Submitted:
03 February 2026
Posted:
05 February 2026
You are already at the latest version
Abstract
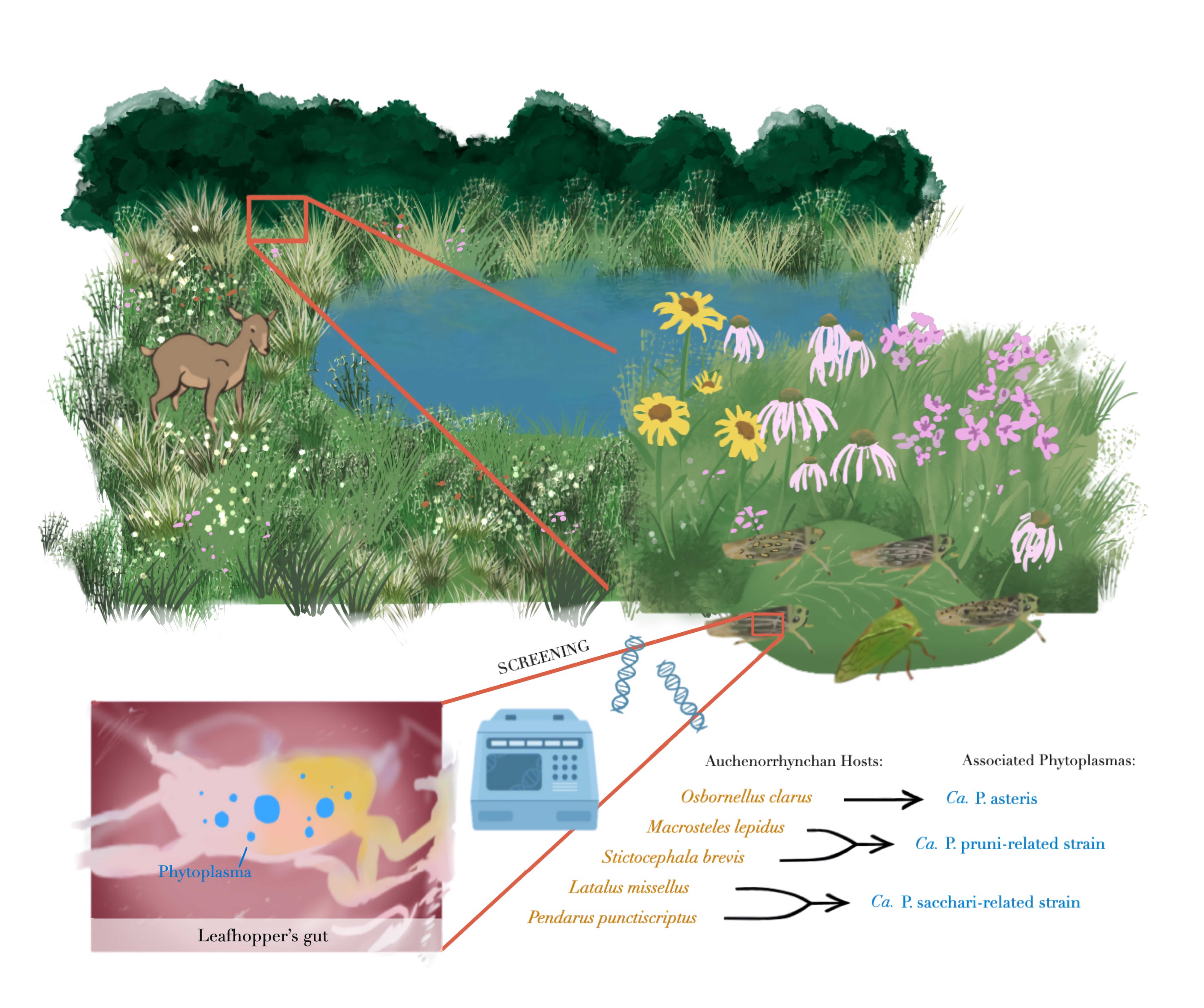
Keywords:
1. Introduction
2. Materials and Methods
2.1. Leveraged Monitoring Programs and Specimen Collection
2.2. Non-Destructive DNA Extraction
2.3. Voucher Preparation, Host Association and Metadata Integration
2.4. Real-Time PCR Screening for Phytoplasmas and Conventional PCR Assays
2.5. Sequencing, Phytoplasma Characterization and Phylogenetic Analysis
3. Results
3.1. Phytoplasma Screening and Insect Museum Vouchers
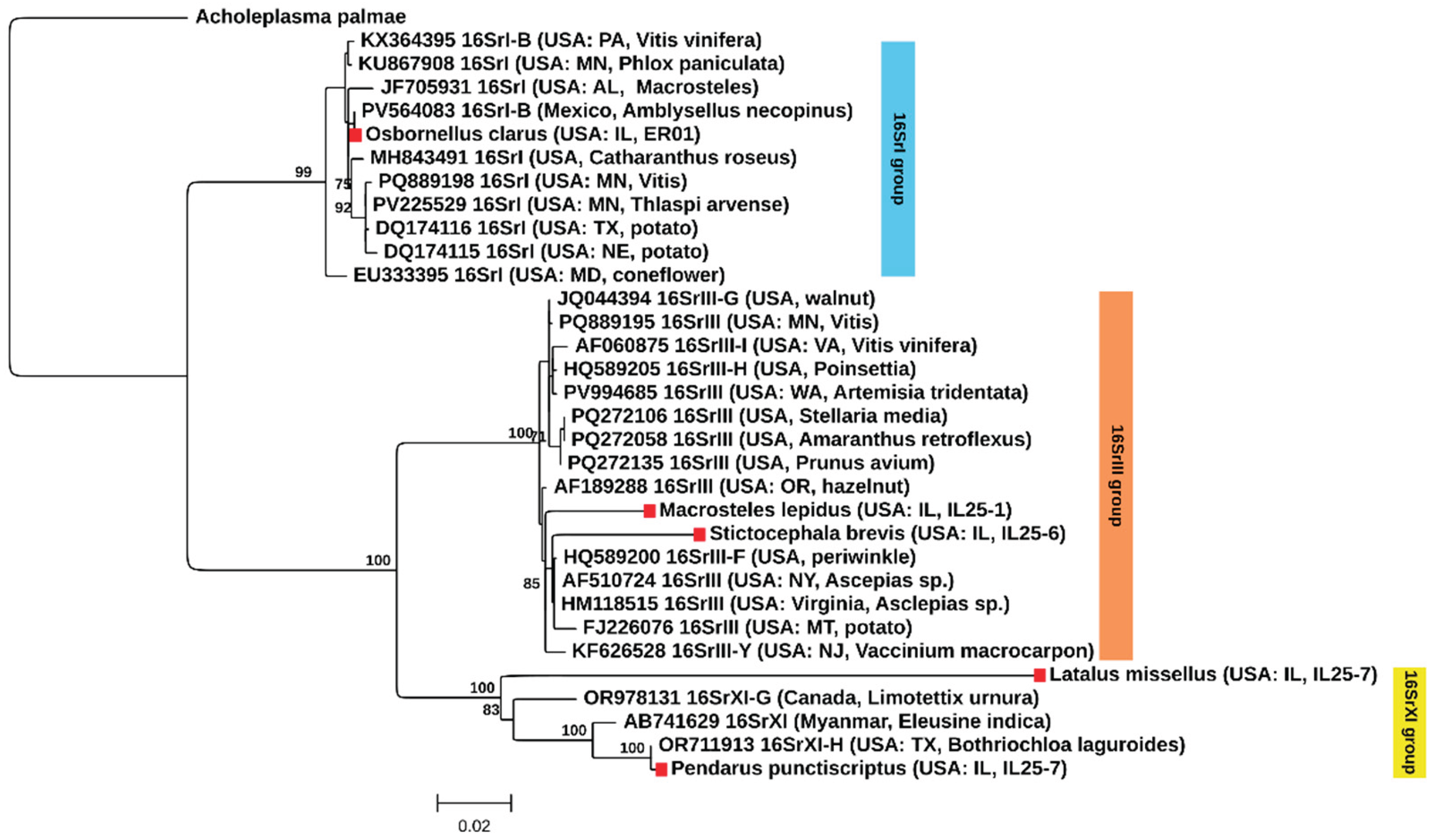
3.2. BLAST Analysis, Phytoplasma Identification, and Phylogenetic Analysis
3.3. Novel Insect-Phytoplasma Associations in Natural Areas
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- One Health Joint Plan of Action (2022‒2026): Working Together for the Health of Humans, Animals, Plants and the Environment Available online: https://www.who.int/publications/i/item/9789240059139 (accessed on 17 December 2025). [CrossRef]
- Giraudoux, P.; Besombes, C.; Bompangue, D.; Guégan, J.-F.; Mauny, F.; Morand, S. One Health or ‘One Health Washing’? An Alternative to Overcome Now More than Ever. CABI One Health 2022, 2022. [CrossRef]
- Panel (OHHLEP), O.H.H.-L.E.; Adisasmito, W.B.; Almuhairi, S.; Behravesh, C.B.; Bilivogui, P.; Bukachi, S.A.; Casas, N.; Becerra, N.C.; Charron, D.F.; Chaudhary, A.; et al. One Health: A New Definition for a Sustainable and Healthy Future. PLOS Pathogens 2022, 18, e1010537. [CrossRef]
- Stockly, O. The Epistemology of Ignorance. Anthós 2011, 3. [CrossRef]
- Furberg, E. Are We Persons or Animals? Exposing an Anthropocentric Bias and Suggesting a Hybrid View. Ethics, Medicine and Public Health 2017, 3, 279–287. [CrossRef]
- Barroso, P.; Relimpio, D.; Zearra, J.A.; Cerón, J.J.; Palencia, P.; Cardoso, B.; Ferreras, E.; Escobar, M.; Cáceres, G.; López-Olvera, J.R.; et al. Using Integrated Wildlife Monitoring to Prevent Future Pandemics through One Health Approach. One Health 2023, 16, 100479. [CrossRef]
- Cardoso, B.; García-Bocanegra, I.; Acevedo, P.; Cáceres, G.; Alves, P.C.; Gortázar, C. Stepping up from Wildlife Disease Surveillance to Integrated Wildlife Monitoring in Europe. Research in Veterinary Science 2022, 144, 149–156. [CrossRef]
- Machalaba, C.; Uhart, M.; Ryser-Degiorgis, M.-P.; Karesh, W.B. Gaps in Health Security Related to Wildlife and Environment Affecting Pandemic Prevention and Preparedness, 2007–2020. Bull. World Health Organ. 2021, 99, 342-350B. [CrossRef]
- Mackay, A.J.; Cara, C.; Kim, C.-H.; Stone, C.M. Update on the Distribution of Culiseta Melanura in Regions of Illinois with Prior Eastern Equine Encephalitis Virus Activity. Journal of Vector Ecology 2022, 47. [CrossRef]
- Brown, M.E.; Ottati, S.; Trivellone, V. A Non-Destructive, Fast, Inexpensive, Non-Toxic Chelating Resin-Based DNA Extraction Protocol for Insect Voucher Specimens and Associated Microbiomes. Journal of Insect Science 2025, 25, 17. [CrossRef]
- McElrath, T. Illinois Natural History Survey Insect Collection. Illinois Natural History Survey. Occurrence Dataset. Available online: (accessed on 2 January 2026). [CrossRef]
- Deng, S.; Hiruki, C. Amplification of 16S rRNA Genes from Culturable and Nonculturable Mollicutes. Journal of Microbiological Methods 1991, 14, 53–61. [CrossRef]
- Smart, C.D.; Schneider, B.; Blomquist, C.L.; Guerra, L.J.; Harrison, N.A.; Ahrens, U.; Lorenz, K.H.; Seemüller, E.; Kirkpatrick, B.C. Phytoplasma-Specific PCR Primers Based on Sequences of the 16S-23S rRNA Spacer Region. Appl Environ Microbiol 1996, 62, 2988–2993.
- Lee, I.M.; Bertaccini, A.; Vibio, M.; Gundersen, D.E. Detection of Multiple Phytoplasmas in Perennial Fruit Trees with Decline Symptoms in Italy. Phytopathology 1995, 85, 728–735. [CrossRef]
- Bonfield, J.K.; Smith, K. f; Staden, R. A New DNA Sequence Assembly Program. Nucleic Acids Res 1995, 23, 4992–4999.
- Wright, E.S. Fast and Flexible Search for Homologous Biological Sequences with DECIPHER V3. The R Journal 2025, 16, 191–200. [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [CrossRef]
- Edgar, R.C. MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Res 2004, 32, 1792–1797. [CrossRef]
- Edler, D.; Klein, J.; Antonelli, A.; Silvestro, D. raxmlGUI 2.0: A Graphical Interface and Toolkit for Phylogenetic Analyses Using RAxML. Methods in Ecology and Evolution 2021, 12, 373–377. [CrossRef]
- Hribar, L.J. Expanding Basic Entomological Knowledge by Using Mosquito Surveillance Bycatch. esaa 2020, 113, 439–446. [CrossRef]
- Grootaert, P.; Loziaková Peňazziová, K.; Csank, T.; Pačanská, Z.; Pivka, S.; Oboňa, J. Bycatch with Benefits: CO₂ Traps Reveal New Hybotidae for Slovakia. Biodiversity & Environment 2025, 17, 15–18.
- Mullin, L.J. The Effects of Infusion Water Type and Fermaentation on Mosquito and Non-Target Organism Collected in the CDC’s Autocidal Gravid Ovitrap. J FL Mosq Control Assoc 2021, 67, 23–27. [CrossRef]
- Barringer, L.E.; Bartlett, C.R. Pennsylvania Planthoppers (Hemiptera: Auchenorrhyncha: Fulgoroidea): Relative Abundance and Incidental Catch Using Novel Trapping Methods. Insecta Mundi 2018.
- Urfer, K.; König, N.; Kranz-Baltensperger, Y.; Klopfstein, S.; Niehuis, O.; Von Virag, A.; Sann, M. Reclaiming the Unwanted: Mining Scientific Gold from Multi-Trap Bycatch. AlpEnt 2025, 9, 91–102. [CrossRef]
- Spears, L.R.; Christman, M.E.; Koch, J.B.U.; Looney, C.; Ramirez, R.A. A Review of Bee Captures in Pest Monitoring Traps and Future Directions for Research and Collaboration. Journal of Integrated Pest Management 2021, 12, 49. [CrossRef]
- Munhoz, E.M.; Pereira, T.B.C.; Silva, E.M.S.; Bedendo, I.P. A Subgroup 16SrIII-J Phytoplasma Associated with Aegiphila Verticillata, a Typical Wild Plant of Brazilian Savanna Region, Exhibiting Witches’ Broom Symptoms. Trop. plant pathol. 2016, 41, 52–56. [CrossRef]
- Jomantiene, R.; Postman, J.D.; Montano, H.G.; Maas, J.L.; Davis, R.E.; Johnson, K.B. First Report of Clover Yellow Edge Phytoplasma in Corylus (Hazelnut). Plant Disease 2000, 84, 102–102. [CrossRef]
- Rios, D.; Ueckert, J.; Ong, K.; Barillas, J.R.; Costanzo, S. First Report of a ‘ Candidatus Phytoplasma Sacchari’-Related Strain Associated with Yellowing and Decline of Silver Bluestem in Texas, U.S.A. Plant Disease 2024, 108, 2216. [CrossRef]
- Beirne, B.P. The Nearctic Species Of. The Canadian Entomologist 1952, 84, 208–232. [CrossRef]
- Wallner, A. Evaluating North American Tallgrass Prairie Quality Using the Auchenorrhyncha Quality Index. PhD dissertation, University of Illinois at Urbana-Champaign: Urbana, 2010.
- Kopp, D.D.; Yonke, T.R. The Treehoppers of Missouri: Part 2. Subfamily Smiliinae; Tribes Acutalini, Ceresini, and Polyglyptini (Homoptera: Membracidae). Journal of the Kansas Entomological Society 1973, 233–276.
- Beamer, R.H. A Review of the Genus Osbornellus in the United States and Canada (Homoptera-Cicadellidae). Journal of the Kansas Entomological Society 1937, 10, 89–112.
- DeLong, D.M. Twelve New Species of Osbornellus (Homoptera-Cicadellidae) from Mexico, Guatemala, and Panama. Bulletin of the Brooklyn Entomological Society 1942, 92–101.
- Blondet, L.; Mouanda Sounda, A.; Fritz, M.; Filloux, D.; Yvon, M.; Blanc, S.; Varsani, A.; Niama, R.F.; Leroy, E.M.; Roumagnac, P. Sentinels Tracking Viruses in Various Ecosystems: Towards a One Health Approach. PLoS Pathog 2025, 21, e1013141. [CrossRef]
- Weintraub, P.G.; Trivellone, V.; Krüger, K. The Biology and Ecology of Leafhopper Transmission of Phytoplasmas. In Phytoplasmas: Plant Pathogenic Bacteria - II: Transmission and Management of Phytoplasma - Associated Diseases; Bertaccini, A., Weintraub, P.G., Rao, G.P., Mori, N., Eds.; Springer: Singapore, 2019; pp. 27–51 ISBN 9789811328329.
- Trivellone, V.; Ripamonti, M.; Angelini, E.; Filippin, L.; Rossi, M.; Marzachí, C.; Galetto, L. Evidence Suggesting Interactions between Immunodominant Membrane Protein Imp of Flavescence Dorée Phytoplasma and Protein Extracts from Distantly Related Insect Species. J. Appl. Microbiol. 2019, 127, 1801–1813. [CrossRef]
- Chiykowski, L.N.; Chapman, R.K. Migration of the Six-Spotted Leafhopper in Central North America. Wisconsin Agricultural Experiment Station Research Bulletin 1965, 261, 21–45.
- Clements, J.; Garcia, M.; Bradford, B.; Crubaugh, L.; Piper, S.; Duerr, E.; Zwolinska, A.; Hogenhout, S.; Groves, R.L. Genetic Variation Among Geographically Disparate Isolates of Aster Yellows Phytoplasma in the Contiguous United States. J Econ Entomol 2020. [CrossRef]
- Frost, K.E.; Esker, P.D.; Van Haren, R.; Kotolski, L.; Groves, R.L. Seasonal Patterns of Aster Leafhopper (Hemiptera: Cicadellidae) Abundance and Aster Yellows Phytoplasma Infectivity in Wisconsin Carrot Fields. Environ Entomol 2013, 42, 491–502. [CrossRef]
- Fontes, E.M.G.; Habeck, D.H.; Slansky Jr, F. Phytophagous Insects Associated with Goldenrods (Solidago Spp.) in Gainesville, Florida. Florida Entomologist 1994, 209–221.
- Rizza, S.; Antonella, P.; Vera, D.; Ernesto, R.; Cristina, M.; Tessitori, M. Transmission of ‘Candidatus Phytoplasma Asteris’ (16SrI) by Osbornellus Horvathi (Matsumura 1908) Co-Infected with “Ca. Phytoplasma Phoenicium” (16SrIX). Phytoparasitica 2016, 44, 491–500. [CrossRef]
- Jensen, D.D. Transmission of Peach Yellow Leaf Roll Virus by Fieberiella Florii (Stål) and a New Vector, Osbornellus Borealis DeL. & M.1. J Econ Entomol 1957, 50, 668–672. [CrossRef]
- Ghauri, M.S.K. The Identity of a Suspected Vector of Coconut Lethal Yellowing Disease in Jamaica and Notes on Caribovia Intensa (Walker) (Homoptera: Cicadelloidea). Bulletin of Entomological Research 1980, 70, 411–415. [CrossRef]
- Trivellone, V.; Cao, Y.; Dietrich, C.H. Comparison of Traditional and Next-Generation Approaches for Uncovering Phytoplasma Diversity, with Discovery of New Groups, Subgroups and Potential Vectors. Biology 2022, 11, 977. [CrossRef]
- Brcak, J. Leafhopper and Planthopper Vectors of Plant Disease Agents in Central and Southern Europe. In Leafhopper Vectors and Plant Disease Agents; Academic Press: London, 1979; pp. 97–146.
- Fránová, J.; Špak, J.; Šimková, M. First Report of a 16SrIII-B Subgroup Phytoplasma Associated with Leaf Reddening, Virescence and Phyllody of Purple Coneflower. Eur J Plant Pathol 2013, 136, 7–12. [CrossRef]
- Fischnaller, S.; Parth, M.; Messner, M.; Stocker, R.; Kerschbamer, C.; Janik, K. Surveying Potential Vectors of Apple Proliferation Phytoplasma: Faunistic Analysis and Infection Status of Selected Auchenorrhyncha Species. Insects 2020, 12, 12. [CrossRef]
- Duduk, B.; Peri, P.; Mar, D.; Drobnjakovi, T.; Picciau, L.; Alma, A.; Bertaccini, A. Phytoplasmas in Carrots: Disease and Potential Vectors in Serbia. Bulletin of Insectology 2008, 61, 327–331.
- Quaglino, F.; Sanna, F.; Moussa, A.; Faccincani, M.; Passera, A.; Casati, P.; Bianco, P.A.; Mori, N. Identification and Ecology of Alternative Insect Vectors of ‘Candidatus Phytoplasma Solani’ to Grapevine. Sci Rep 2019, 9, 19522. [CrossRef]
- Molnar, C.; Shires, M.K.; Wright, A.A.; Hoskins, M.C.; Cowell, S.J.; Nikolaeva, E.V.; Knier, R.; Nouri, M.T.; Black, B.; Harper, S.J. Putting ‘X’ into Context: The Diversity of ‘ Candidatus Phytoplasma Pruni’ Strains Associated with the Induction of X-Disease. Plant Disease 2024, 108, 2677–2687. [CrossRef]
- Harper, S.J.; Northfield, T.D.; Nottingham, L.B.; DuPont, S.T.; Thompson, A.A.; Sallato, B.V.; Serban, C.F.; Shires, M.K.; Wright, A.A.; Catron, K.A.; et al. Recovery Plan for X-Disease in Stone Fruit Caused by ‘ Candidatus Phytoplasma Pruni.’ Plant Health Progress 2023, 24, 258–295. [CrossRef]
- Cooper, W.R.; Marshall, A.T.; Lewis, T.M.; Cook, R.; Horton, D.R.; Pitt, W.J.; Corral, C.R.; Jocson, D.M.I.; Gardner, J.D.; Zack, R.S.; et al. Biosystematic Classification of Colladonus Ball Vectors of “Candidatus Phytoplasma Pruni” in the Pacific Northwest (Hemiptera: Cicadellidae: Deltocephalinae). went 2025, 127, 141–161. [CrossRef]
- Trivellone, V.; Wei, W.; Filippin, L.; Dietrich, C.H. Screening Potential Insect Vectors in a Museum Biorepository Reveals Undiscovered Diversity of Plant Pathogens in Natural Areas. Ecol. Evol. 2021, 11, 6493–6503. [CrossRef]
- Duran-Ahumada, S.; Petersen, V.; Futo, M.; Zamy, M.; Pereira, T.; Burini, B.C. The Toolbox for Mosquito Vector Research. Parasites Vectors 2025, 18, 389. [CrossRef]
- Kim, K.; Huang, Q.; Lei, C. Advances in Insect Phototaxis and Application to Pest Management: A Review. Pest Management Science 2019, 75, 3135–3143. [CrossRef]
- Li, C.-X.; Smith, M.L.; Fulcher, A.; Kaufman, P.E.; Zhao, T.-Y.; Xue, R.-D. Field Evaluation of Three New Mosquito Light Traps Against Two Standard Light Traps to Collect Mosquitoes (Diptera: Culicidae) and Non-Target Insects in Northeast Florida. Florida Entomologist 2015, 98, 114–117. [CrossRef]
- Acevedo, N. COMPARISON OF THE CDC LIGHT TRAP AND THE DYNATRAP® DT2000 FOR COLLECTION OF MOSQUITOES IN SEMI-FIELD AND FIELD SETTINGS. J FL Mosq Control Assoc 2021, 67, 69–72. [CrossRef]
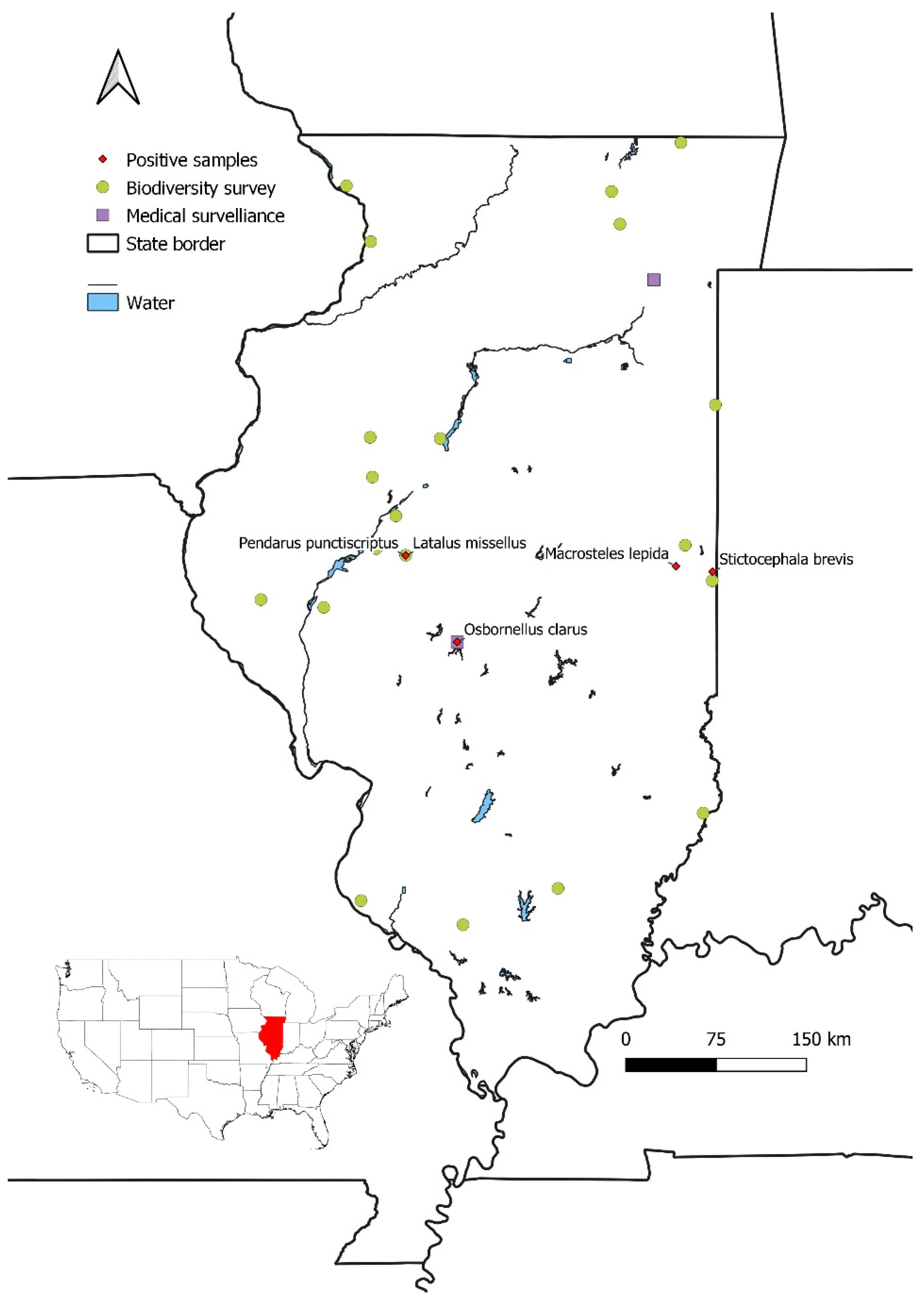
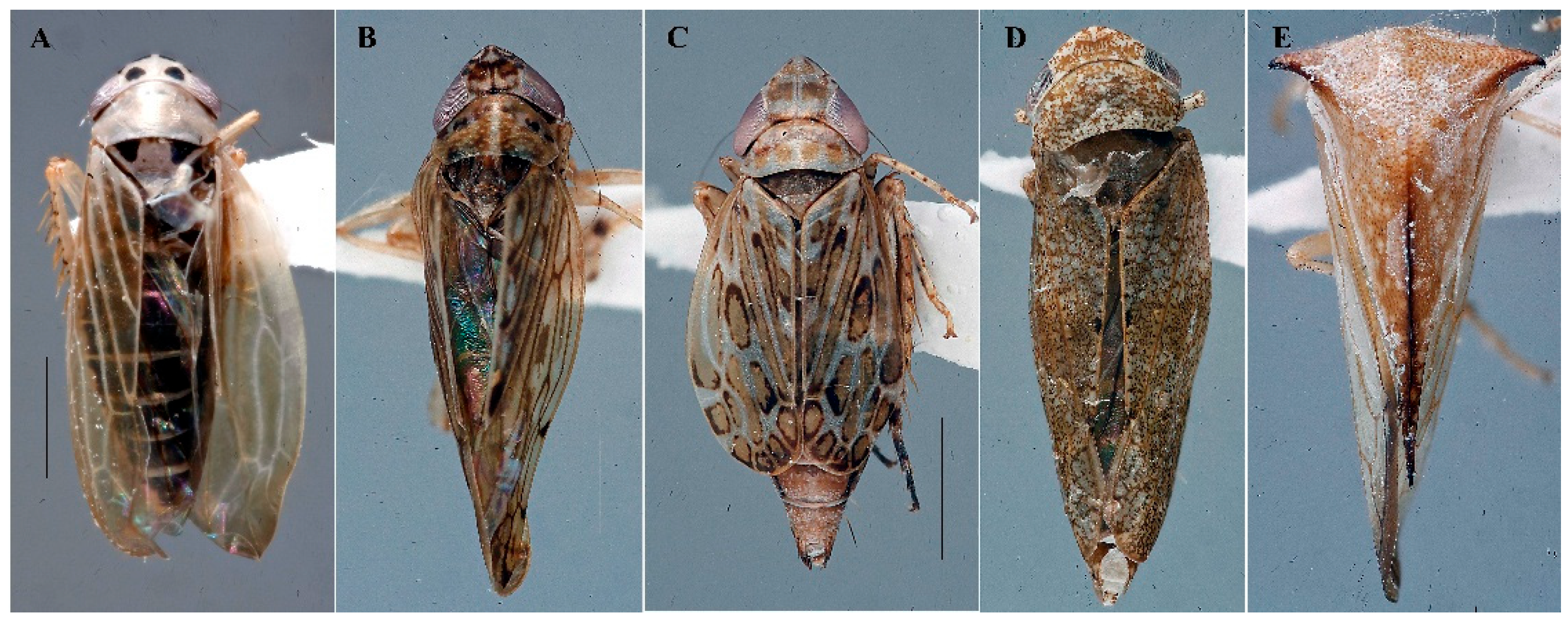
| Species | Coordinates | Date | INHS Insect coll. code | Associated Phy. | qPCR Ct value | NCBI accession |
| Macrosteles lepidus1 | 87°50'11"W 40°05'27"N | 06/12/2025 | 1086365 | 16SrIII | 21.97 | PX789936 |
| Osbornellus clarus2 | 89°28'08.9"W 39°39'26.9"N | 09/02/2022 | 1083960 | 16SrI-B | 24.02 | PX789937 |
| Latalus missellus1 | 89°51'12.5"W 40°09'09.6"N |
08/20/2025 | 1083966 | 16SrXI-H | 25.85 | PX789938 |
| Pendarus punctiscriptus1 | 89°51'12.5"W 40°09'09.6"N |
08/20/2025 | 1083969 | 16SrXI-H | 23.21 | PX789939 |
| Stictocephala brevis1 | 87°33'50.9"W 40°03'35.6"N |
07/29/2025 | 1089637 | 16SrIII | 19.08 | PX789940 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





