Submitted:
05 December 2025
Posted:
08 December 2025
You are already at the latest version
Abstract
Weakened forest stands provide a favorable environment for the development of saprotrophic and necrotrophic fungi. Traditional identification methods for these ecological groups of fungi are not always effective. However, modern DNA barcoding methods based on sequencing DNA regions of the studied fungal specimens and comparing them 31 with the nucleotide sequences of previously classified organisms often allow for precise species identification. Thus, to identify fungi using barcoding in weakened Scots pine (Pinus sylvestris L.) stands in the Central Forest-Steppe of European Russia (Voronezh, Voronezh Region), 55 mycelium and fruiting body samples were collected from conifer litter and Scots pine damaged butt part of the trunk and roots. After morphological and ecological description and systematization of the collection, DNA extraction was performed, the ITS1 and ITS2 regions were sequenced in 21 samples, and 11 species of xylotrophic fungi were identified, of which three species — Hirschioporus fuscoviolaceus (Ehrenb.) Donk, Gymnopilus penetrans (Fr.) Murrill, and Ganoderma applanatum (Pers.) Pat. — infect living wood. The remaining eight representatives of the mycobiota were saprotrophic fungi involved in the mineralization of organic residues. Two phyla, Ascomycota and Basidiomycota, were identified, with the most samples belonging to Basidiomycota and the class Agaricomycetes with seven orders. The predominance of Basidiomycota species indicates stages III-IV mycogenic xylolysis of the weakened stands analyzed. The DNA barcoding method complements the results of morphological and ecological analysis and provides a more complete mycological picture of wood-destroying fungi in weakened pine forests that are difficult to identify.
Keywords:
1. Introduction
2. Materials and Methods
Fungi Sampling
DNA Isolation and PCR
DNA Sequencing and Barcoding
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Xu, J. Fungal DNA barcoding. Genome 2016, 59, 913–932. [Google Scholar] [CrossRef]
- Ćelepirović, N.; Agbaba, S.N.; Vlahović, M.K. DNA barcoding of fungi in the forest ecosystem of the Psunj and Papuk Mountains in Croatia. South-East European forestry 2020, 11, 145–152. [Google Scholar] [CrossRef]
- Guo, M.; Yuan, C.; Tao, L.; Cai, Y.; Zhang, W. Life barcoded by DNA barcodes. Conservation Genetics Resources 2022, 14, 351–365. [Google Scholar] [CrossRef] [PubMed]
- Valentini, A.; Pompanon, F.; Taberlet, P. DNA barcoding for ecologists. Trends in Ecology and Evolution 2009, 24, 110–7. [Google Scholar] [CrossRef]
- BLAST: Basic Local Alignment Search Tool. Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi (accessed on 9 June 2025).
- Gong, S.; Ding, Y.; Wang, Y.; Jiang, G.; Zhu, C. Advances in DNA barcoding of toxic marine organisms. International Journal of Molecular Sciences 2018, 19, 2931. [Google Scholar] [CrossRef] [PubMed]
- Purty, R.S.; Chatterjee, S. DNA barcoding: an effective technique in molecular taxonomy. Austin Journal of Biotechnology & Bioengineering 2016, 3, 1059. Available online: https://austinpublishinggroup.com/biotechnology-bioengineering/fulltext/ajbtbe-v3-id1059.php (accessed on 25 October 2025).
- Ji, Y.; Huotari, T.; Roslin, T.; Schmidt, N.S.; Wang, J.; Yu, D.W.W.; Ovaskainen, O. Spikepipe: a metagenomic pipeline for the accurate quantification of eukaryotic species occurrences and intraspecific abundance change using DNA barcodes or mitogenomes. Molecular Ecology Resources 2020, 20, 256–267. [Google Scholar] [CrossRef]
- Brichta, J.; Vacek, S.; Vacek, Z.; Cukor, J.; Mikeska, M.; Bílek, L.; Šimůnek, V.; Gallo, J.; Brabec, P. Importance and potential of Scots pine (Pinus sylvestris L.) in 21st century. Central European Forestry Journal 2023, 69, 3–20. [Google Scholar] [CrossRef]
- Rosleskhoz Rosleskhoz: ploshchadʹ zemelʹ lesnogo fonda za 2023 god vyrosla na 107 tys. ga [Press release]. Federal Forestry Agency of Russia. 21 May 2024. Available online: https://rosleshoz.gov.ru/news/federal/rosleskhoz-ploshchad-zemel-lesnogo-fonda-za-2023-god-vyrosla-na-107-tysyach-gektarov-n10998 (accessed on 26 October 2025). (In Russian).
- Musievsky, A.L. Trends in the dynamics of the forest fund of the Central forest-steppe of the European part of Russia for the period 1966-2017. Proceedings of the Saint Petersburg Forestry Research Institute 2022, 2, 58–69, (In Russian with English Abstract). [Google Scholar] [CrossRef]
- Lipka, O.N.; Korzukhin, M.D.; Zamolodchikov, D.G.; Dobrolyubov, N.Yu.; Krylenko, S.V.; Bogdanovich, A.Yu.; Semenov, S.M. A Role of Forests in Natural Systems Adaptation to Climate Change. Russian Journal of Forest Science (Lesovedenie) 2021, 5, 531–546, (In Russian with English abstract). [Google Scholar] [CrossRef]
- Zamolodchikov, D.; Shvidenko, A.; Bartalev, S.; Kulikova, E.; Held, A.; Valentini, R.; Lindner, M. State of Russian forests and forestry. Chapter 2. In Russian forests and climate change. What Science Can Tell Us 11; Leskinen, P, Lindner, M, Verkerk, PJ, Nabuurs, GJ, Van Brusselen, J, Kulikova, E, Hassegawa, M, Lerink, B, Eds.; European Forest Institute, 2020; p. 21. [Google Scholar] [CrossRef]
- Varentsova, E.Y.; Mamaev, N.A.; Martirova, M.B. Scots pine and Norway spruce phytopathological condition in forest stands of the Leningrad region. Izvestia Sankt-Peterburgskoj Lesotexnicheskoj Akademii 2023, 244, 131–149, (In Russian with English Abstract). [Google Scholar] [CrossRef]
- Camarero, J.; Gazol, A.; Sangüesa-Barreda, G.; Vergarechea, M.; Sánchez, R.A.; Cattaneo, N.; Vicente-Serrano, S.M. Tree growth is more limited by drought in rear-edge forests most of the times. Forest Ecosystems 2021, 8, 25. [Google Scholar] [CrossRef]
- Boczoń, A.; Hilszczańska, D.; Wrzosek, M.; Szczepkowski, A.; Sierota, Z. Drought in the forest breaks plant–fungi interactions. European Journal of Forest Research 2021, 140, 1301–1321. [Google Scholar] [CrossRef]
- Ryu, M.; Mishra, R.C.; Jeon, J.; Lee, S.K.; Bae, H. Drought-induced susceptibility for Cenangium ferruginosum leads to progression of Cenangium-dieback disease in Pinus koraiensis. Scientific Reports 2018, 8, 16368. [Google Scholar] [CrossRef]
- Garbelotto, M.; Gonthier, P. Biology, epidemiology, and control of Heterobasidion species worldwide. Annual Review of Phytopathology 2013, 51, 39–59. [Google Scholar] [CrossRef]
- Shen, S.; Zhang, X.; Jian, S. The distributional range changes of European Heterobasidion under future climate change. Forests 2024, 15, 1863. [Google Scholar] [CrossRef]
- Jurc, D.; Jurc, M.; Sieber, T.N.; Bojovic, S. Endophytic Cenangium ferruginosum (Ascomycota) as a reservoir for an epidemic of Cenangium dieback in Austrian pine. Phyton – Annales Rei Botanicae (Horn, Austria). 2000, 40, pp. 103–108. Available online: https://www.researchgate.net/profile/Thomas-Sieber-2/publication/279662682_Endophytic_Cenangium_ferruginosum_Ascomycota_as_a_reservoir_for_an_epidemic_of_Cenangium_dieback_in_Austrian_pine/links/560d5abf08aeed9d13751f5a/Endophytic-Cenangium-ferruginosum-Ascomycota-as-a-reservoir-for-an-epidemic-of-Cenangium-dieback-in-Austrian-pine.pdf (accessed on 17 September 2025).
- Storozhenko, V.G. The structure of tree stands and wood-destroying fungi of native pine biogeocoenoses of the Russian plain. Siberian Forest Journal 2015, 4, 30–39, (In Russian with English Abstract). [Google Scholar] [CrossRef]
- BOLD – The Barcode of Life Data Systems. Available online: https://boldsystems.org (accessed on 9 June 2025).
- Mycobank - Fungal Databases, Nomenclature & Species Banks [online]. 6 August 2025. Available online: http://www.mycobank.org.
- López, S.; Theelen, B.; Manserra, S.; Issak, T.Y.; Rytioja, J.; Mäkelä, M.R.; de Vries, R.P. Functional diversity in Dichomitus squalens monokaryons. IMA Fungus 2017, 8, 17–25. [Google Scholar] [CrossRef]
- Ageev, D.V.; Bulkonova, T.M. Dichomitus squalens - Mushrooms of Siberia. 2025. Available online: https://mycology.su/dichomitus-squalens.html (accessed on 9 June 2025).
- Hallenberg, N. A taxonomic analysis of the Sistotrema brinkmannii complex (Corticiaceae, Basidiomycetes). Mycotaxon 1984, 21, 389–411. [Google Scholar] [CrossRef]
- Nagy, L.G.; Riley, R.; Tritt, A.; Adam, C.; Daum, C.; Floudas, D.; Sun, H.; Yadav, J.S.; Pangilinan, J.; Larsson, K.-H.; Matsuura, K.; Barry, K.; Labutti, K.; Kuo, R.; Ohm, R.A.; Bhattacharya, S.S.; Shirouzu, T.; Yoshinaga, Y.; Martin, F.M.; Grigoriev, I.V.; Hibbett, D.S. Comparative genomics of early-diverging mushroom-forming fungi provides insights into the origins of lignocellulose decay capabilities. Molecular Biology and Evolution 2016, 33, 959–970. [Google Scholar] [CrossRef] [PubMed]
- Zmitrovich, I. Genus Athelia Pers. in Russia. Turczaninowia 2004, 7(4), 22–46. (In Russian) [Google Scholar]
- Koukol, O. A new species of Infundichalara from pine litter. Mycotaxon 2012, 120, 343–352. [Google Scholar] [CrossRef]
- Cho, Y.; Kim, J.S.; Dai, Y.; Gafforov, Y.; Lim, Y.W. Taxonomic evaluation of Xylodon (Hymenochaetales, Basidiomycota) in Korea and sequence verification of the corresponding species in GenBank. PeerJ 2021, 9, e12625. [Google Scholar] [CrossRef]
- Spirin, V.; Volobuev, S.; Viner, I.; Miettinen, O. On Sistotremastrum and similar-looking taxa (Trechisporales, Basidiomycota). Mycological Progress 2021, 20, 453–476. [Google Scholar] [CrossRef]
- Lincoff, G.; Mitchel, D. Toxic and hallucinogenic mushroom poisoning; Van Nostrand Reinhold Publishing: New York, 1977; p. 267 p. ISBN -10 0442245807. [Google Scholar]
- Kirk, P.M.; Paul, F.; Cannon, D.; Minter, W.; Stalpers, J.A. Dictionary of the Fungi. In CAB International Publishing; 2008; ISBN -10 0851998267. [Google Scholar]
- Biek, D. The mushrooms of northern California; Spore Prints: Redding, CA, 1984; ISBN 0-9612020-0-9. ISSN OCLC 10870632. [Google Scholar]
- Zmitrovich, I.; Volobuev, S.; Dudka, V.; Zhukova, E.; Sidelnikova, M.; Bondartseva, M. Ganoderma applanatum (Polyporales, Basidiomycota) at the Saint Petersburg area. Mycology and Phytopathology (Mikologiya i Fitopatologiya) 2019, 53, 354–362. [Google Scholar] [CrossRef]
- Begum, H.A.; Ahmad, W.; Rafiq, N.; Ali, H.; Hussain, S.; Ali, B.; Ullah, I.; Ahmed, I.; Khan, B.; Khan, A. Exploring the pharmacological potential of Trametes hirsuta (White Rot Fungi): Analgesic, anti-Inflammatory, antispasmodic and antimicrobial activities. Pure and Applied Biology 2023, 12, 1183–1193. [Google Scholar] [CrossRef]
- Trudell, S.; Ammirati, J. Mushrooms of the Pacific Northwest. In Timber Press Field Guides; Timber Press: Portland, OR, 2009; p. 264. [Google Scholar] [CrossRef]
- GBIF Secretariat. GBIF Backbone Taxonomy. 2023. [Google Scholar] [CrossRef]
- Liu, S.L.; He, S.H.; Wang, X.W.; May, T.W.; He, G.; Chen, S.L.; Zhou, L.W. Trechisporales emended with a segregation of Sistotremastrales ord. nov. (Basidiomycota). Mycosphere 2022, 13, 862–954. [Google Scholar] [CrossRef]
- El-Morsy, E.M.; Nour El-Dein, M.M.; El-Didamoney, S.M.M. Mucor racemosus as a biosorbent of metal ions from polluted water in Northern Delta of Egypt. Mycosphere 2013, 4, 1118–1131. [Google Scholar] [CrossRef]
- Lübbehüsen, T.; Nielsen, J.; McIntyre, M. Morphology and physiology of the dimorphic fungus Mucor circinelloides (syn. M. racemosus) during anaerobic growth. Mycological Research 2003, 107, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Bogar, B.; Szakacs, G.; Pandey, A.; Abdulhameed, S.; Linden, J.; Tengerdy, R. Production of phytase by Mucor racemosus in solid-state fermentation. Biotechnology Progress 2003, 19, 312–319. [Google Scholar] [CrossRef]
- Caldcluvia paniculosa (2022) CABI Compendium; CABI International Publishing, 17 09 2025. [CrossRef]
- Hietala, A.; Agan, A.; Nagy, N.E.; Børja, I.; Timmermann, V.; Drenkhan, R.; Solheim, H. The native Hymenoscyphus albidus and the invasive Hymenoscyphus fraxineus are similar in their necrotrophic growth phase in Ash leaves. Frontiers in Microbiology 2022, 13, 892051. [Google Scholar] [CrossRef]
- Elfstrand, M.; Chen, J.; Cleary, M.; Halecker, S.; Ihrmark, K.; Karlsson, M.; Davydenko, K.; Stenlid, J.; Stadler, M.; Durling, M.B. Comparative analyses of the Hymenoscyphus fraxineus and Hymenoscyphus albidus genomes reveals potentially adaptive differences in secondary metabolite and transposable element repertoires. BMC Genomics 2021, 22, 503. [Google Scholar] [CrossRef] [PubMed]
- Safonov, M.A.; Safonova, T.I. Wood-destroying fungi observed on tree and shrub introduced species in the steppe zone of the Southern Urals (Orenburg oblast). Journal of Agriculture and Environment 2023, 11, 1–5. [Google Scholar] [CrossRef]
- Safonov, M.A.; Safonova, T.I. Xylotrophic fungi on major forest-forming species in Orenburg oblast. International Research Journal 2023b, 4, 1–6, (In Russian with English Abstract). [Google Scholar] [CrossRef]
- Kwaśna, H.; Mazur, A.; Łabędzki, A.; Kuźmiński, R.; Łakomy, P. Communities of fungi in decomposed wood of oak and pine. Forest Research Papers 2016, 77, 261–275. [Google Scholar] [CrossRef]
- Stokland, J.; Larsson, K-H. Legacies from natural forest dynamics: different effects of forest management on wood-inhabiting fungi in pine and spruce forests. Forest Ecology and Management 2011, 261, 707–1721. [Google Scholar] [CrossRef]
- Neklyaev, S.E.; Larina, G.E.; Seraya, L.G. Features of active mycogenic xylolysis on Scots pine in the zone of coniferous-deciduous forests. Izvestiya Sankt-Peterburgskoj Lesotexnicheskoj Akademii 2023, 244, 164–183, (in Russian with English Abstract). [Google Scholar] [CrossRef]
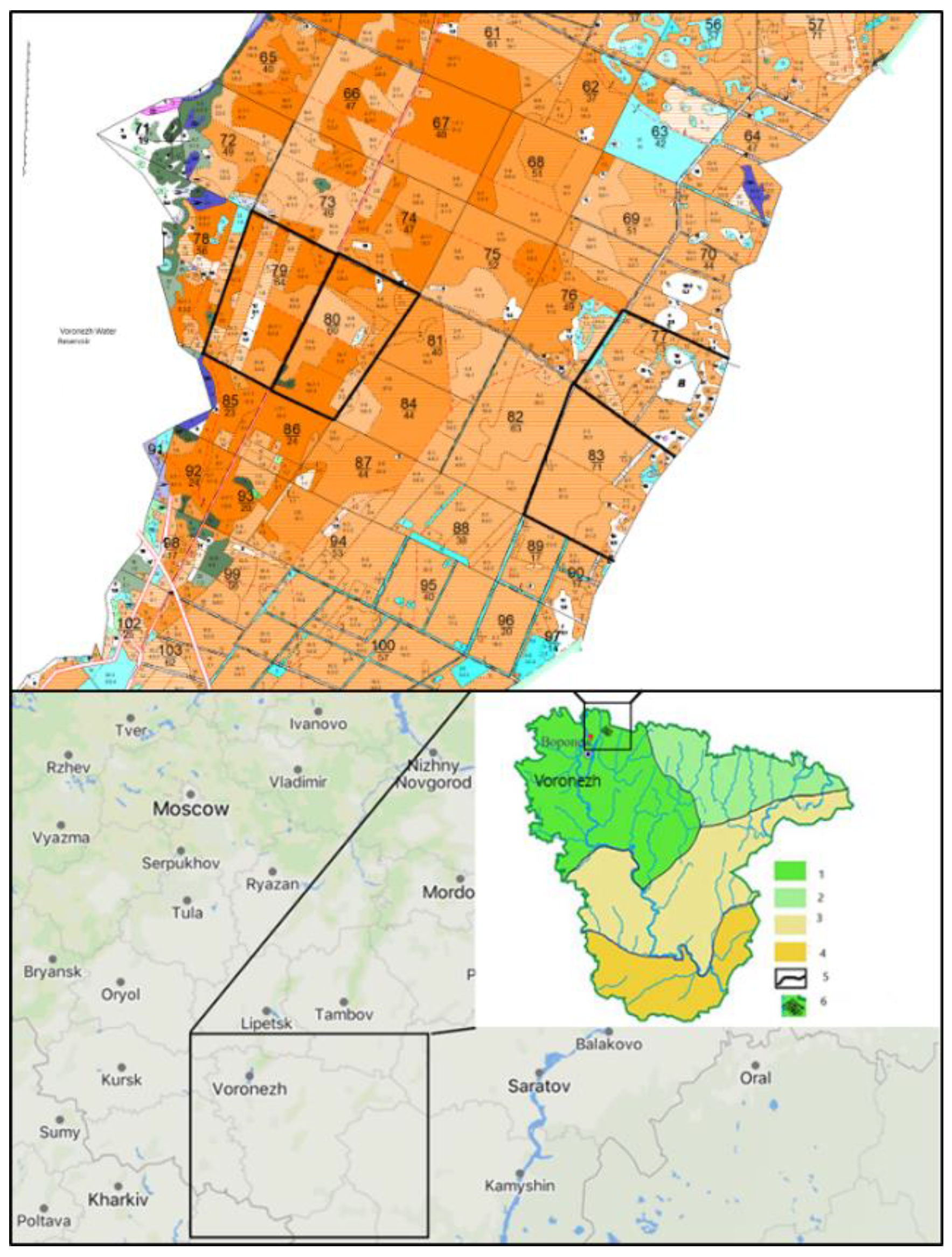
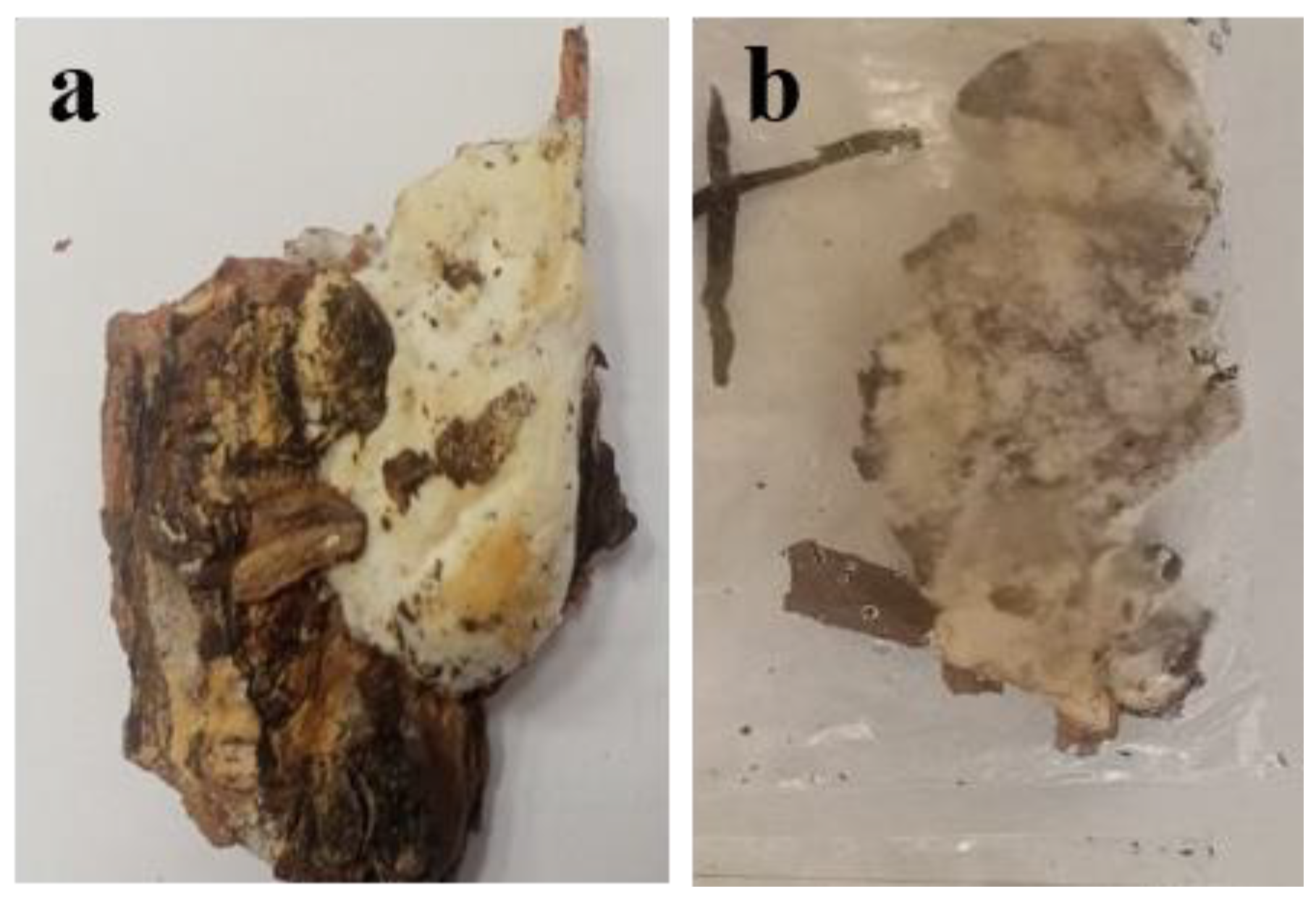
| Permanent sample plot | Species composition | Age, years | Density | Type of vegetation |
|---|---|---|---|---|
| 77 | Only Scots pine trees | 70, 78 | 0.4-0.8 | Fresh pine forest |
| 79 | 40, 48, 55, 66 | 0.6-0.8 | Fresh subor | |
| 80 | Felling after forest fires. At the time of felling, the forest was approximately 120 years old | At the moment of felling − 0.7 | Fresh subor | |
| 83 | 83 | 0.6-0.8 | Fresh subor | |
| “Northern Forest” (NF) | Almost all Scots pine trees, including occasional birch, aspen, and maple trees | planting, average age 50 years | 0.6-0.8 | Fresh pine forest |
| Sample | Identified species | Environmental group | Mycelium collection site | NCBI GenBank accession number |
|---|---|---|---|---|
| 79/5 root | Dichomitus squalens (P. Karst.) D.A. Reid | xylosaprotroph | dead wood of pine trees | PX132320 |
| 79/8 root | PX132322 | |||
| 79/4 root | PX132312 | |||
| 79/6 root; NF 16 trunk | Sistotrema sp. | xylosaprotroph, forest litter saprotroph | dead wood and plant detritus in the forest litter upper layers | PX132321 |
| 83/3.2 root | рoд Athelia Pers. | xylosaprotroph | dead wood and small woody plant residues | PX132313 |
| NF 4 forest litter | Xenopolyscytalum pinea Crous | forest litter saprotroph | pine tree litter | PX132315 |
| NF 13 old windfallen tree | Xylodon flaviporus (Berk. & M.A. Curtis ex Cooke) Riebesehl & E. Langer | xylosaprotroph | dead wood of pine and deciduous trees | PX218715 |
| NF 18 forest litter | Trechispora sp. | xylosaprotroph | dead wood of pine and deciduous trees | PX132316 |
| NF 20 forest litter | PX132317 | |||
| NF 21 butt of a tree | Sertulicium vernale Spirin &Volobuev | xylosaprotroph | dead wood of pine and deciduous trees | PX218716 |
| NF 22 root of old windfallen tree | Gymnopilus penetrans (Fr.) Murrill | facultative xylosaprotroph | live and dead wood of pine and deciduous species | PX132318 |
| NF 23 butt of a deadtree | Ganoderma applanatum (Pers.) Pat. | facultative xylosaprotroph, parasite | live and dead wood of pine and deciduous trees | PX132319 |
| K3, K7 root | Trametes hirsuta | saprotroph | old stumps, deadwood, dying trunks of deciduous trees | PX132323 |
| NF 3 stem | Trichaptum fuscoviolaceus | saprotroph | livie and dead wood of pine trees | PX132314 |
| Phylum | Ascomycota | Basidiomycota | ||||||
| Class | Leotiomycetes | Agaricomycetes | ||||||
| Order | Helotiales | Agaricales | Atheliales | Canthare-llales | Hymeno-chaetales | Polyporales | Sistotrematales | Trechisporales |
| Family | Hamatocanthoscyphaceae | Strophariaceae | Atheliaceae | Hydnaceae | Schizoporaceae | Polyporaceae | Sistotremastraceae | Hydnodontaceae |
| Species | Xenopolyscytalum pinea Crous | Gymnopilus penetrans (Fr.) Murrill | Athelia Pers. | Sistotrema brinkmannii (Bres.) J. Erikss. | Xylodon flaviporus (Berk. & M.A. Curtis ex Cooke) Riebesehl & E. Langer. | 1. Dichomitus squalens (P. Karst.) D.A. Reid. 2. Ganoderma applanatum (Pers.) Pat. 3. Trametes hirsuta (Wulfen) Lloyd. 4. Trichaptum fuscoviolaceum (Ehrenb.) Ryvarden |
Sertulicium vernale Spirin & Volobuev | Trechispora P. Karst. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





