D-Glucose | up | Metabolic pathways;Biosynthesis of secondary metabolites;Amino sugar and nucleotide sugar metabolism;ABC transporters;Galactose metabolism;Glycolysis / Gluconeogenesis;Starch and sucrose metabolism;Pentose phosphate pathway | Energy metabolism |
| Glucose | down | Metabolic pathways;Biosynthesis of secondary metabolites;Biosynthesis of nucleotide sugars;ABC transporters;Amino sugar and nucleotide sugar metabolism;Carbon metabolism;Fructose and mannose metabolism;Galactose metabolism;Glycolysis / Gluconeogenesis;Starch and sucrose metabolism;Pentose phosphate pathway | Energy metabolism |
| L-Asparagine | up | Metabolic pathways;Biosynthesis of secondary metabolites;Cyanoamino acid metabolism;Biosynthesis of amino acids;Alanine, aspartate and glutamate metabolism;Aminoacyl-tRNA biosynthesis | Amino acid metabolism |
| L-Glutamic Acid | up | Metabolic pathways;Glyoxylate and dicarboxylate metabolism;Arginine biosynthesis;ABC transporters;Taurine and hypotaurine metabolism;Biosynthesis of cofactors;2-Oxocarboxylic acid metabolism;Nitrogen metabolism;Carbon metabolism;Arginine and proline metabolism;Glutathione metabolism;Biosynthesis of amino acids;Alanine, aspartate and glutamate metabolism;D-Amino acid metabolism;Aminoacyl-tRNA biosynthesis;Histidine metabolism;Biosynthesis of secondary metabolites;Butanoate metabolism; C5-Branched dibasic acid metabolism;Porphyrin metabolism | Amino acid metabolism Metabolism of short-chain fatty acids |
| Myristic acid | down | Metabolic pathways;Fatty acid biosynthesis | Metabolism of short-chain fatty acids |
| Palmitic Acid | down | Metabolic pathways;Fatty acid degradation;Fatty acid metabolism;Cutin, suberine and wax biosynthesis;Fatty acid biosynthesis;Fatty acid elongation;Biosynthesis of unsaturated fatty acids | Lipid metabolism Metabolism of short-chain fatty acids |
| Ribose | up | Metabolic pathways;Pentose phosphate pathway;ABC transporters | Energy metabolism |
| Stearic Acid | down | Metabolic pathways;Biosynthesis of unsaturated fatty acids;Fatty acid biosynthesis | Metabolism of short-chain fatty acids |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




