Highlights
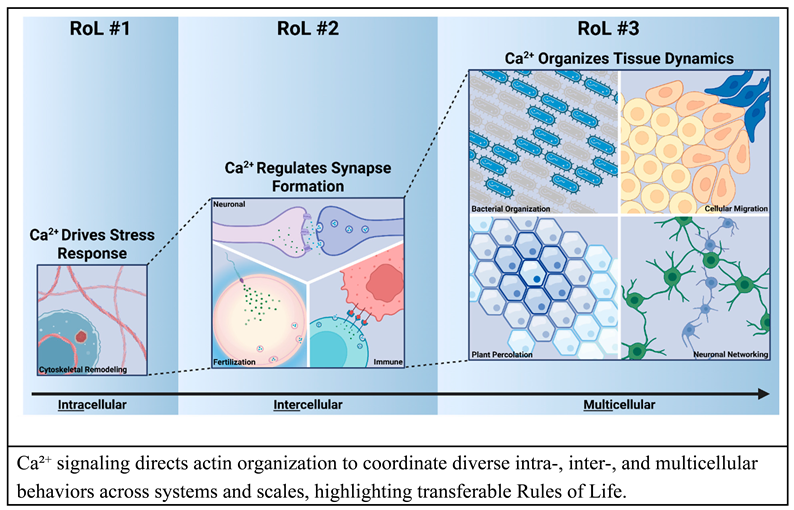
The emergent features of biological robustness, signal integration, and organization are encoded by the dynamics of second messengers such as calcium ions (Ca2+) to regulate cellular processes such as cytoskeletal organization from the cellular to organismal level.
Introduction
Living systems are remarkably adaptive, capable of reorganizing after injury, communicating complex information, and self-organizing for robust performance. Underlying these emergent properties are the modular interactions between mechanical cues and chemical signals, which can be described as “Rules of Life” (RoLs)—principles that define how biological systems behave across contexts and scales. Many such RoLs center on the integration of physical cues with biochemical pathways, and one particular focus is through calcium-mediated regulation of the cytoskeleton [
1,
2,
3,
4,
5,
6]. As ubiquitous second messengers, calcium ions (Ca
2+) link environmental inputs to biological processes and are central for coordinating cellular responses, tissue organization, and organism behavior [
7,
8,
9].
A systems-level understanding of cell and tissue mechanics depends on deciphering the coupling between Ca
2+ signaling and cytoskeletal dynamics. This Ca
2+-cytoskeletal coupling not only reveals how cells process mechanical and chemical inputs, but also further informs the development of computational models for accelerated progress in bioengineered tissues, self-organizing biomaterials, and predictive digital twins [
10,
11,
12,
13,
14]. Such models range from mechanochemical reaction-diffusion systems for signal regulation [
15] to mechanistic models simulating force generation and mechanical behavior [
16]. These frameworks are often built on ordinary and partial differential equations (ODEs/PDEs) and are solved using agent-based finite element simulations [
17,
18]. By integrating with experimental work, these simulations reveal how calcium signaling and cytoskeletal remodeling generate force and direct cellular function across multiple biological systems and spatial scales [
5,
19].
To organize this complexity, we apply the RoLs framework, traditionally used to explain organismal traits (development, function, and environmental response), and distill them into transferable computationally defined rules that map input-output relationships across biological scales [
20,
21,
22]. A major challenge lies in understanding how these RoLs operate across diverse spatial and temporal contexts. Developing predictive, quantitative RoL frameworks allows researchers to simulate how biological systems integrate multiple cues, enabling hypothesis testing, perturbation analysis, and iterative refinement through the integration of modeling and experimentation. Furthermore, multiple RoLs can be combined into a collection of modular computational models that incorporate specific initial and boundary conditions, enabling the generation of hypotheses that are testable through quantitative experimental methods [
22].
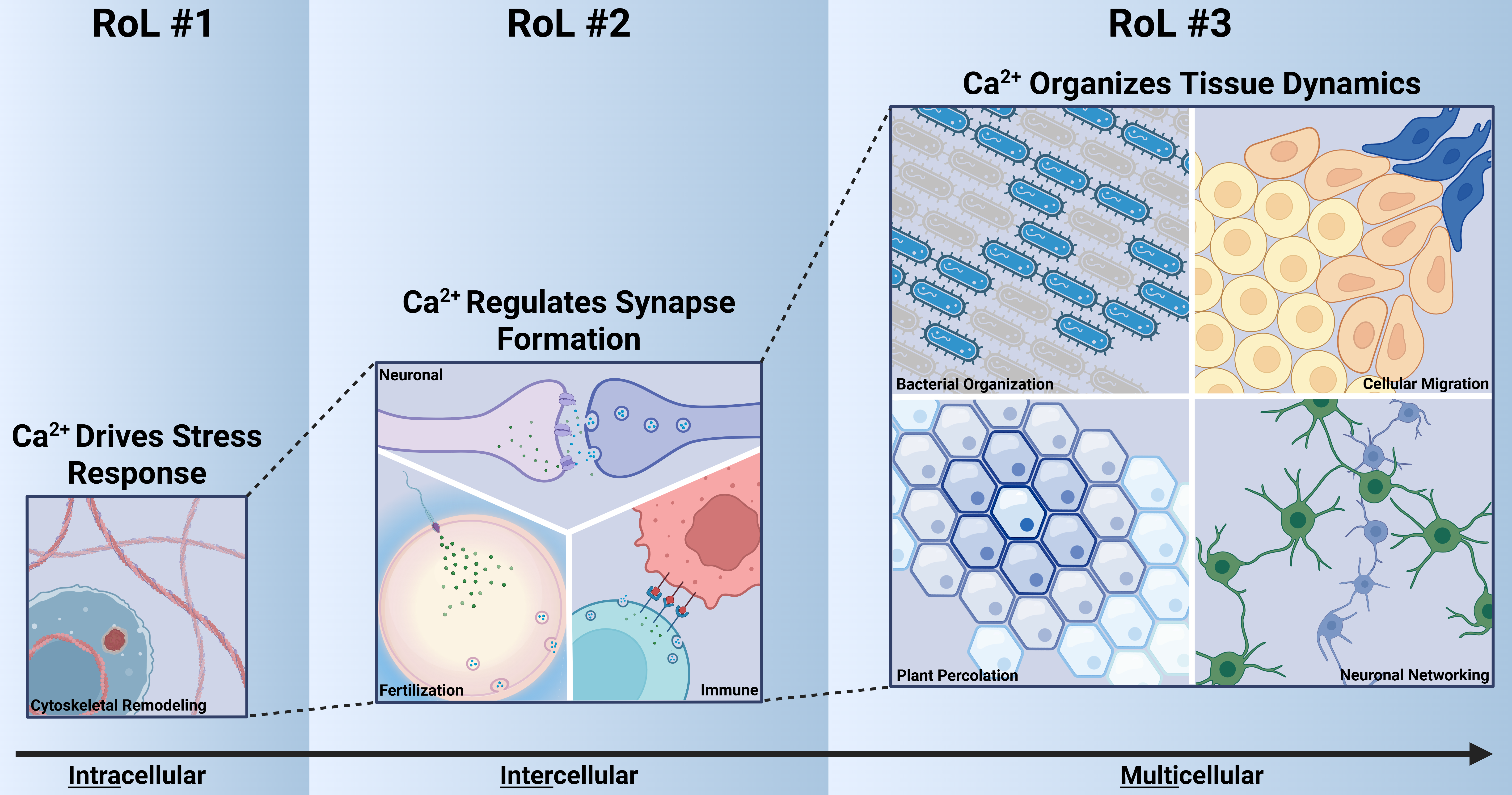
In this review, we examine the functional roles of how Ca2+-cytoskeleton coupling shapes biological functions across multiple contexts and propose three initial Ca2+-cytoskeleton Rules of Life:
RoL #1: Ca2+ regulates cytoskeletal reorganization after stress and damage. A rapid Ca2+ influx initiates cytoskeletal remodeling, which is essential for cell responses such as migration, wound healing, and regeneration.
RoL #2: Ca2+ regulates actin dynamics to control synapse-like processes including neuronal, immune, and fertilization synapses, supporting both synapse formation and exocytosis.
RoL #3: Ca2+ exhibits diverse spatial patterns that both influence and are influenced by cellular dynamics, resulting in a division of cellular roles.
By defining these RoLs and illustrating their cross-context applications, this review describes a conceptual foundation for computational modeling, experimental hypothesis generation, and the engineering of living systems that are robust, adaptive, and responsive to their environments.
Background: The Calcium-Cytoskeleton Toolkit
Calcium ions (Ca
2+) function as critical intracellular messengers, enabling cells to respond to various stimuli [
23,
24]. For instance, ligand binding to plasma membrane receptors such as G protein-coupled receptors (GPCRs) activates signaling pathways that often regulate cytoplasmic Ca
2+ levels with specific spatiotemporal dynamics, termed Ca
2+ signatures [
25]. Upon GPCR stimulation by ligands (neurotransmitters, hormones, growth factors, etc.), phospholipase C (PLC) is recruited to cleave phosphatidylinositol 4,5-bisphosphate (PIP
2) into inositol 1,4,5-trisphosphate (IP
3) and diacylglycerol (DAG). IP
3 then binds its receptor (IP
3R) on the endoplasmic reticulum (ER), triggering the release of Ca
2+ to the cytosol from the ER lumen [
24,
25]. These increases in cytosolic Ca
2+ concentration are essential for a wide range of cellular responses to stimuli, including dynamic actin reorganization, which in turn supports the formation and function of critical cellular and tissue structures [
5,
7,
8,
26,
27,
28].
Proteins that respond to Ca
2+ activities—either by direct binding or via Ca
2+-binding partners like calmodulin—undergo conformational changes that decode Ca
2+ signals to regulate processes such as exocytosis, contraction, metabolism, egg activation during fertilization, and morphogenesis [
9,
29,
30,
31,
32,
33]. This interplay between calcium signaling dynamics and the cytoskeleton spans across kingdoms. For example, between plants and animals, Ca
2+ plays a crucial role in cell growth, development, and response to environmental stimuli [
8,
27,
33,
34,
35]. In the following subsections, we present examples from a range of biological systems to illustrate how organisms transduce stimuli into Ca
2+ signals and interpret those signals to regulate cytoskeletal organization.
Calcium Regulation of Actin Dynamics
As a core cytoskeletal component, actin exists in two primary forms: monomeric globular actin (G-actin) and polymeric filamentous actin (F-actin). F-actin undergoes continuous cycles of polymerization and depolymerization, enabling significant remodeling essential for cellular processes such as migration, division, and maintaining cell shape [
36,
37]. Structurally, monomeric G-actin polymerizes into right-handed helical filaments (F-actin) in an ATP-dependent manner, with growth occurring at the barbed (+) end and disassembly at the pointed (-) end. This process known as actin “treadmilling” facilitates turnover [
36]. Calcium signaling plays a pivotal role in regulating actin filament dynamics, both directly and indirectly, by interacting with actin-binding proteins (ABPs) [
4,
6,
38,
39,
40]. External stimuli create changes in intracellular Ca
2+ levels, and through ABPs, are essential for maintaining cellular morphology and allow for structural reorganization by influencing the severing, capping, and crosslinking of actin [
6,
41,
42].
For example, Ca
2+ binding activates gelsolin by inducing a conformational change that enables it to sever F-actin and cap newly formed barbed ends, halting further elongation [
39,
43]. Similarly, elevated Ca
2+ levels reduce the actin crosslinking ability of α-actinin, altering filament bundling and overall cytoskeletal organization [
40,
44]. Other ABPs, such as plastin/fimbrin and EFHD2/Swiprosin-1, also respond to Ca
2+ via EF-hand domains [
38,
45,
46]. In plastin/fimbrin, Ca
2+ binding induces a conformational change that weakens or misaligns their actin-binding domains, reducing their ability to bundle filaments [
45]. This intricate regulation by Ca
2+-responsive proteins provides cells with a versatile toolkit for rapid assembly and disassembly of the actin cytoskeleton, enabling multiple processes not limited to synaptic plasticity, and tissue morphogenesis [
6,
47,
48,
49].
Beyond direct regulation, Ca²⁺ indirectly controls actin architecture through signaling cascades involving Ca
2+-binding proteins such as calmodulin (CaM), which activates downstream effectors such as Calcium/Calmodulin-dependent protein kinase II (CaMKII), Protein Kinase C (PKC), and phosphatases like calcineurin [
24,
50]. These enzymes regulate, in turn, the activity of small Rho GTPases (RhoA, Rac1, Cdc42) by modulating the phosphorylation of guanine nucleotide exchange factors and GTPase-activating proteins (GAPs), altering actin polymerization, branching, or crosslinking [
51,
52]. For example, back-propagating action potentials in neurons elevate intracellular Ca
2+ concentration levels, activating CaM. In turn, CaM binds proteins like Cordon-Bleu to drive actin branching at or near the submembrane region of neuronal dendrites, facilitating the remodeling and extension of dendritic processes and thereby demonstrating how Ca
2+ governs cytoskeletal flexibility [
53,
54].
In plants, a striking example of Ca
2+-actin crosstalk occurs in organizing actin dynamics during key growth processes, such as the elongation of highly polarized structures like pollen tubes and root hairs. At the growing tip, localized Ca
2+ oscillations drive rapid cell expansion. ABPs such as villin, gelsolin, and fragmin (homologs to animal villin/gelsolin family members) respond to these Ca
2+ fluctuations by severing and capping actin filaments, promoting both filament bundling and depolymerization [
55,
56,
57]. In pollen tubes, a “tip-focused calcium gradient” forms as Ca
2+ levels rise at the growing tip, triggering actin depolymerization through Ca
2+-bound calmodulin, which activates villin. This enhances actin turnover and facilitates continued tip growth [
58]. Ca
2+ also integrates with RhoGTPase signaling pathways, such as the RHO of plant (ROP) family, to regulate actin polymerization and coordinate pollen tube growth [
56]. Additionally, Ca
2+ works with auxin signaling in root hairs to regulate ABPs and support root outgrowth [
26,
59].
While the functional outcomes and the specific intermediary Ca2+-actin interactions vary across species and cell types, the overarching role of Ca2+ in actin regulation is conserved. Animal cells exploit rapid Ca2+ transients to drive dynamic actin reorganization during cell migration, adhesion, and synaptic plasticity processes. While plant cells also exhibit transient Ca2+ signals, they also use stable or oscillatory Ca2+ gradients to support sustained, directional growth and nutrient transport across large, vacuolated cells. Despite these differences, both systems utilize conserved mechanisms involving CaM and RhoGTPases, highlighting the shared importance and signaling network architecture of Ca2+ regulation in actin dynamics and cytoskeletal control across species.
Ca2+ Regulation of Actin-Interacting Motors
Myosins are a family of contractile proteins critical for cytoskeletal organization. Myosin proteins support diverse cellular functions, including apical constriction, cell mobility, cytokinesis, cargo and organelle transport, and muscle contraction. Myosin activity relies on actin binding and is regulated by chemical and mechanical cues, with calcium signaling playing a central role [
60,
61,
62,
63,
64,
65,
66]. Myosin proteins share common structural domains, including motor, neck and tail domains. The neck domains contain IQ motifs defined as a series of isoleucine (I) and glutamine (Q) residues, which bind essential and regulatory light chains (ELCs, RLCs) and Ca
2+-dependent calmodulin (CaM). Both the tail and neck domains vary across families [
60,
67]. Among the 30+ myosin families, myosin II (also known as non-muscle myosin II) is the most studied and relies on Ca
2+-bound CaM to activate myosin light-chain kinase (MLCK) to phosphorylate its RLCs, enabling motor activation and filament assembly [
65,
67]. RhoA-associated Rho-kinase (RhoA/ROK) inhibits myosin light-chain phosphatase (MLCP), which dephosphorylates the RLC and deactivates myosin II.
RhoA/ROK sustains RLC phosphorylation, enhancing myosin II sensitivity to MLCK and Ca
2+ levels [
68]. This regulation is crucial for processes like muscle contraction, cytokinesis, and tension regulation across species, including mammals [
30,
65,
69], zebrafish [
70],
Drosophila melanogaster [
71],
Caenorhabditis elegans [
72],
Dictyostelium discoideum [
73], and yeasts [
74], among others [
65].
Unconventional myosins (myosins other than myosin II), such as myosin V and VI, exhibit distinct Ca
2+-dependent regulatory mechanisms specialized for cargo transport [
75,
76,
77,
78,
79,
80,
81]. Myosin V, a processive motor, transports organelles and vesicles along actin filaments before dissociating [
77,
82,
83,
84]. Its activity is regulated by Ca
2+-CaM binding at IQ motifs, which controls cargo attachment and actin interaction [
75,
85]. Interestingly, reduced Ca
2+ levels during active transport enhance myosin V motility speed [
75]. Myosin V functionality is conserved across various species, including mammals [
77,
86]), yeasts [
82,
87], and
Drosophila [
83,
88], among others. Myosin VI, unique for its reversed polarity and movement toward actin’s pointed (-) end [
66,
89,
90,
91], similarly relies on Ca
2+-CaM to release its tail for cargo binding. However, motility is only initiated once Ca
2+ levels return to a reduced level [
76]. Myosin VI functions in vesicle transport, endocytosis, lysosome maturation, autophagy, cell migration, and cell division [
76,
78,
79,
80,
81], with functional roles conserved across species [
92,
93,
94]. Other myosins, such as myosin I [
65,
95], VII [
65,
96], and plant-specific myosin VIII and XI [
97,
98], also display varying degrees of Ca
2+ sensitivity, underscoring the intricate regulation of myosin activity by calcium signaling and associated factors like MLCK, MLCP, RhoA/ROK, CaM, and IQ motifs. This fine-tuned control of myosin activity via Ca
2+ signatures enables precise cytoskeletal dynamics and intracellular transport.
Computational Models as Tools to Condense and Transfer Essential Biological Mechanisms
As discussed above, the intricate interplay of calcium signaling and cytoskeletal dynamics is pivotal in a wide range of cellular processes. This highlights the versatility of Ca2+ as a second messenger. The ability of Ca2+ signaling to regulate cytoskeletal organization underscores its foundational role in both biological relationships and broader Rules of Life.
Quantitative data-driven representations of hypothesized RoLs, expressed through computational models, can serve as powerful tools for integrating the complexity of biological signaling systems. Such models aim to recapitulate components of a system with functional-interdependent variables. In biological applications, computational modeling enables simulation of system dynamics and the generation of predictive outcomes, often with greater specificity and capability, providing insight that can be difficult or impossible to acquire from wet lab experimentation [
99], such as at the inference of physical parameter values [
100]. These models can test biological hypotheses, perform cross-system optimizations and translations, and transform our understanding of how Ca
2+ governs cytoskeletal dynamics and their underlying mechanisms. To do this, the simulation frameworks should capture spatiotemporal dynamics of calcium signaling, actin reorganization, and motor protein activity. This, in turn, demands a multiscale approach that incorporates both localized Ca
2+ spikes and global cytoskeletal reorganization.
Computational models also enable a comparative analysis across systems, revealing both shared principles and comparing context-specific variations in calcium signaling. Ultimately, they lay the foundation for deeper insight into mechanisms of health and disease linked to Ca2+ dysregulation.
Rules of Life (RoLs), which describe functional relationships between biological components, link systems across hierarchical levels to predict responses to perturbations. By unifying reductionistic mechanisms into a single modeling framework [
21,
22], RoLs condense vast datasets into generalized heuristics, facilitating knowledge transfer across scientific fields. For example,
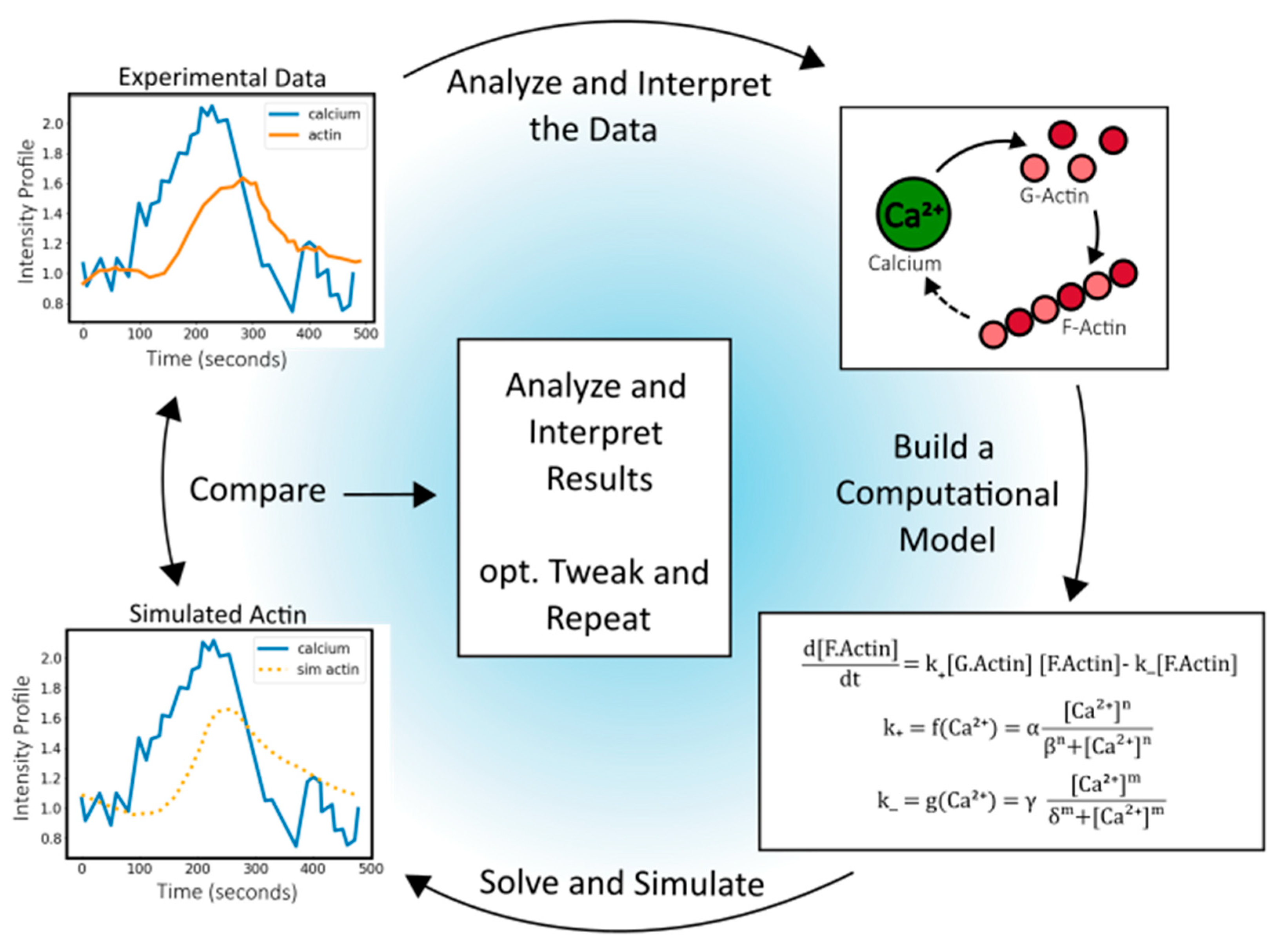
Figure 1 shows a representative pathway connecting experimental data with mathematical models. Here, actin polymerization is modeled as a reversible reaction using rate constants representing polymerization (
) and depolymerization (
), respectively (Eqn. 1). To account for Ca
2+ feedback, these rate constants can be further parametrized as functions of Ca
2+ concentrations (Eqn. 2a, 2b). This often uses linear or Hill equations. These coupling functions constitute relationships, and their parameterization requires data from biological systems. Analyzing these system-specific parameters reveals underlying rules and dynamics associated with Ca
2+ regulation of actin polymerization.
Directly measuring the parameters involved in biological processes remains a key bottleneck for validating computational models of biological systems. Techniques such as Fluorescence Recovery After Photobleaching (FRAP) and Raster Image Correlation Spectroscopy (RICS) provide powerful experimental assays for parameter estimation to extract kinetic parameters like diffusion coefficients and binding rates, respectively [
101,
102,
103,
104]. When integrated with partial differential equation (PDE)-based models of reaction-diffusion dynamics, both FRAP and RICS can help infer the underlying biochemical parameters that define ligand-receptor and protein-protein interactions [
105,
106]. Nonetheless, parameter values can be estimated with quantifiable levels of uncertainty through model calibration to experimentally obtain metrics. Parameter calibration uses sensitivity analysis, prediction testing, and comparison of model results over a variety of scenarios [
107]. Commonly, parameter screening explores defined biologically feasible parameter spaces to find the best-fit parameter value (or parameter distribution) for a given set of experimental data; however, this approach is often computationally expensive depending on the model complexity and size of the parameter space [
108,
109].
Recent advances in Artificial Intelligence (AI) and Machine Learning (ML), including Neural Networks (NNs), Physics-Informed Neural Networks (PINNs), and Bayesian integrative models, have improved the efficiency of parameter optimization in biological-computational modeling, particularly in their application of surrogate models and active parameter range searching [
108,
110,
111,
112]. Multisystem optimization has improved our understanding of how shared biological molecules or mechanisms function across different systems, offering insights into conserved processes and their variability across species [
113,
114,
115,
116]. One study using a core Smad pathway model combined with Pareto front analysis showed that by tuning non-conserved parameters, the BMP-Smad pathway can be optimized for multiple performance objectives, such as response speed, noise amplification, and output sensitivity, but not all objectives can be idealized simultaneously. This tradeoff can instead be addressed through Pareto-optimal balance, which varies across species [
117]. Advances in artificial intelligence and computational power further open new opportunities to bridge experimental and computational approaches, helping transform RoLs into quantifiable and practical frameworks for biological research [
118,
119].
As computational power increases, the formulation of biochemical-biomechanical models enables the quantitative application of RoLs across diverse cellular and tissue processes. By harnessing computational techniques to reverse-engineer the emergent biological mechanisms, we can uncover how the combinatorial coupling of RoLs varies across different biological contexts and systems, providing insights into their impact on robustness and organization in those complex systems.
Figure 1.
Design-test-iterate, coupling computational and experimental workflows. Experimental Ca2+ and actin observations are shown with a simple model to predict actin response from input Ca2+ data. Beginning with the experimental data, relationships among network components can be mapped, such as the impact of Ca2+ on G-actin and F-actin. A hypothesized interplay of F-actin on Ca2+ is included as a dashed line. Next, a system of equations is defined and integrated computationally. Comparison and interpretation of the experimental and computational results can be used for calibration, sensitivity analysis, and model comparison to test predictions across systems.
Figure 1.
Design-test-iterate, coupling computational and experimental workflows. Experimental Ca2+ and actin observations are shown with a simple model to predict actin response from input Ca2+ data. Beginning with the experimental data, relationships among network components can be mapped, such as the impact of Ca2+ on G-actin and F-actin. A hypothesized interplay of F-actin on Ca2+ is included as a dashed line. Next, a system of equations is defined and integrated computationally. Comparison and interpretation of the experimental and computational results can be used for calibration, sensitivity analysis, and model comparison to test predictions across systems.
Candidate Set of Rules of Life for Calcium-Cytoskeleton Coupling
Building on the biological and computational discussions above, we propose candidate Rules of Life pertaining to calcium-cytoskeleton coupling. These proposed rules have wide zones of influence and we discuss specific application contexts for each proposed rule. We also lay a conceptual framework for systematically testing RoL candidates through iteration between quantitative experiments and simulations.
I. RoL 1: Ca2+ Triggers Cytoskeletal Reorganization After Stress and Damage
Ca
2+ mediates cellular responses to various stressors, including physical damage, toxins, environmental extremes, pathogens, and antigens. A disruption in cellular homeostasis triggers an immediate calcium signal, elevating cytosolic Ca
2+ levels and prompting a restructuring of the cell’s actin-myosin cytoskeleton structure. This mechanism is shared across numerous cell types and species [
120,
121]. For instance,
in mammals, stressors like ATP or damage-associated molecules lead to Ca
2+ influx in endothelial cells [
122,
123,
124]
. As additional examples, fungi, and yeast show distinct Ca
2+ signaling patterns in response
to osmotic shock and oxidative stress [
125,
126]
. Similarly, plants rely on Ca
2+ to respond to both abiotic stress, such as drought, salinity, and extreme temperatures, and biotic stresses caused by pathogen invasion or damage from insects and parasites [
127,
128,
129,
130], where Ca
2+ influx initiates systemic defense signaling.
Other perturbations such as stimulation with G-protein-coupled receptor ligands [
131]
, thapsigargin-mediated ER Ca2+ release [
132,
133,
134]
, and mechanical disruptions like atomic force microscopy or laser-induced ablation, induce rapid Ca2+ waves across cells in diverse organisms [
5]
. Thus, calcium signaling regulates stress and injury responses across kingdoms.
Calcium-Mediated Actin Reorganization
Calcium-mediated actin reorganization occurs due to a sudden increase in intracellular Ca
2+ concentration [
5], as observed in multiple mammalian cell types,
including epithelial, mesenchymal, endothelial, and immune cells, and is triggered by diverse physiological stimuli [
5,
135]. In calcium-mediated Actin Reset (CaAR), a sudden Ca
2+ influx induces the disassembly and reassembly of cortical actin around the nucleus (
Figure 2). This dynamic rearrangement alters cell shape and mechanical properties, facilitating essential cellular functions such as the transient immobilization of organelles, cortex repair, cell spreading, wound healing, and regulation of gene expression crucial for cellular repair [
5]. In immune responses, neutrophil activation requires Ca
2+-driven actin reorganization to promote the phagocytosis-promoting complement receptor, in response to a pathogen encounter. This allows the cells to engulf invaders at sites of infection [
136].
In plants and fungi, rapid actin remodeling occurs in response to stress or cell damage. For example, fungal or oomycete pathogen penetration induces dense actin patches and focal arrays [
137], while mechanical stimuli, such as touching the plant surface with a microneedle, trigger Ca
2+ waves and local actin aggregation within 1–2 minutes [
138,
139]. Furthermore, in response to microbe-associated molecular patterns, actin remodeling and an increase in global filament abundance are induced in the
Arabidopsis epidermal cell cortex [
137]. Although direct evidence linking Ca
2+ influx to actin remodeling is lacking in these studies, it likely influences actin dynamics indirectly by modulating reactive oxygen species (ROS), lipid production, or calcium-dependent protein kinases (CPKs). For instance, during plant pattern-triggered immunity, Ca
2+ activates ROS bursts, which along with phosphatidic acid (PA), stimulate actin assembly and regrowth by inhibiting the Capping Protein [
140]. Additionally, CPK3 targets the Actin Depolymerizing Factor 4 during the immune response against bacterial pathogens [
141]. In the pollen self-incompatibility response, which prevents self-fertilization, high Ca
2+ influx triggers actin severing and depolymerization leading to pollen cell death [
142], with a gelsolin-like protein identified as the Ca
2+ target [
143]. Fungi also demonstrate instances of Ca
2+-mediated actin reorganization, specifically in
Aspergillus nidulans, a filamentous fungi in the phylum Ascomycota. Here, pulsed Ca
2+ influxes regulate temporally controlled actin assembly and exocytosis, resulting in stepwise cell extension [
29]. These examples demonstrate how Ca
2+ acts as a conserved and versatile signal that transduces environmental stress into actin remodeling, enabling diverse cell types across organisms to adapt rapidly to stimulus.
Ca2+ Signal Initiation of Wound Healing and Regeneration
Ca
2+ channels also play a crucial role in wound repair by mediating the influx of Ca
2+ into cells) [
144,
145,
146]. In
Caenorhabditis elegans, skin wounding triggers a Ca
2+-dependent signaling cascade with two key functions: facilitating wound closure and activating the innate immune response. After injury, a rapid and sustained elevation of Ca
2+ levels are observed in the epidermis, which is essential for organism survival. This influx relies on the transient receptor potential channel of the melastatin family (TRPM), known as GTL-2, which senses mechanical or chemical stress and mediates Ca
2+ influx from the extracellular space into the cytosol, and is amplified through IP
3-mediated Ca
2+ release from internal stores (
Figure 2C). Xu et al. identified an epidermal signal transduction pathway involving Gα
q-Ca
2+ and its effector PLCβ (EGL-30 and EGL-8 in
C. elegans, respectively), which drives actin-dependent wound closure, independently of known innate immune responses; loss of this pathway impairs survival post-injury [
147].
After injury, mammalian cells, such as fibroblasts, immune cells, and epithelial cells exhibit a rapid increase in intracellular Ca
2+ that can propagate through tissue via signaling receptors or gap junctions [
144]. Similar calcium signaling dynamics occur during tissue-level responses in human wound healing. For example, fibroblasts depend on Ca
2+ influx in response to deep dermal wounds for proper response,
which can influence both healing outcomes and scar formation. Blocking this signaling with the Ca
2+ chelator EGTA, Jiang et al. observed reduced scarring by fascia fibroblasts [
148], suggesting Ca
2+ modulation can influence scarring via the gelsolin family of actin-severing proteins [
149]. Additionally, traditional wound treatments, like honey, induce extracellular Ca²⁺ influx in keratinocytes through TRPM2 and Orai1 channels, an effect that varies with honey’s botanical source due to differential H
2O
2 production [
150]. Moreover, both calcium and ROS species transcriptionally regulate tissue regeneration. The conserved Ca
2+-calcineurin signaling pathway activates transcription factors such as the nuclear factor of activated T-cells (NFAT) and myocyte enhancer factor-2 (MEF2), initiating immune responses [
31,
151,
152,
153,
154]. H
2O
2 can regulate actin cytoskeleton dynamics by modulating actin-binding protein (ABPs) expression such as Profilin, ARP-3, MRLC, and 14-3-3ζ, therefore facilitating wound closure [
155,
156]. Elevated Ca
2+ and H
2O
2 levels activate inflammatory transcription factors, such as activator protein 1 (AP-1) and nuclear factor kappa B (NF-κB), to promote wound healing [
157,
158,
159,
160]. Ultimately, calcium signaling acts as a central regulator of both immediate wound responses and long-term regeneration outcomes, coordinating cytoskeletal remodeling, immune activation, and transcriptional reprogramming across species. All these processes ultimately require coordination between cells and organs, and multicellular calcium signaling enables the systemic-level inter-organ communication through modulation of systemic and hormonal signals [
161,
162].
Feedback Between Actin and Ca2+ in Stress Response
At the molecular scale, Ca
2+ and actin participate in a feedback regulation loop that enhances cellular responses to stress. For instance, actin dynamics significantly influence Ca
2+ levels, playing a pivotal role in a plant’s response to salt stress. The Arp2/3 complex, which initiates actin filament nucleation, is central to modulating this adaptive mechanism. Disrupting actin organization in stressed plants results in elevated Ca
2+ levels [
120,
163].
This actin-Ca2+ connection is highlighted by Mimosa pudica’s well-known rapid leaf-folding response to touch, which serves as a defense against herbivores. The rapid bending results from changes in forces within the plant cell that pushes the plasma membrane against the cell wall (turgor pressure), driven by ion flux and partially regulated by Ca2+-mediated actin dynamics. Electrical stimulation experiments in this species confirm that changes in actin dynamics directly affect Ca
2+ levels, highlighting the interconnectedness of these components in the plant's response to stimuli [
164]. Furthermore, Annexin proteins like AtANN5 and AnxGb6, further mediate Ca
2+-actin interactions by serving as scaffolding proteins for binding F-actin. These proteins associate with membranes displaying negatively charged phospholipids in a Ca
2+-dependent manner, influencing Ca
2+-permeable channel function during stress in cotton plants [
165].
In mammals, actin dynamics contribute to Ca
2+ homeostasis by interacting with ion transport channels on the plasma membrane [
166]. One key calcium pump in this process is the plasma membrane calcium ATPase (PMCA), crucial for maintaining low Ca
2+ concentrations in eukaryotic cells [
166,
167]. In red blood cells and blood platelets, G-actin directly binds to PMCA to release Ca
2+ from the cell [
166,
167]. Maintaining low intracellular Ca
2+ levels in platelets is essential for preventing premature activation, a process facilitated by PMCA pumps [
166,
168]. Upon platelet activation by agonists such as thrombin in response to vascular damage, PMCA is phosphorylated, leading to its inhibition and a subsequent rise in intracellular Ca
2+. This elevation in Ca
2+ triggers conformational changes that promote clot formation [
169]. In contrast, when F-actin binds to PMCAs, it deactivates the channel, preventing Ca
2+ removal, highlighting a complex and dynamic interplay between actin and Ca
2+ [
167]. Together, these findings illustrate a tightly regulated calcium-cytoskeleton coupled mechanism in which actin and actin binding proteins and Ca
2+-related proteins dynamically modulate one another, enabling rapid and precise cellular responses to environmental needs.
Computational Modeling of Actin Dynamics Following Stress
Ca
2+ acts as a central mediator of cellular responses to diverse stressors, triggering actin cytoskeleton remodeling to support regeneration, immune defense, and structural adaptation [
5,
120,
121]. This interplay between calcium signaling and actin dynamics highlights a conserved mechanism for coordinating repair and stress responses across kingdoms, enabling cells to adapt efficiently to environmental and physiological challenges. Computational models have been applied to capture the role of Ca
2+ signals in wound healing, for instance, Stevens et al. presents a multi-scale calcium signaling model for epithelial wound healing in a
Drosophila pupal tissue. It combines a tissue injury model with detailed cellular calcium dynamics [
170]. Meanwhile, mechanical models have been developed to explore how these actin-driven processes contribute to wound closure efficiency. Bai and Zeng introduced a finite-element (FE) based mechanical model of wound healing, treating an epithelial cell sheet as a continuum but with cellular-level resolution [
171]. However, existing modeling approaches often struggle to fully integrate the signaling network with mechanical models [
172]. As noted earlier, one promising strategy is explicitly linking calcium dynamics to actin polymerization rates. For example, as shown in
Figure 2, a single-cell model can be used to test and predict the change in actin concentration following a rapid rise in Ca
2+ levels [
173]. This approach, ultimately, enables mechanistic testing of the factors that regulate cellular behaviors such as cell shape changes, cell migration, and wound repair.
Figure 2.
Calcium signaling and actin dynamics during wound response. A) Ca2+ and actin dynamics at the single-cell level in response to wounding. Localized wounding induces a rapid increase in Ca²⁺ concentration, which triggers the formation of an actin ring around the nucleus, followed by an actin reset to restore normal cytoskeletal organization once healed. B) Ca2+ and actin dynamics at the multi-cell level in response to wounding. Tissue wounding triggers a Ca2+ increase in surrounding cells, propagating a Ca2+ wave across the tissue. This leads to the coordinated formation of an actin ring at the wound site, promoting collective wound closure. C) General intracellular Ca2+ dynamics upon wounding across species. Intracellular Ca2+ levels rise through calcium channel influx across the plasma membrane or release from the endoplasmic reticulum (ER). Meanwhile, G protein-coupled receptors (GPCRs) can detect cellular damage by recognizing Damage-Associated Molecular Patterns (DAMPs) and chemokines, activating G protein pathways. Activated Gα subunits subsequently recruit PLCβ to the plasma membrane, hydrolyzing PIP₂ into IP₃. The generated IP₃ then binds to IP₃ receptors (IP₃R) on the ER membrane, triggering Ca2+ release into the cytosol. The elevated intracellular Ca2+ levels remodel actin dynamics through multiple pathways, including Rho GTPase and ROS signaling. Activated GTPases such as Rac1, RhoA, and Cdc42 regulate actin polymerization and actomyosin contractility. Cytoskeletal remodeling, in turn, can trigger Ca2+ responses, creating a positive feedback loop between calcium signaling and cytoskeletal dynamics. Created in BioRender.
Figure 2.
Calcium signaling and actin dynamics during wound response. A) Ca2+ and actin dynamics at the single-cell level in response to wounding. Localized wounding induces a rapid increase in Ca²⁺ concentration, which triggers the formation of an actin ring around the nucleus, followed by an actin reset to restore normal cytoskeletal organization once healed. B) Ca2+ and actin dynamics at the multi-cell level in response to wounding. Tissue wounding triggers a Ca2+ increase in surrounding cells, propagating a Ca2+ wave across the tissue. This leads to the coordinated formation of an actin ring at the wound site, promoting collective wound closure. C) General intracellular Ca2+ dynamics upon wounding across species. Intracellular Ca2+ levels rise through calcium channel influx across the plasma membrane or release from the endoplasmic reticulum (ER). Meanwhile, G protein-coupled receptors (GPCRs) can detect cellular damage by recognizing Damage-Associated Molecular Patterns (DAMPs) and chemokines, activating G protein pathways. Activated Gα subunits subsequently recruit PLCβ to the plasma membrane, hydrolyzing PIP₂ into IP₃. The generated IP₃ then binds to IP₃ receptors (IP₃R) on the ER membrane, triggering Ca2+ release into the cytosol. The elevated intracellular Ca2+ levels remodel actin dynamics through multiple pathways, including Rho GTPase and ROS signaling. Activated GTPases such as Rac1, RhoA, and Cdc42 regulate actin polymerization and actomyosin contractility. Cytoskeletal remodeling, in turn, can trigger Ca2+ responses, creating a positive feedback loop between calcium signaling and cytoskeletal dynamics. Created in BioRender.

II. RoL 2: Ca2+ Oscillations Drive Actin Reorganization Facilitating Synapse Formation
Synapses connect different biological compartments, facilitating vital interactions between cells [
174,
175]. A prominent example is the neuronal synapse, which transmits and integrates signals between neural cells [
176]. Synapse-like structures are not limited to the nervous system; they also appear in other biological contexts, such as the immunological synapse between T-cells and APCs or target cells, and the fertilization synapse between sperm and egg [
174,
175,
177,
178]. Despite differences in structure and function, these synapse-like structures surprisingly share key organization and molecular features, including adhesion molecules, signaling cascades, secretion, and precise localization of molecular arrangements [
174,
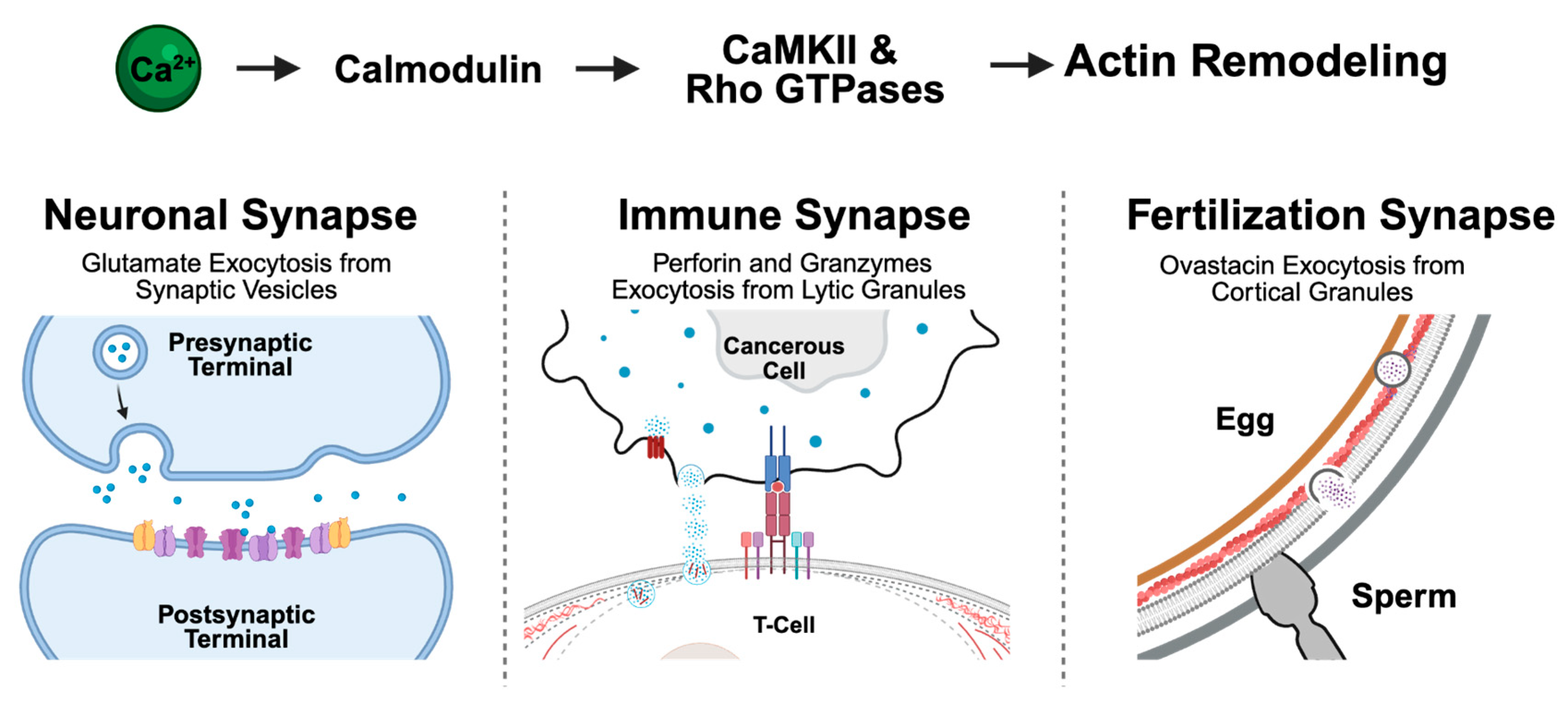
175]. In this section, we review a specific example of common signaling cascades in synapse-like structures, focusing on how calcium signaling dynamics, triggered by synapse formation, induce actin cytoskeleton reorganization in three distinct specialized synapse-like structures: i) neuronal synapse, ii) immune synapse, and iii) fertilization synapse (
Figure 3).
Calcium-mediated Actin Regulation in Neuronal Synapses
In neuronal synapses, communication requires actin reorganization, which allows the synapse to modify cytoskeletal structures in response to stimuli [
179,
180,
181]. While signaling pathways differ between presynaptic and postsynaptic terminals [
182,
183,
184], intracellular Ca
2+ serves as a key modulator of these changes by influencing F-actin dynamics through actin-binding proteins (ABPs) like cofilin [
185], CaMKII
[
186], and drebrin (DBN) [
187]). These interactions are further shaped by signaling cascades involving kinases like LIM Kinase 1, which regulates cofilin activity [
188].
In the presynaptic terminal, Ca
2+-dependent exocytosis of glutamate (Glu) from synaptic vesicles (SV) plays a critical role in signal propagation across the synaptic cleft (
Figure 3). This process includes spontaneous "mini" release, asynchronous release, and action potential-driven neurotransmitter release, regulated by synaptotagmin (Syt) and complexin [
189]. Cytoskeletal proteins such as F-actin and associated motor proteins are essential for SV trafficking and positioning within distinct pools, particularly the reserve and readily releasable pool. The movement of SVs from the reserve pool to the readily releasable pool is a critical step in exocytosis [
190,
191,
192,
193]. Ca
2+-mediated actin filament dynamics, such as actin depolymerization, facilitate SV movement into the active zone (AZ) of the readily releasable pool, where exocytosis occurs [
194]. Further, Rab-3 and SNARE (soluble N-ethyl-maleimide-sensitive fusion protein (NSF) attachment protein receptor) proteins orchestrate vesicle trafficking, with Rab-3 aligning and docking vesicles at the membrane and the SNARE complex (syntaxin, SNAP-25, and VAMP) mediating their fusion. Ca
2+ binding to synaptotagmin rearranges this complex, allowing vesicle and plasma membrane to merge and neurotransmitter release to occur. Following exocytosis, NSF and α-SNAP disassemble the SNARE complex, ensuring vesicle recycling [
195,
196,
197].
At the postsynaptic terminal, Ca
2+ influx from the synapse through NMDA (N-methyl-D-aspartate) receptors (NMDARs) initiates actin remodeling that is critical for dendritic spine morphology. NMDAR activation via glutamate binding dislodges Mg
2+ ions in the receptor and initiates Ca
2+ influx from the extracellular space. This increase in Ca
2+ triggers signaling cascades involving Ca
2+/CaM, RhoGTPase proteins, and actin depolymerizing factors (ADFs) [
198,
199,
200,
201]. Additionally, this Ca
2+ influx recruits NMDARs and AMPA receptors (AMPARs) to the postsynaptic terminal via exocytosis, accelerating future dendritic spine depolarization [
202,
203]. Proper localization of scaffolding proteins like PSD95 [
204], GKAP [
205], Shank [
206,
207], and Homer [
208,
209] at the postsynaptic terminal, is essential for actin cytoskeletal dynamics and cellular morphology, as they support effective synaptic signaling. For example, disruptions in Homer1b through gene silencing show reduced NMDAR- and AMPAR-mediated excitatory postsynaptic currents (EPSCs) and spine density [
207,
209]. Further evidence with knockout studies of ArpC3, a subunit of the actin-related protein 2/3 (Arp2/3) complex involved in F-actin branching, highlights the role of actin in structural plasticity, as these knockouts exhibit abnormal dendritic spine loss following stimulation [
210]. Additionally, splice variants of SNAP-25 in neurons result in fewer primed vesicles compared to their native counterpart [
211]. However, most experiments draw conclusions through either individual protein silencing, knockout, or protein homologs. A key open question relates to the systematic characterization of cytoskeletal dynamics. Consequently, computational models that incorporate experimental data and finetune parameters are needed to test systems-level behavior. For example, recent computational models demonstrate that the geometry of the synaptic cleft itself can influence neurotransmitter diffusion and NMDAR opening timing, highlighting the coordination between Ca
2+ and cytoskeletal proteins for synapse formation [
212]. In summary, precise regulation of Ca
2+ signaling, together with cytoskeletal protein function, is essential for neuronal synapse formation.
The Interplay Between Calcium and Actin in Immune Synapses
The immune synapse forms at the interface between T cells and either antigen-presenting cells (APCs) or target cells such as cancerous or virally infected cells (
Figure 3). The remodeling of the actin cytoskeleton within T cells plays a central role in establishing and maintaining the T-cell receptor (TCR) microclusters, which are critical for initiating downstream immune responses [
213,
214,
215,
216,
217,
218]. TCRs recognize peptides presented on the surface of infected cells or APCs by major histocompatibility complex (MHC) molecules, forming peptide-MHC complexes that trigger a signaling cascade leading to the recruitment of phospholipase C gamma 1 (PLCγ1) and subsequent Ca
2+ release from the ER through the IP₃ pathway [
213,
219,
220,
221]. When ER Ca
2+ stores are sufficiently depleted, the Ca
2+ sensor STIM1 translocates to ER–plasma membrane junctions, where it activates calcium release-activated calcium (CRAC) channels composed of Orai1 subunits [
222,
223]. Elevated cytosolic Ca
2+ then activates calmodulin, CaMKII gamma (CaMKIIγ), calcineurin, and Rho-family GTPases including Rac1 and Cdc42, which are essential for actin remodeling and dynamics [
222,
224,
225,
226]. These Ca
2+-dependent signals promote the activation of actin nucleators including Arp2/3 and formins, which drive the formation of distinct actin networks such as distal (dSMAC), peripheral (pSMAC), and central (cSMAC) supramolecular activation clusters (SMAC), as well as small actin foci distributed within the dSMAC and pSMAC [
213,
227,
228].
TCR microclusters are initially assembled within the branched actin network in the dSMAC and then move further into the pSMAC. An inward retrograde flow of F-actin in the dSMAC drives the centripetal retrograde movement of TCR microclusters toward the center of the cell, where their signaling is terminated and Ca
2+ levels are decreased. This process plays a critical role in modulating the strength and duration of TCR signaling [
213,
227,
229]. Interestingly, while Ca
2+ is essential for actin dynamics and retrograde flow, retrograde flow and the formation of TCR microclusters are, in turn, necessary to sustain Ca²⁺ signaling. Studies have shown that freezing retrograde flow preserves preformed TCR microclusters, but prevents the formation of new ones, leading to the loss of PLCγ1 activation and intracellular Ca
2+ [
221,
230].
One of the major T cell effector functions driven by Ca
2+-actomyosin dynamics in cytotoxic T cells (CTLs) is the polarized secretion of lytic granules (LGs)—specialized lysosomes containing cytotoxic molecules like perforin and granzymes—at the immunological synapse [
231,
232,
233]. LG secretion occurs at the cSMAC, but requires a coordinated sequence of cytoskeletal events. First, integrins cluster and concentric actomyosin arcs form within the pSMAC, creating a tight adhesive seal between the T cell and its target. Second, actin-binding scaffold proteins are cleared from the cSMAC, enabling centrosome polarization and docking at the plasma membrane [
213,
234]. Lytic granules are then transported along microtubules toward the polarized centrosome, transfer onto actin filaments near the synapse under guidance of myosin IIA [
234], and finally dock at the plasma membrane with the assistance of tethering proteins [
213,
234]. In the mature cSMAC, a dense actin meshwork initially acts as a barrier to the granule release. Upon T cell activation, actin remodeling increases the porosity of the network, enabling the final approach and fusion with the membrane, facilitating the targeted secretion of LGs at the immunological synapse [
213]. Once released, cytotoxic molecules enter the target cell either via endocytosis (internalization model) or directly through perforin-formed pores in the plasma membrane (pore formation model), ultimately leading to apoptosis of the target cell [
234]. Natural killer (NK) cells use a similar mechanism for granule release, relying on the same cytoskeletal dynamics in the transition from microtubule-based transport to actin-guided movement and Ca
2+-driven actin remodeling and fusion, despite differing in how they recognize their targets [
235]. As in neuronal synapses, LG exocytosis at the immune synapse involves conserved machinery—such as synaptotagmin, Rab proteins, and SNAREs—which respond to Ca²⁺ dynamics to trigger granule release [
236].
The Role of Calcium on Actin in the Fertilization Synapse
The fertilization synapse is formed by sperm-egg membrane interaction, a process that leads to increased cytosolic Ca
2+ concentrations in the egg (
Figure 3) [
175]. The spatiotemporal cytosolic Ca
2+ signaling dynamics during fertilization vary by species and experimental conditions, particularly in pattern (overall temporal profile), frequency (occurrence of waves or oscillations over time), and amplitude ([Ca
2+] change) [
33,
237,
238,
239]. In many non-mammalian species, fertilization typically triggers one or a few Ca
2+ waves lasting minutes (e.g.,
Caenorhabditis elegans,
Drosophila melanogaster) [
237], while in certain marine invertebrates oscillations can persist 30–60 minutes (e.g., the mollusk
Septifer virgatus), and in mammals, fertilization initiates a large initial transient followed by shorter oscillations lasting several hours with intervals ranging from a few to tens of minutes [
237]. In most invertebrates studied to date, this Ca
2+ rise at fertilization is mediated by an influx through voltage-gated long-lasting (L-type) Ca
2+ channels [
237]. However, in mammals, this fertilization-induced rise of intracellular Ca
2+ originates instead from the ER, triggered by a signal transduction cascade initiated by the sperm. The sperm introduces a unique form of phospholipase C (PLC), PLC zeta (PLCζ), which stimulates Ca
2+ release from the ER through the IP
3 pathway [
240].
This initial release is essential for propagating Ca
2+ waves across the egg; however, sustained Ca
2+ oscillations depend on continued Ca
2+ influx from the extracellular space, which is mediated by different channels depending on the species. During mammalian fertilization, several Ca
2+ entry pathways have been implicated. One of the best characterized pathways is Store-Operated Calcium Entry (SOCE), mediated by STIM1/2 and Orai1 upon ER Ca
2+ depletion [
237,
241]. However, its role in fertilization remains debated and appears species-specific [
237]. In mice, channels such as Cav3.2, TRPM7, and possibly TRPV3 also contribute to Ca
2+ influx [
175,
242,
243,
244]. The resulting rise in cytosolic Ca
2+ acts as a key trigger for both signaling and actin remodeling, driving essential events such as cortical granules exocytosis, meiotic resumption, and cytoplasmic reorganization via effectors like calmodulin, CaMKIIγ, gelsolin, PKC, and myosin II [
245,
246].
Prior to fertilization, Golgi-derived cortical granules (CGs) migrate along actin filaments, nucleated by formin-2 and Spire 1/2, toward the oocyte cortex in preparation for exocytosis [
247,
248]. In metaphase II eggs, which are competent for fertilization, CGs are positioned just beneath the plasma membrane and are anchored to the cortical actin cytoskeleton with the assistance of the subcortical maternal complex (SCMC), which creates passages for CG docking to enable rapid exocytosis at fertilization [
248]. Non-muscle myosin IIA, associated with CGs, generates contractile forces to help clear actin and facilitate CGs docking and exocytosis [
246,
249]. Ca
2+-dependent actin depolymerizing proteins like gelsolin also facilitate actin clearance [
246]. Following sperm-egg fusion, the CGs exocytose the metalloendopeptidase ovastacin, which cleaves zona pellucida (ZP) glycoprotein 2 (ZP2), modifying the ZP to prevent further sperm penetration and thus blocking polyspermy [
250,
251].
Similar to synaptic vesicle (SV) release in neurons, exocytosis of CGs involves proteins such as synaptotagmin, synapsin I, Rab3, Rabphilin-3A, and SNAREs. Synaptotagmin and SNAREs, in particular, regulate vesicle fusion in response to Ca
2+. While these proteins are expressed in oocytes and localized to the egg cortex, their specific roles in CG exocytosis remain to be fully confirmed. Nonetheless, their presence suggests they may be key conserved players in the Ca
2+-dependent exocytotic process during fertilization [
246].
Computational Modeling of the Conserved Characteristics Between Synapses
Although neuronal, immune, and fertilization synapses operate in distinct biological contexts, they share many core features highlighting their conserved organization and function. One example is the use of Ca
2+-triggered exocytosis, which is tightly regulated by actin cytoskeletal dynamics across all three synapse types (
Figure 3). In each case, Ca
2+signaling drives actin remodeling, enabling the transport, positioning, and fusion of secretory vesicles with the plasma membrane—whether releasing glutamate in neurons, lytic granules in cytotoxic T cells, or ovastacin in oocytes. Limitations in a cell’s ability to remodel its structure lead to dysfunction and disease. Understanding the signaling cascades governing Ca
2+-driven cytoskeletal changes enable the development of a generalized computational model based on RoL principles. Recent advancements in super-resolution microscopy techniques, such as single-particle tracking photoactivated localization microscopy (sptPALM), enable researchers to measure a variety of actin-binding protein diffusion rates within dendritic spines [
204]. These quantitative measurements can calibrate cell-to-cell communication models to test how Ca
2+ signaling dynamics impact the cytoskeletal and communication junctions between cells. For example, agent-based models simulate CaMKII phosphorylation dynamics in response to calcium signals, while stochastic models analyze how spine geometry modulates Ca
2+ influx and its effects on structural plasticity [
252,
253]. Additionally, molecular simulations combined with experimental data have reconstructed CaMKII/F-actin bundle architectures to explain Ca
2+-triggered cytoskeletal remodeling [
254].
Figure 3.
A unifying model of Ca²⁺-regulated actin remodeling that leads to vesicle exocytosis at the fertilization, immune, and neuronal synapse. Upon synapse formation, a rise in intracellular Ca²⁺ activates calmodulin, CaMKII, and Rho GTPases, leading to reorganization of the actin cytoskeleton and exocytosis of specialized vesicles: synaptic vesicles in neurons, lytic granules in T cells, and cortical granules in oocytes. This conserved pathway suggests a common Ca2+ dependent mechanistic rule of life for organizing synapses between cells. Created in BioRender.
Figure 3.
A unifying model of Ca²⁺-regulated actin remodeling that leads to vesicle exocytosis at the fertilization, immune, and neuronal synapse. Upon synapse formation, a rise in intracellular Ca²⁺ activates calmodulin, CaMKII, and Rho GTPases, leading to reorganization of the actin cytoskeleton and exocytosis of specialized vesicles: synaptic vesicles in neurons, lytic granules in T cells, and cortical granules in oocytes. This conserved pathway suggests a common Ca2+ dependent mechanistic rule of life for organizing synapses between cells. Created in BioRender.
III. RoL 3: The Functional Subdivision of Cell Populations Enables Improved Cellular Performance
From the pack mentality in wolves to the cellular communication within diverse bacterial communities in biofilms, collective behavior and intelligence [
255] is a fundamental phenomena observed throughout the natural world [
256]. Quantifying collective activities is complex and context-dependent, requiring diverse methodologies. For example, collective intelligence is assessed in human groups by evaluating individual and group task performance across various settings [
257]. Similarly, collective behavior is quantified in biological systems by measuring individual parameters such as cell velocity in cell migration [
258]. These approaches link individual performance to emergent collective properties using meta-analytic models, providing insights into intelligence across different contexts [
257,
258]. Understanding the mechanisms driving collective behavior can lead to innovative approaches for manipulating them, with potential applications in tissue development, wound healing, cancer treatment, and other health-related fields [
259].
In multicellular organisms, ion-based electrochemical signaling delineates cellular roles and facilitates tissue-wide signaling [
28,
260,
261,
262]. Less complex communities, like bacterial biofilms, use potassium ions to facilitate percolation—signal propagation through heterogeneous mediums. This process segregates cellular roles into active and passive cells, enabling coordinated organization and optimized survival [
260]. This phenomenon is preserved in larger complex organisms like metazoans where calcium signaling plays crucial roles in similar biological pathways. This has resulted in a division between cell populations into firing/non-firing [
260,
261], initiator/standby [
28], and leader/follower (
Figure 4) [
262,
263]. Ca
2+ exhibits diverse patterns that shape and are shaped by cellular dynamics, creating functional separations that facilitate signaling within and between cells and between tissues and organs. Because of this, calcium signaling is pivotal in regulating cellular behaviors and coordinating multicellular processes.
Firing and Non-firing
Tissue-wide, organ-to-organ, whole-body signaling is accomplished in metazoans through the use of complex cell types, structures, and agonists like neurons, axons, and chemokines. These provide the framework that facilitates long-range signaling throughout an organism [
28,
146,
260,
263]. However, similar signal coordination and self-organization can occur in simpler yet still complex biological systems, such as diverse cell populations found in bacterial biofilms. This phenomenon of percolation, signal propagation through heterogeneous mediums, arises from a separation of cellular roles between active (firing) and passive (non-firing) cells (
Figure 4A). By self-separating, bacterial communities can achieve the benefits of tissue-wide organization, decreasing the average individual cost of survival. They do this by channeling electrochemical cell-to-cell signaling from the growth center to the biofilm periphery creating a connected cluster of cells and a conduit for signal diffusion of potassium ions. Ultimately, this results in reduced nutrient consumption at the edges, while allowing increased nutrient availability and consumption at the interior [
260]. This is not the only example of role segregation in biology. Ca
2+, a universal messenger with a diverse set of signaling patterns, has been found to facilitate percolation in more complex systems.
Percolation also occurs in plants. Here, plants use Ca
2+ to facilitate cell-to-cell communication with diverse Ca
2+ kinetics and patterns [
261]. This can be modeled using stochastic two-cell and multi-cell scenarios, incorporating the origin of the signal and the number of initiating cells (
Figure 4B) [
261]. Such models properly simulate experimentally observed calcium dynamics of fire-diffuse-fire, where localized Ca
2+ release triggers further release from neighboring cells as the signal diffuses. This can result in a single cell Ca
2+ spark (a local stochastic event compared to more prolonged spiking), propagating through a tissue [
264]. These sparks in Ca
2+ don’t always propagate, however, but demonstrate nonequilibrium phase transitions between propagating and non-propagating, suggesting that the same set of scaling laws defines the fire-diffuse-fire phenomenon as those describing percolation [
265].
In animals, a related signaling behavior is observed in neurons through “sparse coding,” which efficiently translates complex environmental stimuli, such as light and sound, by reducing dimensionality [
266]. Sparse coding achieves this by relying on a small subset of active neurons within a population to represent information (
Figure 4C). These active neurons exhibit changes in membrane potential, producing temporally coordinated spikes of various ions like Ca
2+ [
267,
268]. The percolation patterns of signal propagation governed by sparse coding have been demonstrated in systems like
Drosophila [
269]. This strategy offers advantages similar to bacterial biofilm communication, including energy efficiency, enhanced storage capacity (reduced cross-talk enables greater memory), and improved data processing capabilities. In neurons, processing involves sequentially breaking information into components processed by distinct sets of neurons [
270]. Together, these studies highlight the utility of percolation theory in describing multicellular communication across both prokaryotic and eukaryotic systems.
Initiator and Standby
Cell populations often result in a subdivision into multiple cell classes based on cell signaling or morphological distinctions. For example, in epithelia, quantifying signaling dynamics have revealed the distinction between two types of cells, initiator and standby cells, which are defined by their specific roles in calcium signaling. Calcium’s diverse tissue patterns involve rapid spiking and long transient signaling within single cells and extended wave responses as well as sharp fluttering across tissues [
8,
10,
28,
31,
271]. These dynamic patterns create separations in cellular behavior. For example, Soundarajan et al. 2021 demonstrated the existence of initiator cells—responsible for initiating calcium signals—and standby cells—waiting to propagate these signals—in
ex vivo experiments on developing wing imaginal discs in
Drosophila melanogaster, which is similar to what is observed in plants (
Figure 4B) [
28,
272]. Their findings also suggest a “Goldilocks zone” for calcium signaling, where either too low or too high of a Ca
2+ level, disrupts tissue growth and development. This phenomenon was modeled by mimicking natural observations, with initiator cells identified by increased phospholipase activity, leading to rapid Ca
2+ signaling [
28].
Distinct calcium signaling events in plants have also been reported, each carrying different physiological relevance [
27]. For example, tip-focused Ca
2+ oscillations during pollen tube and root hair elongation provide different signaling cues that facilitate rapid cell growth essential for elongation [
26,
56]. Additionally, in response to infection, transient cytoplasmic increases of Ca
2+ within local cells activate plant defense responses in an initiator vs. standby pattern [
272,
273]. Salt stress signals also rely on high-speed Ca
2+ waves capable of triggering molecular responses in distant parts of the plant and are capable of traveling through root cortical and endodermal tissue layers
(Figure 4B) [
274]. Such phenomena exemplify the role of Ca
2+ signaling in facilitating long-range communication and percolation across cellular networks.
Structures such as gap junctions in animals, insects, and plasmodesmata in plants, facilitate direct intercellular communication and regulate Ca
2+ signaling, representing additional examples of RoL 2 communication beyond classical synapses [
275,
276,
277]. Gap junctions form continuous pathways between adjacent cells by connecting two hemichannels, enabling the exchange of ions and small molecules essential for physiological processes. In
Drosophila, the gap junction protein Inx2 regulates Ca
2+ flux, influencing the cell fate of border cells during oogenesis [
278]. Similarly, plasmodesmata mediate molecular and ionic exchanges between plant cells through cell walls [
279,
280]. Notably, plasmodesmata permeability is regulated by Ca
2+; changes in cytosolic Ca
2+ concentrations lead to plasmodesmata closure, controlling Ca
2+ flux [
281,
282]. This regulation is crucial in plant defense, as studies in
Arabidopsis thaliana show that Ca
2+ influx and plasmodesmata closure are integral to pathogen perception and response [
283]. Ultimately, calcium enables complex signaling within and between cells across tissues and organs by assigning cellular roles.
Leaders and Followers
Following this trend, cellular migration relies on calcium signaling, facilitating cellular organization in many critical areas like wound healing and immunity [
146]. Like the division of firing/nonfiring and initiator/standby roles, multicellular migration divides cells into distinct subpopulations as either leaders or followers. Leader cells are, by definition, located at the leading edge of the migrating ensemble and have the characteristics of directing and attracting other cells while inhibiting any leader activities in the cells that succeed them. In contrast, follower cells are defined as following the leaders, however, they do have the potential of becoming leaders themselves if the leading edge expands or a follower outcompetes the leader(s) (
Figure 4D) [
1,
262,
263,
284]. This type of cellular organization is observed in many different systems from insects, mammals, and zebrafish [
285,
286,
287,
288].
Multicellular migration depends on the separation of roles between leader and follower cells, driven by polarized signaling and physical asymmetry [
262,
263,
284]. Leaders exhibit asymmetric contact with their environment, forming a protrusive leading edge, while followers maintain symmetric contact with surrounding cells (
Figure 4D). Higher cadherin recycling rates along leaders’ lateral sides enhance migration speed and thick myosin-actin bundles prevent followers from overtaking them. Leaders additionally have reduced Notch1 levels due to lower internal tension than higher-tension follower cells [
263].
This polarization of signaling in leader-follower dynamics is mirrored in internal Ca
2+ levels, where a gradient of Ca
2+ across the cell drives persistent forward migration [
1]. Cells maintain reduced Ca
2+ at the front with elevated plasma membrane Ca
2+-ATPase (PMCA) activity compared to the rear. This polarization preserves myosin light chain kinase (MLCK) sensitivity to localized Ca
2+ signals via receptor tyrosine kinase and phospholipase C pathways. Repetitive signaling from these pathways depletes Ca
2+ from the ER, stimulating STIM1 (stromal interaction molecule 1), which is then transported to the cell front by the growing ends of microtubules, facilitating Ca
2+ entry via Orai1. Ca
2+ is then restored to the ER by SERCA (sarcoplasmic/endoplasmic reticulum Ca
2+-ATPase), sustaining local Ca
2+ signaling [
1,
289]. This dynamic signaling supports actin polymerization and recycling at the front and actin turnover at the rear, enabling persistent forward migration.
The leader-follower dynamic seen in multicellular systems can also be observed within single-cell migration. A single cell also exhibits distinct spatial Ca
2+ dynamics to regulate its migration differently from surrounding cells. Similar to multicellular migration, single cells create local Ca
2+ pulses near the leading edge and maintain a back-to-front Ca
2+ gradient [
289]. These calcium signaling events drive the processes of polarization, protrusion, retraction, and adhesion. However, underlying mechanisms driving the leader-follower structure, in which the activity of one cell influences Ca
2+ signal propagation across tissues, remain unclear. For example, during wound healing, ATP (adenosine triphosphate) serves as an early damage signal. A recent computational study revealed that distinct Ca
2+ responses to ATP could be attributed to variations in how Ca
2+ and IP
3 (inositol trisphosphate) are sensed by the IP
3 receptor (IP
3R) [
290]. These variations may arise due to differences in IP
3R expression within the endoplasmic reticulum. In epithelial tissues, functional diversity may arise during wound healing, as cells with unique physiological roles exhibit distinct calcium signaling activity to meet specific functional demands [
291]. These findings suggest that differential expression of Ca
2+ receptors like IP
3R may play a key role in determining which cells initiate and propagate signals during processes like wound healing.
Computational Modeling for Organized Tissue Responses
Multicellular computational models simulate and capture collective behaviors across diverse biological contexts, offering insights into how calcium signaling scales from internal cellular dynamics (RoL 1) to cell-cell interactions (RoL 2) and tissue-wide coordination (RoL 3). While RoL 1 and RoL 2 describe localized, cell-to-cell calcium dynamics that influence direct interactions, expanding to RoL 3 reveals how these signals propagate across larger cellular networks. Computational models developed for
Drosophila wing discs show the importance of variability in PLC production and gap junction permeability in regulating spatiotemporal patterns of Ca
2+ [
28]. Similarly, a multi-scale approach combining agent-based methods and partial differential equation (PDE) models in plant systems illustrate how localized Ca
2+ release propagates through tissues using Ca
2+-activated Ca
2+ channels, generating emergent patterns [
272].
Computational modeling plays a critical role in elucidating the signaling mechanisms in these systems, and the core details of these models can be applied to other biological settings, creating computational building blocks that can be used across species and kingdoms with little to no adaptation [
292,
293]. These models also help identify thresholds and feedback mechanisms determining whether cells function as initiators or standby cells. Since calcium signaling shapes and is shaped by cellular processes including actomyosin remodeling, integrating experimental and computational approaches will be essential for deepening our understanding of calcium's role in regulating cellular behaviors and coordinating multicellular processes. This will simultaneously guide Ca
2+-targeted therapeutic applications in regenerative medicine and disease treatment.
Figure 4.
Subdivision of Cell Populations. A) Firing and nonfiring in bacterial organization. Bacteria demonstrate signal percolation with distinct active (firing - blue) and passive (non-firing - gray) cells, through potassium ion use, organizing their communication. B) Initiator and standby dynamics in plant percolation. In plants, Ca2+ ions are similarly used to propagate signals across tissue layers, generating a Ca2+ wave that travels across the tissue, originating from the initiating cell (darker blue) and diffusing through the standby cells (fading blue). C) Neuronal dimensional reduction. In neurons, complex information is dimensionally reduced through sparse coding, organizing neurons from a single (dense) group into (sparse) sub-populations and networks (denoted by color), each representing unique stimuli or information, reducing cross-communication. D) Leader and follower segregation in multicellular migration. Leading cells (blue) direct cell migration forming a protrusive leading edge, while followers (tan) maintain more symmetric contact with surrounding cells while following the leaders. Created in BioRender.
Figure 4.
Subdivision of Cell Populations. A) Firing and nonfiring in bacterial organization. Bacteria demonstrate signal percolation with distinct active (firing - blue) and passive (non-firing - gray) cells, through potassium ion use, organizing their communication. B) Initiator and standby dynamics in plant percolation. In plants, Ca2+ ions are similarly used to propagate signals across tissue layers, generating a Ca2+ wave that travels across the tissue, originating from the initiating cell (darker blue) and diffusing through the standby cells (fading blue). C) Neuronal dimensional reduction. In neurons, complex information is dimensionally reduced through sparse coding, organizing neurons from a single (dense) group into (sparse) sub-populations and networks (denoted by color), each representing unique stimuli or information, reducing cross-communication. D) Leader and follower segregation in multicellular migration. Leading cells (blue) direct cell migration forming a protrusive leading edge, while followers (tan) maintain more symmetric contact with surrounding cells while following the leaders. Created in BioRender.

Conclusions
Defining the “Rules of Life” (RoLs) as mechanistic, data-driven relationships provides a foundational framework for understanding how calcium signaling and cytoskeletal dynamics coordinate biological responses across systems and scales. This forward-looking review identifies widespread roles for Ca2+ in regulating cytoskeletal remodeling in stress response, cellular communication, and tissue organization—demonstrating how RoLs unify predictive models for cellular responses while remaining adaptable to molecular and environmental contexts.
By developing modular computational functions grounded in experimental data, researchers can build adaptable computational models that simulate system-specific behaviors—capturing both shared principles and specialized mechanisms. In this way, the framework addresses a major challenge in systems biology—translating conserved signaling components into predictive, transferable models despite species specific differences in channel kinetics, effector sensitivity, feedback regulation, and spatial organization, even when homologous or orthologous proteins are involved [
294,
295,
296,
297,
298]. These models also support closed-loop discovery by guiding experimental planning through model-based design of experiments (MBDoE)
techniques [
299,
300].
This computational framework aligns with the RoLs presented here, where modular logic is derived from conserved signaling patterns and adapted to context.
RoL 1 demonstrates how Ca2+ signaling mediates stress responses by triggering cytoskeletal remodeling across organisms. However, the regulatory feedback between Ca²+, actin dynamics, and pathways like reactive oxygen species and G-protein signaling, remain poorly understood. Computational models, integrated with advanced live-imaging, are critically needed to further capture this modularity across stress-response systems.
RoL 2 illustrates how Ca2+-triggered actin remodeling spans neuronal, immune, and fertilization synapses—communication systems that share conserved features, like exocytosis, but also exhibit distinct molecular mechanisms. This combination of specialized signaling dynamics with shared core processes shows how biology implements modular components to accomplish context-specific functions. Replicating this principle in modular computational functions and integrating them with expanding in vivo studies and bioengineering approaches, will enhance our understanding of cellular communication and biological junctions.
RoL 3 underscores the versatility of Ca2+ signaling as a second messenger. Systematic expansion of in vivo studies across a broader range of model organisms, combined with centralized bioinformatic resources, featuring live-imaging data, will accelerate the identification of additional organizational RoLs, supporting flexible and extensible modeling frameworks.
Finally, RoL-based computational modules can inform digital twins for Regulator Science Tools in drug evaluation [
301,
302,
303,
304], or help connect translational medicine between model organisms and humans through predictive systems modeling [
292,
305,
306]. In synthetic biology and tissue engineering, modular RoLs may accelerate the design of engineered tissues and biomaterials with context-aware signaling capabilities that respond to their local environmental conditions [
307,
308,
309]. Finally, as advances and applications of artificial intelligence and advanced machine learning with bioengineering applications raise ethical, legal, and social implications (ELSI) [
310,
311,
312], a RoL framework offers a formal, modular basis for articulating design intent, tracing system behaviors, and guiding responsible innovation in emergent biological technologies [
313,
314,
315]. Overcoming each of these challenges ultimately requires interdisciplinary collaborations [
316]. By unifying experimental insight with modular computational models, this RoL-centered approach lays the groundwork for cross-scale, cross-species understanding of Ca
2+–actin dynamics—advancing both fundamental biology and its engineering applications.
Author Contributions
LL, DG, and JZ led the conception, design, and project management of the work. They contributed specialized expertise, actively participated in journal club discussions, contributed to drafting each section, performed literature review, created visualizations, and were heavily involved in editing and final approval. SMS and NL contributed to conception and design, participated in journal discussions, drafted individual sections, created visualizations, reviewed literature, and contributed to manuscript editing and approval. EK and NK contributed specialized expertise, as well as reviewed literature, and supported final revisions. MSM, MJK, and BS assisted in study conception and design, participated in journal discussions, and supported revisions and approval. CD, MM, AP, QD, EP, JPE, DMU, and JZ contributed to guidance, supervision, and final manuscript approval.
Acknowledgments
This work is based upon efforts supported by the EMBRIO Institute, contract 2120200, a National Science Foundation (NSF) Biology Integration Institute. This research was additionally supported in part by the NSF grant 2422229 awarded to JZ, LL, and DMU. We thank Dr. Christopher Staiger and Dr. Weiwei Zhang for the insightful discussions which greatly improved the manuscript. We are grateful to Dr. Norma Citlalcue Pérez Rosas, Jazzmin Owens and Feyisayo Akande for their participants and constructive input in the early stage journal club sessions.
Declaration of Interests
The authors declare no competing interests.
References
- Tsai, F.-C., Kuo, G.-H., Chang, S.-W. & Tsai, P.-J. Ca2+ Signaling in Cytoskeletal Reorganization, Cell Migration, and Cancer Metastasis. BioMed Res. Int. 2015, 409245 (2015).
- Bascom, C. S., Winship, L. J. & Bezanilla, M. Simultaneous imaging and functional studies reveal a tight correlation between calcium and actin networks. Proc. Natl. Acad. Sci. 115, E2869–E2878 (2018). [CrossRef]
- Hoffman, L., Farley, M. M. & Waxham, M. N. Calcium-calmodulin-dependent protein kinase II isoforms differentially impact the dynamics and structure of the actin cytoskeleton. Biochemistry 52, 1198–1207 (2013). [CrossRef]
- Staiger, C. J. & Blanchoin, L. Actin dynamics: old friends with new stories. Curr. Opin. Plant Biol. 9, 554–562 (2006). [CrossRef]
- Wales, P. et al. Calcium-mediated actin reset (CaAR) mediates acute cell adaptations. eLife 5, e19850 (2016). [CrossRef]
- Furukawa, R. et al. Calcium regulation of actin crosslinking is important for function of the actin cytoskeleton in Dictyostelium. J. Cell Sci. 116, 187–196 (2003). [CrossRef]
- Carafoli, E. & Krebs, J. Why Calcium? How Calcium Became the Best Communicator *. J. Biol. Chem. 291, 20849–20857 (2016). [CrossRef]
- Luan, S. & Wang, C. Calcium Signaling Mechanisms Across Kingdoms. Annu. Rev. Cell Dev. Biol. 37, 311–340 (2021). [CrossRef]
- Rajagopal, S. & Ponnusamy, M. Calcium Ion in Biological Systems. in Calcium Signaling: From Physiology to Diseases (eds. Rajagopal, S. & Ponnusamy, M.) 1–14 (Springer, Singapore, 2017). [CrossRef]
- Ahamad, N. et al. Differential activation of Ca2+ influx channels modulate stem cell potency, their proliferation/viability and tissue regeneration. NPJ Regen. Med. 6, 67 (2021). [CrossRef]
- Park, S.-Y., Park, C.-H., Choi, D.-H., Hong, J. K. & Lee, D.-Y. Bioprocess digital twins of mammalian cell culture for advanced biomanufacturing. Curr. Opin. Chem. Eng. 33, 100702 (2021). [CrossRef]
- Karolak, A., Markov, D. A., McCawley, L. J. & Rejniak, K. A. Towards personalized computational oncology: from spatial models of tumour spheroids, to organoids, to tissues. J. R. Soc. Interface 15, 20170703 (2018). [CrossRef]
- de Pablo, J. J. et al. New frontiers for the materials genome initiative. Npj Comput. Mater. 5, 1–23 (2019).
- Ghosh, S., Bheri, M. & Pandey, G. K. Delineating Calcium Signaling Machinery in Plants: Tapping the Potential through Functional Genomics. Curr. Genomics 22, 404–439 (2021). [CrossRef]
- Mori, Y. A multidomain model for ionic electrodiffusion and osmosis with an application to cortical spreading depression. Phys. Nonlinear Phenom. 308, 94–108 (2015). [CrossRef]
- Regazzoni, F., Dedè, L. & Quarteroni, A. Machine learning of multiscale active force generation models for the efficient simulation of cardiac electromechanics. Comput. Methods Appl. Mech. Eng. 370, 113268 (2020). [CrossRef]
- Van Liedekerke, P., Palm, M. M., Jagiella, N. & Drasdo, D. Simulating tissue mechanics with agent-based models: concepts, perspectives and some novel results. Comput. Part. Mech. 2, 401–444 (2015). [CrossRef]
- Yin, S.-F., Xue, S.-L., Li, B. & Feng, X.-Q. Bio–chemo–mechanical modeling of growing biological tissues: Finite element method. Int. J. Non-Linear Mech. 108, 46–54 (2019).
- Sun, M. & Zaman, M. H. Modeling, signaling and cytoskeleton dynamics: integrated modeling-experimental frameworks in cell migration. WIREs Syst. Biol. Med. 9, e1365 (2017). [CrossRef]
- NSF’s 10 Big Ideas - About NSF | NSF - National Science Foundation. https://www.nsf.gov/about/history/big-ideas.
- Lawing, A. M. et al. A Framework for Investigating Rules of Life by Establishing Zones of Influence. Integr. Comp. Biol. 61, 2095–2108 (2021). [CrossRef]
- Poust, A. W., Bogar, L., Robinson, W. D. & Poole, G. A Framework for Investigating Rules of Life Across Disciplines. Integr. Comp. Biol. 61, 2208–2217 (2022). [CrossRef]
- Brodskiy, P. A. et al. Mapping the calcium signalsome during Drosophila wing development. IFAC-Pap. 51, 108–109 (2018). [CrossRef]
- Berridge, M. J. The Inositol Trisphosphate/Calcium Signaling Pathway in Health and Disease. Physiol. Rev. 96, 1261–1296 (2016). [CrossRef]
- Bagur, R. & Hajnóczky, G. Intracellular Ca2+ Sensing: Its Role in Calcium Homeostasis and Signaling. Mol. Cell 66, 780–788 (2017). [CrossRef]
- Zhang, X. P., Ma, C. X., Sun, L. R. & Hao, F. S. Roles and mechanisms of Ca2+ in regulating primary root growth of plants. Plant Signal. Behav. 15, 1748283 (2020). [CrossRef]
- Tian, W., Wang, C., Gao, Q., Li, L. & Luan, S. Calcium spikes, waves and oscillations in plant development and biotic interactions. Nat. Plants 6, 750–759 (2020). [CrossRef]
- Soundarrajan, D. K., Huizar, F. J., Paravitorghabeh, R., Robinett, T. & Zartman, J. J. From spikes to intercellular waves: Tuning intercellular calcium signaling dynamics modulates organ size control. PLOS Comput. Biol. 17, e1009543 (2021). [CrossRef]
- Takeshita, N. et al. Pulses of Ca2+ coordinate actin assembly and exocytosis for stepwise cell extension. Proc. Natl. Acad. Sci. 114, 5701–5706 (2017). [CrossRef]
- Brito, C. & Sousa, S. Non-Muscle Myosin 2A (NM2A): Structure, Regulation and Function. Cells 9, 1590 (2020). [CrossRef]
- Brodskiy, P. A. & Zartman, J. J. Calcium as a signal integrator in developing epithelial tissues. Phys. Biol. 15, 051001 (2018). [CrossRef]
- Romero-Garcia, S. & Prado-Garcia, H. Mitochondrial calcium: Transport and modulation of cellular processes in homeostasis and cancer (Review). Int. J. Oncol. 54, 1155–1167 (2019). [CrossRef]
- Miao, Y.-L. & Williams, C. J. Calcium signaling in mammalian egg activation and embryo development: The influence of subcellular localization. Mol. Reprod. Dev. 79, 742–756 (2012). [CrossRef]
- Bhar, A., Chakraborty, A. & Roy, A. The captivating role of calcium in plant-microbe interaction. Front. Plant Sci. 14, (2023). [CrossRef]
- Friedl, P. & Gilmour, D. Collective cell migration in morphogenesis, regeneration and cancer. Nat. Rev. Mol. Cell Biol. 10, 445–457 (2009). [CrossRef]
- Blanchoin, L., Boujemaa-Paterski, R., Sykes, C. & Plastino, J. Actin Dynamics, Architecture, and Mechanics in Cell Motility. Physiol. Rev. 94, 235–263 (2014). [CrossRef]
- Schaks, M., Giannone, G. & Rottner, K. Actin dynamics in cell migration. Essays Biochem. 63, 483–495 (2019). [CrossRef]
- Lehne, F. et al. Calcium bursts allow rapid reorganization of EFhD2/Swip-1 cross-linked actin networks in epithelial wound closure. Nat. Commun. 13, 2492 (2022). [CrossRef]
- Nag, S., Larsson, M., Robinson, R. C. & Burtnick, L. D. Gelsolin: The tail of a molecular gymnast. Cytoskeleton 70, 360–384 (2013). [CrossRef]
- Burridge, K. Non-muscle n-actinins are calcium-sensitive actin-binding proteins. 294, (1981). [CrossRef]
- Babich, A. & Burkhardt, J. K. Coordinate control of cytoskeletal remodeling and calcium mobilization during T-cell activation. Immunol. Rev. 256, 80–94 (2013). [CrossRef]
- Janmey, P. A. Phosphoinositides and Calcium as Regulators of Cellular Actin Assembly and Disassembly. Annu. Rev. Physiol. 56, 169–191 (1994). [CrossRef]
- Maekawa, S. et al. Purification and characterization of a Ca2+-dependent actin filament severing protein from bovine adrenal medulla. J. Biol. Chem. 264, 7458–7465 (1989). [CrossRef]
- Pinotsis, N. et al. Calcium modulates the domain flexibility and function of an α-actinin similar to the ancestral α-actinin. Proc. Natl. Acad. Sci. 117, 22101–22112 (2020). [CrossRef]
- Schwebach, C. L., Agrawal, R., Lindert, S., Kudryashova, E. & Kudryashov, D. S. The Roles of Actin-Binding Domains 1 and 2 in the Calcium-Dependent Regulation of Actin Filament Bundling by Human Plastins. J. Mol. Biol. 429, 2490–2508 (2017). [CrossRef]
- Thylur, R. P., Gowda, R., Mishra, S. & Jun, C.-D. Swiprosin-1: Its Expression and Diverse Biological Functions. J. Cell. Biochem. 119, 150–156 (2018). [CrossRef]
- Bamm, V. V., De Avila, M., Smith, G. S. T., Ahmed, M. A. M. & Harauz, G. Structured Functional Domains of Myelin Basic Protein: Cross Talk between Actin Polymerization and Ca2+-Dependent Calmodulin Interaction. Biophys. J. 101, 1248–1256 (2011). [CrossRef]
- Schroeter, M. M., Beall, B., Heid, H. W. & Chalovich, J. M. The actin binding protein, fesselin, is a member of the synaptopodin family. Biochem. Biophys. Res. Commun. 371, 582–586 (2008).
- Renegar, R. H., Chalovich, J. M., Leinweber, B. D., Zary, J. T. & Schroeter, M. M. Localization of the actin-binding protein fesselin in chicken smooth muscle. Histochem. Cell Biol. 131, 191–196 (2009). [CrossRef]
- Li, L., Stefan, M. I. & Le Novère, N. Calcium Input Frequency, Duration and Amplitude Differentially Modulate the Relative Activation of Calcineurin and CaMKII. PLoS ONE 7, e43810 (2012). [CrossRef]
- Lehne, F. & Bogdan, S. Getting cells into shape by calcium-dependent actin cross-linking proteins. Front. Cell Dev. Biol. 11, (2023). [CrossRef]
- Elsaraj, S. M. & Bhullar, R. P. Regulation of platelet Rac1 and Cdc42 activation through interaction with calmodulin. Biochim. Biophys. Acta BBA - Mol. Cell Res. 1783, 770–778 (2008). [CrossRef]
- Landau, A. T. et al. Dendritic branch structure compartmentalizes voltage-dependent calcium influx in cortical layer 2/3 pyramidal cells. eLife 11, e76993 (2022).
- Izadi, M. et al. Cobl-like promotes actin filament formation and dendritic branching using only a single WH2 domain. J. Cell Biol. 217, 211 (2018). [CrossRef]
- Hepler, P. K. The Cytoskeleton and Its Regulation by Calcium and Protons. Plant Physiol. 170, 3–22 (2016). [CrossRef]
- Scheible, N. & McCubbin, A. Signaling in Pollen Tube Growth: Beyond the Tip of the Polarity Iceberg. Plants 8, 156 (2019). [CrossRef]
- Xiang, Y. et al. ACTIN BINDING PROTEIN29 from Lilium Pollen Plays an Important Role in Dynamic Actin Remodeling. Plant Cell 19, 1930–1946 (2007). [CrossRef]
- Qu, X. et al. Organization and regulation of the actin cytoskeleton in the pollen tube. Front. Plant Sci. 5, (2015). [CrossRef]
- Zhu, J. & Geisler, M. Keeping it all together: auxin–actin crosstalk in plant development. J. Exp. Bot. 66, 4983–4998 (2015). [CrossRef]
- Harrington, W. F. & Rodgers, M. E. Myosin. Annu. Rev. Biochem. 53, 35–73 (1984).
- Heissler, S. M. & Sellers, J. R. Kinetic Adaptations of Myosins for their Diverse Cellular Functions. Traffic Cph. Den. 17, 839–859 (2016). [CrossRef]
- Berg, J. S., Powell, B. C. & Cheney, R. E. A Millennial Myosin Census. Mol. Biol. Cell 12, 780–794 (2001). [CrossRef]
- Sellers, J. R. & Goodson, H. V. Motor proteins 2: myosin. Protein Profile 2, 1323–1423 (1995).
- Woolner, S. & Bement, W. M. Unconventional myosins acting unconventionally. Trends Cell Biol. 19, 245–252 (2009). [CrossRef]
- Hartman, M. A. & Spudich, J. A. The myosin superfamily at a glance. J. Cell Sci. 125, 1627–1632 (2012). [CrossRef]
- Ross, J. L., Ali, M. Y. & Warshaw, D. M. Cargo transport: molecular motors navigate a complex cytoskeleton. Curr. Opin. Cell Biol. 20, 41–47 (2008). [CrossRef]
- Bresnick, A. R. Molecular mechanisms of nonmuscle myosin-II regulation. Curr. Opin. Cell Biol. 11, 26–33 (1999). [CrossRef]
- Somlyo, A. P. & Somlyo, A. V. Ca2+ Sensitivity of Smooth Muscle and Nonmuscle Myosin II: Modulated by G Proteins, Kinases, and Myosin Phosphatase. Physiol. Rev. 83, 1325–1358 (2003).
- Collins, K. & Matsudaira, P. Differential regulation of vertebrate myosins I and II. J. Cell Sci. 1991, 11–16 (1991). [CrossRef]
- Williams, M. L. K. & Solnica-Krezel, L. Cellular and molecular mechanisms of convergence and extension in zebrafish. Curr. Top. Dev. Biol. 136, 377–407 (2020).
- Heissler, S. M., Chinthalapudi, K. & Sellers, J. R. Kinetic characterization of the sole nonmuscle myosin-2 from the model organism Drosophila melanogaster. FASEB J. 29, 1456–1466 (2015). [CrossRef]
- Hu, J. et al. Distinct roles of two myosins in C. elegans spermatid differentiation. PLOS Biol. 17, e3000211 (2019). [CrossRef]
- Eliott, S., Vardy, P. H. & Williams, K. L. The distribution of myosin II in Dictyostelium discoideum slug cells. J. Cell Biol. 115, 1267–1274 (1991). [CrossRef]
- May, K. M., Win, T. Z. & Hyams, J. S. Yeast myosin II: A new subclass of unconventional conventional myosins? Cell Motil. Cytoskeleton 39, 195–200 (1998).
- Krementsov, D. N., Krementsova, E. B. & Trybus, K. M. Myosin V : regulation by calcium, calmodulin, and the tail domain. J. Cell Biol. 164, 877–886 (2004).
- Batters, C., Brack, D., Ellrich, H., Averbeck, B. & Veigel, C. Calcium can mobilize and activate myosin-VI. Proc. Natl. Acad. Sci. 113, E1162–E1169 (2016). [CrossRef]
- Lapierre, L. A. et al. Myosin vb is associated with plasma membrane recycling systems. Mol. Biol. Cell 12, 1843–1857 (2001). [CrossRef]
- Buss, F., Arden, S. D., Lindsay, M., Luzio, J. P. & Kendrick-Jones, J. Myosin VI isoform localized to clathrin-coated vesicles with a role in clathrin-mediated endocytosis. EMBO J. 20, 3676–3684 (2001).
- Tumbarello, D. A. et al. Autophagy receptors link myosin VI to autophagosomes to mediate Tom1-dependent autophagosome maturation and fusion with the lysosome. Nat. Cell Biol. 14, 1024–1035 (2012). [CrossRef]
- Kruppa, A. J., Kendrick-Jones, J. & Buss, F. Myosins, Actin and Autophagy. Traffic 17, 878–890 (2016).
- Tumbarello, D. A., Kendrick-Jones, J. & Buss, F. Myosin VI and its cargo adaptors – linking endocytosis and autophagy. J. Cell Sci. 126, 2561–2570 (2013). [CrossRef]
- Tang, F. et al. Regulated degradation of a class V myosin receptor directs movement of the yeast vacuole. Nature 422, 87–92 (2003). [CrossRef]
- Tóth, J., Kovács, M., Wang, F., Nyitray, L. & Sellers, J. R. Myosin V from Drosophila Reveals Diversity of Motor Mechanisms within the Myosin V Family *. J. Biol. Chem. 280, 30594–30603 (2005). [CrossRef]
- Reck-Peterson, S. L., Provance, D. W., Mooseker, M. S. & Mercer, J. A. Class V myosins. Biochim. Biophys. Acta BBA - Mol. Cell Res. 1496, 36–51 (2000).
- Terrak, M., Rebowski, G., Lu, R. C., Grabarek, Z. & Dominguez, R. Structure of the light chain-binding domain of myosin V. Proc. Natl. Acad. Sci. 102, 12718–12723 (2005). [CrossRef]
- Van Gele, M., Dynoodt, P. & Lambert, J. Griscelli syndrome: a model system to study vesicular trafficking. Pigment Cell Melanoma Res. 22, 268–282 (2009). [CrossRef]
- Bookwalter, C. S., Lord, M. & Trybus, K. M. Essential Features of the Class V Myosin from Budding Yeast for ASH1 mRNA Transport. Mol. Biol. Cell 20, 3414–3421 (2009). [CrossRef]
- Ji, H.-H., Zhang, H.-M., Shen, M., Yao, L.-L. & Li, X. The motor function of Drosophila melanogaster myosin-5 is activated by calcium and cargo-binding protein dRab11. Biochem. J. 469, 135–144 (2015). [CrossRef]
- Wells, A. L. et al. Myosin VI is an actin-based motor that moves backwards. Nature 401, 505–508 (1999). [CrossRef]
- Ménétrey, J. et al. The structure of the myosin VI motor reveals the mechanism of directionality reversal. Nature 435, 779–785 (2005). [CrossRef]
- Magistrati, E. & Polo, S. Myomics: myosin VI structural and functional plasticity. Curr. Opin. Struct. Biol. 67, 33–40 (2021). [CrossRef]
- Heintzelman, M. B., Hasson, T. & Mooseker, M. S. Multiple unconventional myosin domains of the intestinal brush border cytoskeleton. J. Cell Sci. 107, 3535–3543 (1994). [CrossRef]
- Hasson, T. et al. Unconventional Myosins in Inner-Ear Sensory Epithelia. J. Cell Biol. 137, 1287–1307 (1997). [CrossRef]
- Finan, D., Hartman, M. A. & Spudich, J. A. Proteomics approach to study the functions of Drosophila myosin VI through identification of multiple cargo-binding proteins. Proc. Natl. Acad. Sci. 108, 5566–5571 (2011). [CrossRef]
- Lewis, J. H., Greenberg, M. J., Laakso, J. M., Shuman, H. & Ostap, E. M. Calcium Regulation of Myosin-I Tension Sensing. Biophys. J. 102, 2799–2807 (2012). [CrossRef]
- Yang, Y., Kovács, M., Xu, Q., Anderson, J. B. & Sellers, J. R. Myosin VIIB from Drosophila Is a High Duty Ratio Motor *. J. Biol. Chem. 280, 32061–32068 (2005). [CrossRef]
- Yokota, E., Muto, S. & Shimmen, T. Inhibitory Regulation of Higher-Plant Myosin by Ca2+Ions1. Plant Physiol. 119, 231–240 (1999). [CrossRef]
- Nebenführ, A. & Dixit, R. Kinesins and Myosins: Molecular Motors that Coordinate Cellular Functions in Plants. Annu. Rev. Plant Biol. 69, 329–361 (2018). [CrossRef]
- Brodland, G. W. How computational models can help unlock biological systems. Semin. Cell Dev. Biol. 47–48, 62–73 (2015). [CrossRef]
- Kumar, N., Mim, M. S., Dowling, A. & Zartman, J. J. Reverse engineering morphogenesis through Bayesian optimization of physics-based models. Npj Syst. Biol. Appl. 10, 1–14 (2024). [CrossRef]
- Sprague, B. L., Pego, R. L., Stavreva, D. A. & McNally, J. G. Analysis of Binding Reactions by Fluorescence Recovery after Photobleaching. Biophys. J. 86, 3473–3495 (2004).
- Ishikawa-Ankerhold, H., Ankerhold, R. & Drummen, G. Fluorescence Recovery After Photobleaching (FRAP). in eLS (John Wiley & Sons, Ltd, 2014). [CrossRef]
- Scipioni, L., Di Bona, M., Vicidomini, G., Diaspro, A. & Lanzanò, L. Local raster image correlation spectroscopy generates high-resolution intracellular diffusion maps. Commun. Biol. 1, 1–10 (2018).
- Brown, C. M. et al. Raster image correlation spectroscopy (RICS) for measuring fast protein dynamics and concentrations with a commercial laser scanning confocal microscope. J. Microsc. 229, 78–91 (2008).
- Williamson, D. E., Sahai, E., Jenkins, R. P., O’Dea, R. D. & King, J. R. Parameter estimation in fluorescence recovery after photobleaching: quantitative analysis of protein binding reactions and diffusion. J. Math. Biol. 83, 1 (2021).
- Digman, M. A. & Gratton, E. Analysis of diffusion and binding in cells using the RICS approach. Microsc. Res. Tech. 72, 323–332 (2009).
- Lillacci, G. & Khammash, M. Parameter Estimation and Model Selection in Computational Biology. PLOS Comput. Biol. 6, e1000696 (2010).
- Linden, N. J., Kramer, B. & Rangamani, P. Bayesian parameter estimation for dynamical models in systems biology. PLOS Comput. Biol. 18, e1010651 (2022).
- Browning, A. P. & Simpson, M. J. Geometric analysis enables biological insight from complex non-identifiable models using simple surrogates. PLOS Comput. Biol. 19, e1010844 (2023).
- Rmus, M., Pan, T.-F., Xia, L. & Collins, A. G. E. Artificial neural networks for model identification and parameter estimation in computational cognitive models. PLOS Comput. Biol. 20, e1012119 (2024).
- Yazdani, A., Lu, L., Raissi, M. & Karniadakis, G. E. Systems biology informed deep learning for inferring parameters and hidden dynamics. PLOS Comput. Biol. 16, e1007575 (2020).
- Li, L. et al. Determining the role of advection in patterning by bone morphogenetic proteins through neural network model-based acceleration of a 3D finite element model of the zebrafish embryo. Front. Syst. Biol. 2, (2022).
- Kristiansson, E. et al. A novel method for cross-species gene expression analysis. BMC Bioinformatics 14, 70 (2013).
- Reynolds, K. A. et al. Accelerating Biological Insight for Understudied Genes. Integr. Comp. Biol. 61, 2233–2243 (2021).
- Merzbacher, C., Mac Aodha, O. & Oyarzún, D. A. Bayesian Optimization for Design of Multiscale Biological Circuits. ACS Synth. Biol. 12, 2073–2082 (2023).
- Li, L., Mdluli, T., Buzzard, G. & Umulis, D. Digital Cousins: Simultaneous Optimization of One Model for BMP Signaling in Distant Relatives Reveals Essential Core. 2025.02.25.640248 Preprint at (2025). [CrossRef]
- Shaikh, R. et al. Optimal performance objectives in the highly conserved bone morphogenetic protein signaling pathway. Npj Syst. Biol. Appl. 10, 1–13 (2024).
- St. Mary, C., Powell, T. H. Q., Kominoski, J. S. & Weinert, E. Rescaling Biology: Increasing Integration Across Biological Scales and Subdisciplines to Enhance Understanding and Prediction. Integr. Comp. Biol. 61, 2031–2037 (2021).
- Hassoun, S. et al. Artificial Intelligence for Biology. Integr. Comp. Biol. Volume 61, Pages 2267-2275 (2021).
- Qian, D. & Xiang, Y. Actin Cytoskeleton as Actor in Upstream and Downstream of Calcium Signaling in Plant Cells. Int. J. Mol. Sci. 20, 1403 (2019).
- DerMardirossian, C. The Role of Calcium in Actin-Dependent Cell Migration and Invasion in Cancer. (IntechOpen, 2024). [CrossRef]
- Yamamoto, K., Korenaga, R., Kamiya, A. & Ando, J. Fluid Shear Stress Activates Ca2+ Influx Into Human Endothelial Cells via P2X4 Purinoceptors. Circ. Res. 87, 385–391 (2000).
- Dull, R. O. & Davies, P. F. Flow modulation of agonist (ATP)-response (Ca2+) coupling in vascular endothelial cells. Am. J. Physiol.-Heart Circ. Physiol. 261, H149–H154 (1991).
- Mo, M., Eskin, S. G. & Schilling, W. P. Flow-induced changes in Ca2+ signaling of vascular endothelial cells: effect of shear stress and ATP. Am. J. Physiol. 260, H1698-1707 (1991).
- Liu, S. et al. Components of the Calcium-Calcineurin Signaling Pathway in Fungal Cells and Their Potential as Antifungal Targets. Eukaryot. Cell 14, 324–334 (2015).
- Giuraniuc, C. V. et al. Dynamic calcium-mediated stress response and recovery signatures in the fungal pathogen, Candida albicans. mBio 14, e01157-23 (2023).
- Liu, J. et al. Ca2+ channels and Ca2+ signals involved in abiotic stress responses in plant cells: recent advances. Plant Cell Tissue Organ Cult. PCTOC 132, 413–424 (2018).
- Park, C.-J. & Shin, R. Calcium channels and transporters: Roles in response to biotic and abiotic stresses. Front. Plant Sci. 13, (2022).
- Ranty, B. et al. Calcium Sensors as Key Hubs in Plant Responses to Biotic and Abiotic Stresses. Front. Plant Sci. 7, (2016).
- Ghosh, P. & Roychoudhury, A. Plant Peptides Involved in Abiotic and Biotic Stress Responses and Reactive Oxygen Species (ROS) Signaling. J. Plant Growth Regul. 43, 1410–1427 (2024).
- WERRY, T. D., WILKINSON, G. F. & WILLARS, G. B. Mechanisms of cross-talk between G-protein-coupled receptors resulting in enhanced release of intracellular Ca2+. Biochem. J. 374, 281–296 (2003).
- Ordenes, V. R., Reyes, F. C., Wolff, D. & Orellana, A. A Thapsigargin-Sensitive Ca2+ Pump Is Present in the Pea Golgi Apparatus Membrane. Plant Physiol. 129, 1820–1828 (2002).
- Yamarnoto, N. et al. A study on thapsigargin-induced calcium ion and cation influx pathways in vascular endothelial cells. Biochim. Biophys. Acta BBA - Mol. Cell Res. 1266, 157–162 (1995).
- Andersen, T., López, C., Manczak, T., Martinez, K. & Simonsen, H. Thapsigargin—From Thapsia L. to Mipsagargin. Molecules 20, 6113–6127 (2015).
- Shao, X., Li, Q., Mogilner, A., Bershadsky, A. D. & Shivashankar, G. V. Mechanical stimulation induces formin-dependent assembly of a perinuclear actin rim. Proc. Natl. Acad. Sci. 112, E2595–E2601 (2015).
- Small, A. G. et al. Neutrophils Require Activation to Express Functional Cell-Surface Complement Receptor Immunoglobulin. Front. Immunol. 13, (2022).
- Li, J. & Staiger, C. J. Understanding Cytoskeletal Dynamics During the Plant Immune Response. Annu. Rev. Phytopathol. 56, 513–533 (2018).
- Hardham, A. R., Takemoto, D. & White, R. G. Rapid and dynamic subcellular reorganization following mechanical stimulation of Arabidopsisepidermal cells mimics responses to fungal and oomycete attack. BMC Plant Biol. 8, 63 (2008).
- Bellandi, A. et al. Diffusion and bulk flow of amino acids mediate calcium waves in plants. Sci. Adv. 8, eabo6693 (2022).
- Cao, L., Wang, W., Zhang, W. & Staiger, C. J. Lipid Signaling Requires ROS Production to Elicit Actin Cytoskeleton Remodeling during Plant Innate Immunity. Int. J. Mol. Sci. 23, 2447 (2022).
- Lu, Y.-J. et al. Arabidopsis calcium-dependent protein kinase 3 regulates actin cytoskeleton organization and immunity. Nat. Commun. 11, 6234 (2020).
- Snowman, B. N., Kovar, D. R., Shevchenko, G., Franklin-Tong, V. E. & Staiger, C. J. Signal-Mediated Depolymerization of Actin in Pollen during the Self-Incompatibility Response. Plant Cell 14, 2613 (2002).
- Huang, S., Blanchoin, L., Chaudhry, F., Franklin-Tong, V. E. & Staiger, C. J. A Gelsolin-like Protein from Papaver rhoeas Pollen (PrABP80) Stimulates Calcium-regulated Severing and Depolymerization of Actin Filaments. J. Biol. Chem. 279, 23364–23375 (2004).
- Handly, L. N. & Wollman, R. Wound-induced Ca2+ wave propagates through a simple release and diffusion mechanism. Mol. Biol. Cell 28, 1457–1466 (2017).
- Lansdown, A. B. G. Calcium: a potential central regulator in wound healing in the skin. Wound Repair Regen. 10, 271–285 (2002).
- Rodrigues, M., Kosaric, N., Bonham, C. A. & Gurtner, G. C. Wound Healing: A Cellular Perspective. Physiol. Rev. 99, 665–706 (2019).
- Xu, S. & Chisholm, A. D. A Gαq-Ca2+ signaling pathway promotes actin-mediated epidermal wound closure in C. elegans. Curr. Biol. CB 21, 1960–1967 (2011).
- Jiang, D. et al. Injury triggers fascia fibroblast collective cell migration to drive scar formation through N-cadherin. Nat. Commun. 11, 5653 (2020).
- Aj, C. Role of the actin cytoskeleton in wound healing and scar formation. 14, (2006).
- Martinotti, S., Laforenza, U., Patrone, M., Moccia, F. & Ranzato, E. Honey-Mediated Wound Healing: H2O2 Entry through AQP3 Determines Extracellular Ca2+ Influx. Int. J. Mol. Sci. 20, 764 (2019).
- Jain, J. et al. The T-cell transcription factor NFATp is a substrate for calcineurin and interacts with Fos and Jun. Nature 365, 352–355 (1993).
- Friday, B. B., Mitchell, P. O., Kegley, K. M. & Pavlath, G. K. Calcineurin initiates skeletal muscle differentiation by activating MEF2 and MyoD. Differentiation 71, 217–227 (2003).
- Sanz-Morejón, A. & Mercader, N. Recent insights into zebrafish cardiac regeneration. Curr. Opin. Genet. Dev. 64, 37–43 (2020).
- Sun, J. et al. Zebrafish heart regeneration after coronary dysfunction-induced cardiac damage. Dev. Biol. 487, 57–66 (2022).
- Ding, C. et al. Reactive Oxygen Species Counteract Wound Contraction and Promote Wound Healing. 2025.03.27.645726 Preprint at (2025). [CrossRef]
- Clarkson, M. R. et al. High Glucose-altered Gene Expression in Mesangial Cells: ACTIN-REGULATORY PROTEIN GENE EXPRESSION IS TRIGGERED BY OXIDATIVE STRESS AND CYTOSKELETAL DISASSEMBLY*. J. Biol. Chem. 277, 9707–9712 (2002).
- Yamanaka, T., Ueki, T., Mase, M. & Inoue, K. Arbitrary Ca2+ regulation for endothelial nitric oxide, NFAT and NF-κB activities by an optogenetic approach. Front. Pharmacol. 13, (2023).
- Ma, T. C., Barco, A., Ratan, R. R. & Willis, D. E. cAMP-responsive element-binding protein (CREB) and cAMP co-regulate activator protein 1 (AP1)-dependent regeneration-associated gene expression and neurite growth. J. Biol. Chem. 289, 32914–32925 (2014).
- Schäfer, M. & Werner, S. Oxidative stress in normal and impaired wound repair. Pharmacol. Res. 58, 165–171 (2008).
- Dunnill, C. et al. Reactive oxygen species (ROS) and wound healing: the functional role of ROS and emerging ROS-modulating technologies for augmentation of the healing process. Int. Wound J. 14, 89–96 (2017).
- O’Connor, J. T. et al. Proteolytic activation of Growth-blocking peptides triggers calcium responses through the GPCR Mthl10 during epithelial wound detection. Dev. Cell 56, 2160-2175.e5 (2021).
- Velagala, V. et al. The multimodal action of G alpha q in coordinating growth and homeostasis in the Drosophila wing imaginal disc. 2023.01.08.523049 Preprint at (2023). [CrossRef]
- Zhao, S. et al. Regulation of Plant Responses to Salt Stress. Int. J. Mol. Sci. 22, 4609 (2021).
- Kanzawa, N. et al. Change in the Actin Cytoskeleton during Seismonastic Movement of Mimosa pudica. Plant Cell Physiol. 47, 531–539 (2006).
- Huang, Y., Wang, J., Zhang, L. & Zuo, K. A Cotton Annexin Protein AnxGb6 Regulates Fiber Elongation through Its Interaction with Actin 1. PLOS ONE 8, e66160 (2013).
- Dalghi, M. G., Ferreira-Gomes, M. & Rossi, J. P. Regulation of the Plasma Membrane Calcium ATPases by the actin cytoskeleton. Biochem. Biophys. Res. Commun. 506, 347–354 (2018).
- Vanagas, L. et al. Differential Effects of G- and F-Actin on the Plasma Membrane Calcium Pump Activity. Cell Biochem. Biophys. 66, 187–198 (2013).
- Dean, W. L. Role of platelet plasma membrane Ca2+-ATPase in health and disease. World J. Biol. Chem. 1, 265–270 (2010).
- Dean, W. L., Chen, D., Brandt, P. C. & Vanaman, T. C. Regulation of Platelet Plasma Membrane Ca2+-ATPase by cAMP-dependent and Tyrosine Phosphorylation*. J. Biol. Chem. 272, 15113–15119 (1997).
- Stevens, A. C., O’Connor, J. T., Pumford, A. D., Page-McCaw, A. & Hutson, M. S. A mathematical model of calcium signals around laser-induced epithelial wounds. Mol. Biol. Cell 34, ar49 (2023).
- Bai, J. & Zeng, X. Computational modeling and simulation of epithelial wound closure. Sci. Rep. 13, 6265 (2023).
- Alber, M. et al. Integrating machine learning and multiscale modeling—perspectives, challenges, and opportunities in the biological, biomedical, and behavioral sciences. Npj Digit. Med. 2, 1–11 (2019).
- Yumura, S. et al. Dynamics of Actin Cytoskeleton and Their Signaling Pathways during Cellular Wound Repair. Cells 11, 3166 (2022).
- Dustin, M. L. Signaling at neuro/immune synapses. J. Clin. Invest. 122, 1149–1155 (2012).
- Krauchunas, A. R., Marcello, M. R. & Singson, A. The molecular complexity of fertilization: Introducing the concept of a fertilization synapse. Mol. Reprod. Dev. 83, 376–386 (2016).
- Südhof, T. C. Towards an Understanding of Synapse Formation. Neuron 100, 276–293 (2018).
- Capitani, N. & Baldari, C. T. The Immunological Synapse: An Emerging Target for Immune Evasion by Bacterial Pathogens. Front. Immunol. 13, (2022).
- Dustin, M. L. The immunological synapse. Cancer Immunol. Res. 2, 1023 (2014).
- Honkura, N., Matsuzaki, M., Noguchi, J., Ellis-Davies, G. C. R. & Kasai, H. The subspine organization of actin fibers regulates the structure and plasticity of dendritic spines. Neuron 57, 719–729 (2008).
- Borovac, J., Bosch, M. & Okamoto, K. Regulation of actin dynamics during structural plasticity of dendritic spines: Signaling messengers and actin-binding proteins. Mol. Cell. Neurosci. 91, 122–130 (2018).
- Gentile, J. E., Carrizales, M. G. & Koleske, A. J. Control of Synapse Structure and Function by Actin and Its Regulators. Cells 11, 603 (2022).
- Malenka, R. C. et al. An essential role for postsynaptic calmodulin and protein kinase activity in long-term potentiation. Nature 340, 554–557 (1989).
- Krapivinsky, G., Medina, I., Krapivinsky, L., Gapon, S. & Clapham, D. E. SynGAP-MUPP1-CaMKII Synaptic Complexes Regulate p38 MAP Kinase Activity and NMDA Receptor- Dependent Synaptic AMPA Receptor Potentiation. Neuron 43, 563–574 (2004).
- Oh, J. S., Manzerra, P. & Kennedy, M. B. Regulation of the neuron-specific Ras GTPase-activating protein, synGAP, by Ca2+/calmodulin-dependent protein kinase II. J. Biol. Chem. 279, 17980–17988 (2004).
- Liu, A. et al. Neuroligin 1 regulates spines and synaptic plasticity via LIMK1/cofilin-mediated actin reorganization. J. Cell Biol. 212, 449–463 (2016).
- Khan, S., Downing, K. H. & Molloy, J. E. Architectural Dynamics of CaMKII-Actin Networks. Biophys. J. 116, 104–119 (2019).
- Yamazaki, H., Koganezawa, N., Yokoo, H., Sekino, Y. & Shirao, T. Super-resolution imaging reveals the relationship between CaMKIIβ and drebrin within dendritic spines. Neurosci. Res. 199, 30–35 (2024).
- Meng, Y. et al. Abnormal Spine Morphology and Enhanced LTP in LIMK-1 Knockout Mice. Neuron 35, 121–133 (2002).
- Südhof, T. C. The presynaptic active zone. Neuron 75, 11–25 (2012).
- Prekeris, R. & Terrian, D. M. Brain myosin V is a synaptic vesicle-associated motor protein: evidence for a Ca2+-dependent interaction with the synaptobrevin-synaptophysin complex. J. Cell Biol. 137, 1589–1601 (1997).
- von Gersdorff, H. & Matthews, G. Dynamics of synaptic vesicle fusion and membrane retrieval in synaptic terminals. Nature 367, 735–739 (1994).
- Gersdorff, H. von & Matthews, G. Depletion and Replenishment of Vesicle Pools at a Ribbon-Type Synaptic Terminal. J. Neurosci. 17, 1919 (1997).
- Wang, L.-Y., Neher, E. & Taschenberger, H. Synaptic Vesicles in Mature Calyx of Held Synapses Sense Higher Nanodomain Calcium Concentrations during Action Potential-Evoked Glutamate Release. J. Neurosci. 28, 14450–14458 (2008).
- Wu, L.-G. & Chan, C. Y. Multiple Roles of Actin in Exo- and Endocytosis. Front. Synaptic Neurosci. 14, (2022).
- Chapman, E. R. Synaptotagmin: a Ca(2+) sensor that triggers exocytosis? Nat. Rev. Mol. Cell Biol. 3, 498–508 (2002).
- Bajjalieh, S. M. Synaptic vesicle docking and fusion. Curr. Opin. Neurobiol. 9, 321–328 (1999).
- Srinivasan, G., Kim, J. H. & von Gersdorff, H. The pool of fast releasing vesicles is augmented by myosin light chain kinase inhibition at the calyx of Held synapse. J. Neurophysiol. 99, 1810–1824 (2008).
- Mainen, Z. F., Malinow, R. & Svoboda, K. Synaptic calcium transients in single spines indicate that NMDA receptors are not saturated. Nature 399, 151–155 (1999).
- Sobczyk, A., Scheuss, V. & Svoboda, K. NMDA receptor subunit-dependent [Ca2+] signaling in individual hippocampal dendritic spines. J. Neurosci. Off. J. Soc. Neurosci. 25, 6037–6046 (2005).
- Noguchi, J., Matsuzaki, M., Ellis-Davies, G. C. R. & Kasai, H. Spine-neck geometry determines NMDA receptor-dependent Ca2+ signaling in dendrites. Neuron 46, 609–622 (2005).
- Sabatini, B. L., Oertner, T. G. & Svoboda, K. The life cycle of Ca(2+) ions in dendritic spines. Neuron 33, 439–452 (2002).
- Béïque, J.-C. et al. Synapse-specific regulation of AMPA receptor function by PSD-95. Proc. Natl. Acad. Sci. 103, 19535–19540 (2006).
- Chen, X. et al. PSD-95 family MAGUKs are essential for anchoring AMPA and NMDA receptor complexes at the postsynaptic density. Proc. Natl. Acad. Sci. U. S. A. 112, E6983-6992 (2015).
- Chazeau, A. et al. Nanoscale segregation of actin nucleation and elongation factors determines dendritic spine protrusion. EMBO J. 33, 2745 (2014).
- Hung, A. Y., Sung, C. C., Brito, I. L. & Sheng, M. Degradation of Postsynaptic Scaffold GKAP and Regulation of Dendritic Spine Morphology by the TRIM3 Ubiquitin Ligase in Rat Hippocampal Neurons. PLoS ONE 5, e9842 (2010).
- Sala, C. et al. Regulation of Dendritic Spine Morphology and Synaptic Function by Shank and Homer. Neuron 31, 115–130 (2001).
- Roussignol, G. et al. Shank Expression Is Sufficient to Induce Functional Dendritic Spine Synapses in Aspiny Neurons. J. Neurosci. 25, 3560–3570 (2005).
- Luo, P., Li, X., Fei, Z. & Poon, W. Scaffold protein Homer 1: Implications for neurological diseases. Neurochem. Int. 61, 731–738 (2012).
- Hayashi, M. K. et al. The Postsynaptic Density Proteins Homer and Shank Form a Polymeric Network Structure. Cell 137, 159–171 (2009).
- Kim, I. H. et al. Disruption of Arp2/3 Results in Asymmetric Structural Plasticity of Dendritic Spines and Progressive Synaptic and Behavioral Abnormalities. J. Neurosci. 33, 6081–6092 (2013).
- Sørensen, J. B. et al. Differential control of the releasable vesicle pools by SNAP-25 splice variants and SNAP-23. Cell 114, 75–86 (2003).
- Mesa, M. H., McCabe, K. J. & Rangamani, P. Synaptic cleft geometry modulates NMDAR opening probability by tuning neurotransmitter residence time. Biophys. J. 0, (2025).
- Hammer, J. A., Wang, J. C., Saeed, M. & Pedrosa, A. T. Origin, Organization, Dynamics, and Function of Actin and Actomyosin Networks at the T Cell Immunological Synapse. Annu. Rev. Immunol. 37, 201–224 (2019).
- Huppa, J. B. & Davis, M. M. T-cell-antigen recognition and the immunological synapse. Nat. Rev. Immunol. 3, 973–983 (2003).
- Grakoui, A. et al. The Immunological Synapse: A Molecular Machine Controlling T Cell Activation. Science 285, 221–227 (1999).
- Piragyte, I. & Jun, C.-D. Actin Engine in Immunological Synapse. Immune Netw. 12, 71–83 (2012).
- Cassioli, C. & Baldari, C. T. Lymphocyte Polarization During Immune Synapse Assembly: Centrosomal Actin Joins the Game. Front. Immunol. 13, 830835 (2022).
- Billadeau, D. D., Nolz, J. C. & Gomez, T. S. Regulation of T-cell activation by the cytoskeleton. Nat. Rev. Immunol. 7, 131–143 (2007).
- Dustin, M. L. Cell adhesion molecules and actin cytoskeleton at immune synapses and kinapses. Curr. Opin. Cell Biol. 19, 529–533 (2007).
- Trebak, M. & Kinet, J.-P. Calcium signalling in T cells. Nat. Rev. Immunol. 19, 154–169 (2019).
- Babich, A. et al. F-actin polymerization and retrograde flow drive sustained PLCγ1 signaling during T cell activation. J. Cell Biol. 197, 775–787 (2012).
- Hartzell, C. A., Jankowska, K. I., Burkhardt, J. K. & Lewis, R. S. Calcium influx through CRAC channels controls actin organization and dynamics at the immune synapse. eLife 5, e14850 (2016).
- Luik, R. M., Wu, M. M., Buchanan, J. & Lewis, R. S. The elementary unit of store-operated Ca2+ entry: local activation of CRAC channels by STIM1 at ER–plasma membrane junctions. J. Cell Biol. 174, 815–825 (2006).
- Wabnitz, G. H. et al. Sustained LFA-1 cluster formation in the immune synapse requires the combined activities of L-plastin and calmodulin. Eur. J. Immunol. 40, 2437–2449 (2010).
- Wang, Y., Shibasaki, F. & Mizuno, K. Calcium Signal-induced Cofilin Dephosphorylation Is Mediated by Slingshot via Calcineurin *. J. Biol. Chem. 280, 12683–12689 (2005).
- Ishiguro, K. et al. Ca2+/Calmodulin-Dependent Protein Kinase II Is a Modulator of CARMA1-Mediated NF-κB Activation. Mol. Cell. Biol. 26, 5497–5508 (2006).
- Blumenthal, D. & Burkhardt, J. K. Multiple actin networks coordinate mechanotransduction at the immunological synapse. J. Cell Biol. 219, e201911058 (2020).
- Calvo, V. & Izquierdo, M. Inducible Polarized Secretion of Exosomes in T and B Lymphocytes. Int. J. Mol. Sci. 21, 2631 (2020).
- Valitutti, S., Coombs, D. & Dupré, L. The space and time frames of T cell activation at the immunological synapse. FEBS Lett. 584, 4851–4857 (2010).
- Short, B. Actin works both sides of the immunological synapse. J. Cell Biol. 208, 383 (2015).
- Lyubchenko, T. A., Wurth, G. A. & Zweifach, A. Role of Calcium Influx in Cytotoxic T Lymphocyte Lytic Granule Exocytosis during Target Cell Killing. Immunity 15, 847–859 (2001).
- Pores-Fernando, A. T. & Zweifach, A. Calcium influx and signaling in cytotoxic T-lymphocyte lytic granule exocytosis. Immunol. Rev. 231, 160–173 (2009).
- Kim, K.-D. et al. Targeted calcium influx boosts cytotoxic T lymphocyte function in the tumour microenvironment. Nat. Commun. 8, 15365 (2017).
- Stinchcombe, J. C., Majorovits, E., Bossi, G., Fuller, S. & Griffiths, G. M. Centrosome polarization delivers secretory granules to the immunological synapse. Nature 443, 462–465 (2006).
- Krzewski, K. & Coligan, J. E. Human NK cell lytic granules and regulation of their exocytosis. Front. Immunol. 3, (2012).
- Chang, H.-F., Schirra, C., Pattu, V., Krause, E. & Becherer, U. Lytic granule exocytosis at immune synapses: lessons from neuronal synapses. Front. Immunol. 14, 1177670 (2023).
- Stein, P., Savy, V., Williams, A. M. & Williams, C. J. Modulators of calcium signalling at fertilization. Open Biol. 10, 200118 (2020).
- Wakai, T., Vanderheyden, V. & Fissore, R. A. Ca2+ Signaling During Mammalian Fertilization: Requirements, Players, and Adaptations. Cold Spring Harb. Perspect. Biol. 3, a006767 (2011).
- Hogben, M., Parrington, J., Shevchenko, V., Swann, K. & Lai, F. A. Calcium oscillations, sperm factors and egg activation at fertilisation. J. Mol. Med. 76, 548–554 (1998).
- Swann, K. & Lai, F. A. Egg Activation at Fertilization by a Soluble Sperm Protein. Physiol. Rev. 96, 127–149 (2016).
- Parekh, A. B. & Putney, J. W. Store-Operated Calcium Channels. Physiol. Rev. 85, 757–810 (2005).
- Bernhardt, M. L. et al. TRPM7 and CaV3.2 channels mediate Ca2+ influx required for egg activation at fertilization. Proc. Natl. Acad. Sci. 115, E10370–E10378 (2018).
- Shafqat, A., Kashir, J., Alsalameh, S., Alkattan, K. & Yaqinuddin, A. Fertilization, Oocyte Activation, Calcium Release and Epigenetic Remodelling: Lessons From Cancer Models. Front. Cell Dev. Biol. 10, 781953 (2022).
- Lee, H. C., Yoon, S.-Y., Lykke-Hartmann, K., Fissore, R. A. & Carvacho, I. TRPV3 channels mediate Ca2+ influx induced by 2-APB in mouse eggs. Cell Calcium 59, 21–31 (2016).
- Backs, J. et al. The gamma isoform of CaM kinase II controls mouse egg activation by regulating cell cycle resumption. Proc. Natl. Acad. Sci. U. S. A. 107, 81–86 (2010).
- Abbott, A. L. & Ducibella, T. Calcium and the control of mammalian cortical granule exocytosis. Front. Biosci. J. Virtual Libr. 6, D792-806 (2001).
- Ducibella, T., Kurasawa, S., Duffy, P., Kopf, G. S. & Schultz, R. M. Regulation of the Polyspermy Block in the Mouse Egg: Maturation-Dependent Differences in Cortical Granule Exocytosis and Zona Pellucida Modifications Induced by Inositol 1,4,5-Trisphosphate and an Activator of Protein Kinase C1. Biol. Reprod. 48, 1251–1257 (1993).
- Vogt, E.-J. et al. Anchoring cortical granules in the cortex ensures trafficking to the plasma membrane for post-fertilization exocytosis. Nat. Commun. 10, 2271 (2019).
- Matson, S., Markoulaki, S. & Ducibella, T. Antagonists of Myosin Light Chain Kinase and of Myosin II Inhibit Specific Events of Egg Activation in Fertilized Mouse Eggs1. Biol. Reprod. 74, 169–176 (2006).
- Wolf, D. P. & Hamada, M. Induction of Zonal and Egg Plasma Membrane Blocks to Sperm Penetration in Mouse Eggs with Cortical Granule Exudate. Biol. Reprod. 17, 350–354 (1977).
- Burkart, A. D., Xiong, B., Baibakov, B., Jiménez-Movilla, M. & Dean, J. Ovastacin, a cortical granule protease, cleaves ZP2 in the zona pellucida to prevent polyspermy. J. Cell Biol. 197, 37–44 (2012).
- Ordyan, M., Bartol, T., Kennedy, M., Rangamani, P. & Sejnowski, T. Interactions between calmodulin and neurogranin govern the dynamics of CaMKII as a leaky integrator. PLoS Comput. Biol. 16, e1008015 (2020).
- Bell, M. K., Holst, M. V., Lee, C. T. & Rangamani, P. Dendritic spine morphology regulates calcium-dependent synaptic weight change. J. Gen. Physiol. 154, e202112980 (2022).
- Wang, Q. et al. Assemblies of calcium/calmodulin-dependent kinase II with actin and their dynamic regulation by calmodulin in dendritic spines. Proc. Natl. Acad. Sci. 116, 18937–18942 (2019).
- McMillen, P. & Levin, M. Collective intelligence: A unifying concept for integrating biology across scales and substrates. Commun. Biol. 7, 1–17 (2024).
- Mehta, P. & Gregor, T. Approaching the molecular origins of collective dynamics in oscillating cell populations. Curr. Opin. Genet. Dev. 20, 574–580 (2010).
- Riedl, C., Kim, Y. J., Gupta, P., Malone, T. W. & Woolley, A. W. Quantifying collective intelligence in human groups. Proc. Natl. Acad. Sci. 118, e2005737118 (2021).
- Loesel, K. E., Hiraki, H. L., Baker, B. M. & Parent, C. A. An adaptive and versatile method to quantitate and characterize collective cell migration behaviors on complex surfaces. Front. Cell Dev. Biol. 11, (2023).
- Zajdel, T. J., Shim, G., Wang, L., Rossello-Martinez, A. & Cohen, D. J. SCHEEPDOG: Programming Electric Cues to Dynamically Herd Large-Scale Cell Migration. Cell Syst. 10, 506-514.e3 (2020).
- Larkin, J. W. et al. Signal Percolation within a Bacterial Community. Cell Syst. 7, 137-145.e3 (2018).
- Plieth, C. Signal percolation through plants and the shape of the calcium signature. Plant Signal. Behav. 5, 379–385 (2010).
- Campanale, J. P. & Montell, D. J. Who’s really in charge: Diverse follower cell behaviors in collective cell migration. Curr. Opin. Cell Biol. 81, 102160 (2023).
- Qin, L., Yang, D., Yi, W., Cao, H. & Xiao, G. Roles of leader and follower cells in collective cell migration. Mol. Biol. Cell 32, 1267–1272 (2021).
- Coombes, S. & Timofeeva, Y. Sparks and waves in a stochastic fire-diffuse-fire model of Ca2+ release. Phys. Rev. E 68, 021915 (2003).
- Timofeeva, Y. & Coombes, S. Directed percolation in a two-dimensional stochastic fire-diffuse-fire model. Phys. Rev. E 70, 062901 (2004).
- Beyeler, M., Rounds, E. L., Carlson, K. D., Dutt, N. & Krichmar, J. L. Neural correlates of sparse coding and dimensionality reduction. PLOS Comput. Biol. 15, e1006908 (2019).
- Xu, Q. et al. Robust Transcoding Sensory Information With Neural Spikes. IEEE Trans. Neural Netw. Learn. Syst. 33, 1935–1946 (2022).
- Sætra, M. J. & Mori, Y. An electrodiffusive network model with multicompartmental neurons and synaptic connections. PLOS Comput. Biol. 20, e1012114 (2024).
- Tian, Y. & Sun, P. Percolation may explain efficiency, robustness, and economy of the brain. Netw. Neurosci. 6, 765–790 (2022).
- Olshausen, B. A. & Field, D. J. Sparse coding of sensory inputs. Curr. Opin. Neurobiol. 14, 481–487 (2004).
- Chabosseau, P. et al. Molecular phenotyping of single pancreatic islet leader beta cells by “Flash-Seq”. Life Sci. 316, 121436 (2023).
- Zhang, W. et al. Local traveling waves of cytosolic calcium elicited by defense signals or wounding are propagated by distinct mechanisms. 2025.01.23.634554 Preprint at (2025). [CrossRef]
- Seybold, H. et al. Ca2+ signalling in plant immune response: from pattern recognition receptors to Ca2+ decoding mechanisms. New Phytol. 204, 782–790 (2014).
- Choi, W.-G., Toyota, M., Kim, S.-H., Hilleary, R. & Gilroy, S. Salt stress-induced Ca2+ waves are associated with rapid, long-distance root-to-shoot signaling in plants. Proc. Natl. Acad. Sci. 111, 6497–6502 (2014).
- Evans, W. H. & and Martin, P. E. M. Gap junctions: structure and function (Review). Mol. Membr. Biol. 19, 121–136 (2002).
- Nielsen, M. S. et al. Gap Junctions. Compr. Physiol. 2, 10.1002/cphy.c110051 (2012).
- Meşe, G., Richard, G. & White, T. W. Gap Junctions: Basic Structure and Function. J. Invest. Dermatol. 127, 2516–2524 (2007).
- Sahu, A., Ghosh, R., Deshpande, G. & Prasad, M. A Gap Junction Protein, Inx2, Modulates Calcium Flux to Specify Border Cell Fate during Drosophila oogenesis. PLOS Genet. 13, e1006542 (2017).
- Robards, A. W. & Lucas, W. J. Plasmodesmata. Annu. Rev. Plant Biol. 41, 369–419 (1990).
- Cilia, M. L. & Jackson, D. Plasmodesmata form and function. Curr. Opin. Cell Biol. 16, 500–506 (2004).
- Tucker, E. B. & Boss, W. F. Mastoparan-Induced Intracellular Ca2+ Fluxes May Regulate Cell-to-Cell Communication in Plants. Plant Physiol. 111, 459–467 (1996).
- Holdaway-Clarke, T. L., Walker, N. A., Hepler, P. K. & Overall, R. L. Physiological elevations in cytoplasmic free calcium by cold or ion injection result in transient closure of higher plant plasmodesmata. Planta 210, 329–335 (2000).
- Xu, B. et al. A calmodulin-like protein regulates plasmodesmal closure during bacterial immune responses. New Phytol. 215, 77–84 (2017).
- Mayor, R. & Etienne-Manneville, S. The front and rear of collective cell migration. (2015).
- Tekotte, H., Tollervey, D. & Davis, I. Imaging the migrating border cell cluster in living Drosophila egg chambers. Dev. Dyn. 236, 2818–2824 (2007).
- Konen, J. et al. Image-guided genomics of phenotypically heterogeneous populations reveals vascular signalling during symbiotic collective cancer invasion. Nat. Commun. 8, 15078 (2017).
- McLennan, R. et al. Multiscale mechanisms of cell migration during development: theory and experiment. Development 139, 2935–2944 (2012).
- Alhashem, Z. et al. Notch controls the cell cycle to define leader versus follower identities during collective cell migration. eLife 11, e73550 (2022).
- Tsai, F.-C. et al. A polarized Ca2+, diacylglycerol and STIM1 signalling system regulates directed cell migration. Nat. Cell Biol. 16, 133–144 (2014).
- Yao, J., Pilko, A. & Wollman, R. Distinct cellular states determine calcium signaling response. Mol. Syst. Biol. 12, 894 (2016).
- Sonnemann, K. J. & Bement, W. M. Wound Repair: Toward Understanding and Integration of Single-Cell and Multicellular Wound Responses. Annu. Rev. Cell Dev. Biol. 27, 237–263 (2011).
- Cheng, K. C. et al. Promoting validation and cross-phylogenetic integration in model organism research. Dis. Model. Mech. 15, dmm049600 (2022).
- Sun, F. et al. Cross-species signaling pathways analysis inspire animal model selections for drug screening and target prediction in vascular aging diseases. Evol. Appl. 17, e13708 (2024).
- Bootman, M. D., Berridge, M. J. & Roderick, H. L. Calcium Signalling: More Messengers, More Channels, More Complexity. Curr. Biol. 12, R563–R565 (2002).
- Konieczny, V., Keebler, M. V. & Taylor, C. W. Spatial organization of intracellular Ca2+ signals. Semin. Cell Dev. Biol. 23, 172–180 (2012).
- Berridge, M. J., Lipp, P. & Bootman, M. D. The versatility and universality of calcium signalling. Nat. Rev. Mol. Cell Biol. 1, 11–21 (2000).
- Schuster, S., Marhl, M. & Höfer, T. Modelling of simple and complex calcium oscillations. Eur. J. Biochem. 269, 1333–1355 (2002).
- Bray, D. Protein molecules as computational elements in living cells. 376, (1995).
- Liu, X. et al. Optimization of Reactive Ink Formulation for Controlled Additive Manufacturing of Copolymer Membrane Functionalization. ACS Appl. Mater. Interfaces 16, 59216–59233 (2024).
- Chakrabarty, A., Buzzard, G. T. & Rundell, A. E. Model-based design of experiments for cellular processes. WIREs Syst. Biol. Med. 5, 181–203 (2013).
- Alazab, M. et al. Digital Twins for Healthcare 4.0—Recent Advances, Architecture, and Open Challenges. IEEE Consum. Electron. Mag. 12, 29–37 (2023).
- Kaul, R. et al. The role of AI for developing digital twins in healthcare: The case of cancer care. WIREs Data Min. Knowl. Discov. 13, e1480 (2023).
- Moingeon, P., Chenel, M., Rousseau, C., Voisin, E. & Guedj, M. Virtual patients, digital twins and causal disease models: Paving the ground for in silico clinical trials. Drug Discov. Today 28, 103605 (2023).
- Research, C. for D. E. and. Promoting Innovation in Medical Product Assessment: A Risk-based Framework for Evaluating Computational Models for Regulatory Decision-Making. FDA (2024).
- ALOISI, M. et al. DROSOPHILA: THE CENTURY-LONG FLIGHT FROM THE WILD TO THE PATIENT. Med. Sci. Pulse 19, 1–15 (2025).
- Green, S., Dam, M. S. & Svendsen, M. N. Mouse avatars of human cancers: the temporality of translation in precision oncology. Hist. Philos. Life Sci. 43, 27 (2021).
- Zarkesh, I. et al. Synthetic developmental biology: Engineering approaches to guide multicellular organization. Stem Cell Rep. 17, 715–733 (2022).
- Trentesaux, C., Yamada, T., Klein, O. D. & Lim, W. A. Harnessing synthetic biology to engineer organoids and tissues. Cell Stem Cell 30, 10–19 (2023).
- Shakiba, N., Jones, R. D., Weiss, R. & Del Vecchio, D. Context-aware synthetic biology by controller design: Engineering the mammalian cell. Cell Syst. 12, 561–592 (2021).
- Ethical, Legal and Social Implications Research Program. https://www.genome.gov/Funded-Programs-Projects/ELSI-Research-Program-ethical-legal-social-implications.
- Huang, P., Kim, K. & Schermer, M. Ethical Issues of Digital Twins for Personalized Health Care Service: Preliminary Mapping Study. J. Med. Internet Res. 24, e33081 (2022).
- Singh, K., Zeeshan, Z. & Karna, S. The Regulation of Biotechnology: Ethical, Legal, and Social Implications. Interdiscip. Stud. Soc. Law Polit. 2, 44–50 (2023).
- Endy, D. Foundations for engineering biology. Nature 438, 449–453 (2005).
- Sun, Y. & Zhong, Q. How modularity influences product innovation: The mediating role of module suppliers’ relationship-specific investments. Manag. Decis. 58, 2743–2761 (2020).
- König, H., Dorado-Morales, P. & Porcar, M. Responsibility and intellectual property in synthetic biology. EMBO Rep. 16, 1055–1059 (2015).
- Gardner, E. Multiscale imaging illuminates the big picture of plant protection. Purdue University - College of Agriculture https://ag.purdue.edu/news/2021/11/multiscale-imaging-illuminates-the-big-picture.html.
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).