Submitted:
22 January 2026
Posted:
22 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. General Transcriptomic and Proteomic Statistics
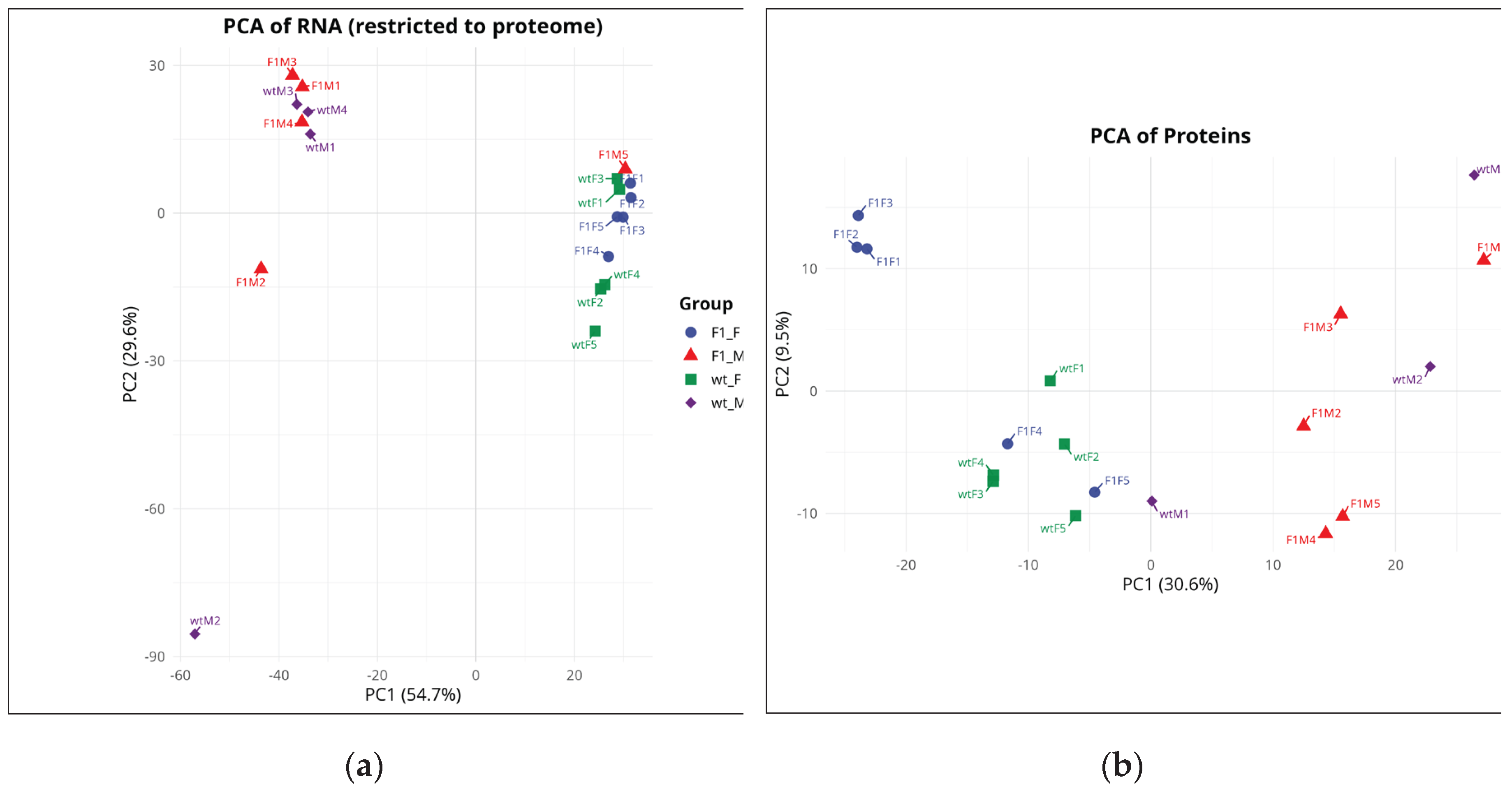
2.2. Principal Component Analysis (PCA) Schemes
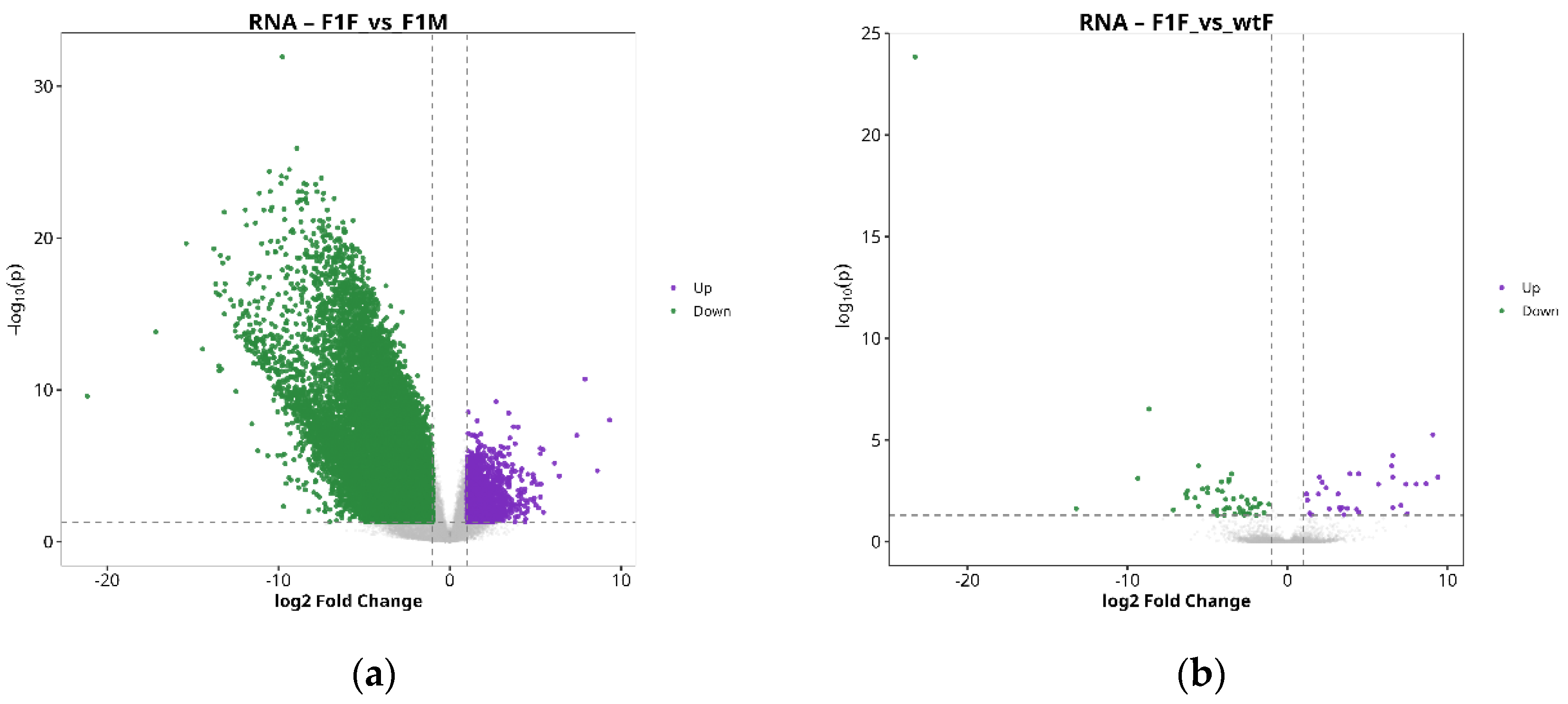
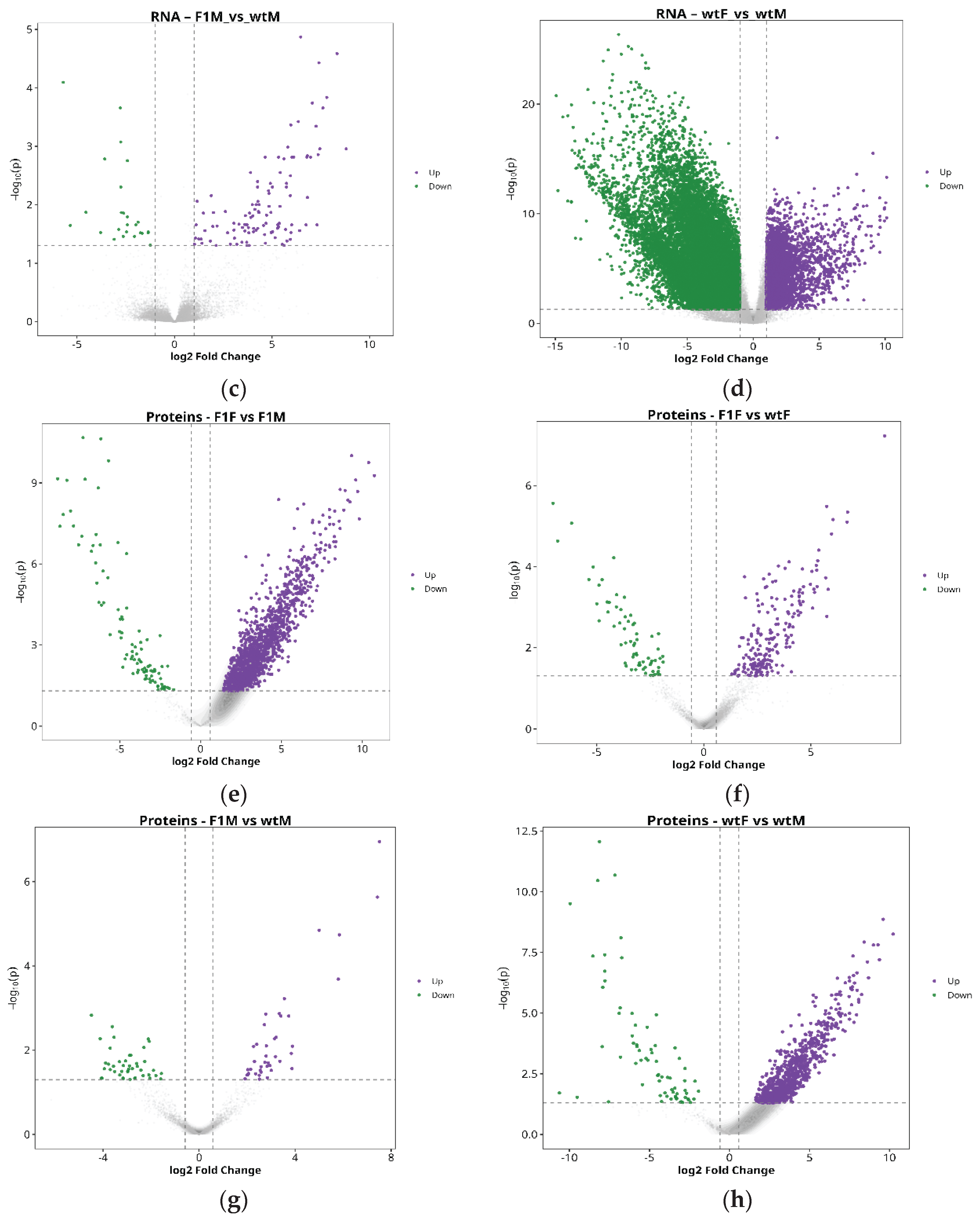
2.3. Identification of DEGs and DEPs
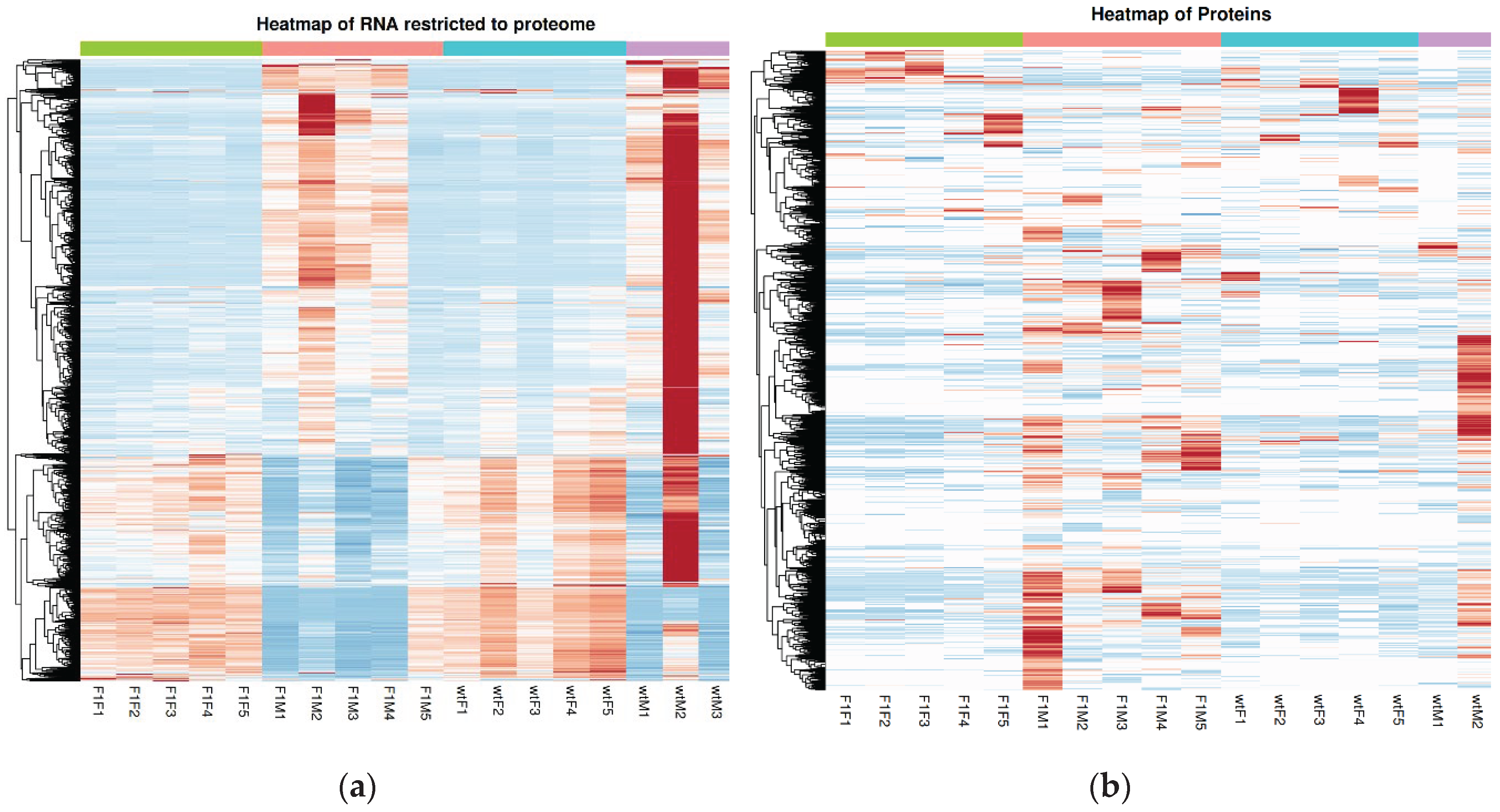
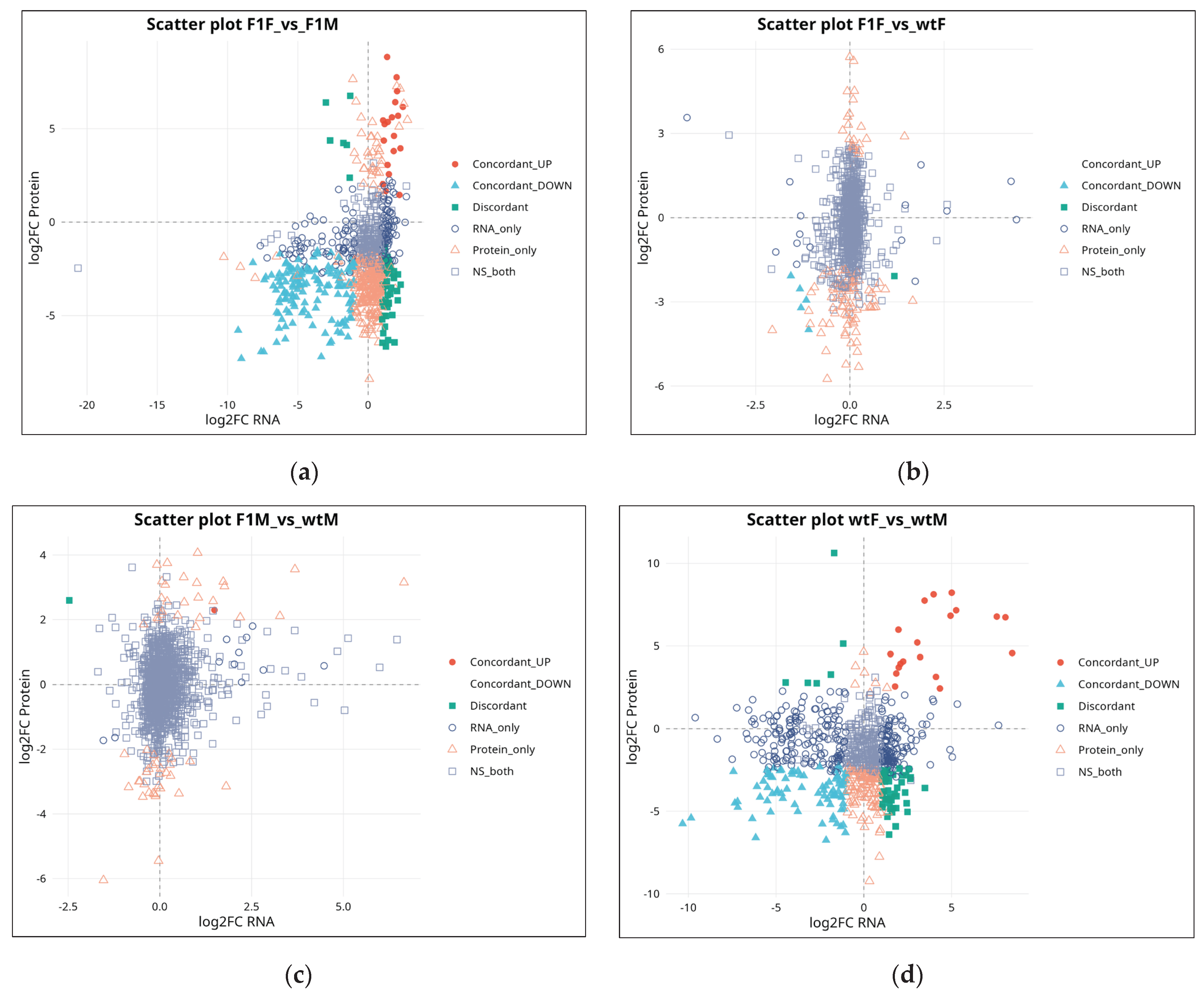
2.4. RNA–Protein Integration Analysis
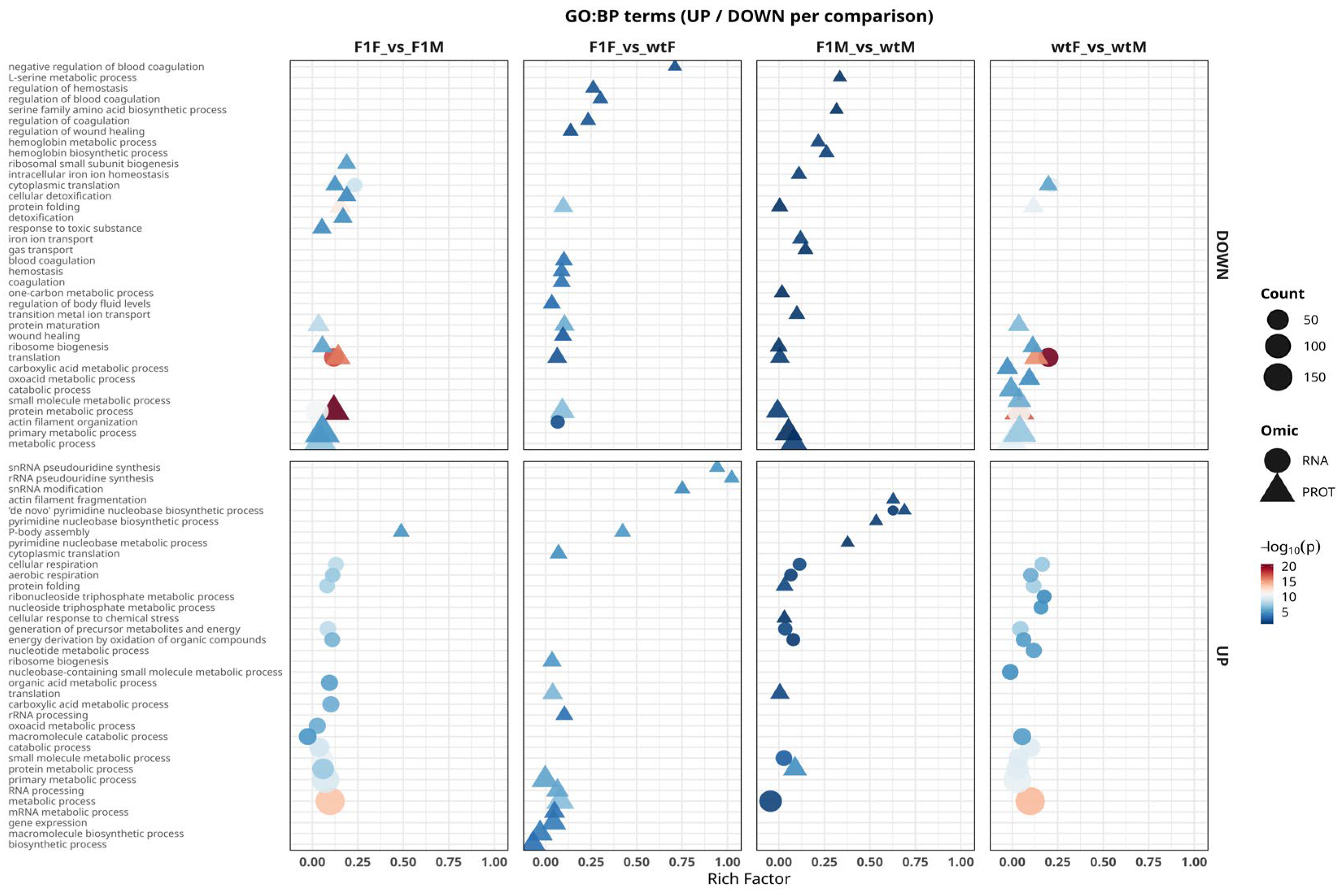
2.5. Functional Analysis Using Gene Ontology
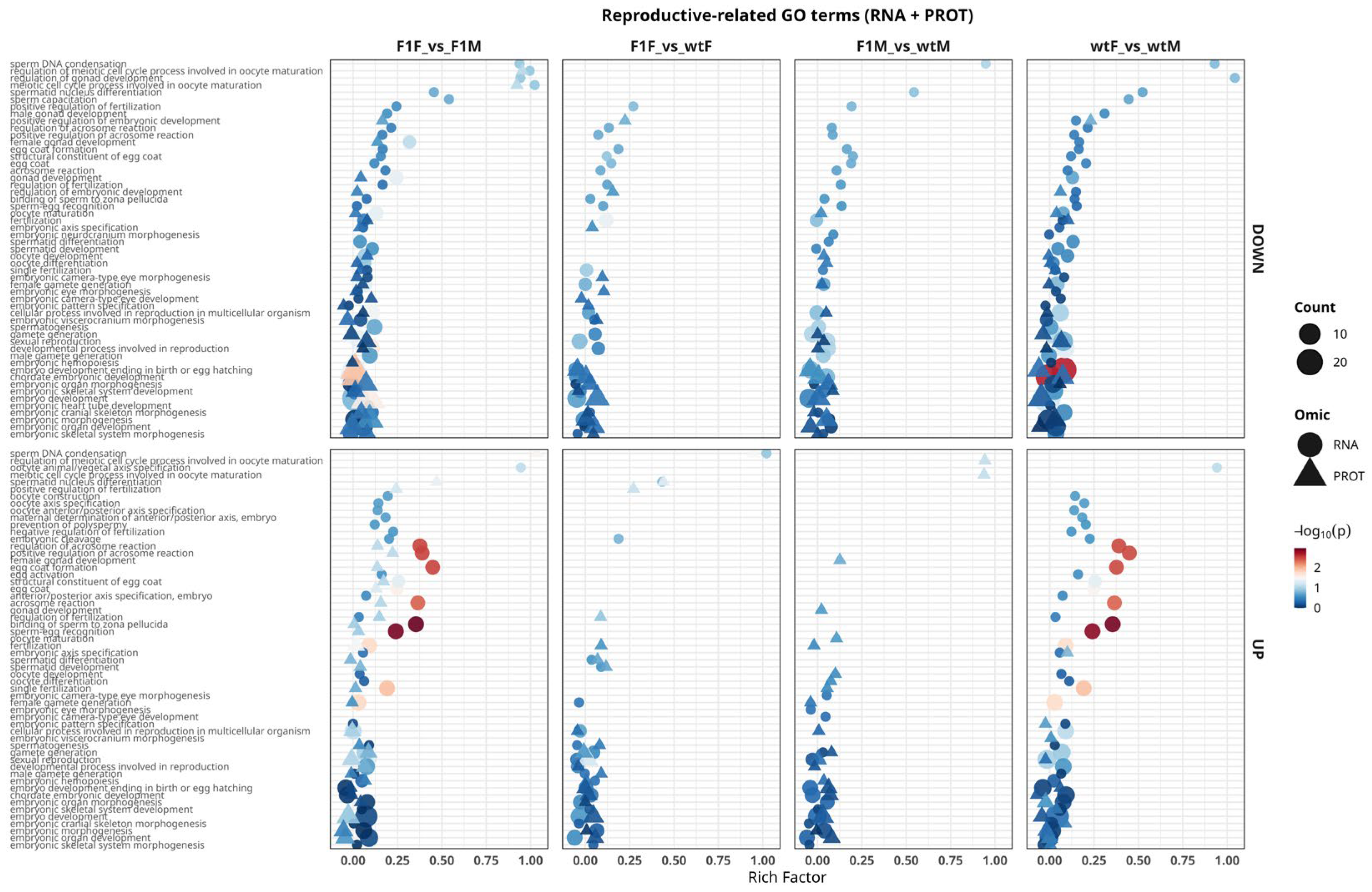
2.5.1. Functional Analysis of GO Terms Related to Reproductive Processes
2.5.2. RNA–Protein Concordance in Enriched GO Terms Associated with Reproductive Processes
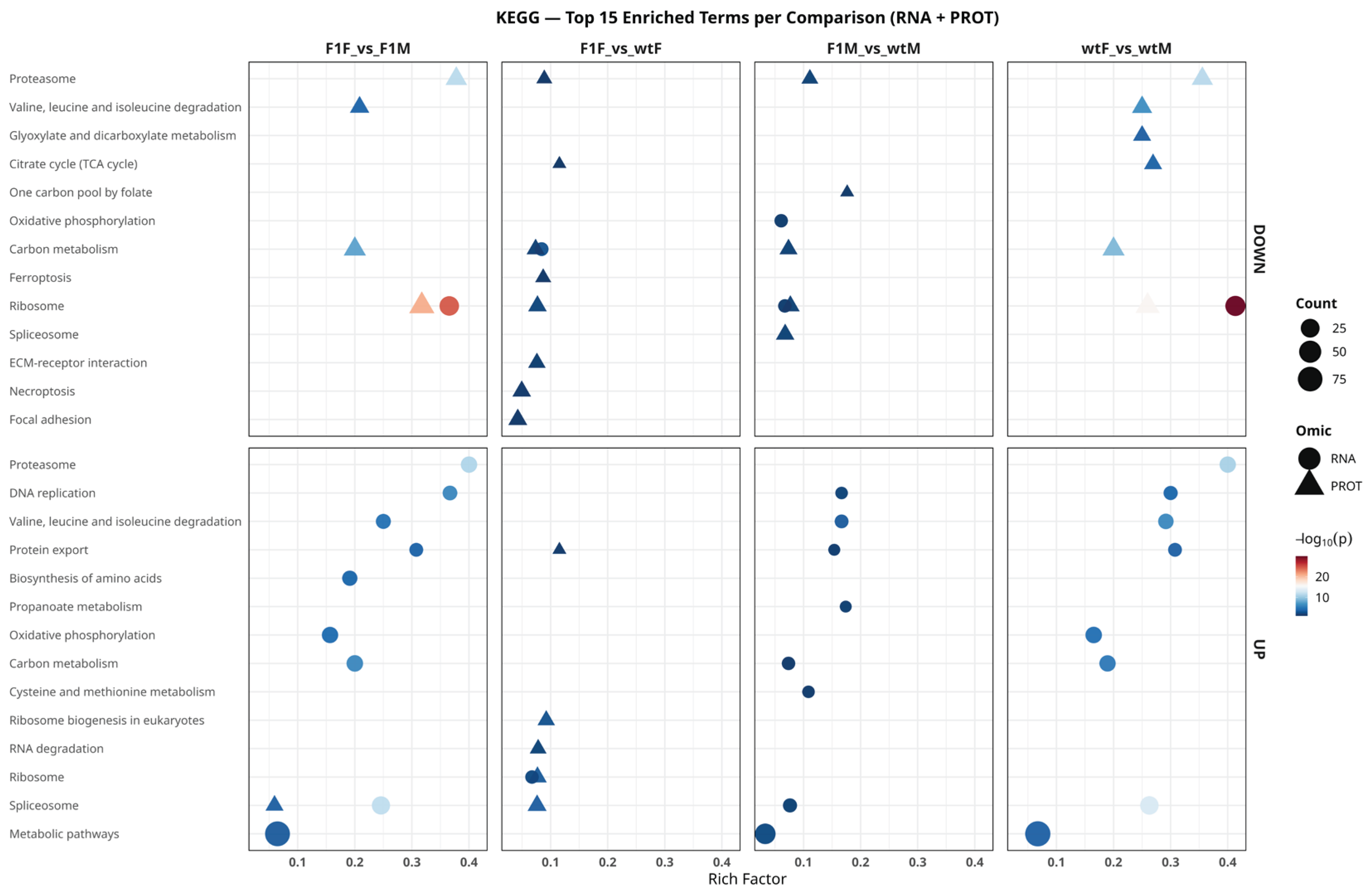
2.6. Functional Analysis Based on KEGG Pathways
2.7. DNA Methylation Landscape in Relation to Transcriptomic–Proteomic Patterns
3. Discussion
4. Materials and Methods
4.1. Extraction of RNA and Proteins from Gonadal Tissue
4.2. RNA Sequencing
4.3. Protein Analysis by LC-MS/MS
4.4. Bioinformatic Analysis for the Omics Integration and Functional Enrichment
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BP BCA |
Biological Process Bicinchoninic Acid |
| CC CpG |
Cellular Component Cytosine–phosphate–guanine dinucleotide |
| DEG | Differentially Expressed Gene |
| DEP DTT |
Differentially Expressed Protein Dithiothreitol |
| F1 F1F F1M |
First-generation captive-bred First-generation captive-bred Female First-generation captive-bred Male |
| FDR FPKM |
False Discovery Rate Fragments per Kilobase of transcript per Million mapped |
| GO IAA |
Gene Ontology Iodoacetamide |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| LC–MS/MS LFQ |
Liquid Chromatography–Tandem Mass Spectrometry Label-Free Quantification |
| MF | Molecular Function |
| NES | Normalized Enrichment Score |
| PCA | Principal Component Analysis |
| RNA-seq RRBS SDS-PAGE |
RNA sequencing Reduced Representation Bisulfite Sequencing Sodium Dodecyl Sulfate – Polyacrylamide Gel Electrophoresis |
| UP/DOWN | Overexpressed / Underexpressed |
| Wt wtF WTM |
Wild-type Wild-type Female Wild-type Male |
References
- Morais, S.; Aragão, C.; Cabrita, E.; Conceição, L.E.C.; Constenla, M.; Costas, B.; Dias, J.; Duncan, N.; Engrola, S.; Estevez, A.; Gisbert, E.; Mañanós, E.; Valente, L.M.P.; Yúfera, M.; Dinis, M.T. New developments and biological insights into the farming of Solea senegalensis reinforcing its aquaculture potential. Rev. Aquac. 2016, 8, 227–263. [Google Scholar] [CrossRef]
- Forné, I.; Castellana, B.; Marín-Juez, R.; Cerdà, J.; Abián, J.; Planas, J.V. Transcriptional and proteomic profiling of flatfish (Solea senegalensis) spermatogenesis. Proteomics 2011, 11, 2195–2211. [Google Scholar] [CrossRef] [PubMed]
- Honji, R.M.; de Mello, P.H.; Araújo, B.C. Reproduction and development in fish: Solving bottlenecks in modern aquaculture. Animals 2025, 15, 145. [Google Scholar] [CrossRef]
- Riesco, M.F.; Valcarce, D.G.; Martínez-Vázquez, J.M.; Martín, I.; Calderón-García, A.Á.; Gonzalez-Nunez, V.; Robles, V. Male reproductive dysfunction in Solea senegalensis: New insights into an unsolved question. Reprod. Fertil. Dev. 2019, 31, 1104–1115. [Google Scholar] [CrossRef] [PubMed]
- Cabrita, E.; Soares, F.; Dinis, M.T. Characterization of Senegalese sole (Solea senegalensis) male broodstock in terms of sperm production and quality. Aquaculture 2006, 261, 967–975. [Google Scholar] [CrossRef]
- de la Herrán, R.; Hermida, M.; Rubiolo, J.A.; Gómez-Garrido, J.; Cruz, F.; Robles, F.; Navajas-Pérez, R.; Blanco, A.; Villamayor, P.R.; Torres, D.; et al. A chromosome-level genome assembly enables the identification of the follicule stimulating hormone receptor as the master sex-determining gene in the flatfish Solea senegalensis. Mol. Ecol. Resour. 2023, 00, 1–19. Available online: http://hdl.handle.net/10347/30019. [CrossRef]
- Ramírez, D.; Anaya-Romero, M.; Rodríguez, M.E.; Arias-Pérez, A.; Mukiibi, R.; D’Cotta, H.; Robledo, D.; Rebordinos, L. Insights into Solea senegalensis reproduction through gonadal tissue methylation analysis and transcriptomic integration. Biomolecules 2025, 15, 54. [Google Scholar] [CrossRef]
- Forné, I.; María, J.A.; Asensio, E.; Abián, J.; Cerdà, J. 2-D DIGE analysis of Senegalese sole (Solea senegalensis) testis proteome in wild-caught and hormone-treated F1 fish. Proteomics 2009, 9, 2171–2181. [Google Scholar] [CrossRef]
- Li, B.; Zhang, Y.; Du, J.; Liu, C.; Zhou, G.; Li, M.; Yan, Z. Application of multi-omics techniques in aquatic ecotoxicology: A review. Toxics 2025, 13, 653. [Google Scholar] [CrossRef]
- Anaya-Romero, M.; Ramírez, D.; Arias-Pérez, A.; Rodríguez, M.E.; Robledo, D.; Rebordinos, L. Comparative transcriptomic profiling of gonads in Solea senegalensis: Exploring sex, maturity, and origin variations. Aquaculture 2025, 604, 742461. [Google Scholar] [CrossRef]
- Anaya-Romero, M.; Arias-Pérez, A.; Rodríguez, M.E.; Merlo, M.A.; Portela-Bens, S.; Cross, I.; Rebordinos, L. Comparative proteomic analysis of gonadal tissue in Solea senegalensis reveals reproductive deregulation associated with F1 individuals. Biomolecules preprint. 2026. [Google Scholar] [CrossRef]
- Tao, W.; Yuan, J.; Zhou, L.; Sun, L.; Sun, Y.; Yang, S.; Li, M.; Zeng, S.; Huang, B.; Wang, D. Characterization of gonadal transcriptomes from Nile tilapia (Oreochromis niloticus) reveals differentially expressed genes. PLoS ONE 2013, 8, e63604. [Google Scholar] [CrossRef] [PubMed]
- Sarropoulou, E.; Kaitetzidou, E.; Papandroulaki, N.; Tsalafouta, A.; Pavlidis, M. Inventory of European sea bass (Dicentrarchus labrax) sncRNAs vital during early teleost development. Front. Genet. 2019, 10, 657. [Google Scholar] [CrossRef]
- Clelland, E.; Peng, C. Endocrine/paracrine control of zebrafish ovarian development. Mol. Cell. Endocrinol. 2009, 312, 42–52. [Google Scholar] [CrossRef] [PubMed]
- Lubzens, E.; Young, G.; Bobe, J.; Cerdà, J. Oogenesis in teleosts: How fish eggs are formed. Gen. Comp. Endocrinol. 2010, 165, 367–389. [Google Scholar] [CrossRef]
- Park, J.-W.; Lee, Y.M.; Noh, J.K.; Kim, H.C.; Park, C.J.; Hwang, I.-J.; Kim, S.-Y.; Lee, J.H. The morphological study of wild and farmed olive flounder (Paralichthys olivaceus): The role of indirect selection within and between populations. Dev. Reprod. 2012, 16, 309–314. [Google Scholar] [CrossRef]
- Ponomarenko, E.A.; Krasnov, G.S.; Kiseleva, O.I.; Kryukova, P.A.; Arzumanian, V.A.; Dolgalev, G.V.; Ilgisonis, E.V.; Lisitsa, A.V.; Poverennaya, E.V. Workability of mRNA sequencing for predicting protein abundance. Genes 2023, 14, 2065. [Google Scholar] [CrossRef]
- Kuchta, K.; Towpik, J.; Biernacka, A.; Kutner, J.; Kudlicki, A.; Ginalski, K.; Rowicka, M. Predicting proteome dynamics using gene expression data. Sci. Rep. 2018, 8, 1–13. [Google Scholar] [CrossRef]
- Munro, V.; Kelly, V.; Messner, C.B.; Kustatscher, G. Cellular control of protein levels: A systems biology perspective. Proteomics 2024, 24, e2200220. [Google Scholar] [CrossRef]
- Liu, F.; Zhang, T.; He, Y.; Zhan, W.; Xie, Q.; Lou, B. Integration of transcriptome and proteome analyses reveals regulatory mechanisms of liver tissue under heat stress. Fish Shellfish Immunol. 2023, 135, 108704. [Google Scholar] [CrossRef]
- Wei, F.; Liang, J.; Tian, W.; Yu, L.; Feng, Z.; Hua, Q. Transcriptomic and proteomic analyses provide insights into adaptive responses to combined salinity and alkalinity stress. Bioresour. Bioprocess. 2022, 9, 1–22. [Google Scholar] [CrossRef]
- O’Donnell, L.; Stanton, P.; de Kretser, D.M. Endocrinology of the male reproductive system and spermatogenesis. Endotext. 2017. Available online: https://www.ncbi.nlm.nih.gov/books/NBK279031/.
- Gaboriau, D.; Howes, E.A.; Clark, J.; Jones, R. Binding of sperm proacrosin/β-acrosin to zona pellucida glycoproteins is sulfate- and stereodependent. Dev. Biol. 2007, 306, 646–657. [Google Scholar] [CrossRef] [PubMed]
- Solovova, O.A.; Chernykh, V.B. Genetics of oocyte maturation defects and early embryo development arrest. Genes 2022, 13, 1920. [Google Scholar] [CrossRef]
- Martín, I.; Carazo, I.; Rasines, I.; Rodríguez, C.; Fernández, R.; Martínez, P.; Norambuena, F.; Chereguini, O.; Duncan, N. Reproductive performance of captive Senegalese sole (Solea senegalensis) according to origin and gender. Span. J. Agric. Res. 2019, 17, e0608. [Google Scholar] [CrossRef]
- Chauvigné, F.; Parhi, J.; Ollé, J.; Cerdà, J. Dual estrogenic regulation of the nuclear progestin receptor and spermatogonial renewal during gilthead seabream spermatogenesis. Comp. Biochem. Physiol. A 2017, 206, 36–46. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, Q.; Xu, S.; Wang, Y.; Feng, C.; Zhao, C.; Song, Z.; Li, J. A deep insight into spermatogenesis and hormone levels of aquacultured turbot (Scophthalmus maximus). Front. Mar. Sci. 2021, 7, 592880. [Google Scholar] [CrossRef]
- Viñas, J.; Piferrer, F. Stage-specific gene expression during fish spermatogenesis as determined by laser-capture microdissection and quantitative-PCR in sea bass gonads. Biol. Reprod. 2008, 79, 738–747. [Google Scholar] [CrossRef]
- Gosz, E.; Horbowy, J.; Ruczyńska, W.; Zietara, M.S. Enzymatic activities in spermatozoa and butyltin concentrations in Baltic turbot (Scophthalmus maximus). Mar. Environ. Res. 2011, 72, 188–195. [Google Scholar] [CrossRef]
- Wu, Q.; Song, Z.; Wang, L.; Wu, Z.; Zou, C.; Shu, C.; Liang, S.; Wang, W.; Sun, Y.; Yue, X.; Peng, Q.; You, F. Comparative study on gonadal development in diploid and artificially induced triploid olive flounder (Paralichthys olivaceus). Aquaculture 2023, 565, 739106. [Google Scholar] [CrossRef]
| Samples | RNA-seq | Proteomics | Groups |
| F1F1, F1F2, F1F3, F1F4, F1F5 | 5 | 5 | F1F |
| F1M1, F1M2, F1M3, F1M4, F1M5 | 5 | 5 | F1M |
| wtF1, wtF2, wtF3, wtF4, wtF5 | 5 | 5 | wtF |
| wtM1, wtM2, wtM3, wtM4 | 4 | 3 | wtM |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




