Submitted:
22 December 2025
Posted:
22 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Design
2.2. Sample
2.3. Recruitment Strategies
2.4. Data Collection
2.5. Measures
2.5.1. Sociodemographic Data at Baseline
2.5.2. Clinical data at baseline
2.5.3. Diet Quality (at Baseline and 6-Month Post-Chemotherapy Initiation)
2.5.4. Fecal Microbiome Profile (at Baseline and 6-Month Post-Chemotherapy Initiation)
2.6. Statistical Analysis
3. Results
3.1. Characteristics of Participants
3.2. Nutritional Profiles Pre- and Post-Chemotherapy
3.3. Microbiome Profiles Pre- and Post-Chemotherapy
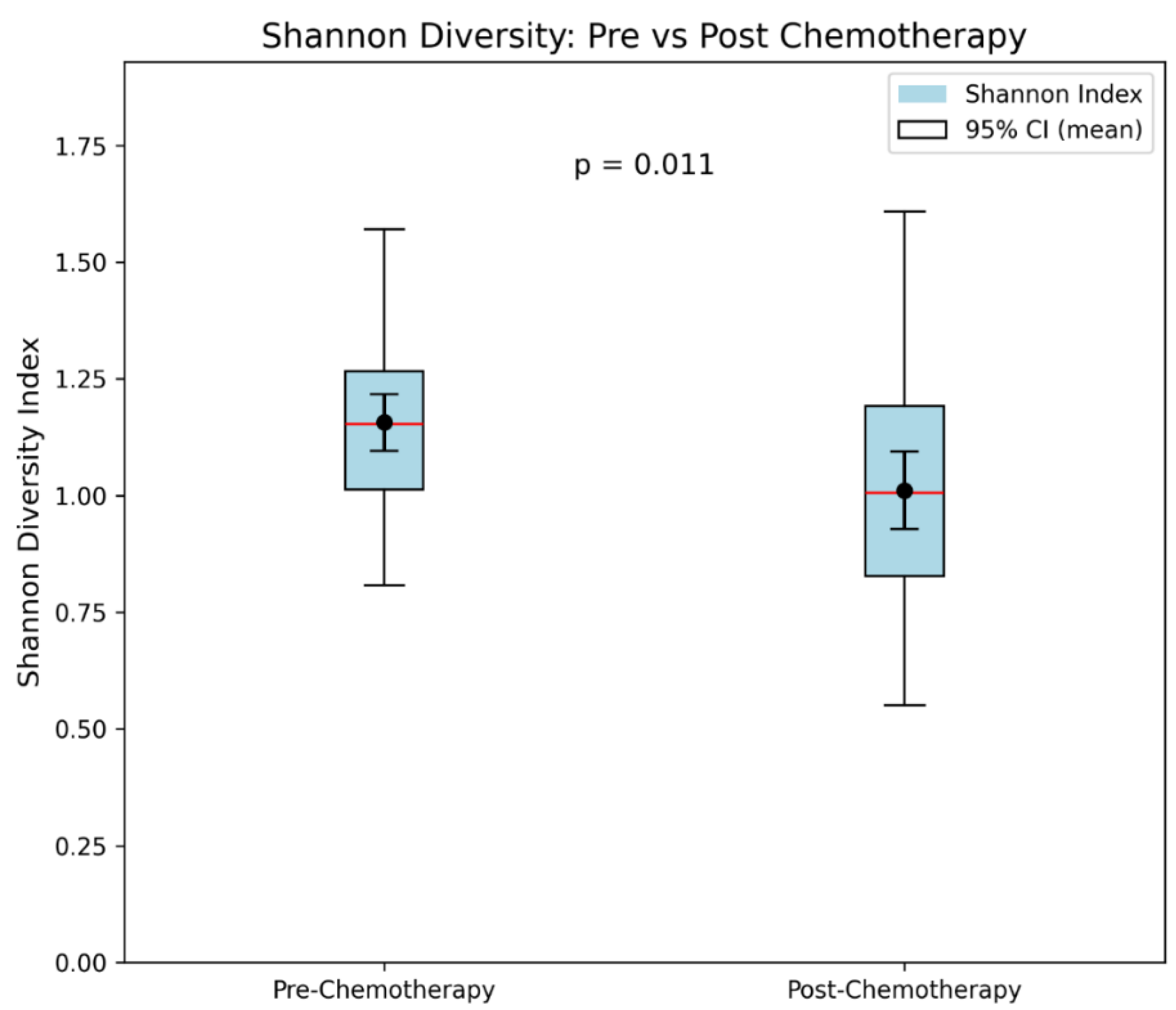
3.3.1. Shannon Diversity
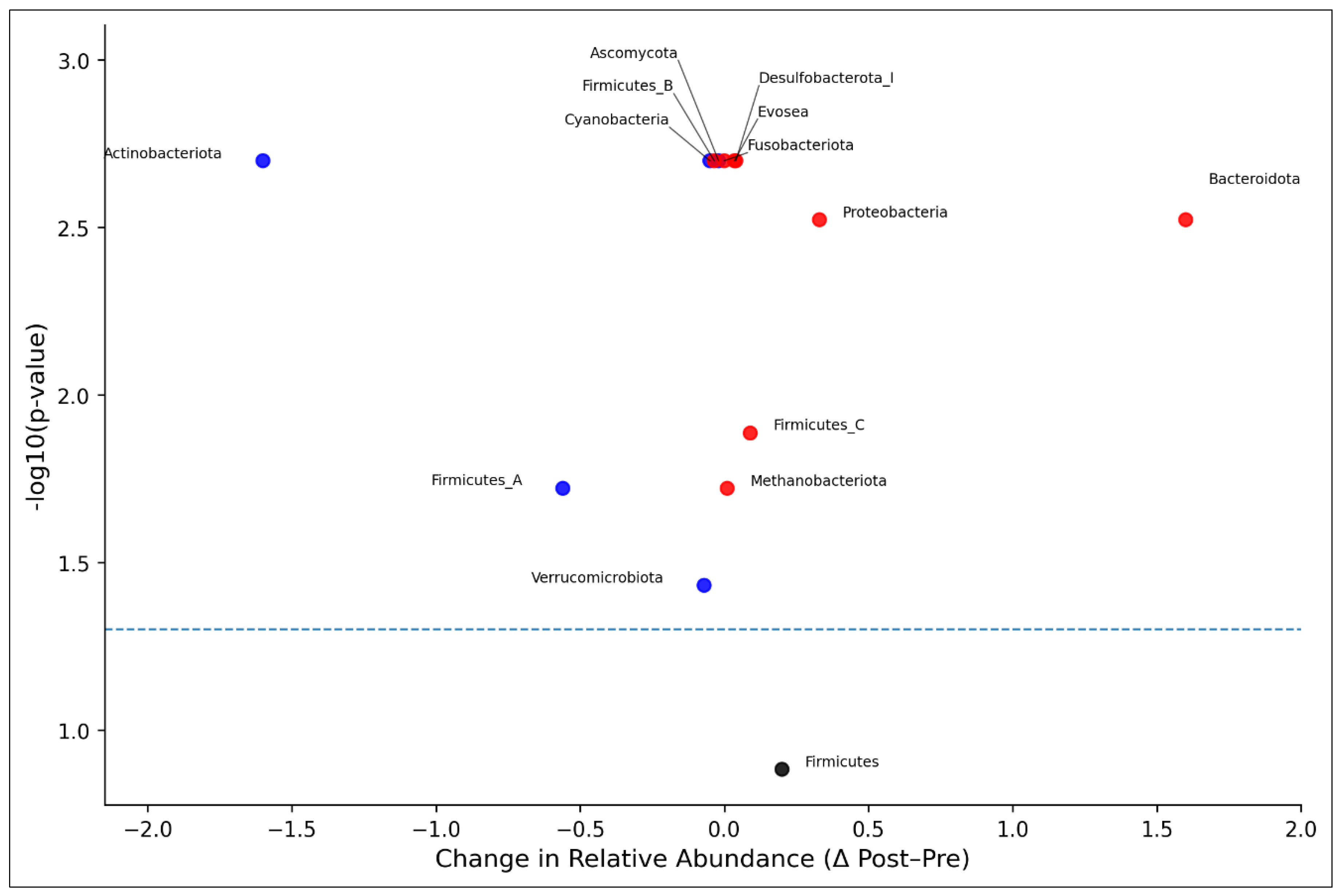
3.3.2. Relative Abundance of Major Microbiome Phylum Pre- and Post-Chemotherapy
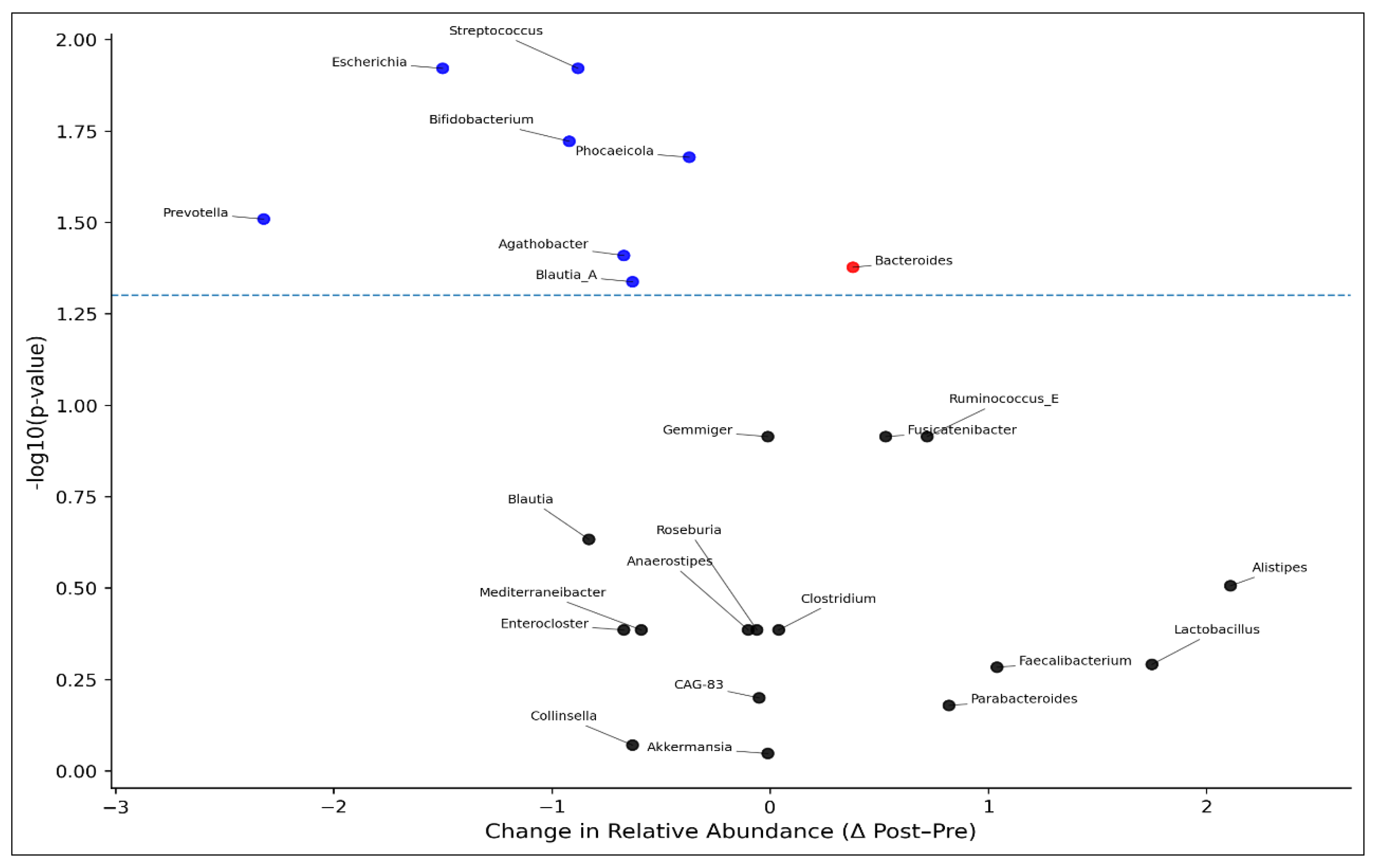
3.3.3. Relative Abundance of Major Microbiome Genus Pre- and Post-Chemotherapy
3.4. Associations Between Diet Quality Measured with HEI and Microbiome Profiles
3.4.1. Diet quality and Shannon Diversity (Table 6)
3.4.2. Diet Quality and Microbial Phyla (Table 6)
3.4.3. Diet Quality and Microbiome Genus (Table 7)
4. Discussion
4.1. Diet Changes
4.2. Microbiome Diversity (Shannon Index)
4.3. Microbiome Composition
4.4. Associations Between Diet Quality (Measured as HEI) and Microbiome Profiles
4.5. Clinical Implications
4.6. Strengths and Limitations
5. Conclusions
Author Contributions
Funding
Acknowledgments
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- Siegel, R.L.; Kratzer, T.B.; Giaquinto, A.N.; Sung, H.; Jemal, A. Cancer statistics, 2025. CA Cancer J. Clin. 2025, 75, 10–45. [Google Scholar] [CrossRef] [PubMed]
- Morgan, E.; Arnold, M.; Gini, A.; et al. Global burden of colorectal cancer in 2020 and 2040: Incidence and mortality estimates from GLOBOCAN. Gut 2023, 72, 338–344. [Google Scholar] [CrossRef] [PubMed]
- Sung, H.; Siegel, R.L.; Laversanne, M.; et al. Colorectal cancer incidence trends in younger versus older adults. Lancet Oncol. 2025, 26, 51–63. [Google Scholar] [CrossRef] [PubMed]
- Spaander, M.C.W.; Zauber, A.G.; Syngal, S.; et al. Young-onset colorectal cancer. Nat. Rev. Dis. Primers 2023, 9, 21. [Google Scholar] [CrossRef]
- Pu, W.; et al. A novel approach in colon cancer treatment using endoscopic rendezvous technique. Photodiagn. Photodyn. Ther. 2025, 104761. [Google Scholar] [CrossRef]
- Balderas-Peña, L.M.; et al. Body composition and nutritional status in colorectal cancer. Nutrients 2020, 12, 2110. [Google Scholar] [CrossRef]
- Gebremedhin, T.K.; et al. Malnutrition among adult cancer patients receiving chemotherapy. Heliyon 2021, 7, e07362. [Google Scholar] [CrossRef]
- Rock, C.L.; Thomson, C.A.; Sullivan, K.R.; Howe, C.L.; Kushi, L.H.; Caan, B.J.; Neuhouser, M.L.; Bandera, E.V.; Wang, Y.; Robien, K.; et al. ACS nutrition and physical activity guideline for cancer survivors. CA Cancer J. Clin. 2022, 72, 230–262. [Google Scholar] [CrossRef]
- Xue, M.; et al. Fasting-mimicking diet in breast cancer patients receiving chemotherapy. Breast Cancer Res. Treat. 2025. [CrossRef]
- Crowder, S.L.; et al. Diet quality indices and cognition during chemotherapy. Support. Care Cancer 2022, 31, 75. [Google Scholar] [CrossRef]
- Kleckner, A.S.; et al. Mediterranean diet intervention and cancer-related fatigue. Cancers 2022, 14, 4202. [Google Scholar] [CrossRef]
- Lazar, D.C.; et al. Gastric cancer and microbiota. Life 2025, 15, 999. [Google Scholar] [CrossRef] [PubMed]
- Yan, R.; et al. Gut microbiome dysbiosis in colorectal cancer. J. Med. Microbiol. 2025, 74, 002042. [Google Scholar] [CrossRef] [PubMed]
- Asseri, A.H.; et al. Gut dysbiosis–cancer axis. Front. Pharmacol. 2023, 14, 1208044. [Google Scholar] [CrossRef]
- Artym, J.; Zimecki, M. Colostrum proteins in therapy-induced injuries. Biomedicines 2023, 11, 114. [Google Scholar] [CrossRef]
- Montassier, E.; et al. Chemotherapy-driven dysbiosis. Aliment. Pharmacol. Ther. 2015, 42, 515–528. [Google Scholar] [CrossRef]
- Goubet, A.G.; et al. Intestinal microbiota and therapeutic responses. C. R. Biol. 2018, 341, 284–289. [Google Scholar] [CrossRef]
- Zmora, N.; Suez, J.; Elinav, E. Diet and the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 35–56. [Google Scholar] [CrossRef]
- Carding, S.; Verbeke, K.; Vipond, D.T.; Corfe, B.M.; Owen, L.J. Dysbiosis of the gut microbiota. Microb. Ecol. Health Dis. 2015, 26, 26191. [Google Scholar] [CrossRef]
- Zhang, P. Influence of foods on the gut microbiome. Int. J. Mol. Sci. 2022, 23, 9588. [Google Scholar] [CrossRef]
- Aarnoutse, R.; et al. Changes in intestinal microbiota in postmenopausal oestrogen receptor-positive breast cancer patients treated with (neo)adjuvant chemotherapy. NPJ Breast Cancer 2022, 8, 89. [Google Scholar] [CrossRef]
- Maddern, A.S.; et al. The association between the gut microbiome and development and progression of cancer treatment adverse effects. Cancers 2023, 15, 4361. [Google Scholar] [CrossRef] [PubMed]
- Chan, W.L.; et al. Prediction models for severe treatment-related toxicities in older adults with cancer: A systematic review. Age Ageing 2025, 54, afaf070. [Google Scholar] [CrossRef] [PubMed]
- Carson, T.L.; et al. Association between the gut microbiota and colorectal cancer. Gut Pathog. 2024, 16, 13. [Google Scholar] [CrossRef] [PubMed]
- He, T.; Cheng, X.; Xing, C. Gut microbial diversity of colon cancer patients. Bioengineered 2021, 12, 7046–7060. [Google Scholar] [CrossRef]
- Colombo, F.; Illescas, O.; Noci, S.; et al. Gut microbiota composition in colorectal cancer patients. Sci. Rep. 2022, 12, 11424. [Google Scholar] [CrossRef]
- Hosein, K.; et al. Short-form food frequency questionnaire vs. 3-day food intake record. Br. J. Nutr. 2025, 1–17. [Google Scholar] [CrossRef]
- Jubayer, A.; et al. Evaluation of Bangladesh Healthy Eating Index (BD-HEI). BMC Nutr. 2025, 11, 159. [Google Scholar] [CrossRef]
- Nutritional Pro. Version 2025; Healthy Living Software Inc.: Chicago, IL, USA, 2024. DOI not available.
- Baughman, C.; Norman, K.; Mukamal, K. Adherence to ACS nutrition and physical activity guidelines. JAMA Oncol. 2024, 10, 789–792. [Google Scholar] [CrossRef]
- Galloway-Peña, J.R.; et al. Gut microbiome signatures predict infectious risk in AML. Clin. Infect. Dis. 2020, 71, 63–71. [Google Scholar] [CrossRef]
- Rashidi, A.; et al. Gut microbiota changes and neutropenic fever. Leukemia 2020, 34, 312–316. [Google Scholar] [CrossRef]
- Al-Rashidi, H.E. Gut microbiota and immunity. Saudi J. Biol. Sci. 2022, 29, 1628–1643. [Google Scholar] [CrossRef] [PubMed]
- Anhê, F.F.; et al. Polyphenol-rich cranberry extract and Akkermansia. Gut 2015, 64, 872–883. [Google Scholar] [CrossRef] [PubMed]
- Hills, R.D., Jr.; et al. Gut microbiome implications for diet and disease. Nutrients 2019, 11, 1613. [Google Scholar] [CrossRef] [PubMed]
- Montassier, E.; et al. Chemotherapy-induced fecal microbiota shifts. Microb. Ecol. 2014, 67, 690–699. [Google Scholar] [CrossRef]
- Kuugbee, E.D.; et al. Probiotic cocktail enhances gut barrier and reduces cancer. Dig. Dis. Sci. 2016, 61, 2908–2920. [Google Scholar] [CrossRef]
- Kober, M.M.; Bowe, W.P. Probiotics and immune regulation. Int. J. Womens Dermatol. 2015, 1, 85–89. [Google Scholar] [CrossRef]
- Rock, C.L.; et al. ACS guideline for diet and physical activity for cancer prevention. CA Cancer J. Clin. 2020, 70, 245–271. [Google Scholar] [CrossRef]
- Guinter, M.A.; et al. Diet quality and mortality after colorectal cancer. J. Clin. Oncol. 2018, 36, JCO1800714. [Google Scholar] [CrossRef]
- Han, C.J.; Reding, K.W.; Kalady, M.F.; Yung, R.; Greenlee, H.; Paskett, E.D. GI symptoms in CRC survivors. PLoS ONE 2023, 18, e0286058. [Google Scholar] [CrossRef]
- Vandebroek, A.J.V.; Schrijvers, D. Nutritional issues in anticancer treatment. Ann. Oncol. 2008, 19, v52–v55. [Google Scholar] [CrossRef]
- Ravasco, P. Nutrition in cancer patients. J. Clin. Med. 2019, 8, 1211. [Google Scholar] [CrossRef] [PubMed]
- Ohnishi, S.; Takeda, H. Herbal medicines for chemotherapy side effects. Front. Pharmacol. 2015, 6, 14. [Google Scholar] [CrossRef] [PubMed]
- Kurt, B.; Oksuzoglu, B.O.C. Taste alterations during cisplatin therapy. Cancer Control 2025, 32, 10732748251363323. [Google Scholar] [CrossRef] [PubMed]
- Cok, O.Y.; et al. Paracetamol and postoperative nausea. Eur. J. Anaesthesiol. 2011, 28, 836–841. [Google Scholar] [CrossRef]
- Thomas, N.S.; et al. Depression, anxiety, fibromyalgia, and ME/CFS. Psychol. Med. 2025, 55, e232. [Google Scholar] [CrossRef]
- Bergerot, C.D.; et al. Distress and emotional support needs in stage IV cancer. Psychooncology 2025, 34, e70232. [Google Scholar] [CrossRef]
- Hasan, N.; et al. Dietary sugars and cancer risk. Cancer Treat. Res. Commun. 2025, 43, 100876. [Google Scholar] [CrossRef]
- Pellegrini, C.A.; et al. Healthy eating habits during COVID-19. Health Psychol. Behav. Med. 2023, 11, 2182307. [Google Scholar] [CrossRef]
- White, E. Bad eating habits of Ras-driven cancers. Genes Dev. 2013, 27, 2065–2071. [Google Scholar] [CrossRef]
- Roca-Saavedra, P.; et al. Food additives and gut microbiota. J. Physiol. Biochem. 2018, 74, 69–83. [Google Scholar] [CrossRef]
- Jiang, Y.; Li, Y. Nutrition intervention and microbiome modulation. Nutrients 2024, 16, 2621. [Google Scholar] [CrossRef] [PubMed]
- Salberg, S.; et al. Sex differences in gut microbiome and nociception. Dev. Neurobiol. 2023, 83, 219–233. [Google Scholar] [CrossRef] [PubMed]
- Mitra, A.; et al. Microbial diversity and toxicity during chemoradiation. Int. J. Radiat. Oncol. Biol. Phys. 2020, 107, 163–171. [Google Scholar] [CrossRef] [PubMed]
- Bohm, D.; et al. Gut microbiota and chemotherapy response. NPJ Precis. Oncol. 2025, 9, 265. [Google Scholar] [CrossRef]
- Yu, Z.K.; et al. Bacterial microbiome in cancer treatment. BMC Cancer 2021, 21, 934. [Google Scholar] [CrossRef]
- Otto-Dobos, L.D.; et al. Alpha diversity predicts GI symptoms. NPJ Breast Cancer 2024, 10, 99. [Google Scholar] [CrossRef]
- Rajagopala, S.V.; et al. Persistent dysbiosis in childhood ALL. Microb. Ecol. 2020, 79, 1034–1043. [Google Scholar] [CrossRef]
- Bai, J.; Behera, M.; Bruner, D.W. Gut microbiome and symptoms in pediatric cancer. Support. Care Cancer 2018, 26, 427–439. [Google Scholar]
- Di Vincenzo, F.; et al. Gut microbiota and systemic inflammation. Intern. Emerg. Med. 2024, 19, 275–293. [Google Scholar] [CrossRef]
- Ribatti, D. Comment on gut microbiota and inflammation. Intern. Emerg. Med. 2024, 19, 1515–1516. [Google Scholar] [CrossRef]
- Oh, B.; et al. Gut microbiome in chemotherapy. Front. Oncol. 2021, 11, 706331. [Google Scholar] [CrossRef] [PubMed]
- Prisciandaro, L.D.; et al. Probiotics for intestinal mucositis. Crit. Rev. Food Sci. Nutr. 2011, 51, 239–247. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.; et al. Postoperative probiotics in CRC. Nutrients 2023, 15, 356. [Google Scholar] [CrossRef]
- Kennedy, M.S.; Chang, E.B. Microbiome composition and locations. Prog. Mol. Biol. Transl. Sci. 2020, 176, 1–42. [Google Scholar] [CrossRef]
- Manor, O.; et al. Health and disease markers and gut microbiome. Nat. Commun. 2020, 11, 5206. [Google Scholar] [CrossRef]
- Ruan, W.; et al. Healthy human gastrointestinal microbiome. Dig. Dis. Sci. 2020, 65, 695–705. [Google Scholar] [CrossRef]
- Oliphant, K.; et al. Early-life microbiome and neurodevelopment. Gut Microbes 2021, 13, 1997560. [Google Scholar] [CrossRef]
- Portincasa, P.; et al. Gut microbiota and short-chain fatty acids. Int. J. Mol. Sci. 2022, 23, 1105. [Google Scholar] [CrossRef]
- Otto-Dobos, L.D.; et al. Chemotherapy-induced microbiome disruption and cognition. Brain Behav. Immun. 2024, 120, 208–220. [Google Scholar] [CrossRef]
- Song, M.; et al. Fiber intake and CRC survival. JAMA Oncol. 2018, 4, 71–79. [Google Scholar] [CrossRef]
- Wu, G.D.; et al. Dietary patterns and gut enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; et al. Gut microbiota and SCFAs in osteoporosis. Medicine (Baltimore) 2024, 103, e40554. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; et al. Short-chain fatty acids as antiviral mediators. Front. Immunol. 2025, 16, 1614879. [Google Scholar] [CrossRef] [PubMed]
- David, L.A.; et al. Diet rapidly alters the gut microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef]
- Huang, F.; et al. Probiotics and chemotherapy-related GI complications. Nutrients 2023, 15, 356. [Google Scholar] [CrossRef]
- Shastry, R.P.; Rekha, P.D. Bacterial cross-talk with gut microbiome. Folia Microbiol. 2021, 66, 15–24. [Google Scholar] [CrossRef]
- Del Chierico, F.; et al. Mediterranean diet and gut microbiota. Int. J. Mol. Sci. 2014, 15, 11678–11699. [Google Scholar] [CrossRef]
- Bielik, V.; Kolisek, M. Minerals and gut microbiome. Int. J. Mol. Sci. 2021, 22, 6803. [Google Scholar] [CrossRef]
- Frame, L.A.; Costa, E.; Jackson, S.A. Nutrition and gut microbiome: Review of reviews. Nutr. Rev. 2020, 78, 798–812. [Google Scholar] [CrossRef]
- Velly, H.; Britton, R.A.; Preidis, G.A. Diet–microbiome–host cross-talk. Gut Microbes 2017, 8, 98–112. [Google Scholar] [CrossRef]
- Wiertsema, S.P.; et al. Gut microbiome and immune system interplay. Nutrients 2021, 13, 886. [Google Scholar] [CrossRef]
- Ross, F.C.; et al. Diet–gut microbiome interactions. Nat. Rev. Microbiol. 2024, 22, 671–686. [Google Scholar] [CrossRef]
- Zhang, P. Influence of foods and nutrition on gut microbiome. Int. J. Mol. Sci. 2022, 23, 9588. [Google Scholar] [CrossRef]
| Characteristic | n (%) or Mean ± Standard Deviation (SD) [Range] |
| Demographic Characteristics | |
| Age (years) | 69.4 ± 6.7 [59–86] |
| Sex | |
| Male | 26 (54.2%) |
| Female | 22 (45.8%) |
| Marital Status | |
| Married | 28 (58.3%) |
| Single | 20 (41.7%) |
| Body Mass Index | 29.4 ± 3.2 [22.1-44.6] |
| Employment Status | |
| Employed | 13 (27.1%) |
| Not Employed | 35 (72.9%) |
| Education Level | |
| High School or Less | 15 (31.3%) |
| Some College | 12 (25.0%) |
| Undergraduate Degree | 14 (29.2%) |
| Graduate Degree | 7 (14.6%) |
| Race | |
| Black/White | 24 (50.0%)/ 24(50.0%) |
| Insurance Type | |
| Private | 18 (37.5%) |
| Public | 28 (58.3%) |
| None | 2 (4.2%) |
| Smoking Status | |
| Never Smoker | 30 (62.5%) |
| Former Smoker | 13 (27.1%) |
| Current Smoker | 5 (10.4%) |
| Income Levels | |
| Low income (<$35,000/year) | 16 (33.3%) |
| Middle income ($35,000–$74,999/year) | 18 (37.5%) |
| High income (≥$75,000/year) | 14 (29.2%) |
| Alcohol Use | |
| Yes/No | 9 (18.8%)/ 39(81.2%) |
| Healthy Diet Adherence | |
| Yes/No | 19 (39.6%)/ 29 (60.4%) |
| Routine Physical Activity | |
| Yes | 17 (35.4%) |
| No | 31 (64.6%) |
| Clinical Characteristics | |
| Colon Cancer Stage | |
| Stage II | 11 (23.1%) |
| Stage III | 37 (76.9%) |
| Years Since Diagnosis | 0.8 ± 0.9 [0.3–4.0] |
| Chemotherapy Regimen | |
| FOLFOX (Folinic acid, Fluorouracil, and Oxaliplatin) | 24 (50.0%) |
| 5-FU (single agent) | 14 (29.2%) |
| FOLFIRI (Folinic acid, Fluorouracil, and Irinotecan) | 10 (20.8%) |
| History of Colon Surgery, Yes | 48 (100.0%) |
| History of Radiation | |
| Yes | 12 (25.0%) |
| No | 36 (75.0%) |
| Comorbidity Index (≥2) | 46 (95.4%) |
| Component | Pre-Chemo (Mean ± SD) | Post-Chemo (Mean ± SD) | Change (Post – Pre) |
% Change | P value |
|---|---|---|---|---|---|
| Macronutrients | |||||
| Total Calories (kcal) | 1925 ± 420 | 1680 ± 390 | –245 | –12.7% | 0.012 |
| Protein (g) | 72.3 ± 15.8 | 64.7 ± 14.2 | –7.6 | –10.5% | 0.021 |
| Carbohydrates (g) | 230.5 ± 50.3 | 205.8 ± 48.1 | –24.7 | –10.7% | 0.033 |
| Total Fat (g) | 78.9 ± 16.7 | 69.1 ± 15.2 | –9.8 | –12.4% | 0.018 |
| Fiber (g) | 21.6 ± 5.4 | 17.9 ± 4.8 | –3.7 | –17.1% | 0.009 |
| Micronutrients | |||||
| Vitamin D (IU) | 310 ± 140 | 245 ± 115 | –65 | –21.0% | 0.041 |
| Calcium (mg) | 925 ± 210 | 812 ± 185 | –113 | –12.2% | 0.027 |
| Folate (mcg) | 380 ± 96 | 345 ± 84 | –35 | –9.2% | 0.085 |
| Iron (mg) | 13.5 ± 3.2 | 11.9 ± 2.9 | –1.6 | –11.9% | 0.034 |
| Sodium (mg) | 2300 ± 460 | 2285 ± 450 | –15 | –0.7% | 0.770 |
| Water Intake (ml) | 1400 ± 310 | 1250 ± 285 | –150 | –10.7% | 0.045 |
| Dietary Intake (Food Groups / Serving per day) | |||||
| Added Sugar (g) | 45.6 ± 14.9 | 38.2 ± 13.2 | –7.4 | –16.2% | 0.026 |
| Red Meat (servings/day) | 1.1 ± 0.5 | 0.9 ± 0.4 | –0.2 | –18.2% | 0.048 |
| Fruit & Vegetable (serving/day) | 3.4 ± 1.2 | 2.5 ± 1.0 | –0.9 | –26.5% | 0.004 |
| Whole Grains (servings/day) | 2.6 ± 1.1 | 2.0 ± 0.9 | –0.6 | –23.1% | 0.007 |
|
HEI score (0-100), the higher the score, the better the diet quality |
62.4 ± 8.5 | 54.2 ± 9.3 | -8.2 | -13.1% | 0.015 |
| Metric. | Pre-Chemotherapy | Post-Chemotherapy | p-value |
|---|---|---|---|
| Mean ± SD | 1.20 ± 0.20 | 1.05 ± 0.21 | 0.011 for mean values |
| Median [IQR] | 1.19 [1.05, 1.34] | 1.03 [0.88, 1.20] | |
| 95% CI (mean) | [1.13, 1.27] | [0.97, 1.13] |
| Phylum | Pre | Post | Δ (Post–Pre) | % Change | Wilcoxon W | p-value |
| Firmicutes_A | 51.19 | 50.63 | -0.56 | -1.1% | 1 | 0.019 |
| Bacteroidota | 29.08 | 30.68 | 1.6 | 5.5% | 1.0 | 0.003 |
| Firmicutes | 6.14 | 6.34 | 0.2 | 3.2% | 12.0 | 0.131 |
| Actinobacteriota | 6.71 | 5.11 | -1.6 | -23.8% | 0.0 | 0.002 |
| Proteobacteria | 3.62 | 3.95 | 0.33 | 9.1% | 1.0 | 0.003 |
| Firmicutes_C | 1.35 | 1.44 | 0.09 | 6.6% | 4.0 | 0.013 |
| Verrucomicrobiota | 1.19 | 1.12 | -0.07 | -5.8% | 7.0 | 0.037 |
| Methanobacteriota | 0.26 | 0.27 | 0.01 | 3.8% | 5.0 | 0.019 |
| Desulfobacterota_I | 0.20 | 0.24 | 0.04 | 20% | 0.0 | 0.002 |
| Cyanobacteria | 0.15 | 0.10 | -0.05 | -33.3% | 0.0 | 0.002 |
| Ascomycota | 0.04 | 0.02 | -0.02 | -50% | 0.0 | 0.002 |
| Firmicutes_B | 0.02 | 0.02 | 0 | 0% | 0.0 | 0.002 |
| Fusobacteriota | 0.01 | 0.01 | 0 | 0% | 0.0 | 0.002 |
| Evosea | 0.01 | 0.01 | 0 | 0% | 0.0 | 0.002 |
| Genus | Pre-Mean | Post Mean | Δ (Post–Pre) | % Change | Wilcoxon W | p-value |
| Bacteroides | 20.67 | 21.05 | +0.38 | +1.8% | 120 | 0.042 |
| Blautia_A | 12.94 | 12.31 | -0.63 | -4.8% | 109 | 0.046 |
| Phocaeicola | 9.62 | 9.26 | -0.37 | -3.8% | 110 | 0.021 |
| Agathobacter | 6.06 | 5.40 | -0.67 | -11.0% | 110 | 0.039 |
| Bifidobacterium | 5.55 | 4.64 | -0.92 | -16.5% | 130 | 0.019 |
| Faecalibacterium | 3.46 | 4.50 | +1.04 | +30.0% | 65 | 0.521 |
| Parabacteroides | 3.13 | 3.95 | +0.82 | +26.1% | 40 | 0.663 |
| Streptococcus | 3.95 | 3.07 | -0.88 | -22.2% | 135 | 0.012 |
| Alistipes | 2.19 | 4.30 | +2.11 | +96.5% | 138 | 0.312 |
| Mediterraneibacter | 3.41 | 2.81 | -0.59 | -17.2% | 55 | 0.412 |
| Prevotella | 4.00 | 1.68 | -2.32 | -58.0% | 125 | 0.031 |
| Enterocloster | 3.17 | 2.51 | -0.67 | -21.1% | 60 | 0.412 |
| Collinsella | 2.78 | 2.15 | -0.63 | -22.6% | 35 | 0.851 |
| Ruminococcus_E | 2.10 | 2.82 | +0.72 | +34.3% | 90 | 0.122 |
| Fusicatenibacter | 1.93 | 2.46 | +0.53 | +27.6% | 88 | 0.122 |
| Clostridium | 2.01 | 2.06 | +0.04 | +2.1% | 60 | 0.412 |
| Gemmiger | 1.82 | 1.82 | -0.01 | -0.2% | 90 | 0.122 |
| Roseburia | 1.71 | 1.65 | -0.06 | -3.5% | 60 | 0.412 |
| Akkermansia | 1.64 | 1.62 | -0.01 | -0.9% | 30 | 0.897 |
| Escherichia | 2.37 | 0.87 | -1.50 | -63.3% | 140 | 0.012 |
| Anaerostipes | 1.66 | 1.56 | -0.10 | -5.9% | 60 | 0.412 |
| CAG-83 | 1.50 | 1.45 | -0.05 | -3.5% | 50 | 0.632 |
| Blautia | 1.82 | 0.99 | -0.83 | -45.7% | 70 | 0.233 |
| Lactobacillus | 0.47 | 2.21 | +1.75 | +374.8% | 75 | 0.512 |
| Diversity/ Phylum |
Outcomes | Input Variables | Unadjusted B (95% CI) | Unadjusted β (p) | Adjusted B (95% CI) |
Adjusted β (p) |
|---|---|---|---|---|---|---|
|
Shanno Diversity |
Baseline Shannon Diversity |
Baseline HEI | 0.23 (0.08,0.38) | +0.42(0.01) | 0.20(0.06,0.34) | +0.38(0.02) |
| Δ Shannon Diversity | Baseline HEI | 0.31 (0.15, 0.48) | +0.61(0.01) | 0.26 (0.09,0.43) | +0.49(0.01) | |
| Δ HEI | Baseline Shannon Diversity |
0.78 (0.07, 1.15) | +0.85(0.31) | 0.75 (0.03, 1.03) | +0.79(0.32) | |
| Δ HEI | Δ Shannon Diversity |
0.19 (0.07, 0.31) | +0.35(0.01) | 0.16 (0.03, 0.29) | +0.29(0.02) | |
| Firmicutes A | Baseline Firmicutes_A | Baseline HEI | 1.12 (0.03, 2.21) | +0.32(0.04) | 0.98 (0.02, 1.94) | +0.28(0.05) |
| Δ Firmicutes_A | Baseline HEI | 0.67(−0.40,1.75) | +0.18(0.21) | 0.81 (0.04, 1.58) | +0.22(0.04) | |
| Δ HEI | Baseline Firmicutes_A |
0.66(−0.10,1.42) | +0.27(0.09) | 0.58(−0.07,1.23) | +0.23(0.08) | |
| Δ HEI | Δ Firmicutes_A | 0.61(−0.18,1.40) | +0.22(0.13) | 0.55(−0.15,1.25) | +0.20(0.12) | |
| Bacteroidota | Baseline Bacteroidota | Baseline HEI | 0.88(−0.05,1.81) | +0.29(0.07) | 0.75(−0.12,1.62) | +0.25(0.09) |
| Δ Bacteroidota | Baseline HEI | 0.55(−0.47,1.56) | +0.14(0.29) | 0.60(−0.25,1.46) | +0.19(0.15) | |
| Δ HEI | Baseline Bacteroidota |
0.52(−0.20,1.24) | +0.21(0.15) | 0.45(−0.20,1.10) | +0.18(0.17) | |
| Δ HEI | Δ Bacteroidota | 0.48(−0.24,1.20) | +0.19(0.18) | 0.42(−0.22,1.06) | +0.17(0.19) | |
| Firmicutes | Baseline Firmicutes | Baseline HEI | 1.50 (0.30, 2.70) | +0.38(0.02) | 1.35 (0.15, 2.55) | +0.34(0.03) |
| Δ Firmicutes | Baseline HEI | 0.91(−0.27,2.08) | +0.22(0.13) | 1.02 (0.05, 1.99) | +0.24(0.04) | |
| Δ HEI | Baseline Firmicutes |
0.57(−0.11,1.25) | +0.24(0.10) | 0.50(−0.12,1.12) | +0.21(0.11) | |
| Δ HEI | Δ Firmicutes | 0.73 (0.01, 1.45) | +0.30(0.05) | 0.65 (0.01, 1.29) | +0.27(0.06) | |
| Actinobacteriota | Baseline Actinobacteriota | Baseline HEI | 0.49(−0.18,1.16) | +0.21(0.15) | 0.42(−0.23,1.07) | +0.18(0.20) |
| Δ Actinobacteriota | Baseline HEI | 0.27(−0.27,0.81) | +0.12(0.33) | 0.43(−0.06,0.91) | +0.17(0.09) | |
| Δ HEI | Baseline Actinobacteriota |
0.40(−0.25,1.06) | +0.16(0.23) | 0.35(−0.27,0.97) | +0.14(0.26) | |
| Δ HEI | Δ Actinobacteriota |
0.31(−0.32,0.94) | +0.12(0.33) | 0.25(−0.34,0.84) | +0.10(0.40) | |
| Proteobacteria | Baseline Proteobacteria | Baseline HEI | −1.11(−1.86,−0.36) | −0.40(0.01) | −1(−1.80,−0.20) | −0.36(0.02) |
| Δ Proteobacteria | Baseline HEI | −0.94(−1.67,−0.21) | −0.32(0.03) | −0.87(−1.68,−0.06) | −0.29(0.04) | |
| Δ HEI | Baseline Proteobacteria |
−0.95(−1.70,−0.20) | −0.38(0.02) | −0.85(−1.62,−0.08) | −0.34(0.03) | |
| Δ HEI | Δ Proteobacteria | −0.88(−1.60,−0.15) | −0.33(0.03) | −0.80(−1.55, −0.05) | −0.30(0.04) |
| Genus | Outcomes | Input Variables | Unadjusted β (95% CI) | p-value | Adjusted B (95% CI) | p-value |
|---|---|---|---|---|---|---|
| Bacteroides | Baseline | Baseline HEI | +0.15 (-0.07, 0.37) | 0.18 | +0.10 (-0.12, 0.32) | 0.37 |
| Δ | Baseline HEI | +0.12 (-0.10, 0.34) | 0.28 | +0.08 (-0.14, 0.30) | 0.47 | |
| Baseline | Δ HEI | +0.08 (-0.14, 0.30) | 0.47 | +0.05 (-0.17, 0.27) | 0.65 | |
| Δ | Δ HEI | +0.21 (-0.03, 0.45) | 0.09 | +0.18 (-0.07, 0.43) | 0.15 | |
| Blautia_A | Baseline | Baseline HEI | -0.22 (-0.42, -0.02) | 0.03 | -0.18 (-0.38, 0.02) | 0.08 |
| Δ | Baseline HEI | -0.15 (-0.35, 0.05) | 0.14 | -0.12 (-0.32, 0.08) | 0.24 | |
| Baseline | Δ HEI | -0.10 (-0.30, 0.10) | 0.33 | -0.07 (-0.27, 0.13) | 0.49 | |
| Δ | Δ HEI | -0.25 (-0.47, -0.03) | 0.03 | -0.21 (-0.43, 0.01) | 0.06 | |
| Phocaeicola | Baseline | Baseline HEI | +0.10 (-0.12, 0.32) | 0.37 | +0.07 (-0.15, 0.29) | 0.54 |
| Δ | Baseline HEI | -0.05 (-0.27, 0.17) | 0.65 | -0.03 (-0.25, 0.19) | 0.79 | |
| Baseline | Δ HEI | +0.03 (-0.19, 0.25) | 0.79 | +0.01 (-0.21, 0.23) | 0.93 | |
| Δ | Δ HEI | -0.12 (-0.34, 0.10) | 0.29 | -0.09 (-0.31, 0.13) | 0.42 | |
| Agathobacter | Baseline | Baseline HEI | -0.18 (-0.38, 0.02) | 0.08 | -0.15 (-0.35, 0.05) | 0.14 |
| Δ | Baseline HEI | -0.22 (-0.42, -0.02) | 0.03 | -0.19 (-0.39, 0.01) | 0.06 | |
| Baseline | Δ HEI | -0.07 (-0.27, 0.13) | 0.49 | -0.05 (-0.25, 0.15) | 0.63 | |
| Δ | Δ HEI | -0.28 (-0.50, -0.06) | 0.01 | -0.24 (-0.46, -0.02) | 0.03 | |
| Bifidobacterium | Baseline | Baseline HEI | +0.19 (0.02, 0.36) | 0.03 | +0.15 (-0.02, 0.32) | 0.09 |
| Δ | Baseline HEI | +0.10 (-0.07, 0.27) | 0.25 | +0.07 (-0.10, 0.24) | 0.42 | |
| Baseline | Δ HEI | +0.12 (-0.05, 0.29) | 0.17 | +0.09 (-0.08, 0.26) | 0.30 | |
| Δ | Δ HEI | -0.10 (-0.26, 0.06) | 0.22 | -0.08 (-0.24, 0.08) | 0.33 | |
| Faecalibacterium | Baseline | Baseline HEI | +0.31 (0.08, 0.54) | 0.01 | +0.26 (0.03, 0.49) | 0.03 |
| Δ | Baseline HEI | +0.25 (0.02, 0.48) | 0.04 | +0.21 (-0.02, 0.44) | 0.07 | |
| Baseline | Δ HEI | +0.18 (-0.05, 0.41) | 0.12 | +0.14 (-0.09, 0.37) | 0.23 | |
| Δ | Δ HEI | +0.38 (0.14, 0.62) | <0.01 | +0.33 (0.09, 0.57) | 0.01 | |
| Parabacteroides | Baseline | Baseline HEI | +0.13 (-0.09, 0.35) | 0.24 | +0.09 (-0.13, 0.31) | 0.41 |
| Δ | Baseline HEI | +0.22 (0.00, 0.44) | 0.05 | +0.18 (-0.04, 0.40) | 0.11 | |
| Baseline | Δ HEI | +0.15 (-0.07, 0.37) | 0.18 | +0.12 (-0.10, 0.34) | 0.28 | |
| Δ | Δ HEI | +0.26 (0.02, 0.50) | 0.03 | +0.22 (-0.02, 0.46) | 0.07 | |
| Streptococcus | Baseline | Baseline HEI | -0.20 (-0.40, 0.00) | 0.05 | -0.16 (-0.36, 0.04) | 0.12 |
| Δ | Baseline HEI | -0.25 (-0.45, -0.05) | 0.01 | -0.21 (-0.41, -0.01) | 0.04 | |
| Baseline | Δ HEI | -0.12 (-0.32, 0.08) | 0.24 | -0.09 (-0.29, 0.11) | 0.38 | |
| Δ | Δ HEI | -0.30 (-0.52, -0.08) | <0.01 | -0.26 (-0.48, -0.04) | 0.02 | |
| Alistipes | Baseline | Baseline HEI | +0.11 (-0.11, 0.33) | 0.32 | +0.07 (-0.15, 0.29) | 0.53 |
| Δ | Baseline HEI | +0.35 (0.13, 0.57) | <0.01 | +0.31 (0.09, 0.53) | <0.01 | |
| Baseline | Δ HEI | +0.14 (-0.08, 0.36) | 0.21 | +0.10 (-0.12, 0.32) | 0.37 | |
| Δ | Δ HEI | +0.42 (0.18, 0.66) | <0.01 | +0.37 (0.13, 0.61) | <0.01 | |
| Mediterraneibacter | Baseline | Baseline HEI | -0.16 (-0.36, 0.04) | 0.12 | -0.13 (-0.33, 0.07) | 0.20 |
| Δ | Baseline HEI | -0.19 (-0.39, 0.01) | 0.06 | -0.16 (-0.36, 0.04) | 0.12 | |
| Baseline | Δ HEI | -0.08 (-0.28, 0.12) | 0.43 | -0.06 (-0.26, 0.14) | 0.56 | |
| Δ | Δ HEI | -0.23 (-0.45, -0.01) | 0.04 | -0.20 (-0.42, 0.02) | 0.08 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




