1. Introduction
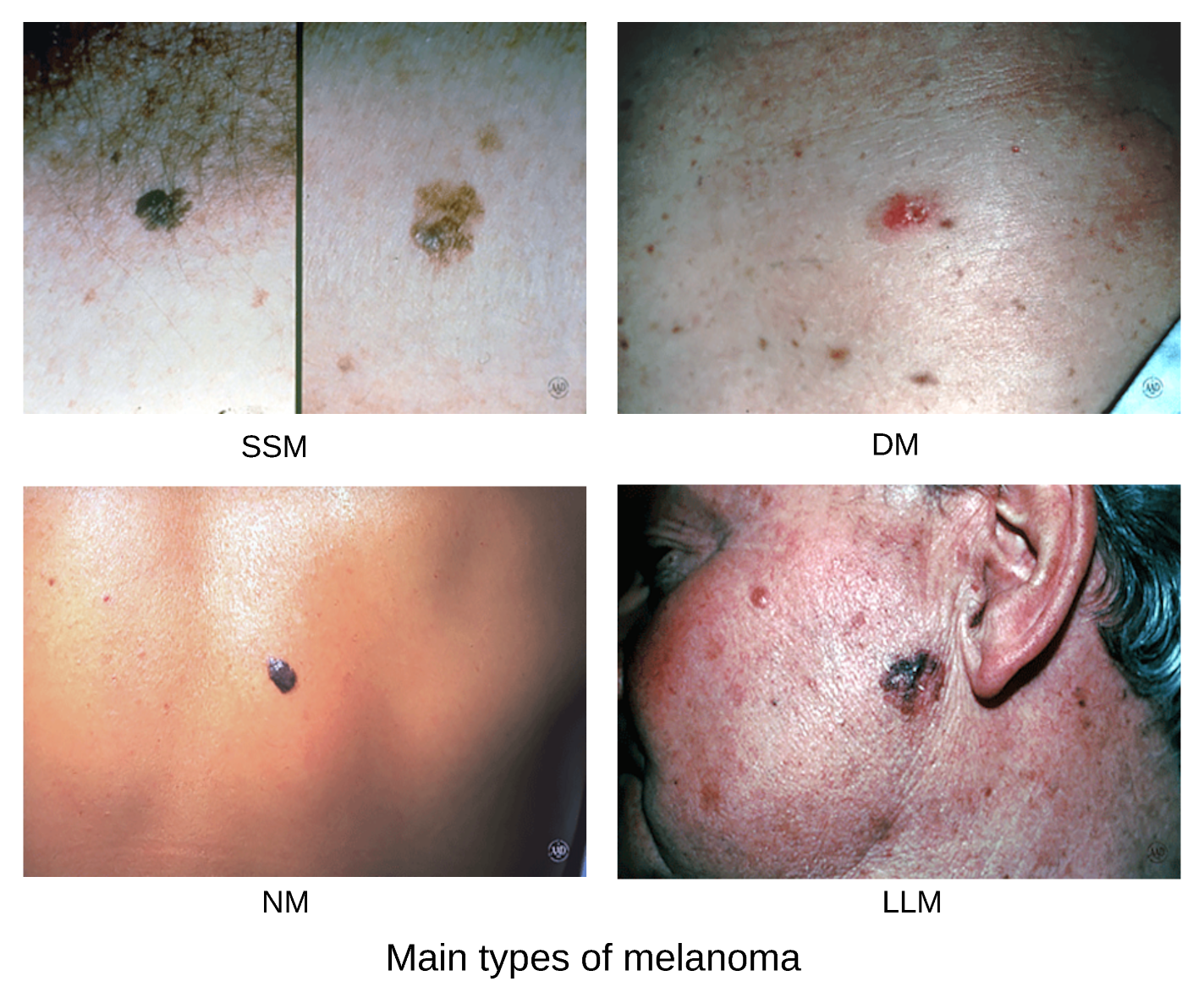
Melanoma is an aggressive malignant tumor that originates from melanocytic cells, which are primarily located in the skin, but can also be found in the mucosa, uvea, and meningeal membranes. The development of melanoma is influenced by several key risk factors, including a history of excessive sun exposure and sunburns, Fitzpatrick skin types I and II, and the presence of atypical nevi. There are several subtypes of melanoma as shown on
Figure 1. Superficial spreading melanoma (SSM) is the most common, accounting for approximately 70% of cutaneous melanomas. Nodular melanoma (NM) represents about 15-20%, while lentigo maligna melanoma (LMM) accounts for 5-10%. Other less common subtypes include acral lentiginous melanoma (ALM), amelanotic melanoma, desmoplastic melanoma (DM), mucosal melanoma, and uveal melanoma. [
1,
2,
3]
According to epidemiological data, the incidence rate of melanoma in the United States was reported at 21.2 cases per 100,000 individuals in 2022 [
4]. Projections from the American Cancer Society estimate that 104,960 new cases will be diagnosed in 2025 [
5]. Despite significant advancements in therapeutic approaches, melanoma continues to exhibit a mortality rate of approximately 8% [
6,
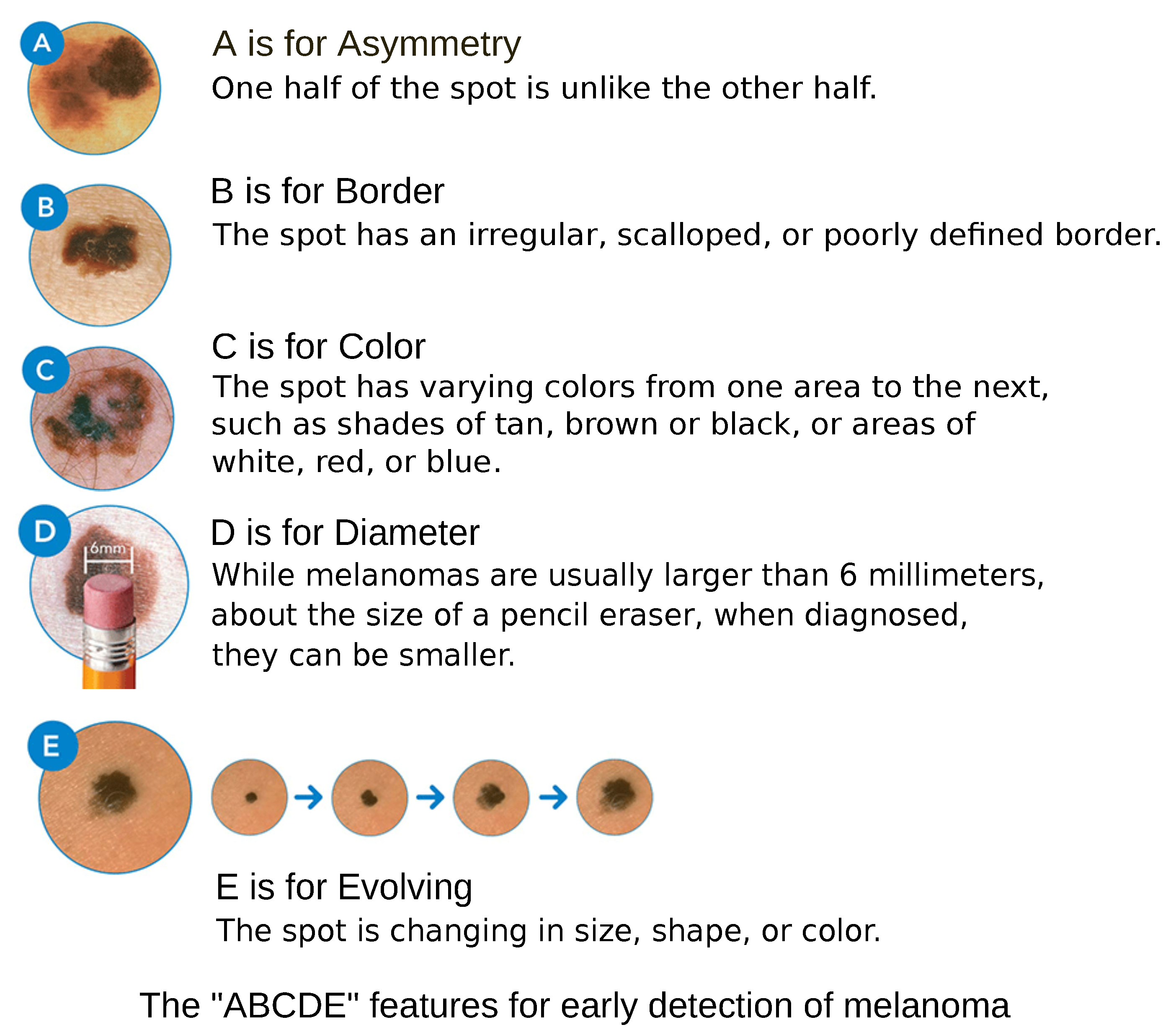
7]. The clinical diagnosis of melanoma relies on the assessment of lesion characteristics based on the ABCDE criteria, as depicted in
Figure 2, which evaluate asymmetry, border irregularity, color variation, diameter greater than 6 mm, and evolution over time [
8]. Other than the diameter, the ABC criteria are subject to subjective interpretation. Furthermore, physicians with more clinical experience tend to demonstrate higher diagnostic accuracy [
9].
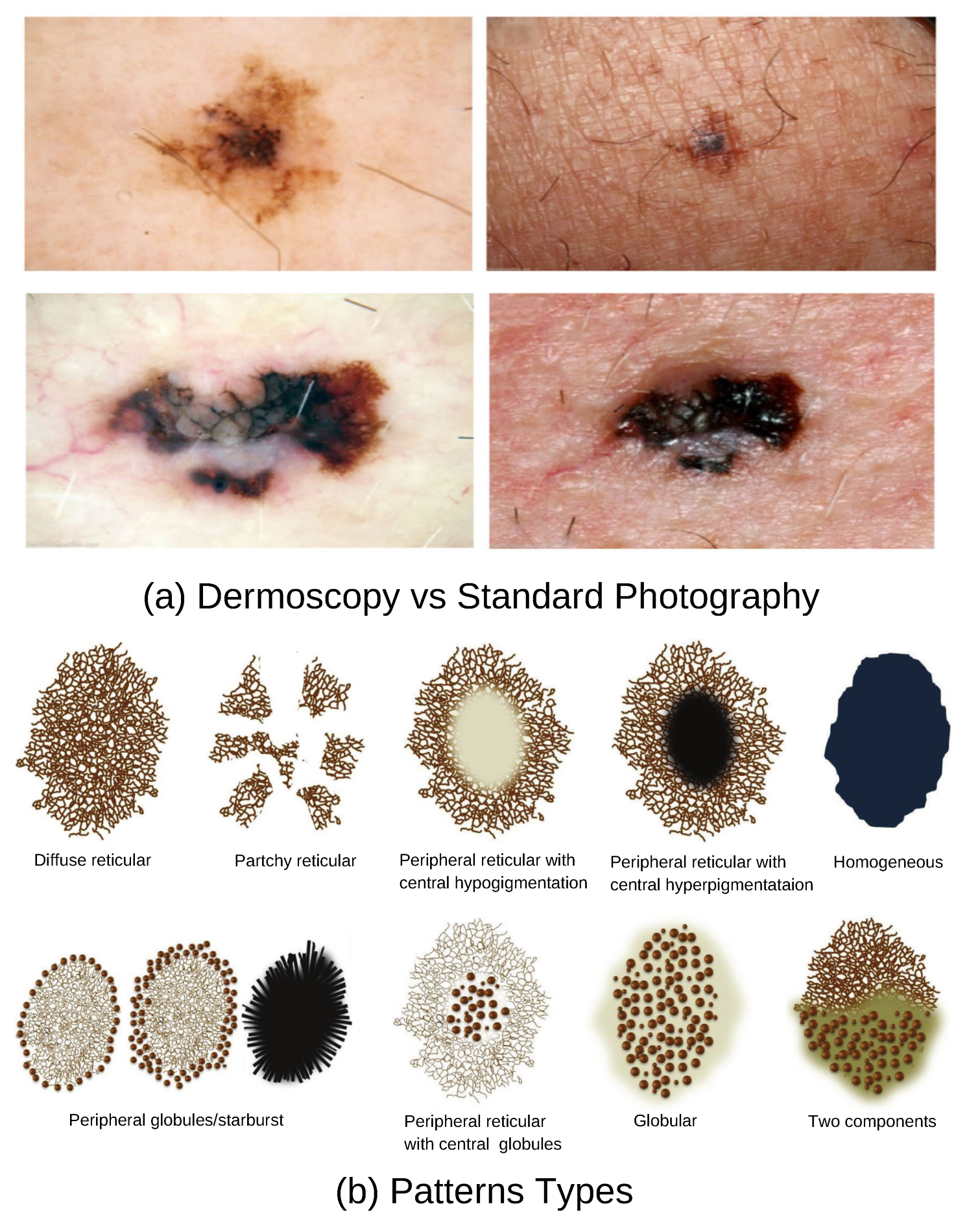
Dermoscopy is a non-invasive diagnostic technique widely used by general practitioners (GP) and dermatologists to improve the detection of skin lesions suspected of malignancy. A dermatoscope, in simple terms, is a magnifying lens equipped with illumination and, in many cases, digital imaging capabilities. It facilitates the visualization of subsurface skin structures that are not visible to the unaided eye as shown in part (a) of
Figure 3. Studies have demonstrated that the incorporation of dermoscopy into the diagnostic workflow enhances accuracy; it increases accuracy from 54% to between 76% and 79%, compared to visual inspection alone [
10]. Dermoscopy captures unique and mixed basic structural patterns such as reticular, streaked, globular, and homogeneous, as illustrated in part (b) of
Figure 3. These structures may be distributed centrally, peripherally, or may be diffused across the lesion, and some of these patterns are observed exclusively in malignant lesions. Additionally, dermoscopy can reveal vascular patterns, which provide further diagnostic value. Although some dermoscopic patterns are clearly identifiable, more complex ones pose a significant challenge, especially for primary care physicians [
11]. This also introduces subjectivity into the diagnostic process as pattern identification relies on clinical experience.
Scoring systems, such as the Total Dermoscopy Score (TDS) and the 7-Point Checklist (7PCL), incorporate these features to enhance the diagnostic accuracy of melanoma detection. [
12] These systems integrate asymmetry, border irregularity, color variation, and diameter to provide a comprehensive risk assessment.
Table 1: outlines the procedures for calculating the most common scoring systems.
Specifically, TDS classifies lesions as follows:
TDS ≤ 4.75: Benign lesion
TDS between 4.8 and 5.45: Suspicious lesion
TDS ≥ 5.45: Malignant lesion
TDS and 7PCL are valuable tools in melanoma diagnosis, however, their reliance on subjective assessments can lead to variability in interpretations. [
13]
For GP, the initial step involves distinguishing melanocytic lesions from non-melanocytic ones through clinical evaluation to ensure accurate preliminary classification. The second step focuses on differentiating nevi from melanoma by applying clinical criteria such as the ABCDE rules, interpreting dermoscopic findings, and utilizing scoring systems. Suspicious lesions are typically referred to a specialist (dermatologist) for further assessment. We have described how human intelligence, through visual inspection, dermoscopic tools, and scoring systems, is used to determine whether a lesion is benign or malignant. Nonetheless, these evaluation methods rely heavily on human judgment, whether through clinical assessment, the use of tools such as dermoscopy, or the application of clinician-derived scoring systems, making them inherently prone to subjectivity.
The diagnostic subjectivity and inter-observer variability associated with melanoma underscore the need for automated and objective computer-assisted diagnosis (CAD) systems. Two primary approaches have been developed for melanoma classification using CAD.
The first approach aligns with the clinical methodology and employs scoring systems based on the ABCD rule. To ensure objectivity, automated methods have been proposed for detecting shape asymmetry, color variegation, and lesion diameter. Among these, Fourier Descriptors are considered the most robust techniques for quantifying asymmetry due to their mathematically rigorous formulation and demonstrated alignment with expert dermatological assessments, achieving up to 92% concordance with dermatologist evaluations [
14,
15]. For border irregularity, combined shape descriptors including fractal dimension, Zernike moments, and convexity coupled with convolutional neural network (CNN) classifiers have achieved state-of-the-art results, with reported classification accuracies reaching 93.6% [
16]. In assessing color variegation, the CIELAB color space combined with the Minkowski distance metric has proven the most effective, offering perceptual uniformity that closely matches human color vision and handling wide color variations reliably [
17]. Regarding lesion diameter, Feret’s diameter has been identified as the most accurate and consistent measure, particularly for lesions with irregular boundaries [
17].
The second approach focuses on direct skin lesion classification, which can be performed using either full images or segmented lesion regions. Studies have shown that classifiers trained on segmented images generally outperform those trained on whole images [
18]. In classical CAD systems, lesion segmentation is typically performed using three main approaches: edge-based, region-based, and threshold-based methods [
19]. For classification, methods such as score averaging (AVGSC), linear SVM, and non-linear SVM using a histogram intersection kernel have demonstrated acceptable performance levels [
20].
The advent of deep learning has significantly enhanced both segmentation and classification accuracy by leveraging CNNs and transformer-based architectures [
21]. Deep supervised learning has led to the development of increasingly sophisticated architectures beyond traditional CNNs.
Popular CNN-based models for melanoma classification include ResNet [
22], VGG16 [
23], MobileNet [
24], DenseNet [
24] and Inception [
25]. Transformer-based architectures have also demonstrated strong performance [
26,
27,
28]. Reported classification performance across these models varies, with accuracy ranging from 80% to 98%, sensitivity from 60% to 90%, and specificity from 86% to 98%. Among these, EfficientNet-B7 [
29] and transformer-based models [
26,
27,
28] exhibit the highest diagnostic accuracy.
Models based solely on CNNs have been increasingly outperformed by architectures incorporating encoder-decoder frameworks, reflecting a shift towards more effective hierarchical feature representations. CNN-based segmentation models fall into two primary categories: pixel-wise upsampling techniques, such as fully convolutional networks (FCNs), and spatial and pyramidal upsampling approaches, such as the DeepLab family of models [
30]. Notably, DeepLabV3+ introduces a decoder module designed to enhance segmentation accuracy and refine feature extraction, particularly at object boundaries [
31]. Encoder-decoder architecture like U-Net and V-Net utilize symmetric upsampling at each layer to preserve spatial resolution during reconstruction. This improves the fidelity of the segmented output [
32] [
33,
34]. Further advancements have been realized through attention-based models, which improve both feature selection and contextual awareness. Prominent architectures in this category include the Vision Transformer (ViT) [
35,
36] and Visual Attention Networks [
37], which leverage self-attention mechanisms to capture long-range dependencies within an image. The segmentation accuracy of deep learning models for skin lesion analysis ranges from 80% to 98%, with recent architectures consistently surpassing their predecessors in both precision and generalization capacity.
Accurate lesion segmentation is essential for the reliable extraction of ABCD features and for effective classification. In most publicly available datasets, lesion boundaries are manually annotated by dermatologists, residents, medical students, or trained personnel. This process is not only time-consuming but also inherently subjective. For lesions with fuzzy or ambiguous boundaries, substantial inter-annotator variability exists. A study by Kittler et al. reported discrepancies of up to 20-30% due to ambiguous lesion edges and variations in annotator expertise [
12,
38,
39]. To address this, some organizations provide standardized segmentation protocols. For instance, the ISIC archive employs multi-rater consensus in cases with ambiguous lesion boundaries [
20]. These practices underscore the degree of subjectivity that remains embedded in CAD systems dependent on manual segmentation for classification or ABCD rule application.
A comprehensive meta-analysis conducted within the medical community evaluated the effectiveness of AI models in skin cancer detection. Following the PRISMA methodology, the review included 272 publications from an initial pool of 14,224 studies, spanning the years 2000 to 2021. The analysis reported a mean F1 score of 0.807 for melanoma detection, with scores ranging from 0.732 to 0.882 [
40].
In summary, we have presented insights from both computer science and medical perspectives. Despite significant advancements in deep learning based melanoma detection, current AI methods remain insufficient for widespread clinical adoption. There exists a discrepancy in reported accuracy results between the medical and engineering communities, which warrants further investigation, particularly with respect to melanoma subtypes, patient demographics, image quality, and dataset provenance. Additionally, it is important to explore the bias introduced by relying on annotated lesion borders, especially in cases involving irregular or ambiguous boundaries.
1.1. Contributions
Although AI models have significantly advanced in recent years, delineating lesions remains a subjective task even for specialists. We propose that the lesion border should not be treated as a single, well-defined line, but rather as a transitional region that separates the background from the core of the lesion. Part (b) of
Figure 5 presents a blended image composed of the ground truth mask overlaid on the original lesion. In our work, the border region is defined as the area between the outer edge of the mask and the visible boundary of the lesion. This region can be more precisely delineated using superpixels, which provide spatially coherent segmentation aligned with local image features. Our objective is to develop a model capable of accurately identifying all three regions. In this paper, we propose a new method called SPADE (Superpixel Adjacency Driven Embedding) for three class melanoma segmentation. The main contributions of SPADE are as follows.
Definition of Three Anatomical Zones: We define three distinct zones in dermoscopy images, the background, the border, and the core of the lesion.
Transformer-based Superpixel Embedding: We train a transformer autoencoder to generate embeddings for each superpixel, enabling the model to capture contextual relationships between spatially distant regions belonging to the same class.
Context-Aware Region Adjacency Graph (RAG): We construct a Region Adjacency Graph using superpixels obtained via the SLIC algorithm to model spatial relationships.
Input Definition Based on the Region Adjacency Graph: Each input vector consists of the embedding of a given superpixel along with the embeddings of its immediate neighbors, effectively capturing local spatial context.
Semantic Representation of Dermoscopy Images: The system effectively captures the semantic content of dermoscopy images, supporting improved lesion characterization.
1.2. Related Work
Fine-tuning AI models for accurate border delineation in blurry or low-contrast dermoscopic images has been addressed through methods that identify and label regions adjacent to lesions. Adegun et al. employ probabilistic models in which each pixel is assigned a value between 0 and 1, representing the likelihood of belonging to the lesion [
41]. The algorithm ultimately produces a lesion boundary represented as a line. Halil et al. utilize region of interest (ROI) bounding, where an initial bounding box or elliptical region is drawn around the lesion, followed by consensus among multiple annotators [
42]. Although the initial stage localizes the lesion broadly, the final boundary is refined using the GrabCut algorithm, resulting in a sharply delimited line. Zahra et al. propose an approach based on multiple expert-generated masks, from which a consensus mask is derived through fusion [
43]. The final output is again a single mask outlined by a distinct boundary line. Li et al. [
44,
45] introduce a method that explicitly marks regions of uncertainty rather than enforcing a hard boundary. They use class activation maps to generate pseudo-labels and apply binary thresholding to create segmentation masks. This approach allows the model to learn from ambiguous regions, thereby improving robustness in uncertain contexts. In contrast to these methods, which ultimately produce a line-based lesion boundary, our approach focuses on defining lesion regions, providing a more granular and context-aware representation.
Graph-based methods also commonly leverage superpixels by constructing a RAG, where each node represents a superpixel and edges encode similarity metrics between neighboring regions such as color, texture, or boundary strength. Once the graph is constructed, segmentation is performed using algorithms such as spectral clustering, normalized cuts, or min-cut/max-flow, and neural networks, which partition the graph into semantically meaningful regions.[
46]. RAGs naturally integrate with Graph Neural Networks (GNNs) and have been employed for image classification tasks. Nazir et al. applied Graph Convolutional Neural Networks (GCNNs) using both spatial and spectral convolution techniques, demonstrating that spectral-based models outperform spatial-based models and classical CNNs while requiring less computational cost [
47]. Avelar et al. explored attention-based GNNs for classification and showed that the feature space can be enhanced by weighting the edges of a superpixel graph using a learned function based solely on geometric information [
46]. Nowosad et al. apply graph-based segmentation to non-imagery geospatial data by grouping superpixels based on dissimilarity measures and pruning the resulting graph to a Minimum Spanning Tree (MST) [
48]. Additionally, Qin et al. propose a superpixel-based and boundary-sensitive CNN model specifically designed for liver segmentation, demonstrating the effectiveness of combining region-based pre-processing with deep learning techniques [
49]. Similar to our work, these studies employed superpixels and RAG to construct graph representations. However, unlike prior work, the current phase of our study does not incorporate graph neural networks or edge attributes for classification.
2. Methods and Algorithm Design
The SPADE pipeline proceeds as follows: the images are first resized; superpixels are then generated using the Simple Linear Iterative Clustering (SLIC) algorithm; a region adjacency graph is constructed; each superpixel is labeled by class (background, border, lesion); class-specific embeddings are generated using a transformer; and finally, a transformer is trained to predict the three classes.
2.1. Resizing Images
All images were initially adjusted to a landscape orientation and resized to a width of 1024 pixels and a height of 768 pixels using a bilinear interpolation algorithm from the OpenCV framework.
2.2. Superpixel Generation Strategy
Superpixel segmentation partitions an image into regions composed of similar and spatially connected pixels providing perceptually meaningful atomic regions.
SLIC is classified as a neighborhood-based clustering algorithm. SLIC is an adaptation of the k-means clustering algorithm, specifically designed for efficient superpixel generation. It introduces two key parameters: a search radius that limits the number of distance computations, and a weighted distance metric that balances color similarity and spatial proximity
S. The distance function is defined as:
where
is the Euclidean distance in the RGB color space,
is the geometric distance between pixels, and
m is a compactness parameter that controls the trade-off between color and spatial proximity. These parameters enable control over the size and compactness of the resulting superpixels. Furthermore, SLIC’s linear computational complexity contributes to its widespread adoption in superpixel generation tasks [
50].
Each pixel in SLIC is represented by a five-dimensional feature vector consisting of three components for color and two for spatial coordinates
. The algorithm employs the CIELAB color space, which offers a perceptually uniform representation of color, ensuring a more meaningful similarity measure. The segmentation process in SLIC involves three main steps. First, clusters are initialized based on the desired number of superpixels. Second, an iterative clustering process assigns pixels to the nearest cluster using a distance function that incorporates both color and spatial information. Finally, a post-processing step enforces spatial connectivity to ensure that each superpixel forms a contiguos region [
50,
51].
SLIC superpixels effectively capture low-level image features like color, position, and depth, while preserving fine-grained boundary details. This is essential for detecting subtle background transitions around lesions, which clinicians often rely on. Moreover, superpixels reduce noise by averaging local pixel variations within each region and help focus computational resources on meaningful structures, since larger segments often carry more semantic information than individual pixels.
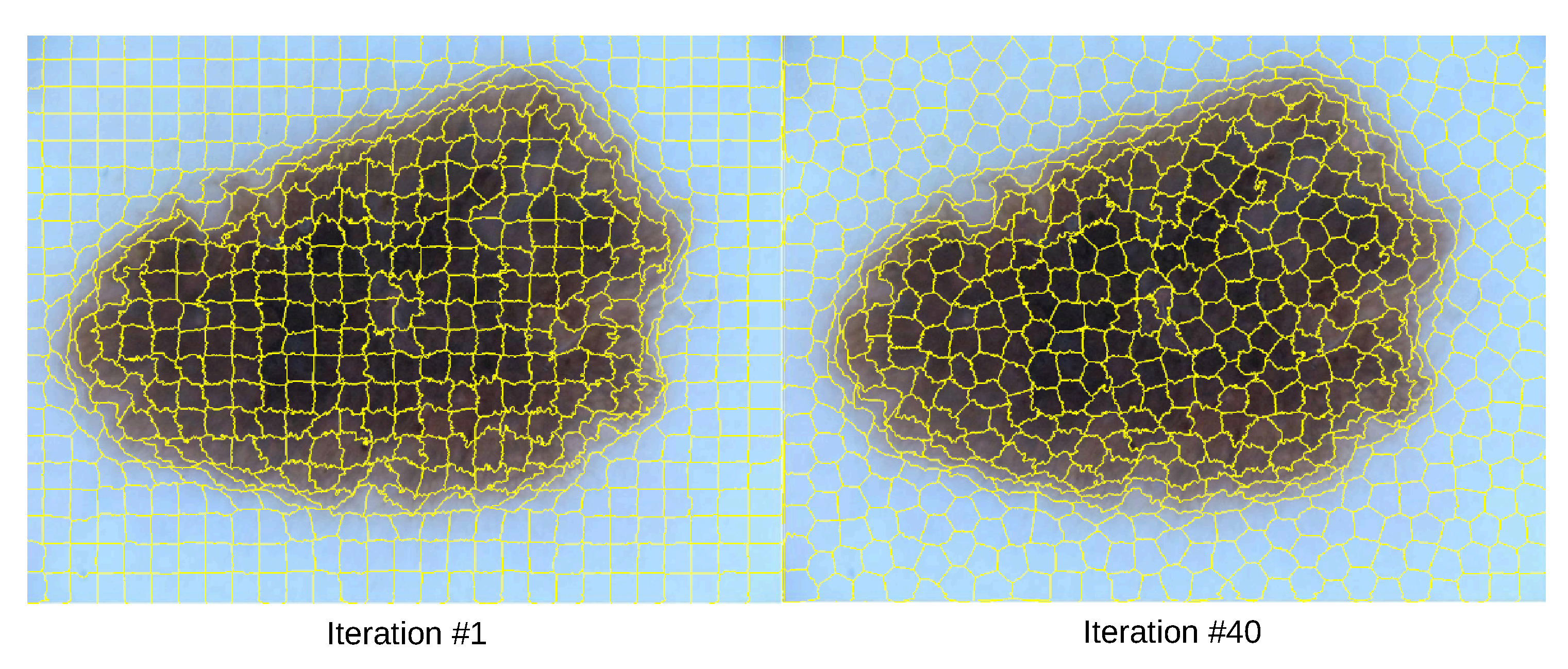
The SLIC algorithm was applied to each image to obtain its segmented regions. Specifically, we employed the SuperpixelSLIC algorithm from OpenCV with the following parameters: a desired number of superpixels
k set to 600, a area size of 1320,
, and a ruler value(
m) of 19220 producing the segmentations shown in
Figure 4. The software suite provides three different algorithms, SLIC, SLICO, and MSLIC, among which SLIC was chosen due to its balance between algorithmic compactness, boundary recall, and computational efficiency. Each superpixel was assigned a unique identifier, which was used to track and analyze the segmented regions [
52]. At the beginning of the algorithm, each superpixel is assigned a unique identifier (ID) when the initial centroids are defined. This ID is stored as an integer value in the database and remains constant throughout the duration of the experiment.
2.3. Constructing the Region Adjacency Graph (RAG)
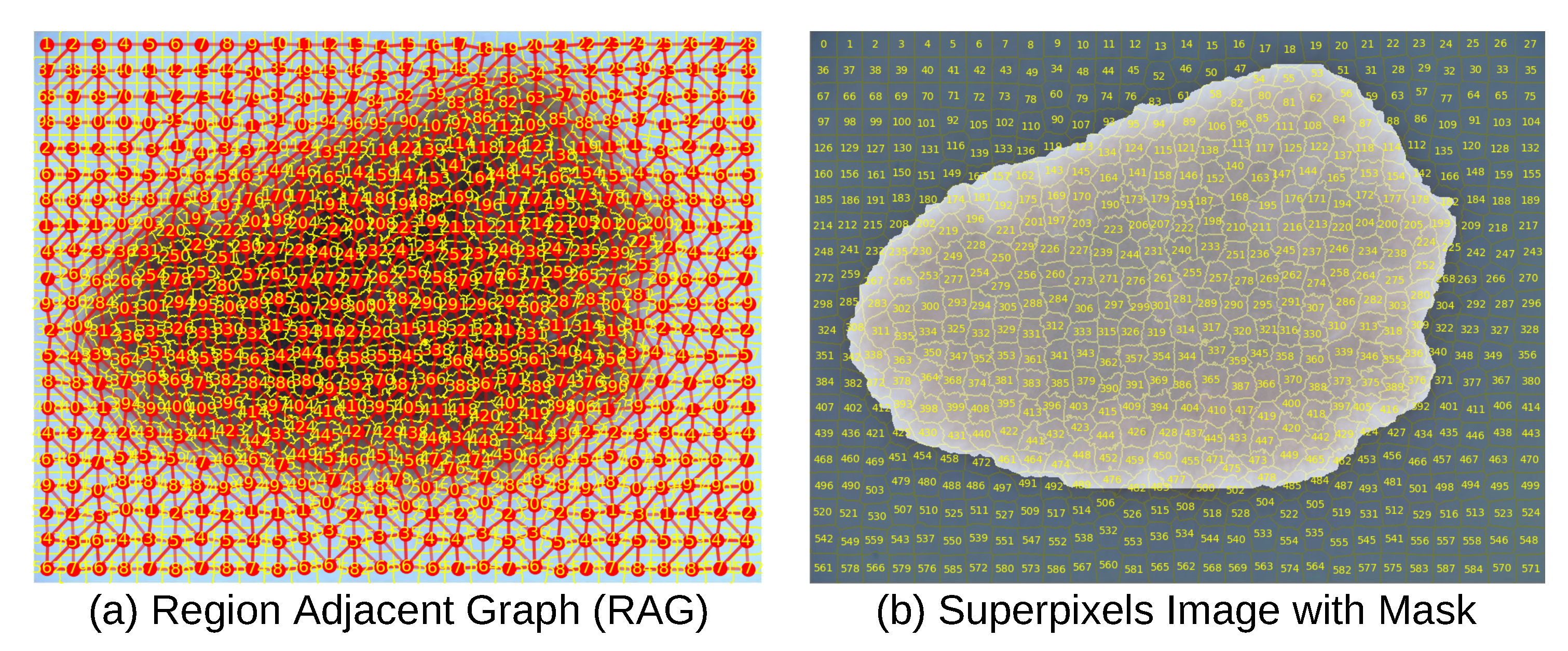
The RAG was constructed by traversing the labeled superpixel matrix and establishing undirected edges between adjacent superpixels with differing labels based on 4-and 8-connected neighborhood criteria, capturing spatial connectivity through horizontal, vertical, and diagonal adjacency as depicted in part (a) of
Figure 5 [
53].
Figure 5.
Dermoscopy Image, Superpixels, RAG, and Mask.
Figure 5.
Dermoscopy Image, Superpixels, RAG, and Mask.
We hypothesize that, similar to how transformers leverage the sequential ordering of words to learn language structure, our Region Adjacency Graph (RAG) explicitly defines "neighbor" relationships among superpixels. Unlike Vision Transformers, where all patches are included as input tokens, our RAG-based approach provides meaningful context to the transformer. In this framework, the transformer’s self-attention mechanism operates over graph nodes, attending preferentially to adjacent regions that share meaningful boundaries or semantic relationships. In contrast to Vision Transformers, which may waste computational resources attending to regions lacking relevant context, our RAG-based model focuses attention on superpixels containing semantically relevant information.
For instance, consider an image with a size pixels, with patches sizes of ; this results in a total of 196 patches. In a Vision Transformer, each transformer layer computes self-attention with a complexity of , where N is the number of patches and D is the embedding dimension. In our superpixel based approach, we observe that each superpixel has, on average, 7 neighbors (ranging from ), leading to a significantly reduced complexity of .
2.4. Creating Features
For each image, we extracted metadata including the filename, height, and width. We also computed the mean and standard deviation for each RGB channel. For each superpixel, we computed the mean, standard deviation, kurtosis, and skewness per RGB channel. Using the segmented region as a binary mask in a 2D image, where pixels belonging to the superpixel were set to 1 and all others to 0, we calculated the area , centroid , centroid , and second central moments using the OpenCV moments function. Additionally, neighboring superpixels were identified and included as part of the superpixel-level information.
To facilitate further analysis, the superpixels were categorized into three classes (background, border, and core lesion) for a subset of 2,000 images from the HAM10000 dataset ("Human Against Machine with 10,000 training images") [
54]. A custom annotation tool was developed to allow manual labeling of superpixels. The supplementary video (Video 1) provides a visual demonstration of how the annotator selects the "border in" and "border out" regions on dermoscopic images. The "border out" class includes superpixels marking the transition between the lesion and the surrounding skin, whereas the "border in" class contains superpixels located just inside the lesion boundary.
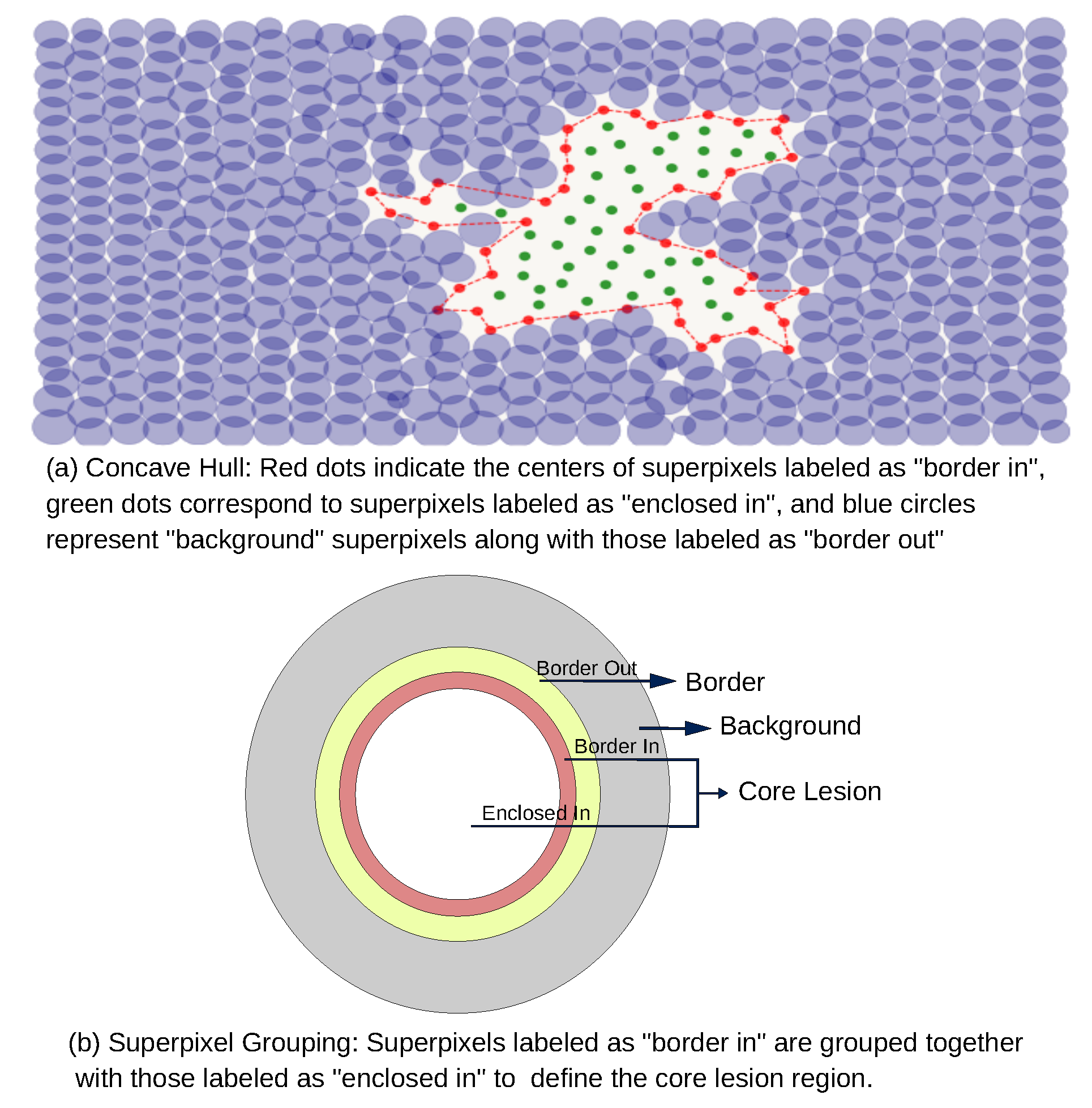
To delineate the lesion interior, a concave hull algorithm was applied to enclose all superpixels within the "border in" region. Unlike the convex hull, which may encompass unrelated background superpixels, the concave hull prevents the inclusion of superpixels that do not belong to the lesion, as illustrated in part (a) of
Figure 6.
Based on the proposed classification scheme, we define the core lesion as the union of superpixels labeled as "border in" and those enclosed by them. The border class corresponds to the "border out" superpixels and all the remaining superpixels are assigned to the background class, as shown in part (b) of
Figure 6
Table 2 presents the total number of superpixels per category, where class labels 0, 1, and 2 correspond to Background, Core Lesion, and Border, respectively.
2.5. Transformer Autoencoders for Generating Embeddings
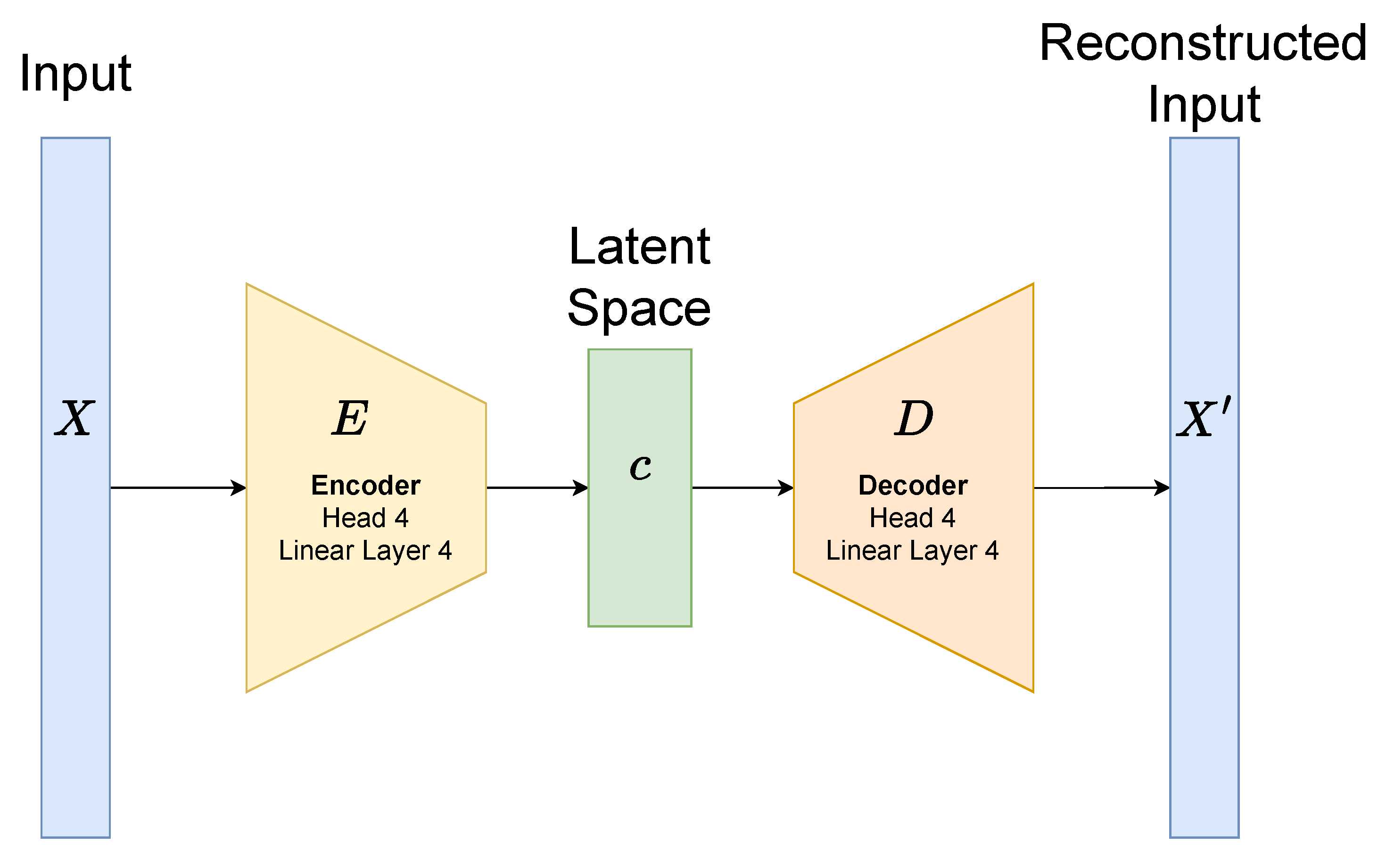
To generate embeddings for each superpixel, we train a transformer autoencoder independently for each semantic class (Background, Border, or Core). Each superpixel is resized to contain 700 pixels per channel, and the latent space dimensionality is set to 256. The model architecture is illustrated in
Figure 7. The encoder consists of four multi-head self-attention layers (with four heads each), interleaved with four feed-forward linear layers. This encoder maps the input patch tokens to a 256-dimensional latent vector. The decoder is symmetric, consisting of four self-attention layers and four linear layers, and reconstructs the original patch from the latent representation. Each attention and feed-forward sub-layer is followed by residual connections and layer normalization to stabilize training and improve gradient flow.
We train a separate autoencoder for each class using only superpixels labeled accordingly. The reconstruction loss is measured using Mean Squared Error (MSE) between the original input and the decoder’s output. Optimization is performed using the Adam optimizer with early stopping, based on an 80/20 training-validation split (random seed = 40).
By leveraging the transformer’s self-attention mechanism, these autoencoders effectively learn to encode irregular, shape-conforming superpixels. This allows them to capture not only intra-region content, but also inter-region semantic patterns—capabilities that fixed-grid patches or traditional CNN encoders cannot achieve as efficiently. We hypothesize that combining these learned embeddings with our Region Adjacency Graph (RAG) structure (Section II.C) enables a context-aware segmentation framework built upon semantically meaningful, spatially grounded building blocks.
The extracted information was stored in a Neo4j graph database [
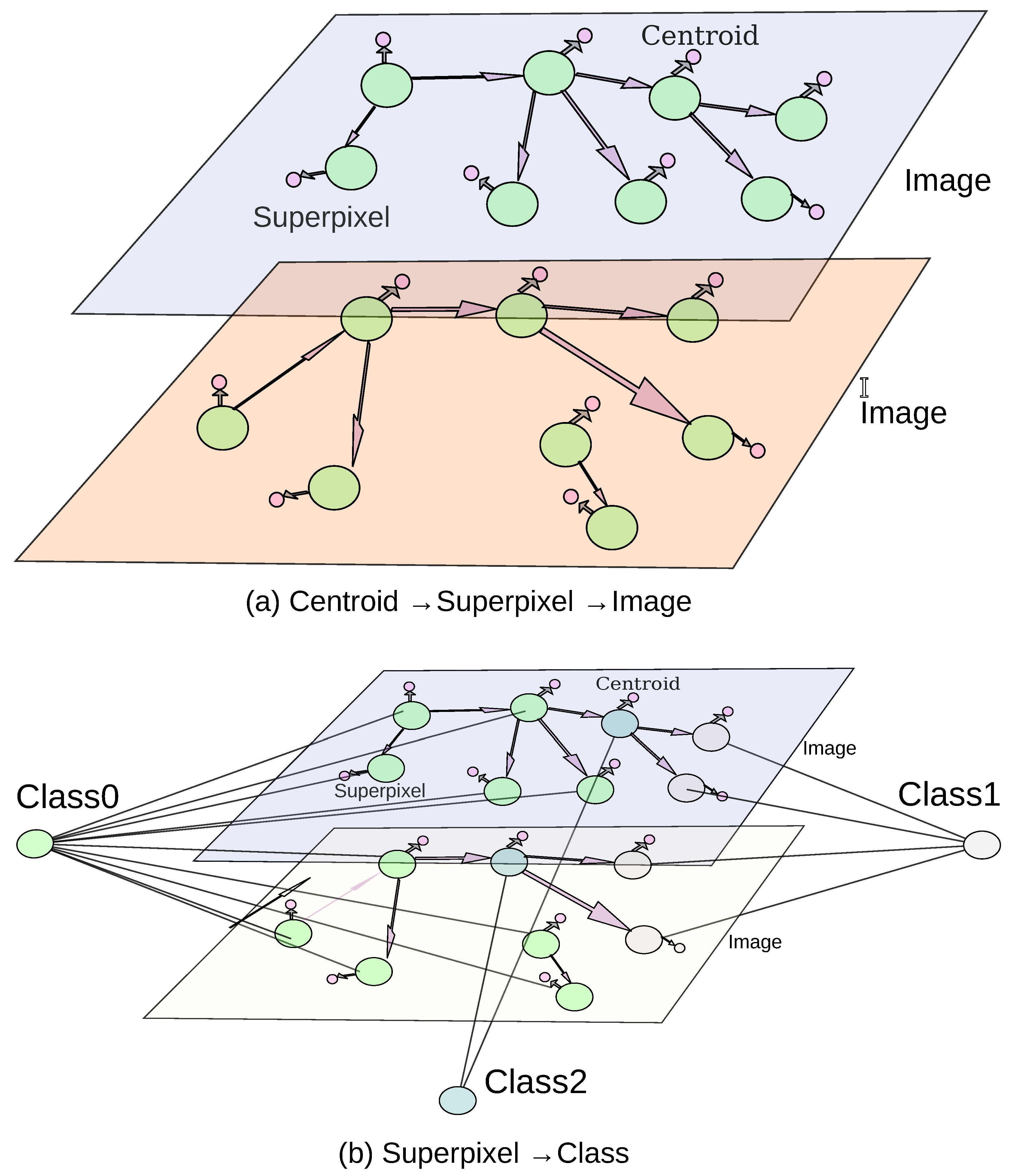
55], where the data was structured into three main entities Images, Superpixels, and Centroids as shown in part (a,b)
Figure 8. The Image entity contained all image-related properties, while the Superpixel entity stored superpixel-specific characteristics including the embeddings generated by the latent space. The Centroid entity recorded the centroid’s
x and
y coordinates. Within the database, IN_IMAGE relationships linked each Image entity to its corresponding Superpixel entities. NEIGHBOR_OF relationships connected Superpixel entities with their neighboring superpixels, such that edge weights represented the Euclidean distance between RGB mean values. Finally, CENTER_AT relationships associated each Superpixel entity with its respective Centroid.
PyTorch version 2.2.2 and CUDA 12 was used to create and run the Deep Learning models for training and inference. The data was normalized and split amongst training and validation sets, 80% and 20% respectively. We set the random seed to 40 for all experiments.
2.6. Algorithm
|
Algorithm 1: Obtain Embeddings from Superpixels and Neighbors |
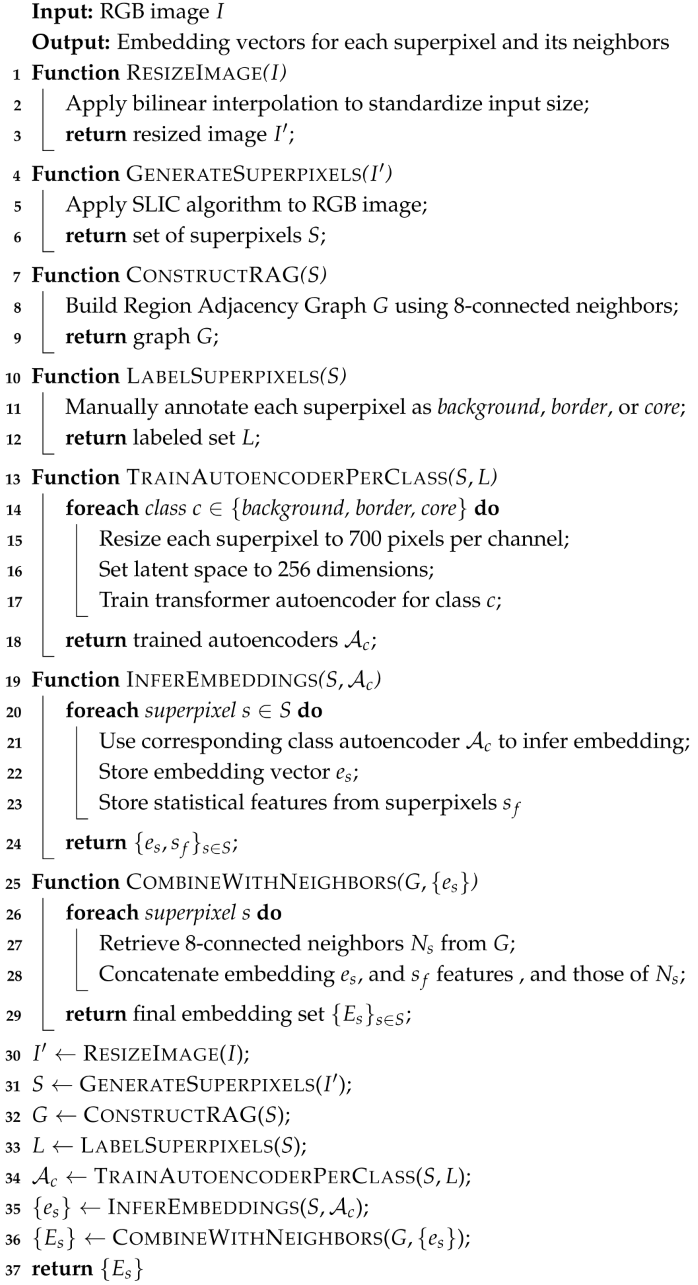 |
3. Results
3.1. Experiments Phase One
HAM10000 dataset contains dermatoscopic images of both malignant and benign pigmented lesions, with diagnoses confirmed through histopathology, dermatologist consensus, and clinical evolution.
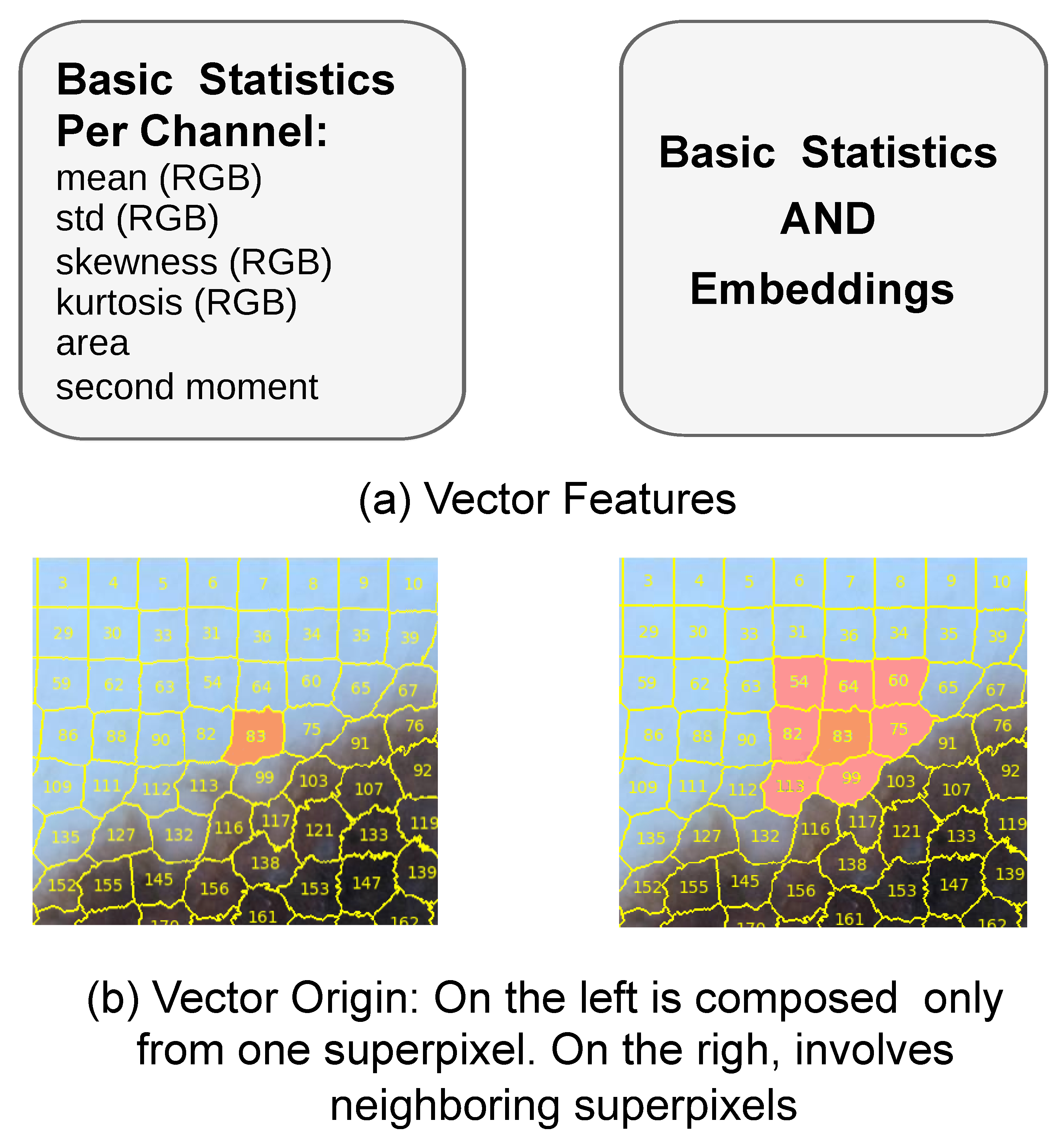
The features can be originated solely from basic statistics extracted from each superpixel, or from a combination of basic statistics and learned embeddings, as illustrated in part (a) of
Figure 9. Additionally, feature vectors were constructed either using information from a single superpixel or from its neighboring superpixels, as shown in part (b) of
Figure 9. In the latter case, the order of concatenation was determined by sorting the superpixels according to their ID numbers in ascending order. The combination of these features and their spatial origin forms the basis for constructing our input vectors.
In the first phase of our experiments, we constructed four distinct types of input-output feature vectors to support different modes of training and inference:
Type I: The input vector consists of basic region descriptors mean, standard deviation, skewness, and kurtosis of the RGB channels; area; centroid; and shape moments extracted from the target superpixel. The output is the class label associated with the target superpixel.
Type II: The input vector includes the basic descriptors of the target superpixel as well as those of its neighboring superpixels. The output remains the class label of the target superpixel.
Type III: Similar to Type II, the input concatenates descriptors of both the target and neighboring superpixels. However, the output vector extends to predict the class label for every superpixel included in the input group, encompassing both the target and its neighbors.
Type IV: This configuration consists of two sets of input-output vectors. The first input vector encodes the statistical descriptors of the neighboring superpixels, with the corresponding output vector specifying their class labels. The second input vector captures the descriptors of the target superpixel alone, with a dedicated output vector indicating its class.
The following neural architectures were used to model these input-output relationships:
Linear Neural Network: A fully connected feed-forward network with three hidden layers of sizes 128, 64, and 32, applied to the Type I and Type II feature vectors.
GRU RNN (Many-to-One): A recurrent neural network utilizing four Gated Recurrent Units (GRUs), applied to the sequential embedding of neighbor features (Type II). The final hidden state is passed to a softmax classifier for target class prediction.
Transformer Encoder–Decoder (Many-to-One): The encoder, composed of four self-attention heads, processes the sequence of target and neighbor superpixels (Type IV). A decoder with four attention heads attends to the encoded context and predicts the target superpixel’s class.
Transformer Encoder (Many-to-Many): A transformer encoder block (with four self-attention heads) jointly processes the concatenated feature set from multiple superpixels (Type III). It outputs class predictions for each region simultaneously.
In our first experiment, we trained a linear Neural Network on the first vector type to predict the class of each superpixel. As shown in
Table 3 the Linear Neural Network performs adequately given the limited feature set, particularly for Classes 0 and 1 with F1 Scores of 0.8333 and 0.8077 respectively. Class 1 exhibits better differentiation from other classes, with the highest accuracy (0.9091) and competitive recall (0.7915). The model performs poorly on class 2 with a F1 Score of 0.2937.
In the second experiment, we trained a linear Neural Network on the second vector type to predict the class of the target superpixel. As depicted in
Table 3 compared to the previous model, the values are higher; however, the behaviour is similar. The model is good enough for class 0 and 1, but under-performs for class 2. Class 1 is better differentiated than the other classes. These results suggest that incorporating neighboring superpixel properties enhances classification especially for class 0, and 1.
In the third experiment, we modeled the data as a sequential input of superpixels in ascending order, and trained a RNN, specifically a GRU, on the second vector type to predict the class of the target superpixel. As illustrated in
Table 3 this model, compared to the previous Linear Neural Networks experiments, performs marginally lower for classes 0 and 1 implying that sequential modeling may not fully exploit the contextual dependencies in the third vector type.
In the fourth experiment, we implemented a many-to-one RNN architecture using a transformer model with encoder-decoder components, trained on the fourth vector type to predict the class of the target superpixel. The encoder processes multiple superpixels (including their class information), while the decoder focuses on the target superpixel to generate its predicted class. As portrayed in
Table 3, Class 0 achieves exceptional performance, with an F1 score of 0.9377, precision of 0.9229, and recall of 0.9529, indicating highly reliable predictions. Class 1 demonstrates strong results (F1: 0.8828, precision: 0.8823) and the highest accuracy (0.9432), suggesting effective utilization of contextual information from neighboring superpixels. Class 2, however, continues to underperform, with an F1 score of 0.3572 and remarkably low recall (0.2976), despite moderate precision (0.4468).
In the fifth experiment, we employed a many-to-many RNN architecture, specifically a transformer model, trained on the third vector type to predict the classes of all superpixels simultaneously. As detailed in
Table 3, Class 0 achieves near-perfect performance, with an F1 score of 0.9816, precision of 0.9753, and recall of 0.9880, demonstrating robust alignment between predictions and ground truth. Class 1 also exhibits strong performance (F1: 0.9196, precision: 0.9199, recall: 0.9193), with accuracy peaking at 0.9812. However, Class 2 remains a challenge, with an F1 score of 0.4816 and low recall (0.4011), despite moderate precision (0.6026). These results underscore the efficacy of many-to-many transformers in leveraging global dependencies for multi-superpixel classification tasks, though Class 2 may require targeted architectural or data-centric interventions.
3.2. Experiment Phase Two
In the second phase of our experiments, we selected the best-performing model from the initial evaluation, the transformer encoder model, as the basis for further analysis. To enhance the feature representation, we expanded the Type III input vectors by incorporating additional embeddings generated by the transformer autoencoders. This augmentation resulted in a total of 270 features per input instance.
The updated feature vectors were trained using a many-to-many transformer encoder architecture, consisting of a single encoder block with four self-attention heads. The model demonstrated perfect classification performance, achieving an accuracy and F1 score of 1.0 across all semantic classes.
To assess the robustness and generalizability of this performance, we conducted a five-fold cross-validation. The model consistently produced the same results across all folds, confirming the stability and reliability of the learned representations.
As observed in the transformer many-to-many encoder model, using input vectors without embeddings yields strong classification performance for class 0 and class 1, with F1 scores of 0.98 and 0.91, and accuracies of 0.96 and 0.97, respectively. These results suggest that simple statistical features are sufficient for accurately identifying superpixels corresponding to the background and core of the lesion. In other words, distinguishing these two classes is not particularly challenging for the model. Therefore, incorporating embeddings naturally aligns with the goal of maintaining or improving performance.
In contrast, classification of class 2 (the lesion border) proves significantly more difficult, with an F1 score of 0.48 despite a high accuracy of 0.97, showing the class imbalance and mis-classification When embeddings are added and the input is expanded to include information from neighboring superpixels, performance improves substantially.
These findings strongly suggest that the border superpixels carry the most significant information in the entire image. Moreover, the border regions may be crucial for distinguishing between benign and malignant lesions.
We acknowledge that our dataset consists of 2,000 images, and it is likely that the variation in lesion morphology across diverse skin types and racial backgrounds is limited, an inherent limitation of the dataset.
4. Discussion
At this stage of our research, achieving high performance in either lesion classification (benign vs. malignant) or segmentation is not our primary objective. For the medical community to adopt AI in clinical settings, it is essential to address several challenges related to data quality, methodological rigor, and most importantly interpretability [
40].
In our study, unlike previous approaches which focus on segmentation or disease classification, the primary role of superpixels is to capture the semantic of the superpixels. Each superpixel is labeled as a background, border, or lesion, corresponding to the classes identified in the image. Superpixels from each class are used to train transformer-based autoencoders in order to produce meaningful embeddings that capture the context of all superpixels of each class regardless of distance between them. We proved that these embeddings capture the semantic content of the classes more effectively than relying solely on the statistical properties of the superpixels.
To enhance classification performance, we utilize not only the embedding of the target superpixel but also incorporate embeddings from its neighboring superpixels as input to the classifier. The transformer captures the context of the near superpixels (spatial locality). We consider our approach to be novel, with the main contribution being a model that learns to interpret the semantic structure of an image through superpixel-based embeddings.
We hypothesize that superpixels provide more natural and semantically meaningful context for the classification transformer compared to the fixed
square patches used in the Vision Transformer (ViT), while maintaining the same number of features [
56].
We establish the originality of our methodology in several key aspects:
Defining the border as a region instead of a well defined line.
Generating embeddings for every superpixel using a transformer autoencoder
Incorporating those embeddings as features for further training
Taking into account the neighborhood to create the input vectors
However, our study has several limitations. These include the dependency on SLIC segmentation quality, the need for manual annotations, and the requirement to train the transformer autoencoder as a separate task for embedding generation.