1. Introduction
By shipping away bioactive compounds, Gram-negative bacteria are able to communicate, relieve stress, and attack foreign entities [
1,
2]. Proteins, nucleic acids, and compounds found in the periplasm are packaged into spheres made of their own outer membrane (OM) material. Their cargo, protected during travel by the lipid bilayer, can reach host organisms more reliably than compounds secreted directly to the environment. Proteins displayed on the OM can become a part of outer membrane vesicles (OMVs) and provoke adherence with other organisms [
3]. Manipulation of membrane proteins and OMV cargo can be leveraged to target selected organisms and inject a desired bioactive material [
4,
5].
While the effectiveness of widely used antibiotics is declining, naturally occurring phenomena continue to inspire antibiotic therapies, alternative antibiotic sources, and drug delivery [
1].
Pseudomonas aeruginosa produces OMVs that have bacterial toxicity. They attack other bacteria using a periplasmic autolysin encapsulated in OMVs [
6].
Lisobacter spp. have also shown similar activity with bioactive compounds present in its OMVs cargo [
7]. Other gram-negative bacteria, such as
Escherichia coli have a high OMVs production potential [
8]. This presents an opportunity to use
E. coli as a platform for the production of OMVs for drug delivery, the development of novel vaccines [
9], or to emulate an OMVs-based antibacterial system.
Several factors affect the production of OMVs. Mechanisms affecting their biogenesis are related to deformations in the OM including: 1.) disrupted links between the inner membrane, the peptidoglycan layer, and the outer membrane; 2.) accumulation of periplasmic proteins; and 3.) curvatures induced by OM proteins [
2,
3,
9,
10]. Kulp et al. suggest that vesicle production is affected by structural characteristics of the periplasmic membrane while metabolic pathways for the production of membrane proteins, lipopolysaccharides (LPS), enterobacterial common antigen (ECA), and stress response pathways also affect their formation [
11]. However, the metabolic pathways involved in the production of OMVs have yet to be elucidated. Metabolic network models connect enzymatic reactions predicted from the genome to describe the metabolic states and biological capabilities of an organism. Finding what compounds are needed by the cells to achieve a certain state, growth rate, or compound production can help accomplish a greater in vivo productivity. Further exploring the effect of gene editing in the metabolic space can increase our understanding of OMV formation and improve its yield.
Being able to produce OMVs in large quantities is an important target. Additionally, the activity and stability of OMVs are important when considering their therapeutic use [
12]. Proteins in the cargo and displayed on the membrane affect their bioactivity [
3]. Toxic LPS in the outer membrane can also affect their medical use [
13].The OMVs proteome analyses of various Gram-negative bacteria, including
E. coli, have been reported and lipidomic profiles for
Klebsiella pneumoniae OMVs [
14,
15,
16,
17] have given an insight into their composition. However, there are currently no lipidomic profiles reported for
E. coli OMVs.
In this study, we adapted a genome-scale metabolic network model for E. coli K12 strain and employed constraint-based optimization methods to develop strategies that enhance OMVs production in E. coli JC8031. Description of the vesicle production was achieved through the model constraints and the objective functions driving the optimization problem. We found four E. coli JC8031 mutant strains with enhanced vesicle production through in silico gene knockouts. To our knowledge, this is the first use of metabolic network models to study OMVs production and the first report on the lipid composition of isolated E. coli OMVs.
3. Discussion
Deletion analysis performed on the refined model showed that knock-outs enhance vesicles production as modeled by the objective function. Interestingly, the gene deletions suggested by the OptKnock algorithm are not directly related to phospholipid metabolism. However, two genes have a direct relation to the outer membrane (
poxB,
gmhA).
poxB encodes for a membrane protein that acts as a pyruvate dehydrogenase. This protein is related to membrane binding and presents interactions with lipids [
26].
gmhA encodes for a phosphoheptose isomerase involved in the lipopolysaccharide biosynthesis and Δ
gmhA have shown lipopolysaccharides lacking heptose [
27]. The involvement of these genes with the formation of OM components supports previously proposed mechanisms of vesicle biogenesis through perturbation of membrane proteins, induced curvatures, and membrane integrity [
3]. However, the results show a lack of significance in a Δ
gmhA alteration over the protein concentration. The other two deleted genes
(allD,
sgbE) showed an indirect relation to vesicle production.
sgbE encodes for an enzyme from an L-ascorbate utilization operon and is involved in the metabolism of nucleotide and aromatic amino acid precursors [
28].
allD gene encodes for an enzyme involved in the assimilation of allantoin and allantoin use as a nitrogen source [
29]. These single deletion targets are also indirectly related to vesicle production through the alteration of biomass growth. The use of the Warburg-Christian equation through spectrophotometric methods shows limitation to quantify vesicle production. This method can be complemented with LPS measurements through the Purpald test [
23].
Results from in silico deletion experiments showed that there is a high biomass growth rate, multiple deletions did not reduce growth rate to an extent where the designed strain is no longer valuable. In silico vs. in vitro differences could be observed experimentally through deletion suggested genes. This implies a challenge in generating multiple subsequent deletions and achieving the expected biomass growth. Here, we found that deletions associated to a reduction on cellular growth also showed a reduced yield for vesicle recovery.
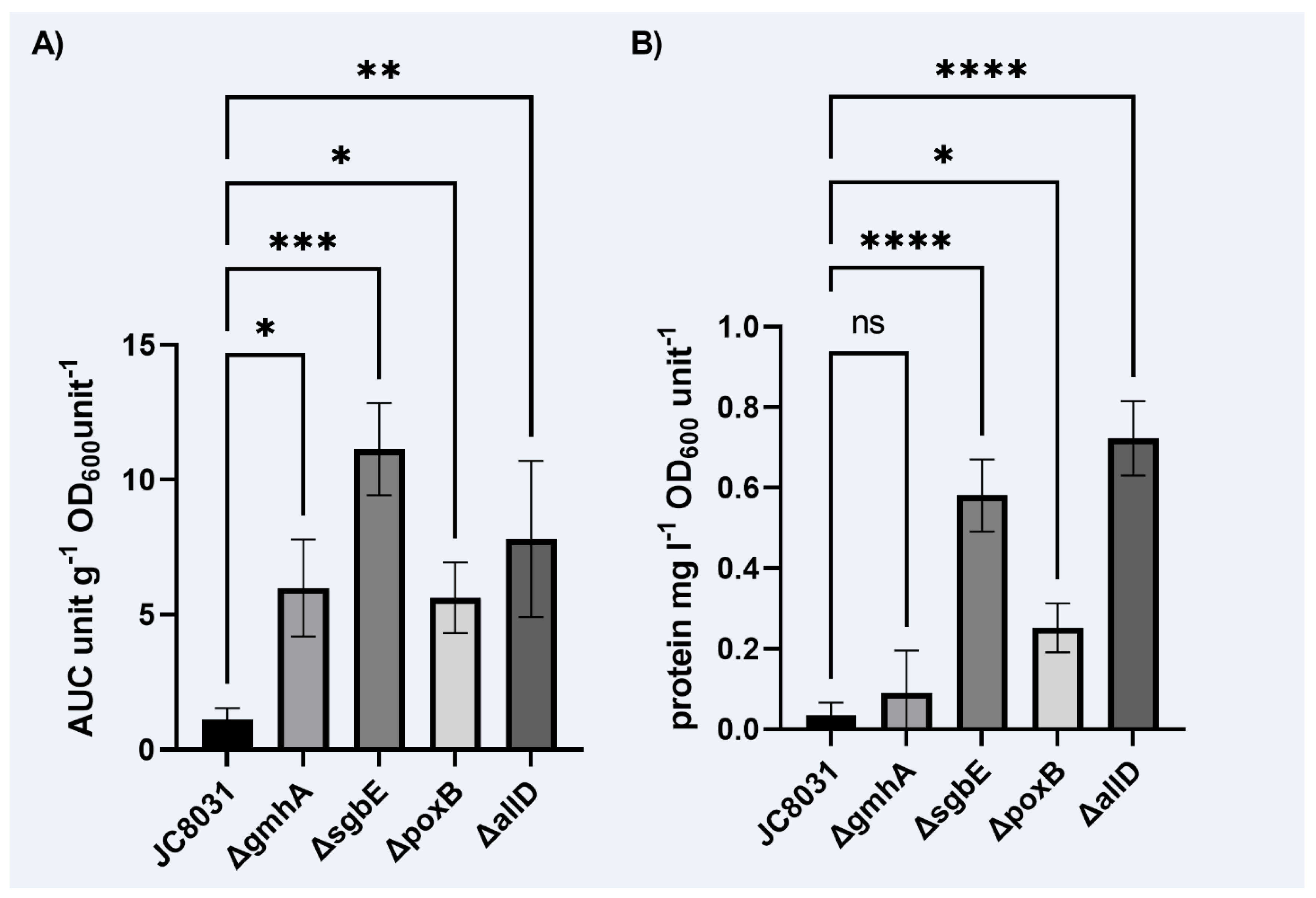
On the other hand, from the industrial point of view biomass growth was not significantly affected for all mutants while OMVs generation was enhanced up to 10 times in the single deletion mutant strain ΔsgbE, therefore more attractive when scaling up the process from an economical perspective.
The composition of the produced OMVs is also important when considering their use in therapeutic applications. Previous studies have raised concern on the effect of the recovery method on the size distribution, morphology and composition of OMVs. However, a comparison of isolation methods has shown that different methods produce a similar effect on vesicle morphology [
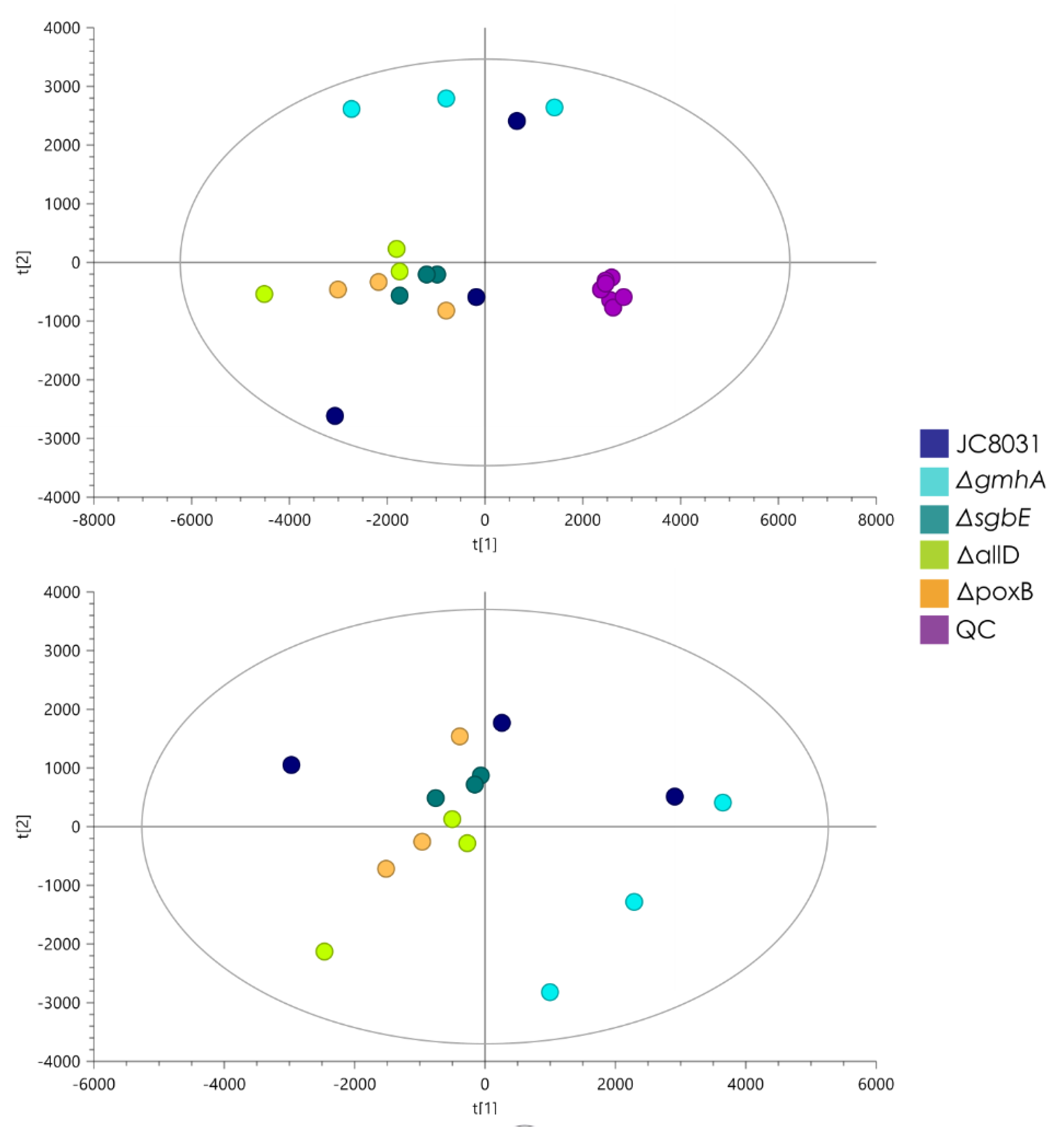
5]. The lipidomic analysis showed no statistically significant change in composition that separated the single deletion mutants from the JC8031 strain. This gives an insight into the response of general OMVs composition with respect to genome alteration. The lack of change in the lipid profile could be attractive for biomedical applications as it facilitates the study and analysis of material stability of OMVs as envelopes for the desired cargo and the effect of external factors on their structural integrity. Compared to detergent
To the best of our knowledge, the studies reporting OMVs composition are relatively limited and they are mostly focused on proteomics analysis rather than lipidomics. Besides the information that can be obtained from searching in academic databases, it is also possible to consult specialized compendiums available online, such as vesiclepedia [
30]. A search in this compendium can also show how information of bacterial OMVs lipidomics is very limited and suggests that the data obtained during the present study may be one of few attempts to characterize
E. coli OMVs from the lipidomics point of view (See Additional file 3).
On the other hand, regardless of the relative lack of information, it is possible to make further analysis of the data if assuming that OMVs lipid composition is similar to the
E. coli outer membrane composition, as suggested by previous studies [
31]. In that case, special attention should be paid to compounds like PGs and PEs, as these are reported to be the prevalent lipid components of the
E. coli outer membrane [
20,
25], particularly for the K12 strain, which is the strain from which the JC8031 strain was derived [
19]. Indeed, PGs and PEs were detected in our OMVs samples although no significant changes in lipid composition can be inferred from the results when comparing OMVs from the designed single deletion mutants against OMVs from the JC8031 strain. This result is consistent with a previous study where all possible single gene deletions, in
E. coli, were analyzed with respect to their effect on the membranes lipid composition [
32]; none of the genes that were deleted during the present study (
poxB,
sgbE,
gmhA, and
allD) were reported previously as responsible of causing a significant change in the lipid composition of the bacterium and it might be assumed that they should not cause significant changes in OMVs lipid composition either.
Another important aspect to take into account is about the prevalent fatty acyl chains forming part of the OMVs. In our case, the prevalent species are vaccenic acid (18:1) and palmitic acid (16:0) (Additional file 3), both of them previously reported as prevalent in
E. coli membranes [
25]. In this last report, it was also stressed that palmitic acid was slightly more prevalent in the outer membrane. Our results show that palmitic acid is indeed slightly more prevalent when analyzed in the PEs context, but a more quantitative analysis is necessary to obtain conclusive data. This kind of analysis will be particularly important if considering that OMVs are expected to be relatively rigid, thanks to a higher content of saturated fatty acyl chains [
31].
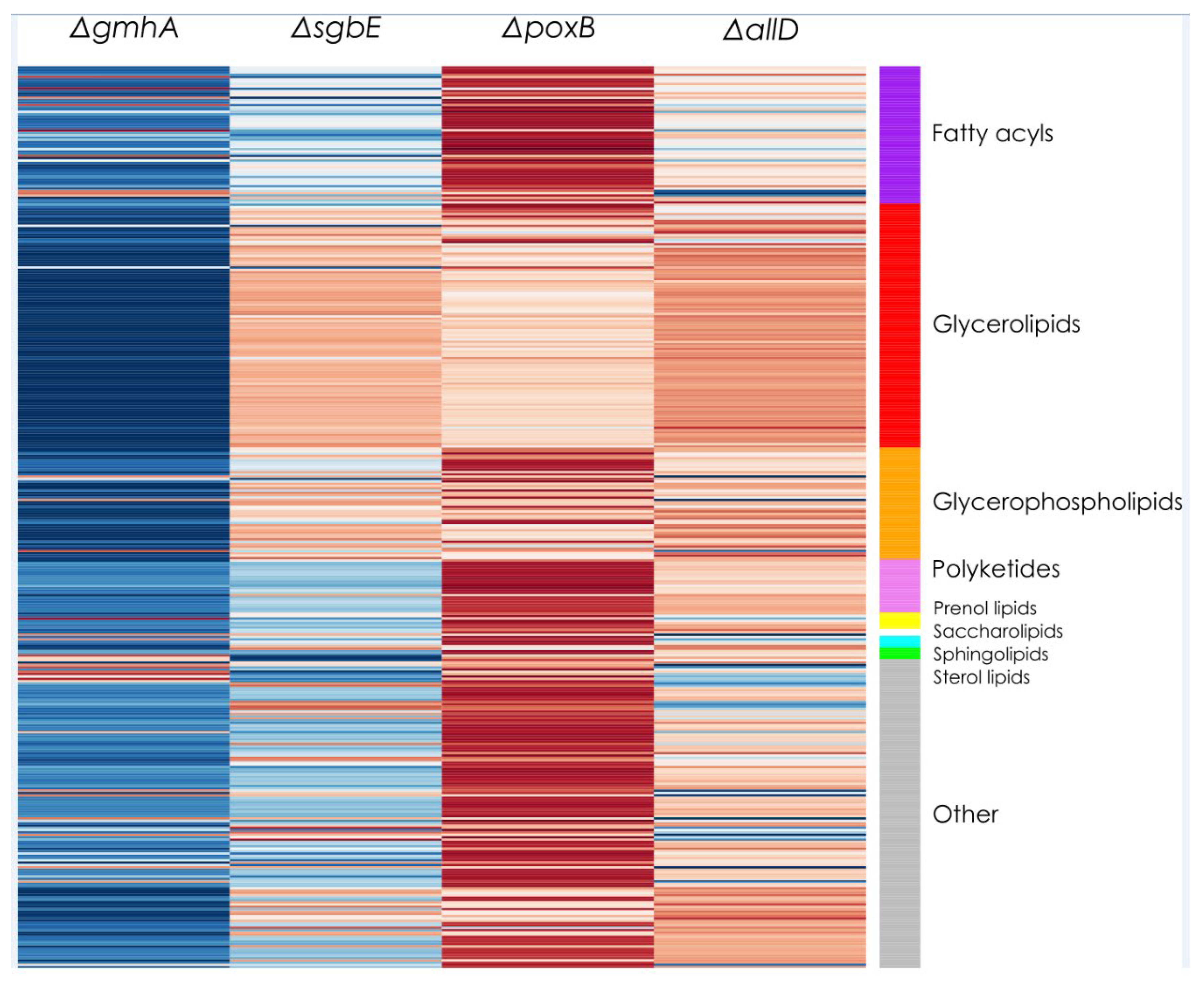
The lipid profile of
E. coli OMVs presented an unexpected major component in glycerolipids, while expected major components glycerophospholipids were found in second place. While glycerophospholipids are usually present in the outer membrane in large proportions, it is not uncommon for membranes composed of glycerolipids [
33]. The lack of phosphorus in membrane lipids is associated with the depletion of phosphorus in the growth media [
34]. In this case, LB culture media presented a rich nutritious environment for cell growth which provided phosphorous. This implies small participation of glycerophospholipids in the
E. coli OMV lipid profile is not due to phosphorus availability. It was not possible to determine if OMV lipid profile matches the outer membrane profile in whole
E. coli cells, or if vesicle formation was preferential in regions low in phosphorus or with higher amounts of glycerolipids. Additionally, proteomic studies have found an enhanced carryover of inner membrane to the generated vesicles [
5]. This could have implications for the engineering of membrane proteins for interaction with host cells and could partly account for the unexpected high glycerolipids content in the isolated OMVs. Further work is needed to analyze differences in OMVs from the originating outer membrane and the effect of phosphorus availability on the lipid profile.
Genomic advances have helped to determine the set of genes in an organism. Annotation of the genome of
E. coli has produced a map of enzymes expressed by these bacteria. A network of these reactions comprises the metabolic states the cells are capable of undertaking. A key limitation in metabolic modeling arises from genome annotation. Missing pieces in annotation results in disconnections between reactions which alter the distribution of metabolic fluxes and could result in blocked reactions, or no in silico growth. In the case of
E. coli, several network models that tackle these limitations have been developed [
18]. The
E. coli model has increased in the number of metabolites and reactions and expanded from the core metabolism to secondary metabolism and other functions. Disconnection of reactions still exists. Algorithms such as GrowMatch allow closing the gap between the computational model and the observed biological results. Results from GrowMatch suggest that this gap was somewhat but not fully closed in the proposed model of a high vesicle yield
E. coli strain. Even though this step is not completely successful in restoring growth-no growth consistency in the model, it brings it closer to the biological growth data available.
Vesicle generation is mediated by structural mechanisms. Some metabolic pathways have been related to the production of OMVs [
11]. However, these pathways are related to changes in the structure of phospholipids that cause instability of the cell envelope and higher vesicle production. In this study, we focused on exploring the metabolic pathways involved in the generation of OMV precursors: cell envelope precursors, lipidA and glycerophospholipids. This results in modeling a system of multiple metabolites. Connectivity in metabolic networks made the optimization of the selected objective function harder. In comparison, enhancement of the production of a single metabolite is easier to study as the effects of changes in the metabolic network can be traced back easier. To improve the predictive power of the metabolic model, additional information is required as input to further constrain the model and contextualize it to experimental data. Future work should include the improvement of the model through the integration of lipidomic and transcriptomic data. Integration of lipidomic data is challenging as lipids are usually expressed in generic metabolites in metabolic models and experimental data is not easily associated with the network.
4. Materials and Methods
4.1. Vesicle Recovery and Quantification
High vesicle yield
E. coli JC8031 strain was selected as the base strain (also referred to as WT strain).
E. coli JC8031 is a tol/pal mutant [
19]. This strain was modeled in silico and used for experimental procedures, vesicle recovery, and gene editing.
E. coli JC8031 was transformed with a GFP production plasmid with an ampicillin resistance cassette for selection.
E. coli was cultured in 50mL supplemented LB medium, with ampicillin for GFP plasmid selection (100 ug/mL ampicillin final concentration), and incubated for 24 hours at 37ºC in a shaker incubator at 250 rpm to obtain OMVs at a late log phase and avoid cell debris collection [
11]. The culture was transferred to 50mL falcon tubes and centrifuged at 4500 rpm and 20ºC for 30 minutes to remove cells. After this, 30 mL of supernatant was filtered through a 45um polyamide membrane in a filter housing using a 50 mL syringe to remove any debris present in the sample. The filtrate was ultracentrifuged at 50,000G for 3 hours at 4ºC [
5]. Presence of vesicles was verified as a translucent pellet with green fluorescence under blue light. Vesicles were resuspended in 1000uL of PBS with 10uL of spectinomycin antibiotic and stored at -80ºC until further analysis. Vesicle solution in PBS was inoculated in LB medium agar with ampicillin to check that no viable cells were still present.
Vesicle production was quantified indirectly through UV absorption spectra. Absorption was obtained in a UV spectrophotometer in a spectrum ranging from 200nm to 320nm at intervals of 2nm. Vesicle production values were calculated as the area under the UV absorption curve in the specified spectrum range. Additionally, the protein concentration in each sample was semi-quantitatively estimated using the Warburg-Christian equation using absorbance values at 280nm for protein content and 260nm to account for the influence of nucleic acid content [
23]. Vesicle production and protein concentration was normalized to the mass of sample and OD
600 to account for varying growth rates between sample groups.
4.2. Metabolic Network Model
A metabolic network model for
E. coli K12 was used as initial input to assess the OMVs production and to find optimal culture conditions
in silico. This model was initially developed by Monk et al. [
18] as iML1515 and modified to ensure the presence of biomass precursors and a functional biomass growth reaction. The model was adapted to strain JC8031 to account for different mutations.
4.3. Model Refinement
Vesicle production values and growth rates of 3908
E. coli mutants in the Keio collection were reported by Kulp et al. [
11]. Flux balance analysis (FBA) [
35] was used to simulate biomass growth of 150 Keio mutants reported by Kulp to exhibit high vesicle production phenotype. The results of the FBA were used to compare simulated and experimental cell growth rates and check the consistency of in silico results and reported in vitro data. Subsets of this dataset were used to perform improvements through constraint based optimization methods [
36]. Matching results from in silico predictions and reported experimental values are present as either growth-growth (GG), or no growth-no growth (NGNG). When inconsistencies appear, in silico predictions display growth while experimental results do not, or vice versa. These are cataloged as growth-no growth (GNG) or no growth-growth (NGG) inconsistencies. GrowMatch [
37] was used to calibrate in silico predictions to cell growth data of the Keio collection through the exclusion of model reactions, while NGG inconsistencies were resolved by the inclusion of reactions from an external database.
4.4. Model Objective Function Definition
The objective function for vesicle production is key to properly model biological behavior. As the objective function is maximized or minimized, the distribution of metabolic fluxes changes. This has the effect of guiding the model towards a biologically relevant solution. The objective functions found by using available algorithms [
38] were linear combinations of reactions found in the model but do not appear to be related to known mechanisms of vesicle production, important metabolic steps in vesicle production, or tied to the composition of vesicles and were over-fitted to the data. The objective function was described according to the expected composition of the
E. coli OM, glycerophospholipids and lipopolysaccharides [
34] as well as the expectation for an increased phospholipid production to increase OMV formation [
39]. This approach is similar to the constitution of biomass objective functions which are also key to the model [
40].
4.5. Strain Design
Targeted deletions to improve vesicle production were studied using the calibrated model with the defined objective function for vesiculation. The constraint-based method OptKnock [
21] was used to simulate gene deletions by inactivating reaction fluxes while maximizing vesiculation and biomass production. This approach ensures lethal deletions were not evaluated. OptKnock was implemented to simulate gene deletions on the refined model taking into account that LB medium metabolites are available as active exchange reactions. OptKnock algorithm was used limited to 5 deletions.
4.6. Genome Editing
Target genes were knocked-out by complete gene deletion by using CRISPR-Cas9. Guide RNAs were designed for the genes obtained through strain design using the tools from Benchling [
41] and a reference
E. coli genome (NCBI Accession: NC_000913.3). Synthetic cassettes were designed including the N20 from gRNA design (
Table 2) and two segments of 500bp corresponding to sequence flanking the target gene to provide template availability for homology directed repair of the double stranded DNA lesion thereby completely eliminating the targeted gene from the bacterial genome. A system of two plasmids was employed: a plasmid including Cas9 genes and a plasmid for gRNA expression [
42].
4.7. Lipidomics
Lipidomic analysis was carried out at MetCore Uniandes (Bogota, Colombia). 40 uL of type I water and 160 uL of HPLC-grade methanol were added to the lyophilized samples. Samples were resuspended in a vortex for two minutes. 400 uL of MTBE were added and samples were vortexed for 60 minutes. 250 uL of type I water were added and samples were vortexed for two minutes. Samples were centrifuged at 6190 rpm and 25°C for 10 minutes. 20 uL of organic phase were transferred to HPLC fixed-insert vials and diluted with 80 uL of MTBE.
Quality control samples were prepared by mixing equal volumes of metabolite extract from each sample. Quality control runs were performed to stabilize the analytic platform. Subsequent quality control runs were employed every five randomized samples.
Lipidomic analysis was implemented in an Agilent Technologies 1260 Liquid Chromatography system coupled to a quadrupole time of flight mass analyzer and ionization by electrospray (LC-ESI-QTOF-MS). 1 uL of each sample was injected in a C8 column (InfinityLab Poroshell 120 EC-C8 (150 x 3.0 mm, 2.7 µm)) at 60°C. A gradient elution was employed composed of 5 mM ammonium formiate in Milli-Q water (Phase A) and 5 mM of ammonium formiate in isopropanol-methanol 15:85 (Phase B) with a constant flux of 0,4 mL/min. Mass spectrometry detection was performed in ESI positive mode in full scan from 100 to 1100 m/z. Mass correction was employed throughout the analysis with two reference masses: m/z 121,0509 (C5H4N4) and m/z 922,0098 (C18H18O6N3P3F24).
Lipidomic profiles were obtained by using Agilent Mass Hunter Profinder 10 software employing the Recursive Feature Extraction (RFE) with extraction conditions: 0-31 min and 10000 counts, positive ion species: (-H, +Cl, +NH4), and no additional mass or species filters. Molecular characteristics found in the solvent blank control were eliminated. Data from alignment, deconvolution and integration were filtered by calculation of a variation coefficient (VC) of the area in QC samples. Characteristics with CV > 20% were filtered out. Data was normalized based on vesicle sample information: quantity and sample volume.
Selection of statistically significant molecular characteristics was performed through multivariate statistical analysis (MVA) and univariate (UVA). MVA was performed using SIMCA-P + 16.0 software (Umetrics) and UVA was performed by implementing the MetaboAnalyst 4.0 tool (#). Annotation of molecular characteristics was performed using the CEU MASS MEDIATOR tools (#) by batch analysis with the following parameters: tolerance=10 ppm; databases=LipidMaps; Metabolites=OnlyLipids; input masses mode= m/z masses; ionization mode=positive; adducts= (M+H, M+Cl, M+NH4). Additionally, molecular formulas were generated for statistically significant molecular characteristics by using the Agilent MassHunter Qualitative 10 software with positive ions= (-H, +Cl, +NH4) and elements= (H, C, O, N, P) as parameters.
4.8. Statistical Analysis
Data was analysed using GraphPad Prism version 9.3.1 (GraphPad Software). An ordinary one-way ANOVA was used to estimate the statistical significance of differences within groups. Sidak’s multiple comparison test was used to evaluate statistical difference with wild type group JC8031. Differences were considered significant at P values of <0.05.
5. Conclusions
In this study, we generated four single deletion mutants through in silico metabolic network model strain design and CRISPR-Cas9 genome editing (ΔpoxB, ΔsgbE, ΔgmhA, ΔallD). The four designed strains showed an enhanced vesicle production of up to 10 times that of the wild-type JC8031 strain while maintaining cell growth, while only three displayed a greater protein concentration (ΔpoxB, ΔsgbE, ΔallD). This is a step ahead to obtain OMVs in a large-scale setting and evidences it is plausible to use metabolic network modelling the complex vesicle formation process while targeting single genes.
Lipidomic analysis though LC-ESI-QTOF-MS showed that there were no significant changes in lipid profile that separated the four mutants and WT in different groups. OMV lipid profile remained stable across strains. Recovered E. coli OMVs displayed a lack of phosphorus in their composition as glycerolipids were the majority component while glycerophospholipids were present in a lesser quantity. Knowledge of E. coli OMV lipid profile and lack of alterations in different single deletion strains is useful to employ OMVs as therapeutic alternatives and drug transport and delivery.
Strain design using metabolic network models encounters difficulties when the target for production is a complex structure such as OMVs instead of a single metabolite. Integration of multiple source omic data should be used to improve the model.
Author Contributions
Conceptualization: AFGB, JDV, HAR, Data curation: HAR, MPC, JDV, Formal analysis: HAR, JDV, MPC, Funding acquisition: JDV, AFGB, Investigation: HAR, MPC, MF, JDV, Methodology: HAR, JDV, MPC, MF, AFGB, Project administration: JDV, AFGB,Supervision & Validation: AFGB, JDV, Visualization: HAR, MPC, Writing; original draft: HAR, JDV, MPC, Writing; review & editing: HAR, JDV, MPC, MF, AFGB.