1. Introduction
Filamin С (FLNC, γ-filamin) is an actin-binding protein, a member of the filamins family, localized at Z-discs of striated muscles sarcomeres. In normal dimerized state it interacts with numerous FLNC-binding proteins, including those responsible for normal sarcomere structure organization, signaling cascades members and cell junctions proteins [
1,
2].
Similarly to other filamins, FLNC is a structural protein that has actin-binding domain (ABD) made up of two calponin homology domains (CH), followed by 24 immunoglobulin-like (Ig-like) domains divided into ROD1 and ROD2 regions, and C-terminal dimerization domain [
3]. ROD2 domain contains the majority of binding sites for FLNC binding partners and is required for mechanosensing and muscle function support [
4].
A list of mutations in the
Flnc gene have been shown to be associated with pathology of various degree of severity [
5,
6,
7,
8]. For example, heterozygous truncations of
Flnc cause dilated cardiomyopathy and arrhythmia [
4]. Aberrant protein folding and dimerization failure leads to aggregate formation, negatively affecting muscle function, which is characteristic of hypertrophic myopathy and myofibrillar myopathy. ROD2 domain mutations are often associated with hypertrophic cardiomyopathy [
4].
Myopathy is a chronic progressive neuromuscular disorder characterized by primary muscle lesions. An example of myopathies is cardiomyopathy (CM) — any structural or functional pathologies of cardiac muscle not related to a particular condition, such as ischemic heart disease, congenital heart defect or valve anomaly [
9]. The major types of cardiomyopathies are dilation, hypertrophic and restrictive, all characterized by different pathological changes. The most common CM type is hypertrophic cardiomyopathy, it is diagnosed in 1 in 500 adults and is one of primary causes of death [
10].
Previously published work performed on mouse models has reported that deletion of exons 41–48 in
Flnc gene causes neonatal lethality in homozygotes due to breathing disorder, while heterozygous animals had lower muscle weight and reduced count of primary muscle fibers [
11]. Other studies have shown that deletion of 16 C-terminal amino acid residues in FLNC protein prevents normal dimerization process, leading to muscle weakness and myofibers instability in heterozygous mice [
12,
13].
In this work we have generated two genetically edited mouse strains that carry distinct mutations in the ROD2 domain-coding region of the Flnc gene. We have performed morphological and microscopic analysis of knockout mouse embryos, phenotype assessment of adult animals with filamin C mutations, including histological analysis of cardiac muscle and comparison of muscle performance. We have also conducted bioinformatics analysis of possible structural changes in mutant protein forms. Interestingly, compound heterozygous animals (who carry different mutant alleles of Flnc gene) successfully develop and do not exhibit any cardiomyopathy signs, neither on histology level, nor in ECG (electrocardiography) and functional diagnostic assays.
2. Results
2.1. Two Mutant Mouse Strains Were Generated.
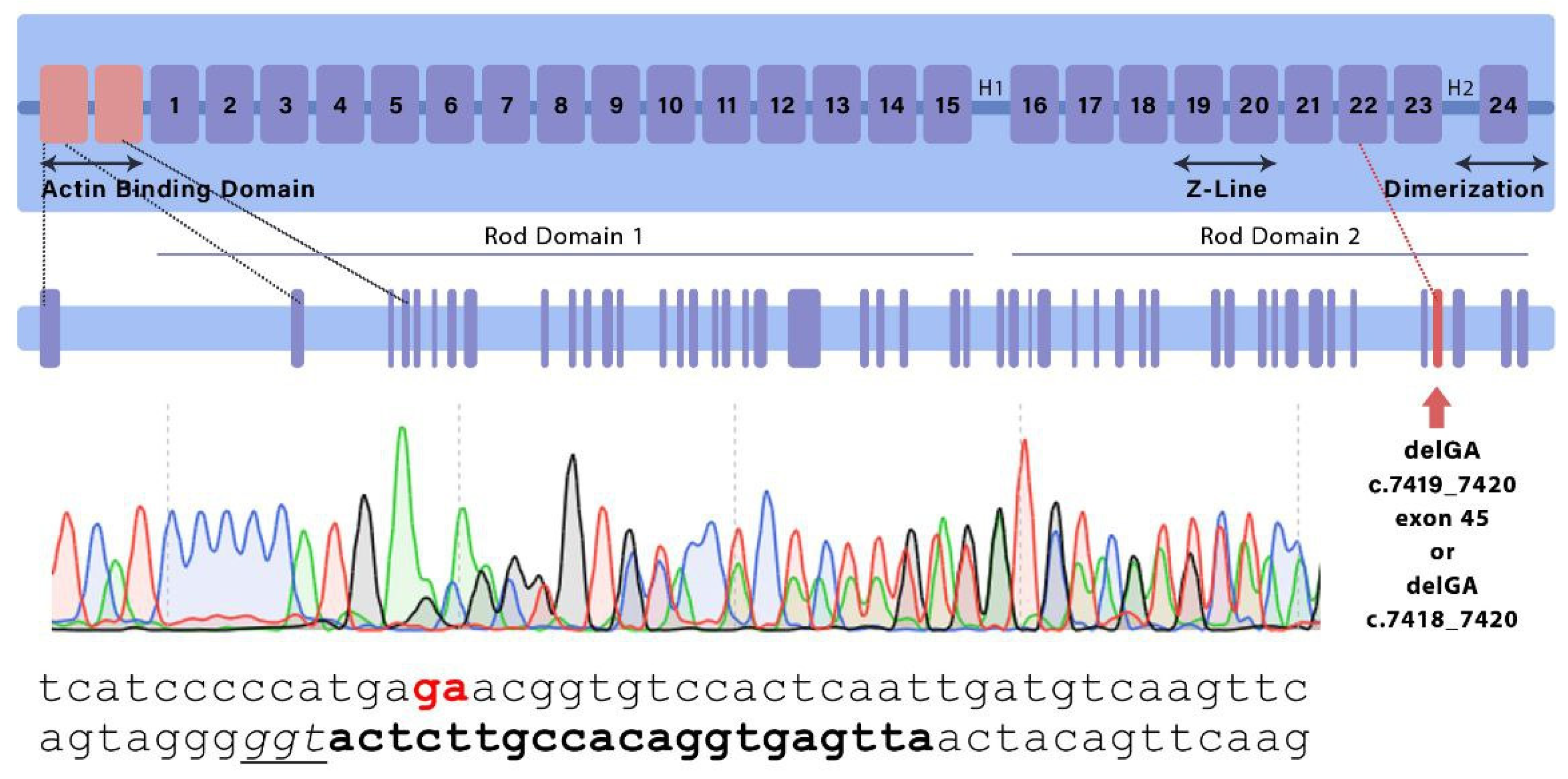
From 460 transplanted embryos, injected with the gene editing system, 36 F0 generation pups were obtained, of which 2 had desired genetic modifications. Two animal strains were used in subsequent experiments — one of them carried a deletion of three nucleotides in exon 45 at position NM_001081185.2:c.7418_7420, and, thus, an amino acid substitution p.E2472D and Asn2473 deletion, and in another one GA deletion at NM_001081185.2:c.7419_7420 led to a frameshift mutation and a premature stop codon. The mutant alleles were designated Flnc
AGA and Flnc
GA, respectively. Schematic representation of mutations position and Sanger sequencing results of F1 animals are shown on
Figure 1.
To assess the viability of hetero- and homozygous carriers of novel mutations, monohybrid crossings of heterozygous animals of both mutant strains were set up: Flnc
wt/GA x Flnc
wt/GA, Flnc
wt/AGA x Flnc
wt/AGA and Flnc
wt/AGA x Flnc
wt/GA. As a result, a total of 79 Flnc
wt/wt, 98 Flnc
wt/GA, 42 Flnc
wt/AGA and 18 Flnc
AGA/GA animals were born (
Table 1a, b) and genotyped in F1–F4 generations. No homozygous animals were born, suggesting homozygosity of either allele is embryonically lethal. Surprisingly, compound heterozygous (Flnc
AGA/GA) mice were viable and distribution of genotypes in offspring of Flnc
wt/AGA x Flnc
wt/GA crossing agreed with expected (
Table 1c).
2.2. Homozygous FlncGA/GA Embryos Do Not Survive Beyond E11.5 –E12.5
To determine the developmental stage when homozygous embryos die, dated pregnancies from Flnc
wt/GA x Flnc
wt/GA crossing were set up. As according to Mendelian segregation law 25% of offspring should be homozygotes, genotyping of embryos was performed to determine which
Flnc alleles they carry. For each embryo, a genotype and a microphotograph captured before tissue sample removal were matched. A comparison of phenotypes of embryos with Flnc
GA/GA, Flnc
wt/GA and Flnc
wt/wt genotypes was performed. Morphology features, characteristic of each developmental stage, such as presence and development of various brain regions, eyes, limb buds and other was assessed and compared to atlases of normal development [
14].
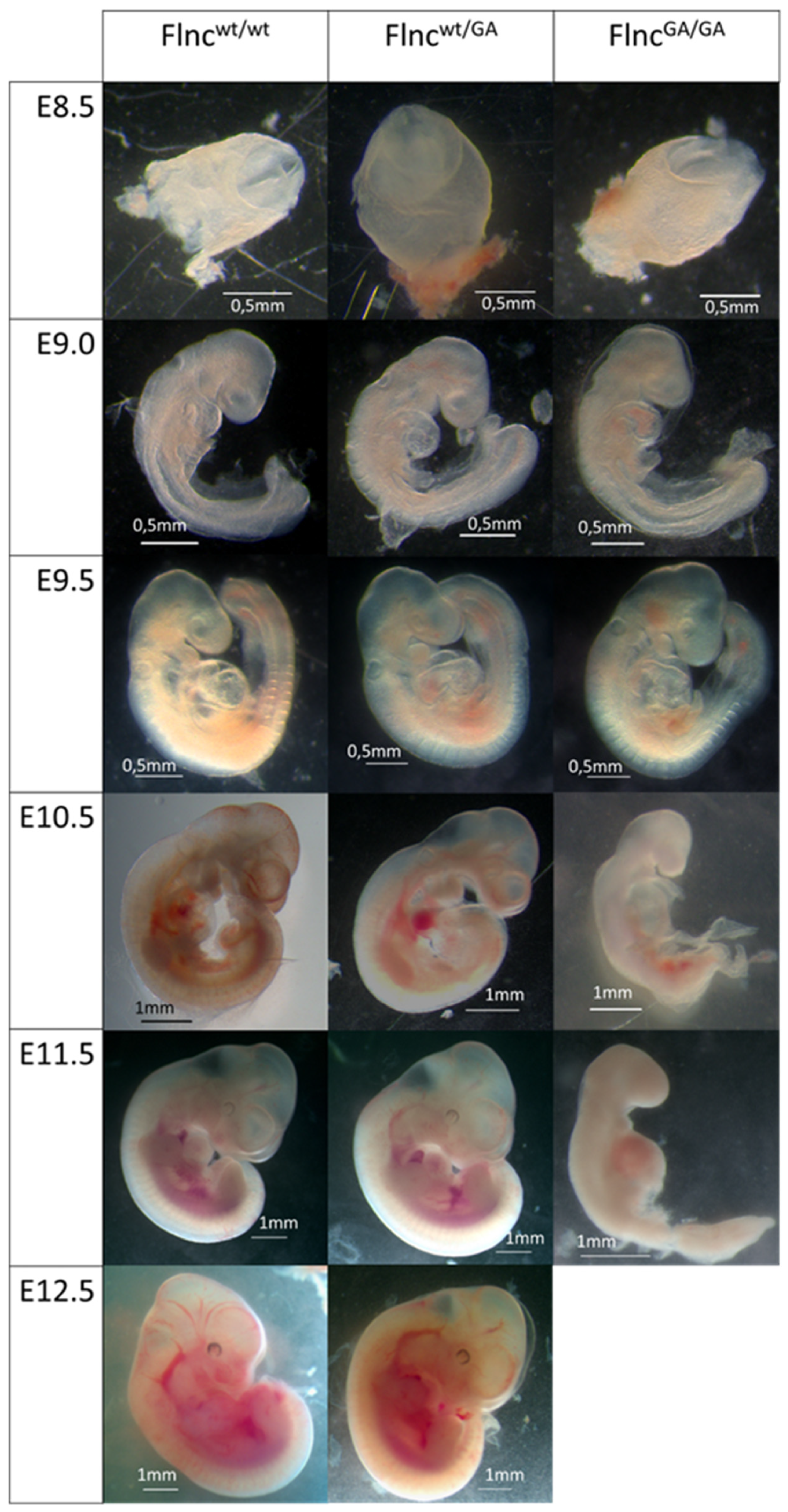
No difference was observed between wild-type and Flnc
wt/GA groups, embryos of both genotypes were developing normally. At the same time, Flnc
GA/GA homozygotes develop at a normal rate only until embryonic day 9.5 (E9.5) (
Figure 2). At E10.5 Flnc
GA/GA embryos lack properly developed forebrain hemispheres and forelimb and hindlimb buds that are characteristic of this developmental stage. At E11.5 developmental delay of homozygous knockout is even more pronounced, general embryo tissue degradation is observed, and by E12.5 homozygous embryos are presumably completely resorbed — no Flnc
GA/GA samples were found at this stage, while empty deciduae were observed.
2.3. Indicators of Grip Strength and Endurance of Mice with Genotypes Flncwt/AGA, Flncwt/GA, FlncAGA/GA Distinguish from Wild-Type Animals
Next, we have studied the influence of mutant FLNC protein expression on development of pathological phenotype in vivo. Taking into consideration the fact that FLNC protein is normally expressed and present not only in cardiac Z-discs, but also in sarcomeres of striated skeletal muscles, we have hypothesized that introduced mutations may affect muscle activity of mutant animals. To test the hypothesis, we have performed two physiology tests — endurance and grip strength tests. It is important to note that assessment of muscle system functions, such as endurance and strength, in homozygous FlncGA/GA and FlncAGA/AGA animals was impossible due to embryonic lethality.
Four groups of 1.5–3 months old males were used for both tests: Flncwt/wt, Flncwt/GA, Flncwt/AGA, FlncAGA/GA.
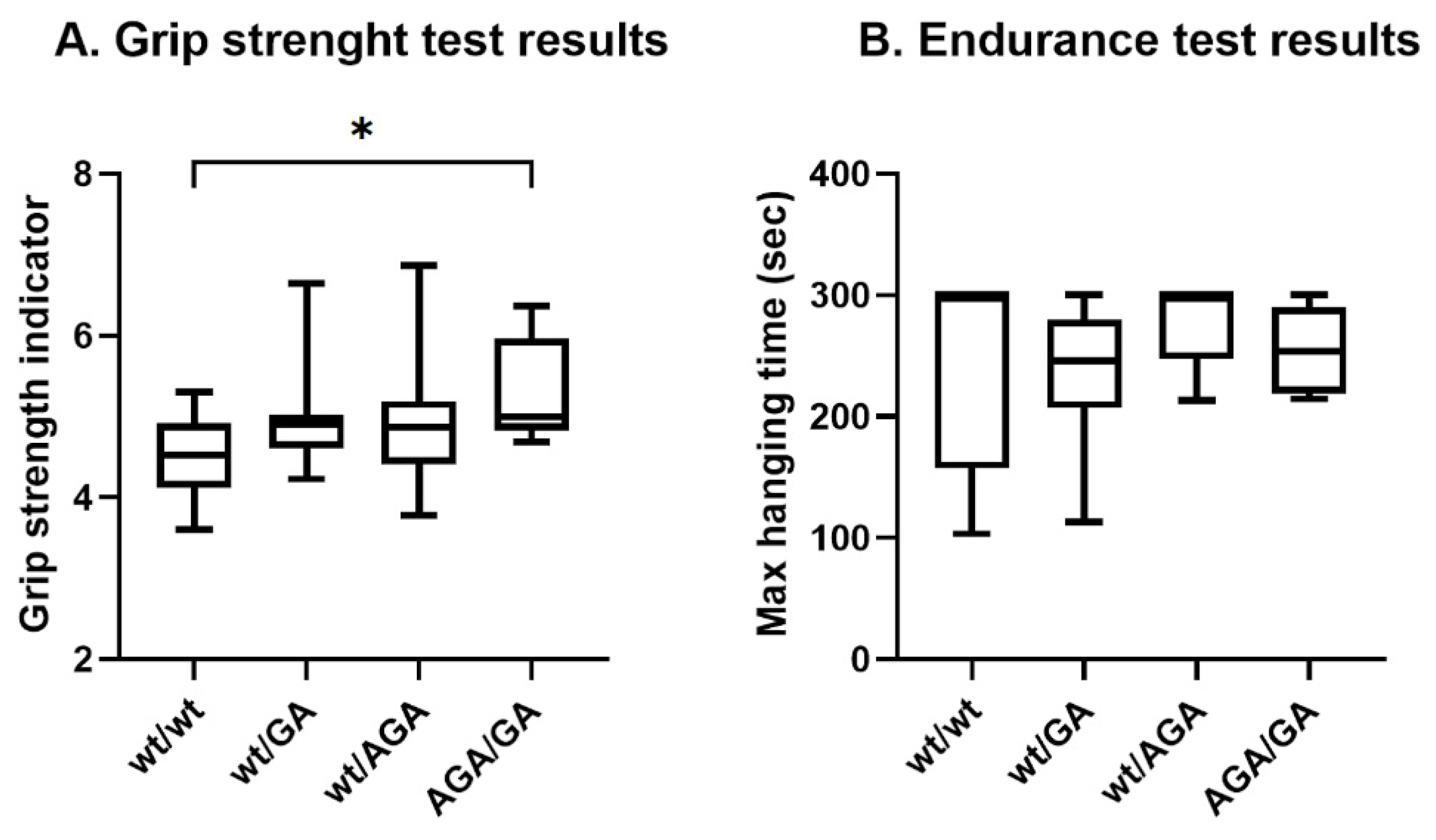
Surprisingly, quantitative analysis has revealed statistically significant increase in grip strength in Flnc
AGA/GA mice as compared to wild-type animals (p = 0.0017, Mann-Whitney U-test) (
Figure 3). Muscle endurance values were similar in all experimental groups and wild-type animals (p = 0.53, Kruskal-Wallis test).
2.4. Protein Alterations Do Not Impair Heart Function
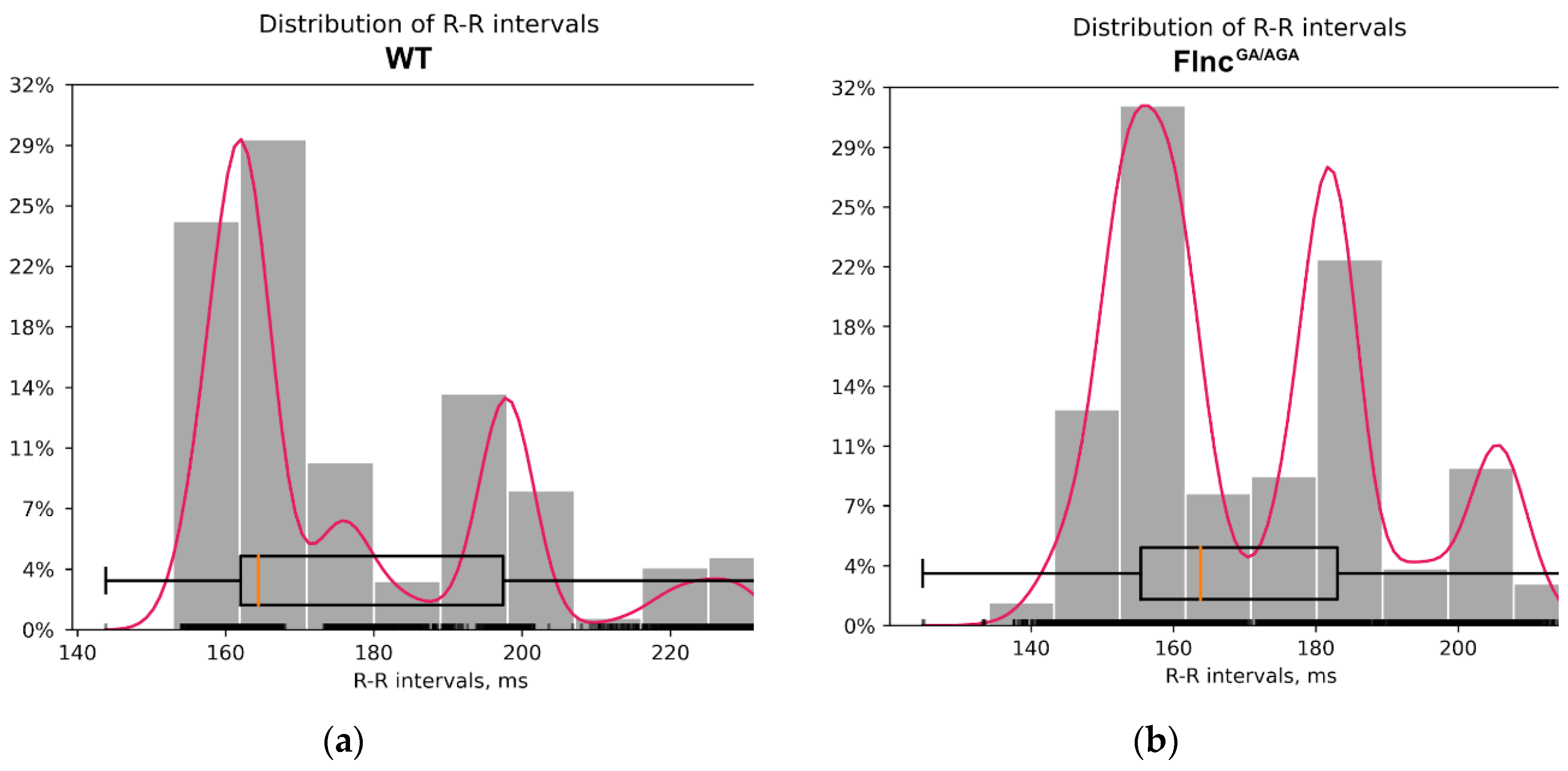
All mice exhibited a regular sinus rhythm in their ECG data. The mean values (MeanNN) and standard deviations (SDNN) of the R-R intervals did not show significant differences between the two groups (p-value MeanNN = 0.49, p-value SDNN = 0.69; Mann-Whitney U-test) (
Table 2,
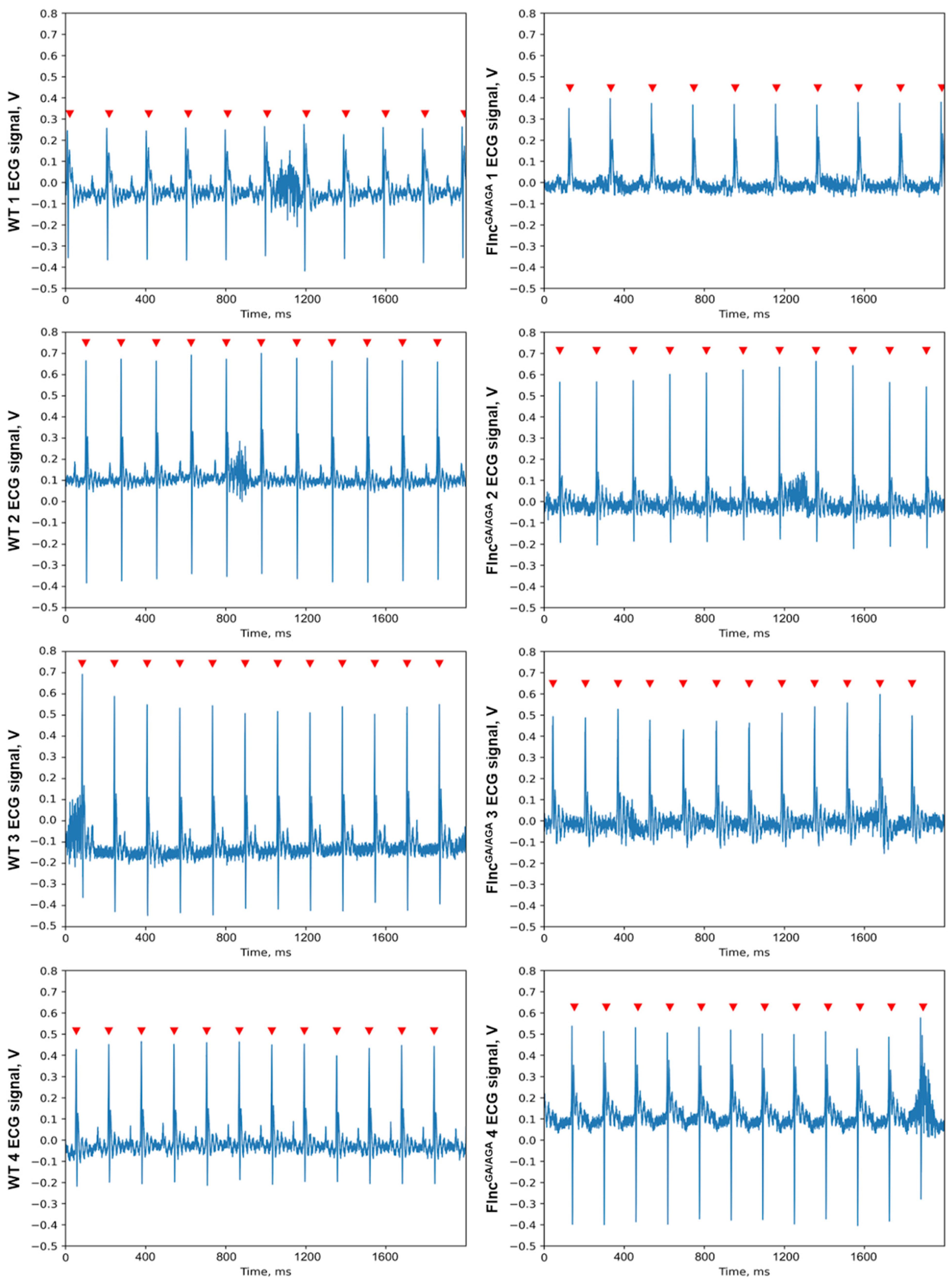
Figure 4). Thus, heart rate variability was comparable across groups. No specific qualitative differences were observed in the configuration of the atrioventricular complexes between the knockout and control mice, with amplitudes remaining within normal ranges and exhibiting only individual variations. Comprehensive ECG analysis revealed no abnormalities in excitability or conduction (
Figure 5). Consequently, the ECG findings indicated no significant changes in the electrophysiological state of the myocardium in the knockout mice.
The comparison of heart rate variability revealed no significant differences between the wild-type mice (WT) and the knockout mice (FlncGA/AGA). The following metrics were analyzed: MeanNN, which is the average of the R-R intervals; SDNN, the standard deviation of the R-R intervals; RMSSD, the square root of the mean of the squared differences between successive R-R intervals; SDSD, the standard deviation of successive differences between R-R intervals; CVNN, the coefficient of variation of R-R intervals, calculated as the standard deviation of the R-R intervals (SDNN) divided by the mean of the R-R intervals (MeanNN); CVSD, the root mean square of successive differences (RMSSD) divided by the mean of the R-R intervals (MeanNN); and MedianNN, the median of the R-R intervals.
2.5. Myocardium and Skeletal Muscle Histological Sections of Flncwt/GA, Flncwt/AGA and FlncAGA/GA Mice Do Not Exhibit Pathological Features Characteristic of Myopathy
We have prepared histological sections of cardiac and skeletal muscle of adult Flnc
wt/GA, Flnc
wt/AGA and Flnc
AGA/GA animals. Representative images of heart and skeletal muscle sections from animals of different genotypes stained with H&E (Hematoxylin and Eosin) and HBFP (Haematoxylin-Basic Fuchsin-picric acid) are shown on
Figure 6.
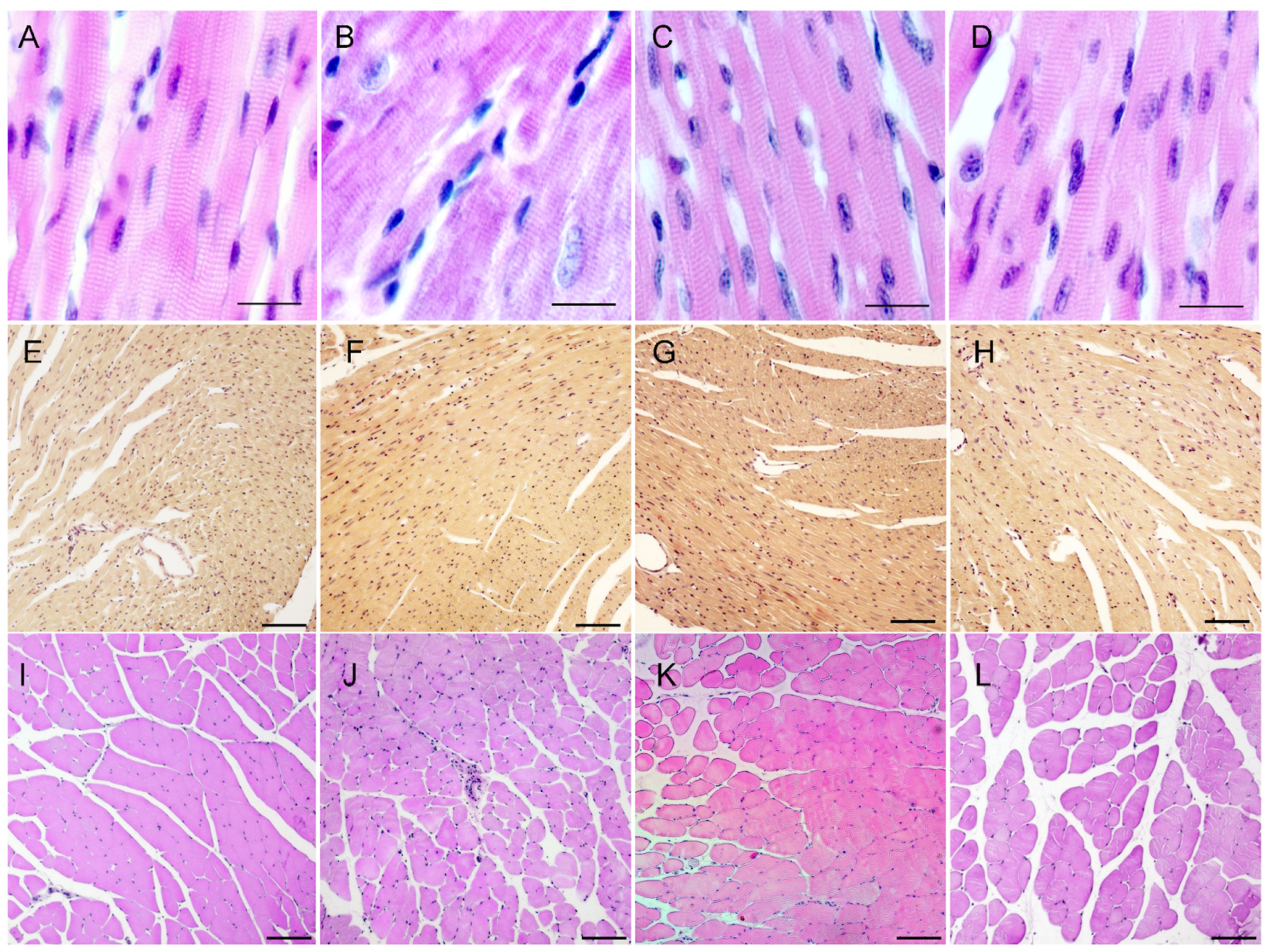
Sections show normal cross-striated structure, absence of obvious cellular aggregates and fibrosis. Cardiomyocytes’ nuclei have normal peripheral cellular localization. HBFP method is used to visualize fibrin and connective tissue structures in ischemia-damaged myocardium — fuchsin stains damaged regions red, while intact myocardial tissue is yellow to yellow-brown, allowing to thoroughly evaluate cardiac muscle condition. Using HBFP staining we have found that heterozygous Flncwt/GA animals do not show any difference as compared to wild-types, and their cardiac tissue is unaffected.
2.6. Computational Prediction of Mutant Proteins Structure
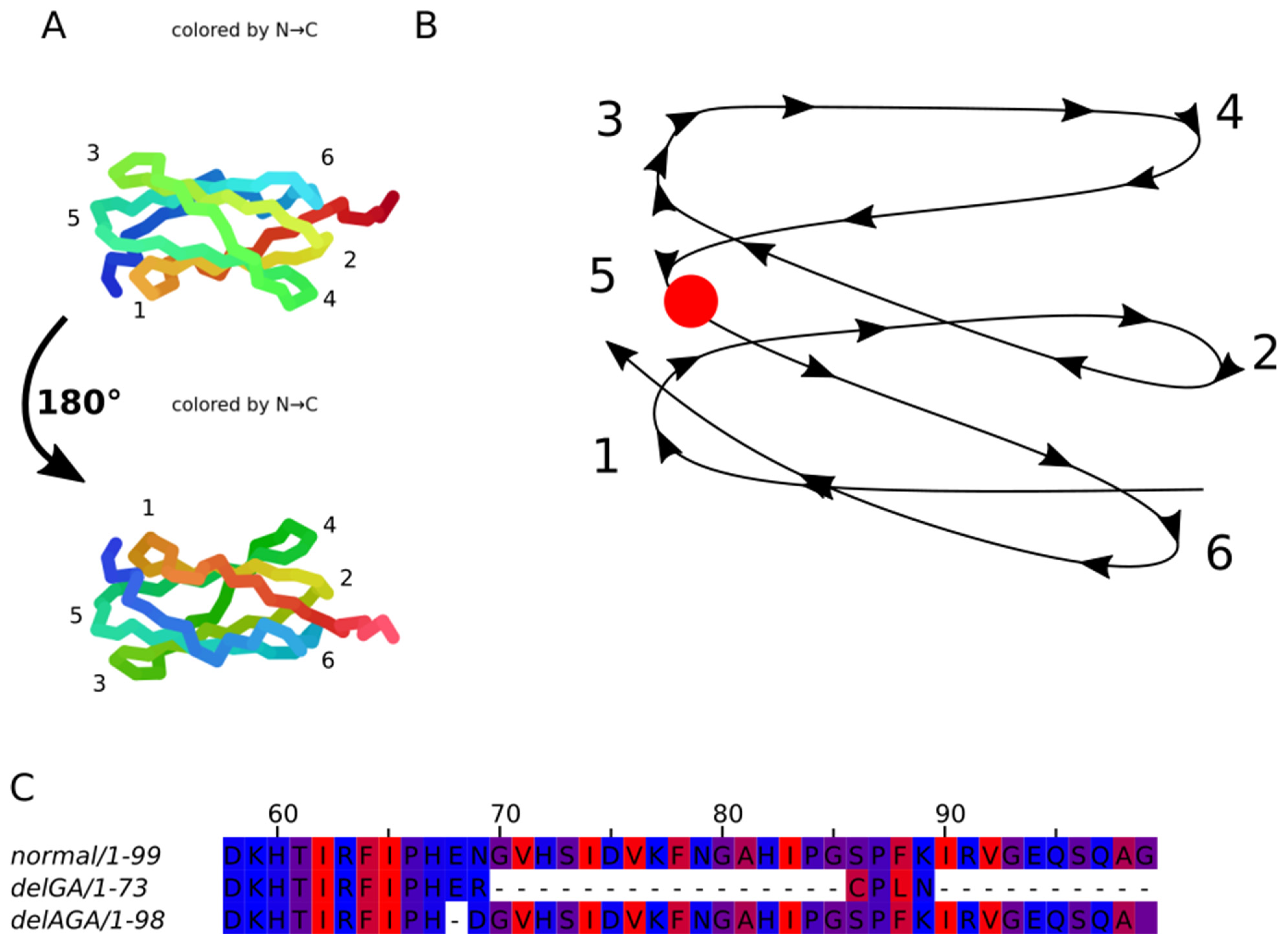
We have performed computational analysis of possible effects of introduced mutations on the structure and function of FLNC protein. Using AlphaFold2 software a 3D-model of spatial arrangement of native filamin С domain 22 loops was built (
Figure 7A). Based on previously solved and published domain 14–15 structure (PDB ID: 7OUU), as well as known filamin A structure (PDB ID: 6D8C), it can be assumed that loops 1, 3 and 5 are located at the region of interaction between domains 22 and 23. This interaction might be due to two hydrophobic surfaces being in close proximity or formation of hydrogen bonds between amino acids of closely located alpha helices.
DelGA variant leads to a frame shift and translation termination six amino acid residues downstream of the mutation site, potentially affecting Ig-like domain 22 folding. Homodimerization domain is missing in this variant due to being downstream of the premature stop codon.
Variant delAGA does not lead to a frame shift, but it results in a residue substitution p.E2472D and deletion of Asn2473. Further protein synthesis is not affected in this case.
Both mutations cause structural changes in loop 5 (
Figure 7B). Variant delAGA leads to a loss of a single hydrophilic amino acid, possibly resulting in a reduced number of hydrogen bonds as compared to a native protein. As exact position of Asn2473 side chain is unclear, its deletion may reduce the interaction between loop 5 and loop 1 or loop 3 or weaken the interaction between domains 22 and 23. In both cases weakening of the interaction leads to an insufficiently strong binding, potentially making it difficult to form a functional homodimer.
FlncGA mutation does not cause a reduction in the number of hydrophilic amino acids in loop 5, although loop 6 and domains 23–24 disappear from the protein due to frame-shift (
Figure 7C).
3. Discussion
In humans heterozygous mutations in
FLNC gene can cause various types of cardiomyopathies [
15,
16]. However, modelling this pathology in mouse strains is challenging as homozygous animals are either embryonically lethal [
11,
17], or die soon after birth at P0 [
18], whereas heterozygous animals do not have any pathological phenotype [
18].
The phenotype of animal strains reported in this study is in agreement with previously published data — both mutations are embryonically lethal. Flnc
GA/GA embryos start displaying developmental delay between Е9.5 and Е10.5, while heterozygous Flnc
wt/GA embryos develop normally. At Е9.5 Flnc
GA/GA embryos are morphologically indistinguishable from controls, suggesting that the earliest stages of cardiogenesis starting from cardiogenic mesoderm specification up to at least heart tube rotation are unaffected. At later stages mutation in the FLNC gene starts to interfere with the normal development and a pathological phenotype develops — at E10.5 homozygous Flnc
GA/GA embryos display developmental delay, and by Е12.5 homozygous embryos are not found, suggestive of their death and absorption. Most probably, the reason behind Flnc
GA/GA embryo lethality is cardiac dysfunction, as the timing of developmental delay onset is consistent with the start of heart functioning in normally developing mouse embryos [
19]. Similarly to Flnc
GA mutant strain, analysis of offspring genotypes in Flnc
wt/AGA x Flnc
wt/AGA crossing revealed absence of homozygous Flnc
AGA/AGA animals, suggesting embryonic lethality. Moreover, observed heterozygous to wild-type genotypes ratio is 1:1, while expected would be 2:1, suggesting Flnc
wt/AGA are also prone to embryonic death.
Both physiology tests and histology analyses of skeletal and cross-striated muscles performed on heterozygous Flncwt/GA and Flncwt/AGA animals did not reveal any difference as compared to wild-type mice. This suggests that presence of a single normal allele of Flnc is sufficient for normal muscle functioning.
Surprisingly, compound heterozygous animals (FlncGA/AGA) developed normally, were born alive, reached puberty, and displayed statistically significantly higher grip strength as compared to wild-type animals. The DNA samples of these FlncGA/AGA mice were re-sequenced to confirm the compound heterozygous genotype and absence of wild-type allele. ECG has revealed the absence of any difference between compound heterozygous and wild-type mice.
Based on the data on relative positions of native protein domains during dimerization [
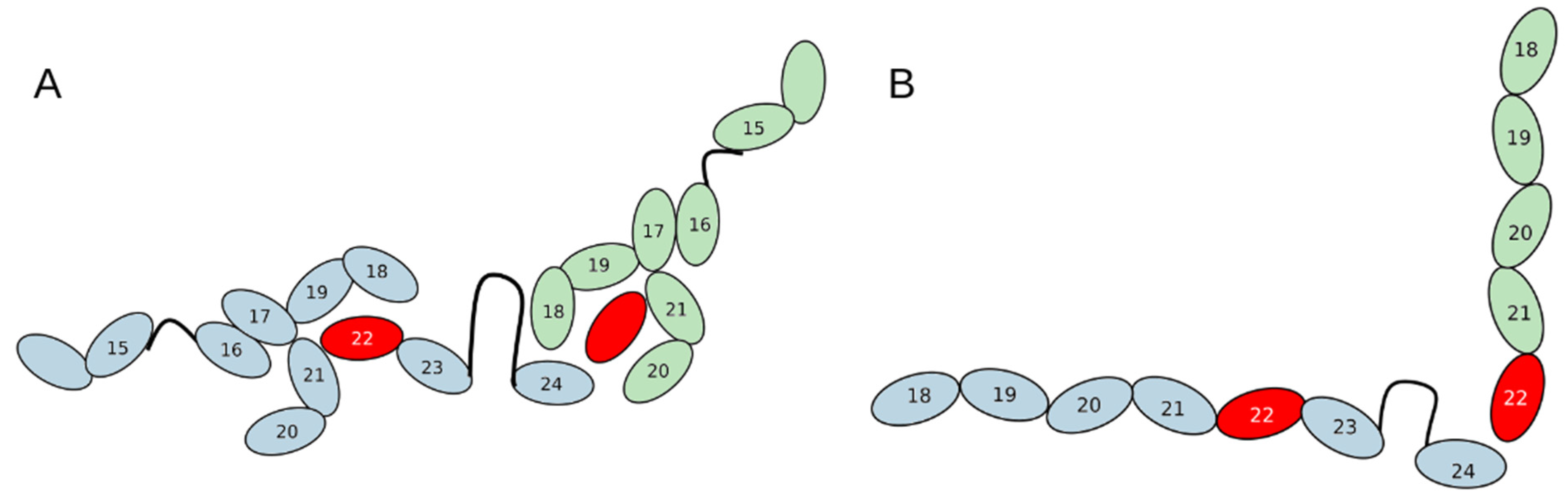
6] we proposed a hypothetical model of “heterodimerization” of two mutant proteins, where mutual compensation of their “weak points” occur. FLNCdelGA variant lacks domains 23–24, required for dimerizations, while homodimerization of FLNCdelAGA variant is not strong enough for a functional dimer formation. Possible interaction between FLNCdelAGA and FLNCdelGA in case of heterodimerization in relaxed and tense is shown in
Figure 8.
Normally, domain 24 is responsible for dimerization, while in a protein molecule synthesized from delGA allele domains 23 and 24 are missing.
Such domains’ position may explain the fact that heterozygous delGA mice perform better in physical tests — instead of a single domain, three domains are involved in dimerization in truncated protein in a proposed model.
Although hypotheses, proposed here, require further experimental confirmation, all obtained results can be used in the future to develop gene therapy to treat pathology related to FLNC gene mutations.
4. Materials and Methods
4.1. Animals
4.1.1. Animal Housing
Females of the CD1 strain were used as surrogate mothers for genetically edited embryos, and F1 CBA х C57Bl/6J hybrids were used as embryo donors. To get pseudopregnant surrogate mothers females were mated with CD1 strain vasectomized males. C57Bl/6J males were used as breeders. The animals were given ad libitum access to water and specialized feed. The dark-light cycle was 14/10, air temperature in the facility — 23±1℃.
All experiments involving animals were carried out in adherence with local regulations and approved by the IGB RAS bioethics committee.
4.1.2. Transgenic Animals’ Generation
Transgenic mice were obtained by standard protocol as described previously [
20]. Briefly, zygotes were obtained by superovulation protocol, genetic construction was injected into the cytoplasm of zygotes which were cultured after microinjection for 1 day
in vitro and then embryos which cleaved for 2 blastomeres were transferred into the oviducts of the pseudopregnant females.
On day 19 after embryo transfer, cesarean section and extraction of survived embryos was performed if natural labor did not occur. In case of natural delivery pups were left with their biological mother for rearing. In case of absence of signs of labor pregnant mice were sacrificed by cervical dislocation, cesarean section was performed and newborn pups were placed with a female who was rearing a litter of a similar age.
4.2. Genetic Construct Production
The sgRNA prediction was performed using online service CRISPOR TEFOR (
http://crispor.tefor.net/, accessed 12.09.2022) considering subsequent SpCas9 nuclease usage. An efficient sgRNA that cleaves the intended site of mutation was selected.
The sgRNA was cloned into a px330 vector (Addgene;
https://www.addgene.org/42230/) as per manufacturer’s protocol for plasmid microinjection. Vector was cleaved using BbsI-HF restriction enzyme (New England Biolabs) and dephosphorylated with FastAP enzyme (Thermofisher Scientific, USA), followed by agarose gel electrophoresis for product separation and extraction using Monarch DNA Gel Extraction Kit (New England Biolabs, MA, USA). Oligonucleotides were synthesized by Evrogen company (Russian Federation) follows: 5’-CACCATTGAGTGGACACCGTTCTCA-3’, 5’-ATTGAGTGACACCGTTCTCAGTTT-3’. Oligonucleotides were phosphorylated by polynucleotide kinase (Thermofisher Scientific, USA), annealed and ligated with previously prepared linearized vector using T4 DNA ligase (Thermofisher Scientific, USA). Ligation mixture was transformed into competent XL1blue strain cells (Evrogen, Russian Federation). Colony screening was performed using a PCR kit (Isogen Lifescience B.V., The Netherlands) and U6-forv primer (GAGGGCCTATTTCCCATGATT) and a reverse primer used for cloning. Two sgRNA-containing colonies were cultured overnight and plasmid DNA was extracted using Monarch Plasmid Miniprep Kit. Plasmid sequences were analyzed by Sanger sequencing, analysis was performed by Evrogen company using U6-forv primer. Sequencing results were analyzed with Chromas software (Chromas Lite).
Microinjection mix contained 1 ng/µL px330 vector with sgRNA insertion.
4.3. Genotyping
4.3.1. F0 Genotyping
F0 genotyping was carried out by sequencing of genomic DNA. Region of interest was amplified using Flnc seq F (GTGCCTGACCTACCAAGGCA) and Flnc seq R (CATCTTTCGCGGTGCAGCTT) primers, obtained fragment was sequenced using Flnc seq R primer.
Genotyping of F1 and further generations was carried out using real-time PCR with fluorescent probes.
4.3.2. F1 and Further Generations Genotyping.
Mice and embryos were genotyped using real-time PCR. Ear samples were taken from 1–4 weeks old animals for genotyping purposes. DNA was extracted using an alkaline lysis method. Tissue samples were placed in 200 µL of lysis buffer (25 mM NaOH, 0.2 mM EDTA, pH 12) and incubated at 95 °C for 1.5 h, followed by removal of undissolved tissues by centrifugation for 30 s (Microspin FV-2400, bioSan, Latvia). DNA-containing supernatant was used as a PCR template, nucleic acid concentration was measured spectrophotometrically (IMPLEN NanoPhotometer P300, IMPLEN, Germany). DNA samples were diluted approximately 100x with milli-Q water to reach optimal concentration of 10 ng/µL DNA per reaction. The diluted DNA sample (3 µL) was added per well in 96-well plate (SSI Bio 3401-00, USA). In case of embryo genotyping 50 µL of lysis buffer was enough to lyse the tissue, sample taken from the embryo, and 10x dilution was sufficient. Following primers were used for real-time PCR: forward 5’-GAAGAAGCATGGAGGCCCAC-3’, reverse 5’-GCTCCCCAACACGGATCTTG-3’ and fluorescent probes FAM-CCCATGAGAACGGTGTCCAC-bhq1 and ROX-CCCATGACGGTGTCCACTCA-bhq2 (for delAGA allele detection), or ROX-CCCATGAACGGTGTCCACTCA-bhq2 (for delGA allele detection).
4.4. Physiology Test Performing
Tests were carried out on two groups of 1.5–3 months old males, 8–9 animals in each: wild-type controls and experimental group of heterozygous animals.
4.4.1. Endurance Test
A metal wire (2 mm thick, 50 cm long) was used in the test. The wire was fixed 30cm above the cage filled with soft bedding to prevent mice from hurting their limbs in case of falling (
Figure S1). All mice were weighed prior to experiment. Mice were grabbed by the tail and hung on the wire with its forelimbs, not allowed to use hind limbs. Duration of mouse hanging was assessed. Maximal hanging time was limited to 300 s, and if the mouse did not fall within this time, the experiment with the animal was terminated for the day. If the mouse did fall after less than 300 s, the duration of hanging was recorded and the mouse was hung again; 10 attempts in total were given, after which the experiment with the animal was terminated for the day. The test was performed three times with each male from all experimental groups with one day breaks to let mice rest (at least 1 min) between independent repeats.
As a result, we got data on maximal hanging duration for each animal in triplicates and on the number of attempts that were analyzed statistically.
4.4.2. Grip Strength Test
Highly sensitive Grip Strength Test Meter for Mice and Rats (“IITC Life Science”, CA, USA) was used to conduct a grip strength assessment. Animals were grabbed by the tail and brought close to grabbing metal device, fixed on a sensor (
Figure S2). Mouse was allowed to grab the grid only with its forelimbs, after which the animal was gently and slowly pulled backwards until it released the grid. Sensor recorded the maximal grip strength in grams. Each animal was given three to five attempts with at least one-minute rest time in between. The test was performed three times with at least one day of rest. Obtained results were normalized to animal body weight measured prior to each test and were analyzed statistically.
4.5. Electrocardiography
The functional state of the myocardium in knockout mice was evaluated using standard ECG under isoflurane anesthesia [
21]. R-R intervals and their variability were analyzed, alongside a qualitative comparison of the atrioventricular complexes between two groups: four knockout mice (Flnc
GA/AGA) and four control mice (Flnc
wt/wt), all approximately one month old.
ECG recordings were performed at a frequency of 10000 Hz using the PowerGraph 3.x application. A 5-minute recording at a sampling rate of 5000 Hz was analyzed using a custom application developed with Python 3.11 libraries, including neurokit, pandas, numpy, scipy, seaborn, and others. R-R interval data outside the interquartile range (IQR) were filtered. Heart rate variability, defined as the variability of R-R intervals between consecutive heartbeats, was calculated using key metrics [
22].
4.6. Tissue Preparation and Histology
For histological study, 4 groups of mice were involved, 5–10 individuals of each genotype: Flnc
wt/wt, Flnc
wt/GA, Flnc
wt/AGA, Flnc
AGA/GA. After intraperitoneal sedation using Vezotil (VETSTEM® pharma & cell, Russian Federation) and Xyla (Interchemie werken "De Adelaar" B.V.,The Netherlands) transcardiac perfusion with saline followed by 10% neutral buffered formalin (NBF) was performed [
23]. For the sedation the mix containing 0.6 ml of Vezotil, 0.3 ml of Xyla and 9 ml of saline (0.9% NaCl) was used. Each mouse received 0.2 ml of that mix for sedation. Samples of limb muscles and hearts were collected and placed into NBF for 24 hours. After the completion of fixation, samples were processed via isopropanol dehydrating solution and mineral oil to paraffin wax according to manufacturer's instructions (Medix, Russian Federation). Tissue sectioning was conducted via a rotary microtome RMD-3000 (Medtehnikapoint, Russian Federation). Paraffin sections (5 µm) were cut both for H&E staining and HBPF staining. For H&E staining Mayer's Hematoxylin and Eosin aqueous solution (1%) were used (BioVitrum, Russian Federation). HBFP staining (BioVitrum, Russian Federation) was used for early detection of myocardial damage and necrosis. Images were captured on Nikon ECLIPSE Ti (“Nikon”, Japan).
4.7. Embryo Dissections for Macroscopic and Histology Analysis
Genetically edited female mice were mated with males carrying the same mutation until the discovery copulation plug, thus obtaining dated pregnancies. At E8.0 – E12.5 pregnant females were sacrificed by cervical dislocation and embryos were collected in saline. After decidua tissue and extraembryonic membranes removal embryo images were captured under NikonSMZ800N stereomicroscope (Japan), Basler camera (Germany) connected via Nikon LT-TV adapter (Japan) using Basler microscopy software (version: 2.1). Next, a tail region of the embryo posterior to hind limb buds was removed and used for DNA extraction for genotyping purposes. The remaining embryo was fixed for histology studies.
4.8. Statistical Analysis
Statistical analysis of the data was performed in GraphPad Prism (GraphPad Software 8.0). Average was calculated from independent technical repeats (n of repeats ≥ 3). Statistical significance was assessed using non-parametric analysis, as normal distribution could not be confirmed (as according to Kolmogorov-Smirnov criterion). To detect the difference between two groups Mann-Whitney U-test and the Bonferroni correction were applied. Result was considered significant at p < 0.008.
Bioinformatics analysis was conducted using AlphaFold2 Colab [
24], alignment was performed utilizing Jalview [
25], using Mafft algorithm with defaults.
5. Conclusions
In humans, heterozygous mutations of
FLNC gene cause cardiomyopathy [
26], and animal models of hereditary cardiomyopathy are required to study possible therapeutic agents and approaches. In this work we studied two strains of genetically edited mice that carry mutation in
Flnc gene to assess if they can be used as a hereditary cardiomyopathy model. Although both mutations turned out to be embryonically lethal in homozygous state (homozygous Flnc
GA/GA embryos die at E10.5), heterozygous animals develop successfully and do not display any cardiomyopathy or skeletal muscle dysfunction signs in adulthood. Apparently, this study serves to illustrate that mice are not always a good model of human pathological conditions. The reasons behind this discrepancy between mouse and human could be lifespan difference or physiological features, such as circulating blood volume. Surprisingly, compound heterozygosity (Flnc
AGA/GA genotype) is beneficial as compared to homozygosity of either mutant alleles. Together, our data suggest that even partial recovery of
Flnc allele function may restore muscle function, opening up a novel field for development of gene therapy to treat myopathies caused by mutations in this gene.
Supplementary Materials
The following supporting information can be downloaded at:
Preprints.org, Figure S1: Endurance test performing. Mouse hanging on a wire.; Figure S2: Grip strength test performing. Mouse is grabbing the grid with its forelimbs, while it is slowly pulled backwards.
Author Contributions
Conceptualization, L.I., K.K., A.B., M.K., A.R., A.M., V.B., M.F., Y.S; methodology, L.I., K.K., I.B., P.S., A.B., M.K., Y.O., N.K., A.R., A.M, V.B., M.F., Y.S.; writing—original draft preparation, T.M., L.I., K.K., M.F., Y.S.; writing—review and editing, M.F., D.D., Y.S., L.I, K.K.; visualization, T.M., L.I., I.B.; project administration, Y.S.; funding acquisition, Y.S. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Ministry of Science and Higher Education of the Russian Federation, grant number 075-15-2019-1661.
Institutional Ethics Committee Statement
The animal study protocol was approved by the Institutional Ethics Committee of IGB RAS (protocol code #15 from March 15, 2023).
Data Availability Statement
The data supporting the findings of this study are available within the article and its supplementary materials.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Lu, J., et al., Filamin B mutations cause chondrocyte defects in skeletal development. Hum Mol Genet, 2007. 16(14): p. 1661-75. [CrossRef]
- Nakamura, F., T.P. Stossel, and J.H. Hartwig, The filamins: organizers of cell structure and function. Cell Adh Migr, 2011. 5(2): p. 160-9.
- van der Flier, A. and A. Sonnenberg, Structural and functional aspects of filamins. Biochim Biophys Acta, 2001. 1538(2-3): p. 99-117.
- Verdonschot, J.A.J., et al., A mutation update for the FLNC gene in myopathies and cardiomyopathies. Hum Mutat, 2020. 41(6): p. 1091-1111. [CrossRef]
- Baldassarre, M., et al., Filamins regulate cell spreading and initiation of cell migration. PLoS One, 2009. 4(11): p. e7830. [CrossRef]
- Mao, Z. and F. Nakamura, Structure and Function of Filamin C in the Muscle Z-Disc. Int J Mol Sci, 2020. 21(8). [CrossRef]
- Ortiz-Genga, M.F., et al., Truncating FLNC Mutations Are Associated With High-Risk Dilated and Arrhythmogenic Cardiomyopathies. J Am Coll Cardiol, 2016. 68(22): p. 2440-2451. [CrossRef]
- van der Flier, A., et al., Different splice variants of filamin-B affect myogenesis, subcellular distribution, and determine binding to integrin [beta] subunits. J Cell Biol, 2002. 156(2): p. 361-76.
- Kent H. Rehfeldt, W.J.M., Gregory A. Nuttall, William C. Oliver, Perioperative Transesophageal Echocardiography, ed. G.W.F. David L. Reich. 2014.
- Brieler, J., M.A. Breeden, and J. Tucker, Cardiomyopathy: An Overview. Am Fam Physician, 2017. 96(10): p. 640-646.
- Dalkilic, I., et al., Loss of FilaminC (FLNc) results in severe defects in myogenesis and myotube structure. Mol Cell Biol, 2006. 26(17): p. 6522-34. [CrossRef]
- Chevessier, F., et al., Myofibrillar instability exacerbated by acute exercise in filaminopathy. Hum Mol Genet, 2015. 24(25): p. 7207-20. [CrossRef]
- Schuld, J., et al., Homozygous expression of the myofibrillar myopathy-associated p.W2710X filamin C variant reveals major pathomechanisms of sarcomeric lesion formation. Acta Neuropathol Commun, 2020. 8(1): p. 154. [CrossRef]
- Theiler, K., The House Mouse Atlas of Embryonic Development. 1989: Springer Berlin, Heidelberg.
- Ohiri, J.C., et al., Reduction of Filamin C Results in Altered Proteostasis, Cardiomyopathy, and Arrhythmias. J Am Heart Assoc, 2024. 13(10): p. e030467. [CrossRef]
- Shatunov, A., et al., In-frame deletion in the seventh immunoglobulin-like repeat of filamin C in a family with myofibrillar myopathy. Eur J Hum Genet, 2009. 17(5): p. 656-63.
- Wu, T., et al., Filamin C is Essential for mammalian myocardial integrity. PLoS Genet, 2023. 19(1): p. e1010630. [CrossRef]
- Zhou, Y., et al., Loss of Filamin C Is Catastrophic for Heart Function. Circulation, 2020. 141(10): p. 869-871. [CrossRef]
- Savolainen, S.M., J.F. Foley, and S.A. Elmore, Histology atlas of the developing mouse heart with emphasis on E11.5 to E18.5. Toxicol Pathol, 2009. 37(4): p. 395-414. [CrossRef]
- Kalinina, A.A., et al., Physiological and Functional Effects of Dominant Active TCRα Expression in Transgenic Mice. Int J Mol Sci, 2023. 24(7). [CrossRef]
- Alexander V. Balatskiy, A.M.N., Aleksandr A. Lanin, Vera S. Ovechkina, Semyon S. Sabinin, , et al., Thermogenetics for cardiac pacing. 2024.
- Pham, T., et al., Heart Rate Variability in Psychology: A Review of HRV Indices and an Analysis Tutorial. Sensors, 2021. 21(12): p. 3998. [CrossRef]
- Gage, G.J., D.R. Kipke, and W. Shain, Whole animal perfusion fixation for rodents. J Vis Exp, 2012(65).
- Mirdita, M., et al., ColabFold: making protein folding accessible to all. Nat Methods, 2022. 19(6): p. 679-682. [CrossRef]
- Waterhouse, A.M., et al., Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics, 2009. 25(9): p. 1189-1191. [CrossRef]
- Melnik O.V., K.A.A., Poluboyarinova O.Y., Vakhrushev Yu.A., Fomisheva Yu.V., Sitnikova M.Yu., Simonenko M.A., Vershinina T.L., Kozyreva A.A., Polyakova A.A., Vasichkina T.S., Gudkova A.Ya., Kostareva А.А. , Clinical and morphological characteristics of restrictive cardiomyopathy associated with mutations in the filamin c gene. (Article in Russian). Translational Medicine, 2018. 5(3): p. 15-22.
Figure 1.
Schematic representation of FLNC monomer indicating the location of introduced mutations as confirmed by sequencing. Bold and underlined text annotate sgRNA and PAM-sequence, respectively.
Figure 1.
Schematic representation of FLNC monomer indicating the location of introduced mutations as confirmed by sequencing. Bold and underlined text annotate sgRNA and PAM-sequence, respectively.
Figure 2.
Mouse embryos of different genotypes at Е8.5 – Е12.5 stages; stereomicroscopy.
Figure 2.
Mouse embryos of different genotypes at Е8.5 – Е12.5 stages; stereomicroscopy.
Figure 3.
Physiology test results. А. Grip strength test; grip strength indicator was calculated as an average of grip strength (g) to mouse body weight (g) rates, average value were calculated from three technical repeats; Mann-Whitney U-test, p = 0.0161 (Flncwt/wt(N=21)/ Flncwt/GA(N=18)), p = 0.2226 (Flncwt/wt(N=21)/ Flncwt/AGA(N=11)), p = 0.0017 (Flncwt/wt(N=21)/ FlncAGA/GA(N=9)); B. Endurance test; Mann-Whitney U-test, p = 0.6147 (Flncwt/wt(N=9)/ Flncwt/GA(N=8)), p = 0.5105 (Flncwt/wt(N=9)/ Flncwt/AGA(N=5)), p = 0.9414 (Flncwt/wt(N=9)/ FlncAGA/GA(N=7)). Wiskers on plots refer to min to max, the plot represent IQR (interquartile range) and the line inside the plot indicates median, * corresponds to adjusted p < 0.008.
Figure 3.
Physiology test results. А. Grip strength test; grip strength indicator was calculated as an average of grip strength (g) to mouse body weight (g) rates, average value were calculated from three technical repeats; Mann-Whitney U-test, p = 0.0161 (Flncwt/wt(N=21)/ Flncwt/GA(N=18)), p = 0.2226 (Flncwt/wt(N=21)/ Flncwt/AGA(N=11)), p = 0.0017 (Flncwt/wt(N=21)/ FlncAGA/GA(N=9)); B. Endurance test; Mann-Whitney U-test, p = 0.6147 (Flncwt/wt(N=9)/ Flncwt/GA(N=8)), p = 0.5105 (Flncwt/wt(N=9)/ Flncwt/AGA(N=5)), p = 0.9414 (Flncwt/wt(N=9)/ FlncAGA/GA(N=7)). Wiskers on plots refer to min to max, the plot represent IQR (interquartile range) and the line inside the plot indicates median, * corresponds to adjusted p < 0.008.
Figure 4.
Distribution of R-R intervals: A – in the wild type mice group (WT); B – in the knockout mice group (FlncGA/AGA).
Figure 4.
Distribution of R-R intervals: A – in the wild type mice group (WT); B – in the knockout mice group (FlncGA/AGA).
Figure 5.
The single lead ECG segments of wild-type (WT) mice (left) and knockout mice (FlncGA/AGA) (right). The peaks of the R waves in the ventricular complexes are indicated by red arrows. No specific qualitative differences were observed in the configuration of the atrioventricular complexes between the knockout and control mice, with amplitudes remaining within normal ranges and exhibiting only individual variations.
Figure 5.
The single lead ECG segments of wild-type (WT) mice (left) and knockout mice (FlncGA/AGA) (right). The peaks of the R waves in the ventricular complexes are indicated by red arrows. No specific qualitative differences were observed in the configuration of the atrioventricular complexes between the knockout and control mice, with amplitudes remaining within normal ranges and exhibiting only individual variations.
Figure 6.
Frontal myocardium sections (A-H), stained with H&E (A-D) and HBFP (E-H), and sagittal sections of gastrocnemius muscles (I-L), stained with H&E. А, E, I — wild-type male; B, F, J — Flncwt/GA; C, G, K — male Flncwt/AGA; D, H, L — female FlncAGA/GA. A-D — scale bar: 20 μm. E-L — scale bar: 100 μm.
Figure 6.
Frontal myocardium sections (A-H), stained with H&E (A-D) and HBFP (E-H), and sagittal sections of gastrocnemius muscles (I-L), stained with H&E. А, E, I — wild-type male; B, F, J — Flncwt/GA; C, G, K — male Flncwt/AGA; D, H, L — female FlncAGA/GA. A-D — scale bar: 20 μm. E-L — scale bar: 100 μm.
Figure 7.
Three-dimensional arrangement of the loops of filamin С domain 22. Structure predicted using AlphaFold2: A — AlphaFold2 output; B — schematic loop arrangement. Loops are numbered from N- to C-terminus; C — domain 22 sequences alignment from FLNCwt and mutant FLNCGA and FLNCAGA. Amino acids are colored according to their hydrophobic/hydrophilic properties: red are the most hydrophobic residues, blue are the most hydrophilic ones; the alignment was done for the domain 22 sequence only.
Figure 7.
Three-dimensional arrangement of the loops of filamin С domain 22. Structure predicted using AlphaFold2: A — AlphaFold2 output; B — schematic loop arrangement. Loops are numbered from N- to C-terminus; C — domain 22 sequences alignment from FLNCwt and mutant FLNCGA and FLNCAGA. Amino acids are colored according to their hydrophobic/hydrophilic properties: red are the most hydrophobic residues, blue are the most hydrophilic ones; the alignment was done for the domain 22 sequence only.
Figure 8.
Possible folding of two mutant domains in case of heterodimerization: A — relaxed state; B — tense state. Mutant domain is highlighted in red. DelAGA variant is blue, delGA variant is green.
Figure 8.
Possible folding of two mutant domains in case of heterodimerization: A — relaxed state; B — tense state. Mutant domain is highlighted in red. DelAGA variant is blue, delGA variant is green.
Table 1.
Number of animals of each genotype observed and expected following (a) Flncwt/GA x Flncwt/GA, (b) Flncwt/AGA x Flncwt/AGA and (c) Flncwt/AGA x Flncwt/GA crossings.
Table 1.
Number of animals of each genotype observed and expected following (a) Flncwt/GA x Flncwt/GA, (b) Flncwt/AGA x Flncwt/AGA and (c) Flncwt/AGA x Flncwt/GA crossings.
| (a) Flncwt/GA x Flncwt/GA |
| |
Flncwt/wt |
Flncwt/GA |
FlncGA/GA |
Total |
| Observed* |
32 (28.57%) |
80 (71.43%) |
0 (0%) |
112 |
| Expected |
28 (25%) |
56 (50%) |
28 (25%) |
112 |
| (b) Flncwt/AGA x Flncwt/AGA |
| |
Flncwt/wt |
Flncwt/AGA |
FlncAGA/AGA |
Total |
| Observed* |
29 (51.79%) |
27 (48.21%) |
0 (0%) |
56 |
| Expected |
14 (25%) |
28 (50%) |
14 (25%) |
56 |
| (c) Flncwt/AGA x Flncwt/GA |
| |
Flncwt/wt |
Flncwt/GA |
Flncwt/AGA |
FlncGA/AGA |
Total |
| Observed |
18 (26.09%) |
18 (26.09%) |
15 (21.74%) |
18 (26.09%) |
69 |
| Expected |
17.25 (25%) |
17.25 (25%) |
17.25 (25%) |
17.25 (25%) |
69 |
Table 2.
Heart Rate Variability Metrics.
Table 2.
Heart Rate Variability Metrics.
| Type of mice |
WT 1 |
WT 2 |
WT 3 |
WT 4 |
FlncGA/AGA 1 |
FlncGA/AGA 2 |
FlncGA/AGA 3 |
FlncGA/AGA 4 |
| Age |
40 days |
40 days |
30 days |
30 days |
36 days |
36 days |
32 days |
32 days |
| Sex |
female |
male |
male |
male |
male |
female |
male |
male |
| MeanRR |
197.48 |
196.38 |
162.54 |
160.50 |
197.11 |
180.67 |
156.28 |
155.60 |
| SDNN |
4.29 |
23.04 |
1.66 |
2.72 |
10.23 |
3.00 |
7.28 |
3.34 |
| RMSSD |
3.29 |
2.41 |
0.90 |
0.39 |
2.67 |
0.74 |
2.68 |
1.12 |
| SDSD |
3.29 |
2.42 |
0.90 |
0.39 |
2.67 |
0.74 |
2.68 |
1.12 |
| CVNN |
0.02 |
0.12 |
0.01 |
0.02 |
0.05 |
0.02 |
0.05 |
0.02 |
| CVSD |
0.02 |
0.01 |
0.01 |
0.002 |
0.01 |
0.004 |
0.02 |
0.01 |
| MedianNN |
197.8 |
182.8 |
162.6 |
161.6 |
201.4 |
181 |
157.8 |
155 |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).