1. Introduction
In a large part of ecological niches, bacterial species are united in communities and develop together in the form of heterogeneous structures called biofilms, which are formed on surfaces of different nature and nature with the participation and as a result of the interaction of two or more bacterial species [
1]. At the basis of their formation and their resistance to the various effects of the external environment are the relationships between microorganisms, which include competition for nutrients from the composition of the medium for cultivating biofilms, antagonism and symbiosis. The structure of biofilms and the composition of the microbial community in them are determined by the fluctuation in the values of environmental factors [
2]. Changes in the values of environmental factors affect the physiological state of the cells in the structure of the forming biofilm. In the process of development, conditions are created for contacts between bacterial cells and contacts of cells with the surface on surfaces that are different in nature and morphology. Changes in environmental conditions have an impact on the properties of the bacterial cell, expressed in a change in the level of gene regulation for the implementation of the process of biofilm formation or in the charge of the cell surface, as well as on the physicochemical characteristics of the substrate [
3].
The growth and development of microbial biofilms is accompanied by the biosynthesis of exocellular polysaccharides, which ensure intercellular contacts in them and their attachment to the surface of the substrate, protecting microbial cells from the influence of adverse environmental factors - the formation and secretion of surfactants produced by other microbial species , which stimulate the transition from attached state to plankton, toxic compounds, oxidative and acid stress [
4]. The formation of biofilms and the composition of the matrix forming them depend on the characteristics of the growth medium and the bacterial surface, its course is affected by the pH-value and temperature of the environment [
5].
The pH-value of the environment affects the surface charge of the cells of the microbial species and the substrate, and the interaction between them in the process of biofilm formation is reversible and is based on the formation of hydrogen bonds and vander-Waals interactions. At pH 7.0, the bacterial cell and the surface layer of the substrate are distinguished by a different charge, which creates prerequisites for the adhesion of microbial cells on the surface and the formation of biofilms [
6]. Maximum cell adhesion of
E. coli O157:H7 strains to biotic surfaces was observed at a pH value of the medium of pH 7.0, which was reduced when the pH decreased to pH 5.5 [
7]. The expression of the ycfR gene in the biofilm, encoding the formation of the YcfR protein from the composition of the outer side of the cell wall, occurs when the pH value of the culture medium is changed from pH 8.0 to pH 5.5 [
8]. It includes in its structure the low molecular weight subunits YcfR, YahO, YbiJ, YbiM, YdgH, YhcN, YjfN, YjfO and YjfY [
8], and its formation inhibits the process of development of E. coli strains to biofilms as a result of the inhibition of intercellular contacts and the contact of cells with the surface layer of the substrate [
9], the increase in the level of secreted indole in the intercellular space [
4] and the reduction of cell hydrophobicity through the expression of other cell surface proteins such as lipoprotein OsmB [
10]. Deletion in the ycfR gene in
E. coli strains is accompanied by the expression of 30 genes that encode the biosynthesis of proteins from the structure of the periplasmic space and the outer layer of the cell wall, which leads to a change in the character of the bacterial cell surface [
11] and the increase in the level of cell aggregation and the biomass of the formed biofilms [
9]. The course of these regulatory processes during the formation of biofilms is also a result of the increase in the intracellular level of acetyl-phosphate from the change in pH, as a result of which conditions are created for the formation of type 1 pili and fringes, ensuring the adhesion of cells on the surface layer of the substrate and their development to the formation of biofilms [
12].
The regulation of acid stress in
B.subtilis strains in the process of their development into biofilms is carried out by extracytoplasmic σ-factors, the increase in whose activity creates conditions for increasing the resistance of their cells to the effects of antibiotic agents of a protein nature and other agents inhibiting cell wall biosynthesis [
13]. Their structure contains the subunits SigO and RsoA [
14], the formation of which is initiated by the decrease in the acidity value of the medium [
15], while the increase in pH to pH 7.0 leads to the activation of
rsiO gene responsible for the synthesis of the anti-sigma factor RsiO [
6]. RsiO binds to the N-terminal region of SigO and thus inhibits the transcription of genes encoding the biosynthesis of alkaline stress proteins [
14]. Its activation is the result of the formation of the RSiO-SigO-RsoA complex, which occurs under conditions of high acidity (pH 5.4) [
13]. The formation of the σ-factors σ
X and σ
W during the development of
B.subtilis strains on the surface layer of substrates of different nature and structure, which is stimulated by the decreasing pH value of the culture medium, leads to the activation of
abh gene responsible for the formation of the Abh regulatory protein, which positively regulates the expression of
eps and
yqxM operons by reducing the level of repression initiated by the regulator AbrB, carrying the information about the biosynthesis of exocellular polysaccharides and TasA protein from the matrix structure of biofilms. At the basis of this mechanism lie the processes of expression of
slrR gene, encoding the positive regulator for the transcription of
yqxM operon, and activation of the eps operon in the way of increasing the expression of
slrR gene for the transcriptional regulator SlrR, positively controlled by Abh [
16].
The biosynthesis of cytoplasmic σ-factors under conditions of high acidity is accompanied by the production of antibiotic compounds of a protein nature by strains of
B.subtilis [
17], and the regulation of its synthesis takes place with the participation of the regulatory proteins AbrB, Abh and Rok [
18]. Acid stress leads to an increase in the intracellular level of the phosphorylated form of the regulator Spo0A, which specifically binds to the promoter region of
abrB gene [
19], encoding the master regulator AbrB, which represses the formation of the antibacterial peptide toxins SdpC and factors of cannibalism Skf [
20]. Negative regulation on the activity of the promoter regions of
skfA and
sdpA genes is also carried out by the regulator Abh [
21], which positively affects the synthesis of suplancin [
22]. The operon for its formation during the development of
B.subtilis strains to biofilms is part of the SPβ prophage genome, including sunA, sunT, bdbA, yolJ and bdbB genes [
23,
24]. Suplancin, secreted by
B.subtilis strains in the process of biofilm formation, is characterized by an oligopeptide nature and includes in its structure β-methyllanthoid and disulfide bonds [
25], thanks to which it is distinguished by its antagonistic activity against gram -positive bacterial species such as
Bacillus cereus,
Staphylococcus aureus and
Streptococcus pyogenes [
26]. The structural
sunA gene encodes the biosynthesis of the starting peptide containing 56 amino acid residues [
24]. Transcription of the adjacent
sunT gene determines the activity of an ABC-type transport system, including a domain with proteolytic activity, which catalyzes the hydrolysis of the peptide structure of suplancin in the process of its transfer across the cytoplasmic membrane [
25]. The BdbB and BdbA domains possess a region with thiol oxidoreductase activity, thanks to which they take part in the post-translational regulation of coplancin synthesis by
B.subtilis strains during their development into biofilms [
26].
The expression of extracytoplasmic factors creates conditions when the pH of the medium decreases to increase the activity of the regulatory protein Abh, which positively affects the formation of suplancin by
B.subtilis strains during their development into biofilms [
27]. The transcription of
abh gene increases upon the activation of σ
X or σ
M, which leads to an increase in the amount of the synthesized antibiotic compound [
29] and ensures the competitive advantage of
B.subtilis strains in the spatial structures formed on the surface layer of the substrate at their association with other gram-positive and gram-negative bacterial species [
29]. The basis of their antagonistic effect, determined by the synthesis of suplancin, is the increase in the activity of
sunA operon, which limits the repressive influence of the negative regulator AbrB on the operon for the σ
W factor [
28]. The increase in its intracytoplasmic level ensures the resistance of the antibiotics of
B.subtilis strains in the process of biofilm formation against the effect of their secreted suplancin [
30], which inhibits the spread of cells of
S. epidermidis strains [
29],
E. coli [
26] and
S. aureus in the structure of biofilms [
29]. The induction of extracytoplasmic factors in
B.subtilis strains during the formation of biofilms, their role in the synthesis of the compound with antimicrobial effect in their association with other microbial species is carried out by different pathways and appears to be a mechanism by which they adapt to the unfavorable factors of environment. Their clarification necessitates the need to look for a relationship between the oxygen content and the formation of biofilms with the participation of two or more bacterial species. This circumstance is complemented by the peculiarities of the synthesis of the YcfR protein in
E. coli strains, which is induced by lowering the pH-value of the medium, increasing the temperature of cultivation of vegetative cells and biofilms, the formation of H
2O
2 as a result of the course of intracellular oxidative processes or in the presence of heavy metal ions [
30]. Most natural biofilms are polymicrobial in composition, but the mechanism remains unclear about how pH regulates polymicrobial biofilm development and the biofilm matrix component and spores in their structures.
The main objective of the present work is to study the effects of pH-value in the culture medium on the biofilm development and architecture of biofilms of the bacterium (B. subtilis) during their interactions with Escherichia coli K-12 1655 strain. Effects of pH-value in the culture medium on bacterial biofilm formation and the mechanisms were analyzed by the crystal violet staining method combined with cultivated microbial analysis, and confocal laser scanning microscopy. Taken together, the results of this study demonstrate a close link between biofilm formation and phosphates acquisition in B. subtilis and E.coli strains, allowing a better comprehension of how bacteria can cope with unfavorable pH-value under environmental conditions.
2. Material and Methods
2.1. Experimental Design
In the first part of this research, a dual-species model biofilm consisting of Bacillus subtilis and Escherichia coli was developed. In order to obtain a strongly adherent and mature model biofilm, different (incubation) conditions were altered, i.e., growth medium. The adherence of the biofilm at each of the different (incubation) conditions was quantified by means of crystal violet staining and subsequent optical density (OD) measurements. To determine the cell density/maturity of the biofilm, viable plate counts were used. General and selective media were applied to determine the total biofilm cell density and the contribution of each individual species to this total cell density.
2.2. Bacterial Strains and Culture Media
In this research, Bacillus subtilis and Escherichia coli, both acquired from the BCCM/LMG bacteria collection of NBIMCC, NCIPD and the “Stephan Angeloff”Institute of Microbiology in Sofia, were used. Stock-cultures were stored at −80°C in Luria Bertani Broth (LB, NCIPD, Sofia), which were both supplemented with 20 (v/v) % glycerol (NCIPD, Sofia). For every experiment, a purity plate was prepared by spreading a loopful of stock-culture onto a LB agar plate [Plate Count Agar (NCIPD, Sofia). The purity plates for Bacillus subtilis and Escherichia coli were incubated for 24 h at 37°C.
Starting from the purity plates, pre-cultures were prepared by transferring one colony into an Erlenmeyer flask containing 20 mL of LB medium (LB, NCIPD, Sofia). Bacillus subtilis and Escherichia coli pre-cultures were incubated for 24 h at 37°C. Following this incubation period, stationary phase cultures with a cell density of ~109 CFU/mL were obtained.
2.3. Biofilm Development Conditions
The stationary phase pre-cultures were used to develop a 100-fold diluted inoculum with a cell density of ~107 CFU/mL. The investigated pre-culture ratios (Bacillus subtilis and Escherichia coli) were 1:1 and the growth media was Luria Bertani Broth(NCIPD, Sofia), which proved to be the optimal media for single-species and multispecies biofilm development by Bacillus subtilis and Escherichia coli, respectively.
To develop the biofilms, 1.2 mL of the inoculum was transferred to a small Petri dish made out of polystyrene (50 mm diameter, 9 mm height, Simport, Canada). After inoculation, Petri dishes were closed and gently shaken to make sure the inoculum covered the entire surface. Dependent on the applied (incubation) conditions, Petri dishes were incubated for 24 h at 20°C, which were the optimal temperatures for Bacillus subtilis and Escherichia coli single-species and multispecies biofilm formation, respectively.
2.4. Crystal Violet Assay
Bacterial biofilms were developed into 96-well microtiter plates (Greiner Bio-One, Kremsmünster, Austria) with 100 μL of bacteria in post-exponential growth phase in VNSS per well. In parallel, 100 μL of cell-free culture supernatant of another strain were collected in VNSS at the beginning of the stationary growth phase and were added on the bacterial biofilms (the addition of VNSS alone was used as a control). The final OD600 nm was 0.4 into each well. After 24 h of growth in static conditions and a temperature of 20°C, samples were washed thrice with NaCl (36 g.l-1) and dried during 30 min at room temperature. Biofilms were stained during 15 min with 200 μL of Crystal Violet at 0.01% (w/v) and rinsed thrice with NaCl (36 g.l-1) and dried for 10 min. The quantification of biofilm was evaluated by releasing the stain from the biofilm with absolute ethanol for 10 min at 20°C, at 120 rpm and measuring the absorbance of the Crystal Violet solution at 595 nm. The final OD595 nm of each sample was divided by the blank (i.e., VNSS medium only treated with Cristal Violet).
2.5. Quantification of Colonies
Each specimen was individually placed in a centrifuge tube containing 4.5 mL of sterile physiological solution, and these tubes were vortexed for 1 min to detach the biofilms from the acrylic samples. After this, aliquots of 25 μL of serial dilutions (10−1 , 10−2 , 10−3 , and 10−4) were seeded in duplicate on Plate Count Agar (NCIPD, Sofia) and MacCocey Agar (NCIPD, Sofia) for the identification of Bacillus subtilis and Escherichia coli, respectively. Red and pink colonies grown on MacCocey Agar (NCIPD, Sofia) were presumptively identified as Escherichia coli. After incubation at 37 °C for 24 h, the colony-forming unit per milliliter (CFU/ml) was determined and log-transformed (log10).
2.6. Multispecies Biofilm Formation
For the dual-species biofilm, bacteria in post-exponential growth phase were suspended in ASW and inoculated in 24 well plates (Corning Incorporated Costar®, New York, NY, United States) to a final OD600 nm of 0.3 (0.15 per strain). For the biofilms involving two bacterial strains, the final OD600 nm was 0.3 or 0.4 (0.1 per strain). Controls included single species biofilms formed in the same concentrations and conditions than the multispecies biofilm. To study the influence of the pH value of the medium on biofilm growth, nutrient media were prepared with different ratios of KH2PO4 and K2HPO4 so that the pH values were 5.0; 6.0; 7.0; 8.0. For the purposes of the task, 18-hour cultures of B.subtilis 170, B.subtilis 168 and E.coli k-12 1655 strains were prepared in advance in a medium broth (BulBio Laboratory - Sofia). 50 μl of the liquid cultures were inoculated into 5 cm3 liquid medium M63 (0.02 M KH2PO4, 0.04 M K2HPO4, 0.02 M (NH4)2SO4, 0.1 mM MgSO4 and 0.04 M glucose) (pH 7.0) and M63 at different ratio of KH2PO4 and K2HPO4 so that the pH of the medium is pH 5.0, pH 6.0, and pH 8.0. For each factor, the experiment was carried out in five test tubes, in the first one only 50 μl of the liquid culture of B.subtilis 170 strain was seeded, in the second - B.subtilis 168, in the third - E.coli K-12 1655, in the fourth tube inoculate 50 μl of the liquid culture of B.subtilis 170 strain and 50 μl of the liquid culture of E.coli K-12 1655 strain, in the fifth place inoculate 50 μl of the liquid culture of B.subtilis 168 strain and 50 μl of the liquid culture culture of E.coli K-12 1655 strain. Fresh cultures are distributed in 96-well plates. Each fresh culture was dispensed into 12 wells, placing 150 μl of the liquid fresh culture in each well. 150 μl of distilled water is dripped into the final unseeded wells. The first plate is placed at a temperature of 20 °C. Cultivation of the biofilms on the plates was carried out for a duration of 24 h. After that, the plankton was separated from each well and washed three times with saline (0.85% NaCl), 150 μl of saline was placed in half of the wells, and the biofilm was peeled off using a knife previously burned and cooled in sterile saline of them, and for each variant of the experiment, the suspension is collected from 6 wells in one eppendorf.
2.7. Matrix Components Staining
For the matrix staining, a static biofilm of 48 h was performed. Each biofilm was stained with DAPI at 5 μg/mL (Sigma-Aldrich, Darmstadt, Germany) and one of the following matrix dyes. Exopolysaccharides were stained with the Wheat Germ Agglutinin (WGA) associated with the Alexa FluorTM 555 conjugate (Thermo Fisher Scientific, Waltham, MA, United States) at 100 μg/mL to label N-acetyl-glucosamine. After 30 min of incubation of each probe, each coverslip was washed 3 times in PBS 1×. Finally, the coverslips were mounted with a drop of ProlongTM Diamond Antifade before observation with confocal laser scanning microscopy Leica TCS SPE at wavelength of 540 nm.
2.8. Data Extraction from Images and Statistics
At least three replicates and five pictures per replicate were performed and used for data extraction. The pictures have been acquired by epifluorescence microscopy or by CLSM. The percentages of recovery of epifluorescence microscopy were determined using an algorithmic method with RStudio 0.98.1025 (RStudio, Boston, MA, United States), where the brightness pixel was determined by threshold’s definition of small area of picture around the pixel and against the background. Each picture has been divided in 36 pieces, that permitted empirically to render negligible the distortion of objective and the threshold was defined as two multiplied by percentile 5th of pixel values on a small area in which the pixel was measured (Supplementary Figure S2). The biovolume, the average thickness and the evaluation of the maximum coverage in the CLSM pictures was determined with the COMSTAT software developed in MATLAB R2015a (MathWorks, Natick, MA, United States) as previously performed [
31].
To test for statistically significant differences (P < 0.05) between two conditions a t-test was performed and between different time points, a two-way analysis of variance including the Bonferroni post-test were performed using SPSS 13.0 (IBM, Armonk, NY, United States).
3. Results and Discussion
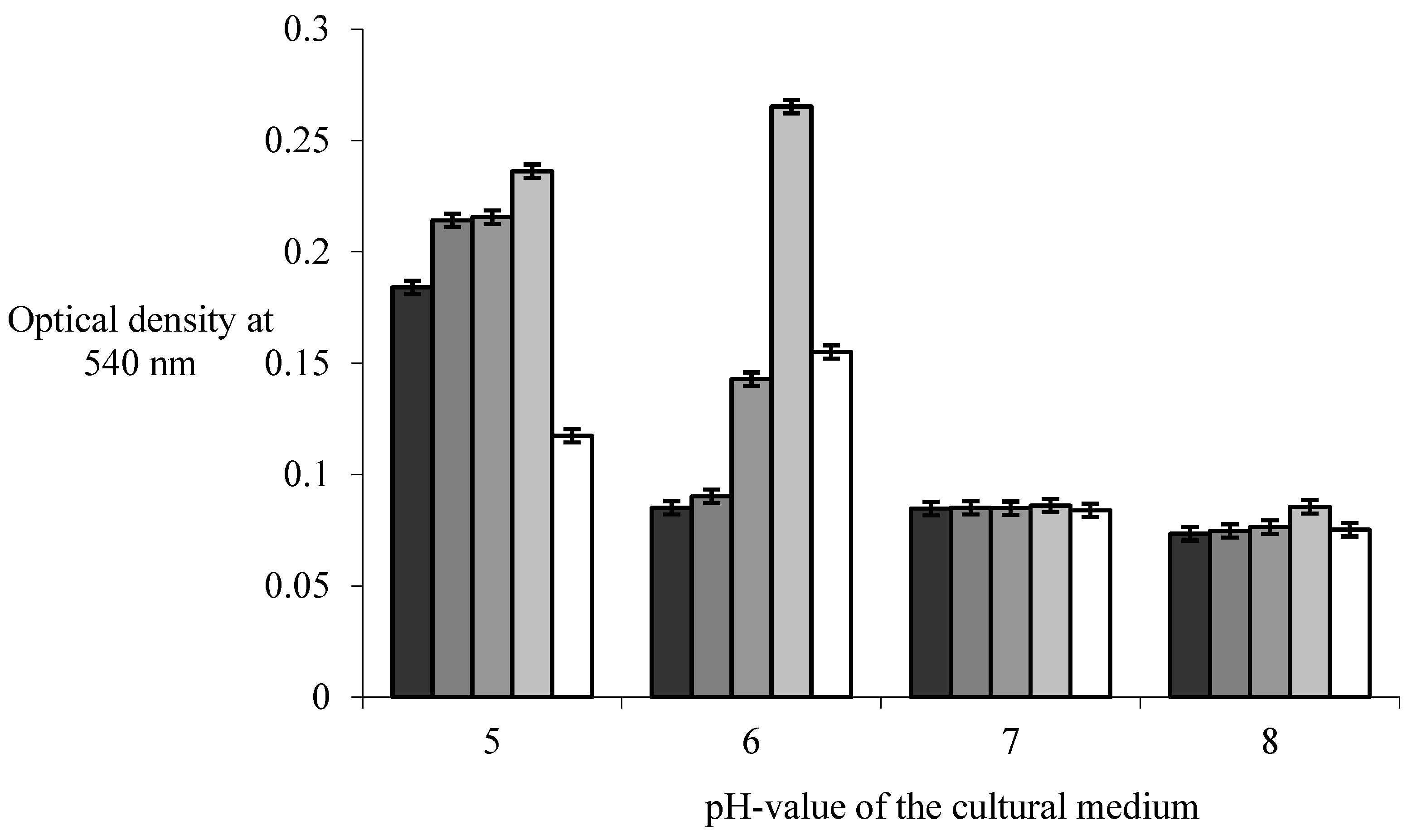
The increase in the pH value of the medium between pH 5.0 and pH 6.0 in the conducted study did not create conditions for a statically significant increase in the optical density at 540 nm (p>0.05) in the co-cultivation of
B. subtilis and
E. coli strains. Increasing its value to pH 8.0 leads to a statistically significant decrease in the biomass of mixed biofilms (p<0.05). A maximum optical density value of 0.236±0.11 for the co-culture of
B. subtilis 170 and
E. coli K-12 1655 strains and 0.155±0.63 for the co-cultivation of
B. subtilis 168 and
E. coli K-12 1655 strains was reached at pH value of the medium pH 6 ,0, which significantly (p<0.05) exceeds their indicators in monospecies biofilms of
B. subtilis and
E. coli strains (
Figure 1).
A number of extra cytoplasmic σ-factors take part in the regulation mechanism of the formation of biofilms by
B. subtilis strains under conditions of changing pH-value of the medium, which affect the activity of a number of gene regions encoding resistance to the effects of antibiotic substances [
24,
26], the low acidity of the environment [
8], regulation of the biosynthesis of the extracellular polymers that make up the matrix [
16]. The increase in the activity of the σm-factor in
B. subtilis 168 strain is activated at low acidity of the medium [
19,
21]. Its activation plays an important role in the development of strains to form biofilms [
24]. Deletion in the σ
m-factor gene in
B. subtilis 168 strain led to a decrease in biofilm biomass in the study by Nagorska et al. [
32]. These results confirm the role of the σ-factor in the process of biofilm formation at low acidity [
24,
26], which can explain the exponential decrease in the optical density value at 540 after crystal violet staining of the biofilms of
B. subtilis 170,
B. subtilis 168 strains and in the process of their association with
E .coli K-12 1655 strain when changing the pH value of the culture medium in the range from pH 6.0 to pH 8.0. The higher value of optical density of biofilms of
B. subtilis 168 strain and in its association with
E. coli K-12 1655 strain compared to
B. subtilis 170 strain was associated with the different expression level of genes for σ- and anti -σ factor in the process of their cultivation at pH 5.0-6.0.
The low pH value of the biofilm culture medium of pH 4.0-5.0 inhibits the process of indole formation [
5], increasing its value to pH 9.0 induces the expression of
tnaA gene responsible for the biosynthesis of indole [
5,
8], which reduced the biomass of biofilms of
E. coli BW25113 strains [
24]. The optical density at 540 nm after staining the biofilms of
E. coli K-12 1655 strain was distinguished by a value of 0.215±0.09, which significantly decreased to a level of 0.085 , when the pH of the culture medium was changed from pH 5.0 to pH 8.0 in the present study and confirms the results of a study of Leo et al. [
24]. The formation of indole by
E. coli strains was initiated by the presence of antibiotics in the medium [
4], which in the present study appeared to be the result of the development of
B. subtilis strains. In other studies, the accumulation of indole creates conditions for inhibiting the process of biofilm formation by inhibiting cell motility [
24] and inhibiting chemotaxic movement [
24,
25,
26].
The pH value of the culture medium affected the nature of the relationships between
B.subtilis 170 and
E.coli K-12 1655,
B.subtilis 168 and
E.coli K-12 1655 strains in the process of their co-development into biofilms in the study conducted . At pH 5.0, the microbial population of the mixed biofilm structure was dominated by
B.subtilis 170 strains with a number reached of 9.55±0.13.10
6 cfu/cm
3, while for
B.subtilis 168strain the value was 6.38±0. 29.10
6 cfu/cm
3 and statistically does not differ from their value in monospecies biofilms (p>0.05). A similar regularity between the number of colonies in the structure of monospecies and mixed biofilms was found in
E.coli K-12 1655 strain, whose population number was lower compared to
B.subtilis strains. The pH-value of the medium in the range of pH 5.0 to pH 6.0 ensures the presence of cells of
E. coli K-121655 strains in the population structure of biofilms in their association with
B. subtilis 170 and
B. subtilis 168 strains, as their numbers decreased from 3.16±0.12.10
6 cfu/cm
3 to 0.40±0.05.10
6 cfu/cm
3 in the structure of mixed biofilms when co-cultivated with
B. subtilis 170 strain, as well as from 5.21±0.22.10
6 cfu /cm
3 to 0.63±0.04.10
6 cfu/cm
3 in their interaction with
B. subtilis 168 strain. Their values are higher than their numbers in the structure of monospecies biofilms of
E.coli K-12 1655 strain (
Table 1 and
Table 2).
The biosynthesis of surfactin by
B. subtilis BS5 strain is favored by an increase in the pH value of the culture medium in the range of pH 5.0 to pH 6.8 according to the study of Abdel-Mawgoud et al. [
33], which corresponds to the significant increase in the number of colonies of
B.subtilis 170 and
B.subtilis 168 strains in their monospecies culture and in their interaction with
E.coli K-12 1655 strain during the formation of biofilms in the present study.
B.subtilis strains are distinguished by their antagonistic activity in their association with other microbial species in the rhizosphere layer of different crops thanks to their ability to synthesize surface-active agents of a lipoprotein nature that specifically inhibit cell motility and colonization of the cells of the companion species on the surface of the substrate [
19,
26,
27]. The ability to secrete surfactants with anti-adhesion actions underlies the antagonistic activity of
B.subtilis strains against
Pseudomonas syringae strains in the rhizosphere layer of tomato plants [
19].
At higher pH values of the culture medium above pH 6.0, conditions are created for the occurrence of antagonistic relationships, as a result of which biofilms are formed only with the participation of cells of
B. subtilis strains. However, a significant decrease in population numbers of
B.subtilis 170 and
B.subtilis 168 strains was reported, which was found to values of 1.48±0.07.10
6 cfu/cm
3 and 6.18±0.20.10
6 cfu/cm
3 at reaching the pH-value of the environment of pH 8.0, which exceeds their value in monospecies biofilms, where a similar regularity is established in the investigated pH-interval. Media pH above 7.0 inhibits the formation and secretion of surfactin in
B. subtilis strain [
33], which appears to be a signaling molecule during its development in a surface-attached state [
26] and contributes to the maturation of the formed structures [
1]. Inhibition of surfactin synthesis at pH in the range of pH 7.0 to pH 8.0 explains the decrease in population numbers of
B.subtilis 170 and
B.subtilis 168 strains in the structure of mixed biofilms in their interaction with
E.coli K -12 1655 strain in the present study.
Indole, formed by
E. coli strains in its association with other bacterial species, represses the formation of biofilms [
24], which is determined by its binding to SdiA, which is distinguished by the property of forming complexes with homoserine lactones [
8,
24], and exerts a stimulating effect on the formation of biofilms by
Pseudomonas strains, which are characterized by oxygenase activity, allowing the reduction of its concentration in the process of biofilm formation [
24]. An increase in the activity of SdiA is accompanied by a decrease in the transcription level of
ftsQ2p gene, responsible for the acid resistance of
E.coli strains [
24,
25], which explains the lower number of their population in the structure of biofilms compared to
E. coli K-12 1655 strains, when changing the pH in the range of pH 5.0 to pH 6.0.
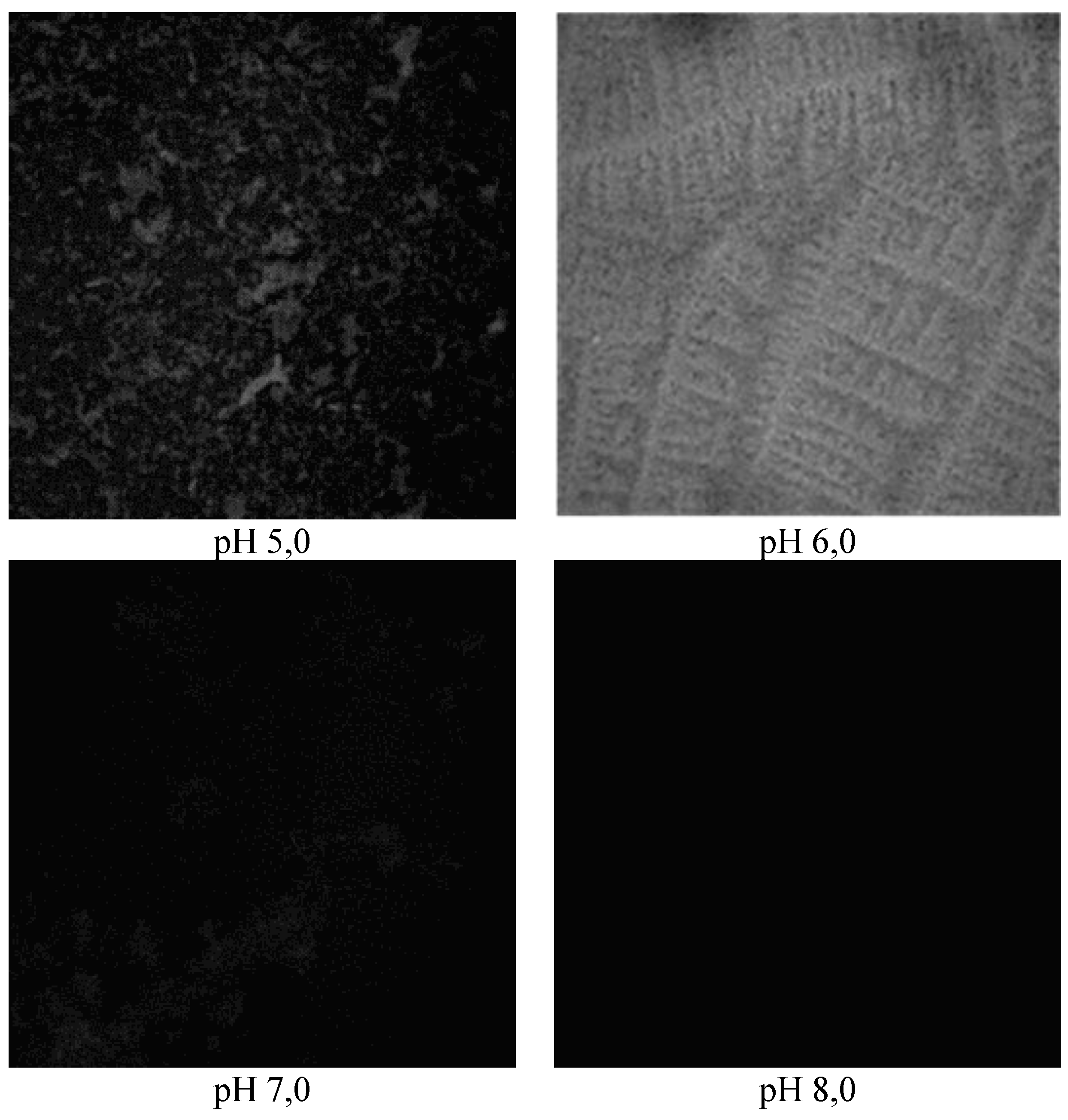
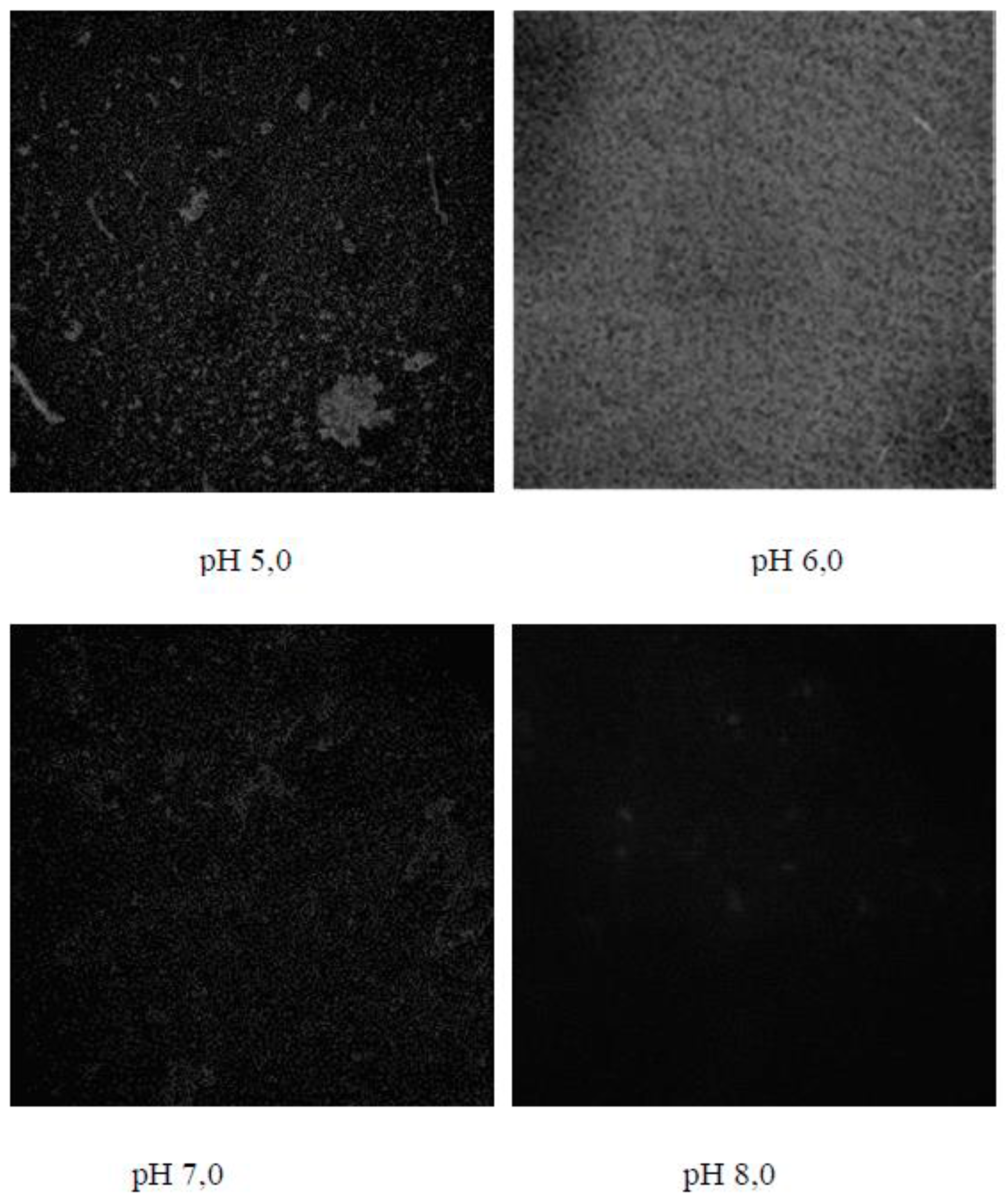
In acidic media (pH 5.0),
B. subtilis and
E. coli strains formed biofilms with a rough surface with an average thickness of 8.14±0.48 μm to 8.46±0.37 μm at a roughness factor of 0, 04±0.03 to 0.03±0.01, which increased to a value of 10.35±0.41 μm upon reaching the pH-value of the medium of pH 6.0 in the co-cultivation of the pair of
B. subtilis 170 and
E. coli K-12 1655 strains, and the inequality coefficient kept a relatively constant value (p<0.05) (
Figure 2 and
Figure 3). The average thickness of the structures formed as a result of the development of the pair of
B. subtilis 170 and
E. coli K-12 1655 strains is characterized by a value of 10.78±0.37 μm under the same cultivation conditions. In neutral and alkaline media, the relative proportion of biofilm dispersion zones increases, which correlates with a decrease in the diameter of the structures in the narrow range of 3.02±0.23 μm and 3.01±0.05 μm in the pair of
B. subtilis 170 and
E. coli K-12 1655 strains, while in
B. subtilis 168 and
E. coli K-12 1655 strains single structures with the same size of the average thickness were found at the pH of the medium in the range of pH 7.0 to pH 8 .0 in the conducted survey. These results positively correlate with optical density values at 570 nm after crystal violet staining and with the change in population numbers of
B. subtilis strains in the structure of mixed biofilms, which changes inversely with the magnitude of the ratio of the area of the structures formed to their volume during the conducted survey. It reached a minimum value of 0.09±0.006 μm
2.μm
-3 for the pair of
B. subtilis 170 and
E. coli K-12 1655 strains and 0.09±0.009 μm
2.μm
-3 for
B. subtilis 168 and
E. coli K-12 1655 strains at a pH of 6.0 of the biofilm culture medium, immediately after which an increase in its size was observed as a result of the change in the pH value to pH 7.0, which remained at the same level at pH 8 ,0 (
Table 3).
An important feature in the formation of biofilms, involving cells of
B.subtilis strains, is the biosynthesis of the exocellular matrix [
34]the formation of which is encoded by
epsA-O and
tapA-sipW-tasA operons [
7,
35]. Their transcription is determined by the activity of the regulatory proteins AbrB and Abh, whose activity is determined by the intracellular expression level of σ-factors [
8], which increases under conditions of alkaline stress [
24]. The Abh regulatory protein positively affects the biosynthesis of the biofilm matrix, and the activation of the transcription of the abh gene encoding it occurs as a result of the interaction of the σ-factor with the RNA polymerase to activate the transcription of
abh gene [
25]. The transcription of the operons is not carried out directly by the regulatory protein, but with the help of the intermediary molecule RsiW, which by its nature is a protease and performs the role of an anti-sigma factor [
27], as a similar regulatory mechanism is also observed in
E.coli strains under the conditions of alkaline stress with the participation of the anti-sigma factor RseP [
19]. The occurrence of this genetically complex regulatory mechanism explains the decrease in the average thickness and relative volume of the structures formed as a result of the co-development of
B. subtilis 170 and
E. coli K-12 1655,
B. subtilis 168 and
E. coli K- 12 1655 strains when changing the pH value of the culture medium in the range of pH 6.0 to pH 8.0 in the present study. The reduction of the relative volume per unit area of the formed structures is the result of the inhibition of the synthesis of the exocellar matrix, which determines the hydrophobic nature of the biofilms and ensures their adhesion to the surface of the substrate. The established regularity is in agreement with the results of the study by Helman et al. [
27], according to which mutations in
yoaW and
sipW genes, responsible for the biosynthesis of the exocellular proteins of the biofilm matrix composition and negatively regulated by the regulator AbrB, do not lead to a change in the adhesion properties of cells of
B. subtilis 170 strain in comparison with the starting strain. These strains form biofilms with a thickness of 8.0±0.5 μm, while in the original strain this value is of the order of 14±0.7 μm [
27]. A similar mechanism of reduction of the thickness of the formed structures was observed in the development of the
wcaF mutant
Escherichia coli strain in the attached state [Danese et al., 2000]. Between
B. subtilis and
E. coli strains there is a similarity in terms of inhibition of the process of biofilm formation, expressed in suppression of the biosynthesis of exocellular polysaccharides when the pH-value of the medium is increased [
27].
The relative spreading area of the mixed biofilms formed by the co-cultivation of the pair of B. subtilis 170 and E. coli K-12 1655, B. subtilis 168 and E. coli K-12 1655 strains increases with a change in the pH value of the medium for cultivation from pH 5.0 to pH 6.0, immediately after which a strong decrease occurs when pH 7.0 is reached (p<0.05). Its value at pH 6.0 is 0.99±0.12 μm2 for the biofilms of the pair of B. subtilis 170 and E. coli K-12 1655 strains and 0.99±0.00 μm2 for the pair of B. subtilis 168 and E. coli K-12 1655 strains and was approximately fourfold higher as the pH of the medium increased to pH 7.0. Reaching its value to pH 8.0 is accompanied by a statistically insignificant decrease in the size of the relative area of spread of the formed structures of 0.15±0.02 μm2 in the dual-species biofilms of the co-culture of B. subtilis 170 and E. coli K-12 1655 strains, while in B. subtilis 168 and E. coli K-12 1655 strains it keeps a constant value.
The main function of the regulatory protein AbrB is reduced to the inhibition of
sipW gene, responsible for the biosynthesis of the signal peptidase SipW, whose role in the process of biofilm formation is reduced to the secretion of proteins to the cell surface, which contributes to the adhesion and spreading of cells on the surface layer of the substrate [
27] and can explain the decrease in the value of the relative spreading area of the formed biofilms as a result of the co-cultivation of the co-culture of
B. subtilis 170 and
E. coli K-12 1655,
B. subtilis 168 and
E. coli K-12 1655 strains, which was linear as the pH of the medium changed from pH 6.0 to pH 8.0 in the study. According to other studies, peptidase SipW plays an important role in the formation of adhesin of peptide nature, which ensures intercellular contacts and the biosynthesis of structures for cell motility in B. subtilis strains [
19]. Inhibition of its biosynthesis explains the correlation found in the present study between the change in optical density at 540 nm after crystal violet staining of the formed mixed biofilms in the co-cultivation of the pair of
B. subtilis 170 and
E. coli K-12 1655,
B. subtilis 168 and
E. coli K-12 1655 strains, their average thickness and relative volume per unit area. According to other studies, regulatory proteins take part in the process of cell differentiation until the development of competence associated with the acceptance of exogenous DNA from the lysis of part of the cell population and their germination into spores [
27].
The obtained results for the morphological characteristics of biofilms when the pH-value of the culture medium is changed do not correlate with the number of spores in their structure. The number of spores with the structure of multispecies biofilms of B.subtilis and E.coli strains remains relatively constant when the pH changes in the range from pH 5.0 to pH 6.0, while under the same cultivation conditions a statically insignificant increase in numbers is observed them from 0.2.103 cfu/cm3 to 0.7.103 cfu/cm3 in the co-culture of B. subtilis and E. coli strains. The increase in its value is accompanied by a significant increase in the number of spores in the structure of the mixed biofilms during the course of the study (p<0.05), which at the pH of the medium of pH 8.0 reaches a value of 2.0.103 cfu/cm3 in the biofilms of B.subtilis 170 and E.coli K-12 1655 strains and 2.5.103 cfu/cm3 in the co-culture of B. subtilis 168 and E.coli K-12 1655 strains. The number of spores changed inversely proportional to the biomass of the structures formed in the conducted study, and statistically significant differences were found in the number of spores in the structure of monospecies and mixed biofilms in the present study (p<0.05).
The results of the dependence of the number of spores in the structure of biofilms in the present study, formed with the participation of
B. subtilis strains, depending on the pH value of the culture medium confirm the studies of Illades-Aguiar and Setlow [
36], in which low pH ensured vegetative growth of the studied strains and inhibited sporulation. The indole formed by
E. coli K-12 1655 strain in the process of its co-cultivation with
B. subtilis 170 and
B. subtilis168 strains at pH - the value of the medium in the range from pH 7.0 to pH 8.0 has an inducing effect on the formation of spores in the structure of biofilms [
24]. The regulatory protein AbrB negatively affects the expression of genes for the formation of the biofilm matrix, and the gene encoding it is activated by a high intracellular level of the core regulatory protein SpoOA, which is formed when the pH of the culture medium increases. The course of this regulatory mechanism leads to the activation of
abrB gene, which is accompanied by an increase in the activity of
spoIIE,
spoIIE and
spoIIG genes encoding the processes occurring during the second stage of ferrospore formation in
B. subtilis strains [
22,
23]. Mutations in the genes encoding the σ-factors of the mother cell and for ferrospore formation did not significantly affect the biofilm-forming ability compared to the original or wild-type strains, suggesting that spore formation is not an obligatory mechanism for biofilm formation when the factors are changed of the environment [
22], which explains the lack of correlation between the optical density at 540 nm of monospecies and mixed biofilms and the number of spores in them when the pH value of the culture medium is changed in the range of pH 5, 0 to pH 8.0 in the present study.
Table 4.
Morphometric features of dual-species biofilms of Bacillus subtilis 170 and Escherichia coli K-12 1655, Bacillus subtilis 168 and Escherichia coli K-12 1655 strains, depending on pH-value of the cultural medium.
Table 4.
Morphometric features of dual-species biofilms of Bacillus subtilis 170 and Escherichia coli K-12 1655, Bacillus subtilis 168 and Escherichia coli K-12 1655 strains, depending on pH-value of the cultural medium.
| рН |
Mean thickness, μm* |
Coefficient of unevenness |
Relative area, μm2**
|
Relationship area/volume, μm2.μm-3***
|
| Dual-species biofilms of Bacillus subtilis 170 and Escherichia coli K-12 1655 strains |
| 5,0 |
8,14±0,48 |
0,04±0,03 |
0,65±0,01 |
0,12±0,007 |
| 6,0 |
10,35±0,41 |
0,02±0,06 |
0,99±0,12 |
0,09±0,006 |
| 7,0 |
3,02±0,23 |
0,05±0,02 |
0,23±0,01 |
0,33±0,002 |
| 8,0 |
3,01±0,05 |
0,06±0,04 |
0,15±0,02 |
0,34±0,001 |
| Dual-species biofilms of Bacillus subtilis 168 and Escherichia coli K-12 1655 strains |
| 5,0 |
8,46±0,37 |
0,03±0,01 |
0,70±0,01 |
0,11±0,004 |
| 6,0 |
10,78±0,37 |
0,03±0,01 |
0,99±0,00 |
0,09±0,009 |
| 7,0 |
3,39±0,48 |
0,06±0,07 |
0,23±0,01 |
0,30±0,004 |
| 8,0 |
3,39±0,48 |
0,06±0,07 |
0,23±0,01 |
0,30±0,004 |
|
|
|
|
|