Submitted:
13 January 2026
Posted:
16 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
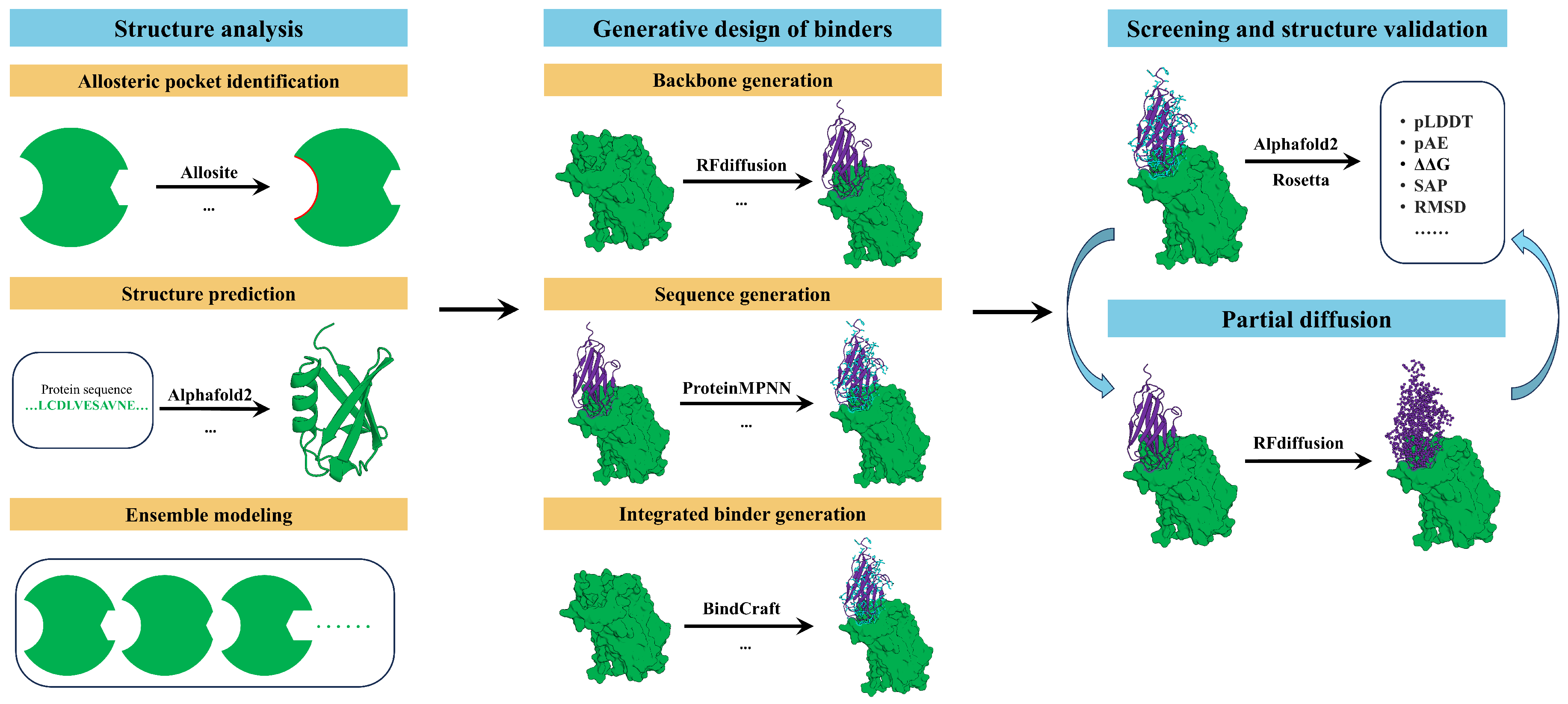
2. AI-Driven Pipeline for Designing Allosteric Miniprotein Modulators
2.1. Structure Analysis
2.1.1. Allosteric Pocket Identification
2.1.2. Structure Prediction and Ensemble Modeling
2.2. Generative Design of Binders
2.2.1. Backbone Generation
2.2.2. Sequence Design
2.2.3. Integrated Binder Generation
2.3. Screening and Structure Validation
2.4. Partial Diffusion
3. Latest Case Study in AI-Driven Design of Miniprotein Modulators
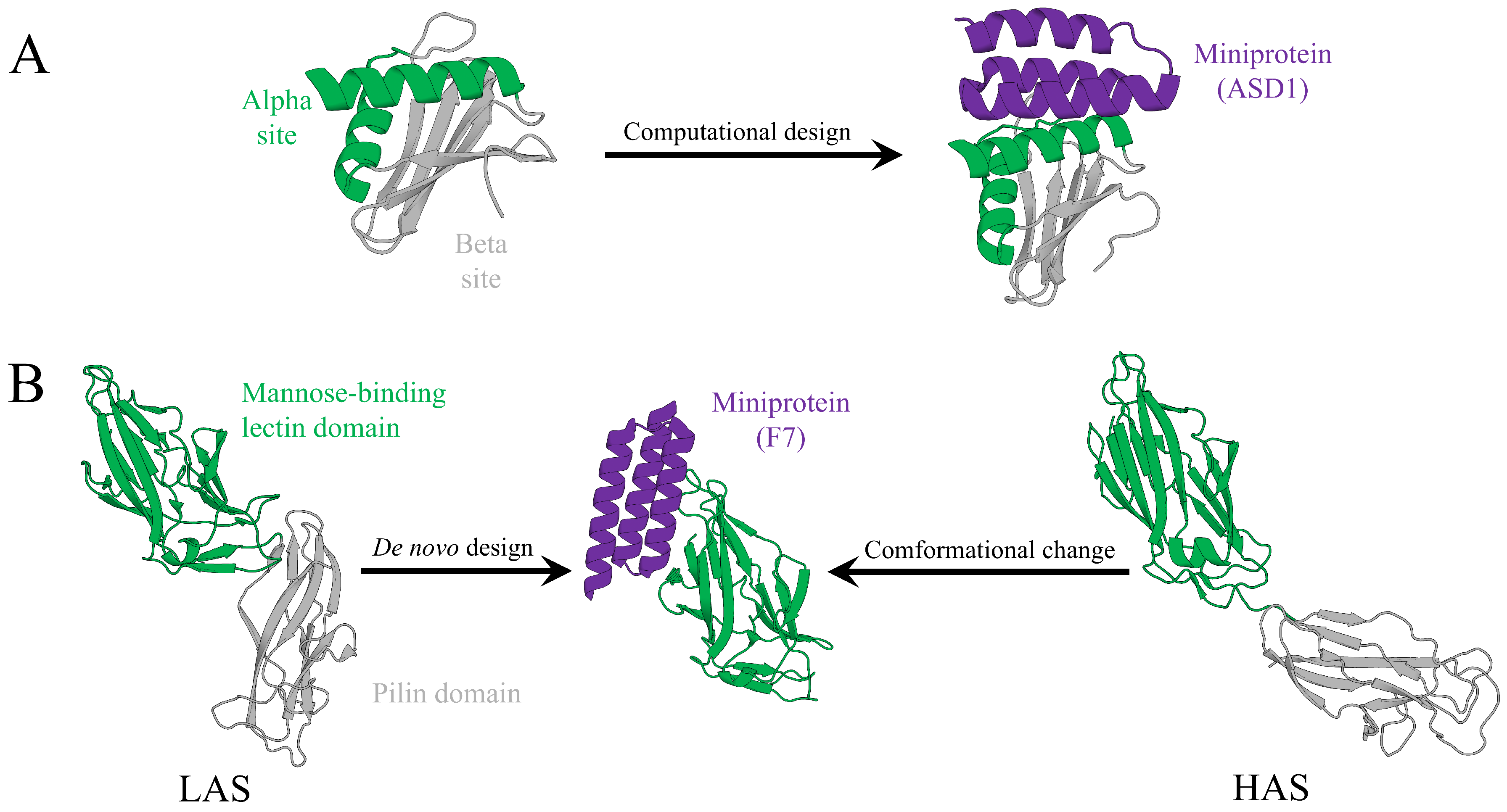
3.1. Case 1: High-Affinity Binders to the Flpp3 Virulence Factor
3.2. Case 2: Miniprotein Inhibitors of Bacterial Adhesins
Conclusions and Future Prospects
Funding
References
- Fenton, A.W. Allostery: an illustrated definition for the ‘second secret of life’. Trends in biochemical sciences 2008, 33, 420–425. [Google Scholar] [CrossRef] [PubMed]
- Astore, M.A.; Pradhan, A.S.; Thiede, E.H.; Hanson, S.M. Protein dynamics underlying allosteric regulation. Current Opinion in Structural Biology 2024, 84, 102768. [Google Scholar] [CrossRef]
- Kenakin, T.; Christopoulos, A. Signalling bias in new drug discovery: detection, quantification and therapeutic impact. Nature reviews Drug discovery 2013, 12, 205–216. [Google Scholar] [CrossRef]
- Mannes, M.; Martin, C.; Menet, C.; Ballet, S. Wandering beyond small molecules: peptides as allosteric protein modulators. Trends in Pharmacological Sciences 2022, 43, 406–423. [Google Scholar] [CrossRef]
- Olson, K.M.; Traynor, J.R.; Alt, A. Allosteric modulator leads hiding in plain site: Developing peptide and peptidomimetics as GPCR allosteric modulators. Frontiers in Chemistry 2021, 9, 671483. [Google Scholar] [CrossRef]
- Fournier, L.; Guarnera, E.; Kolmar, H.; Becker, S. Allosteric antibodies: a novel paradigm in drug discovery. Trends in Pharmacological Sciences 2025, 46, 311–323. [Google Scholar] [CrossRef]
- Kar, G.; Keskin, O.; Gursoy, A.; Nussinov, R. Allostery and population shift in drug discovery. Current opinion in pharmacology 2010, 10, 715–722. [Google Scholar] [PubMed]
- Yan, Z.; Wang, J. Funneled energy landscape unifies principles of protein binding and evolution. Proceedings of the National Academy of Sciences 2020, 117, 27218–27223. [Google Scholar] [CrossRef]
- Yan, Z.; Li, Y.; Cao, Y.; Tao, X.; Wang, J.; Jiang, Y. Binding Specificity and Local Frustration in Structure-based Drug Discovery. Current Medicinal Chemistry 2025. [Google Scholar] [CrossRef]
- Nussinov, R. Introduction to protein ensembles and allostery. Chemical reviews 2016, 116, 6263–6266. [Google Scholar] [CrossRef] [PubMed]
- Lewis, S.; Hempel, T.; Jiménez-Luna, J.; Gastegger, M.; Xie, Y.; Foong, A.Y.; Satorras, V.G.; Abdin, O.; Veeling, B.S.; Zaporozhets, I.; et al. Scalable emulation of protein equilibrium ensembles with generative deep learning. Science 2025, 389, eadv9817. [Google Scholar] [CrossRef] [PubMed]
- Shen, S.; Zhao, C.; Wu, C.; Sun, S.; Li, Z.; Yan, W.; Shao, Z. Allosteric modulation of G protein-coupled receptor signaling. Frontiers in Endocrinology 2023, 14, 1137604. [Google Scholar] [CrossRef] [PubMed]
- Chatzigoulas, A.; Cournia, Z. Rational design of allosteric modulators: Challenges and successes. Wiley Interdisciplinary Reviews: Computational Molecular Science 2021, 11, e1529. [Google Scholar] [CrossRef]
- Asada, N.; Krebs, C.F.; Panzer, U. Miniproteins may have a big impact: new therapeutics for autoimmune diseases and beyond. Signal Transduction and Targeted Therapy 2024, 9, 298. [Google Scholar] [CrossRef]
- Maeda, S.; Xu, J.; N. Kadji, F.M.; Clark, M.J.; Zhao, J.; Tsutsumi, N.; Aoki, J.; Sunahara, R.K.; Inoue, A.; Garcia, K.C.; et al. Structure and selectivity engineering of the M1 muscarinic receptor toxin complex. Science 2020, 369, 161–167. [Google Scholar] [CrossRef]
- Cao, L.; Goreshnik, I.; Coventry, B.; Case, J.B.; Miller, L.; Kozodoy, L.; Chen, R.E.; Carter, L.; Walls, A.C.; Park, Y.J.; et al. De novo design of picomolar SARS-CoV-2 miniprotein inhibitors. Science 2020, 370, 426–431. [Google Scholar] [CrossRef]
- Berger, S.; Seeger, F.; Yu, T.Y.; Aydin, M.; Yang, H.; Rosenblum, D.; Guenin-Macé, L.; Glassman, C.; Arguinchona, L.; Sniezek, C.; et al. Preclinical proof of principle for orally delivered Th17 antagonist miniproteins. Cell 2024, 187, 4305–4317. [Google Scholar] [CrossRef]
- Roy, A.; Shi, L.; Chang, A.; Dong, X.; Fernandez, A.; Kraft, J.C.; Li, J.; Le, V.Q.; Winegar, R.V.; Cherf, G.M.; et al. De novo design of highly selective miniprotein inhibitors of integrins αvβ6 and αvβ8. Nature Communications 2023, 14, 5660. [Google Scholar] [CrossRef]
- Nussinov, R.; Zhang, M.; Liu, Y.; Jang, H. AlphaFold, artificial intelligence (AI), and allostery. The Journal of Physical Chemistry B 2022, 126, 6372–6383. [Google Scholar] [CrossRef]
- Agajanian, S.; Alshahrani, M.; Bai, F.; Tao, P.; Verkhivker, G.M. Exploring and learning the universe of protein allostery using artificial intelligence augmented biophysical and computational approaches. Journal of chemical information and modeling 2023, 63, 1413–1428. [Google Scholar] [CrossRef]
- Xiao, S.; Verkhivker, G.M.; Tao, P. Machine learning and protein allostery. Trends in biochemical sciences 2023, 48, 375–390. [Google Scholar] [CrossRef]
- Huang, J.; Tang, G.; Liu, N.; Li, X.; Lu, S. Recent advances in computational strategies for allosteric site prediction: Machine learning, molecular dynamics, and network-based approaches. Drug Discovery Today 2025, 104466. [Google Scholar] [CrossRef]
- Koh, H.Y.; Zheng, Y.; Yang, M.; Arora, R.; Webb, G.I.; Pan, S.; Li, L.; Church, G.M. AI-driven protein design. Nature Reviews Bioengineering 2025, 1–23. [Google Scholar] [CrossRef]
- Bouchiba, Y.; Ruffini, M.; Schiex, T.; Barbe, S. Computational design of miniprotein binders. In Computational Peptide Science: Methods and Protocols; Springer, 2022; pp. 361–382. [Google Scholar]
- Ożga, K.; Berlicki, Ł. Design and engineering of miniproteins. ACS bio & med Chem Au 2022, 2, 316–327. [Google Scholar]
- Kortemme, T. De novo protein design—From new structures to programmable functions. Cell 2024, 187, 526–544. [Google Scholar] [CrossRef]
- Fox, D.R.; Taveneau, C.; Clement, J.; Grinter, R.; Knott, G.J. Code to complex: AI-driven de novo binder design. Structure 2025, 33, 1631–1642. [Google Scholar] [CrossRef] [PubMed]
- Zhu, R.; Wu, C.; Zha, J.; Lu, S.; Zhang, J. Decoding allosteric landscapes: computational methodologies for enzyme modulation and drug discovery. RSC Chemical Biology 2025, 6, 539–554. [Google Scholar] [CrossRef]
- Hollingsworth, S.A.; Kelly, B.; Valant, C.; Michaelis, J.A.; Mastromihalis, O.; Thompson, G.; Venkatakrishnan, A.; Hertig, S.; Scammells, P.J.; Sexton, P.M.; et al. Cryptic pocket formation underlies allosteric modulator selectivity at muscarinic GPCRs. Nature Communications 2019, 10, 3289. [Google Scholar] [CrossRef]
- Bemelmans, M.P.; Cournia, Z.; Damm-Ganamet, K.L.; Gervasio, F.L.; Pande, V. Computational advances in discovering cryptic pockets for drug discovery. Current Opinion in Structural Biology 2025, 90, 102975. [Google Scholar] [CrossRef] [PubMed]
- Wallerstein, J.; Han, X.; Levkovets, M.; Lesovoy, D.; Malmodin, D.; Mirabello, C.; Wallner, B.; Sun, R.; Sandalova, T.; Agback, P.; et al. Insights into mechanisms of MALT1 allostery from NMR and AlphaFold dynamic analyses. Communications biology 2024, 7, 868. [Google Scholar] [CrossRef] [PubMed]
- Duan, J.; He, X.H.; Li, S.J.; Xu, H.E. Cryo-electron microscopy for GPCR research and drug discovery in endocrinology and metabolism. Nature Reviews Endocrinology 2024, 20, 349–365. [Google Scholar] [CrossRef]
- Huang, W.; Lu, S.; Huang, Z.; Liu, X.; Mou, L.; Luo, Y.; Zhao, Y.; Liu, Y.; Chen, Z.; Hou, T.; et al. Allosite: a method for predicting allosteric sites. Bioinformatics 2013, 29, 2357–2359. [Google Scholar] [CrossRef]
- Greener, J.G.; Sternberg, M.J. AlloPred: prediction of allosteric pockets on proteins using normal mode perturbation analysis. BMC bioinformatics 2015, 16, 335. [Google Scholar] [CrossRef] [PubMed]
- Song, K.; Liu, X.; Huang, W.; Lu, S.; Shen, Q.; Zhang, L.; Zhang, J. Improved method for the identification and validation of allosteric sites. Journal of chemical information and modeling 2017, 57, 2358–2363. [Google Scholar] [CrossRef]
- Tian, H.; Jiang, X.; Tao, P. PASSer: Prediction of allosteric sites server. Machine learning: science and technology 2021, 2, 035015. [Google Scholar] [CrossRef]
- Xiao, S.; Tian, H.; Tao, P. PASSer2. 0: accurate prediction of protein allosteric sites through automated machine learning. Frontiers in molecular biosciences 2022, 9, 879251. [Google Scholar] [CrossRef]
- Tian, H.; Xiao, S.; Jiang, X.; Tao, P. PASSerRank: Prediction of allosteric sites with learning to rank. Journal of computational chemistry 2023, 44, 2223–2229. [Google Scholar] [CrossRef]
- Ugurlu, S.Y.; McDonald, D.; He, S. Mef-allosite: An accurate and robust multimodel ensemble feature selection for the allosteric site identification model. Journal of Cheminformatics 2024, 16, 116. [Google Scholar] [CrossRef] [PubMed]
- Nerin-Fonz, F.; Cournia, Z. Machine learning approaches in predicting allosteric sites. Current Opinion in Structural Biology 2024, 85, 102774. [Google Scholar] [CrossRef]
- Li, R.; He, X.; Wu, C.; Li, M.; Zhang, J. Advances in structure-based allosteric drug design. Current Opinion in Structural Biology 2025, 90, 102974. [Google Scholar] [CrossRef] [PubMed]
- Hu, G.; Doruker, P.; Li, H.; Demet Akten, E. Understanding protein dynamics, binding and allostery for drug design. 2021. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Baek, M.; DiMaio, F.; Anishchenko, I.; Dauparas, J.; Ovchinnikov, S.; Lee, G.R.; Wang, J.; Cong, Q.; Kinch, L.N.; Schaeffer, R.D.; et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 2021, 373, 871–876. [Google Scholar] [CrossRef] [PubMed]
- Du, Z.; Su, H.; Wang, W.; Ye, L.; Wei, H.; Peng, Z.; Anishchenko, I.; Baker, D.; Yang, J. The trRosetta server for fast and accurate protein structure prediction. Nature protocols 2021, 16, 5634–5651. [Google Scholar] [CrossRef]
- Evans, R.; O’Neill, M.; Pritzel, A.; Antropova, N.; Senior, A.; Green, T.; Žídek, A.; Bates, R.; Blackwell, S.; Yim, J.; et al. Protein complex prediction with AlphaFold-Multimer. biorxiv 2021, 2021–10. [Google Scholar]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Niazi, S.K.; Yang, J. A comprehensive application of FiveFold for conformation ensemble-based protein structure prediction. Scientific Reports 2025, 15, 33498. [Google Scholar] [CrossRef]
- Cui, X.; Ge, L.; Chen, X.; Lv, Z.; Wang, S.; Zhou, X.; Zhang, G. Beyond static structures: protein dynamic conformations modeling in the post-AlphaFold era. Briefings in bioinformatics 2025, 26, bbaf340. [Google Scholar] [CrossRef]
- Winnifrith, A.; Outeiral, C.; Hie, B. Generative artificial intelligence for de novo protein design. arXiv 2023, arXiv:2310.09685. [Google Scholar] [CrossRef]
- Das, U. Generative AI for drug discovery and protein design: the next frontier in AI-driven molecular science. Medicine in Drug Discovery 2025, 100213. [Google Scholar] [CrossRef]
- Watson, J.L.; Juergens, D.; Bennett, N.R.; Trippe, B.L.; Yim, J.; Eisenach, H.E.; Ahern, W.; Borst, A.J.; Ragotte, R.J.; Milles, L.F.; et al. De novo design of protein structure and function with RFdiffusion. Nature 2023, 620, 1089–1100. [Google Scholar] [CrossRef]
- Ahern, W.; Yim, J.; Tischer, D.; Salike, S.; Woodbury, S.M.; Kim, D.; Kalvet, I.; Kipnis, Y.; Coventry, B.; Altae-Tran, H.R.; et al. Atom-level enzyme active site scaffolding using RFdiffusion2. Nature Methods 2025, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Butcher, J.; Krishna, R.; Mitra, R.; Brent, R.I.; Li, Y.; Corley, N.; Kim, P.T.; Funk, J.; Mathis, S.; Salike, S.; et al. De novo design of all-atom biomolecular interactions with rfdiffusion3. bioRxiv 2025. [Google Scholar] [CrossRef]
- Bose, A.J.; Akhound-Sadegh, T.; Huguet, G.; Fatras, K.; Rector-Brooks, J.; Liu, C.H.; Nica, A.C.; Korablyov, M.; Bronstein, M.; Tong, A. Se (3)-stochastic flow matching for protein backbone generation. arXiv 2023, arXiv:2310.02391. [Google Scholar]
- Ingraham, J.B.; Baranov, M.; Costello, Z.; Barber, K.W.; Wang, W.; Ismail, A.; Frappier, V.; Lord, D.M.; Ng-Thow-Hing, C.; Van Vlack, E.R.; et al. Illuminating protein space with a programmable generative model. Nature 2023, 623, 1070–1078. [Google Scholar] [CrossRef]
- Dauparas, J.; Anishchenko, I.; Bennett, N.; Bai, H.; Ragotte, R.J.; Milles, L.F.; Wicky, B.I.; Courbet, A.; de Haas, R.J.; Bethel, N.; et al. Robust deep learning–based protein sequence design using ProteinMPNN. Science 2022, 378, 49–56. [Google Scholar] [CrossRef]
- Gao, Z.; Tan, C.; Chacón, P.; Li, S.Z. Pifold: Toward effective and efficient protein inverse folding. arXiv 2022, arXiv:2209.12643. [Google Scholar]
- Hsu, C.; Verkuil, R.; Liu, J.; Lin, Z.; Hie, B.; Sercu, T.; Lerer, A.; Rives, A. Learning inverse folding from millions of predicted structures. In Proceedings of the International conference on machine learning. PMLR, 2022; pp. 8946–8970. [Google Scholar]
- Zambaldi, V.; La, D.; Chu, A.E.; Patani, H.; Danson, A.E.; Kwan, T.O.; Frerix, T.; Schneider, R.G.; Saxton, D.; Thillaisundaram, A.; et al. De novo design of high-affinity protein binders with AlphaProteo. arXiv 2024. arXiv:2409.08022. [CrossRef]
- Jendrusch, M.A.; Yang, A.L.; Cacace, E.; Bobonis, J.; Voogdt, C.G.; Kaspar, S.; Schweimer, K.; Perez-Borrajero, C.; Lapouge, K.; Scheurich, J.; et al. AlphaDesign: A de novo protein design framework based on AlphaFold. Molecular Systems Biology 2025, 1–24. [Google Scholar] [CrossRef]
- Zhang, O.; Zhang, X.; Lin, H.; Tan, C.; Wang, Q.; Mo, Y.; Feng, Q.; Du, G.; Yu, Y.; Jin, Z.; et al. ODesign: A World Model for Biomolecular Interaction Design. arXiv 2025, arXiv:2510.22304. [Google Scholar] [CrossRef]
- Pacesa, M.; Nickel, L.; Schellhaas, C.; Schmidt, J.; Pyatova, E.; Kissling, L.; Barendse, P.; Choudhury, J.; Kapoor, S.; Alcaraz-Serna, A.; et al. BindCraft: one-shot design of functional protein binders. bioRxiv 2024, 2024–09. [Google Scholar] [CrossRef]
- Stark, H.; Faltings, F.; Choi, M.; Xie, Y.; Hur, E.; O’Donnell, T.J.; Bushuiev, A.; Uçar, T.; Passaro, S.; Mao, W.; et al. BoltzGen: Toward Universal Binder Design. bioRxiv 2025, 2025–11. [Google Scholar] [CrossRef]
- Ren, M.; Sun, J.; Guan, J.; Liu, C.; Gong, C.; Wang, Y.; Wang, L.; Cai, Q.; Chen, X.; Xiao, W.; et al. Pxdesign: Fast, modular, and accurate de novo design of protein binders. bioRxiv 2025, 2025–08. [Google Scholar] [CrossRef]
- Song, Z.; Li, T.; Li, L.; Min, M.R. PPDiff: Diffusing in Hybrid Sequence-Structure Space for Protein-Protein Complex Design. arXiv arXiv:2506.11420.
- Bennett, N.R.; Coventry, B.; Goreshnik, I.; Huang, B.; Allen, A.; Vafeados, D.; Peng, Y.P.; Dauparas, J.; Baek, M.; Stewart, L.; et al. Improving de novo protein binder design with deep learning. Nature Communications 2023, 14, 2625. [Google Scholar] [CrossRef] [PubMed]
- Jussupow, A.; Kaila, V.R. Effective molecular dynamics from neural network-based structure prediction models. Journal of Chemical Theory and Computation 2023, 19, 1965–1975. [Google Scholar] [CrossRef]
- Das, R.; Baker, D. Macromolecular modeling with rosetta. Annu. Rev. Biochem. 2008, 77, 363–382. [Google Scholar] [CrossRef] [PubMed]
- Sgrignani, J.; Buscarini, S.; Locatelli, P.; Guerra, C.; Furlan, A.; Chen, Y.; Zoppi, G.; Cavalli, A. AI assisted design of ligands for Lipocalin-2. bioRxiv 2025, 2025–05. [CrossRef]
- Gokce-Alpkilic, G.; Huang, B.; Liu, A.; Kreuk, L.S.; Wang, Y.; Adebomi, V.; Bueso, Y.F.; Bera, A.K.; Kang, A.; Gerben, S.R.; et al. De Novo Design of High-Affinity Miniprotein Binders Targeting Francisella Tularensis Virulence Factor. Angewandte Chemie International Edition 2025, e202516058. [Google Scholar]
- Chazin-Gray, A.M.; Thompson, T.R.; Lopatto, E.D.; Magala, P.; Erickson, P.W.; Hunt, A.C.; Manchenko, A.; Aprikian, P.; Tchesnokova, V.; Basova, I.; et al. De Novo Design of Miniprotein Inhibitors of Bacterial Adhesins. bioRxiv 2025. [Google Scholar] [CrossRef]
- Heling, L.W.; van der Veen, J.; Rofe, A.; West, E.; Jiménez-Panizo, A.; Alegre-Martí, A.; Sheikhhassani, V.; Ng, J.; Schmidt, T.; Estébanez-Perpiñá, E.; et al. Deciphering the allosteric control of androgen receptor DNA binding by its disordered N-terminal domain. Molecular and Cellular Endocrinology 2025, 112634. [Google Scholar] [CrossRef]
- Govindaraj, R.G.; Thangapandian, S.; Schauperl, M.; Denny, R.A.; Diller, D.J. Recent applications of computational methods to allosteric drug discovery. Frontiers in Molecular Biosciences 2023, 9, 1070328. [Google Scholar] [CrossRef]
- Yang, D.; Zhou, Q.; Labroska, V.; Qin, S.; Darbalaei, S.; Wu, Y.; Yuliantie, E.; Xie, L.; Tao, H.; Cheng, J.; et al. G protein-coupled receptors: structure-and function-based drug discovery. Signal transduction and targeted therapy 2021, 6, 7. [Google Scholar] [CrossRef] [PubMed]
- Moore, M.N.; Person, K.L.; Robleto, V.L.; Alwin, A.R.; Krusemark, C.L.; Foster, N.; Ray, C.; Inoue, A.; Jackson, M.R.; Sheedlo, M.J.; et al. Designing allosteric modulators to change GPCR G protein subtype selectivity. Nature 2025, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Meijer, F.A.; Leijten-van de Gevel, I.A.; de Vries, R.M.; Brunsveld, L. Allosteric small molecule modulators of nuclear receptors. Molecular and cellular endocrinology 2019, 485, 20–34. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Hawkins, B.A.; Du, J.J.; Groundwater, P.W.; Hibbs, D.E.; Lai, F. A guide to in silico drug design. Pharmaceutics 2023, 15, 49. [Google Scholar] [CrossRef]
| Tool a | Machine-learning strategy and key features | Year | Ref |
|---|---|---|---|
| Allosite | Support vector machine classifier trained on static structural descriptors to discriminate allosteric from non-allosteric pockets | 2013 | [33] |
| AlloPred | Perturbation-guided machine-learning scoring of candidate pockets combined with normal mode analysis | 2016 | [34] |
| AllositePro | Structure-based machine-learning framework integrating multiple physicochemical and geometric features for improved robustness | 2017 | [35] |
| PASSer | Ensemble machine-learning approach trained on curated allosteric datasets for large-scale pocket identification | 2021 | [36] |
| PASSer 2.0 | AutoML-driven framework enabling automated feature selection, model optimization, and improved generalization | 2022 | [37] |
| PASSerRank | Learning-to-rank strategy for prioritizing predicted allosteric pockets rather than binary classification | 2023 | [38] |
| MEF-AlloSite | Multi-model ensemble learning with optimized feature selection for accurate identification of allosteric sites and pockets | 2024 | [39] |
| Tool a | Method and Key Features | Year | Ref |
|---|---|---|---|
| trRosetta | Deep-learning model predicting inter-residue distances and orientations from MSA-derived features; early high-throughput deep predictor for fold inference. | 2020 | [45] |
| RoseTTAFold | Three-track neural network integrating sequence, pairwise distances, and 3D coordinates; uses MSAs for accurate monomer and multimer predictions. | 2021 | [44] |
| AlphaFold2 | Deep-learning model using MSA and Evoformer architecture; delivers high-accuracy monomer and complex structure predictions with confidence metrics. | 2021 | [43] |
| AlphaFold-Multimer | Extension of AlphaFold2 for protein complex modeling; incorporates paired MSAs to capture inter-chain co-evolutionary signals. | 2022 | [46] |
| AlphaFold3 | Updated deep-learning model with diffusion refinement and broader capability for complexes including proteins and other biomolecules, while still using MSA information. | 2024 | [47] |
| Category | Tool | Core capability | Year | Ref |
|---|---|---|---|---|
| Backbone generation | RFdiffusion | Diffusion-based backbone generation conditioned on target interfaces for stable miniprotein scaffolds | 2023 | [52] |
| Sequence generation | ProteinMPNN | Inverse folding–based sequence design for fixed backbone miniproteins | 2022 | [57] |
| ESM-IF1 | Protein language model–based inverse folding for sequence design on fixed miniprotein backbones | 2022 | [59] | |
| PiFold | Graph neural network–based inverse folding enabling efficient miniprotein sequence design | 2022 | [58] | |
| Integrated design of backbone and sequence | AlphaProteo | AlphaFold-assisted binder design emphasizing functional interaction motifs | 2024 | [60] |
| BindCraft | Automated one-shot de novo miniprotein binder design with high experimental hit rates | 2024 | [63] | |
| O-design | Objective-driven interface refinement via energy-based and deep learning–assisted sequence optimization | 2025 | [62] | |
| Integrated design of backbone and sequence | AlphaDesign | AlphaFold-guided hallucination with diffusion-based sequence optimization for multistate binder design | 2025 | [61] |
| BoltzGen | All-atom generative model unifying structure and sequence for universal binder design, including miniproteins | 2025 | [64] | |
| PXDesign | End-to-end de novo binder design pipeline (generation plus confidence filtering) with high experimental success rates | 2025 | [65] | |
| PPDiff | Joint sequence–structure diffusion framework for direct generation of protein–protein complexes and miniprotein binders | 2025 | [66] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




