Submitted:
19 December 2025
Posted:
19 December 2025
You are already at the latest version
Abstract
Background/Objectives: Bacterial biofilms formed by Escherichia coli pose a significant challenge in veterinary medicine due to their intrinsic resistance to antibiotics. Antimicrobial peptides (AMPs) represent a promising alternative. AMPs exert their bactericidal activity by binding to negatively charged phospholipids in bacterial membranes via electrostatic interactions, leading to membrane disruption and rapid cell lysis. Methods: In vitro assays included MIC determination, biofilm eradication testing (crystal violet, colony counts, CLSM), swimming motility, and EPS quantification. CRISPR/Cas9 was used to construct and complement a kduD mutant. A transposon mutagenesis library was screened for biofilm-defective mutants. In vivo, a murine excisional wound infection model was treated with CRAMP-34, with wound closure and bacterial burden monitored. Gene expression changes were analyzed via RT-qPCR. Results: The mouse-derived AMP (abbreviation CRAMP-34) effectively eradicates pre-formed biofilms of a clinically relevant, porcine-origin E.coli strain and promotes wound healing in a murine infection model. We conducted a genome-wide transposon mutagenesis screen, which identified kduD, as a critical gene for robust biofilm formation. Functional characterization revealed that kduD deletion drastically impairs flagellar motility and alters exopolysaccharide production, leading to defective biofilm architecture without affecting growth. Notably, the anti-biofilm activity of CRAMP-34 phenocopied aspects of the kduD deletion, including motility inhibition and transcriptional repression of a common set of biofilm-related genes. Conclusions: The research highlight CRAMP-34 as a potent anti-biofilm agent and unveil kduD as a previously unrecognized regulator of E.coli biofilms development, whose associated pathway is implicated in the mechanism of action of CRAMP-34.
Keywords:
1. Introduction
2. Results
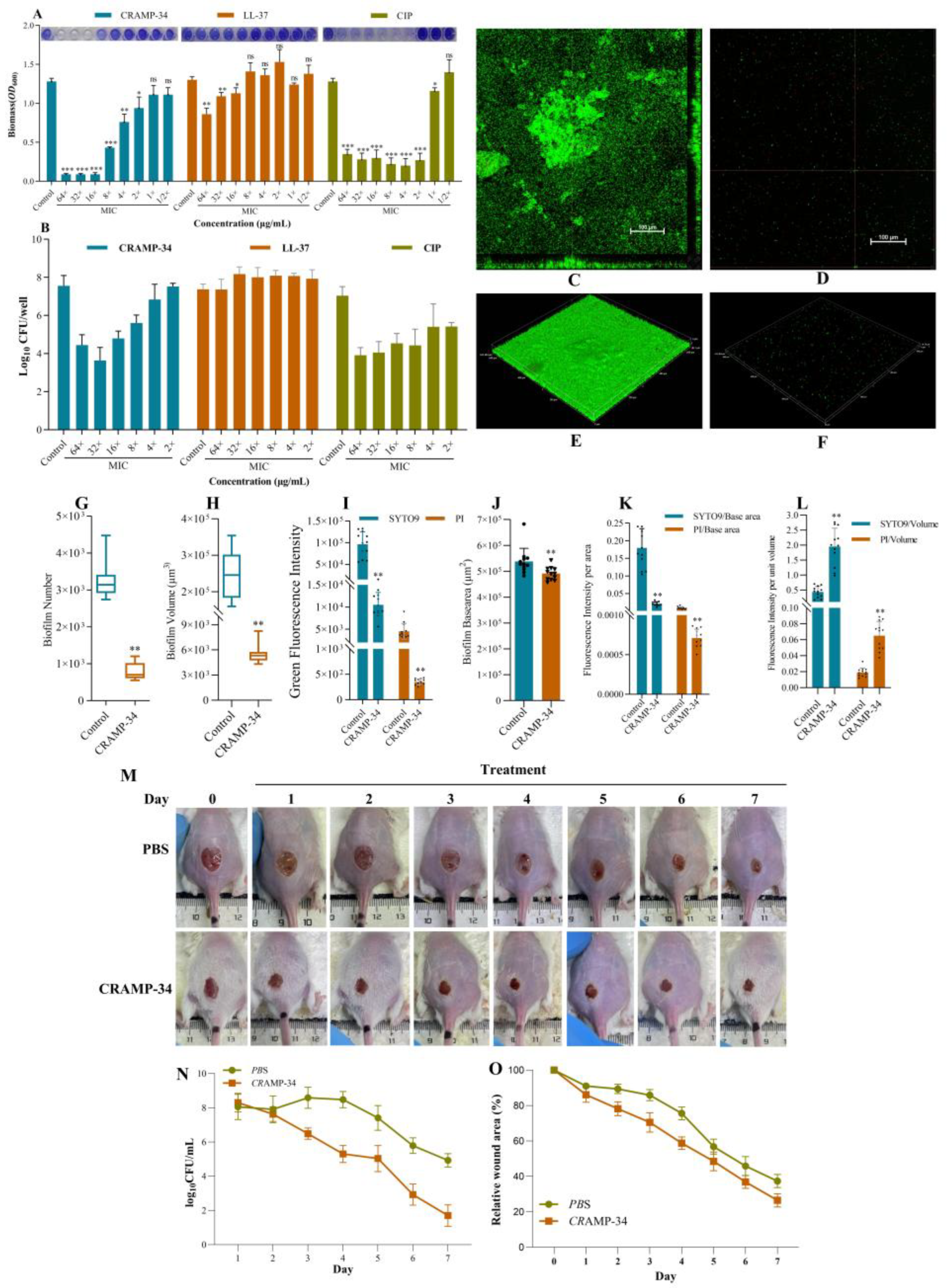
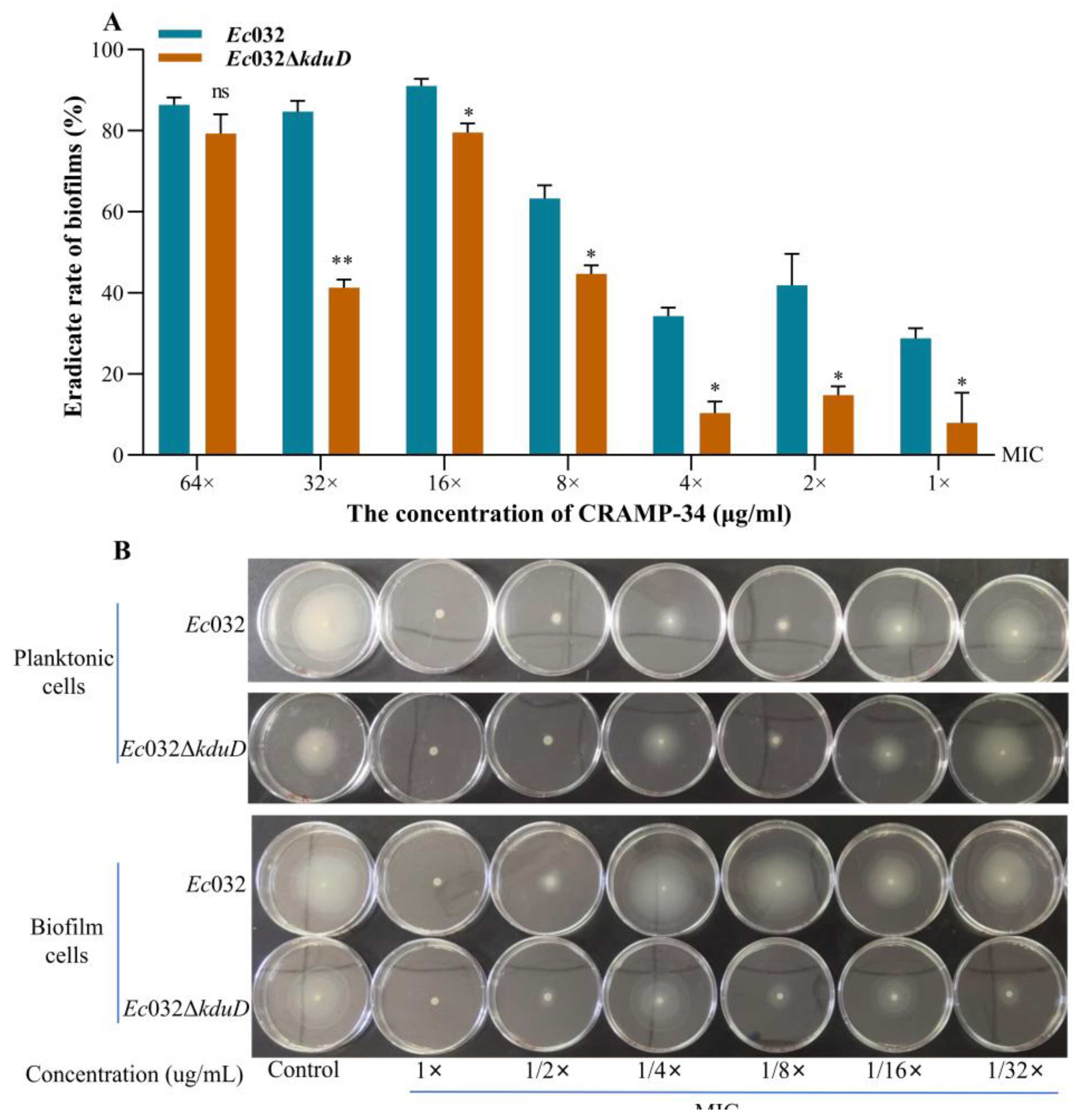
2.1. CRAMP-34 Demonstrates Potent Biofilm-Eradicating Activity and Promotes Wound Healing In Vivo
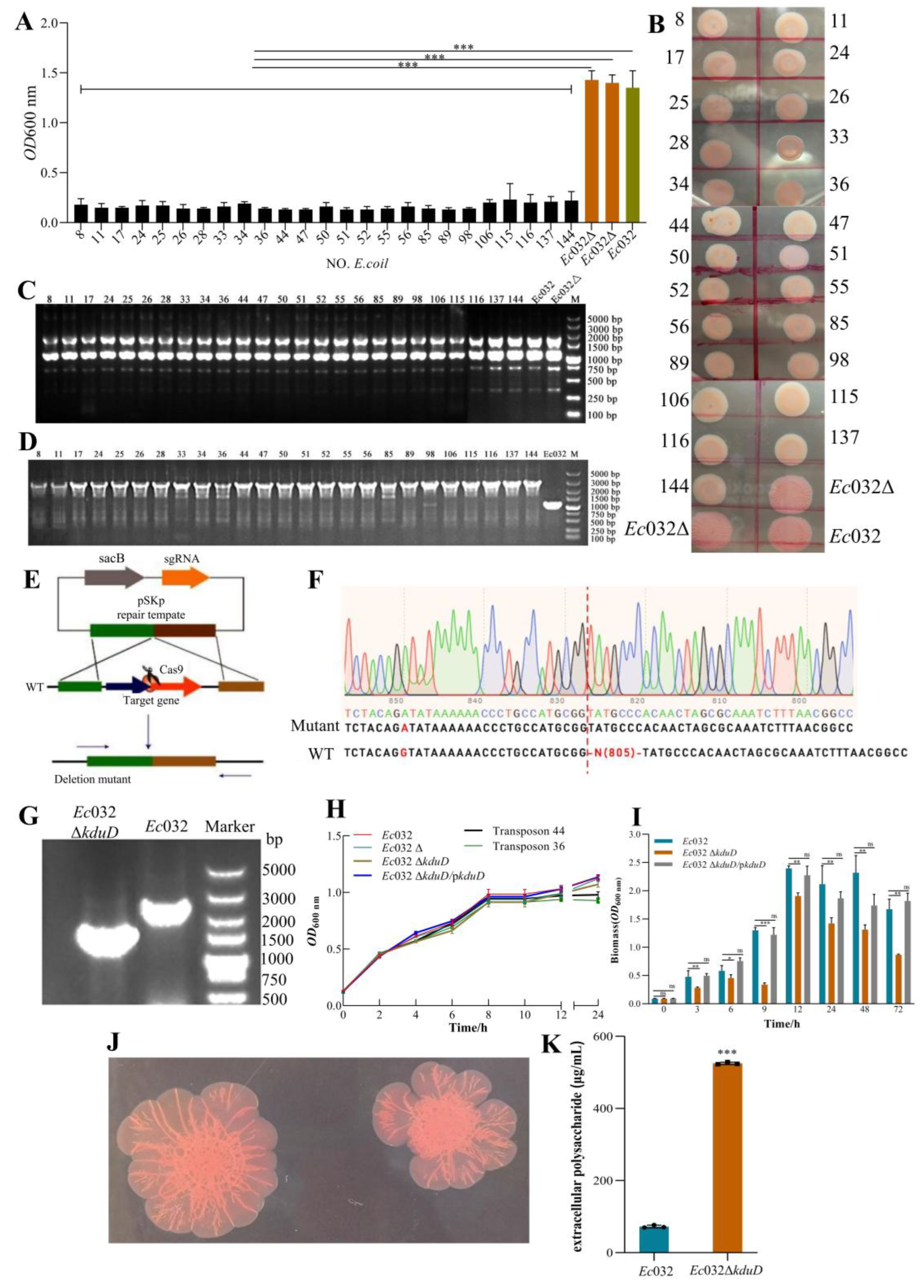
2.2. A Genome-Wide Screen Identifies kduD as a Novel Gene Essential for Robust Biofilm Formation
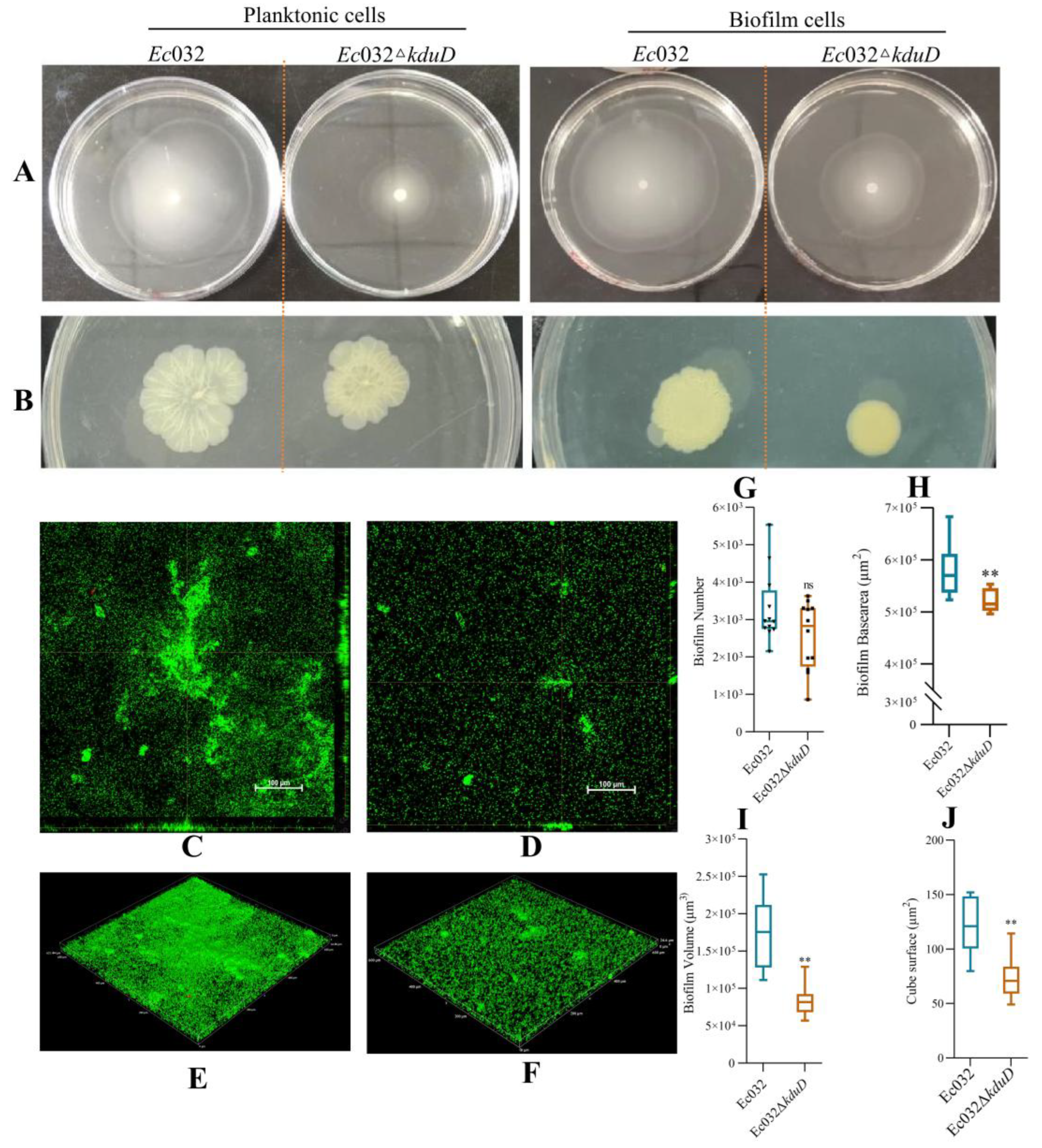
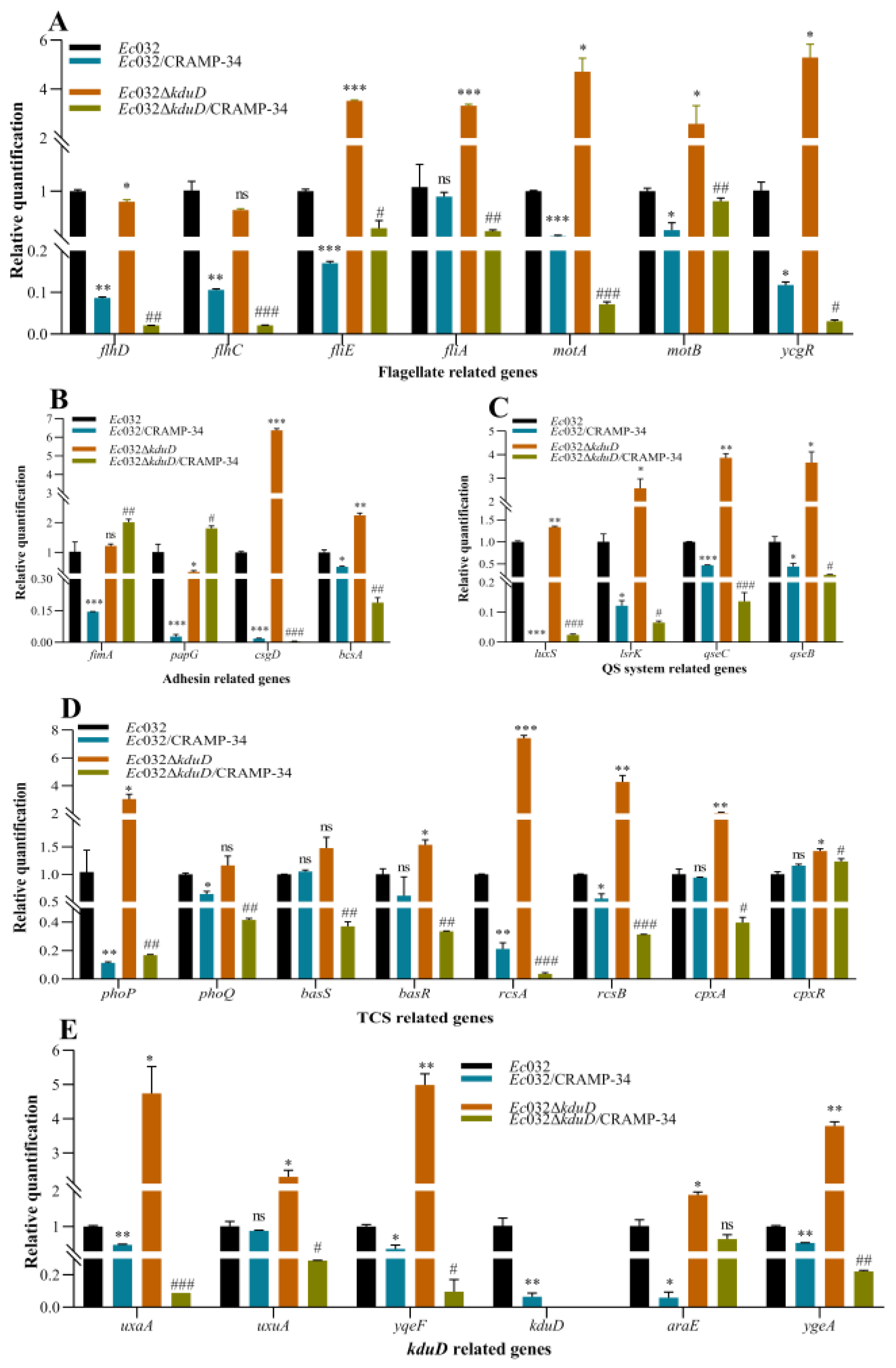
2.3. kduD Regulates Biofilm by Modulating Flagellar Motility and EPS Production
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains, Plasmids, Primers, and Growth Conditions
4.2. Determination of Minimum Inhibitory Concentration (MIC)
4.3. Biofilms Formation and Antibiofilm Assay
4.4. Skin Wound Infection Model Was Established
4.5. Crystal Violet Staining and Colony Count Assay
4.6. Swimming Motility Assay
4.7. Confocal Laser Scanning Microscopy (CLSM)
4.8. Screening of Gene Mutants with Decreased Biofilms Formation
4.9. Generation of a Mutagenesis Library of Ec032
4.10. Identification of Transposon Insertion Sites
4.11. kduD Mutant Construction Using CRISPR/Cas9 System
4.12. The Plasmid Deletion Strains Ec032ΔkduD Were Obtained
4.13. Growth Curves
4.14. Congo Red-Binding Assay and EPS Assay
4.15. Real-Time Fluorescence Quantitative PCR (RT-qPCR)
4.16. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| E.coli | Escherichia coli |
| CRAMP-34 | Cathelicidins related antimicrobial peptide |
| EPS | exopolysaccharides |
| AMP | Antibacterial peptide |
| PCR | Polymerase chain reaction |
| QS | Quorum sensing |
| TCS | Two omponents system |
| c-di-GMP | Cyclic diguanosine monophosphate |
| KduD | 2-dehydro-3-deoxy-D-gluconate 5-dihydrogenase |
| CIP | Ciprofloxacin |
| MIC | Minimum inhibitory concentration |
| CLSM | Confocal Laser Scanning Microscope |
| RT-qPCR | Real-time fluorescent quantitative PCR |
References
- Yang H, Liang Y, Yang Z, Liu L, Ran L, Liu J, Ma C, Wei W, Zhang S, Zhu M et al: Paeonol eradicates biofilm in porcine-source Escherichia coli by targeting the quorum sensing system. BMC VET RES 2025, 21(1):644. [CrossRef]
- Long J, Yang C, Liu J, Ma C, Jiao M, Hu H, Xiong J, Zhang Y, Wei W, Yang H et al: Tannic acid inhibits Escherichia coli biofilm formation and underlying molecular mechanisms: Biofilm regulator CsgD. BIOMED PHARMACOTHER 2024, 175:116716. [CrossRef]
- Liu L, Li H, Ma C, Liu J, Zhang Y, Xu D, Xiong J, He Y, Yang H, Chen H: Effect of anti-biofilm peptide CRAMP-34 on the biofilms of Acinetobacter lwoffii derived from dairy cows. FRONT CELL INFECT MI 2024, 14:1406429. [CrossRef]
- Ma C, Mei C, Liu J, Li H, Jiao M, Hu H, Zhang Y, Xiong J, He Y, Wei W et al: Effect of baicalin on eradicating biofilms of bovine milk derived Acinetobacter lwoffii. BMC VET RES 2024, 20(1):212. [CrossRef]
- Tang A, Li C, Feng D, Li A: Deciphering the code of temperature rise on aerobic granular sludge stability: A DSF-c-di-GMP mediated regulatory mechanism. ENVIRON RES 2025, 267:120705. [CrossRef]
- Perrin C, Briandet R, Jubelin G, Lejeune P, Mandrand-Berthelot M, Rodrigue A, Dorel C: Nickel promotes biofilm formation by Escherichia coli K-12 strains that produce curli. APPL ENVIRON MICROB 2009, 75(6):1723-1733. [CrossRef]
- Manobala T: Peptide-based strategies for overcoming biofilm-associated infections: a comprehensive review. CRIT REV MICROBIOL 2025, 51(4):563-580. [CrossRef]
- Gong H, He L, Zhao Z, Mao X, Zhang C: The specific effect of (R)-(+)-pulegone on growth and biofilm formation in multi-drug resistant Escherichia coli and molecular mechanisms underlying the expression of pgaABCD genes. BIOMED PHARMACOTHER 2021, 134:111149. [CrossRef]
- Shi Z, Zhai J, Yu J, Wang Z, Liu H, Yang X, Wang X: Biofilm formation by Gallibacterium anatis depends on TolC-mediated initial attachment of cells. Veterinary journal (London, England : 1997) 2025, 314:106488. [CrossRef]
- Dweba Y, Eleojo Aruwa C, Sabiu S: In Silico Bioprospection of Daniellia oliveri-Based Products as Quorum Sensing Modulators of Escherichia coli SdiA. BIOCHEM RES INT 2025, 2025:7191508. [CrossRef]
- Niu N, Dou L, Yang S, Wang H, Zhuang S, Fan Y, Liu Y, Zhang W, Ma W: Drug resistance detection of canine origin Escherichia coli in China and inhibition by genipin. Veterinary journal (London, England : 1997) 2025, 310:106307. [CrossRef]
- Wang S, Ma C, Long J, Cheng P, Zhang Y, Peng L, Fu L, Yu Y, Xu D, Zhang S et al: Impact of CRAMP-34 on Pseudomonas aeruginosa biofilms and extracellular metabolites. FRONT CELL INFECT MI 2023, 13:1295311. [CrossRef]
- Wang Z, Tang Y, Li H, Li J, Chi X, Ma X, Liu Z: ArgR regulates motility and virulence through positive control of flagellar genes and inhibition of diguanylate cyclase expression in Aeromonas veronii. COMMUN BIOL 2024, 7(1):1720. [CrossRef]
- Sun G, Yu Z, Li Q, Zhang Y, Wang M, Liu Y, Liu J, Liu L, Yu X: Mechanism of Escherichia coli Lethality Caused by Overexpression of flhDC, the Flagellar Master Regulator Genes, as Revealed by Transcriptome Analysis. International journal of molecular sciences 2023, 24(18):14058. [CrossRef]
- Visnardi AB, Ribeiro RA, de Souza AS, Churasacari Vinces TG, Llontop EE, de Almeida Ferrari AS, França Henrique PA, Valdivieso D, Sánchez-Limache DE, Silva GR et al: Insertion of a Divergent GAF-like Domain Defines a Novel Family of YcgR Homologues That Bind c-di-GMP in Leptospirales. ACS OMEGA 2025, 10(4):3988-4006. [CrossRef]
- Chen Y, Liu Z, Chen B, Tam H, Shia W, Yu H, Chen P: Effects of Heat-Killed Probiotic Strains on Biofilm Formation, Transcription of Virulence-Associated Genes, and Prevention of UTIs in Mice. PROBIOTICS ANTIMICRO 2025, 17(6):4619-4634. [CrossRef]
- Wang Y, Zhang R, Wang P, Zhang W, Li Z, Pang X, Huang F, Wang S, Liu X, Zhang H: Biofilm-mediated resistance to berberine in Escherichia coli. FRONT CELL INFECT MI 2025, 15:1565714. [CrossRef]
- Chen S, Feng Z, Sun H, Zhang R, Qin T, Peng D: Biofilm-Formation-Related Genes csgD and bcsA Promote the Vertical Transmission of Salmonella Enteritidis in Chicken. FRONT VET SCI 2021, 7:625049. [CrossRef]
- Li S, Bi C, Xiang B, Wang Z, Yang H, Fu C, Chen L, Chen Y: The role of cyclic di-GMP in biomaterial-associated infections caused by commensal Escherichia coli. PLOS ONE 2025, 20(8):e330229. [CrossRef]
- Shrestha S, Bista S, Byanjankar N, Prasai Joshi T: Evaluation of bottled drinking water and occurrence of multidrug-resistance and biofilm producing bacteria in Nepal. Environmental pollution (Barking, Essex : 1987) 2024, 341:122896. [CrossRef]
- Li F, Cao L, Bähre H, Kim S, Schroeder K, Jonas K, Koonce K, Mekonnen SA, Mohanty S, Bai F et al: Patatin-like phospholipase CapV in Escherichia coli - morphological and physiological effects of one amino acid substitution. NPJ BIOFILMS MICROBI 2022, 8(1):39. [CrossRef]
- Vianney A, Jubelin G, Renault S, Dorel C, Lejeune P, Lazzaroni JC: Escherichia coli tol and rcs genes participate in the complex network affecting curli synthesis. Microbiology (Reading, England) 2005, 151(Pt 7):2487-2497. [CrossRef]
- Chen C, Nguyen LT, Cottrell BJ, Irwin PL, Uhlich GA: Multiple mechanisms responsible for strong Congo-red-binding variants of Escherichia coli O157:H7 strains. PATHOG DIS 2016, 74(2):ftv123. [CrossRef]
- Tubeleviciute A, Teese MG, Jose J: Escherichia coli kduD encodes an oxidoreductase that converts both sugar and steroid substrates. APPL MICROBIOL BIOT 2014, 98(12):5471-5485. [CrossRef]
- Maruyama Y, Oiki S, Takase R, Mikami B, Murata K, Hashimoto W: Metabolic fate of unsaturated glucuronic/iduronic acids from glycosaminoglycans: molecular identification and structure determination of streptococcal isomerase and dehydrogenase. The Journal of biological chemistry 2015, 290(10):6281-6292.
- Hwang Y, Perez M, Holzel R, Harshey RM: c-di-GMP is required for swarming in E. coli, producing colanic acid that acts as surfactant. MBIO 2025, 16(6):e91625. [CrossRef]
- Hodge-Hanson KM, Zoino A, Downs DM: Expression of Pyridoxal 5'-Phosphate-Independent Racemases Can Reduce 2-Aminoacrylate Stress in Salmonella enterica. J BACTERIOL 2018, 200(9):e717-e751. [CrossRef]
- Miyamoto T, Saitoh Y, Katane M, Sekine M, Homma H: YgeA is involved in L- and D-homoserine metabolism in Escherichia coli. FEMS MICROBIOL LETT 2022, 369(1):fnac96. [CrossRef]
- Zhang Y, Cheng P, Wang S, Li X, Peng L, Fang R, Xiong J, Li H, Mei C, Gao J et al: Pseudomonas aeruginosa biofilm dispersion by the mouse antimicrobial peptide CRAMP. VET RES 2022, 53(1):80. [CrossRef]
- Monikadevi G, Vijayakumar S, Vanathi P, Prathipkumar S, Al-Sadoon MK, Srinivasan P, Vidhya E: Fruit Extract Mediated Formation of Luminescent Titanium Dioxide Nanometer-Sized Particles: An Innovative Strategy in Domain of Photodecomposition and Germicidal Properties. LUMINESCENCE 2024, 39(11):e70025. [CrossRef]
| Strains/Plasmids | Description |
|---|---|
| Ec032 | Clinical isolation of IncX4 plasmid strain carrying mcr-1, Colr |
| Ec032Δ | plasmid was cured |
| Ec032ΔkduD | kduD gene deletd |
| E. coli DH5α | cloning vectors |
| E. coli WM3064 | a diaminopimelic acid (DAP) auxotroph strain |
| pCure-oriT-GFP-MCR | Aprr, oriT+, GFP+, sacB+ |
| pCat-arr | Mariner transposon, Rifr, oriT+ |
| pCasKp-OriT | Aprr, oriT+, bacterial expression of Cas9 nuclease and λ-Red recombination system, with temperature-sensitive replication |
| pSGKp-arr2 | Rifr, sacB+, a sgRNA expression plasmid for targeting a specific sequence |
| pSGKp-Ec032-kduD | pSGKp derivative with the spacer of the kduD gene |
| pSGKp-Ec032-kduD-500 | pSGKp derivative with the repair arms of the kduD gene |
| Primer | Sequence (5’-3’) |
|---|---|
| Eric-F | ATGTAAGCTCCTGGGGATTCAC |
| Eric-R | AAGTAAGTGACTGGGGTGAGCG |
| Ec032-F-XJ | GTCGAGAATTTCCGCGCTAC |
| Ec032-R-XJ | GTATTGATACCGGCACTCCG |
| infu-kduD-N20F | CACATAATCTGAAGCGCTGGGTTTTAGAGCTAGAAATAGCAAGTTAAAATAAGGC |
| infu-kduD-N20R | CCAGCGCTTCAGATTATGTGACTAGTATTATACCTAGGACTGAGCTAGC |
| kduD-N20-F | CACATAATCTGAAGCGCTGG |
| kduD-F1 | TCGAATTCCTGCAGCCCGGGGGATCCTATTTACGCCCCAGGCGGAA |
| kduD-R1 | CCGCATGGCAGGGTTTTTTA |
| kduD-F2 | TAAAAAACCCTGCCATGCGGTATGCCCACAACTAGCGCAA |
| kduD-R2 | TCCACCGCGGTGGCGGCCGCTCTAGATTACTGTCGATGGCCAATGC |
| kduD-F | TGCCGAGTGTGACCATCAAC |
| kduD-R | CGGAGGTTGATGTGGACGTA |
| Gene | Forward primer sequence | Reverse primer sequence |
|---|---|---|
| gapA | GTTGTCGCTGAAGCAACTGG | CGATGTCCTGGCCAGCATAT |
| flhC | ATGCTGCCATTCTCAACCGA | GCTTGTGGGCACTGTTCAAG |
| fliE | GACCATTAGTTTTGCCGGGC | ACGCACCTGAATCCCCATTT |
| fliA | GAACGCTATGACGCCCTACA | TCCAGTTGCCCTATTGCCTG |
| motA | CGCCGAAACCAGCAAAATGA | TCCTCGGTTGTCGTCTGTTG |
| motB | TGACTGCGATGATGGCCTTT | CCCCTGGCTTTGGGTGTAAT |
| ycgR | GGGGCAATGGGGTGTTTTTC | CTTTGTCCGCTTTTTCCCGG |
| fimA | TTGTTCTGTCGGCTCTGTCC | ACTGGTTGCTCCTTCCTGTG |
| papG | TTCGCATCGTGAAACAGCAC | TACGTTTCGCTTCCATGGCT |
| csgD | GATTACCCGTACCGCGACAT | GCGTAATCAGGTAGCTGGCA |
| bcsA | AACGAAGGCACGCTGTTCTA | GAGGTATAGCCACGACGGTG |
| luxS | TTGGTACGCCAGATGAGCAG | ACGTCACGTTCCAGAATGCT |
| lsrK | CAGATTACTTTGGCTGGCGC | CGTAGGCCAGCCATATCCAG |
| qseC | CGTGACCCTGACTCGGAAAA | TTCGGTTTGCACTTTCAGCG |
| qseB | ATTGGCGACGGCATCAAAAC | CACGCTGACCTTTTTCTCGC |
| phoP | GCGCGTACTGGTTGTTGAAG | CTGGCAATCCGAGATCGACA |
| basS | CCTGCTGCGGATGTTATTGC | ACGCTTTACTCAACTCCCCG |
| basR | GTACTGATCCTCACCGCTCG | GACCCATGTTCAGCGTCAGA |
| rcsA | ATTGAGCCGAACCGAATCGA | GTCAGTCGGACGACATGGTA |
| rcsB | ATCAGTGCTGGTGGTTACGG | TCAGCAGGGCGATATCGTTC |
| cpxA | CGCAGGTGCCAGTTTTAACC | CAGTTCCTTGCTTTCACCGC |
| cpxR | TTAGTGCTGAATCCAGGCCG | CGTTTGCCCAACACTTCCTG |
| kduD | AACCATCGAGCAGGTCACAG | TCGCTGAACTCGAGAGCATC |
| uxuA | ACCAGATCGAATTCGCTGCA | CCCGGAAGACCAGCAATGAT |
| uxaA | GTGATTGGTCTGGGCTGTGA | CTGGCTCGCGTTTATCGTTG |
| araE | TTACCTGTTCGACCACCACG | GGTTTCCGGAATGAGCCAGA |
| ygeA | ACGATGCATAAAGTGGCGGA | AAAATTGTTCCGTCAGCCGC |
| yqeF | TTGCCAGCGTTGGTGTAGAT | ATTGACATTGACCCGACGCT |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




