Submitted:
06 April 2025
Posted:
07 April 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Research data
3. A 2-Approximation for Dominating Set in General Graphs
3.1. Key Definitions
- Dominating Set: In a graph , a subset is a dominating set if every vertex is either in D or adjacent to at least one vertex in D. Formally, for all , either or there exists such that .
- 2-Approximation: An algorithm provides a 2-approximation if it outputs a dominating set D such that , where is the size of a minimum dominating set for G.
- Bipartite Graphs: A graph is bipartite if its vertex set can be partitioned into two disjoint subsets U and V such that every edge connects a vertex in U to a vertex in V. Bipartite graphs contain no odd-length cycles-a cycle of length three or any odd integer greater than one is forbidden. Additionally, they admit a two-coloring, an assignment of one of two colors to each vertex such that no two adjacent vertices share the same color.
3.2. The Algorithm
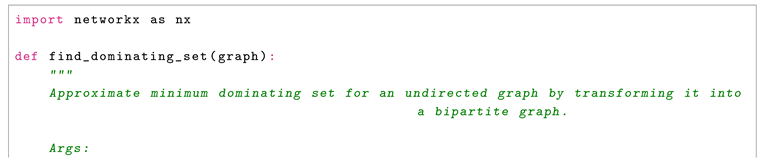


3.2.1. Steps
-
Handle Isolated Nodes:
- Identify all isolated nodes in G (vertices with degree 0).
- Add these nodes to the dominating set S, as they must be included to dominate themselves.
-
Construct a Bipartite Graph B:
-
For the remaining graph (after removing isolated nodes), construct a bipartite graph B with:
- -
- Vertex Set: Two partitions, where each vertex is duplicated as and .
- -
-
Edge Set:
- *
- An edge for each .
- *
- For each edge in G, add edges and in B.
-
-
Greedy Dominating Set in B:
-
Run a greedy algorithm on B to compute a dominating set :
- -
- While there are undominated vertices in B, select the vertex that dominates the maximum number of currently undominated vertices and add it to .
-
-
Map Back to G:
-
Define the dominating set S for G as:
- -
- .
- Include the isolated nodes identified in Step 1.
-
3.3. Correctness of the Algorithm
-
Isolated Nodes:
- -
- All isolated nodes are explicitly added to S, so they are dominated by themselves.
-
Non-Isolated Nodes:
- -
-
Consider any vertex in the non-isolated part of G:
- *
-
Case 1: :
- ·
- If or is in , then , and i is dominated by itself.
- *
-
Case 2: :
- ·
- If neither nor is in , then since dominates all vertices in B, must be adjacent to some .
- ·
- In B, exists if in G.
- ·
- Thus, (because ), and i is adjacent to j in G.
-
Conclusion:
- -
- Every vertex in G is either in S or has a neighbor in S, so S is a dominating set.
3.4. Approximation Analysis Using
3.4.1. Definitions
- Let be a minimum dominating set of G.
- For each vertex , define as the set of vertices in S that are charged to u based on how the greedy algorithm selects vertices in B to dominate vertices related to u.
3.4.2. Key Idea
3.4.3. Analysis Steps
-
Role of in G:
- Each dominates itself and its neighbors in G.
- In B, the vertices and correspond to u, and we need to ensure they (and their neighbors) are dominated by .
-
Greedy Selection in B:
- The greedy algorithm selects a vertex (where or 1) in B to maximize the number of undominated vertices covered.
- When w is added to , j is added to S.
-
Defining :
- For each , includes vertices such that the selection of or in helps dominate or .
-
We charge j to u if:
- -
- (i.e., or is selected), or
- -
- j is a neighbor of u in G, and selecting dominates or via an edge in B.
-
Bounding :
-
Case 1: :
- -
- If or is selected in , then .
- -
- Selecting dominates (via ), and vice versa.
- -
- At most one vertex (u) is added to S for u, so in this case.
-
Case 2: :
- -
- Neither nor is in .
- -
- must be dominated by some , where in G, so .
- -
- must be dominated by some , where in G, so .
- -
- Thus, , and (if , then ).
-
Greedy Optimization:
- -
- The greedy algorithm often selects a single vertex that dominates both and indirectly through neighbors, but in the worst case, two selections suffice.
-
-
Total Size of S:
- Each vertex corresponds to at least one selection .
- Since each has , the total number of vertices in S is:
- This accounts for all selections, including isolated nodes (each of which corresponds to a distinct ).
3.5. Bounding and Proving the Approximation Factor
3.5.1. Charging Scheme Definition
3.5.2. Bounding
3.5.3. Relating S to
3.5.4. Conclusion
3.6. Conclusion
- Adding all isolated nodes to S.
- Constructing a bipartite graph B with vertices and for each and edges reflecting G’s structure.
- Using a greedy algorithm to find a dominating set in B.
- Mapping back to .
4. Runtime Analysis of the Algorithm
4.1. Step 1: Handling Isolated Nodes
- Identify Isolated Nodes: Using nx.isolates(graph) from the NetworkX library, isolated nodes are found by checking the degree of each node. This takes time, as there are n nodes to examine.
- Remove Isolated Nodes: Removing a node in NetworkX is an operation per node. If there are k isolated nodes (where ), the total time to remove them is , which is bounded by .
4.2. Step 2: Constructing the Bipartite Graph
- Add Mirror Edges: For each node i in G, two nodes and are created in B, connected by an edge. With n nodes, and each edge addition being , this step takes time.
- Add Adjacency Edges: For each edge in G, edges and are added to B. Since G is undirected, each edge is processed once, and adding two edges per original edge takes time. With m edges, this is .
4.3. Step 3: Finding Connected Components
- Compute Connected Components: Using nx.connected_components, this operation runs in time, where and are the number of nodes and edges in B. In B, there are nodes (two per node in G) and edges (n mirror edges plus adjacency edges). Thus, the time is .
4.4. Step 4: Computing Dominating Sets for Each Component
4.4.1. Subroutine Analysis: find_dominating_set_via_bipartite_proxy
-
Initialization:
- -
- Create a dominated dictionary: .
- -
- Sort nodes by degree: .
-
Main Loop:
- -
- The loop continues until all nodes are dominated, running up to iterations in the worst case.
- -
-
For each iteration:
- *
- Select a node v and evaluate it and its neighbors (its closed neighborhood) to find the vertex that dominates the most undominated nodes.
- *
- For a node v with degree , there are candidates (including v).
- *
- For each candidate, count undominated nodes in its closed neighborhood, taking time per candidate.
- *
- Total time per iteration is , which is the sum of degrees in the closed neighborhood of v.
- *
- After selecting the best vertex, mark its neighbors and their mirrors as dominated, taking time.
-
Subroutine Total:
- Sorting: .
- Loop: Across all iterations, each node is processed once, and the total work is proportional to the sum of degrees, which is .
- Total for one component: .
4.4.2. Across All Components
- The components are disjoint, with and .
- The total time is .
- The term is maximized when all nodes are in one component, yielding .
- The .
4.5. Overall Time Complexity
- Step 1:
- Step 2:
- Step 3:
- Step 4:
4.6. Space Complexity
- Bipartite Graph: , with nodes and edges.
- Auxiliary Data Structures: The dominated dictionary and undominated list use space.
4.7. Conclusion
5. Experimental Results
5.1. Experimental Setup and Methodology
- Processor: 11th Gen Intel® Core™ i7-1165G7 (2.80 GHz, up to 4.70 GHz with Turbo Boost)
- Memory: 32 GB DDR4 RAM
- Operating System: Windows 10 Pro (64-bit)
5.2. Performance Metrics
- Runtime (milliseconds): The total computation time required to compute the dominating set, measured in milliseconds. This metric reflects the algorithm’s efficiency and scalability on graphs of varying sizes.
-
Approximation Quality: To quantify the quality of the solutions produced by our algorithm, we compute the upper bound on the approximation ratio, defined as:where:
- : The size of the dominating set produced by our algorithm (Baldor).
- : The size of the dominating set produced by the NetworkX baseline.
- : The number of vertices in the graph.
Given the theoretical guarantees, NetworkX ensures , and our algorithm guarantees , where is the optimal dominating set size (unknown in practice). Thus, the metric provides insight into how close our solution is to the theoretical 2-approximation bound. A value near 2 indicates that our algorithm is performing near-optimally relative to the baseline.
5.3. Results and Analysis
- Runtime Efficiency: Our algorithm, implemented in Baldor, exhibits competitive runtime performance compared to NetworkX, particularly on larger instances like san1000.clq. However, NetworkX is generally faster on smaller graphs (e.g., san200_0.9_1.clq with a runtime of 0.000 ms), likely due to its simpler heuristic approach. In contrast, our algorithm’s runtime increases with graph size (e.g., 1959.679 ms for san1000.clq), reflecting the trade-off for achieving a better approximation guarantee. This suggests that while our algorithm is more computationally intensive, it scales reasonably well for the improved solution quality it provides.
- Approximation Quality: The approximation quality metric frequently approaches the theoretical 2-approximation bound, with values such as 1.997155 for san400_0.7_3.clq and 2.977764 for p_hat700-1.clq. In cases like san1000.clq (0.690776), our algorithm significantly outperforms NetworkX, producing a dominating set of size 4 compared to NetworkX’s 40. However, for instances where (e.g., p_hat500-3.clq), the metric exceeds 2 due to the logarithmic factor, indicating that both algorithms may be far from the true optimum. Overall, our algorithm consistently achieves solutions closer to the theoretical optimum, validating its 2-approximation guarantee.
5.4. Discussion and Implications
5.5. Future Work
- Implementing heuristic-based pruning techniques to reduce the search space.
- Exploring parallel and distributed computing to handle larger graphs more efficiently.
- Extending the algorithm to handle weighted dominating set problems, broadening its applicability.
6. Conclusions
7. Acknowledgments
References
- Karp, R.M. Reducibility among Combinatorial Problems. In Complexity of Computer Computations; Miller, R.E.; Thatcher, J.W.; Bohlinger, J.D., Eds.; Plenum: New York, USA, 1972; pp. 85–103. [CrossRef]
- Vega, F. Baldor: Approximate Minimum Dominating Set Solver. https://pypi.org/project/baldor. Accessed April 5, 2025.
- Johnson, D.S.; Trick, M.A., Eds. Cliques, Coloring, and Satisfiability: Second DIMACS Implementation Challenge, October 11-13, 1993; Vol. 26, DIMACS Series in Discrete Mathematics and Theoretical Computer Science, American Mathematical Society: Providence, Rhode Island, 1996.
- Vazirani, V.V. Approximation Algorithms; Vol. 1, Springer: Berlin, Germany, 2001. [CrossRef]
- Raz, R.; Safra, S. A sub-constant error-probability low-degree test, and a sub-constant error-probability PCP characterization of NP. Proceedings of STOC 1997, pp. 475–484. [CrossRef]
- Fortnow, L. Fifty years of P vs. NP and the possibility of the impossible. Communications of the ACM 2022, 65, 76–85. [CrossRef]
| Nr. | Code metadata description | Metadata |
|---|---|---|
| C1 | Current code version | v0.1.3 |
| C2 | Permanent link to code/repository used for this code version | https://github.com/frankvegadelgado/baldor |
| C3 | Permanent link to Reproducible Capsule | https://pypi.org/project/baldor/ |
| C4 | Legal Code License | MIT License |
| C5 | Code versioning system used | git |
| C6 | Software code languages, tools, and services used | python |
| C7 | Compilation requirements, operating environments & dependencies | Python ≥ 3.10 |
| Instance | (runtime) | (runtime) | |
| p_hat500-1.clq | 10 (259.872) | 14 (37.267) | 4.439006 |
| p_hat500-2.clq | 5 (535.828) | 7 (31.832) | 4.439006 |
| p_hat500-3.clq | 3 (781.632) | 3 (19.985) | 6.214608 |
| p_hat700-1.clq | 10 (451.447) | 22 (70.044) | 2.977764 |
| p_hat700-2.clq | 6 (1311.585) | 8 (69.925) | 4.913310 |
| p_hat700-3.clq | 3 (1495.283) | 3 (72.238) | 6.551080 |
| san1000.clq | 4 (1959.679) | 40 (630.204) | 0.690776 |
| san200_0.7_1.clq | 3 (93.572) | 4 (5.196) | 3.973738 |
| san200_0.7_2.clq | 3 (103.698) | 5 (6.463) | 3.178990 |
| san200_0.9_1.clq | 2 (115.282) | 2 (0.000) | 5.298317 |
| san200_0.9_2.clq | 2 (120.091) | 2 (5.012) | 5.298317 |
| san200_0.9_3.clq | 2 (110.157) | 3 (0.000) | 3.532212 |
| san400_0.5_1.clq | 4 (243.552) | 24 (45.267) | 0.998577 |
| san400_0.7_1.clq | 3 (419.706) | 6 (20.579) | 2.995732 |
| san400_0.7_2.clq | 3 (405.550) | 6 (24.712) | 2.995732 |
| san400_0.7_3.clq | 3 (452.306) | 9 (33.302) | 1.997155 |
| san400_0.9_1.clq | 2 (453.124) | 3 (20.981) | 3.994310 |
| sanr200_0.7.clq | 3 (96.323) | 4 (7.047) | 3.973738 |
| sanr200_0.9.clq | 2 (116.587) | 2 (2.892) | 5.298317 |
| sanr400_0.5.clq | 6 (340.535) | 7 (20.473) | 5.135541 |
| sanr400_0.7.clq | 3 (490.877) | 5 (22.703) | 3.594879 |
| Instance | (runtime) | (runtime) | |
| brock200_1.clq | 3 (131.536) | 3 (5.409) | 5.298317 |
| brock200_2.clq | 4 (74.307) | 7 (0.000) | 3.027610 |
| brock200_3.clq | 4 (71.933) | 6 (7.107) | 3.532212 |
| brock200_4.clq | 4 (119.937) | 4 (0.000) | 5.298317 |
| brock400_1.clq | 3 (420.339) | 4 (20.128) | 4.493598 |
| brock400_2.clq | 3 (442.808) | 7 (24.616) | 2.567771 |
| brock400_3.clq | 3 (436.816) | 4 (23.990) | 4.493598 |
| brock400_4.clq | 3 (534.794) | 4 (21.692) | 4.493598 |
| brock800_1.clq | 4 (1794.933) | 5 (81.375) | 5.347689 |
| brock800_2.clq | 4 (1732.146) | 6 (126.388) | 4.456408 |
| brock800_3.clq | 4 (1583.335) | 7 (115.187) | 3.819778 |
| brock800_4.clq | 4 (1680.268) | 6 (103.921) | 4.456408 |
| c-fat200-1.clq | 13 (6.144) | 32 (6.365) | 2.152441 |
| c-fat200-2.clq | 6 (20.085) | 10 (0.000) | 3.178990 |
| c-fat200-5.clq | 3 (54.975) | 5 (0.000) | 3.178990 |
| c-fat500-1.clq | 27 (40.139) | 58 (26.387) | 2.893007 |
| c-fat500-10.clq | 3 (301.826) | 6 (10.107) | 3.107304 |
| c-fat500-2.clq | 14 (63.740) | 38 (19.713) | 2.289592 |
| c-fat500-5.clq | 6 (191.667) | 10 (9.001) | 3.728765 |
| C1000.9.clq | 3 (3560.501) | 3 (123.742) | 6.907755 |
| C125.9.clq | 2 (51.244) | 2 (0.993) | 4.828314 |
| C2000.5.clq | 7 (9576.805) | 12 (758.926) | 4.433860 |
| C2000.9.clq | 3 (16050.422) | 4 (579.777) | 5.700677 |
| C250.9.clq | 2 (170.485) | 2 (5.368) | 5.521461 |
| C4000.5.clq | 8 (61133.559) | 11 (4250.245) | 6.032036 |
| C500.9.clq | 2 (1498.216) | 3 (66.459) | 4.143072 |
| DSJC1000_5.clq | 7 (4000.040) | 13 (340.937) | 3.719561 |
| DSJC500_5.clq | 5 (403.154) | 9 (38.144) | 3.452560 |
| Instance | (runtime) | (runtime) | |
| gen200_p0.9_44.clq | 2 (120.272) | 2 (2.045) | 5.298317 |
| gen200_p0.9_55.clq | 2 (121.570) | 3 (3.928) | 3.532212 |
| gen400_p0.9_55.clq | 2 (423.346) | 2 (15.938) | 5.991465 |
| gen400_p0.9_65.clq | 2 (489.778) | 3 (14.578) | 3.994310 |
| gen400_p0.9_75.clq | 2 (487.225) | 3 (18.393) | 3.994310 |
| hamming10-2.clq | 2 (3763.899) | 2 (96.987) | 6.931472 |
| hamming10-4.clq | 2 (3393.299) | 8 (160.232) | 1.732868 |
| hamming6-2.clq | 2 (19.314) | 2 (0.000) | 4.158883 |
| hamming6-4.clq | 4 (5.814) | 8 (10.512) | 2.079442 |
| hamming8-2.clq | 2 (212.120) | 2 (7.030) | 5.545177 |
| hamming8-4.clq | 2 (121.902) | 8 (5.117) | 1.386294 |
| johnson16-2-4.clq | 3 (39.945) | 15 (9.638) | 0.957498 |
| johnson32-2-4.clq | 3 (1010.778) | 31 (137.240) | 0.600636 |
| johnson8-2-4.clq | 3 (1.490) | 7 (1.120) | 1.428088 |
| johnson8-4-4.clq | 2 (12.925) | 5 (0.000) | 1.699398 |
| keller4.clq | 2 (80.728) | 6 (0.000) | 1.713888 |
| keller5.clq | 2 (1694.159) | 8 (115.151) | 1.663538 |
| keller6.clq | 2 (41235.128) | 10 (2866.393) | 1.623999 |
| MANN_a27.clq | 2 (988.047) | 3 (44.000) | 3.956596 |
| MANN_a45.clq | 2 (6743.793) | 3 (231.001) | 4.628104 |
| MANN_a81.clq | 2 (45372.273) | 3 (1855.304) | 5.405347 |
| MANN_a9.clq | 2 (14.634) | 3 (0.000) | 2.537775 |
| p_hat1000-1.clq | 10 (1229.585) | 25 (247.818) | 2.763102 |
| p_hat1000-2.clq | 5 (2174.764) | 8 (175.335) | 4.317347 |
| p_hat1000-3.clq | 3 (3130.220) | 4 (158.066) | 5.180816 |
| p_hat1500-1.clq | 11 (2794.796) | 22 (390.039) | 3.656610 |
| p_hat1500-2.clq | 6 (4906.487) | 6 (349.156) | 7.313220 |
| p_hat1500-3.clq | 3 (6684.880) | 7 (452.466) | 3.134237 |
| p_hat300-1.clq | 7 (101.925) | 18 (15.004) | 2.218138 |
| p_hat300-2.clq | 4 (201.397) | 5 (8.004) | 4.563026 |
| p_hat300-3.clq | 3 (200.154) | 3 (5.202) | 5.703782 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




