1. Introduction
Malaria remains the most prevalent parasitic disease worldwide, with an estimated 263 million cases and 597,000 deaths reported in 2023 [
1]. Among its complications, cerebral malaria (CM) – a severe manifestation of
Plasmodium falciparum infection – continues to pose a significant global health challenge. CM is associated with high mortality rates (20–30%) and long-term neurocognitive impairments in up to 50% of survivors [
2]. Despite decades of research, the pathogenesis of CM is not yet fully understood. There are currently no validated biomarkers or effective adjunctive therapies, highlighting the urgent need for better predictive and mechanistic insights [
3,
4].
Animal models, particularly the experimental cerebral malaria (ECM) model using
Plasmodium berghei ANKA (PbA)-infected mice, have been instrumental in advancing our understanding of CM pathophysiology [
5]. However, this model presents several limitations, including variability in ECM incidence (50–100%) and heterogeneity in the timing of onset, which ranges from day 5 to day 12 post-infection [
4,
6,
7,
8,
9]. Furthermore, hallmark neurological symptoms – such as ataxia, convulsions, and coma – typically appear only hours before death, limiting the window for detecting early indicators of disease severity [
10]. Consequently, relying on post-infection day as a surrogate for disease stage may introduce significant bias, as animals at the same time point may exhibit markedly different levels of disease progression [
11].
Given that early prediction of ECM onset is critical for optimizing the timing of experimental interventions and therapeutic trials, efforts have been made to identify early markers of ECM development [
6,
11,
12,
13,
14,
15,
16,
17]. These studies have highlighted potential predictors such as neutrophil behavior, temperature, behavioral changes, and shifts in the T-cell receptor (TCR)-β repertoire. However, no existing study has developed a practical, predictive tool based solely on parasitemia data to anticipate ECM in individual animals before clinical deterioration.
In this study, we developed a supervised machine learning approach to predict ECM onset in PbA-infected mice using daily parasitemia measurements. We hypothesized that a model capable of integrating temporal changes in parasitemia could more accurately capture the complex dynamics underlying ECM development. Our approach offers a novel and accessible framework for reducing variability and bias in ECM studies, improving experimental design, and enhancing the identification of early pathophysiological changes. Moreover, a reliable method to forecast ECM development holds promise for increasing the translational value of this widely used animal model and deepening our understanding of CM pathogenesis.
2. Materials and Methods
Animals, Parasite, and Infection
This report was prepared according to the ARRIVE guidelines. Parasitemia data from 153 C57BL/6, 164 CBA, and 53 Swiss Webster mice were pooled from previous experiments done at the Laboratory of Malaria Research, IOC-Fiocruz, Rio de Janeiro, Brazil. Infection and parasitemia measurement protocols were described before in detail [
11,
16]. Briefly, five- to eight-week-old female mice were obtained from ICTB-Fiocruz (Rio de Janeiro, Brazil), housed in groups of five maximum, and had free access to food and water. Mice were infected intraperitoneally with 1x 106 PbA-infected erythrocytes coming from a donor mouse of the same strain, age, and sex. Thin blood smears were made daily with a blood drop collected from the tip of the tail, stained according to the Panoptic method (Laborclin, Brazil) and examined under a light microscope (BH2, Olympus: Melville, New York, USA) with an oil immersion lens (1000× final magnification). Parasitemia was determined by counting the number of parasitized RBCs in 2000 RBCs. Mice were observed three times a day and ECM diagnosis was made based on presentation of neurological clinical signs (ataxia, disorientation, paraplegia, roll-over and coma) on days 5 - 12 post-infection. Mice that did not develop ECM were euthanized 15 days post-infection.
Data and Variables
The outcome variable in the study was the development of ECM from day 5 to day 11 post-infection. The predictor variables consisted of parasitemia data from day 1 to day 4 post-infection and mouse strain (C57BL/6, CBA or Swiss Webster).
Statistical Analysis
The statistical plan was devised prior to accessing the data. All analysis was performed using GraphPad Prism Version 10.5.0 (GraphPad Software. Boston, MA) or Python Version 2.3.1. (Anaconda Software Distribution. Austin, TX). Log-rank test was used to compare survival curves. The following add-ons were used for analysis: pandas [
18], numpy [
19], itertools [
20], sklearn.model_selection, sklearn.ensemble, sklearn.linear, sklearn.metrics, sklean.preprocessing, sklearn.impute [
21], seaborn [
22], and matplotlib.pyplot. [
23].
As an initial step, any missing parasitemia values were imputed using k-nearest neighbor. Then, a logistic regression was performed as such:
Let Y ij denote the development of ECM (1 = ECM; 0 = no ECM) for the ith participant at the ith time point, where Yij = 1 indicates the presence of the event and Y ij = 0 indicates its absence. For each participant i, the response Y ij is observed over a series of day pairs (t1, t2).
logit(P(Yij = 1))= β0 + β1ΔPij + bi
Where:
(P(Yij = 1)) is the probability that the outcome occurs for participant i at day pair j.
ΔPij = Pij(t2) − Pij(t1) represents the parasitemia change between two time points.
β0 is the intercept term.
β1 is the coefficient capturing the effect of parasitemia change.
bi ∼ N(0,σ2) is a participant-specific random effect to account for within-subject correlation.
Optimal parasitemia threshold τ for parasitemia change is identified by maximizing the average precision (AP) score:
τ =argτ max AP (Y, Ŷ (ΔP ≥ τ))
Where Ŷ is the predicted outcome based on whether the parasitemia change exceeds the threshold:
Ŷij = 1, if ΔPij ≥ τ
Ŷij = 0, if ΔPij < τ
Resultant probability was calculated for each day pair (t1, t2) and threshold value τ, as follows:
P(Yij = 1) = σ(β0+β1 (ΔPij − τ))
The performance of the logistic regression model was assessed using sensitivity, specificity, AP, recall, area under the precision recall curve (PR-AUC), and f1-score for each day pair.
We also hypothesized that a limitation of the model could be differences between strains. To assess this the data was split by strain and analysis was repeated as described above. Model performance was again assessed for each day pair for each strain.
Random Forest Regressor Model
Although our initial analysis focused solely on predicting the occurrence of ECM as a binary outcome (1 or 0), we later hypothesized that the dynamics of parasitemia could be leveraged to predict the specific day on which ECM onset occurs. Using pandas, data was uploaded with the columns ‘ID’, ‘Strain’, day of ECM onset, and daily parasitemia data for day 1 to day 10. The data was reshaped to long format, the target variable was defined, feature engineering was used to create a column ‘Parasitemia_Change’ and rolling mean of the 3 days prior to ECM onset ‘Parasitemia_MA3’. The data was saved for modeling. Features were defined as ‘Parasitemia’, ‘Parasitemia_Change’, and ‘Parasitemia_MA3’ with the target variable defined as ‘CM_Risk.’ Data was split into training and testing sets with a split of 80% train, 20% testing and a random state of 42. Any missing data was imputed using sklearn SimpleImputer. The model was initialized using rf_model with 100 estimators, a random state of 42, and a balanced class weight. The model was fitted using rf_model.fit and used to make predictions using rf.model.predict. Performance was assessed using classification_report with precision, recall, f1-score, and support. To optimize model performance, sklearn GridSearchCV was used to identify optimal model parameters based upon best mean square error (MSE). Performance was then assessed using MSE, R2, mean absolute error (MAE), and cross-validated MSE and learning curve was plotted.
3. Results
3.1. ECM Incidence and Parasitemia Dynamics
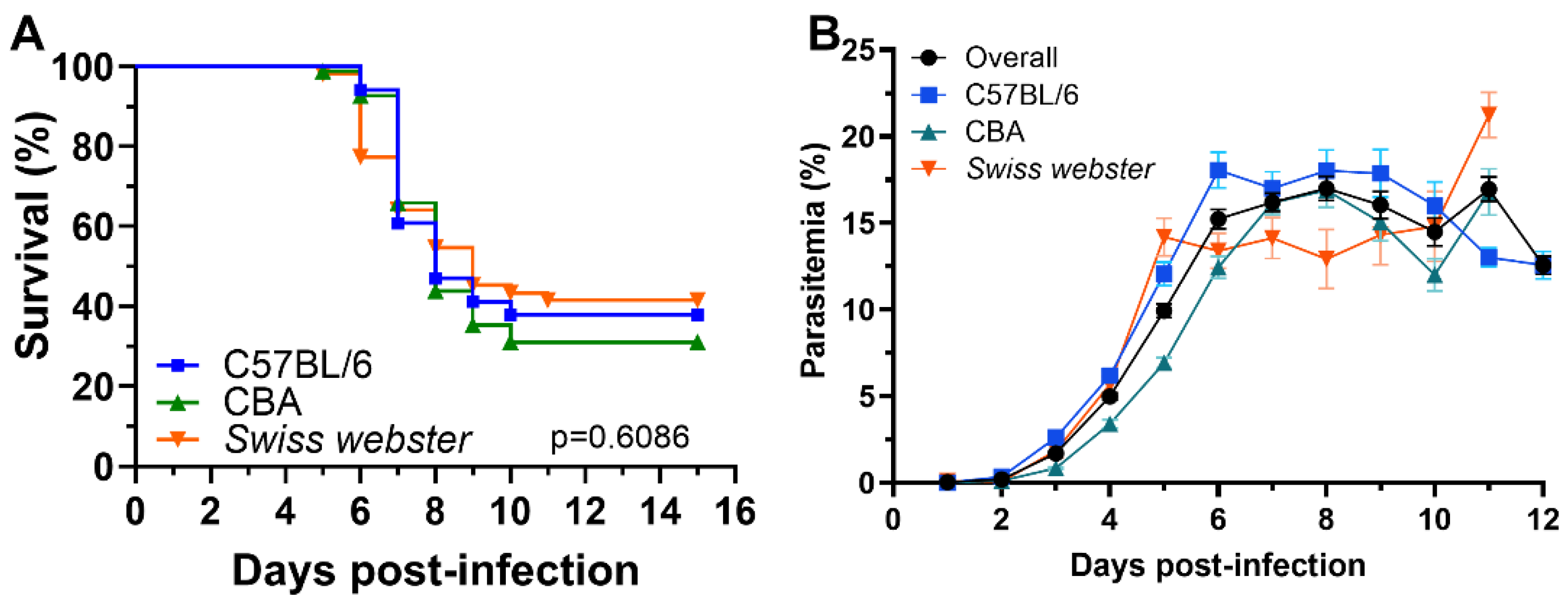
The overall incidence of experimental cerebral malaria (ECM) was 64.5% (64% in C57BL/6, 69% in CBA, and 51% in Swiss Webster mice). Survival curves did not significantly differ between strains. Mice developed ECM between days 5–11 post-infection, with strain-specific ranges: 6–10 days in C57BL/6, 5–10 in CBA, and 5–11 in Swiss Webster (
Figure 1A).
Parasitemia levels increased most rapidly between days 1 and 3, rising from a mean of 0.04% to 1.69%. This was followed by a more gradual increase from days 4 to 6, reaching 15.22% from an initial mean 4.98%. Between days 7 and 9, parasitemia plateaued, remaining relatively stable (mean parasitemia from 16.19% to 16.03%), before gradually declining from 14.49% to 12.57% between days 12 and 14 in the mice that survived (
Figure 1B).
3.2. Logistic Regression Model: Cohort-Level Analysis
We first trained a logistic regression model on the full dataset (all strains combined) to predict ECM risk from early parasitemia changes. This resulted in optimal changes in parasitemia of - 0.05% between day 1 to day 3 (AP: 0.67), 0.16% between day 1 to day 4 (AP: 0.67), 0.12% between day 2 to day 3 (AP: 0.66), 0.14% between day 2 to day 4 (AP: 0.67), and 2.34% between day 3 to day 4 (AP: 0.66). The resultant probability of CM was highest for a change of 0.05% or greater between day 1 and day 3 (93%). To assess model performance, a precision, recall/sensitivity, specificity, and f1-score were calculated for each day pair. Sensitivity exceeded 90% across several day pairs (days 1–3, 1–4, 2–3, and 2–4), while specificity was modest, peaking at 0.43 for days 3–4. Precision and f1-score were highest for day pair day 1 to day 3 (precision: 0.67, f1-score: 0.79), day 1 to day 4 (precision: 0.67, f1-score: 0.79), and day 2 to day 4 (precision: 0.67, f1-score: 0.79) (
Table 1). The model was converted into a usable tool within RedCap (publicly available at
https://redcapsurvey.slu.edu/surveys/?s=7NH7FR3FRN8D3ERW) which will identify the likelihood of a mouse developing ECM, based upon parasitemia data from day 1 to day 4.
3.3. Strain-Specific Logistic Regression
To assess if performance could be improved considering lineage, we took a two-fold approach. First, the model was trained with strain included, however, overall performance was minimally impacted by the inclusion of this variable. This resulted in optimal changes in parasitemia of 0.05% between day 1 to day 3 (AP: 0.67, f1-score: 0.77), 0.16% between day 1 to day 4 (AP: 0.67, f1-score: 0.79), 0.12% between day 2 to day 3 (AP: 0.66, f1-score: 0.76), 0.14% between day 2 to day 4 (AP: 0.67, f1-score: 0.79), and 2.34% between day 3 to day 4 (AP: 0.66, f1-score: 0.74). Alternatively, we assessed if performance could be improved when the mice were separated by strain. Overall performance was again minimally impacted. The highest AP for any day pair was 0.68 in the C57BL/6 mice cohort, 0.72 in the CBA cohort, and 0.60 within the Swiss Webster cohort. The highest f1-score was 0.79 in the C57BL/6 mice cohort, 0.83 in the CBA cohort, and 0.67 in the Swiss Webster cohort. The cutoffs values for all the respective day pair-strain combinations and their respective performance parameters can be seen in
Table 1.
3.4. Predicting Day of ECM Onset with Random Forest Regression
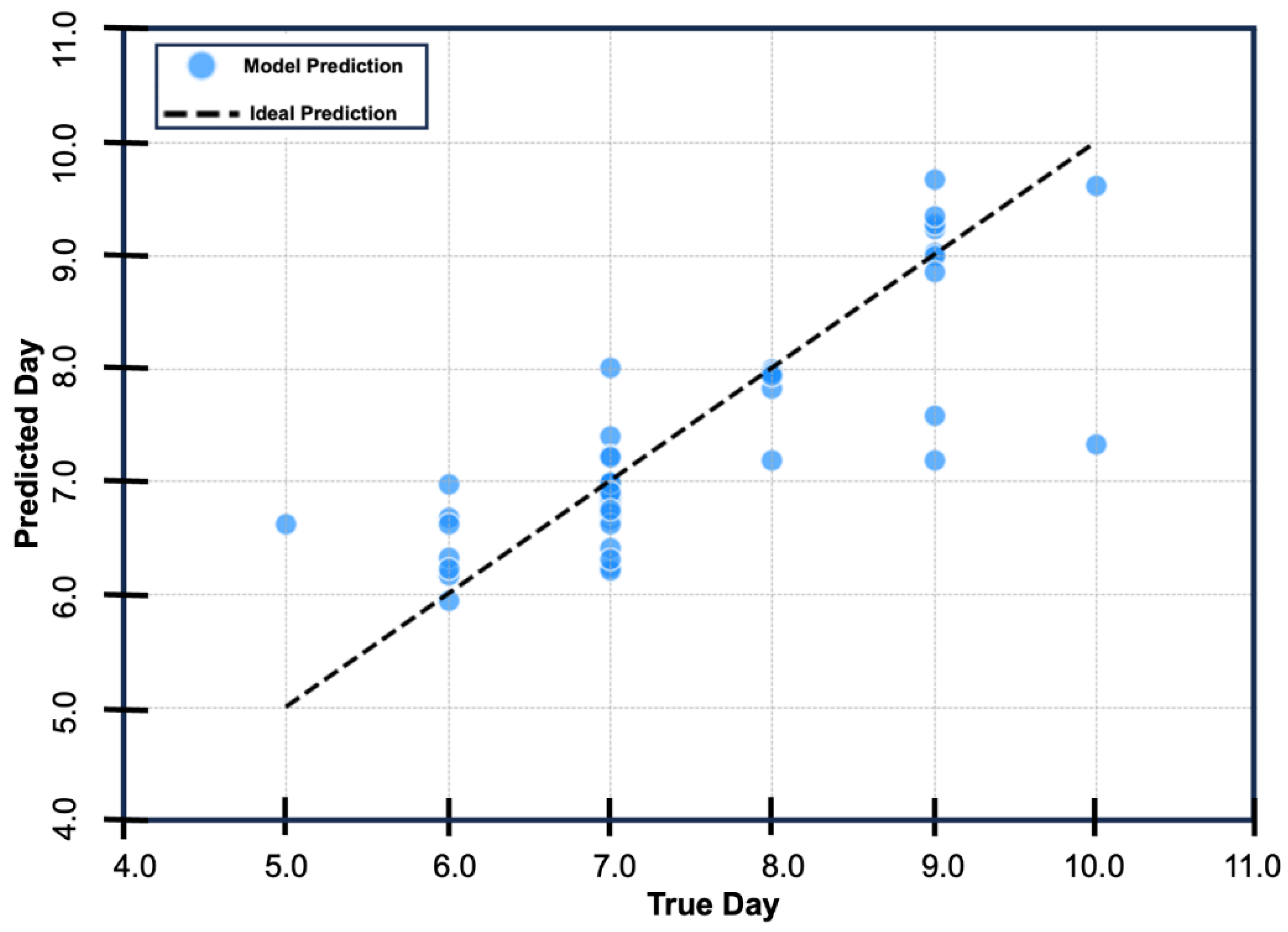
As a secondary step, we sought to train and fit a Random Forest Regressor model to predict the day of ECM onset using available parasitemia data from the preceding days. The model takes all available parasitemia data (from day 1 through day 11), imputes any future values or missing values, and provide a precise estimate of the day of ECM onset. Importantly, the model predicted day of onset of ECM within 0.54 ± 0.38 days across the test dataset, with a cross-validated MSE = 0.2924 ± 0.1461, R2 = 0.6378, and MAE = 0.4301 (
Figure 2). To facilitate the broader application of our model in ECM research, we developed an extended version that enables users to input parasitemia measurements and obtain a predicted day of ECM onset. The model accepts parasitemia data from days 1 through 11 of infection. To accommodate incomplete datasets, we incorporated an imputation algorithm that permits accurate prediction based on as few as two data points (e.g., parasitemia levels from days 1 and 3). The random forest model, the imputer, and the predictor extension are publicly available at:
https://github.com/pmurin29/cm-risk-prediction.
4. Discussion
An ideal model for the identification of ECM would: (1) identify high risk mice before the onset of neurological signs (prior to day 5), (2) predict approximate day of onset facilitating timing of interventions and accurate comparison of disease stages, and (3) be easily applied without need for time-consuming tests or complex assays. In the present study, we applied supervised machine learning, in the form of a multivariable logistic regression, to predict the development of ECM in mice based on parasitemia dynamics over day 1 to day 4. This resulted in parasitemia change threshold values predicting the onset of ECM with near perfect sensitivity at multiple day pairs, achieving aim 1 and aim 3. However, logistic regression models are limited to binary outcome predictions, making it ineffective to predict day of onset. To overcome this limitation, we applied supervised machine learning in the form of a Random Forest Regressor to determine if we could predict the exact day of ECM, achieving robust performance, with the model predicting day of ECM within 0.54 ± 0.38 days across the dataset (MSE: 0.2924 ± 0.1461), thereby achieving aims 1 – 3. Therefore, we demonstrated that parasitemia levels can be an efficacious and robust predictor for the development of ECM in PbA-infected mice.
There have been several previous attempts to address the challenge of predicting ECM onset in mice. A symptom based approach using the SHIRPA protocol was able to predict the development of ECM with high positive predictive value (>89%) on days 5 or 6 post-infection [
24]. However, such a model requires a time consuming and skilled assessment and is limited in its ability to detect ECM earlier than 24 hours before onset of clinical signs [
25]. Several biomarker-based approaches have been utilized. In a study of neutrophil dynamics in PbA-infected mice, a higher percentage of neutrophils was noted in the spleen and peripheral blood in ECM susceptible C57BL/6 mice compared to ECM resistant BALB/c mice at day 4 post-infection [
13]. However, while these differences suggest neutrophils may offer potential as a biomarker for ECM, it is unclear if these preliminary findings could be utilized to predict ECM development and/or time of onset. In a study of T-cell receptor signature alterations is mice infected with PbA, alterations in the T-cell receptor repertoires were noted in the spleen and peripheral blood on day 5 and day 6 of mice with ECM compared to mice without ECM [
15]. This is supported by the well-established role of T-cells in ECM and malaria pathogenesis [
26,
27,
28]. However, such an approach is again limited in its ability to predict ECM early in the course of the disease, prior to ECM onset [
25].
Our study was not without limitations. Not all parasitemia data was available for all days, requiring the use of imputation. While the logistic regression model was highly sensitive for the identification of ECM, specificity was limited. Not all mice were of the same strain. While strain was considered in the development of the logistic regression model, inclusion of strain as a covariate did not appear to improve performance. Given the small sample sizes within individual strains and the associated risk of overfitting, we did not attempt to train the Random Forest model by strain. Future work will focus on further training and refining the Random Forest model to enhance predictive accuracy.
Author Contributions
Conceptualization, Y.C.M.; methodology, P.J.M., C.T.D.R. and L.J.M.C.; formal analysis, P.J.M, Y.C.M.; investigation, C.T.D.R., L.J.M.C. and Y.C.M.; resources, C.T.D.R. and L.J.M.C.; data curation, Y.C.M.; writing—original draft preparation, P.J.M. and Y.C.M.; writing—review and editing, C.T.D.R., L.J.M.C. and Y.C.M.; visualization, P.J.M. and Y.C.M.; supervision, C.T.D.R., L.J.M.C. and Y.C.M.; project administration, Y.C.M.; funding acquisition, L.J.M.C., C.T.D.R. All authors have read and agreed to the published version of the manuscript.
Funding
P.J.M. receives grant support from the American Academy of Neurology. C.T.D.R and L.J.M.C are supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, Brazil 310445/2017-5, and 316462/2021-7) through a Productivity Research Fellowship, and receive a Cientista do Nosso Estado fellowship by the Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro (Faperj, E-26/202.921/2018 and E-26/200.314), respectively. L.J.M.C. is also supported by a “Universal” grant from CNPq (422430/2016-1). The Laboratório de Pesquisa em Malária (LPM-IOC, Fiocruz) is an Associate Laboratory of the Instituto Nacional de Ciência e Tecnologia em Neuroimunomodulação of the CNPq (INCT-NIM/CNPq Project 465489/2014-1) and of the Rede de Neuroinflamação da Faperj (Redes/Faperj, Project 26010.002418/2019) and receives financial support of Faperj (Project SEI-260003/001169/2020). The LPM-IOC, Fiocruz receive support through the POM.
Institutional Review Board Statement
The animal study protocol was approved by the Fiocruz Ethics Committee on Animal Use (protocol codes P-0155-03 and L-079/08).
Data Availability Statement
The original data presented in the study are openly available in Zenodo at doi: 10.5281/zenodo.15557233.
Conflicts of Interest
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Abbreviations
The following abbreviations are used in this manuscript:
| AP |
Average Precision |
| ARRIVE |
Animal Research: Reporting of In Vivo Experiments (guidelines) |
| CNPq |
Conselho Nacional de Desenvolvimento Científico e Tecnológico (Brazil) |
| CM |
Cerebral Malaria |
| ECM |
Experimental Cerebral Malaria |
| Faperj |
Fundação Carlos Chagas Filho de Amparo à Pesquisa do Estado do Rio de Janeiro |
| Fiocruz |
Fundação Oswaldo Cruz |
| ICTB |
Instituto de Ciência e Tecnologia em Biomodelos |
| INCT-NIM |
Instituto Nacional de Ciência e Tecnologia em Neuroimunomodulação |
| IOC |
Instituto Oswaldo Cruz |
| LPM |
Laboratório de Pesquisa em Malária |
| MAE |
Mean Absolute Error |
| MSE |
Mean Squared Error |
| PbA |
Plasmodium berghei ANKA |
| PR-AUC |
Precision-Recall Area Under the Curve |
| R² |
Coefficient of Determination (R-squared) |
| RBC |
Red Blood Cell |
| RedCap |
Research Electronic Data Capture |
| SHIRPA |
SmithKline Beecham, Harwell, Imperial College, Royal London Hospital, Phenotype Assessment (protocol) |
| TCR |
T-cell Receptor |
References
- WHO. World malaria report 2024: addressing inequity in the global malaria response; World Health Organization: Geneva, 2024; p. 320. [Google Scholar]
- Muppidi, P.; Wright, E.; Wassmer, S.C.; Gupta, H. Diagnosis of cerebral malaria: Tools to reduce Plasmodium falciparum associated mortality. Front Cell Infect Microbiol 2023, 13, 1090013. [Google Scholar] [CrossRef]
- Bensalel, J.; Gallego-Delgado, J. Exploring adjunctive therapies for cerebral malaria. Front Cell Infect Microbiol 2024, 14, 1347486. [Google Scholar] [CrossRef] [PubMed]
- Cha, S.J.; Yu, X.; Gregory, B.D.; Lee, Y.S.; Ishino, T.; Opoka, R.O.; John, C.C.; Jacobs-Lorena, M. Identification of Key Determinants of Cerebral Malaria Development and Inhibition Pathways. mBio 2022, 13, e0370821. [Google Scholar] [CrossRef] [PubMed]
- Wassmer, S.C.; de Koning-Ward, T.F.; Grau, G.E.R.; Pai, S. Unravelling mysteries at the perivascular space: a new rationale for cerebral malaria pathogenesis. Trends Parasitol 2024, 40, 28–44. [Google Scholar] [CrossRef] [PubMed]
- Baptista, F.G.; Pamplona, A.; Pena, A.C.; Mota, M.M.; Pied, S.; Vigario, A.M. Accumulation of Plasmodium berghei-infected red blood cells in the brain is crucial for the development of cerebral malaria in mice. Infect Immun 2010, 78, 4033–4039. [Google Scholar] [CrossRef]
- Basir, R.; Rahiman, S.F.; Hasballah, K.; Chong, W.; Talib, H.; Yam, M.; Jabbarzare, M.; Tie, T.; Othman, F.; Moklas, M.; et al. Plasmodium berghei ANKA Infection in ICR Mice as a Model of Cerebral Malaria. Iran J Parasitol 2012, 7, 62–74. [Google Scholar]
- Hortle, E.; Starrs, L.; Brown, F.C.; Jane, S.M.; Curtis, D.J.; McMorran, B.J.; Foote, S.J.; Burgio, G. KCC1 Activation protects Mice from the Development of Experimental Cerebral Malaria. Sci Rep 2019, 9, 6356. [Google Scholar] [CrossRef]
- Stevenson, M.M.; Gros, P.; Olivier, M.; Fortin, A.; Serghides, L. Cerebral malaria: human versus mouse studies. Trends Parasitol 2010, 26, 274–275. [Google Scholar] [CrossRef]
- Hearn, J.; Rayment, N.; Landon, D.N.; Katz, D.R.; de Souza, J.B. Immunopathology of cerebral malaria: morphological evidence of parasite sequestration in murine brain microvasculature. Infect Immun 2000, 68, 5364–5376. [Google Scholar] [CrossRef]
- Martins, Y.C.; Werneck, G.L.; Carvalho, L.J.; Silva, B.P.; Andrade, B.G.; Souza, T.M.; Souza, D.O.; Daniel-Ribeiro, C.T. Algorithms to predict cerebral malaria in murine models using the SHIRPA protocol. Malar J 2010, 9, 85. [Google Scholar] [CrossRef]
- Desruisseaux, M.S.; Gulinello, M.; Smith, D.N.; Lee, S.C.; Tsuji, M.; Weiss, L.M.; Spray, D.C.; Tanowitz, H.B. Cognitive dysfunction in mice infected with Plasmodium berghei strain ANKA. J Infect Dis 2008, 197, 1621–1627. [Google Scholar] [CrossRef] [PubMed]
- Freire-Antunes, L.; Ornellas-Garcia, U.; Rangel-Ferreira, M.V.; Ribeiro-Almeida, M.L.; de Sousa, C.H.G.; Carvalho, L.J.M.; Daniel-Ribeiro, C.T.; Ribeiro-Gomes, F.L. Increased Neutrophil Percentage and Neutrophil-T Cell Ratio Precedes Clinical Onset of Experimental Cerebral Malaria. Int J Mol Sci 2023, 24. [Google Scholar] [CrossRef] [PubMed]
- Ketema, T.; Yohannes, M.; Alemayehu, E.; Ambelu, A. Effect of chronic khat (Catha edulis, Forsk) use on outcome of Plasmodium berghei ANKA infection in Swiss albino mice. BMC Infect Dis 2015, 15, 170. [Google Scholar] [CrossRef] [PubMed]
- Mariotti-Ferrandiz, E.; Pham, H.P.; Dulauroy, S.; Gorgette, O.; Klatzmann, D.; Cazenave, P.A.; Pied, S.; Six, A. A TCRbeta Repertoire Signature Can Predict Experimental Cerebral Malaria. PLoS One 2016, 11, e0147871. [Google Scholar] [CrossRef]
- Martins, Y.C.; Smith, M.J.; Pelajo-Machado, M.; Werneck, G.L.; Lenzi, H.L.; Daniel-Ribeiro, C.T.; Carvalho, L.J. Characterization of cerebral malaria in the outbred Swiss Webster mouse infected by Plasmodium berghei ANKA. Int J Exp Pathol 2009, 90, 119–130. [Google Scholar] [CrossRef]
- Santos, L.C.; Abreu, C.F.; Xerinda, S.M.; Tavares, M.; Lucas, R.; Sarmento, A.C. Severe imported malaria in an intensive care unit: a review of 59 cases. Malar J 2012, 11, 96. [Google Scholar] [CrossRef]
- McKinney, W. Data structures for statistical computing in Python. SciPy 2010, 445, 51–56. [Google Scholar]
- Harris, C.R.; Millman, K.J.; van der Walt, S.J.; Gommers, R.; Virtanen, P.; Cournapeau, D.; Wieser, E.; Taylor, J.; Berg, S.; Smith, N.J.; et al. Array programming with NumPy. Nature 2020, 585, 357–362. [Google Scholar] [CrossRef]
- Van Rossum, G. The python library reference, release 3.8. 2. Python Software Foundation 2020, 36. [Google Scholar]
- Fabian Pedregosa, G.V. , Alexandre Gramfort, Vincent Michel, Bertrand Thirion, Olivier Grisel, Mathieu Blondel, Peter Prettenhofer, Ron Weiss, Vincent Dubourg, Jake Vanderplas, Alexandre Passos, David Cournapeau, Matthieu Brucher, Matthieu Perrot, Édouard Duchesnay. Scikit-learn: Machine Learning in Python. Journal of Machine Learning Research 2011, 12, 2825–2830. [Google Scholar]
- Waskom, M.L. seaborn: statistical data visualization. Journal of Open Source Software 2021, 6. [Google Scholar] [CrossRef]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Computing in Science and Engineering 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Martins, Y.C.; Carvalho, L.J.; Daniel-Ribeiro, C.T. Challenges in the determination of early predictors of cerebral malaria: lessons from the human disease and the experimental murine models. Neuroimmunomodulation 2009, 16, 134–145. [Google Scholar] [CrossRef] [PubMed]
- Royo, J.; Camara, A.; Bertrand, B.; Batigne, P.; Coste, A.; Pipy, B.; Aubouy, A.; Neuro, C.M.G. Kinetics of monocyte subpopulations during experimental cerebral malaria and its resolution in a model of late chloroquine treatment. Front Cell Infect Microbiol 2022, 12, 952993. [Google Scholar] [CrossRef]
- Collette, A.; Bagot, S.; Ferrandiz, M.E.; Cazenave, P.A.; Six, A.; Pied, S. A profound alteration of blood TCRB repertoire allows prediction of cerebral malaria. J Immunol 2004, 173, 4568–4575. [Google Scholar] [CrossRef]
- Haque, A.; Best, S.E.; Unosson, K.; Amante, F.H.; de Labastida, F.; Anstey, N.M.; Karupiah, G.; Smyth, M.J.; Heath, W.R.; Engwerda, C.R. Granzyme B expression by CD8+ T cells is required for the development of experimental cerebral malaria. J Immunol 2011, 186, 6148–6156. [Google Scholar] [CrossRef]
- Fain, C.E.; Zheng, J.; Jin, F.; Ayasoufi, K.; Wu, Y.; Lilley, M.T.; Dropik, A.R.; Wolf, D.M.; Rodriguez, R.C.; Aibaidula, A.; et al. Discrete class I molecules on brain endothelium differentially regulate neuropathology in experimental cerebral malaria. Brain 2024, 147, 566–589. [Google Scholar] [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).