Submitted:
20 June 2025
Posted:
24 June 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
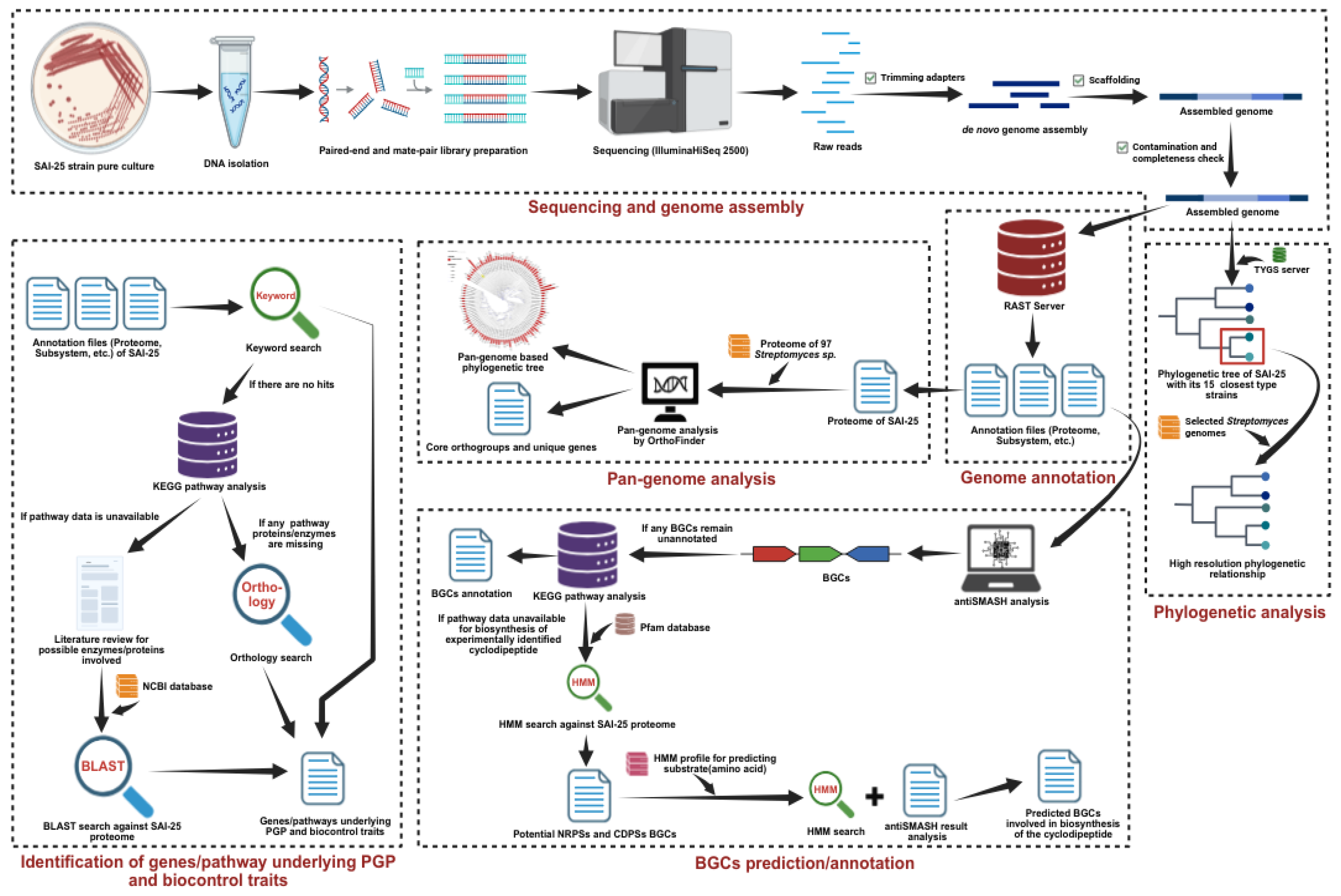
2. Materials and Methods
2.1. Microbial Strain Used in This Study
2.2. Culture of PGP Strain and Genomic DNA Isolation
2.3. Library Preparation and Sequencing
2.4. Genome Assembly and Annotation
2.5. Phylogenetic Relationship of SAI-25
2.6. Pan-Genome Analysis
2.7. Identification of Biosynthetic Gene Clusters (BGCs) in SAI-25 Strain
2.8. Identification of Potential Genes/Enzymes Responsible for Biosynthesis of an Insecticidal Diketopiperazine Derivative, Cyclo(Trp-Phe)
2.9. Genes/Pathways Underlying PGP Features
3. Results
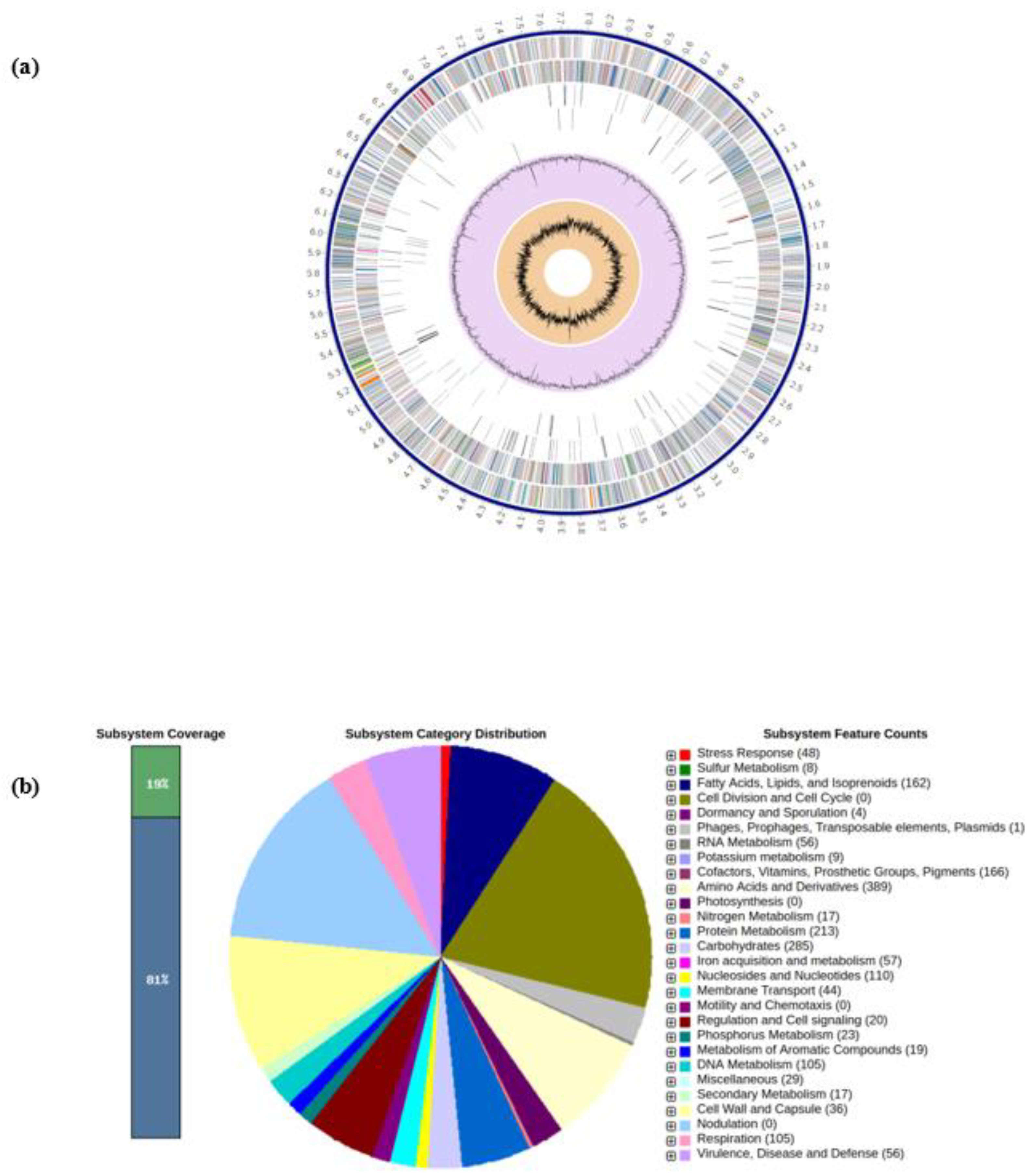
3.1. Features of Genome Assembly of Streptomyces sp. SAI-25
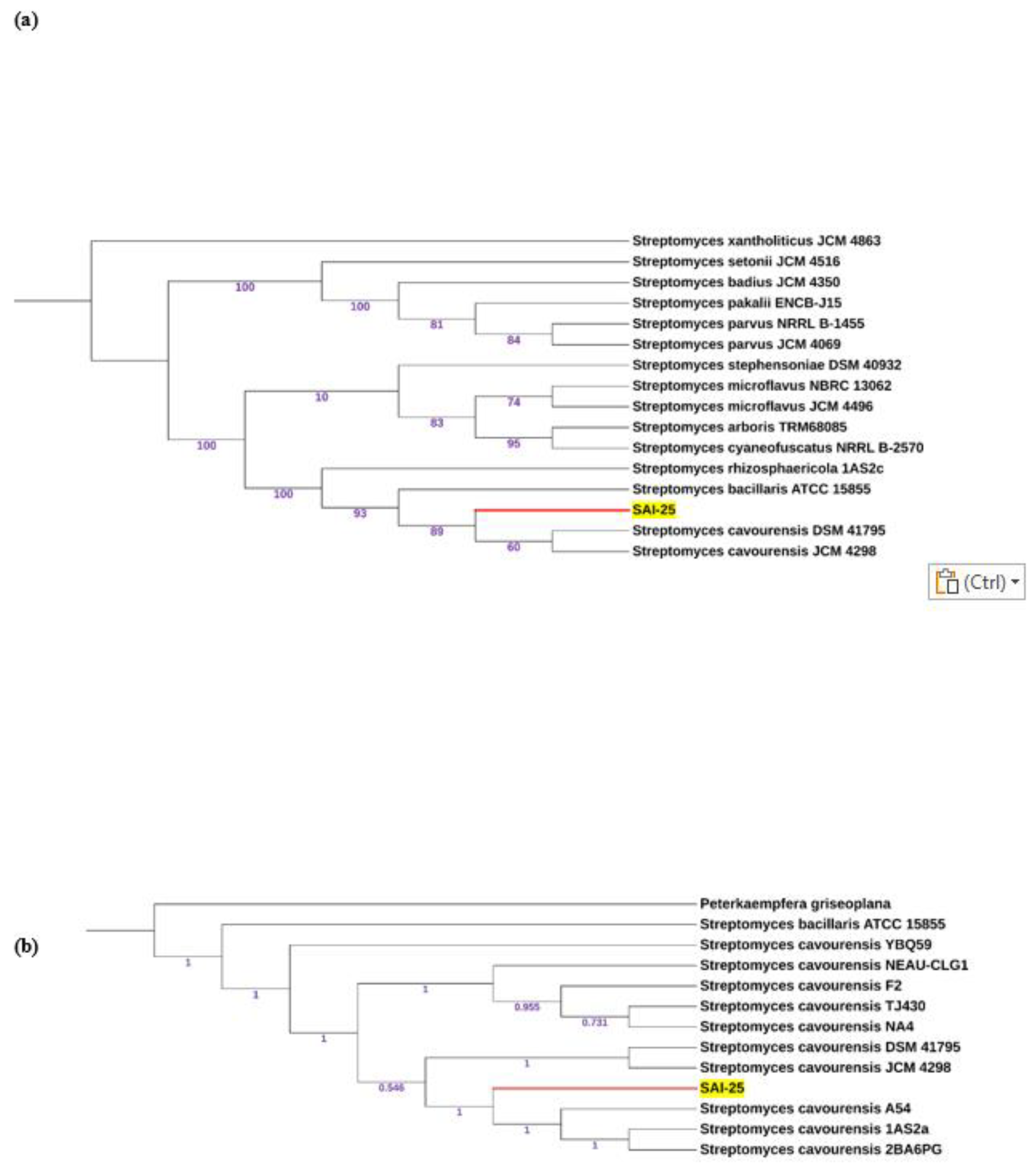
3.2. Phylogenetic Relationship and Taxonomic Positioning of SAI-25:
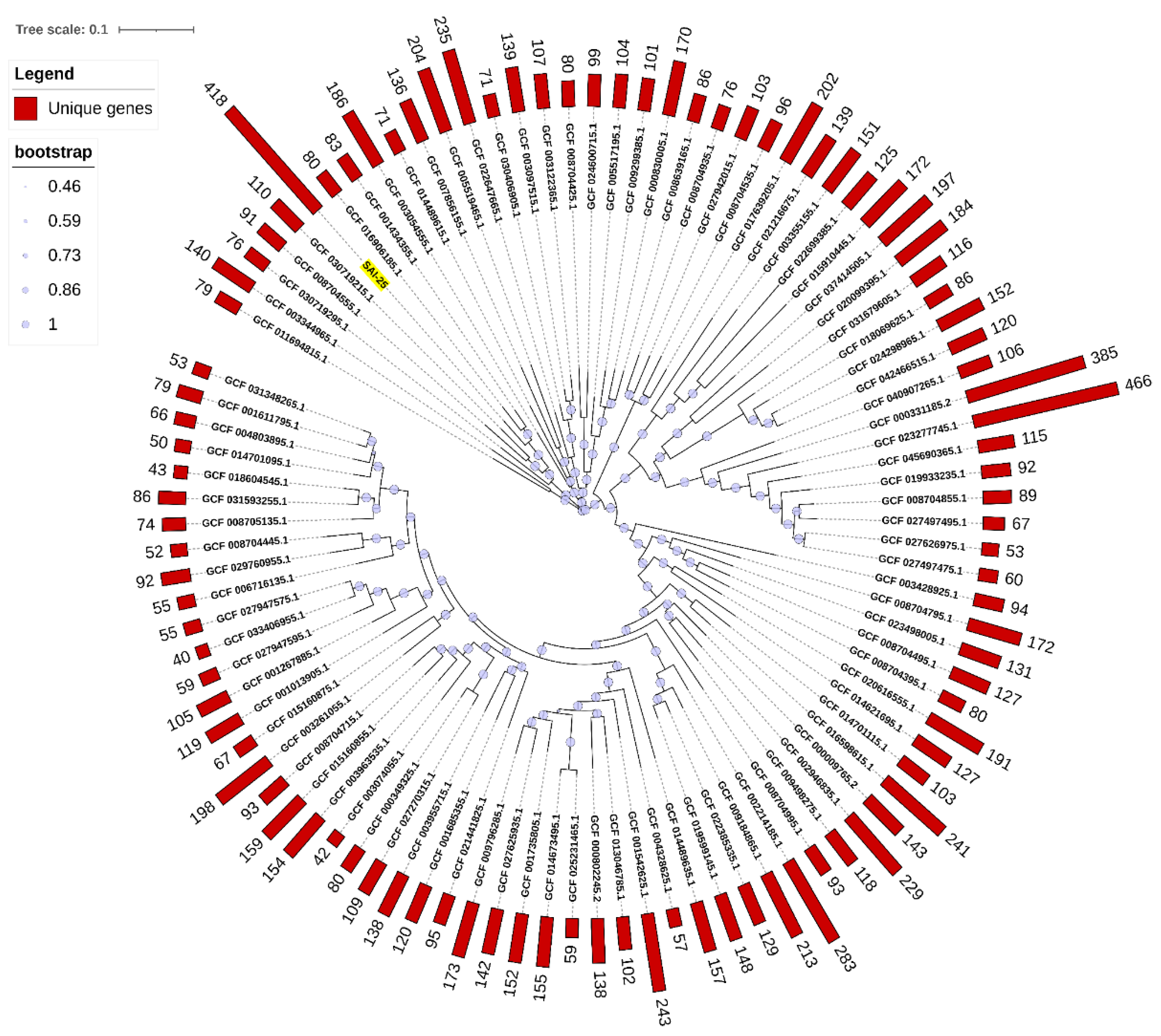
3.3. Core Ortho-Groups and Unique Genes of SAI-25
3.4. Secondary Metabolite Potential of SAI-25
3.5. Potential Genes/Enzymes Responsible for Biosynthesis of an Insecticidal Diketopiperazine Derivative, Cyclo(Trp-Phe)
3.6. Genes/Pathway underlying PGP features
4. Discussion
4.1. SAI-25’s Chromosome-Level Assembly with High Completeness Will Be A Valuable Resource for Streptomyces Genome Mining
4.2. Phylogenetic Analysis Corrected the Species Name From S. Griseoplanus to S. Cavourensis:
4.3. Presence of Sixteen Annotated And The Same Unannotated BGCS Highlights its PGP and Industrial Potential
4.4. Limited Success in Prediction of Genes/Bgcs for Cyclo(Trp-Phe) Biosynthesis Opens the Scope for Further Characterizatio
4.5. Limitations and Future Directions:
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Declarations
Clinical trial number
Permission to publish
References
- Aggarwal N, Thind SK, Sharma S (2016) Role of Secondary Metabolites of Actinomycetes in Crop Protection. In: Subramaniam G, Arumugam S, Rajendran V (eds) Plant Growth Promoting Actinobacteria. Springer Singapore, Singapore, pp 99–121.
- An J, Kim SH, Bahk S, Vuong UT, Nguyen NT, Do HL, Kim SH, Chung WS (2021) Naringenin Induces Pathogen Resistance Against Pseudomonas syringae Through the Activation of NPR1 in Arabidopsis. Front Plant Sci 12:672552. [CrossRef]
- Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. Journal of Computational Biology 19(5):455–477. [CrossRef]
- Belknap KC, Park CJ, Barth BM, Andam CP (2020) Genome mining of biosynthetic and chemotherapeutic gene clusters in Streptomyces bacteria. Sci Rep 10(1):2003. [CrossRef]
- Bellotti D, Remelli M (2021) Deferoxamine B: A Natural, Excellent and Versatile Metal Chelator. Molecules 26(11):3255. [CrossRef]
- Bhattacharyya C, Bakshi U, Mallick I, Mukherji S, Bera B, Ghosh A (2017) Genome-Guided Insights into the Plant Growth Promotion Capabilities of the Physiologically Versatile Bacillus aryabhattai Strain AB211. Front Microbiol 8. [CrossRef]
- Blin K, Shaw S, Augustijn HE, Reitz ZL, Biermann F, Alanjary M, Fetter A, Terlouw BR, Metcalf WW, Helfrich EJN, van Wezel GP, Medema MH, Weber T (2023) antiSMASH 7.0: new and improved predictions for detection, regulation, chemical structures and visualisation. Nucleic Acids Research 51(W1):W46–W50. [CrossRef]
- Blum M, Andreeva A, Florentino LC, Chuguransky SR, Grego T, Hobbs E, Pinto BL, Orr A, Paysan-Lafosse T, Ponamareva I, Salazar GA, Bordin N, Bork P, Bridge A, Colwell L, Gough J, Haft DH, Letunic I, Llinares-López F, Marchler-Bauer A, Meng-Papaxanthos L, Mi H, Natale DA, Orengo CA, Pandurangan AP, Piovesan D, Rivoire C, Sigrist CJA, Thanki N, Thibaud-Nissen F, Thomas PD, Tosatto SCE, Wu CH, Bateman A (2025) InterPro: the protein sequence classification resource in 2025. Nucleic Acids Research 53(D1):D444–D456. [CrossRef]
- Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120. [CrossRef]
- Bosi E, Donati B, Galardini M, Brunetti S, Sagot M-F, Lió P, Crescenzi P, Fani R, Fondi M (2015) M e D u S a : a multi-draft based scaffolder. Bioinformatics 31(15):2443–2451. [CrossRef]
- Bowman EJ, Siebers A, Altendorf K (1988) Bafilomycins: a class of inhibitors of membrane ATPases from microorganisms, animal cells, and plant cells. Proc Natl Acad Sci USA 85(21):7972–7976. [CrossRef]
- Brettin T, Davis JJ, Disz T, Edwards RA, Gerdes S, Olsen GJ, Olson R, Overbeek R, Parrello B, Pusch GD, Shukla M, Thomason JA, Stevens R, Vonstein V, Wattam AR, Xia F (2015) RASTtk: A modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep 5(1):8365. [CrossRef]
- Bruce TJ, Birkett MA, Blande J, Hooper AM, Martin JL, Khambay B, Prosser I, Smart LE, Wadhams LJ (2005) Response of economically important aphids to components of Hemizygia petiolata essential oil. Pest Management Science 61(11):1115–1121. [CrossRef]
- Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10(1):421. [CrossRef]
- Chen Y, Guo M, Yang J, Chen J, Xie B, Sun Z (2019) Potential TSPO Ligand and Photooxidation Quencher Isorenieratene from Arctic Ocean Rhodococcus sp. B7740. Marine Drugs 17(6):316. [CrossRef]
- Cho G, Kim J, Park CG, Nislow C, Weller DM, Kwak Y-S (2017) Caryolan-1-ol, an antifungal volatile produced by Streptomyces spp., inhibits the endomembrane system of fungi. Open Biol 7(7):170075. [CrossRef]
- Creencia AR, Alcantara EP, Diaz MGQ, Monsalud RG (2021) Draft genome sequence of a Philippine mangrove soil actinomycete with insecticidal activity reveals potential as a source of other valuable secondary metabolites. 14.
- Emms DM, Kelly S (2019) OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol 20(1):238. [CrossRef]
- Gandham P, Vadla N, Saji A, Srinivas V, Ruperao P, Selvanayagam S, Saxena RK, Rathore A, Gopalakrishnan S, Thakur V (2024) Genome assembly, comparative genomics, and identification of genes/pathways underlying plant growth-promoting traits of an actinobacterial strain, Amycolatopsis sp. (BCA-696). Sci Rep 14(1):15934. [CrossRef]
- Garbeva P, Avalos M, Ulanova D, Van Wezel GP, Dickschat JS (2023) Volatile sensation: The chemical ecology of the earthy odorant geosmin. Environmental Microbiology 25(9):1565–1574. [CrossRef]
- Gopalakrishnan S, Sharma R, Srinivas V, Naresh N, Mishra SP, Ankati S, Pratyusha S, Govindaraj M, Gonzalez SV, Nervik S, Simic N (2020) Identification and Characterization of a Streptomyces albus Strain and Its Secondary Metabolite Organophosphate against Charcoal Rot of Sorghum. Plants 9(12):1727. [CrossRef]
- Gopalakrishnan S, Srinivas V, Alekhya G, Prakash B (2015a) Effect of plant growth-promoting Streptomyces sp. on growth promotion and grain yield in chickpea (Cicer arietinum L). 3 Biotech 5(5):799–806. [CrossRef]
- Gopalakrishnan S, Srinivas V, Alekhya G, Prakash B, Kudapa H, Rathore A, Varshney RK (2015b) The extent of grain yield and plant growth enhancement by plant growth-promoting broad-spectrum Streptomyces sp. in chickpea. SpringerPlus 4(1):31. [CrossRef]
- Gopalakrishnan S, Srinivas V, Chand U, Pratyusha S, Samineni S (2022) Streptomyces consortia-mediated plant growth-promotion and yield performance in chickpea. 3 Biotech 12(11):318. [CrossRef]
- Gopalakrishnan S, Srinivas V, Naresh N, Pratyusha S, Ankati S, Madhuprakash J, Govindaraj M, Sharma R (2021) Deciphering the antagonistic effect of Streptomyces spp. and host-plant resistance induction against charcoal rot of sorghum. Planta 253(2):57. [CrossRef]
- Gopalakrishnan S, Upadhyaya H, Vadlamudi S, Humayun P, Vidya MS, Alekhya G, Singh A, Vijayabharathi R, Bhimineni RK, Seema M, Rathore A, Rupela O (2012) Plant growth-promoting traits of biocontrol potential bacteria isolated from rice rhizosphere. SpringerPlus 1(1):71. [CrossRef]
- Gopalakrishnan S, Vadlamudi S, Apparla S, Bandikinda P, Vijayabharathi R, Bhimineni RK, Rupela O (2013) Evaluation of Streptomyces spp. for their plant-growth-promotion traits in rice. Can J Microbiol 59(8):534–539. [CrossRef]
- Gopalakrishnan S, Vadlamudi S, Bandikinda P, Sathya A, Vijayabharathi R, Rupela O, Kudapa H, Katta K, Varshney RK (2014) Evaluation of Streptomyces strains isolated from herbal vermicompost for their plant growth-promotion traits in rice. Microbiological Research 169(1):40–48. [CrossRef]
- Gurevich A, Saveliev V, Vyahhi N, Tesler G (2013) QUAST: quality assessment tool for genome assemblies. Bioinformatics 29(8):1072–1075. [CrossRef]
- Hou L, Liu Z, Yu D, Li H, Ju J, Li W (2020) Targeted isolation of new polycyclic tetramate macrolactams from the deepsea-derived Streptomyces somaliensis SCSIO ZH66. Bioorganic Chemistry 101:103954. [CrossRef]
- Huang S, Liu Y, Liu W-Q, Neubauer P, Li J (2021) The Nonribosomal Peptide Valinomycin: From Discovery to Bioactivity and Biosynthesis. Microorganisms 9(4):780. [CrossRef]
- Jayasekara S, Ratnayake R (2019) Microbial Cellulases: An Overview and Applications. In: Rodríguez Pascual A, E. Eugenio Martín M (eds) Cellulose. IntechOpen.
- Kaaniche F, Hamed A, Elleuch L, Chakchouk-Mtibaa A, Smaoui S, Karray-Rebai I, Koubaa I, Arcile G, Allouche N, Mellouli L (2020) Purification and characterization of seven bioactive compounds from the newly isolated Streptomyces cavourensis TN638 strain via solid-state fermentation. Microbial Pathogenesis 142:104106. [CrossRef]
- Kanehisa M, Sato Y (2020) KEGG Mapper for inferring cellular functions from protein sequences. Protein Science 29(1):28–35. [CrossRef]
- Kanehisa M, Sato Y, Kawashima M (2022) KEGG mapping tools for uncovering hidden features in biological data. Protein Science 31(1):47–53. [CrossRef]
- Kanehisa M, Sato Y, Morishima K (2016) BlastKOALA and GhostKOALA: KEGG Tools for Functional Characterization of Genome and Metagenome Sequences. Journal of Molecular Biology 428(4):726–731. [CrossRef]
- Klosterman HJ, Lamoureux GL, Parsons JL (1967) Isolation, Characterization, and Synthesis of Linatine. A Vitamin B6 Antagonist from Flaxseed (Linum usitatissimum)*. Biochemistry 6(1):170–177. [CrossRef]
- Larsen MV, Cosentino S, Lukjancenko O, Saputra D, Rasmussen S, Hasman H, Sicheritz-Pontén T, Aarestrup FM, Ussery DW, Lund O (2014) Benchmarking of Methods for Genomic Taxonomy. J Clin Microbiol 52(5):1529–1539. [CrossRef]
- Lautru S, Deeth RJ, Bailey LM, Challis GL (2005) Discovery of a new peptide natural product by Streptomyces coelicolor genome mining. Nat Chem Biol 1(5):265–269. [CrossRef]
- Laville J, Blumer C, Von Schroetter C, Gaia V, Défago G, Keel C, Haas D (1998) Characterization of the hcnABC Gene Cluster Encoding Hydrogen Cyanide Synthase and Anaerobic Regulation by ANR in the Strictly Aerobic Biocontrol Agent Pseudomonas fluorescens CHA0. J Bacteriol 180(12):3187–3196. [CrossRef]
- Lee N, Hwang S, Kim J, Cho S, Palsson B, Cho B-K (2020) Mini review: Genome mining approaches for the identification of secondary metabolite biosynthetic gene clusters in Streptomyces. Computational and Structural Biotechnology Journal 18:1548–1556. [CrossRef]
- Letunic I, Bork P (2024) Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Research 52(W1):W78–W82. [CrossRef]
- Liu J, Yu H, Li S-M (2018) Expanding tryptophan-containing cyclodipeptide synthase spectrum by identification of nine members from Streptomyces strains. Appl Microbiol Biotechnol 102(10):4435–4444. [CrossRef]
- Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, He G, Chen Y, Pan Q, Liu Y, Tang J, Wu G, Zhang H, Shi Y, Liu Y, Yu C, Wang B, Lu Y, Han C, Cheung DW, Yiu S-M, Peng S, Xiaoqian Z, Liu G, Liao X, Li Y, Yang H, Wang J, Lam T-W, Wang J (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1(1):2047-217X-1–18. [CrossRef]
- Madhaiyan M, Saravanan VS, See-Too W-S, Volpiano CG, Sant’Anna FH, Faria Da Mota F, Sutcliffe I, Sangal V, Passaglia LMP, Rosado AS (2022) Genomic and phylogenomic insights into the family Streptomycetaceae lead to the proposal of six novel genera. International Journal of Systematic and Evolutionary Microbiology 72(10). [CrossRef]
- Meier-Kolthoff JP, Göker M (2019) TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun 10(1):2182. [CrossRef]
- Mishra A, Choi J, Choi S-J, Baek K-H (2017) Cyclodipeptides: An Overview of Their Biosynthesis and Biological Activity. Molecules 22(10):1796. [CrossRef]
- Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer ELL, Tosatto SCE, Paladin L, Raj S, Richardson LJ, Finn RD, Bateman A (2021) Pfam: The protein families database in 2021. Nucleic Acids Research 49(D1):D412–D419. [CrossRef]
- Nazarov AV, Anan’ina LN, Gorbunov AA, Pyankova AA (2022) Bacteria Producing Ectoine in the Rhizosphere of Plants Growing on Technogenic Saline Soil. Eurasian Soil Sc 55(8):1074–1081. [CrossRef]
- Orhan F, Parlak KU, Tabay D, Bozarı S (2023) Alleviation of the Cadmium Toxicity by Application of a Microbial Derived Compound, Ectoine. Water Air Soil Pollut 234(8):534. [CrossRef]
- Ozfidan-Konakci C, Yildiztugay E, Alp FN, Kucukoduk M, Turkan I (2020) Naringenin induces tolerance to salt/osmotic stress through the regulation of nitrogen metabolism, cellular redox and ROS scavenging capacity in bean plants. Plant Physiology and Biochemistry 157:264–275. [CrossRef]
- Pan H-Q, Yu S-Y, Song C-F, Wang N, Hua H-M, Hu J-C, Wang S-J (2015) Identification and Characterization of the Antifungal Substances of a Novel Streptomyces cavourensis NA4. Journal of Microbiology and Biotechnology 25(3):353–357. [CrossRef]
- Papini E, Bernard M, Bugnoli M, Milia E, Rappuoli R, Montecucco C (1993) Cell vacuolization induced by Helicobacter pylori : Inhibition by bafilomycins A1, B1, C1 and D. FEMS Microbiology Letters 113(2):155–159. [CrossRef]
- Patzer SI, Braun V (2010) Gene Cluster Involved in the Biosynthesis of Griseobactin, a Catechol-Peptide Siderophore of Streptomyces sp. ATCC 700974. J Bacteriol 192(2):426–435. [CrossRef]
- Pettit GR, Tan R, Melody N, Kielty JM, Pettit RK, Herald DL, Tucker BE, Mallavia LP, Doubek DL, Schmidt JM (1999) Antineoplastic agents. Part 409: Isolation and structure of montanastatin from a terrestrial actinomycete[1]1Dedicated to the memory of Professor Sir Derek H. R. Barton (1918–1998), a great chemist and friend.1. Bioorganic & Medicinal Chemistry 7(5):895–899. [CrossRef]
- Prieto C, García-Estrada C, Lorenzana D, Martín JF (2012) NRPSsp: non-ribosomal peptide synthase substrate predictor. Bioinformatics 28(3):426–427. [CrossRef]
- Sambangi P, Gopalakrishnan S (2023) Streptomyces-mediated synthesis of silver nanoparticles for enhanced growth, yield, and grain nutrients in chickpea. Biocatalysis and Agricultural Biotechnology 47:102567. [CrossRef]
- Sathya A, Vijayabharathi R, Kumari BR, Srinivas V, Sharma HC, Sathyadevi P, Gopalakrishnan S (2016) Assessment of a diketopiperazine, cyclo(Trp-Phe) from Streptomyces griseoplanus SAI-25 against cotton bollworm, Helicoverpa armigera (Lepidoptera: Noctuidae). Appl Entomol Zool 51(1):11–20. [CrossRef]
- Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31(19):3210–3212. [CrossRef]
- Snyder EE, Kampanya N, Lu J, Nordberg EK, Karur HR, Shukla M, Soneja J, Tian Y, Xue T, Yoo H, Zhang F, Dharmanolla C, Dongre NV, Gillespie JJ, Hamelius J, Hance M, Huntington KI, Jukneliene D, Koziski J, Mackasmiel L, Mane SP, Nguyen V, Purkayastha A, Shallom J, Yu G, Guo Y, Gabbard J, Hix D, Azad AF, Baker SC, Boyle SM, Khudyakov Y, Meng XJ, Rupprecht C, Vinje J, Crasta OR, Czar MJ, Dickerman A, Eckart JD, Kenyon R, Will R, Setubal JC, Sobral BWS (2007) PATRIC: The VBI PathoSystems Resource Integration Center. Nucleic Acids Research 35(Database):D401–D406. [CrossRef]
- Srinivas V, Naresh N, Pratyusha S, Ankati S, Govindaraj M, Gopalakrishnan S (2022) Exploring plant growth-promoting. Crop & Pasture Science 73(5):484–493. [CrossRef]
- Subramaniam G (2016) Plant growth promoting actinobacteria: a new avenue for enhancing the productivity and soil fertility of grain legumes. Springer Berlin Heidelberg, New York, NY.
- Subramaniam G, Thakur V, Saxena RK, Vadlamudi S, Purohit S, Kumar V, Rathore A, Chitikineni A, Varshney RK (2020) Complete genome sequence of sixteen plant growth promoting Streptomyces strains. Sci Rep 10(1):10294. [CrossRef]
- Subramaniam Gopalakrishnan (2011) Biocontrol of charcoal-rot of sorghum by actinomycetes isolated from herbal vermicompost. Afr J Biotechnol 10(79). [CrossRef]
- Sun M, Li L, Wang C, Wang L, Lu D, Shen D, Wang J, Jiang C, Cheng L, Pan X, Yang A, Wang Y, Zhu X, Li B, Li Y, Zhang F (2022) Naringenin confers defence against Phytophthora nicotianae through antimicrobial activity and induction of pathogen resistance in tobacco. Molecular Plant Pathology 23(12):1737–1750. [CrossRef]
- Tang J, Li Y, Zhang L, Mu J, Jiang Y, Fu H, Zhang Y, Cui H, Yu X, Ye Z (2023) Biosynthetic Pathways and Functions of Indole-3-Acetic Acid in Microorganisms. Microorganisms 11(8):2077. [CrossRef]
- Tietz JI, Schwalen CJ, Patel PS, Maxson T, Blair PM, Tai H-C, Zakai UI, Mitchell DA (2017) A new genome-mining tool redefines the lasso peptide biosynthetic landscape. Nat Chem Biol 13(5):470–478. [CrossRef]
- Ueda K, Oinuma K-I, Ikeda G, Hosono K, Ohnishi Y, Horinouchi S, Beppu T (2002) AmfS, an Extracellular Peptidic Morphogen in Streptomyces griseus. J Bacteriol 184(5):1488–1492. [CrossRef]
- Vargas Hoyos HA, Santos SN, Padilla G, Melo IS (2019) Genome Sequence of Streptomyces cavourensis 1AS2a, a Rhizobacterium Isolated from the Brazilian Cerrado Biome. Microbiol Resour Announc 8(18):e00065-19. [CrossRef]
- Vijayabharathi R, Gopalakrishnan S, Sathya A, Srinivas V, Sharma M (2018a) Deciphering the tri-dimensional effect of endophytic Streptomyces sp. on chickpea for plant growth promotion, helper effect with Mesorhizobium ciceri and host-plant resistance induction against Botrytis cinerea. Microbial Pathogenesis 122:98–107. [CrossRef]
- Vijayabharathi R, Gopalakrishnan S, Sathya A, Vasanth Kumar M, Srinivas V, Mamta S (2018b) Streptomyces sp. as plant growth-promoters and host-plant resistance inducers against Botrytis cinerea in chickpea. Biocontrol Science and Technology 28(12):1140–1163. [CrossRef]
- Vijayabharathi R, Kumari BR, Sathya A, Srinivas V, Abhishek R, Sharma HC, Gopalakrishnan S (2014) Biological activity of entomopathogenic actinomycetes against lepidopteran insects (Noctuidae: Lepidoptera). Can J Plant Sci 94(4):759–769. [CrossRef]
- Vijayabharathi R, Sathya A, Gopalakrishnan S (2018c) Extracellular biosynthesis of silver nanoparticles using Streptomyces griseoplanus SAI-25 and its antifungal activity against Macrophomina phaseolina, the charcoal rot pathogen of sorghum. Biocatalysis and Agricultural Biotechnology 14:166–171. [CrossRef]
- Wang P, Liu Z, Huang Y (2018) Complete genome sequence of soil actinobacteria Streptomyces cavourensis TJ430. J Basic Microbiol 58(12):1083–1090. [CrossRef]
- Widodo WS, Billerbeck S (2023) Natural and engineered cyclodipeptides: Biosynthesis, chemical diversity, and engineering strategies for diversification and high-yield bioproduction. Engineering Microbiology 3(1):100067. [CrossRef]
- Yildiztugay E, Ozfidan-Konakci C, Kucukoduk M, Turkan I (2020) Flavonoid Naringenin Alleviates Short-Term Osmotic and Salinity Stresses Through Regulating Photosynthetic Machinery and Chloroplastic Antioxidant Metabolism in Phaseolus vulgaris. Front Plant Sci 11:682. [CrossRef]
- Zabolotneva AA, Shatova OP, Sadova AA, Shestopalov AV, Roumiantsev SA (2022) An Overview of Alkylresorcinols Biological Properties and Effects. Journal of Nutrition and Metabolism 2022:1–12. [CrossRef]
- Zhang Y, Yao T, Jiang Y, Li H, Yuan W, Li W (2021) Deciphering a Cyclodipeptide Synthase Pathway Encoding Prenylated Indole Alkaloids in Streptomyces leeuwenhoekii. Appl Environ Microbiol 87(11):e02525-20. [CrossRef]

| ID | Annotation/Function | Source | Evidence |
| fig|1472664.5.peg.3363 | Ribokinase (EC 2.7.1.15) | RAST server | Code: idu(2);D-ribose_utilization idu(2);Deoxyribose_and_Deoxynucleoside_Catabolism |
| fig|1472664.5.peg.4738 | PE-PGRS FAMILY PROTEIN | RAST server | Not provided |
| fig|1472664.5.peg.5531 | Xanthine dehydrogenase, molybdenum binding subunit (EC 1.17.1.4) | RAST server | Code: icw(2);Purine_Utilization icw(2);Xanthine_dehydrogenase_subunits |
| fig|1472664.5.peg.1814 | ligA protein [Mycobacterium pseudoshottsii JCM 15466] | NCBI BLAST followed by Reciprocal Best BLAST | Accession: GAQ32343.1 e-value: 2.36E-04; Alignment length: 461 Percentage identity: 32.936 Query coverage (fig|1472664.5.peg.1814): 81%; Subject coverage (GAQ32343.1): 85% |
| GenBank accession no.: | KF770901 |
| Source of isolation: | Rice rhizosphere soil |
| Temperature tolerance: | 20–40°C |
| PGP and biocontrol traits: | Siderophore+, chitinase+, cellulase+, lipase+, protease+, indole-3-acetic acid+ and hydrocyanic acid+ |
| Entomopathogenic traits: | Helicoverpa armigera, Spodoptera litura and Chilo partellus |
| Metabolite identified: | Cyclo(Trp-Phe), a diketopiperazine derivative with insecticidal activity on H. armigera. |
| Paired-end (100 bp) | Mate-pair (250 bp) | |
| Raw reads | 50,83,396 | 1,34,97,102 |
| Clean reads | 48,81,906 | 1,03,71,537 |
| Features | Value | |
| Assembly details | Contig count | 1 |
| Genome length | 7,733,723 bp | |
| No. of plasmids | 1 | |
| Total no. of non-ATCG bases | 19,290 (0.25%) | |
| Number of Ns per 100kb | 249.43 | |
| GC content | 72.12% | |
| Contig L50 | 1 | |
| Contig N50 | 7,733,723 | |
| Annotated genome | Coding density | 88.91% |
| Coding seq. count | 6,923 | |
| Coding seq. mean length | 989.9 bp | |
| tRNA gene count | 74 | |
| tRNA mean length | 76.01 | |
| rRNA gene count | 3 | |
| rRNA mean length | 1,595.33 | |
| Count of repeats | 152 | |
| Repeat mean length | 126.87 | |
| CRISPR spacer count | 81 | |
| CRISPR spacer mean length | 32.42 | |
| Proteins | Count of Hypothetical proteins | 2,363 (34.13%) |
| Count of proteins with functional assignment | 4,560 (65.86%) | |
| Count of proteins with EC number assignment | 1,207 |
| Property | Source DB | No. of genes |
| Antibiotic resistance | PATRIC | 48 |
| Drug targets | Drug Bank | 6 |
| Drug targets | TTD | 1 |
| Transporter | TCDB | 36 |
| Virulence factors | PATRIC_VF | 3 |
| AMR mechanism | Genes |
| Antibiotic activation enzyme | katG |
| Antibiotic inactivation enzymes | AAC(2’)-I |
| Antibiotic target in susceptible genes | Alr,Ddl, dxr, EF-G, EF-Tu, folA, Dfr, folP, gyrA, gyrB, inhA, Fabl, Iso-tRNA, kasA, MurA, rho, rpoB, rpoC, S10p, S12p |
| Antibiotic target replacement protein | FabG, HtdX |
| Efflux pump conferring antibiotic resistance | CmIV family, Otr(C) |
| Gene conferring resistance via absence | gldB |
| Protein-altering cell wall charge | GdpD, MprF, PgsA |
| Regulator modulating expression of antibiotic resistance genes | LpqB, MtrA, MtrB, OxyR |
| ID | Annotation/Function | Source | Evidence |
| fig|1472664.5.peg.3363 | Ribokinase (EC 2.7.1.15) | RAST server | Code: idu(2);D-ribose_utilization idu(2);Deoxyribose_and_Deoxynucleoside_Catabolism |
| fig|1472664.5.peg.4738 | PE-PGRS FAMILY PROTEIN | RAST server | Not provided |
| fig|1472664.5.peg.5531 | Xanthine dehydrogenase, molybdenum binding subunit (EC 1.17.1.4) | RAST server | Code: icw(2);Purine_Utilization icw(2);Xanthine_dehydrogenase_subunits |
| fig|1472664.5.peg.1814 | ligA protein [Mycobacterium pseudoshottsii JCM 15466] | NCBI BLAST followed by Reciprocal Best BLAST | Accession: GAQ32343.1 e-value: 2.36E-04; Alignment length: 461 Percentage identity: 32.936 Query coverage (fig|1472664.5.peg.1814): 81%; Subject coverage (GAQ32343.1): 85% |
| Metabolites | Biosynthetic gene cluster | Functions | References |
| Geosmin | Region 5 | Regulates seed germination and acts as a chemical repellent/attractant to predators (nematodes and protists) and insects | (Garbeva et al. 2023) |
| Griseobactin | Region 6 | Siderophore | (Patzer and Braun 2010) |
| Coelichelin | Region 7 | Siderophore | (Lautru et al. 2005) |
| Naringenin | Region 8 | Alleviates abiotic stress (osmotic and salinity stress) and also contributes to pathogen resistance in plants | (Yildiztugay et al. 2020; Ozfidan-Konakci et al. 2020; An et al. 2021; Sun et al. 2022) |
| Desferrioxamine B | Region 11 | Siderophore | (Bellotti and Remelli 2021) |
| Ectoine | Region 16 | An osmoprotectant that alleviates cadmium-induced stress in plants | (Nazarov et al. 2022; Orhan et al. 2023) |
| AmfS | Region 17 | Whose derivative acts as an extracellular morphogen for the onset of aerial mycelium | (Ueda et al. 2002) |
| Biosynthesis of type II polyketide backbone | Region 18* | It is utilised for the biosynthesis of type II polyketide products | Figure S3 |
| Keywimysin | Region 19 | A lasso peptide whose biological function remains unknown | (Tietz et al. 2017) |
| Terpenoid backbone biosynthesis | Region 20* | It is utilised in sesquiterpenoids and triterpenoids biosynthesis | Figure S4 |
| D-Amino acid metabolism | Region 21* | It plays a role in the production of D-proline, which is utilised for biosynthesis of linatine (a vitamin B6 antagonist) | Figure S5 and (Klosterman et al. 1967) |
| Bafilomycin B1 | Region 25 | A macrolide antibiotic that inhibits vacuolar-type ATPase (V-ATPase) | (Bowman et al. 1988; Papini et al. 1993) |
| 10-epi-HSAF and its analogues | Region 26 | Shows antifungal activities against plant pathogens | (Hou et al. 2020) |
| Valinomycin and Montanastatin | Region 28 | Valinomycin is a potassium ionophore which demonstrates a diverse spectrum of biological activities (antibacterial, antifungal, insecticidal, etc.), and Montanastatin is a cancer cell growth inhibitory cyclooctadepsipeptide | (Pettit et al. 1999; Huang et al. 2021) |
| Alkylresorcinol | Region 30 | A polyketide which exhibits a wide range of bioactivities (antimicrobial, anti-cancer, antilipidemic, antioxidant, etc.) | (Zabolotneva et al. 2022) |
| Isorenieratene | Region 31 | A natural antioxidant and photo/UV damage inhibitor | (Chen et al. 2019) |
| Protein ID | RAST annotation | Biosynthetic gene cluster | Number of A-domains |
| fig|1472664.5.peg.6606 | hypothetical protein | Region 28 | 2 |
| fig|1472664.5.peg.6542 | Polyketide synthase modules and related proteins | Region 27 | 2 |
| fig|1472664.5.peg.481 | Siderophore biosynthesis non-ribosomal peptide synthetase modules | Region 7 | 3 |
| fig|1472664.5.peg.6541 | Siderophore biosynthesis non-ribosomal peptide synthetase modules | Region 27 | 2 |
| fig|1472664.5.peg.429 | Siderophore biosynthesis non-ribosomal peptide synthetase modules | Region 6 | 2 |
| fig|1472664.5.peg.5776 | Polyketide synthase modules and related proteins | Region 22 | 1 |
| fig|1472664.5.peg.2757 | Polyketide synthase modules and related proteins | Region 13 | 1 |
| fig|1472664.5.peg.6452 | Capsular polysaccharide biosynthesis fatty acid synthase WcbR | Region 26 | 1 |
| fig|1472664.5.peg.6605 | hypothetical protein | Region 28 | 2 |
| fig|1472664.5.peg.2758 | Polyketide synthase modules and related proteins | Region 13 | 1 |
| fig|1472664.5.peg.6533 | Polyketide synthase modules and related proteins | Region 27 | 1 |
| fig|1472664.5.peg.392 | Polyketide synthase modules and related proteins | Region 6 | 1 |
| fig|1472664.5.peg.6862 | Polyketide synthase modules and related proteins | Region 32 | 2 |
| fig|1472664.5.peg.2755 | Polyketide synthase modules and related proteins | Region 13 | 1 |
| fig|1472664.5.peg.5774 | hypothetical protein | Region 22 | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




