Submitted:
03 December 2025
Posted:
04 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Verticillium Wilt Disease
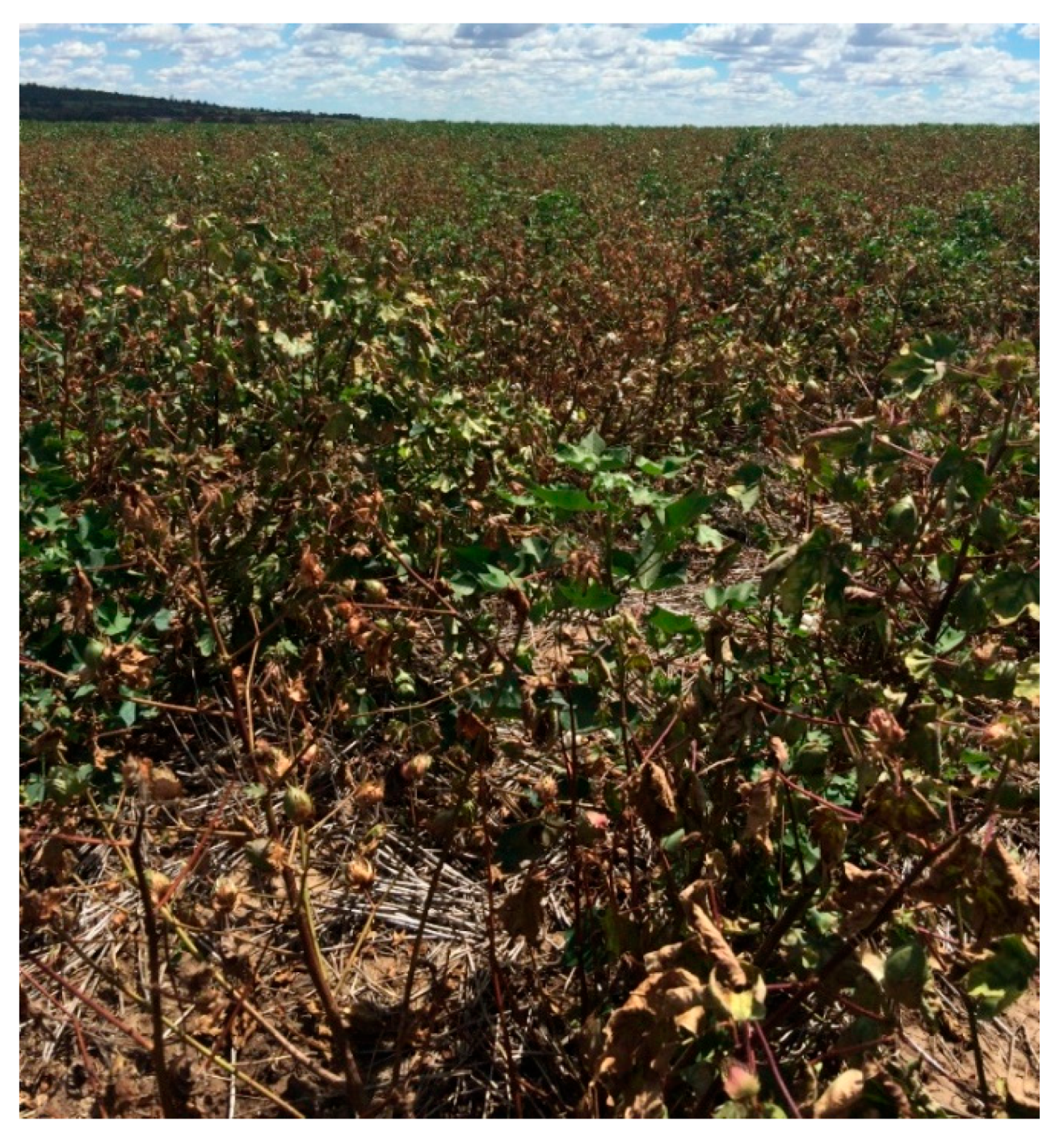
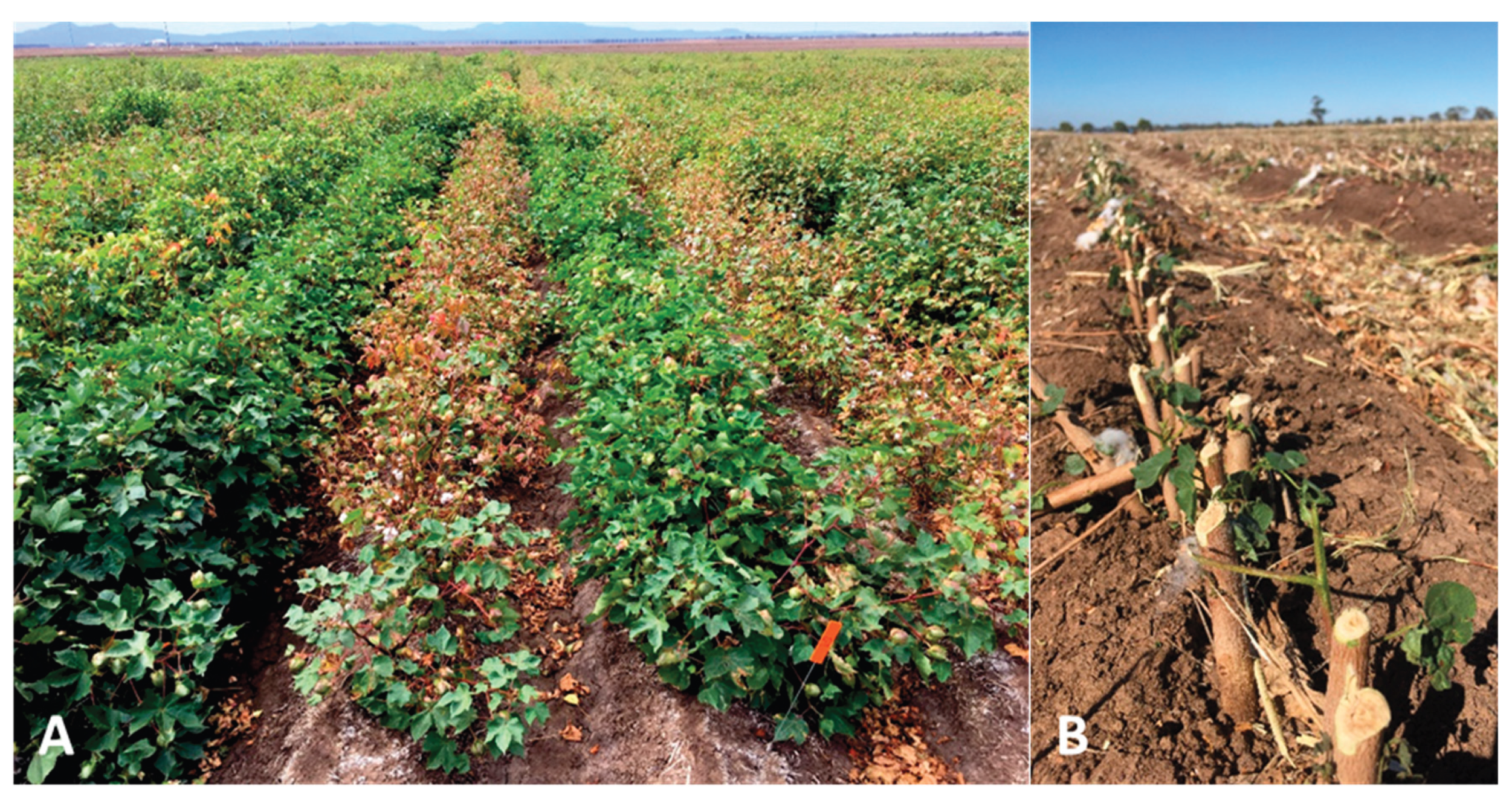
2.1. Symptoms
2.2. Economic Impacts
2.3. Ecological Impacts
3. Verticillium Wilt Pathogens
3.1. Taxonomy and Identification
3.2. Population Biology
3.2.1. VC Groups
3.2.2. Mating Types
3.2.3. Pathogenicity
4. Detection and Its Advances
4.1. Culture-Based Techniques
4.2. Culture-Independent Techniques
5. Control Strategies
5.1. Chemical
5.2. Cultural
5.3. Biological
5.4. Induced Resistance
5.5. Host Resistance
5.5.1. Mechanisms of Resistance
5.5.2. Sources and Inheritance of Resistance
5.5.3. Genetics of Resistance and Mapping of Quantitative Trait Loci
5.5.4. Breeding for Resistance
5.5.5. Screening and Selection Methods
5.6. Integrated Strategies
6. Future Perspectives
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wendel, J.F.; Cronn, R.C. Polyploidy and the evolutionary history of cotton. Advances in Agronomy. 2002, 87, 139–186. [Google Scholar]
- Naranjo, S.E. Impacts of Bt transgenic cotton on integrated pest management. Journal of Agricultural and Food Chemistry 2011, 59, 5842–5851. [Google Scholar] [CrossRef]
- Rocha-Munive, M.G.; Soberón, M.; Castañeda, S.; Niaves, E.; Scheinvar, E.; Eguiarte, L.E.; Mota-Sánchez, D.; Rosales-Robles, E.; Nava-Camberos, U.; Martínez-Carrillo, J.L. Evaluation of the impact of genetically modified cotton after 20 years of cultivation in Mexico. Frontiers in Bioengineering and Biotechnology 2018, 6, 82. [Google Scholar] [CrossRef]
- Kirkpatrick, T.L.; Rockroth, C. Compendium of Cotton Diseases; American Phytopathological Society (APS Press). 2001.
- Kirkby, K.; Lonergan, P.; Allen, S. Three decades of cotton disease surveys in NSW, Australia. Crop and Pasture Science 2013, 64, 774–779. [Google Scholar] [CrossRef]
- Zhengjun, X.; Achar, P.; Benkang, G. Vegetative compatibility groupings of Verticillium dahliae from cotton in mainland China. European Journal of Plant Pathology 1998, 104, 871–876. [Google Scholar] [CrossRef]
- Göre, M.E.; Erdoğan, O.; Caner, Ö.K.; Aydın, M.H.; Berk, S. VCG diversity and virulence of Verticillium dahliae from commercially available cotton seed lots in Turkey. European Journal of Plant Pathology 2014, 140, 689–699. [Google Scholar] [CrossRef]
- Land, C.; Lawrence, K.; Burmester, C.; Meyer, B. Cultivar, irrigation, and soil contribution to the enhancement of Verticillium wilt disease in cotton. Crop Protection 2017, 96, 1–6. [Google Scholar] [CrossRef]
- Palmateer, A.; McLean, K.; Morgan-Jones, G.; Van Santen, E. Frequency and diversity of fungi colonizing tissues of upland cotton. Mycopathologia 2004, 157, 303–316. [Google Scholar] [CrossRef] [PubMed]
- Inderbitzin, P.; Bostock, R.M.; Davis, R.M.; Usami, T.; Platt, H.W.; Subbarao, K.V. Phylogenetics and taxonomy of the fungal vascular wilt pathogen Verticillium, with the descriptions of five new species. PloS one 2011, 6, e28341. [Google Scholar] [CrossRef]
- Göre, M.E.; Erdoğan, O.; Altin, N.; Aydın, M.; Caner, Ö.; Filizer, F.; Büyükdöğerlioğlu, A. Seed transmission of Verticillium wilt of cotton. Phytoparasitica 2011, 39, 285–292. [Google Scholar] [CrossRef]
- Smith, L.J.; Scheikowski, L.; Kafle, D.; Le, D.; Gregson, A.; Green, T.; Holman, S. Regional Disease Update In Proceedings of the CSD 2018-19 Variety Trial Results, Wee Waa, NSW Australia, 2019; pp. 100-103. [Google Scholar]
- Wang, Y.; Liang, C.; Wu, S.; Zhang, X.; Tang, J.; Jian, G.; Jiao, G.; Li, F.; Chu, C. Significant improvement of cotton Verticillium wilt resistance by manipulating the expression of Gastrodia antifungal proteins. Molecular plant 2016, 9, 1436–1439. [Google Scholar] [CrossRef] [PubMed]
- Erdogan, O.; Sezener, V.; Ozbek, N.; Bozbek, T.; Yavas, I.; Unay, A. The effects of verticillium wilt (Verticillium dahliae Kleb.) on cotton yield and fiber quality. Asian Journal of Plant Science 2006, 5, 867–870. [Google Scholar] [CrossRef]
- Lawrence, K.; Hagan, A.; Norton, R.; Hu, J.; Faske, T.; Hutmacher, R.; Muller, J.; Small, I.; Grabau, Z.; Kemerait, R. Cotton disease loss estimate committee report, 2015. In Proceedings of the Proceedings of the 2016 Beltwide Cotton Conference, 2016; pp. 113-115.
- Melero--Vara, J.; Blanco--López, M.; Bejarano--Alcázar, J.; Jiménez--Díaz, R. Control of Verticillium wilt of cotton by means of soil solarization and tolerant cultivars in southern Spain. Plant Pathology 1995, 44, 250–260. [Google Scholar] [CrossRef]
- Göre, M.E.; Caner, Ö.K.; Altın, N.; Aydın, M.H.; Erdoğan, O.; Filizer, F.; Büyükdöğerlioğlu, A. Evaluation of cotton cultivars for resistance to pathotypes of Verticillium dahliae. Crop Protection 2009, 28, 215–219. [Google Scholar] [CrossRef]
- Wheeler, T.A.; Woodward, J.E. Field assessment of commercial cotton cultivars for Verticillium wilt resistance and yield. Crop Protection 2016, 88, 1–6. [Google Scholar] [CrossRef]
- Wheeler, T.; Woodward, J.; Mullinix, B., Jr. Effect of seeding rate on Verticillium wilt incidence, yield, and value for three cotton cultivars. Journal of Cotton Science 2010, 14, 173–180. [Google Scholar]
- Wheeler, T.; Bordovsky, J.; Keeling, J.; Mullinix Jr, B. Effects of crop rotation, cultivar, and irrigation and nitrogen rate on Verticillium wilt in cotton. Plant Disease 2012, 96, 985–989. [Google Scholar] [CrossRef]
- DeVay, J.; Weir, B.; Wakeman, R.; Stapleton, J. Effects of Verticillium dahliae infection of cotton plants (Gossypium hirsutum) on potassium levels in leaf petioles. Plant Disease 1997, 81, 1089–1092. [Google Scholar] [CrossRef]
- Wei, F.; Fan, R.; Dong, H.; Shang, W.; Xu, X.; Zhu, H.; Yang, J.; Hu, X. Threshold microsclerotial inoculum for cotton Verticillium wilt determined through wet-sieving and real-time quantitative PCR. Phytopathology 2015, 105, 220–229. [Google Scholar] [CrossRef]
- Bejarano-Alcázar, J.; Melero-Vara, J.; Blanco-López, M.; Jimenez-Diaz, R. Influence of inoculum density of defoliating and nondefoliating pathotypes of Verticillium dahliae on epidemics of Verticillium wilt of cotton in southern Spain. Phytopathology 1995, 85, 1474–1481. [Google Scholar] [CrossRef]
- Pullman, G.; DeVay, J. Epidemiology of Verticillium wilt of cotton: a relationship between inoculum density and disease progression. Phytopathology 1982, 72, 549–554. [Google Scholar] [CrossRef]
- Le, D.P.; Nguyen, C.P.; Kafle, D.; Scheikowski, L.; Montgomery, J.; Lambeth, E.; Thomas, A.; O’Keeffe, K.; Shakeshaft, B.; Young, A. Surveillance, Diversity and Vegetative Compatibility Groups of Fusarium oxysporum f. sp. vasinfectum Collected in Cotton Fields in Australia (2017 to 2022). Pathogens 2022, 11, 1537. [Google Scholar] [CrossRef] [PubMed]
- Le, D.P.; Gregson, A.; Madden, A.; Jackson, R. Occurrence of Eutypella species on cotton (Gossypium hirsutum L.) in New South Wales, Australia. Journal of Plant Pathology 2023, 105, 517–525. [Google Scholar] [CrossRef]
- Le, D.P.; Nguyen, C.P.T.; Manjarrez, M.; Giblot-Ducray, D. A diagnostic guide for Berkeleyomyces rouxiae causing black root rot of cotton. Plant Health Progress 2024. [Google Scholar] [CrossRef]
- Le, D.P.; Gregson, A.; Tran, T.T.; Jackson, R. Co-occurrence of defoliating and non-defoliating pathotypes of Verticillium dahliae in field-grown cotton plants in New South Wales, Australia. Plants 2020, 9, 750. [Google Scholar] [CrossRef] [PubMed]
- Klosterman, S.J.; Atallah, Z.K.; Vallad, G.E.; Subbarao, K.V. Diversity, pathogenicity, and management of Verticillium species. Annual Review of Phytopathology 2009, 47, 39–62. [Google Scholar] [CrossRef]
- Allen, S.J.; McNamara, G. Cotton Pathology 2007/2008. In 2008.
- Li, X.; Zhang, Y.n.; Ding, C.; Xu, W.; Wang, X. Temporal patterns of cotton Fusarium and Verticillium wilt in Jiangsu coastal areas of China. Scientific reports 2017, 7, 12581. [Google Scholar] [CrossRef]
- Holman, S.; Kirkby, K.; Smith, L.; Hartnett, H. Vert update: The latest in Vert research. Factsheet. CottonInfo Australia, Narrabri, NSW Australia, 2016, 5 pp.
- CSD. Seeds for Thought. Verticillium wilt update, Vol 10, Wee Waa, NSW Australia, 2011, 3 pp.
- Barker, J.; Coggan, L. Qualitative report on the 2023-24 cotton season: a survey of consultants. Crop Consultants Australia, 2024; pp. 1-36.
- Hawke, M.A. Studies on the Survival of Microsclerotia of Verticillium dahliae. PhD thesis, The University of Western Ontario, 1994, pp. 1-162.
- Klimes, A.; Amyotte, S.G.; Grant, S.; Kang, S.; Dobinson, K.F. Microsclerotia development in Verticillium dahliae: regulation and differential expression of the hydrophobin gene VDH1. Fungal Genetics and Biology 2008, 45, 1525–1532. [Google Scholar] [CrossRef]
- Wilhelm, S. Longevity of the Verticillium wilt fungus in the laboratory and field. Phytopathology 1955, 45, 180–181. [Google Scholar]
- Frederick, Z.; Cummings, T.; Johnson, D. Susceptibility of weedy hosts from Pacific Northwest potato production systems to crop-aggressive isolates of Verticillium dahliae. Plant Disease 2017, 101, 1500–1506. [Google Scholar] [CrossRef]
- Vallad, G.E.; Bhat, R.G.; Koike, S.T.; Ryder, E.J.; Subbarao, K.V. Weedborne reservoirs and seed transmission of Verticillium dahliae in lettuce. Plant Disease 2005, 89, 317–324. [Google Scholar] [CrossRef]
- Yildiz, A.; Doğan, M.N.; Boz, Ö.; Benlioğlu, S. Weed hosts of Verticillium dahliae in cotton fields in Turkey and characterization of V. dahliae isolates from weeds. Phytoparasitica 2009, 37, 171–178. [Google Scholar] [CrossRef]
- El-Zik, K.M. Integrated control of Verticillium wilt of cotton. Plant Disease 1985, 69, 1025–1032. [Google Scholar] [CrossRef]
- Nees von Esenbeck, C.G. Das system der pilze und schwämme. Ein versuch; In der Stahelschen buchhandlung, 1816.
- Rasoul, Z.; Walter, G. The type species of Verticillium is not congeneric with the plant-pathogenic species placed in Verticillium and it is not the anamorph of ‘Nectria’inventa. Mycological Research 2004, 108, 576–582. [Google Scholar]
- Qin, Q.-M.; Vallad, G.E.; Wu, B.M.; Subbarao, K.V. Phylogenetic analyses of phytopathogenic isolates of Verticillium spp. Phytopathology 2006, 96, 582–592. [Google Scholar] [CrossRef]
- Short, D.P.; Gurung, S.; Hu, X.; Inderbitzin, P.; Subbarao, K.V. Maintenance of sex-related genes and the co-occurrence of both mating types in Verticillium dahliae. PLoS One 2014, 9, e112145. [Google Scholar] [CrossRef]
- Usami, T.; Itoh, M.; Amemiya, Y. Asexual fungus Verticillium dahliae is potentially heterothallic. Journal of General Plant Pathology 2009, 75, 422–427. [Google Scholar] [CrossRef]
- Usami, T.; Itoh, M.; Amemiya, Y. Mating type gene MAT1-2-1 is common among Japanese isolates of Verticillium dahliae. Physiological and Molecular Plant Pathology 2008, 73, 133–137. [Google Scholar] [CrossRef]
- Collado-Romero, M.; Mercado-Blanco, J.; Olivares-García, C.; Jiménez-Díaz, R.M. Phylogenetic analysis of Verticillium dahliae vegetative compatibility groups. Phytopathology 2008, 98, 1019–1028. [Google Scholar] [CrossRef]
- Jiménez-Gasco, M.d.M.; Malcolm, G.M.; Berbegal, M.; Armengol, J.; Jiménez-Díaz, R.M. Complex molecular relationship between vegetative compatibility groups (VCGs) in Verticillium dahliae: VCGs do not always align with clonal lineages. Phytopathology 2014, 104, 650–659. [Google Scholar] [CrossRef] [PubMed]
- Milgroom, M.G.; del Mar Jiménez-Gasco, M.; Olivares-García, C.; Jiménez-Díaz, R.M. Clonal expansion and migration of a highly virulent, defoliating lineage of Verticillium dahliae. Phytopathology 2016, 106, 1038–1046. [Google Scholar] [CrossRef]
- Collado-Romero, M.; Mercado-Blanco, J.; Olivares-García, C.; Valverde-Corredor, A.; Jiménez-Díaz, R.M. Molecular variability within and among Verticillium dahliae vegetative compatibility groups determined by fluorescent amplified fragment length polymorphism and polymerase chain reaction markers. Phytopathology 2006, 96, 485–495. [Google Scholar] [CrossRef]
- Pegg, G.F.; Brady, B.L. Verticillium wilts; CABI Publishing: Wallingford, 2002. [Google Scholar]
- Bhat, R.; Smith, R.; Koike, S.; Wu, B.; Subbarao, K. Characterization of Verticillium dahliae isolates and wilt epidemics of pepper. Plant Disease 2003, 87, 789–797. [Google Scholar] [CrossRef]
- Joaquim, T.R.; Rowe, R. Vegetative compatibility and virulence of strains of Verticillium dahliae from soil and potato plants. Phytopathology 1991, 81, 552–558. [Google Scholar] [CrossRef]
- Martin, F.N. Mitochondrial haplotype analysis as a tool for differentiating isolates of Verticillium dahliae. Phytopathology 2010, 100, 1231–1239. [Google Scholar] [CrossRef]
- Chapman, T.A.; Chambers, G.A.; Kirkby, K.; Jiménez-Díaz, R.M. First report of the presence of Verticillium dahliae VCG1A in Australia. Australasian Plant Disease Notes 2016, 11, 13. [Google Scholar] [CrossRef]
- Korolev, N.; Pérez-Artés, E.; Bejarano-Alcázar, J.; Rodríguez-Jurado, D.; Katan, J.; Katan, T.; Jiménez-Díaz, R.M. Comparative study of genetic diversity and pathogenicity among populations of Verticillium dahliae from cotton in Spain and Israel. European Journal of Plant Pathology 2001, 107, 443–456. [Google Scholar] [CrossRef]
- Elena, K. Genetic relationships among Verticillium dahliae isolates from cotton in Greece based on vegetative compatibility. European Journal of Plant Pathology 1999, 105, 609–616. [Google Scholar] [CrossRef]
- ElSharawy, A.A.; Yang, G.; Hu, X.; Yang, J. Genetic relationships between virulence, vegetative compatibility and ISSR marker of Verticillium dahliae isolated from cotton. Archives of Phytopathology and Plant Protection 2015, 48, 646–663. [Google Scholar] [CrossRef]
- Korolev, N.; Katan, J.; Katan, T. Vegetative compatibility groups of Verticillium dahliae in Israel: their distribution and association with pathogenicity. Phytopathology 2000, 90, 529–536. [Google Scholar] [CrossRef]
- Korolev, N.; Pérez-Artés, E.; Mercado-Blanco, J.; Bejarano-Alcázar, J.; Rodríguez-Jurado, D.; Jiménez-Díaz, R.M.; Katan, T.; Katan, J. Vegetative compatibility of cotton-defoliating Verticillium dahliae in Israel and its pathogenicity to various crop plants. European Journal of Plant Pathology 2008, 122, 603–617. [Google Scholar] [CrossRef]
- Dervis, S.; Kurt, S.; Soylu, S.; Erten, L.; Mine Soylu, E.; Yıldız, M.; Tok, F.M. Vegetative compatibility groups of Verticillium dahliae from cotton in the southeastern Anatolia region of Turkey. Phytoparasitica 2008, 36, 74–83. [Google Scholar] [CrossRef]
- Göre, M.E. Vegetative compatibility and pathogenicity of Verticillium dahliae isolates from the Aegean Region of Turkey. Phytoparasitica 2007, 35, 222–231. [Google Scholar] [CrossRef]
- Rafiei, V.; Banihashemi, Z.; Jiménez--Díaz, R.; Navas--Cortés, J.; Landa, B.; Jiménez--Gasco, M.; Turgeon, B.; Milgroom, M. Comparison of genotyping by sequencing and microsatellite markers for unravelling population structure in the clonal fungus Verticillium dahliae. Plant Pathology 2018, 67, 76–86. [Google Scholar] [CrossRef]
- Rafiei, V.; Banihashemi, Z.; Bautista-Jalon, L.S.; del Mar Jiménez-Gasco, M.; Turgeon, B.G.; Milgroom, M.G. Population genetics of Verticillium dahliae in Iran based on microsatellite and single nucleotide polymorphism markers. Phytopathology 2018, 108, 780–788. [Google Scholar] [CrossRef] [PubMed]
- Arseneault, T.; Lafond-Lapalme, J.; Gonzalez, C.E.Q.; Jordan, K.S.; Yevtushenko, D.; Barrett, R.; Van der Heyden, H.; Chen, D.; Tenuta, M.; Mimee, B. A genome resource for 192 Verticillium dahliae isolates infecting potatoes across Canada. PhytoFrontiers™ 2023, 3, 753–757. [Google Scholar] [CrossRef]
- Gardiner, D.M.; Smith, L.J.; Rusu, A.; Aitken, E.A. The genomes of two Australian isolates of Verticillium dahliae recovered from cotton fields. Australasian Plant Pathology 2024, 53, 435–441. [Google Scholar] [CrossRef]
- Ninkuu, V.; Liu, Z.; Liu, H.; Li, C.; Zhou, Y.; Zhao, Q.; Qin, A.; Li, M.; Gao, P.; Yan, L. Genome sequencing of a novel Verticillium dahliae strain (huangweibingjun). Scientific Reports 2025, 15, 15143. [Google Scholar] [CrossRef]
- Webster, J.; Dadd--Daigle, P.; Chapman, T.A.; Kirkby, K. Investigating the defoliating--like (DL) VCG2A pathotype of Verticillium dahliae through identification and prediction of secreted proteins from genomes of Australian isolates. Plant Pathology 2023, 72, 334–341. [Google Scholar] [CrossRef]
- Milgroom, M.G.; Jimenez-Gasco, M.d.M.; Olivares García, C.; Drott, M.T.; Jimenez-Diaz, R.M. Recombination between clonal lineages of the asexual fungus Verticillium dahliae detected by genotyping by sequencing. PloS one 2014, 9, e106740. [Google Scholar] [CrossRef] [PubMed]
- Huang, X. Horizontal transfer generates genetic variation in an asexual pathogen. PeerJ 2014, 2, e650. [Google Scholar] [CrossRef] [PubMed]
- Gurung, S.; Short, D.; Atallah, Z.; Subbarao, K. Clonal expansion of Verticillium dahliae in lettuce. Phytopathology 2014, 104, 641–649. [Google Scholar] [CrossRef]
- Dung, J.K.; Peever, T.L.; Johnson, D.A. A. Verticillium dahliae populations from mint and potato are genetically divergent with predominant haplotypes. Phytopathology 2013, 103, 445–459. [Google Scholar] [CrossRef]
- Usami, T.; Itoh, M.; Morii, S.; Miyamoto, T.; Kaneda, M.; Ogawara, T.; Amemiya, Y. Involvement of two different types of Verticillium dahliae in lettuce wilt in Ibaraki Prefecture, Japan. Journal of General Plant Pathology 2012, 78, 348–352. [Google Scholar] [CrossRef]
- Zhang, Y.-D.; Ji, X.-B.; Zong, J.; Dai, X.-F.; Klosterman, S.J.; Subbarao, K.V.; Zhang, D.-D.; Chen, J.-Y. Functional analysis of the mating type genes in Verticillium dahliae. BMC biology 2024, 22, 108. [Google Scholar] [CrossRef] [PubMed]
- Inderbitzin, P.; Subbarao, K.V. Verticillium systematics and evolution: how confusion impedes Verticillium wilt management and how to resolve it. Phytopathology 2014, 104, 564–574. [Google Scholar] [CrossRef]
- Bao, J.; Katan, J.; Shabi, E.; Katan, T. Vegetative-compatibility groups in Verticillium dahliae from Israel. European Journal of Plant Pathology 1998, 104, 263–269. [Google Scholar] [CrossRef]
- Dervis, S.; Erten, L.; Soylu, S.; Tok, F.M.; Kurt, S.; Yıldız, M.; Soylu, E.M. Vegetative compatibility groups in Verticillium dahliae isolates from olive in western Turkey. European Journal of Plant Pathology 2007, 119, 437–447. [Google Scholar] [CrossRef]
- Bejarano-Alcázar, J.; Blanco-López, M.; Melero-Vara, J.; Jiménez-Díaz, R.M. Etiology, importance, and distribution of Verticillium wilt of cotton in southern Spain. Plant Disease 1996, 80, 1233–1238. [Google Scholar] [CrossRef]
- Dadd-Daigle, P.; Labbate, M.; Chowdhury, P.R.; Kirkby, K.; Chapman, T. Australian Verticillium dahliae goes against the group–VCG 2A causes severe disease in Australian cotton. Phytopathology 2018, 108, S1.218. [Google Scholar]
- Subbarao, K.; Chassot, A.; Gordon, T.; Hubbard, J.; Bonello, P.; Mullin, R.; Okamoto, D.; Davis, R.; Koike, S. Genetic relationships and cross pathogenicities of Verticillium dahliae isolates from cauliflower and other crops. Phytopathology 1995, 85, 1105–1112. [Google Scholar] [CrossRef]
- Bhat, R.; Subbarao, K. Host range specificity in Verticillium dahliae. Phytopathology 1999, 89, 1218–1225. [Google Scholar] [CrossRef] [PubMed]
- Douhan, L.I.; Johnson, D. Vegetative compatibility and pathogenicity of Verticillium dahliae from spearmint and peppermint. Plant Disease 2001, 85, 297–302. [Google Scholar] [CrossRef]
- Strausbaugh, C.A.; Eujayl, I.A.; Martin, F.N. Pathogenicity, vegetative compatibility and genetic diversity of Verticillium dahliae isolates from sugar beet. Canadian Journal of Plant Pathology 2016, 38, 492–505. [Google Scholar] [CrossRef]
- Bhat, R.; Subbarao, K. Reaction of broccoli to isolates of Verticillium dahliae from various hosts. Plant Disease 2001, 85, 141–146. [Google Scholar] [CrossRef]
- Ikeda, K.; Banno, S.; Furusawa, A.; Shibata, S.; Nakaho, K.; Fujimura, M. Crop rotation with broccoli suppresses Verticillium wilt of eggplant. Journal of General Plant Pathology 2015, 81, 77–82. [Google Scholar] [CrossRef]
- Evans, G. Influence of weed hosts on the ecology of Verticillium dahliae in newly cultivated areas of the Namoi Valley, New South Wales. Annals of Applied Biology 1971, 67, 169–175. [Google Scholar] [CrossRef]
- Demirci, E.; Genc, T. Vegetative compatibility groups of Verticillium dahliae isolates from weeds in potato fields. Journal of Plant Pathology 2009, 671–676. [Google Scholar]
- Mpofu, S.; Hall, R. Accuracy and precision of population estimates of Verticillium dahliae on growth media in quantitative soil assays. Canadian Journal of Botany 2003, 81, 294–306. [Google Scholar] [CrossRef]
- Nadakuvukaren, M.; Hoener, C. An alcohol agar medium selective for determining Verticillium microsclerotia in soil. Phytopathology 1959, 49, 527–528. [Google Scholar]
- Ausher, R.; Katan, J.; Ovadia, S. An improved selective medium for the isolation of Verticillium dahliae. Phytoparasitica 1975, 3, 133–137. [Google Scholar] [CrossRef]
- Mansoori, B. An improved ethanol medium for efficient recovery and estimation of Verticillium dahliae populations in soil. Canadian Journal of Plant Pathology 2011, 33, 88–93. [Google Scholar] [CrossRef]
- Menzies, J.; Griebel, G. Survival and saprophytic growth of Verticillium dahliae in uncropped soil. Phytopathology 1967, 57, 703–709. [Google Scholar]
- Kabir, Z.; Bhat, R.; Subbarao, K.V. Comparison of media for recovery of Verticillium dahliae from soil. Plant Disease 2004, 88, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Hawke, M.A.; Lazarovits, G. Production and manipulation of individual microsclerotia of Verticillium dahliae for use in studies of survival. Phytopathology 1994, 84, 883–890. [Google Scholar] [CrossRef]
- Jordan, V. Estimation of the distribution of Verticillium populations in infected strawberry plants and soil. Plant Pathology 1971, 20, 21–24. [Google Scholar] [CrossRef]
- Christen, A. A Selective medium for isolating Verticillium albo-atrum from soil. Phytopathology 1982, 72, 47–49. [Google Scholar] [CrossRef]
- Atallah, Z.; Bae, J.; Jansky, S.; Rouse, D.; Stevenson, W. Multiplex real-time quantitative PCR to detect and quantify Verticillium dahliae colonization in potato lines that differ in response to Verticillium wilt. Phytopathology 2007, 97, 865–872. [Google Scholar] [CrossRef]
- Li, W.; Liu, L.; Li, J.; Yang, W.; Guo, Y.; Huang, L.; Yang, Z.; Peng, J.; Jin, X.; Lan, Y. Spectroscopic detection of cotton Verticillium wilt by spectral feature selection and machine learning methods. Frontiers in Plant Science 2025, 16, 1519001. [Google Scholar] [CrossRef]
- Mercado-Blanco, J.; Rodríguez-Jurado, D.; Parrilla-Araujo, S.; Jiménez-Díaz, R. Simultaneous detection of the defoliating and nondefoliating Verticillium dahliae pathotypes in infected olive plants by duplex, nested polymerase chain reaction. Plant Disease 2003, 87, 1487–1494. [Google Scholar] [CrossRef]
- Moradi, A.; Almasi, M.A.; Jafary, H.; Mercado--Blanco, J. A novel and rapid loop--mediated isothermal amplification assay for the specific detection of Verticillium dahliae. Journal of Applied Microbiology 2014, 116, 942–954. [Google Scholar] [CrossRef]
- Pasche, J.; Mallik, I.; Anderson, N.; Gudmestad, N. Development and validation of a real-time PCR assay for the quantification of Verticillium dahliae in potato. Plant Disease 2013, 97, 608–618. [Google Scholar] [CrossRef]
- Nazar, R.; Hu, X.; Schmidt, J.; Culham, D.; Robb, J. Potential use of PCR-amplified ribosomal intergenic sequences in the detection and differentiation of Verticillium wilt pathogens. Physiological and Molecular Plant Pathology 1991, 39, 1–11. [Google Scholar] [CrossRef]
- Wang, J.; Wang, Q.; Anchieta, A.G.; Chen, R.; Dai, X.-F.; Klosterman, S.J.; Chen, J.-Y.; Subbarao, K.V.; Zhang, D.-D. Species-specific primers derived from comparative genomics for PCR detection of Verticillium species. Plant Disease 2025, PDIS-10-24-2227-RE. [Google Scholar] [CrossRef]
- Wang, Q.; Qin, M.; Coleman, J.J.; Shang, W.; Hu, X. Rapid and sensitive detection of Verticillium dahliae from complex samples using CRISPR/Cas12a technology combined with RPA. Plant Disease 2023, 107, 1664–1669. [Google Scholar] [CrossRef]
- Triantafyllopoulou, A.; Tzanaki, A.; Balomenou, O.I.; Jiménez--Díaz, R.M.; Tzima, A.; Paplomatas, E. Development of a robust, VdNEP gene--based molecular marker to differentiate between pathotypes of Verticillium dahliae. Plant Pathology 2022, 71, 1404–1416. [Google Scholar] [CrossRef]
- Li, K.; Rouse, D.; German, T. PCR primers that allow intergeneric differentiation of ascomycetes and their application to Verticillium spp. Applied and Environmental Microbiology 1994, 60, 4324–4331. [Google Scholar] [CrossRef] [PubMed]
- Carder, J. Detection and differentiation by PCR of subspecific groups within two Verticillium species causing vascular wilts in herbaceous hosts. In Modern Assays for Plant Pathogenic Fungi: Identification, Detection and Quantification, Schots, A., Dewey, F.M., Oliver, R., Eds.; CAB International.: Oxford, 1994; pp. 91–97. [Google Scholar]
- Mercado--Blanco, J.; Rodríguez--Jurado, D.; Pérez--Artés, E.; Jiménez--Díaz, R.M. Detection of the nondefoliating pathotype of Verticillium dahliae in infected olive plants by nested PCR. Plant Pathology 2001, 50, 609–619. [Google Scholar] [CrossRef]
- Pérez-Artés, E.; García-Pedrajas, M.D.; Bejarano-Alcázar, J.; Jiménez-Díaz, R.M. Differentiation of cotton-defoliating and nondefoliating pathotypes of Verticillium dahliae by RAPD and specific PCR analyses. European Journal of Plant Pathology 2000, 106, 507–517. [Google Scholar] [CrossRef]
- Mercado-Blanco, J.; Rodríguez-Jurado, D.; Pérez-Artés, E.; Jiménez-Díaz, R.M. Detection of the defoliating pathotype of Verticillium dahliae in infected olive plants by nested PCR. European Journal of Plant Pathology 2002, 108, 1–13. [Google Scholar] [CrossRef]
- Tzelepis, G.; Bejai, S.; Sattar, M.N.; Schwelm, A.; Ilbäck, J.; Fogelqvist, J.; Dixelius, C. Detection of Verticillium species in Swedish soils using real-time PCR. Archives of Microbiology 2017, 199, 1383–1389. [Google Scholar] [CrossRef]
- Fang, Y.; Liu, L.; Zhao, W.; Dong, L.; He, L.; Liu, Y.; Yin, J.; Zhang, Y.; Miao, W.; Chen, D. Rapid and sensitive detection of Verticillium dahliae from soil using LAMP-CRISPR/Cas12a technology. International Journal of Molecular Sciences 2024, 25, 5185. [Google Scholar] [CrossRef]
- Paplomatas, E.; Bassett, D.; Broome, J.; DeVay, J. Incidence of Verticillium wilt and yield losses of cotton cultivars (Gossypium hirsutum) based on soil inoculum density of Verticillium dahliae. Phytopathology 1992, 82, 1417–1420. [Google Scholar] [CrossRef]
- Wei, F.; Shang, W.; Yang, J.; Hu, X.; Xu, X. Spatial pattern of Verticillium dahliae microsclerotia and cotton plants with wilt symptoms in commercial plantations. PloS one 2015, 10, e0132812. [Google Scholar] [CrossRef] [PubMed]
- Kang, X.; Huang, C.; Zhang, L.; Yang, M.; Zhang, Z.; Lyu, X. Assessing the severity of cotton Verticillium wilt disease from in situ canopy images and spectra using convolutional neural networks. The Crop Journal 2023, 11, 933–940. [Google Scholar] [CrossRef]
- Li, W.; Guo, Y.; Yang, W.; Huang, L.; Zhang, J.; Peng, J.; Lan, Y. Severity assessment of cotton canopy Verticillium wilt by machine learning based on feature selection and optimization algorithm using uav hyperspectral data. Remote Sensing 2024, 16, 4637. [Google Scholar] [CrossRef]
- Li, X.; Liang, Z.; Yang, G.; Lin, T.; Liu, B. Assessing the severity of Verticillium wilt in cotton fields and constructing pesticide application prescription maps using unmanned aerial vehicle (uav) multispectral images. Drones 2024, 8, 176. [Google Scholar] [CrossRef]
- Cal, A.D.; Martinez-Treceño, A.; Lopez-Aranda, J.; Melgarejo, P. Chemical alternatives to methyl bromide in Spanish strawberry nurseries. Plant Disease 2004, 88, 210–214. [Google Scholar] [CrossRef]
- Duniway, J.M. Status of chemical alternatives to methyl bromide for pre-plant fumigation of soil. Phytopathology 2002, 92, 1337–1343. [Google Scholar] [CrossRef]
- Pecchia, S.; Franceschini, A.; Santori, A.; Vannacci, G.; Myrta, A. Efficacy of dimethyl disulfide (DMDS) for the control of chrysanthemum Verticillium wilt in Italy. Crop Protection 2017, 93, 28–32. [Google Scholar] [CrossRef]
- Atallah, Z.K.; Hayes, R.J.; Subbarao, K.V. Fifteen years of Verticillium wilt of lettuce in America’s salad bowl: a tale of immigration, subjugation, and abatement. Plant Disease 2011, 95, 784–792. [Google Scholar] [CrossRef]
- Kurt, S.; Dervis, S.; Sahinler, S. Sensitivity of Verticillium dahliae to prochloraz and prochloraz–manganese complex and control of Verticillium wilt of cotton in the field. Crop Protection 2003, 22, 51–55. [Google Scholar] [CrossRef]
- Zhao, W.; Li, S.; Dong, L.; Wang, P.; Lu, X.; Zhang, X.; Su, Z.; Guo, Q.; Ma, P. Effects of different crop rotations on the incidence of cotton Verticillium wilt and structure and function of the rhizospheric microbial community. Plant and Soil 2023, 485, 457–474. [Google Scholar] [CrossRef]
- Nguyen, C.P.; Vo, B.; Le, D.P. Efficacy of commercial disinfectants against black root rot (Berkeleyomyces rouxiae) and Verticillium wilt (Verticillium dahliae) pathogens of cotton. Agronomy 2024, 14, 2502. [Google Scholar] [CrossRef]
- Wheeler, T.A.; Bordovsky, J.P.; Keeling, J.W. The effectiveness of crop rotation on management of Verticillium wilt over time. Crop Protection 2019, 121, 157–162. [Google Scholar] [CrossRef]
- Butterfield, E.; DeVay, J.; Garber, R. The influence of several crop sequences on the incidence of Verticillium wilt of cotton and on the populations of Verticillium dahliae in field soil. Phytopathology 1978, 68, 1217–1220. [Google Scholar] [CrossRef]
- Chen, A.; Morrison, S.; Gregson, A.; Le, D.P.; Urquhart, A.S.; Smith, L.J.; Aitken, E.A.; Gardiner, D.M. Fluorescently Tagged Verticillium dahliae to understand the infection process on cotton (Gossypium hirsutum) and weed plant species. Pathogens 2024, 13, 442. [Google Scholar] [CrossRef] [PubMed]
- Allen, S.J. Diseases of Cotton. Final Report (DAN26L); Cotton Research Council, Narrabri, NSW Australia 1989, pp. 1-37.
- Ligoxigakis, E.; Vakalounakis, D.; Thanassoulopoulos, C. Weed hosts of Verticillium dahliae in Crete: Susceptibility, symptomatology and significance. Phytoparasitica 2002, 30, 511–518. [Google Scholar] [CrossRef]
- Huisman, O.; Ashworth, L., Jr. Influence of crop rotation on survival of Verticillium albo-atrum in soils. 1976, 978-981.
- Khan, F.H.; Tie, Z.; Zhang, X.; Ma, Y.; Yu, Y.; Zhao, S.; Zhang, X.; Xi, H. Short-term maize rotation suppresses Verticillium wilt and restructures soil microbiomes in Xinjiang cotton fields. Microorganisms 2025, 13, 1968. [Google Scholar] [CrossRef]
- Huang, J.; Li, H.; Yuan, H. Effect of organic amendments on Verticillium wilt of cotton. Crop Protection 2006, 25, 1167–1173. [Google Scholar] [CrossRef]
- Lang, J.; Hu, J.; Ran, W.; Xu, Y.; Shen, Q. Control of cotton Verticillium wilt and fungal diversity of rhizosphere soils by bio-organic fertilizer. Biology and Fertility of Soils 2012, 48, 191–203. [Google Scholar] [CrossRef]
- Li, R.; Tao, R.; Ling, N.; Chu, G. Chemical, organic and bio-fertilizer management practices effect on soil physicochemical property and antagonistic bacteria abundance of a cotton field: Implications for soil biological quality. Soil and Tillage Research 2017, 167, 30–38. [Google Scholar] [CrossRef]
- Goicoechea, N. To what extent are soil amendments useful to control Verticillium wilt? Pest Management Science: Formerly Pesticide Science 2009, 65, 831–839. [Google Scholar] [CrossRef]
- Ochiai, N.; Powelson, M.; Crowe, F.; Dick, R. Green manure effects on soil quality in relation to suppression of Verticillium wilt of potatoes. Biology and Fertility of Soils 2008, 44, 1013–1023. [Google Scholar] [CrossRef]
- Wheeler, T.A.; Bordovsky, J.P.; Keeling, J.W.; Keeling, W.; McCallister, D. The effects of tillage system and irrigation on Verticillium wilt and cotton yield. Crop Protection 2020, 137, 105305. [Google Scholar] [CrossRef]
- Albers, C. Vertical distribution of Verticillium dahliae and the impact of nitrate nitrogen on cotton growth, yield, quality and disease development. Master Thesis, Texas Tech University, 2013; pp. 1–105.
- Bell, A.A. Verticillium wilt. In Cotton Diseases, J.Hillocks, R., Ed.; CAB International: Wallingford, UK, 1992; pp. 87–122. [Google Scholar]
- Zhang, L.; Su, X.; Meng, H.; Men, Y.; Liu, C.; Yan, X.; Song, X.; Sun, X.; Mao, L. Cotton stubble return and subsoiling alter soil microbial community, carbon and nitrogen in coastal saline cotton fields. Soil and Tillage Research 2023, 226, 105585. [Google Scholar] [CrossRef]
- Zhang, G.; Meng, Z.; Ge, H.; Yuan, J.; Qiang, S.; Jiang, P.a.; Ma, D. Investigating Verticillium wilt occurrence in cotton and its risk management by the direct return of cotton plants infected with Verticillium dahliae to the field. Frontiers in Plant Science 2023, 14, 1220921. [Google Scholar] [CrossRef] [PubMed]
- CRDC. Cotton Pest Management Guide 2019/2020. Cotton Research and Corporation Development: Narrabri, NSW Australia, 2019, pp. 1-152.
- Davis, R.; Colyer, P.; Rothrock, C.; Kochman, J. Fusarium wilt of cotton: population diversity and implications for management. Plant Disease 2006, 90, 692–703. [Google Scholar] [CrossRef]
- Erdogan, O.; Benlioglu, K. Biological control of Verticillium wilt on cotton by the use of fluorescent Pseudomonas spp. under field conditions. Biological Control 2010, 53, 39–45. [Google Scholar] [CrossRef]
- Shan, Y.; Wang, D.; Zhao, F.-H.; Song, J.; Zhu, H.; Li, Y.; Zhang, X.-J.; Dai, X.-F.; Han, D.; Chen, J.-Y. Insights into the biocontrol and plant growth promotion functions of Bacillus altitudinis strain KRS010 against Verticillium dahliae. BMC Biology 2024, 22, 116. [Google Scholar] [CrossRef] [PubMed]
- Xue, L.; Xue, Q.; Chen, Q.; Lin, C.; Shen, G.; Zhao, J. Isolation and evaluation of rhizosphere actinomycetes with potential application for biocontrol of Verticillium wilt of cotton. Crop Protection 2013, 43, 231–240. [Google Scholar] [CrossRef]
- Yang, W.; Zheng, L.; Liu, H.-X.; Wang, K.-B.; Yu, Y.-Y.; Luo, Y.-M.; Guo, J.-H. Evaluation of the effectiveness of a consortium of three plant-growth promoting rhizobacteria for biocontrol of cotton Verticillium wilt. Biocontrol Science and Technology 2014, 24, 489–502. [Google Scholar] [CrossRef]
- Zhu, H.Q.; Feng, Z.L.; Li, Z.F.; Shi, Y.Q.; Zhao, L.H.; Yang, J.R. Characterization of two fungal isolates from cotton and evaluation of their potential for biocontrol of Verticillium wilt of cotton. Journal of Phytopathology 2013, 161, 70–77. [Google Scholar] [CrossRef]
- Li, C.H.; Shi, L.; Han, Q.; Hu, H.L.; Zhao, M.W.; Tang, C.M.; Li, S.P. Biocontrol of verticillium wilt and colonization of cotton plants by an endophytic bacterial isolate. Journal of Applied Microbiology 2012, 113, 641–651. [Google Scholar] [CrossRef]
- Liu, L.; Medison, R.G.; Zheng, T.-w.; Meng, X.-j.; Sun, Z.-x.; Zhou, Y. Biocontrol potential of Bacillus amyloliquefaciens YZU-SG146 from Fraxinus hupehensis against Verticillium wilt of cotton. Biological Control 2023, 183, 105246. [Google Scholar] [CrossRef]
- Tang, Y.; Xue, Q.; Yu, J.; Zhang, Z.; Wang, Z.; Wang, L.; Feng, H. Biocontrol and growth-promoting potential of antagonistic strain yl84 against Verticillium dahliae. Agronomy 2025, 15, 1997. [Google Scholar] [CrossRef]
- Zhao, F.; Wang, D.; Liu, H.; Shan, Y.; Qi, H.Y.; Dai, X.; Chen, J.; Lesueur, D.; Han, D.; Zhang, X. The biocontrol strain Bacillus mojavensis KRS009 confers resistance to cotton Verticillium wilt and improves tolerance to salt stress. New Plant Protection 2024, 1, e20. [Google Scholar] [CrossRef]
- Zhang, L.; Wang, Y.; Lei, S.; Zhang, H.; Liu, Z.; Yang, J.; Niu, Q. Effect of volatile compounds produced by the cotton endophytic bacterial strain Bacillus sp. T6 against Verticillium wilt. BMC Microbiology 2023, 23, 8. [Google Scholar] [CrossRef]
- Song, J.; Wang, D.; Han, D.; Zhang, D.-D.; Li, R.; Kong, Z.-Q.; Dai, X.-F.; Subbarao, K.V.; Chen, J.-Y. Characterization of the endophytic Bacillus subtilis KRS015 strain for its biocontrol efficacy against Verticillium dahliae. Phytopathology 2024, 114, 61–72. [Google Scholar] [CrossRef]
- Zhang, F.; Li, X.-L.; Zhu, S.-J.; Ojaghian, M.R.; Zhang, J.-Z. Biocontrol potential of Paenibacillus polymyxa against Verticillium dahliae infecting cotton plants. Biological Control 2018, 127, 70–77. [Google Scholar] [CrossRef]
- Yuan, Y.; Feng, H.; Wang, L.; Li, Z.; Shi, Y.; Zhao, L.; Feng, Z.; Zhu, H. Potential of endophytic fungi isolated from cotton roots for biological control against Verticillium wilt disease. PLoS One 2017, 12, e0170557. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Xue, Q.-Y.; Xu, L.-L.; Xu, Q.; Lu, S.; Gu, C.; Guo, J.-H. A screening strategy of fungal biocontrol agents towards Verticillium wilt of cotton. Biological Control 2011, 56, 209–216. [Google Scholar] [CrossRef]
- Wei, F.; Zhang, Y.; Shi, Y.; Feng, H.; Zhao, L.; Feng, Z.; Zhu, H. Evaluation of the biocontrol potential of endophytic fungus Fusarium solani CEF559 against Verticillium dahliae in cotton plant. BioMed Research International 2019, 2019, 3187943. [Google Scholar] [CrossRef]
- Cook, R.J. Biological control of plant pathogens: theory to application. Phytopathology 1985, 75, 25–29. [Google Scholar] [CrossRef]
- Deketelaere, S.; Tyvaert, L.; França, S.C.; Höfte, M. Desirable traits of a good biocontrol agent against Verticillium wilt. Frontiers in Microbiology 2017, 8, 1186. [Google Scholar] [CrossRef]
- Weller, D.M. Pseudomonas biocontrol agents of soilborne pathogens: looking back over 30 years. Phytopathology 2007, 97, 250–256. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Sanogo, S.; Flynn, R.; Baral, J.B.; Bajaj, S.; Hughs, S.; Percy, R.G. Germplasm evaluation and transfer of Verticillium wilt resistance from Pima (Gossypium barbadense) to Upland cotton (G. hirsutum). Euphytica 2012, 187, 147–160. [Google Scholar] [CrossRef]
- Göre, M.E.; Erdoğan, O.; Altın, N. Searching for resistance sources to Verticillium wilt of cotton in seedlings from Gossypium spp. Tropical Plant Pathology 2017, 42, 28–31. [Google Scholar] [CrossRef]
- Colson-Hanks, E.; Allen, S.; Deverall, B. Effect of 2, 6-dichloroisonicotinic acid or benzothiadiazole on Alternaria leaf spot, bacterial blight and Verticillium wilt in cotton under field conditions. Australasian Plant Pathology 2000, 29, 170–177. [Google Scholar] [CrossRef]
- Dong, H.; Zhang, X.; Choen, Y.; Zhou, Y.; Li, W.; Li, Z. Dry mycelium of Penicillium chrysogenum protects cotton plants against wilt diseases and increases yield under field conditions. Crop Protection 2006, 25, 324–330. [Google Scholar] [CrossRef]
- Hanson, L.E. Reduction of Verticillium wilt symptoms in cotton following seed treatment with Trichoderma virens. Journal of Cotton Science 2000, 4, 224–231. [Google Scholar]
- Howell, C.R. Understanding the mechanisms employed by Trichoderma virens to effect biological control of cotton diseases. Phytopathology 2006, 96, 178–180. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Cohen, Y. Dry mycelium of Penicillium chrysogenum induces resistance against Verticillium wilt and enhances growth of cotton plants. Phytoparasitica 2002, 30, 147–157. [Google Scholar] [CrossRef]
- Smit, F.; Dubery, I.A. Cell wall reinforcement in cotton hypocotyls in response to a Verticillium dahliae elicitor. Phytochemistry 1997, 44, 811–815. [Google Scholar] [CrossRef]
- Dong, H.; Li, W.; Zhang, D.; Tang, W. Differential expression of induced resistance by an aqueous extract of killed Penicillium chrysogenum against Verticillium wilt of cotton. Crop Protection 2003, 22, 129–134. [Google Scholar] [CrossRef]
- Chen, Z.; Wang, J.; Li, Y.; Zhong, Y.; Liao, J.; Lu, S.; Wang, L.; Wang, X.; Chen, S. Dry mycelium of Penicillium chrysogenum activates defense via gene regulation of salicylic acid and jasmonic acid signaling in Arabidopsis. Physiological and Molecular Plant Pathology 2018, 103, 54–61. [Google Scholar] [CrossRef]
- Ni, H.; Kong, W.-L.; Zhang, Q.-Q.; Wu, X.-Q. Volatiles emitted by Pseudomonas aurantiaca ST-TJ4 trigger systemic plant resistance to Verticillium dahliae. Microbiological Research 2024, 287, 127834. [Google Scholar] [CrossRef]
- Dann, E.; Diers, B.; Byrum, J.; Hammerschmidt, R. Effect of treating soybean with 2, 6-dichloroisonicotinic acid (INA) and benzothiadiazole (BTH) on seed yields and the level of disease caused by Sclerotinia sclerotiorum in field and greenhouse studies. European Journal of Plant Pathology 1998, 104, 271–278. [Google Scholar] [CrossRef]
- McFadden, H.; Chapple, R.; Feyter, R.D.; Dennis, E. Expression of pathogenesis-related genes in cotton stems in response to infection by Verticillium dahliae. Physiological and Molecular Plant Pathology 2001, 58, 119–131. [Google Scholar] [CrossRef]
- Bell, A.A.; Mace, M.E. Biochemistry and physiology of resistance. In Fungal Wilt Diseases of Plants; Elsevier: 1981, pp. 431-486.
- Daayf, F.; Nicole, M.; Boher, B.; Pando, A.; Geiger, J.-P. Early vascular defense reactions of cotton roots infected with a defoliating mutant strain of Verticillium dahliae. European Journal of Plant Pathology 1997, 103, 125–136. [Google Scholar] [CrossRef]
- Xu, L.; Zhu, L.; Tu, L.; Liu, L.; Yuan, D.; Jin, L.; Long, L.; Zhang, X. Lignin metabolism has a central role in the resistance of cotton to the wilt fungus Verticillium dahliae as revealed by RNA-Seq-dependent transcriptional analysis and histochemistry. Journal of Experimental Botany 2011, 62, 5607–5621. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, X.; Rong, W.; Yang, J.; Li, Z.; Wu, L.; Zhang, G.; Ma, Z. Histochemical analyses reveal that stronger intrinsic defenses in Gossypium barbadense than in G. hirsutum are associated with resistance to Verticillium dahliae. Molecular Plant-Microbe Interactions 2017, 30, 984–996. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Zhao, M.; Li, T.; Wang, L.; Liao, C.; Liu, D.; Zhang, H.; Zhao, Y.; Liu, L.; Ge, X. Interactions between Verticillium dahliae and cotton: pathogenic mechanism and cotton resistance mechanism to Verticillium wilt. Frontiers in Plant Science 2023, 14, 1174281. [Google Scholar] [CrossRef]
- Mace, M.E. Contributions of tyloses and terpenoid aldehyde phytoalexins to Verticillium wilt resistance in cotton. Physiological Plant Pathology 1978, 12, 1–11. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, Y.; Zhao, T.; Yang, J.; Feng, S.; Nazeer, W.; Zhang, T.; Zhou, B. A new synthetic amphiploid (AADDAA) between Gossypium hirsutum and G. arboreum lays the foundation for transferring resistances to Verticillium and drought. PloS One 2015, 10, e0128981. [Google Scholar] [CrossRef] [PubMed]
- Harrison, N.; Beckman, C. Time/space relationships of colonization and host response in wilt-resistant and wilt-susceptible cotton (Gossypium) cultivars inoculated with Verticillium dahliae and Fusarium oxysporum f. sp. vasinfectum. Physiological Plant Pathology 1982, 21, 193–207. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhao, L.; Li, D.; Li, Z.; Feng, H.; Feng, Z.; Wei, F.; Zhou, J.; Ma, Z.; Yang, J. A comprehensive review on elucidating the host disease resistance mechanism from the perspective of the interaction between cotton and Verticillium dahliae. Journal of Cotton Research 2025, 8, 5. [Google Scholar] [CrossRef]
- Wilhelm, S.; Sagen, J.E.; Tietz, H. Resistance to Verticillium wilt in cotton: sources, techniques of identification, inheritance trends, and the resistance. Phytopathology 1974, 64, 924–931. [Google Scholar] [CrossRef]
- Bell, A.A.; Stipanovic, R.D.; Mace, M.E.; Kohel, R.J. Genetic manipulation of terpenoid phytoalexins in Gossypium: effects of disease resistance. In Genetic Engineering of Plant Secondary Metabolism, Ellis, B.E., Kuroki, G.W., Stafford, H.A., Eds. Ellis, B.E., Kuroki, G.W., Stafford, H.A., Eds.; Springer: 1994, pp. 231-249.
- Egamberdiyev, A.E. Amphidiploids of cotton as a new source of wilt resistance. In Proceedings of the the Fifth International Verticillium Symposium, Leningrad, Russia 1990; p. 69. 69.
- Fradin, E.F.; Thomma, B.P. Physiology and molecular aspects of Verticillium wilt diseases caused by V. dahliae and V. albo--atrum. Molecular Plant Pathology 2006, 7, 71–86. [Google Scholar] [CrossRef]
- Talboys, P. Resistance to vascular wilt fungi. Proceedings of the Royal Society of London. Series B. Biological Sciences 1972, 181, 319–332. [Google Scholar]
- Wilhelm, S.; Sagen, J.; Tietz, H. Resistance to Verticillium wilt transferred from Gossypium barbadense to upland cotton phenotype. Phytopathology 1972, 62, 798–799. [Google Scholar]
- Aguado, A.; De Los Santos, B.; Blanco, C.; Romero, F. Study of gene effects for cotton yield and Verticillium wilt tolerance in cotton plant (Gossypium hirsutum L.). Field Crops Research 2008, 107, 78–86. [Google Scholar] [CrossRef]
- Bolek, Y.; Bell, A.; El--Zik, K.; Thaxton, P.; Magill, C. Reaction of cotton cultivars and an F2 population to stem inoculation with isolates Verticillium dahliae. Journal of Phytopathology 2005, 153, 269–273. [Google Scholar] [CrossRef]
- Mert, M.; Kurt, S.; Gencer, O.; Akiscan, Y.; Boyaci, K.; Tok, F. Inheritance of resistance to Verticillium wilt (Verticillium dahliae) in cotton (Gossypium hirsutum L.). Plant Breeding 2005, 124, 102–104. [Google Scholar] [CrossRef]
- Zhang, J.; Fang, H.; Zhou, H.; Sanogo, S.; Ma, Z. Genetics, breeding, and marker--assisted selection for Verticillium wilt resistance in cotton. Crop Science 2014, 54, 1289–1303. [Google Scholar] [CrossRef]
- Kawchuk, L.M.; Hachey, J.; Lynch, D.R.; Kulcsar, F.; Van Rooijen, G.; Waterer, D.R.; Robertson, A.; Kokko, E.; Byers, R.; Howard, R.J. Tomato Ve disease resistance genes encode cell surface-like receptors. Proceedings of the National Academy of Sciences 2001, 98, 6511–6515. [Google Scholar] [CrossRef]
- Song, Y.; Liu, L.; Wang, Y.; Valkenburg, D.J.; Zhang, X.; Zhu, L.; Thomma, B.P. Transfer of tomato immune receptor Ve1 confers Ave1--dependent Verticillium resistance in tobacco and cotton. Plant Biotechnology Journal 2018, 16, 638–648. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, X.; Yang, S.; Chi, J.; Zhang, G.; Ma, Z. Cloning and characterization of a Verticillium wilt resistance gene from Gossypium barbadense and functional analysis in Arabidopsis thaliana. Plant Cell Reports 2011, 30, 2085–2096. [Google Scholar] [CrossRef]
- Li, N.Y.; ma, X.F.; Short, D.P.; Li, T.G.; Zhou, L.; Gui, Y.J.; Kong, Z.Q.; Zhang, D.D.; Zhang, W.Q.; Li, J.J. The island cotton NBS--LRR gene GbaNA1 confers resistance to the non--race 1 Verticillium dahliae isolate Vd991. Molecular Plant Pathology 2018, 19, 1466–1479. [Google Scholar] [CrossRef]
- Chen, J.Y.; Liu, C.; Gui, Y.J.; Si, K.W.; Zhang, D.D.; Wang, J.; Short, D.P.; Huang, J.Q.; Li, N.Y.; Liang, Y. Comparative genomics reveals cotton--specific virulence factors in flexible genomic regions in Verticillium dahliae and evidence of horizontal gene transfer from Fusarium. New Phytologist 2018, 217, 756–770. [Google Scholar] [CrossRef]
- Duan, X.; Zhang, Z.; Wang, J.; Zuo, K. Characterization of a novel cotton subtilase gene GbSBT1 in response to extracellular stimulations and its role in Verticillium resistance. PloS one 2016, 11, e0153988. [Google Scholar] [CrossRef]
- Gao, W.; Long, L.; Zhu, L.-F.; Xu, L.; Gao, W.-H.; Sun, L.-Q.; Liu, L.-L.; Zhang, X.-L. Proteomic and virus-induced gene silencing (VIGS) analyses reveal that gossypol, brassinosteroids, and jasmonic acid contribute to the resistance of cotton to Verticillium dahliae. Molecular and Cellular Proteomics 2013, 12, 3690–3703. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.-B.; Han, L.-B.; Wang, H.-Y.; Zhang, J.; Sun, S.-T.; Feng, D.-Q.; Yang, C.-L.; Sun, Y.-D.; Zhong, N.-Q.; Xia, G.-X. The thioredoxin GbNRX1 plays a crucial role in homeostasis of apoplastic reactive oxygen species in response to Verticillium dahliae infection in cotton. Plant Physiology 2016, 170, 2392–2406. [Google Scholar] [CrossRef] [PubMed]
- Mo, H.; Wang, X.; Zhang, Y.; Zhang, G.; Zhang, J.; Ma, Z. Cotton polyamine oxidase is required for spermine and camalexin signalling in the defence response to Verticillium dahliae. The Plant Journal 2015, 83, 962–975. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Ma, Q.; Zhang, Y.; Wang, X.; Zhang, G.; Ma, Z. Molecular cloning and functional analysis of GbRVd, a gene in Gossypium barbadense that plays an important role in conferring resistance to Verticillium wilt. Gene 2016, 575, 687–694. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wang, X.F.; Ding, Z.G.; Ma, Q.; Zhang, G.R.; Zhang, S.L.; Li, Z.K.; Wu, L.Q.; Zhang, G.Y.; Ma, Z.Y. Transcriptome profiling of Gossypium barbadense inoculated with Verticillium dahliae provides a resource for cotton improvement. BMC Genomics 2013, 14, 637. [Google Scholar] [CrossRef]
- Hill, M.K.; Lyon, K.J.; Lyon, B.R. Identification of disease response genes expressed in Gossypium hirsutum upon infection with the wilt pathogen Verticillium dahliae. Plant Molecular Biology 1999, 40, 289–296. [Google Scholar] [CrossRef]
- Abdelraheem, A.; Liu, F.; Song, M.; Zhang, J.F. A meta-analysis of quantitative trait loci for abiotic and biotic stress resistance in tetraploid cotton. Molecular Genetics and Genomics 2017, 292, 1221–1235. [Google Scholar] [CrossRef]
- Said, J.I.; Lin, Z.; Zhang, X.; Song, M.; Zhang, J. A comprehensive meta QTL analysis for fiber quality, yield, yield related and morphological traits, drought tolerance, and disease resistance in tetraploid cotton. BMC Genomics 2013, 14, 776. [Google Scholar] [CrossRef]
- Bernardo, R. Bandwagons I, too, have known. Theoretical and Applied Genetics 2016, 129, 2323–2332. [Google Scholar] [CrossRef]
- Li, C.-Q.; Liu, G.-S.; Zhao, H.-H.; Wang, L.-J.; Zhang, X.-F.; Liu, Y.; Zhou, W.-Y.; Yang, L.-L.; Li, P.-B.; Wang, Q.-L. Marker-assisted selection of Verticillium wilt resistance in progeny populations of upland cotton derived from mass selection-mass crossing. Euphytica 2013, 191, 469–480. [Google Scholar] [CrossRef]
- Zhang, Z.; Ge, X.; Luo, X.; Wang, P.; Fan, Q.; Hu, G.; Xiao, J.; Li, F.; Wu, J. Simultaneous editing of two copies of Gh14-3-3d confers enhanced transgene-clean plant defense against Verticillium dahliae in allotetraploid upland cotton. Frontiers in Plant Science 2018, 9, 842. [Google Scholar] [CrossRef]
- Zhao, Y.; Chen, W.; Cui, Y.; Sang, X.; Lu, J.; Jing, H.; Wang, W.; Zhao, P.; Wang, H. Detection of candidate genes and development of KASP markers for Verticillium wilt resistance by combining genome-wide association study, QTL-seq and transcriptome sequencing in cotton. Theoretical and Applied Genetics 2021, 134, 1063–1081. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, J.; Chen, Q.; Zheng, K.; Deng, X.; Gao, W.; Pei, W.; Geng, S.; Deng, Y.; Li, C. Quantitative trait locus mapping and identification of candidate genes for resistance to Verticillium wilt in four recombinant inbred line populations of Gossypium hirsutum. Plant Science 2023, 327, 111562. [Google Scholar] [CrossRef]
- Wilson, I.W.; Moncuquet, P.; Yuan, Y.; Soliveres, M.; Li, Z.; Stiller, W.; Zhu, Q.-H. Genetic mapping and characterization of Verticillium wilt resistance in a recombinant inbred population of upland cotton. International Journal of Molecular Sciences 2024, 25, 2439. [Google Scholar] [CrossRef] [PubMed]
- Bayles, M.B.; Verhalen, L.M.; Johnson, W.M.; Barnes, B.R. Trends over time among cotton cultivars released by the Oklahoma Agricultural Experiment Station. Crop Science 2005, 45, 966–980. [Google Scholar] [CrossRef]
- Zhou, H.; Fang, H.; Sanogo, S.; Hughs, S.E.; Jones, D.C.; Zhang, J. Evaluation of Verticillium wilt resistance in commercial cultivars and advanced breeding lines of cotton. Euphytica 2014, 196, 437–448. [Google Scholar] [CrossRef]
- Liu, S.; Constable, G.; Reid, P.; Stiller, W.; Cullis, B. The interaction between breeding and crop management in improved cotton yield. Field Crops Research 2013, 148, 49–60. [Google Scholar] [CrossRef]
- Ayyaz, M.; Chang, Z.; Ding, S.; Han, P.; Xu, L.; Abudukeyoumu, A.; Siddho, I.A.; Li, Z.; Lin, H.; Xu, J. QTL mapping associated with Verticillium wilt resistance in cotton based on MAGIC population. Journal of Cotton Research 2025, 8, 1–15. [Google Scholar] [CrossRef]
- Katsantonis, D.; Hillocks, R.J.; Gowen, S. Comparative effect of root-knot nematode on severity of Verticillium and Fusarium wilt in cotton. Phytoparasitica 2003, 31, 154–162. [Google Scholar] [CrossRef]
- Wilhelm, S.; Sagen, J.; Tietz, H. Phenotype modification in cotton for control of Verticillium wilt through dense plant population culture. Plant Disease 1985, 69, 283–288. [Google Scholar] [CrossRef]
- Devey, M.; Rosielle, A. relationship between field and greenhouse ratings for tolerance to verticillium wilt on cotton 1. Crop Science 1986, 26, 1–4. [Google Scholar] [CrossRef]
- Khaskheli, M.I.; Sun, J.L.; He, S.P.; Du, X.M. Screening of cotton germplasm for resistance to Verticillium dahlia Kleb. under greenhouse and field conditions. European Journal of Plant Pathology 2013, 137, 259–272. [Google Scholar] [CrossRef]
- Varo, A.; Moral, J.; Lozano-Tóvar, M.D.; Trapero, A. Development and validation of an inoculation method to assess the efficacy of biological treatments against Verticillium wilt in olive trees. BioControl 2016, 61, 283–292. [Google Scholar] [CrossRef]
- Bell, A. Tannin concentrations in VerticiIlium-infected cotton: relationships to strain virulence and cultivar resistance. In Proceedings of the Proceedings of the Beltwide Cotton Production Research Conference, Memphis, Tennessee, 1991, p. 187.
- Phillips, D.J.; Wilhelm, S. Root distribution as a factor influencing symptom expression of Verticillium wilt of Cotton. Phytopathology 1971, 61, 1312–1313. [Google Scholar] [CrossRef]
- Schnathorst, W.; Cooper, J. Anomalies in Field and Greenhouse Reaction of Certain Cotton Cultivars in Fected with Verticillium dahliae. In Proceedings of the Proc. Beltwide Cotton Prod. Conf; 1975; pp. 148–149. [Google Scholar]
- Bell. AA, B.A.; Presley. JT, P.J. Temperature effects upon resistance and phytoalexin synthesis in cotton inoculated with Verticillium albo-atrum. Phytopathology 1969, 59, 1141–1151. [Google Scholar]
- Tan, F.; Gao, X.; Cang, H.; Wu, N.; Di, R.; Yan, J.; Li, C.; Gao, P.; Lv, X. Early detection of Verticillium wilt in cotton by using hyperspectral imaging combined with recurrence plots. Agronomy 2025, 15, 213. [Google Scholar] [CrossRef]
- Bennett, R. Survival of Fusarium oxysporum f. sp. vasinfectum chlamydospores under solarization temperatures. Plant Disease 2012, 96, 1564–1568. [Google Scholar] [CrossRef]
- Ben-Yephet, Y.; Melero-Vera, J.; DeVay, J. Interaction of soil solarization and metham-sodium in the destruction of Verticillium dahliae and Fusarium oxysporum f. sp. vasinfectum. Crop Protection 1988, 7, 327–331. [Google Scholar] [CrossRef]
- Blok, W.J.; Lamers, J.G.; Termorshuizen, A.J.; Bollen, G.J. Control of soilborne plant pathogens by incorporating fresh organic amendments followed by tarping. Phytopathology 2000, 90, 253–259. [Google Scholar] [CrossRef]
- Shennan, C.; Muramoto, J.; Koike, S.; Baird, G.; Fennimore, S.; Samtani, J.; Bolda, M.; Dara, S.; Daugovish, O.; Lazarovits, G. Anaerobic soil disinfestation is an alternative to soil fumigation for control of some soilborne pathogens in strawberry production. Plant Pathology 2018, 67, 51–66. [Google Scholar] [CrossRef]
- Neubauer, C.; Heitmann, B.; Müller, C. Biofumigation potential of Brassicaceae cultivars to Verticillium dahliae. European Journal of Plant Pathology 2014, 140, 341–352. [Google Scholar] [CrossRef]
- Neubauer, C.; Hüntemann, K.; Heitmann, B.; Müller, C. Suppression of Verticillium dahliae by glucosinolate-containing seed meal amendments. European Journal of Plant Pathology 2015, 142, 239–249. [Google Scholar] [CrossRef]
- Wei, F.; Passey, T.; Xu, X. Effects of individual and combined use of bio-fumigation-derived products on the viability of Verticillium dahliae microsclerotia in soil. Crop Protection 2016, 79, 170–176. [Google Scholar] [CrossRef]
- Gharbi, Y.; Barkallah, M.; Bouazizi, E.; Cheffi, M.; Krid, S.; Triki, M.A.; Gdoura, R. Development and validation of a new real-time assay for the quantification of Verticillium dahliae in the soil: A comparison with conventional soil plating. Mycological Progress 2016, 15, 54. [Google Scholar] [CrossRef]
- Nizamani, M.M.; Zhang, Q.; Muhae-Ud-Din, G.; Wang, Y. High-throughput sequencing in plant disease management: a comprehensive review of benefits, challenges, and future perspectives. Phytopathology Research 2023, 5, 1–17. [Google Scholar] [CrossRef]
- Suster, C.J.; Pham, D.; Kok, J.; Sintchenko, V. Emerging applications of artificial intelligence in pathogen genomics. Frontiers in Bacteriology 2024, 3, 1326958. [Google Scholar] [CrossRef]
- Klosterman, S.J.; Subbarao, K.V.; Kang, S.; Veronese, P.; Gold, S.E.; Thomma, B.P.; Chen, Z.; Henrissat, B.; Lee, Y.-H.; Park, J. Comparative genomics yields insights into niche adaptation of plant vascular wilt pathogens. PLoS pathogens 2011, 7, e1002137. [Google Scholar] [CrossRef]
- Yu, J.; Li, T.; Tian, L.; Tang, C.; Klosterman, S.J.; Tian, C.; Wang, Y. Two Verticillium dahliae MAPKKKs, VdSsk2 and VdSte11, have distinct roles in pathogenicity, microsclerotial formation, and stress adaptation. Msphere 2019, 4. [Google Scholar] [CrossRef]
- Depotter, J.R.; Shi--Kunne, X.; Missonnier, H.; Liu, T.; Faino, L.; van den Berg, G.C.; Wood, T.A.; Zhang, B.; Jacques, A.; Seidl, M.F. Dynamic virulence--related regions of the plant pathogenic fungus Verticillium dahliae display enhanced sequence conservation. Molecular Ecology 2019, 28, 3482–3495. [Google Scholar] [CrossRef]
- Wang, H.; Tang, C.; Deng, C.; Li, W.; Klosterman, S.J.; Wang, Y. Septins regulate virulence in Verticillium dahliae and differentially contribute to microsclerotial formation and stress responses. Phytopathology Research 2022, 4, 40. [Google Scholar] [CrossRef]
- Eisermann, I.; Garduño--Rosales, M.; Talbot, N.J. The emerging role of septins in fungal pathogenesis. Cytoskeleton 2023, 80, 242–253. [Google Scholar] [CrossRef]
- Wang, D.; Zhao, Z.; Long, Y.; Fan, R. Protein Kinase C Is Involved in Vegetative Development, Stress Response and Pathogenicity in Verticillium dahliae. International Journal of Molecular Sciences 2023, 24, 14266. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, D.-D.; Usami, T.; Liu, L.; Yang, L.; Huang, J.-Q.; Song, J.; Li, R.; Kong, Z.-Q.; Li, J.-J. Functional genomics and comparative lineage-specific region analyses reveal novel insights into race divergence in Verticillium dahliae. Microbiology Spectrum 2021, 9, e01118–01121. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Liu, R.; Lv, J.; Feng, Z.; Wei, F.; Zhao, L.; Zhang, Y.; Zhu, H.; Feng, H. The glycoside hydrolase 28 member VdEPG1 is a virulence factor of Verticillium dahliae and interacts with the jasmonic acid pathway--related gene GhOPR9. Molecular Plant Pathology 2023, 24, 1238–1255. [Google Scholar] [CrossRef] [PubMed]
- Bui, T.T.; Harting, R.; Braus--Stromeyer, S.A.; Tran, V.T.; Leonard, M.; Höfer, A.; Abelmann, A.; Bakti, F.; Valerius, O.; Schlüter, R. Verticillium dahliae transcription factors Som1 and Vta3 control microsclerotia formation and sequential steps of plant root penetration and colonisation to induce disease. New Phytologist 2019, 221, 2138–2159. [Google Scholar] [CrossRef] [PubMed]
- Maurus, I.; Harting, R.; Herrfurth, C.; Starke, J.; Nagel, A.; Mohnike, L.; Chen, Y.-Y.; Schmitt, K.; Bastakis, E.; Süß, M.T. Verticillium dahliae Vta3 promotes ELV1 virulence factor gene expression in xylem sap, but tames Mtf1-mediated late stages of fungus-plant interactions and microsclerotia formation. PLoS Pathogens 2023, 19, e1011100. [Google Scholar] [CrossRef]
- Wang, Q.; Pan, G.; Wang, X.; Sun, Z.; Guo, H.; Su, X.; Cheng, H. Host-induced gene silencing of the Verticillium dahliae thiamine transporter protein gene (VdThit) confers resistance to Verticillium wilt in cotton. Journal of Integrative Agriculture 2024, 23, 3358–3369. [Google Scholar] [CrossRef]
- Tao, Y.; Liu, T.; Cai, L.; You, C.; Lin, H.; Guo, Y.; Dai, T. Controlling verticillium wilt in Acer mollo maxim with a star polycation dsRNA delivery system targeting Verticillium dahliae VDAG_03333. Journal of Cleaner Production 2025, 492, 144827. [Google Scholar] [CrossRef]



| Media | Constituents for 1 L of medium | References |
| Ethanol agar | Ethanol (0.5 mL), streptomycin (100 mg), agar (7.5 g) | [90] |
| Modified ethanol agar | Ethanol (5 mL), sucrose (7.5 g), KCl (0.5 g), K2HPO4 (1 g), NaNO3 (2 g), MgSO2.7H2O (0.5 g), FeSO4. 7H2O (0.01 g), streptomycin (100 mg), chloramphenicol (250 mg), agar (20 g) | [91] |
| Ethanol potassium amoxicillin agar (EPAA) | Ethanol (2 mL), KH2PO4 (1.5 g), K2HPO4 (4 g), amoxicillin (50 mg), agar (15 g) | [92] |
| Soil extract agar | Soil extract1 (25 mL), KH2PO4 (1.5 g), K2HPO4 (4 g), chloramphenicol (50 mg), streptomycin (50 mg), chlorotetracycline (50 mg), agar (15 g) | [93] |
| Modified soil extract agar | Soil extract agar medium, polygalacturonic acid (5 g), ethanol (0.5 mL), tergitol (0.5 g) | [95] |
| Sorbose agar | Sorbose (2 g), streptomycin (0.1 g), agar (10 g) | [96] |
| Sorbose asparagine agar | L-sorbose (2 g), L-asparagine (2 g), K2HPO4 (1 g), MgSO2.7H2O (0.5 g), quintozene (1 g), NaB4O7.10H2O (0.3 g), KCL (0.5 g), Fe-Na-EDTA (0.01 g), oxgall (0.5 g), agar (20 g) | [97] |
| Sorensen’s NP-10 | Tergitol (0.5 mg), KH2PO4 (1 g), KNO3 (0.5 g), MgSO2.7H2O (0.5 g), NaOH (0.025 N), polygalacturonic acid2 (5 g), chloramphenicol (50 mg), streptomycin (50 mg), chlorotetracycline (50 mg), agar (15 g) | [94] |
| Primers | Sequences 5’-> 3’ | Detection assays | Targets1 | References |
| VMSP1 | CATAAAAGACTGCCTAC GCCG | Simplex | V. dahliae | [107] |
| VMSP2 | AAGGGTACTCAAACGGTCAG | |||
| DB19 | CGGTGACATAATACTGAGAG | Simplex | V. dahliae | [108] |
| DB22 | GACGATGCGGATTGAACGAA | |||
| Vd-F | GGGTAGGCTGGCCATATGTG | Simplex | V. dahliae | [104] |
| Vd-R | GTTCTATCCATCGCGGAAAC | |||
| NDf | ATCAGGGGATACTGGTACGAGA | Simplex | ND pathotype | [109] |
| NDr | GAGTATTGCCGATAAGAACATG | |||
| INTND2f | CTCTTCGTACATGGCCATAGATGTGC | Nested | ND pathotype | [109] |
| INTND2r | CAATGACAATGTCCTGGGTGTGCCA | |||
| D1 | CATGTTGCTCTGTTGACTGG | Simplex | D pathotype | [110] |
| D2 | GACACGGTATCTTTGCTGAA | |||
| INTD2f | ACTGGGTATGGATGGCTTTCAGGACT | Nested | D pathotype | [111] |
| INTD2r | TCTCGACTATTGGAAAATCCAGCGAC | |||
| espdef01 | TGAGACTCGGCTGCCACAC | Duplex with DB19/DB22 | D and ND pathotypes | [100] |
| NEP_InPC4 | GGGACTGGGACAGGATGGACA | Duplex | D and ND pathotypes | [106] |
| Dnep286_R | CAAGACCAAATTCGACAGGCAGAG | |||
| NDnep482_R | CCTATTACGAGGTACTTACGGGGACTCTT | |||
| VdUni-F | TCCTAGGCAGGCGAGCAG | qPCR | V. dahliae in soil | [112] |
| VdUni-R | TAGGGCTGTCTGTCGGTGA | |||
| VertBt-F | AAC AAC AGT CCG ATG GAT AAT TC | qPCR | V. dahliae in plant tissue | [98] |
| VertBt-R | GTA CCG GGC TCG AGA TCG | |||
| VTP1-2F | CTC GAT CGT CGT CAA CC | qPCR | V. dahliae in plant tissue | [102] |
| VTP1-2R | TGG TGG TGA GAG TGT TG | |||
| FIP | CGTGATGCTCCGTTTAGGTGGATTTTCGCCATGTTCGGTGCTAG | LAMP | V. dahliae in soil | [101] |
| BIP | TGGCACGTGTGGCGTAAGACTTTTCGATGTCGAGTCTGACACTG | |||
| F3 | TGGCAGCTTCTGATTCAGTT | |||
| B3 | ACAGCGATTTGGATTCCCTC | |||
| LF | CTGACAACCAACGTCTAGATCTCA | |||
| LB | GGCTATTGAGTTCTGCACTCTGTC | |||
| RPA-F | CTTCATTGAGACCAAGTACGCTGTAAGTAACC | RPA-CRISPR/Cas12 | V. dahliae in soil and plant tissue | [105] |
| RPA-R | CAGTTGTCGTGAAGGGGTCATCTTGACTGC | |||
| crRNA1R | GAACCCCAGCACATGATAGAATCTACACTTAGTAGAAATTA | |||
| ssDNA-FQ | 5′FAM-/TTATTATT/-3′DBQ1 | |||
| VD-FIP | TCTCCGTGGATGTTCTCGGGAATAATGGCTGCCGTGACTGTC | LAMP-CRISPR/Cas12 | V. dahliae in soil | [113] |
| VD-BIP | TAGGGACGCAACAATGAGCTGTGCACGGCGCCAAAGTTC | |||
| VD-F3 | AGCGGAAGGGGCACTAG | |||
| VD-B3 | CAAAGACCACGACCATAGGC | |||
| VD-LF | ACGATTGGCAGTCACGGTT | |||
| VD-LB | TTGACGGCTTTACCACAGTCT | |||
| crRNA | UAAUUUCUACUAAGUGUAGAUCCACAGUCUUCUCGGCCAAGU | |||
| ssDNA-1 | (FAM)CCACGGGAGGAATACCAACCCAGTG(BHQ1) |
| Bio-agents | Isolates | In vitro efficacy | In vivo efficacy (GH and field) | Plant growth promotion | References |
| Bacillus altitudinis | KRS010 | 74-80% | 93.6% (GH) | Yes | [146] |
| B. amyloliquefaciens | YZU-SG146 | 68.2-89.2% | 84.2% (GH) | Yes | [151] |
| B. atrophaeus | YL84 | 72.2-84.1% | 66.7% (GH) | Yes | [152] |
| B. mojavensis | KRS009 | 90.2% | 88.6% (GH) | Yes | [153] |
| B. subtilis | T6 | 63.8% | 92.6% (GH) | Not tested | [154] |
| B. subtilis | KRS015 | 59.2-97.1% | 62% (GH) | Yes | [155] |
| B. subtilis | SM21 | 48.2% | 45.7% (GH) | Not tested | [148] |
| Bacillus cereus | AR156 | 67.3% | 74.3% (GH) | Not tested | [148] |
| Serratia sp. | XY21 | 41.3% | 61% (GH) | Not tested | [148] |
| Consortium | SM21:AR156:XY21 | 77.3% | 86.1% (GH), 43.3-76.9% (field) | yes | [148] |
| Enterobacter cancerogenus | HA02 | Not tested | 72.4% (GH), 45.9% (field) | Yes | [150] |
| Paenibacillus polymyxa | ShX301 | 87% | 40.3-71.1% (GH) | Yes | [156] |
| Penicillium simplicissimum | CEF-818 | Not tested | 41.5-60.7% (GH), 62.4-69.5% (field) | No | [157] |
| Pseudomonas spp. | FP22, FP23, FP30, FP35 | 43.9-56% | 32.5-50% (GH), 22.1-50.9% (field) | Yes | [145] |
| Streptomyces kanamyceticu | B-49 | 68.9-85.6% | 65.8% GH), 28.4% (field) |
No | [147] |
| Acremonium sp. | CEF-193 | Not tested | 52.4-47% (GH), 38.5-54.6% (field) | No | [157] |
| F. oxysporum | By125 | Not tested | 69% (GH) | Yes | [158] |
| F. solani | CEF559 | 75-80% | 60% (GH), 30.1-56.3% (field) |
No | [159] |
| Gibellulopsis nigrescens | CVn-WHg | Not tested | 40.6-95% (GH) | No | [149] |
| Leptosphaeria sp. | CEF-714 | Not tested | 47.6-58.2% (GH), 50.2-69.2% (field) | No | [157] |
| Nectria haematococca | Bx247 | Not tested | 69.8% (GH) | Yes | [158] |
| Phomopsis sp. | By231 | Not tested | 63.4% (GH) | Yes | [158] |
| Talaromyces flavus | CEF-642 | Not tested | 29.5-26% (GH), 23.2-45.7% (field) | No | [157] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




