Submitted:
04 December 2025
Posted:
08 December 2025
You are already at the latest version
Abstract
Snakes are becoming increasingly popular as companion animals in Bulgaria, yet their potential as reservoirs for zoonotic pathogens remains underexplored. This study investigates bacterial carriage in domesticated snakes and evaluates the public health risks associated with their handling. Materials and methods: Bacteriological analysis was performed on 29 snake samples from various species, including pythons, boas, corn snakes, and a venomous species. All individuals were raised in controlled environments and primarily fed frozen rodents. Standard microbiological techniques were employed for bacterial isolation and identification, and the results were confirmed using the Vitek 2 Compact System. Results: Salmonella enterica (group B) was isolated from 8 samples, indicating a notable prevalence of this pathogen among the examined snakes. Additionally, several other bacterial genera were identified, including Pseudomonas spp., Staphylococcus spp., Proteus spp., Enterobacteriaceae spp., Sphingomonas paucimobilis, Enterococcus spp., Bacillus spp., Achromobacter denitrificans, Citrobacter koseri, and Klebsiella pneumoniae. Conclusion: The study highlights the zoonotic risks associated with snake ownership, particularly through direct contact with the reptiles or exposure to contaminated environments. Food sources may act as vectors for microbial contamination. To mitigate the risk of bacterial transmission between snakes and humans, it is crucial to adopt rigorous hygiene practices, conduct regular health monitoring of the snakes, and implement proper protocols for feeding and cleaning.
Keywords:
1. Introduction
2. Materials and Methods
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| RAS | reptile-associated salmonellosis |
References
- Warwick, C.; Arena, P.C.; Steedman, C. Health implications associated with exposure to farmed and wild sea turtles. JRSM short reports 2013, 4, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Mermin, J.; Hutwagner, L.; Vugia, D.; Shallow, S.; Daily, P.; Bender, J. & Emerging Infections Program FoodNet Working Group. Reptiles, amphibians, and human Salmonella infection: a population-based, case-control study. Clinical infectious diseases 2004, 38 Suppl. S3, S253–S261. [Google Scholar]
- Whiley, H.; Gardner, M.G.; Ross, K. A review of Salmonella and squamates (lizards, snakes and amphisbians): implications for public health. Pathogens 2017, 6, 38. [Google Scholar] [CrossRef]
- Sodagari, H.R.; Habib, I.; Shahabi, M.P.; Dybing, N.A.; Wang, P.; Bruce, M. Public health challenges of salmonella and turtles. Veterinary Sciences 2020, 7, 56. [Google Scholar] [CrossRef]
- Lin, W.H.; Tsai, T.S. Comparisons of the oral microbiota from seven species of wild venomous snakes in Taiwan using the high-throughput amplicon sequencing of the full-length 16S rRNA gene. Biology 2023, 12, 1206. [Google Scholar] [CrossRef]
- Pasmans, F.; Martel, A.N.; Jacobson, E.R. Bacterial diseases of reptiles. Infectious diseases and Pathology of reptiles 2020, 705–794. [Google Scholar]
- Pees, M.; Brockmann, M.; Steiner, N.; Marschang, R.E. Salmonella in reptiles: Occurrence, interactions, shedding, and human infection risk. Frontiers in Cell and Developmental Biology 2023, 11, 1251036. [Google Scholar] [CrossRef]
- Meletiadis, A.; Biolatti, C.; Mugetti, D.; Zaccaria, T.; Cipriani, R.; Pitti, M.; Decastelli, L.; Cimino, F.; Dondo, A.; Maurella, C.; Bozzetta, E.; Acutis, P.L. Surveys on Exposure to Reptile-Associated Salmonellosis (RAS) in the Piedmont Region—Italy. Animals 2022, 12, 906. [Google Scholar] [CrossRef] [PubMed]
- JACOBSON ER. Bacterial diseases of reptiles. In Infectious diseases and pathology of reptiles; CRC Press, 2007; pp. 475–540. [Google Scholar]
- Pasmans, F.; Bogaerts, S.; Braeckman, J.; Cunningham, A.A.; Hellebuyck, T.; Griffiths, R.A.; Martel, A. Future of keeping pet reptiles and amphibians: towards integrating animal welfare, human health and environmental sustainability. Veterinary Record 2017, 181, 450–450. [Google Scholar] [CrossRef] [PubMed]
- Tagesu, A. Examination of feces. J. Int. Vet. Sci. Res 2018, 1, 45–50. [Google Scholar]
- Smith, J.; Thompson, R.; Walker, L. Enrichment media for pathogen detection in clinical laboratories. Journal of Microbial Science 2019, 15, 221–229. [Google Scholar]
- Sanders, E.R. Aseptic laboratory techniques: plating methods. Journal of Visualized Experiments (JoVE) 2012, (63), e3064. [Google Scholar]
- Mariam, S.H. Isolation and Characterization of Gram-Negative Bacterial Species from Pasteurized Dairy Products: Potential Risk to Consumer Health. J Food Qual. 2021, 17, 1‒10. [Google Scholar] [CrossRef]
- Boneva-Marutsova, B.; Marutsov, P.; Geisler, M.L.; Zhelev, G. Salmonellosis Outbreak in a Rottweiler Kennel Associated with Raw Meat-Based Diets. Animals 2025, 15, 3196. [Google Scholar] [CrossRef]
- Funke, G.; Funke-Kissling, P. Evaluation of the new VITEK 2 card for identification of clinically relevant gram-negative rods. Journal of clinical microbiology 2004, 42, 4067–4071. [Google Scholar] [CrossRef]
- Krishnankutty, S.P.; Muraleedharan, M.; Perumal, R.C.; Michael, S.; Benny, J.; Balan, B.; Zachariah, A. Next-generation sequencing analysis reveals high bacterial diversity in wild venomous and non-venomous snakes from India. Journal of Venomous Animals and Toxins Including Tropical Diseases 2018, 24, 41. [Google Scholar] [CrossRef] [PubMed]
- Su, H.Y.; Hussain, B.; Hsu, B.M.; Lee, K.H.; Mao, Y.C.; Chiang, L.C.; Chen, J.S. Bacterial community analysis identifies Klebsiella pneumoniae as a native symbiotic bacterium in the newborn Protobothrops mucrosquamatus. BMC microbiology 2023, 23, 213. [Google Scholar] [CrossRef]
- Muslin, C.; Salas-Brito, P.; Coello, D.; Morales-Jadán, D.; Viteri-Dávila, C.; Coral-Almeida, M. Salmonella prevalence and serovar distribution in reptiles: a systematic review and meta-analysis. Gut Pathogens 2025, 17, 52. [Google Scholar] [CrossRef]
- Chen, C.Y.; Chen, W.C.; Chin, S.C.; Lai, Y.H.; Tung, K.C.; Chiou, C.S.; Chang, C.C. Prevalence and antimicrobial susceptibility of salmonellae isolates from reptiles in Taiwan. Journal of Veterinary Diagnostic Investigation 2010, 22, 44–50. [Google Scholar] [CrossRef]
- Marin, C.; Martelli, F.; Rabie, A.; Davies, R. Commercial frozen mice used by owners to feed reptiles are highly externally contaminated with Salmonella Enteritidis PT8. Vector-Borne and Zoonotic Diseases 2018, 18, 453–457. [Google Scholar] [CrossRef] [PubMed]
- Varela, K.; Brown, J.A.; Lipton, B.; Dunn, J.; Stanek, D.; NASPHVCommittee, C.o.n.s.u.l.t.a.n.t.s.; Yager, C.M. A review of zoonotic disease threats to pet owners: a compendium of measures to prevent zoonotic diseases associated with non-traditional pets such as rodents and other small mammals, reptiles, amphibians, backyard poultry, and other selected animals. Vector-borne and zoonotic diseases 2022, 22, 303–360. [Google Scholar] [CrossRef]
- Waltenburg, M.A.; Perez, A.; Salah, Z.; Karp, B.E.; Whichard, J.; Tolar, B.; Nichols, M. Multistate reptile-and amphibian-associated salmonellosis outbreaks in humans, United States, 2009–2018. Zoonoses and Public Health 2022, 69, 925–937. [Google Scholar] [CrossRef]
- Ebani, V.V. Domestic reptiles as source of zoonotic bacteria: A mini review. Asian Pacific Journal of Tropical Medicine 2017, 10, 723–728. [Google Scholar] [CrossRef]
- Dégi, J.; Herman, V.; Radulov, I.; Morariu, F.; Florea, T.; Imre, K. Surveys on pet-reptile-associated multi-drug-resistant Salmonella spp. in the Timișoara metropolitan region—Western Romania. Antibiotics 2023, 12, 1203. [Google Scholar] [CrossRef] [PubMed]
- Hoffbeck, C.; Middleton, D.M.; Nelson, N.J.; Taylor, M.W. 16S rRNA gene-based meta-analysis of the reptile gut microbiota reveals environmental effects, host influences and a limited core microbiota. Molecular Ecology 2023, 32, 6044–6058. [Google Scholar] [CrossRef] [PubMed]
- Shivambu, N.; Shivambu, T.C.; Chimimba, C.T. Zoonotic pathogens associated with pet and feeder murid rodent species: a global systematic review. Vector-Borne and Zoonotic Diseases 2023, 23, 551–560. [Google Scholar] [CrossRef]
- Marques, I.; Pinto, A.R.; Martins, J.J.; Alvura, N.; Telinhos, P.; Mendes, P.; Saavedra, M.J. Assessing Potential Reservoir of Multidrug-Resistant Bacteria in the Oral Microbiota of Captive Burmese and Royal Pythons. Life 2025, 15, 442. [Google Scholar] [CrossRef] [PubMed]
- Artavia-León, A.; Romero-Guerrero, A.; Sancho-Blanco, C.; Rojas, N.; Umaña-Castro, R. Diversity of aerobic bacteria isolated from oral and cloacal cavities from free-living snake species in Costa Rica rainforest. International Scholarly Research Notices 2017, 2017, 8934285. [Google Scholar] [CrossRef]
- Chuang, P.C.; Tsai, Y.H.; Hsu, W.H.; Lin, C.C.; Chiou, C.S.; Hsu, T.C. Oral bacteria and their antibiotic susceptibilities in Taiwanese venomous snakes. Microorganisms 2022, 10, 951. [Google Scholar] [CrossRef]
- Young Yusty, S.; Prescilla-Ledezma, A. From culture to metagenomics: Advances in oral microbiota of venomous snakes. Microbiology Research 2025, 16, 233. [Google Scholar] [CrossRef]
- Dehghani, R.; Sharif, M.R.; Moniri, R.; Sharif, A.; Haddad Kashani, H. The identification of bacterial flora in oral cavity of snakes. Comparative Clinical Pathology 2016, 25, 279–283. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Salmonella and reptiles and amphibians. Available online: https://www.cdc.gov.
- European Food Safety Authority. Public health risks associated with Salmonella in reptiles. EFSA Journal 2019, 17, e05656. [Google Scholar]
- Lamichhane, B.; Mawad, A.M.; Saleh, M.; Kelley, W.G.; Harrington, P.J.; Lovestad, C.W.; Helmy, Y.A. Salmonellosis: an overview of epidemiology, pathogenesis, and innovative approaches to mitigate the antimicrobial resistant infections. Antibiotics 2024, 13, 76. [Google Scholar] [CrossRef] [PubMed]
| Species | Family | Common name | N (29) |
|---|---|---|---|
| Python regius | Pythonidae | Ball python | 3 |
| Python curtus | Pythonidae | Sumatran short-tailed python | 1 |
| Python buvittatus | Pythonidae | Burmese python | 2 |
| Morelia spilota | Pythonidae | Carpet python | 3 |
| Acrantophis dumerili | Boidae | Dumerili’s boa | 2 |
| Boa constrictor | Boidae | Common boa | 1 |
| Boaedon fuliginosus | Lamprophilidae | African house snake | 1 |
| Pantherophis guttatus | Colubridae | Corn snake | 15 |
| Bothrops insularis | Viperidae | Golden lancehead | 1 |
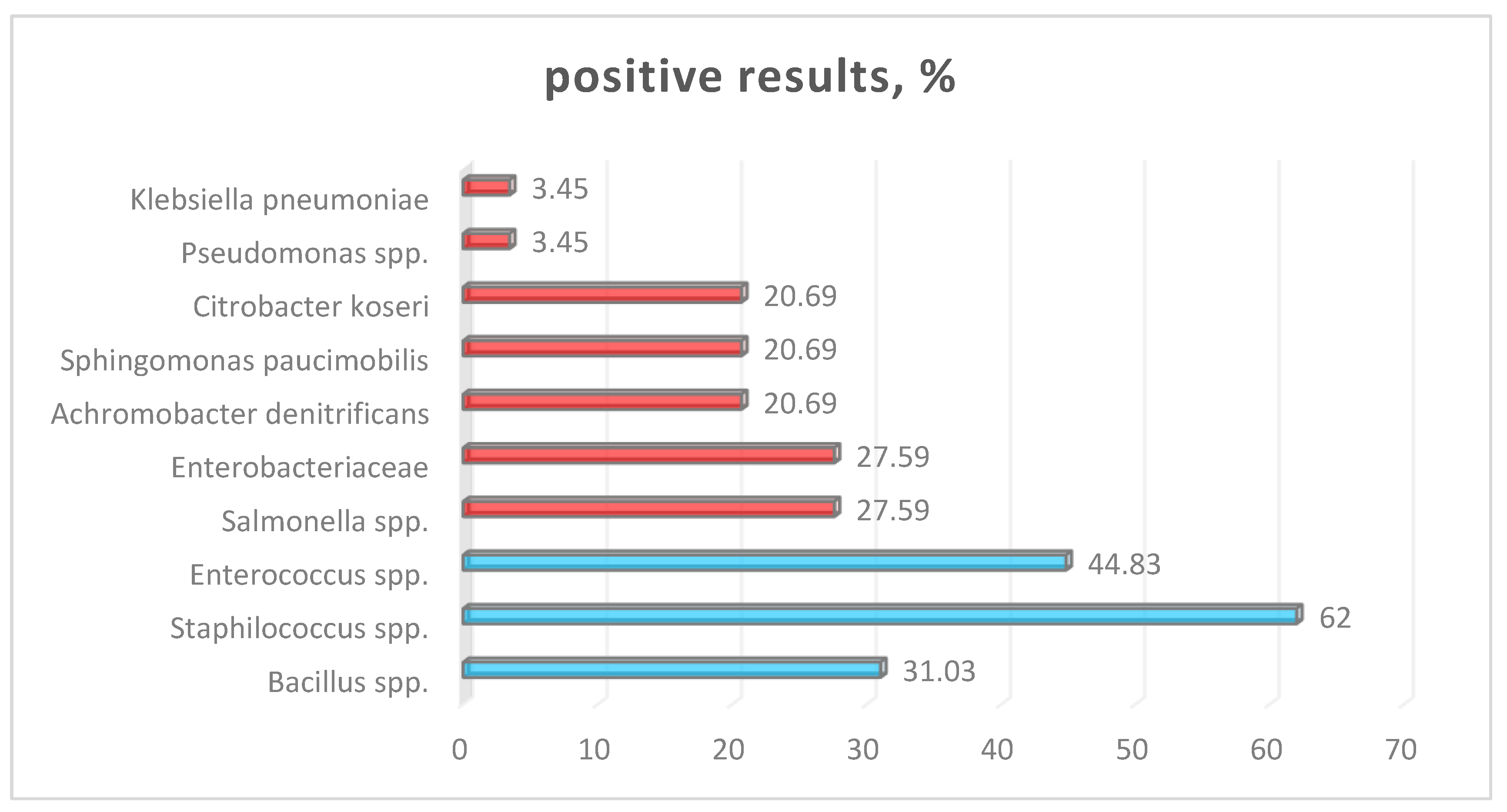
| Species | Samples (n) | Positive (n) | % Positive | SE | 95% CI (lower–upper) |
|---|---|---|---|---|---|
| Salmonella enterica | 29 | 8 | 27.59% | 0.084 | 12.8% – 42.3% |
| Staphylococcus spp. | 29 | 18 | 62.07% | 0.091 | 44.1% – 80.1% |
| Enterococcus spp. | 29 | 13 | 44.83% | 0.092 | 26.6% – 63.0% |
| Bacillus spp | 29 | 9 | 31.03% | 0.086 | 15.1% – 46.9% |
| Enterobacteriaceae spp. | 29 | 8 | 27.59% | 0.084 | 12.8% – 42.3% |
| A. denitrificans | 29 | 6 | 20.69% | 0.076 | 6.7% – 34.6% |
| S. paucimobilis | 29 | 6 | 20.69% | 0.076 | 6.7% – 34.6% |
| C. koseri | 29 | 6 | 20.69% | 0.076 | 6.7% – 34.6% |
| Pseudomonas spp. | 29 | 8 | 27.59% | 0.084 | 12.8% – 42.3% |
| Klebsiella pneumoniae | 29 | 6 | 20.69% | 0.076 | 6.7% – 34.6% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




