1. Introduction
Verticillium wilt caused by the soil-borne fungus
Verticillium dahliae Kleb. is among the most devastating plant diseases affecting more than 350 plant species worldwide [
1]. The disease causes severe losses to several economically important crops including vegetables, fruits, ornamentals, oilseed crops, fiber crops and woody perennials, and is also capable of invading several weeds [
2]. Due to its broad host range and capacity to survive for up to fifteen years in the soil forming resistant structures named ‘microsclerotia’ and benefiting from the lack of effective fungicides,
V. dahliae is especially difficult to manage [
3]. Therefore, management strategies are focused on preventive measures such as solarization of heavily infested soils, the use of resistant/tolerant genotypes, attempts for biological control practices including the use of compost soil amendments or direct application of microorganisms with proven protective activity against the pathogen, or combinations of the above.
On the other hand, agricultural-oriented countries produce massive quantities of agro-industrial by-products and face the challenge of managing such materials in an environmentally friendly and agronomically efficient manner. This is the case for the significant amounts of olive mill wastewater (OMW) that are generated annually during the olive oil extraction process in major olive-producing countries [
4]. OMW constitutes a serious environmental problem in the Mediterranean regions that account for approximately 95% of the world olive oil production due to its high organic load, phytotoxicity and antimicrobial effects. Amongst other agro-industrial by-products, OMW has been proposed to be used in beneficial ways as biofertilizers in several crops or biological protectants against a broad range of soil-borne and aerial pathogens [
4].
In the last two decades OMW has been verified as a valuable source of resistance for plants against numerous pathogens. In respect of verticillium wilt, several studies demonstrated the suppressive effect of OMW-derived composts, extracts or microorganisms against
V. dahliae in-vitro and in vegetable crops
in-planta [
5,
6,
7,
8,
9]. Phenolic substances are among the abiotic factors that have been found to contribute to the suppressive activity of OMW-associated compost amendments and extracts against
V. dahliae [
5,
7,
9], whereas the biological nature of the disease suppressive mechanism has also been addressed by isolating and screening certain microbial strains [
8]. However, the potential suppressive effect of raw OMW material against verticillium wilt
in-planta has not been investigated previously.
Land spreading of OMW is an alternative management practice in several Mediterranean countries that have introduced specific legislation, having several beneficial effects in crops when applied in proper dosage and timing [
4,
10]. OMW application in soil induces changes in soil physical and chemical properties with cascading effects on the structure of the soil bacterial communities [
10,
11]. Moreover, OMW is a microbial-rich material containing many distinct groups of microorganisms including numerous bacterial, yeast and fungal genera [
12,
13,
14]. Despite the microbial nature of OMW and the observed alterations in the structure of soil bacterial communities upon OMW application, little is known regarding the effects of OMW land application on the plant microbiome of treated plants and particularly on its role in disease suppression. Therefore, the main objectives of the present study were to: i) evaluate the suppressive effect of raw OMW against verticillium wilt in vegetable crops, ii) investigate the disease suppressive mechanisms and iii) study the potential effect of OMW on the eukaryotic and prokaryotic plant microbiome in OMW-treated plants in respect to disease suppression.
3. Discussion
Verticillium wilt caused by
V. dahliae constitutes a serious threat for a broad range of economically important crops worldwide, since no effective chemical control is available. Therefore, alternative control strategies are intensively investigated. On the other hand, olive-producing countries face the challenge of managing olive mill waste (OMW) in an environmentally friendly and agronomically beneficial manner. Scientifically guided application of OMW to land has been proposed as a feasible way to successfully manage olive-derived wastes and plant diseases [
4,
10]. Herein, we tested whether soil application of raw OMW can protect vegetable crops against
V. dahliae and investigated the potential disease suppressive mechanisms.
Apart from microsclerotial area, OMW reduced or nullified all fungal parameters evaluated, indicating a significant antifungal activity of OMW against
V. dahliae in-vitro. Correlation analyses revealed that the effects of OMW on growth rate, sporulation, spore germination and microsclerotium germination were dose-depended. Such fungicidal activity of raw OMW, processed OMW (e.g. hydroxytyrosol-rich OMW, composted olive waste extracts) and individual components that are contained in OMW (e.g. the phenolic compounds oleuropein and hydroxytyrosol) against
V. dahliae has been shown in several studies [
5,
9,
15,
16,
17,
18]. However, in all the above studies,
in-vitro inhibitory effects were assessed only on mycelial growth and/or conidial germination of
V. dahliae. In the present study, several microbial parameters were affected with inhibition of microsclerotium germination being the most important. Microsclerotia constitute the primary inoculum structures in soil which germinate and infect host plants in nature [
1]. Therefore, inhibition of microsclerotia germination due to OMW application in soil, could lead to decreased disease incidence and thus, effective disease management in the field.
Root-drench application of OMW provided significant protection of eggplant and tomato against
V. dahliae in-planta. Likewise, reduction of verticillium wilt symptom severity in horticultural crops and cotton by applying olive derivatives (like OMW composts, hydroxytyrosol-rich OMW and OMW-originating microorganisms) has also been reported [
6,
7,
8,
9,
18,
19]. However, the suppressive effect of raw OMW material against verticillium wilt
in-planta has never been shown previously. The aspect of direct application of raw OMW in the regions of origin is an important matter since high regional scattering of olive mills in several countries like Greece could make the re-use of OMW completely unaffordable due to increased transportation and processing costs [
20].
Τhe high OMW-conferred protection of eggplant in November 2019 gradually decreased in the following experiments conducted with the same OMW material, possibly due to the decreased phenolic content in OMW over time. Indeed, total phenols content in non-sterilized OMW at the end (October 2020) was significantly lower than the one measured at the beginning (July 2019) of the experiments. Feki et al. [
21] also reported a remarkable decrease in concentrations of several phenolic compounds occurring in OMW after 5 months of storage whereas hydroxytyrosol was the only compound that increased significantly in the above meantime. A positive association between the total phenols content in composts and their suppressive effect against verticillium wilt in eggplant was also demonstrated by Markakis et al. [
7].
Interestingly, re-isolations performed from OMW-treated plants inoculated with
V. dahliae revealed several fungi (mostly
Trichoderma-like strains) emerging from xylem chips solely or simultaneously with
V. dahliae. Such fungal contaminants were not observed in
V. dahliae-inoculated plants that had not been treated with OMW, indicating a differential texture of cultivable endophytic fungal inhabitants in OMW-treated plants compared to the non-treated ones. Although alterations in the structure of soil microbiome consequential of OMW application have been shown in the past [
10,
11,
22], alterations in cultivable plant microbiome have never been reported previously.
Based on microbiome analysis, no significant differences were revealed among treatments that could be associated with disease suppressiveness. Some notable differences were identified between non-inoculated and
V. dahliae-inoculated plants, irrespective of the treatment with OMW. This is the case of the bacterial ASV0020 belonging to the genus
Flexivigra that was identified at significantly higher abundance in the control tomato plants compared to their
V. dahliae-inoculated counterparts, suggesting a decreasing abundance pattern upon
V. dahliae infection. Indeed, Snelders et al. [
23] showed that
V. dahliae secrets plant natriuretic peptide-like effectors that reduce the abundance of species of
Actinobacteria (including
Flexivigra), which otherwise reduces pathogen invasion. Likewise, fungal ASVs representing
Apiotrichum (ASV0003),
Exobasidium (ASV00123) and
Alternaria (ASV0028) were found at higher abundance in control tomato plants compared to the
V. dahliae-inoculated ones, suggesting a similar decreasing pattern as that mentioned above. In the case of eggplant, the abundance of the genera
Candidata (ASV00030),
Simplicillium (ASV00005) and
Cladosporium (ASV00036) was significantly higher in the control plants, whereas
Nadsonia (ASV00077) and
Verticillium (ASV00002) were comparatively higher in plants that have been inoculated with
V. dahliae. The fungal genera
Alternaria,
Cladosporium and
Simplicillium have been associated with plant welfare and have been pointed out either as potential biocontrol/plant growth promoting agents or as indicators of the asymptomatic status in plants [
24,
25,
26]. Differential abundance analysis along with fungal re-isolation and qPCR revealed a significantly lower
V. dahliae colonization in OMW-treated eggplants inoculated with the fungus compared to the non-treated ones. The observed lower fungal colonization was associated with decreased symptom severity.
Bioassays with sterilized OMW and/or soil suggest an unlikely association of soil- and OMW-originated microbiome with disease suppressive mechanism, since heat sterilization did not affect the protective activity of OMW
in-planta. This hypothesis is strengthened by the fact that the microbial strains isolated from OMW-treated soil and from asymptomatic OMW-treated plants, could not provide sufficient protection of plants in pathogenicity assays. Contrarywise, other studies demonstrated that heat sterilization of olive derivatives resulted in complete or partial loss of their suppressive activity against
V. dahliae, pointing out the microbial nature of their disease suppressive mechanism [
8,
17]. Moreover, in contrast to studies that speculate the induction of defense response mechanism in the plant upon OMW application [
4,
9], the non-significant protection of plants in split-root set ups presented here, indicates a non-systemic OMW-induced resistance mechanism in plants.
In view of all the above, data of the present study suggest that root-drench application of raw OMW can be an efficient practice to protect vegetable crops against V. dahliae and a sustainable way to manage OMW in olive-producing countries that have enacted specific legislation. It is evident that further study under field and controlled conditions is needed to optimize the beneficial effect of OMW and minimize the potential environmental risk, in order to utilize this valuable resource in modern cropping systems. Future research should address the suppressive activity of OMWs against certain plant pathogens in respect to their composition, the olive mill system (two- and three-phase), the processed fruit (maturity, variety) and operating conditions. One step forward would be to elucidate the disease suppressive mechanism and develop feasible delivery and application systems adapted to the crops of interest. The proper use of OMW (in terms of application method, dosage and timing) should be supported by scientific research focusing on the single-field level, considering the soil properties, cultivated species and phytopathological issues existing in individual fields.
4. Materials and Methods
4.1. Determination of mineral nutrients and phenolic content in olive mill wastewater
Olive mill wastewater (OMW) was collected from a three-phase olive mill in the region of Tympaki, Heraklion, Greece, in spring 2019 and kept in an airtight plastic container at room temperature throughout the experiments.
Two samples of about 200 ml from each OMW treatment (sterilized by autoclaving at 121 °C for 30 min and non-sterilized OMW) that were used in the experiment were dried at 105 °C to constant weight, with weight measurements before and after drying used for calculation of the moisture percentage. A subsample of 0.1 g of the dried material was used to colorimetrically determine total N, after wet digestion [
27]. Another subsample of 1 g was dried to ash at 550 °C and then solubilised with 5 ml of 20% HCl and diluted to 25 ml. The solution was then dispersed into 2 vessels, each one used for determination of the following: a) Total concentration of P, K, Ca, Mg, Fe, Zn, Mn and Cu, using a dual view ICP-OES (Optima 8300, Perkin Elmer, USA). b) Total B concentration, after addition of an azomethin-H/EDTA reagent and colorimetric determination at 410 nm, using a visual spectrum photometer (PhotoLab 6100, WTW, Germany). Results of macronutrients were expressed as a percentage (%) of OMW fresh weight whereas micronutrients were expressed as ppm of OMW fresh weight.
Total phenols were recovered by non-sterilized and sterilized OMW samples according to a previous method [
7]. Each sample of sterile or non-sterile OMW (2g) was mixed thoroughly with 40 ml of water in a falcon tube at 220 rpm for 12 hours in a shaking device. The mixture was then centrifuged at 3500 rpm for 10 min and the resulted aqueous phase was filtered directly in a 50 ml-volumetric flask, which was subsequently filled to volume with water. This procedure was repeated four times. Total phenol content of each diluted filtrate was determined twice according to a method described by Markakis et al. [
28]. In brief, an aliquot (1 ml) of the diluted phenol extract, 10 ml of an aqueous solution of Folin–Ciocalteau reagent (10% v/v) and 9 mL of an aqueous solution of sodium carbonate (7.5 % w/v) were successively added to a screw-cup glass vial. The mixture was shaken well and remained in a dark place for 2 hours. The absorbance was then measured at 765 nm, using a U-2900 Hitachi spectrophotometer (Tokyo, Japan). Each sample was measured thrice and total phenolic content expressed in terms of gallic acid equivalent (g of gallic acid/L of OMW).
4.2. In-vitro evaluation of olive mill wastewater against V. dahliae
The mycelial growth of V. dahliae was examined by transferring PDA agar discs of actively-growing mycelium (5 mm in diameter) into the center of new 92-mm-diameter PDA plates (one disc per plate) containing either pure PDA (designated as V. d. + 0.00% OMW) or PDA supplemented with 0.01, 0.05, 0.1, 0.25, 0.50, 1.00 or 5.00 % (vol/vol) sterilized OMW (V. d. + 0.01% OMW, V. d. + 0.05% OMW, V. d. + 0.1% OMW, V. d. + 0.25% OMW, V. d. + 0.5% OMW, V. d. + 1.00% OMW or V. d. + 5.00% OMW, respectively). Plates were incubated at 24 °C in the dark and the colonies’ diameter was measured at 3, 7, 10 and 14 days post inoculation (dpi). The growth rate of V. dahliae in different treatments was expressed in mm/day. At the end of the bioassays (14 dpi) the underside of the plates was scanned using a Samsung Xpress SL-M2875ND Laser Multifunction Printer at 1200 dots per inch and microsclerotial (black-pigmented) area on each plate image was determined manually using the image processing software ImageJ 1.46r (National Institutes of Health, USA). Then, the number of spores was estimated by transferring a 5-mm-diameter disc taken from the periphery of each culture into a 1.5 ml Eppendorf tube with 1 ml of sterilized distilled water, and vortexed vigorously for 30 seconds. The number of spores was measured with the use of a haematocytometer under a light microscope and expressed as the number of spores per 5-mm-diameter disc. Moreover, actively growing mycelia from cultures’ periphery were prepared and microscopic observations (30 readings per culture) were carried out to estimate hyphae width. Three replicated plates per OMW concentration were prepared and the overall experiment repeated twice (six replicated plates per OMW concentration in total).
To evaluate the effect of OMW on spore germination, conidial suspension was prepared as described previously for estimation of spores number, and the concentration was adjusted to 1 spore μl
-1. Eighteen 1-μl-volume drops were deposited onto PDA medium supplemented with OMW at the abovementioned concentrations (18 drops per plate). Plates were incubated at 24 ˚C in the dark for 3 days. Spore germination was assessed visually and estimated as percentage of the emerging colonies. Likewise, to evaluate the effect of OMW on microsclerotium germination, microsclerotial suspension was prepared according to Markakis et al. [
7]. In brief, microsclerotia were prepared by growing
V. dahliae in sucrose sodium nitrate (SSN) liquid cultures in an orbital incubator for 3 weeks. Then, microsclerotia were centrifuged at 3.000 × g for 10 min at room temperature to remove growth medium and air dried into a laminar flow cabinet aseptically. Microsclerotia were resuspended in sterilized distilled water and their concentration was adjusted to 0.5 microslcerotia μl
-1. Eighteen 2-μl-volume drops were deposited onto PDA medium (18 drops per plate) supplemented with OMW at the abovementioned concentrations. Plates were incubated at 24 ˚C in the dark for 2 days. Microsclerotia were examined under a light microscope and microsclerotium germination was estimated as percentage of the germinated microsclerotia. A microsclerotium was considered germinated when the germ tube was ≥ than the microsclerotium diameter. Three replicated plates per OMW concentration were prepared and the overall experiment repeated twice (six replicated plates per OMW concentration in total).
4.3. Plant material
Plant material consisted of eggplant (cv Black Beauty) and tomato (hybrid Belladonna F1) seedlings at the one-true-leaf stage. Both genotypes are susceptible to
V. dahliae [
29]. To prepare seedlings, germinated seeds were sown in vermiculite and when reaching the cotyledon stage seedlings were transplanted in 100 ml-capacity pots containing non-sterilized or heat-sterilized (autoclaved twice at 121°C for one h) soil substrate (HuminSubstrat, Klasmann-3 Deilmann GmbH, Germany).
4.4. Optimal dosage of OMW application
Eggplant and tomato plants grown in 100 ml-capacity pots were root-drenched with 0%, 5%, 10%, 25% or 50% (vol/vol) aqueous OMW solution (20 ml of each OMW solution per plant) and phytotoxicity was determined by assessing symptoms (i.e., leaf chlorosis, yellowing and wilting) and plant growth (plant height and fresh weight) (data not shown). It was revealed that 5% OMW did not cause phytotoxicity and thus, considered to be the optimal dosage for the following experiments.
4.5. OMW, fungal strains and V. dahliae preparation for pathogenicity assays
Non-sterilized and heat sterilized (autoclaved at 121 °C for 30 min) OMW as well as five fungal strains isolated from the disease suppressive soil substrate (designated as KX1, KX2, KX3, KX6 and KX7) and one (KF8) isolated from an asymptomatic
V. dahliae-inoculated eggplant in the experiment I, were evaluated in
in-planta bioassays. The six fungal strains and the highly virulent
V. dahliae isolate 999-1 with proven pathogenicity on eggplant and tomato were prepared according to Markakis et al. [
29]. In brief, conidia were produced by growing each fungus in potato dextrose broth (PDB) at 160 rpm and 24 ˚C in the dark for 5 days. Then, conidia were harvested by filtration through three layers of cheesecloth and the suspensions centrifuged at 3.000 × g for 10 min.
Verticillium dahliae spores were re-suspended in sterilized distilled water and their concentration was adjusted to 5 × 10
6 conidia ml
-1 or 2 × 10
6 conidia ml
-1 for high or low inoculum density, respectively, whereas spores of the rest fungal strains were adjusted to 1 × 10
7 conidia ml
-1.
4.6. In-planta bioassays
Plants at the one-true-leaf stage, grown in 100 ml-capacity pots containing non-sterilized or heat-sterilized soil substrate (designated as ‘Steril. Soil’) were root-drenched with 20 ml of 5% non-sterilized OMW each (designated as ‘OMW’) or with sterilized OMW (designated as ‘Steril. OMW’), or with 20 ml of 1 × 107 conidia ml-1 suspension of each fungal strain individually (designated as ‘KX1’, ‘KX2’, ‘KX3’, ‘KX6’, ‘KX7’ or ‘KF8’) or mixed (‘MIXED’). The commercial biofungicide TRIANUM-P (Koppert B.V. Hellas) was applied at the appropriate dosage according to manufacturer’s instruction (20 ml of 3 × 107 cfu ml-1 per plant) and served as a V. dahliae-suppressive reference treatment in the present study (designated as ‘TRIANUM-P’). Plants that served as negative controls (no OMW/no fungal strain/no TRINANUM-P, designated as ‘Control-’) and those that served as positive controls (no OMW/no fungal strain/no TRIANUM-P/plus V. dahliae, designated as ‘V.d.’), were treated with 20 ml of water. One week later, plants (at the second-true-leaf stage) were inoculated with V. dahliae by drenching the soil substrate in each pot with 20 ml of high (5 × 106 conidia ml-1 suspension) (designated as ‘V.d. high’) or low (2 × 106 conidia ml-1 suspension) (designated as ‘V.d. low’) inoculum density of V. dahliae. Plants that served as negative controls (Control-) were treated with 20 ml of water. Plants were maintained under controlled conditions at 23 ± 2 ˚C with a 12-h light and dark cycle.
To evaluate the suppressive effect of OMW against
V. dahliae in-planta and investigate the potential disease suppressive mechanisms, five independent experiments (experiments Ι, ΙΙ, III, IV and V) were conducted. In experiment Ι, bioassays with eggplant were set up in November 2019 and the following six treatments were applied: Control-, V.d. high,V.d. low, V.d. high+OMW, V.d. low+OMW and V.d. high+TRIANUM-P. In experiment ΙΙ, bioassays with eggplant were set up in April 2020, and fourteen treatments were conducted: Control-, V.d. high, V.d. low, V.d. high+OMW, V.d. high_Steril. soil+OMW, V.d. low+OMW, V.d. high+KX1, V.d. high+KX2, V.d. high+KX3, V.d. high+KX6, V.d. high+KX7, V.d. high+KF8, V.d. high+MIXED and V.d. high+TRIANUM-P. In experiment III, bioassays with tomato were set up in June 2020, and three treatments were included: Control-, V.d., V.d.+OMW), with
V. dahliae to be applied at the high inoculum density only (thereafter designated as V.d.). In experiment IV, bioassays with eggplant were set up in July 2020, and six treatments were applied: Control-, V.d., V.d.+OMW, V.d.+Steril.OMW, V.d./-, V.d./OMW). In this experiment, apart from the typical single-pot set ups (treatments: Control-, V.d., V.d.+OMW and V.d.+Steril.OMW), split-root set ups were also conducted according to Markakis et al. [
30], to investigate direct and/or indirect effect of OMW on
V. dahliae in-planta. In the split-root set ups,
V. dahliae was applied to one side of the split-root system while the other side received water (treatment V.d./-) or the pathogen was applied to one side of the split-root system and the other side received OMW (treatment V.d./OMW). Finally, in experiment V, bioassays with eggplant were set up in September 2020, including three treatments: Control-, V.d. and V.d.+OMW.
Within each experiment, each treatment consisted of seven plants and each experiment was replicated three times (21 plants per treatment and experiment in total).
4.7. Disease assessment
Verticillium wilt symptoms on eggplant were recorded at 2-, 3-, 4- or 7-day intervals from 8 to 35 days post inoculation with
V. dahliae (dpi) in different experiments. Bioassays were evaluated by estimating disease severity, disease incidence, mortality, and relative area under disease progress curve (RAUDPC). Disease parameters were recorded according to Markakis et al. [
7]. In brief, disease severity at each observation was calculated from the number of wilting leaves, as a percentage of total number of leaves of each plant. Disease ratings were plotted over time to generate disease progress curves. Subsequently the area under disease progress curve (AUDPC) was calculated by the trapezoidal integration method [
31]. Disease was expressed as a percentage of the maximum possible area with reference to the maximum value potential reached over the whole period of each experiment and is referred to as relative AUDPC (RAUDPC). Disease incidence was estimated as the percentage of infected plants. Only plants with a final disease severity of ≥20% were considered infected, in order to discriminate between
V. dahliae-associated disease symptoms and other weak symptoms occasionally observed. Mortality was estimated as the percentage of dead plants.
4.8. Plant growth
Growth parameters were evaluated at the end of bioassays (at 30, 32, 35, 32 and 32 dpi for experiments Ι, II, III, IV and V, respectively). To estimate the effect of the above-mentioned treatments on plant growth, all plants were clipped off at the soil surface level and their height, fresh weight and number of leaves were measured.
4.9. Isolation of fungal strains from disease suppressive soil and asymptomatic plants
Isolation of the fungal strains (encoded as KX1, KX2, KX3, KX6 and KX7) from disease suppressive soil (treatment V.d. high+OMW in experiment I) was performed either directly by detaching the emerging fungal fructifications that developed on soil surface and streaking them on APDA, or by macerating 100 g of soil in 300 ml sterilized distilled water, shaking at 100 rpm at 25 °C for 45 min and preparing 10-fold dilution series. In the latter case, 200 μl of each series were plated in APDA and incubated at 25 °C in the dark for 2 days. The plant-originated strain (code KF8) was isolated during pathogen re-isolation performed from an asymptomatic OMW-treated eggplant artificially infested with V. dahliae. Isolated fungal strains were sub-cultured onto new PDA plates to obtain pure fungal colonies and stored at 4°C until use.
4.10. Verticillium dahliae re-isolation
To verify the presence of the applied
V. dahliae strain in plant tissues, the leaves of plants that had been cut above the soil level previously were removed and their stems were surface-disinfected by spraying with 95% ethyl alcohol and passing them quickly through a flame, thrice. For each plant, 3 xylem chips taken from different sites along the stem (base, middle and upper part of the stem) were aseptically placed onto acidified potato dextrose agar (PDA) after the removal of the phloem. Plates were then incubated at 24 ˚C in the dark for 14 days. The emerging fungi that grew out of tissue excisions were examined visually and under a light microscope and identified as
V. dahliae according to their morphological characteristics [
1]. Pathogen isolation ratio was expressed as the frequency of positive
V. dahliae isolations of each plant.
4.11. DNA extraction
To quantify
V.
dahliae DNA in vascular tissues of eggplant (treatments: Control-, V.d. high and V.d. high+OMW in experiment II) and tomato plants (treatments: Control-, V.d. and V.d.+OMW in experiment III), the stems of 21 plants per treatment and experiment were destructively sampled (seven composite samples consisting of 3 pooled stems each) by cutting to 2-3 mm long pieces after the removal of the phloem, and stored at -20 °C. Plant tissues were freeze-dried and ground to a fine powder by using an autoclaved mortar and pestle, in the presence of liquid nitrogen. Total DNA was isolated according to the Cetyltetramethyl ammonium bromide (CTAB) method [
32] with slight modifications. Briefly, 100 mg of plant tissue were homogenized with the use of a mortar and pestle in the presence of liquid nitrogen. Tissue powder was transferred in a 1.5 ml Eppendorf tube and 500 μl of 2X CTAB extraction buffer (100mMTris-HCl, 20mM EDTA, 1.4M NaCl, 2% CTAB, 0.5% v/v β-mecraproethanol) were added and homogenized. The samples were incubated at 65 °C for 45 min with periodical vortexing and centrifuged at 10.000 rpm for 10 min. The supernatant (~250 μl) was transferred to new tubes and equal amounts (~250 μl) of phenol:chloroform:isoamyl alcohol (25:24:1) were added and mixed by vortexing. Samples were centrifuged at 13.000 rpm for 15 min. The aqueous phase (~200 μl) was transferred into a new tube, equal amount of chilled isopropanol was added, followed by quick and gentle inversion, and incubated at -20°C overnight. The DNA pellet was precipitated at 13000 rpm for 20 min, washed with 500 μl of 70% ethanol and precipitated at 13.000 rpm for 5 min. The DNA pellet was then suspended in 40μl of Tris-HCL (10 mM, pH=8). Then, 2 μl RNAse A (5 mg ml
-1) were added followed by incubation at 50°C for 15 min. Purity and quantity of DNA were determined using a Q5000 UV-Vis Spectrophotometer (Quawell, San Jose, CA, USA). Final DNA concentration of each isolate was adjusted to 20 ng ml
-1 and stored at -20 °C until use.
4.12. Verticillium dahliae qPCR quantification
Real-time quantitative PCR (qPCR) assays were conducted for detection and quantification of
V. dahliae DNA in plant tissues. The
V. dahliae ITS region was amplified with the use of primers ITS1-F (5′-CCGCCGGTCCATCAGTCTCTCTGTTTATAC-3′) and ITS2-R (5′-CGCCTGCGGGACTCCGATGCGAGCTGTAAC-3′) [
33]. The
actin gene was used as an internal standard to normalize small differences in total DNA template amounts. The amplification of
actin gene was performed with the primer pair SolACT-F (5′-TTCCGTTGCCCAGAGGTCCT-3′) και SolACT-R (5′- TCGCCCTTTGAAATCCACATC-3′) [
34]. All qPCR assays were carried out in a QuantStudio 3 Real-Time PCR System (ThermoFisher, Waltham, USA) by using the PowerUp™ SYBR® Green Master Mix kit (ThermoFisher, Waltham, USA). The qPCR performance included an initial denaturation at 95 °C for 3 min; followed by 40 cycles of 30 sec of denaturation at 95 °C, 30 sec of annealing at 60 °C, and 30 sec of extension at 72 °C; and a final extension step at 60 °C for 1 min. The relative DNA quantity of
V. dahliae was determined by using the 2
-ΔΔCT method [
35]. The real time qPCR reactions were performed in duplicate, and the absence of nonspecific products and primer dimers was confirmed by the analysis of melting curves. For data analysis, average threshold cycle (Ct) values were calculated for each gene of interest on the basis of seven independent biological samples (seven composite samples consisting of 3 pooled plants each).
4.13. Statistics
Analysis of variance (ANOVA) was employed to determine the effects of replication (1, 2 or 3), treatment (Control-, V.d. high, V.d. low, V.d. high+OMW, V.d. low+OMW and V.d. high+TRIANUM-P in experiment I, Control-, V.d. high, V.d. low, V.d. high+OMW, V.d. high_Steril. soil+OMW, V.d. low+OMW, V.d. high+KX1, V.d. high+KX2, V.d. high+KX3, V.d. high+KX6, V.d. high+KX7, V.d. high+KF8, V.d. high+MIXED, V.d. high+TRIANUM-P in experiment II, Control-, V.d. and V.d.+OMW in experiment III, Control-, V.d., V.d.+OMW, V.d.+Steril.OMW, V.d./-, V.d./OMW in experiment IV, and Control-, V.d. and V.d.+OMW in experiment V), and their interaction on disease incidence (DI), final disease severity (FDS), mortality (M), relative area under disease progress curve (Relative AUDPC), isolation ratio (IR) and
V. dahliae DNA quantity (qPCR), and on leaf number (L), plant height (H) and plant fresh weight (FW) (
Supplementary Table S6). Prior to ANOVA, homogeneity of variance across treatments was evaluated and an arcsine transformation was applied to normalize variance. When a significant
F test was obtained for treatments (
P ≤ 0.05), the data were subjected to means separation by Tukey’s honestly significant difference test. Morphological and physiological characteristics of
V. dahliae in
in-vitro assays were also analysed by Tukey’s test (
P ≤ 0.05). Moreover, standard errors of means were calculated. Association among different fungal parameters (growth rate, sporulation, hyphae width, microslerotial area, spore germination, microsclerotium germination) and OMW content in PDA plates was checked by determining the Pearson’s r correlation coefficients (
P ≤ 0.05).
4.14. Analysis of the microbial community composition of the eggplant and tomato plants
Apart from
V. dahliae qPCR quantification, DNA samples extracted from eggplant (experiment II) and tomato (experiment III) plants, were employed for microbiome analysis. DNA quality was then assessed via electrophoresis on 0.8 % agarose gels, while the DNA concentration was determined via fluorometer measurement with Qubit v.2 Bacterial and fungal community changes in the plants were tracked using multiplex amplicon sequencing, following our internal protocol [
36]. Sequencing was performed by Admera Health (South Plainfield, NJ, USA) on an Illumina NovaSeq platform (San Diego, CA, USA), employing the SP sequencing kit to generate 250 bp paired-end reads. The V4 region of the bacterial 16S rRNA gene was amplified with primers 515f–806r [
37]], and the ITS2 region of fungi was amplified using primers fITS7–ITS4 [
38]. Thermal cycling conditions and primer details are provided in
Supplementary Table S7.
After the acquisition of the NGS sequencing data, a pre-analysis of the sequences was performed by de-multiplexing the samples using the Flexbar program, version 3.0.3 [
39]. Quality control and chimera identification of the sequences was then conducted, followed by error correction in the DNA sequences and assembly of the sequencing inserts from the two reads of each insert to form amplicon sequence variants (ASVs) using the dada2 package [
40] in R software version 4.2.2 (R Core Team, 2020).
Phylogenetic classification was carried out by comparing each ASV with reference sequences from databases. In this case, the Silva SSU v138 database [
41] formatted for dada2 was used for bacterial ASV classification, and the Unite version 9 database [
42] was used for fungal ASVs. Using the phyloseq v1.38.0 package in R, which allows the creation of objects containing a range of information such as the ASV read count table, DNA sequences of the ASVs (via the Biostrings v2.62.0 package), the taxonomic classification table, and the experimental design information for the samples [
43], data management was performed during the subsequent statistical analysis.
In this data analysis, we initially calculted alpha- and beta-diversity indices like Shannon, incerse Simpson, Observed Richeness, Pielou, Low abundance, and the Good’s coverage index for the prokaryotic and fungal communities in tomato and eggplant plants using the phyloseq v1.38.0 package and the entropart v1.6.10 package [
44]. Between samples statistical differences were determined via ANOVA or its non-parametric counterpart, the Kruskal-Wallis test, using the agricolae v1.3.5 package [
45].
Further the effects of the different treatments on the beta-diversity of the plant bacterial and fungal communities was carried out via Canonical Correspondence Analysis (CCA) [
46] or Redundancy Analysis (RDA) [
47], depending on the range of values on the first axis of Detrended Correspondence Analysis (DCA) [
48], and Permutational Analysis of Variance (PERMANOVA) [
49] using the Adonis version 0.0.1 package [
50].
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on Preprints.org. Figure S1: Verticillium wilt disease severity on eggplant mock inoculated (control-) or inoculated with 20 ml of high (5 × 106 conidia ml-1) or low (2 × 106 conidia ml-1) inoculum density of Verticillium dahliae, treated with olive mill wastewater (OMW) or TRIANUM-P or non treated at 12, 14, 16, 19, 23, 26, 28 and 30 days post inoculation (experiment I).; Figure S2: Verticillium wilt disease severity on eggplant mock inoculated (control-) or inoculated with 20 ml of high (5 × 106 conidia ml-1) or low (2 × 106 conidia ml-1) inoculum density of Verticillium dahliae, treated with olive mill wastewater (OMW), separated or mixed microbial strains (KX1, KX2, KX3, KX6, KX7, KF8 or MIXED), TRIANUM-P or non-treated controls at 11, 18, 25, and 32 days post inoculation (experiment II). Plants transplanted in sterilized soil and treated with OMW were also inoculated with high inoculum density (V.d. high_Steril. soil+OMW).; Figure S3: Verticillium wilt disease severity on tomato mock inoculated (control-) or inoculated with 20 ml of high (5 × 106 conidia ml-1) inoculum density of Verticillium dahliae, treated with olive mill wastewater (OMW) or non treated at 11, 14, 18, 21, 25, 28, 32 and 35 days post inoculation (experiment III).; Figure S4: Verticillium wilt disease severity index on eggplant mock inoculated (control-) or inoculated with 20 ml of high (5 × 106 conidia ml-1) inoculum density of Verticillium dahliae, treated with sterilized or non-sterilized olive mill wastewater (OMW) and on eggplant in a split-root set-up with the half of the split-root system being inoculated with the fungus and the other half with water (V.d./-) or the half of the split-root system being inoculated with the fungus and the other half being treated with olive mill wastewater (V.d./OMW), at 8, 11, 14, 18, 21, 25, 28 and 32 days post inoculation (experiment IV). Figure S5: Temporal patterns of the α-diversity indices of the bacterial communities in the tomato (a) and eggplant (b) plants of the study.; Figure S6: Heatmap showing the dominant differentially abundant ASVs of the bacteria community in the tomato plants (a), and the fungal community in the tomato (b) and eggplant (d) plants of the study.; Figure S7: Temporal patterns of the α-diversity indices of the fungal communities in the tomato (a) and eggplant (b) plants of the study.; Table S1: Chemical analysis of non-sterilized and heat-sterilized olive mill wastewater (OMW) used in the study at the beginning (July 2019) and the end (October 2020) of the experiments.; Table S2: The sequence numbers per sample for the bacteria, obtained and remaining from the various quality control steps, and the successful coverage of the sample by the resulting high-quality sequences.; Table S3: Per-sample α-diversity indices for the bacterial microbial communities for both plants of the study.; Table S4: The sequence numbers per sample for the fungi, obtained and remaining from the various quality control steps, and the successful coverage of the sample by the resulting high-quality sequences.; Table S5: Per-sample α-diversity indices for the fungal microbial communities for both plants of the study.; Table S6: Analysis of variance (ANOVA) for disease and plant growth parameters on eggplant and tomato plants artificially inoculated with Verticillium dahliae and treated with various applications, in five different experiments (Ι, II, III, IV and V).; Table S7: Primers, sequences and thermocycling conditions used to amplify the 16S rRNA and ITS2 genes for bacteria and fungi, respectively.
Author Contributions
Conceptualization, E.A.M; methodology, E.A.M. and D.G.K.; validation, S.K.S., A.C., P.A.K. and A.A.P.; formal analysis, S.K.S., P.A.K., E.A.M., D.G.K., G.P. and G.S.K..; investigation, S.K.S., A.C., P.A.K., A.A.P., G.S.K. and G.P.; resources, E.A.M.; data curation, E.A.M., S.K.S., P.A.K. and D.G.K.; writing—original draft preparation, E.A.M.; writing—review and editing, E.A.M., S.K.S., P.A.K., D.G.K., G.P. and D.E.G.; visualization, E.A.M. and D.G.K.; supervision, E.A.M. and D.G.K.; project administration, E.A.M. All authors have read and agreed to the published version of the manuscript.
Figure 1.
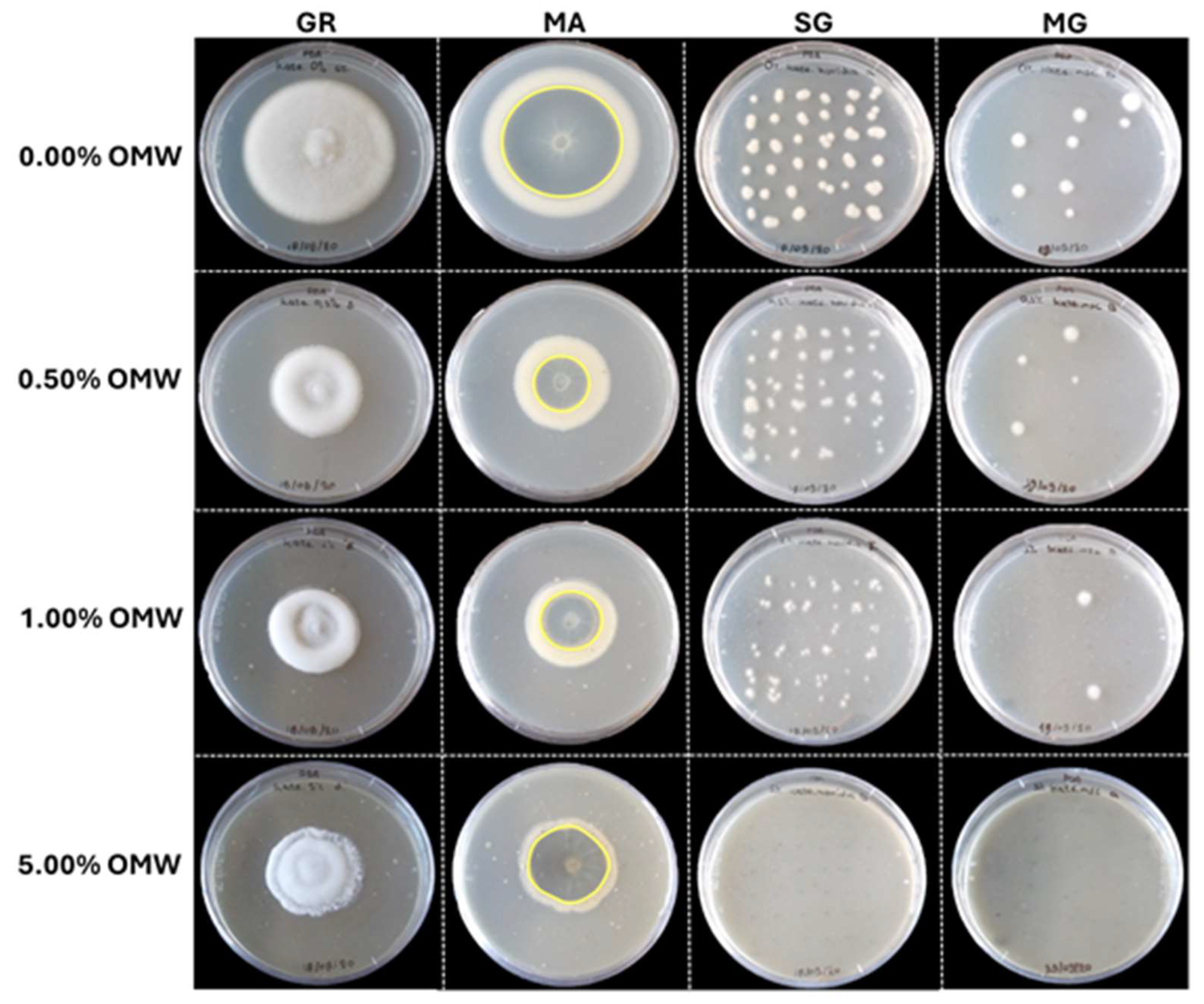
Effect of sterilized olive mill wastewater (OMW) added at various concentrations (0.00%, 0.50%, 1.00% and 5.00%) in PDA, on fungal parameters (GR: growth rate; MA: microsclerotial area; SG: spore germination; MG: microsclerotium germination) of Verticillium dahliae. Yellow lines on the underside of PDA cultures indicates the border of the microsclerotial area among the different treatments in-vitro.
Figure 1.
Effect of sterilized olive mill wastewater (OMW) added at various concentrations (0.00%, 0.50%, 1.00% and 5.00%) in PDA, on fungal parameters (GR: growth rate; MA: microsclerotial area; SG: spore germination; MG: microsclerotium germination) of Verticillium dahliae. Yellow lines on the underside of PDA cultures indicates the border of the microsclerotial area among the different treatments in-vitro.
Figure 2.
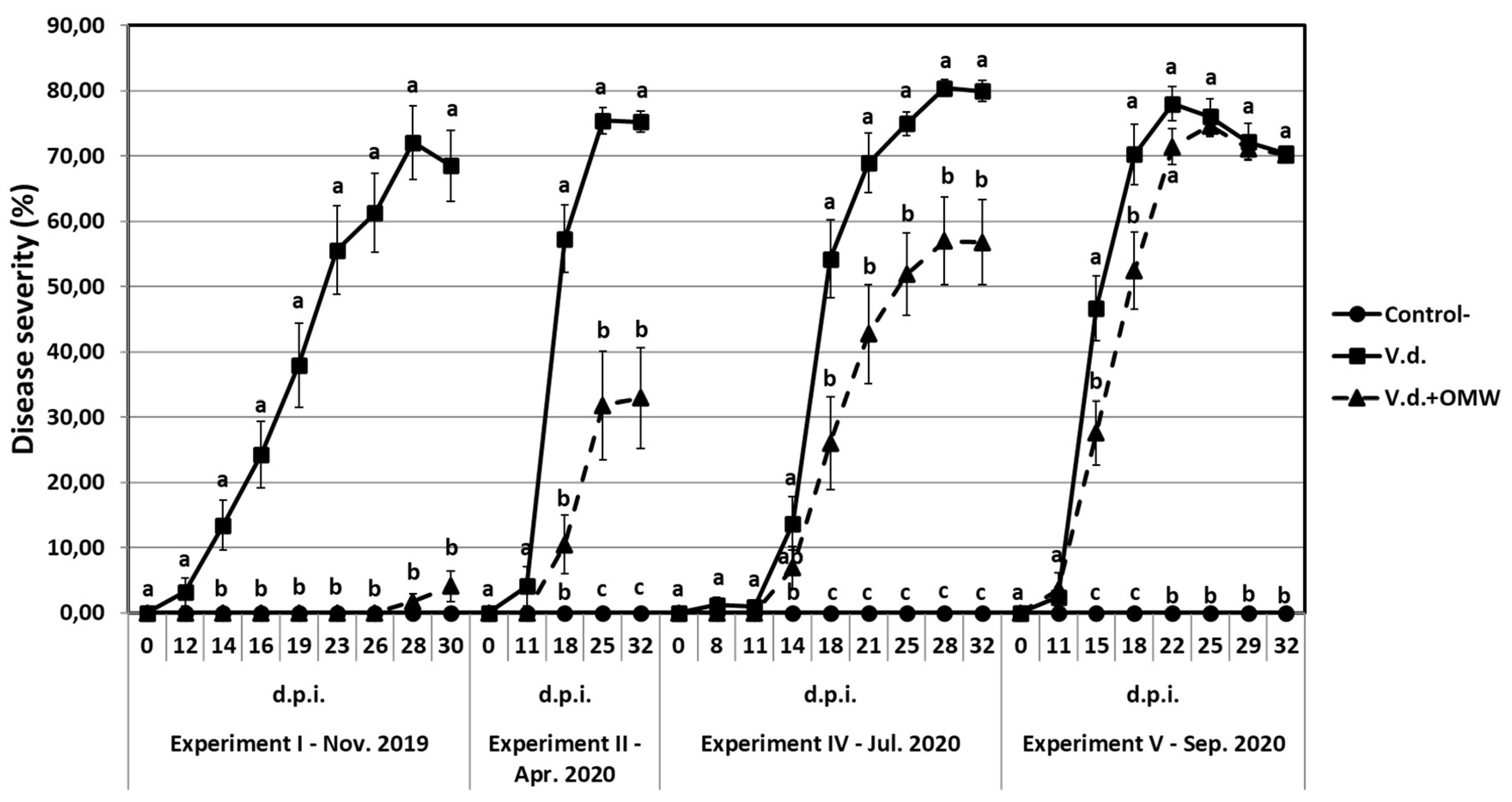
Verticillium wilt disease severity on eggplant mock inoculated (control-) or inoculated with 20 ml of high (5 × 106 conidia ml-1) inoculum density of Verticillium dahliae, treated with olive mill wastewater (OMW) or non treated, in different experiments (I, II, IV and V) that conducted at different periods under the same controlled conditions. Each marker represents the mean of 21 plants. Within each experiment, markers followed by the same letter at each observation time point are not significantly different according to Tukey’s HSD test at P ≤ 0.05. Vertical bars indicate standard errors.
Figure 2.
Verticillium wilt disease severity on eggplant mock inoculated (control-) or inoculated with 20 ml of high (5 × 106 conidia ml-1) inoculum density of Verticillium dahliae, treated with olive mill wastewater (OMW) or non treated, in different experiments (I, II, IV and V) that conducted at different periods under the same controlled conditions. Each marker represents the mean of 21 plants. Within each experiment, markers followed by the same letter at each observation time point are not significantly different according to Tukey’s HSD test at P ≤ 0.05. Vertical bars indicate standard errors.
Figure 3.
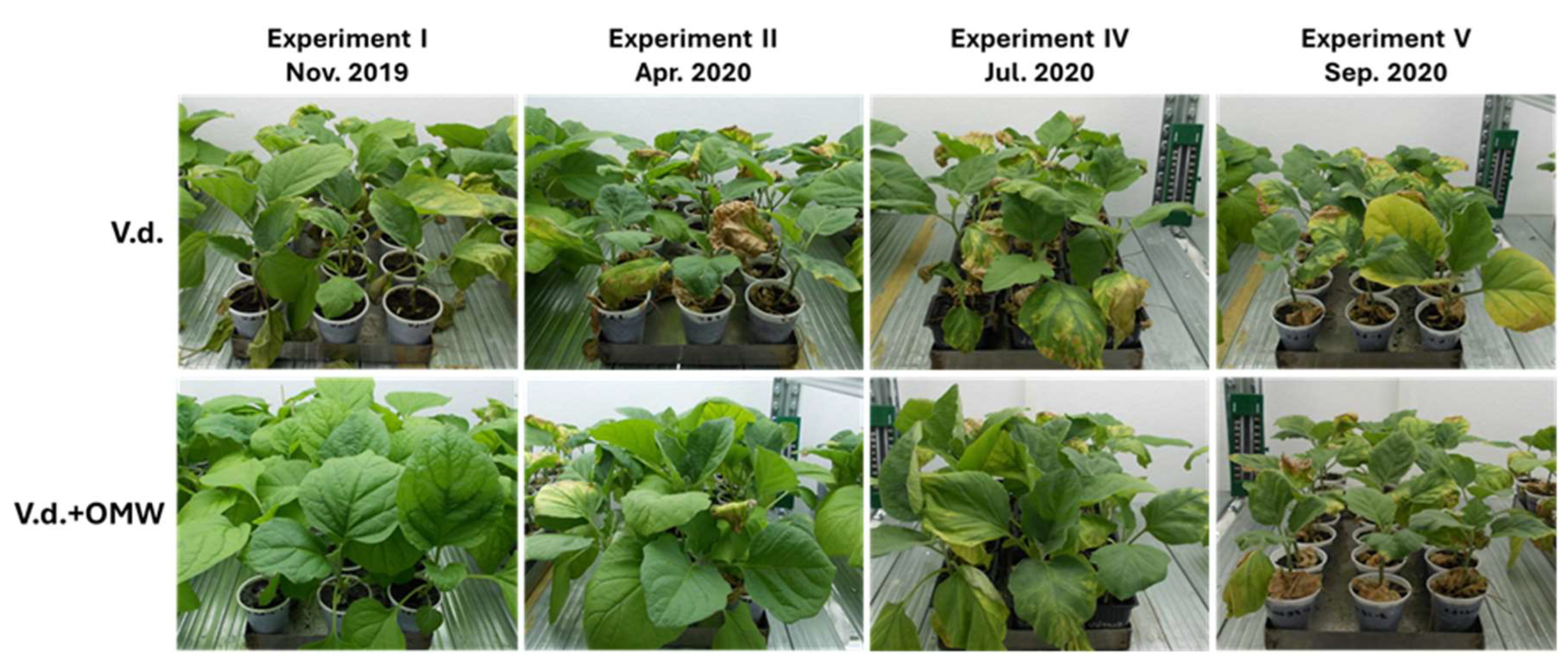
Disease symptoms on eggplants inoculated with Verticillium dahliae, previously treated with olive mill wastewater (V.d.+OMW) or not (V.d.), in different experiments (I, II, IV and V) that conducted at different time periods (November 2019, April 2020, July 2020 and September 2020) under the same conditions.
Figure 3.
Disease symptoms on eggplants inoculated with Verticillium dahliae, previously treated with olive mill wastewater (V.d.+OMW) or not (V.d.), in different experiments (I, II, IV and V) that conducted at different time periods (November 2019, April 2020, July 2020 and September 2020) under the same conditions.
Figure 4.
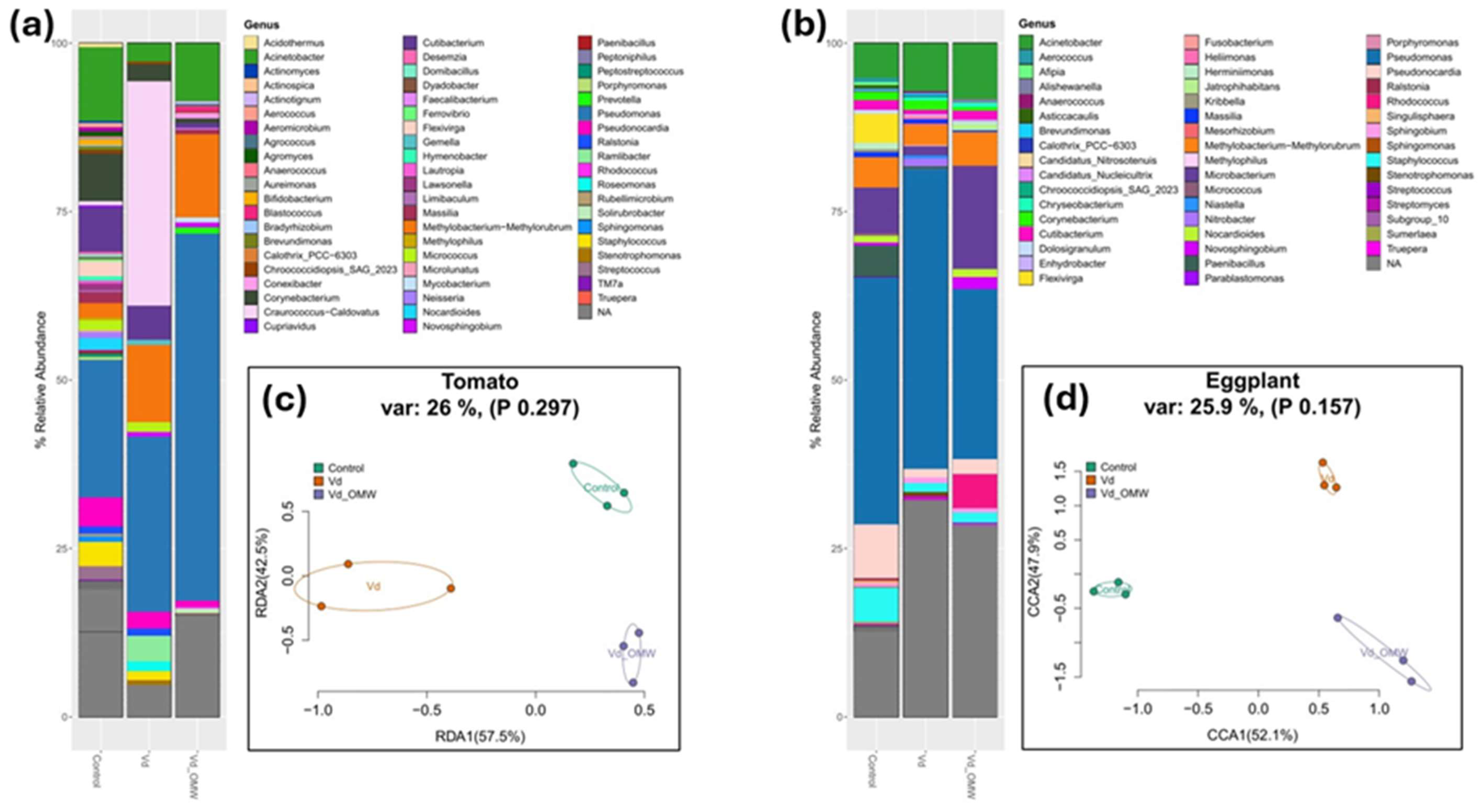
Stacked bar plots showing the relative abundance of the most dominant members of the bacterial community phylogenetically assigned to the genus levels in the tomato (a) and eggplant (b) plants, along with Redundancy analysis (RDA) and Canonical correspondence analysis (CCA) ordination plots of the bacterial community in tomato (c) and eggplant (d) plants, respectively. The relative abundance was calculated with the 200 most dominant ASVs in the dataset. Ellipses are denoting the grouping of the samples according to the treatment. Model parameters, variance and significance levels are provided in the panel above the graphs.
Figure 4.
Stacked bar plots showing the relative abundance of the most dominant members of the bacterial community phylogenetically assigned to the genus levels in the tomato (a) and eggplant (b) plants, along with Redundancy analysis (RDA) and Canonical correspondence analysis (CCA) ordination plots of the bacterial community in tomato (c) and eggplant (d) plants, respectively. The relative abundance was calculated with the 200 most dominant ASVs in the dataset. Ellipses are denoting the grouping of the samples according to the treatment. Model parameters, variance and significance levels are provided in the panel above the graphs.
Figure 5.
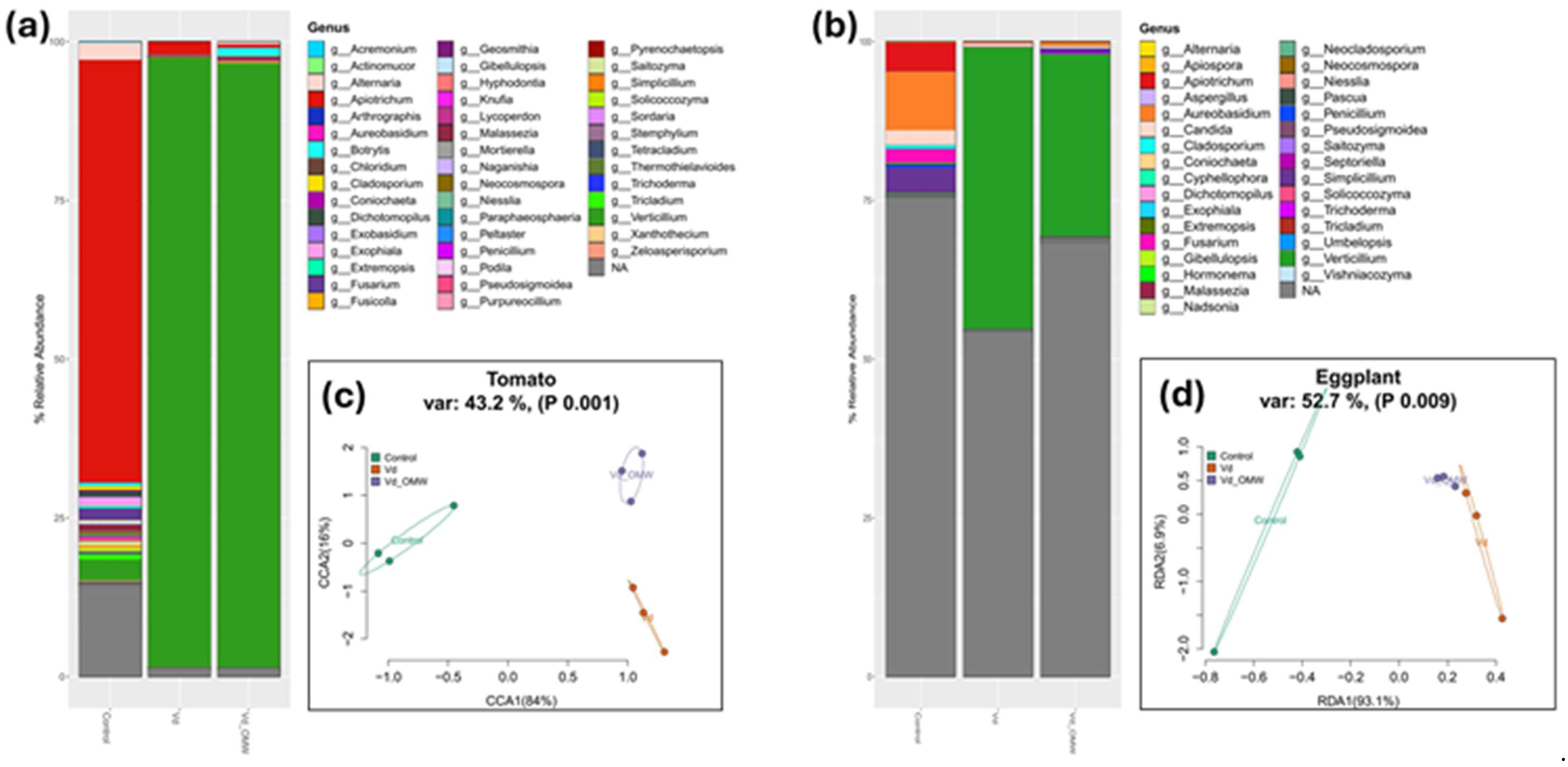
Stacked bar plots showing the relative abundance of the most dominant members of the fungal community phylogenetically assigned to the genus levels in the tomato (a) and eggplant (b) plants, along with Redundancy analysis (RDA) and Canonical correspondence analysis (CCA) ordination plots of the fungal community in tomato (c) and eggplant (d) plants, respectively. The relative abundance was calculated with the 200 most dominant ASVs in the dataset. Ellipses are denoting the grouping of the samples according to the treatment. Model parameters, variance and significance levels are provided in the panel above the graphs.
Figure 5.
Stacked bar plots showing the relative abundance of the most dominant members of the fungal community phylogenetically assigned to the genus levels in the tomato (a) and eggplant (b) plants, along with Redundancy analysis (RDA) and Canonical correspondence analysis (CCA) ordination plots of the fungal community in tomato (c) and eggplant (d) plants, respectively. The relative abundance was calculated with the 200 most dominant ASVs in the dataset. Ellipses are denoting the grouping of the samples according to the treatment. Model parameters, variance and significance levels are provided in the panel above the graphs.
Table 1.
Values (±standard errors) of fungal parameters of Verticillium dahliae grown on PDA amended with sterilized olive mill wastewater (OMW) at various concentrations (0.00%,0.001%, 0.05%, 0.10%, 0.25%, 0.50%, 1.00% and 5.00%).
Table 1.
Values (±standard errors) of fungal parameters of Verticillium dahliae grown on PDA amended with sterilized olive mill wastewater (OMW) at various concentrations (0.00%,0.001%, 0.05%, 0.10%, 0.25%, 0.50%, 1.00% and 5.00%).
| Treatment |
Fungal parametersa
|
| |
Growth rate (mm/day)b
|
Spores (×105) per 5-mm-diameter discc
|
Hyphae width (μm)d
|
Microsclerotial area (cm2)e
|
Spore germination (%)f
|
Microsclerotium germination (%)g
|
| V. d. + 0.00% OMW |
3.84±0.18 a |
27.92±7.87 ab |
3.35±0.13 a |
12.72±2.09 a |
63.66±4.28 a |
37.03±4.97 a |
| V. d. + 0.01% OMW |
3.46±0.22 ab |
21.08±2.16 abcd |
2.58±0.13 b |
8.39±2.31 a |
59.03±2.70 ab |
26.88±5.92 ab |
| V. d. + 0.05% OMW |
3.24±0.23 abc |
38.00±5.18 a |
2.17±0.11 bc |
7.67±2.31 a |
56.01±4.22 abc |
23.14±5.37 ab |
| V. d. + 0.10% OMW |
3.11±0.21 abc |
25.42±4.97 abc |
1.78±0.08 cd |
4.93±1.70 a |
50.23±2.20 abc |
28.19±3.60 ab |
| V. d. + 0.25% OMW |
2.95±0.24 bc |
11.83±2.07 bcd |
2.02±0.09 cd |
5.59±1.98 a |
54.40±3.80 abc |
24.56±3.96 ab |
| V. d. + 0.50% OMW |
2.66±0.10 bc |
10.25±1.25 cd |
1.86±0.08 cd |
5.14±1.28 a |
47.92±2.50 bc |
20.52±4.47 abc |
| V. d. + 1.00% OMW |
2.61±0.12 c |
8.92±0.97 cd |
1.72±0.08 d |
5.09±1.28 a |
43.98±2.82 c |
10.94±5.11 bc |
| V. d. + 5.00% OMW |
2.45±0.08 c |
6.17±0.95 d |
2.07±0.11 cd |
7.93±0.38 a |
0.00±0.00 d |
5.89±2.82 c |
Table 2.
Values (±standard errors) of disease and growth parameters for eggplants mock inoculated (control-) or artificially inoculated with high or low inoculum density of Verticillium dahliae and treated with olive mill wastewater (OMW) or TRIANUM-P or non treated, in experiment Ι.
Table 2.
Values (±standard errors) of disease and growth parameters for eggplants mock inoculated (control-) or artificially inoculated with high or low inoculum density of Verticillium dahliae and treated with olive mill wastewater (OMW) or TRIANUM-P or non treated, in experiment Ι.
| Treatment |
Disease parametersa
|
Plant growth parametersb
|
| |
DI (%) |
FDS (%) |
M (%) |
RAUDPC (%) |
IR (0-1) |
L |
H (cm) |
FW (gr) |
| Control- |
0.00±0.00 c |
0.00±0.00 c |
0.00±0.00 b |
0.00±0.00 c |
0.00±0.00 c |
6.76±0.14 a |
18.31±0.34 a |
6.56±0.17 a |
| V.d. high |
90.48±6.15 a |
68.49±5.43 a |
47.62±9.91 a |
26.76±2.95 a |
0.59±0.08 a |
6.24±0.14 ab |
17.00±0.39 ab |
2.89±0.31 b |
| V.d. low |
52.38±6.14 b |
34.27±7.91 b |
9.52±6.15 b |
6.80±2.01 bc |
0.29±0.14 b |
6.52±0.13 ab |
18.29±0.39 a |
6.19±0.49 a |
| V.d. high+OMW |
14.29±14.29 c |
4.06±2.36 c |
0.00±0.00 b |
0.25±0.15 c |
0.10±0.06 bc |
6.25±0.14 ab |
16.75±0.34 ab |
6.47±0.37 a |
| V.d. low+OMW |
4.76±4.76 c |
1.90±1.90 c |
0.00±0.00 b |
0.66±0.66 c |
0.13±0.07 bc |
6.05±0.13 b |
17.71±0.48 ab |
5.96±0.37 a |
| V.d. high+TRIANUM-P |
52.38±6.74 b |
33.45±7.44 b |
4.76±4.76 b |
7.88±1.94 b |
0.25±0.16 bc |
6.48±0.11 ab |
16.14±0.56 b |
5.87±0.42 a |
Table 3.
Values (±standard errors) of disease and growth parameters for eggplants mock inoculated (control-) or artificially inoculated with high or low inoculum density of Verticillium dahliae and treated with olive mill wastewater (OMW) in non-sterilized or sterilized soil, various microbial strains (KX1, KX2, KX3, KX6, KX7 or KF8), TRIANUM-P or non-treated (control-), in experiment ΙI.
Table 3.
Values (±standard errors) of disease and growth parameters for eggplants mock inoculated (control-) or artificially inoculated with high or low inoculum density of Verticillium dahliae and treated with olive mill wastewater (OMW) in non-sterilized or sterilized soil, various microbial strains (KX1, KX2, KX3, KX6, KX7 or KF8), TRIANUM-P or non-treated (control-), in experiment ΙI.
| Treatment |
Disease parametersa
|
Plant growth parametersb
|
| |
DI (%) |
FDS (%) |
M (%) |
RAUDPC (%) |
IR (0-1) |
qPCR |
L |
H (cm) |
FW (gr) |
| Control- |
0.00±0.00 e |
0.00±0.00 c |
0.00±0.00 b |
0.00±0.00 e |
0.00±0.00 d |
0.00±0.00 b |
7.10±0.41 a |
15.22±0.84 ab |
7.73±0.47 a |
| V.d. high |
100.00±0.00 a |
75.22±1.57 a |
33.33±10.29 ab |
38.40±1.48 a |
0.98±0.02 a |
3.90±1.10 a |
6.95±0.26 a |
11.57±0.37 c |
2.53±0.16 c |
| V.d. low |
85.72±6.73 ab |
66.81±5.06 a |
23.81±14.02 ab |
29.63±3.28 ab |
0.84±0.06 a |
ne |
7.05±0.29 a |
13.21±0.48 bc |
4.24±0.55 bc |
| V.d. high+OMW |
42.86±14.02 c |
32.91±7.73 b |
9.52±6.15 ab |
12.84±3.37 cd |
0.37±0.09 c |
1.12±0.35 b |
6.90±0.15 a |
15.12±0.62 ab |
6.26±0.62 ab |
| V.d. high_Steril. soil+OMW |
57.14±15.79 bc |
35.96±7.20 b |
0.00±0.00 b |
10.23±2.74 de |
0.49±0.10 bc |
ne |
7.10±0.23 a |
16.74±0.65 a |
6.27±0.53 ab |
| V.d. low+OMW |
42.86±11.98 c |
28.34±7.55 b |
14.28±6.73 ab |
8.20±2.78 de |
0.30±0.09 cd |
ne |
6.90±0.15 a |
15.14±0.57 ab |
6.12±0.61 ab |
| V.d. high+KX1 |
100.00±0.00 a |
72.80±1.16 a |
19.05±9.91 ab |
30.81±1.92 ab |
ne |
ne |
6.62±0.15 a |
12.21±0.41 c |
2.78±0.28 c |
| V.d. high+KX2 |
90.48±6.15 ab |
69.54±5.30 a |
52.38±12.30 a |
30.29±3.09 ab |
ne |
ne |
6.62±0.33 a |
13.38±0.31 bc |
3.00±0.45 c |
| V.d. high+KX3 |
100.00±0.00 a |
70.68±2.40 a |
23.81±9.52 ab |
31.78±2.17 ab |
ne |
ne |
7.10±0.28 a |
13.00±0.43 bc |
3.29±0.25 c |
| V.d. high+KX6 |
100.00±0.00 a |
76.15±1.35 a |
42.86±9.52 ab |
33.15±1.57 ab |
ne |
ne |
6.48±0.13 a |
12.81±0.41 bc |
2.52±0.14 c |
| V.d. high+KX7 |
100.00±0.00 a |
75.28±2.32 a |
42.86±9.52 ab |
32.19±2.08 ab |
ne |
ne |
6.52±0.11 a |
12.17±0.27 bcd |
2.80±0.27 c |
| V.d. high+KF8 |
95.24±4.76 ab |
70.16±3.37 a |
33.33±10.29 ab |
26.55±2.60 ab |
ne |
ne |
6.71±0.18 a |
14.05±0.59 bcd |
3.74±0.45 c |
| V.d. high+MIXED |
85.71±9.91 ab |
63.32±6.26 a |
23.76±9.52 ab |
28.28±3.53 ab |
ne |
ne |
6.62±0.41 a |
12.26±0.54 c |
3.48±0.51 c |
| V.d. high+TRIANUM-P |
80.95±9.91 abc |
61.58±7.00 a |
37.95±13.48 ab |
22.91±3.37 bc |
0.76±0.08 ab |
0.56±0.09 b |
6.57±0.44 a |
12.67±0.69 bc |
3.76±0.59 c |
Table 4.
Values (±standard errors) of disease and growth parameters for tomato plants mock inoculated (control-) or artificially inoculated with high inoculum density of Verticillium dahliae and treated with olive mill wastewater (OMW) or non-treated, in experiment ΙII.
Table 4.
Values (±standard errors) of disease and growth parameters for tomato plants mock inoculated (control-) or artificially inoculated with high inoculum density of Verticillium dahliae and treated with olive mill wastewater (OMW) or non-treated, in experiment ΙII.
| Treatment |
Disease parametersa
|
Plant growth parametersb
|
| |
DI (%) |
FDS (%) |
M (%) |
RAUDPC (%) |
IR (0-1) |
qPCR |
L |
H (cm) |
FW (gr) |
| Control- |
0.00±0.00 c |
0.00±0.00 c |
0.00±0.00 a |
0.00±0.00 c |
0.00±0.00 c |
0.002±0.001 c |
7.71±0.10 a |
38.39±0.78 a |
11.26±0.34 a |
| V.d. |
95.27±4.76 a |
42.76±1.89 a |
0.00±0.00 a |
25.14±1.43 a |
0.62±0.07 a |
0.182±0.026 a |
7.67±0.13 a |
34.74±0.83 b |
8.98±0.23 c |
| V.d.+OMW |
52.38±9.91 b |
22.36±4.87 b |
0.00±0.00 a |
11.54±2.62 b |
0.38±0.09 b |
0.097±0.021 b |
7.50±0.11 a |
36.13±0.75 ab |
10.21±0.37 b |
Table 5.
Values (±standard errors) of disease and growth parameters for eggplants mock inoculated (control-) or artificially inoculated with high inoculum density of Verticillium dahliae and treated with sterilized or non-sterilized olive mill wastewater (OMW) or non-treated, and for eggplants in a split-root set-up with the half of the split-root system being inoculated with the fungus and the other half with water (V.d./-) or the half of the split-root system being inoculated with the fungus and the other half being treated with olive mill wastewater (V.d./OMW), in experiment ΙV.
Table 5.
Values (±standard errors) of disease and growth parameters for eggplants mock inoculated (control-) or artificially inoculated with high inoculum density of Verticillium dahliae and treated with sterilized or non-sterilized olive mill wastewater (OMW) or non-treated, and for eggplants in a split-root set-up with the half of the split-root system being inoculated with the fungus and the other half with water (V.d./-) or the half of the split-root system being inoculated with the fungus and the other half being treated with olive mill wastewater (V.d./OMW), in experiment ΙV.
| Treatment |
Disease parametersa
|
Plant growth parametersb
|
| |
DI (%) |
FDS (%) |
M (%) |
RAUDPC (%) |
IR (0-1) |
L |
H (cm) |
FW (gr) |
| Control- |
0.00±0.00 c |
0.00±0.00 c |
0.00±0.00 b |
0.00±0.00 c |
0.00±0.00 b |
7.57±0.22 ab |
17.79±0.72 a |
10.67±0.54 a |
| V.d. |
100.00±0.00 a |
79.91±1.64 a |
57.14±14.02 a |
37.24±1.39 a |
0.78±0.06 a |
6.62±0.22 b |
13.67±0.52 b |
2.93±0.18 c |
| V.d.+OMW |
80.95±6.73 ab |
56.78±6.51 b |
23.81±6.15 ab |
23.72±3.03 b |
0.48±0.11 ab |
6.95±0.30 ab |
16.31±0.68 a |
5.01±0.71 bc |
| V.d.+Steril.OMW |
76.19±9.52 b |
53.32±7.08 b |
14.28±6.73 b |
21.77±3.12 b |
0.63±0.09 a |
7.29±0.27 ab |
15.83±0.65 ab |
5.50±0.75 bc |
| V.d./- |
85.72±6.73 ab |
58.54±6.50 ab |
14.28±6.73 b |
28.15±4.05 ab |
0.70±0.10 a |
7.57±0.21 ab |
18.05±0.60 a |
8.48±1.43 ab |
| V.d./OMW |
85.72±6.73 ab |
62.13±5.83 ab |
14.29±9.91 b |
28.71±3.11 ab |
0.74±0.09 a |
7.71±0.23 a |
16.86±0.70 a |
7.60±1.25 ab |
Table 6.
Values (±standard errors) of disease and growth parameters for eggplants mock inoculated (control-) or artificially inoculated with high inoculum density of Verticillium dahliae and treated with olive mill wastewater (OMW) or non-treated, in experiment V.
Table 6.
Values (±standard errors) of disease and growth parameters for eggplants mock inoculated (control-) or artificially inoculated with high inoculum density of Verticillium dahliae and treated with olive mill wastewater (OMW) or non-treated, in experiment V.
| Treatment |
Disease parametersa
|
Plant growth parametersb
|
| |
DI (%) |
FDS (%) |
M (%) |
RAUDPC (%) |
IR (0-1) |
L |
H (cm) |
FW (gr) |
| Control- |
0.00±0.00 b |
0.00±0.00 b |
0.00±0.00 a |
0.00±0.00 b |
0.00±0.00 b |
7.29±0.22 a |
15.65±0.40 a |
7.87±0.32 a |
| V.d. |
100.00±0.00 a |
70.35±1.97 a |
28.57±11.34 a |
41.37±1.71 a |
0.92±0.04 a |
6.19±0.15 b |
11.86±0.36 b |
2.46±0.26 b |
| V.d.+OMW |
100.00±0.00 a |
70.16±1.41 a |
19.05±9.91 a |
36.62±1.49 b |
0.86±0.06 a |
6.19±0.13 b |
11.79±0.27 b |
2.26±0.11 b |