Submitted:
10 February 2026
Posted:
11 February 2026
You are already at the latest version
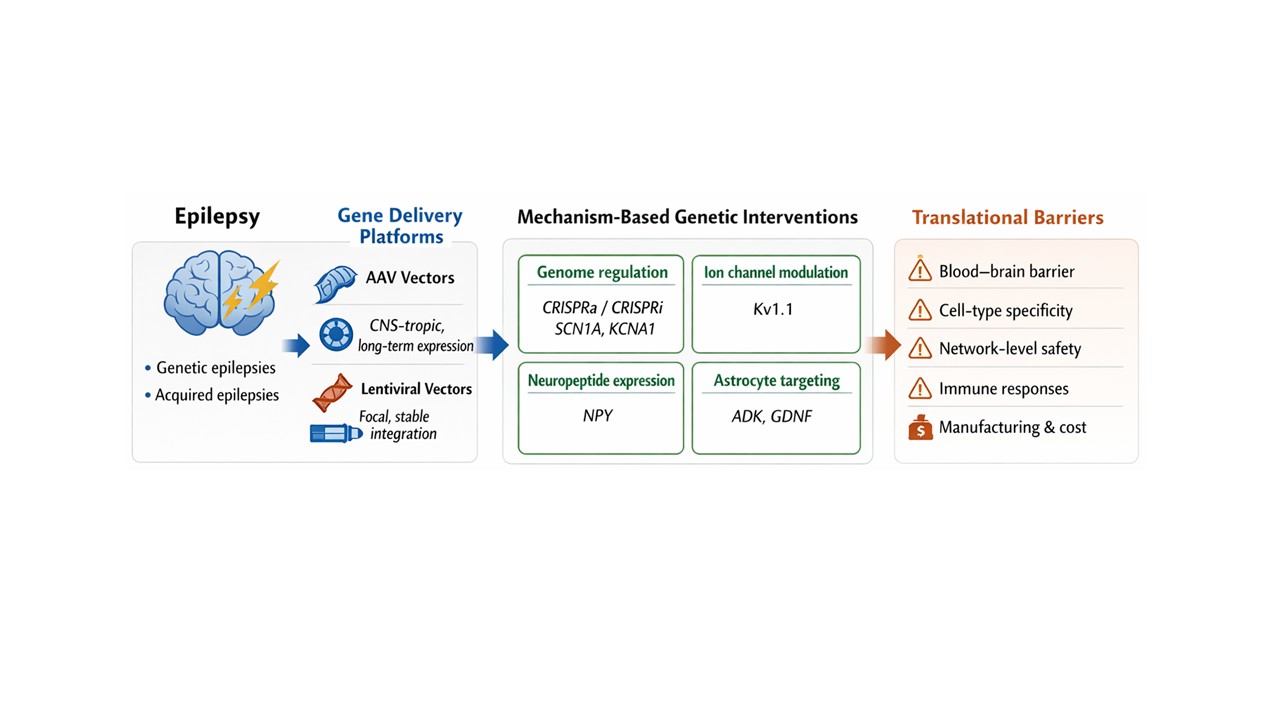
Abstract
Keywords:
Introduction
2. Vector Platforms for Epilepsy Gene Therapy
2.1. Viral Vectors for CNS Delivery
2.2. Lentiviral Vectors and Stable Gene Integration
2.3. Non-Viral Delivery Systems
2.4. Implications for Epilepsy-Focused Gene Therapy
3. Genome Editing and Gene Regulation Strategies in Epilepsy
3.1. Rationale for Genome Editing in Epilepsy
3.2. CRISPR/Cas9-Mediated Gene Disruption
3.3. CRISPR-Based Transcriptional Regulation (CRISPRa and CRISPRi)
3.4. Base Editing and Prime Editing
3.5. Implications for Clinical Translation
4. Mechanism-Based Gene Therapy Strategies for Epilepsy
4.1. Targeting Neuronal Excitability Through Ion Channel Modulation
4.2. Restoring Inhibitory Neurotransmission in Genetic Epilepsies
4.3. Neuropeptide-Based Suppression of Excitatory Neurotransmission
4.4. Astrocyte-Targeted Gene Therapy and Metabolic Modulation
4.5. Summary of Mechanism-Based Approaches
5. Translational and Clinical Challenges in Epilepsy Gene Therapy
5.1. Delivery Constraints and Cell-Type Specificity
5.2. Timing of Intervention and Disease Heterogeneity
5.3. Long-Term Safety and Network-Level Effects
5.4. Immune Responses and Repeat Dosing
5.5. Manufacturing, Cost, and Regulatory Considerations
5.6. Summary of Translational Challenges
6. Future Directions and Outlook
Author Contributions
Funding
Conflicts of Interest
References
- Stafstrom, CE; Carmant, L. Seizures and epilepsy: an overview for neuroscientists. Cold Spring Harb Perspect Med. 2015, 5, a022426. [Google Scholar] [CrossRef] [PubMed]
- Thijs, RD; Surges, R; O’Brien, TJ; Sander, JW. Epilepsy in adults. Lancet 2019, 393, 689–701. [Google Scholar] [CrossRef] [PubMed]
- Perucca, P; Bahlo, M; Berkovic, SF. The genetics of epilepsy. Annu Rev Genomics Hum Genet. 2020, 21, 205–230. [Google Scholar] [CrossRef] [PubMed]
- Sheng, J; Liu, S; Qin, H; Li, B; Zhang, X. Drug-resistant epilepsy and surgery. Curr Neuropharmacol. 2018, 16, 17–28. [Google Scholar] [CrossRef]
- Kay, MA; Glorioso, JC; Naldini, L. Viral vectors for gene therapy. Nat Med. 2001, 7, 33–40. [Google Scholar] [CrossRef]
- Naldini, L. Gene therapy returns to center stage. Nature 2015, 526, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Hudry, E; Vandenberghe, LH. Therapeutic AAV gene transfer to the nervous system: a clinical reality. Neuron 2019, 101, 839–862. [Google Scholar] [CrossRef] [PubMed]
- Simonato, M; Bennett, J; Boulis, NM; Castro, MG; Fink, DJ; Goins, WF; et al. Progress in gene therapy for neurological disorders. Nat Rev Neurol. 2013, 9, 277–291. [Google Scholar] [CrossRef] [PubMed]
- Street, JS; Qiu, Y; Lignani, G. Are genetic therapies for epilepsy ready for the clinic? Epilepsy Curr. 2023, 23, 245–250. [Google Scholar] [CrossRef]
- Daya, S; Berns, KI. Gene therapy using adeno-associated virus vectors. Clin Microbiol Rev. 2008, 21, 583–593. [Google Scholar] [CrossRef] [PubMed]
- Deverman, BE; Pravdo, PL; Simpson, BP; Kumar, SR; Chan, KY; Banerjee, A; et al. Cre-dependent selection yields AAV variants for widespread gene transfer to the adult brain. Nat Biotechnol. 2016, 34, 204–209. [Google Scholar] [CrossRef]
- Hsu, HL; Brown, A; Loveland, AB; Lotun, A; Xu, M; Luo, L; et al. Structural characterization of a novel human adeno-associated virus capsid with neurotropic properties. Nat Commun. 2020, 11, 3279. [Google Scholar] [CrossRef]
- Mathiesen, IH; et al. Development of AAV-CNS3 for enhanced brain transduction in large mammals. Mol Ther. 2023, 31, 1135–1149. [Google Scholar] [CrossRef]
- Wang, D; Tai, PWL; Gao, G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat Rev Drug Discov. 2019, 18, 358–378. [Google Scholar] [CrossRef]
- Dimidschstein, J; Chen, Q; Tremblay, R; Rogers, SL; Saldi, GA; Guo, L; et al. A viral strategy for targeting and manipulating interneurons across vertebrate species. Nat Neurosci. 2016, 19, 1743–1749. [Google Scholar] [CrossRef]
- Kantor, B; McCown, T; Leone, P; Gray, SJ. Clinical applications involving CNS gene transfer. Adv Genet. 2014, 87, 71–124. [Google Scholar]
- Wykes, RC; Heeroma, JH; Mantoan, L; Zheng, K; MacDonald, DC; Deisseroth, K; et al. Optogenetic and potassium channel gene therapy in a rodent model of focal neocortical epilepsy. Sci Transl Med. 2012, 4, 161ra152. [Google Scholar] [CrossRef]
- Snowball, A; Chabrol, E; Wykes, RC; Shekh-Ahmad, T; Cornford, JH; Lieb, A; et al. Epilepsy gene therapy using an engineered potassium channel. J Neurosci. 2019, 39, 3159–3169. [Google Scholar] [CrossRef]
- Colasante, G; et al. dCas9-based Scn1a gene activation restores inhibitory interneuron excitability and attenuates seizures in Dravet syndrome mice. Mol Ther. 2020, 28, 235–253. [Google Scholar] [CrossRef]
- Colasante, G; Qiu, Y; Massimino, L; Di Berardino, C; Cornford, JH; Snowball, A; et al. In vivo CRISPRa decreases seizures and rescues cognitive deficits in a rodent epilepsy model. Brain 2020, 143, 891–905. [Google Scholar] [CrossRef]
- Jinek, M; Chylinski, K; Fonfara, I; Hauer, M; Doudna, JA; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Doudna, JA; Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 2014, 346, 1258096. [Google Scholar] [CrossRef]
- Hsu, PD; Lander, ES; Zhang, F. DNA targeting CRISPR-Cas9: from the laboratory to the clinic. Nat Rev Mol Cell Biol. 2014, 15, 321–334. [Google Scholar] [CrossRef]
- Swiech, L; et al. In vivo interrogation of gene function in the mammalian brain using CRISPR-Cas9. Nat Biotechnol. 2015, 33, 102–106. [Google Scholar] [CrossRef]
- Staahl, BT; et al. Efficient genome editing in the mouse brain by local delivery of engineered Cas9 ribonucleoprotein complexes. Nat Biotechnol. 2017, 35, 431–434. [Google Scholar] [CrossRef]
- Gaudelli, NM; et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 2017, 551, 464–471. [Google Scholar] [CrossRef]
- Anzalone, AV; et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 2019, 576, 149–157. [Google Scholar] [CrossRef]
- Chen, PJ; Liu, DR. Prime editing for precise and highly versatile genome manipulation. Nat Rev Genet. 2023, 24, 161–177. [Google Scholar] [CrossRef]
- Noe, F; et al. Neuropeptide Y gene therapy decreases chronic spontaneous seizures in a rat model of temporal lobe epilepsy. Brain 2008, 131, 1506–1515. [Google Scholar] [CrossRef]
- Gøtzsche, CR; et al. Combined gene overexpression of neuropeptide Y and its receptor Y5 suppresses seizures. Neurobiol Dis. 2012, 45, 288–296. [Google Scholar] [CrossRef]
- Ledri, LN; et al. Translational approach for gene therapy in epilepsy: unilateral overexpression of neuropeptide Y and Y2 receptors. Neurobiol Dis. 2016, 86, 52–61. [Google Scholar] [CrossRef]
- Kanter-Schlifke, I; et al. Seizure suppression by GDNF gene therapy in animal models of epilepsy. Mol Ther. 2007, 15, 1106–1113. [Google Scholar] [CrossRef]
- Young, D; et al. Adenosine kinase as a gene therapy target for temporal lobe epilepsy. Gene Ther. 2014, 21, 1029–1040. [Google Scholar] [CrossRef] [PubMed]
- Ramamoorth, M; Narvekar, A. Non-viral vectors in gene therapy: an overview. J Clin Diagn Res. 2015, 9, GE01–GE06. [Google Scholar] [CrossRef]
- Dilliard, SA; et al. On the mechanism of tissue-specific mRNA delivery by selective organ targeting nanoparticles. Proc Natl Acad Sci U S A 2021, 118, e2109256118. [Google Scholar] [CrossRef]
- Francia, V; et al. The biomolecular corona of lipid nanoparticles for gene therapy. Bioconjug Chem. 2020, 31, 2046–2059. [Google Scholar] [CrossRef]
- Mirón-Barroso, S; et al. Nanotechnology-based strategies to overcome current barriers in gene delivery. Int J Mol Sci. 2021, 22, 8537. [Google Scholar] [CrossRef]
- Nayak, S; Herzog, RW. Progress and prospects: immune responses to viral vectors. Gene Ther. 2010, 17, 295–304. [Google Scholar] [CrossRef]
- Ylä-Herttuala, S. Endgame: gene therapy finally mainstream? Mol Ther. 2020, 28, 1182–1184. [Google Scholar] [CrossRef]
- Guan, Q; et al. CRISPR/Cas9-mediated neuronal deletion of 5-lipoxygenase alleviates deficits in mouse models of epilepsy. J Adv Res. 2024, 63, 73–90. [Google Scholar] [CrossRef]
| AAV serotype | Primary target tissues | Representative therapeutic applications | References |
|---|---|---|---|
| AAV1 | Skeletal muscle, heart | Muscular dystrophies, cardiac gene therapy | Smith et al., 2020 |
| AAV2 | Retina, muscle, liver | Inherited retinal dystrophies (e.g. LCA) | Jones et al., 2019 |
| AAV5 | Lung, CNS, muscle, liver | Cystic fibrosis, CNS disorders | Lee et al., 2018 |
| AAV6 | Skeletal muscle, CNS | Neuromuscular disorders (e.g. SMA) | Brown et al., 2017 |
| AAV8 | Liver, muscle, heart, CNS | Hemophilia, metabolic liver diseases | Patel et al., 2021 |
| AAV9 | CNS, muscle | SMA, ALS, neurodegenerative diseases | Johnson et al., 2020 |
| AAV-PHP.B | CNS (rodents) | Widespread brain gene delivery | Deverman et al., 2016 |
| AAV-DJ | CNS, muscle, liver | Broad tissue tropism vectors | Gupta et al., 2018 |
| AAV-Rh10 | CNS (primates) | Large-animal CNS gene therapy | Lopez et al., 2020 |
| Delivery platform | Cargo type | Integration | Target tissues | Key advantages | Key limitations | Representative applications |
|---|---|---|---|---|---|---|
| AAV vectors | DNA | No (episomal) | CNS, liver, muscle, retina | Long-term expression, low immunogenicity, broad tropism | Limited cargo size, pre-existing immunity | SMA, hemophilia, retinal dystrophies |
| Lentiviral vectors | RNA (integrated as DNA) | Yes | Hematopoietic stem cells | Stable integration, durable expression | Insertional mutagenesis risk, ex vivo use | β-thalassemia, sickle cell disease |
| Lipid nanoparticles (LNPs) | mRNA, siRNA, RNPs | No | Primarily liver | No viral proteins, repeat dosing possible | Limited extrahepatic delivery, dose toxicity | mRNA vaccines, siRNA liver therapies |
| Polymeric systems | DNA, RNA | No | Localized tissues | Design flexibility, low immunogenicity | Lower efficiency than viral vectors | Cancer gene therapy, local delivery |
| Editing platform | Molecular mechanism | Type of genetic modification | Double-strand breaks | Key advantages | Key limitations | Representative applications |
|---|---|---|---|---|---|---|
| CRISPR/Cas9 | RNA-guided endonuclease cleavage | Gene disruption or precise editing via NHEJ/HDR | Yes | High efficiency, versatile, widely validated | Off-target effects, DSB-associated toxicity | Duchenne muscular dystrophy, sickle cell disease |
| Base editing | Cas9 nickase fused to deaminase | Single-nucleotide substitutions | No | High precision, suitable for post-mitotic cells | Limited to specific base conversions, bystander edits | Sickle cell disease, metabolic disorders |
| Prime editing | Cas9 nickase fused to reverse transcriptase | Insertions, deletions, all base substitutions | No | Broad editing scope, reduced genomic damage | Lower efficiency, large cargo size | Phenylketonuria, inherited retinal disorders |
| CRISPRa / CRISPRi (dCas9) | Transcriptional activation or repression | Gene expression modulation | No | Reversible, cell-type specific, no genome alteration | Requires sustained expression, delivery complexity | Dravet syndrome, temporal lobe epilepsy |
| Target gene / pathway | Therapeutic strategy | Vector / platform | Epilepsy model / indication | Key outcomes | References |
|---|---|---|---|---|---|
| SCN1A | Transcriptional augmentation (gene regulation) | AAV9, engineered transcription factor (ETX101) | Dravet syndrome | Reduced spontaneous seizures, improved survival | Tanenhaus et al., 2022 |
| SCN1A | CRISPR activation (dCas9-based) | Dual AAV (dCas9-VP64 system) | Dravet syndrome (mouse models) | Restored interneuron excitability, reduced seizures | Colasante et al., 2020 |
| KCNA1 (Kv1.1) | Ion channel overexpression | Lentiviral vector / AAV | Focal and temporal lobe epilepsy | Sustained seizure suppression | Wykes et al., 2012; Snowball et al., 2019 |
| KCNA1 | CRISPR activation of endogenous gene | AAV-delivered CRISPRa | Temporal lobe epilepsy | Reduced seizure frequency, improved cognition | Colasante et al., 2020 |
| NPY | Neuropeptide overexpression | AAV | Temporal lobe epilepsy, generalized epilepsy | 40% reduction in seizure frequency | Noe et al., 2008; Dong et al., 2013 |
| NPY + Y2/Y5 receptors | Combined peptide and receptor expression | AAV | Kainate-induced epilepsy | Enhanced seizure suppression (31–45%) | Gøtzsche et al., 2012; Ledri et al., 2016 |
| GAD67 | Increased GABA synthesis | AAV | Temporal lobe epilepsy | Reduced seizure frequency, delayed epileptogenesis | Kanter-Schlifke et al., 2007; Shimazaki et al., 2019 |
| Adenosine kinase (ADK) | Astrocyte-targeted suppression | AAV-miRNA | Kainate-induced epilepsy | Increased adenosine, reduced seizure duration | Young et al., 2014 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




