1. Introduction
The conversion of tropical forests to agricultural uses is thought to negatively impact the structure of the soil microbial communities associated with decomposition and efficient conversion of C into biomass C (Arai et al. 2014; Lammel et al. 2015; Rodrigues et al. 2013). This is largely due to reduction in plant-derived resource inputs into the soils (Eaton et al. 2019, 2020a; Kardol & Wardle, 2010; Lammel et al. 2015). Although remediation strategies are commonly used to restore tropical forests (Aide et al. 2013; Schelhas & Sánchez-Azofeifa 2006; Chazdon, 2008), minimal information is available on how these strategies influence recovery of the damaged microbial communities that are critical for restoration of the soil C cycle activities and C capture. Such information would help maximize the efficacy of restoration methods used to restore tropical forest soil ecosystem health (van der Heijden et al. 2008; Berg & Smalla, 2009; Mendes et al. 2018; Eaton & Hamilton, 2022).
The decomposition of organic matter is an important component of soil ecosystem processes that are damaged during deforestation and requires restoration to fully recuperate the soil C cycle activities. Although bacteria are considered important for decomposition during early stages of soil recovery involving oxidation of less complex, more labile forms of organic C compounds (Wallenstein& Burns, 2011; Xu et al. 2021), they are less efficient than fungi at decomposition of the more complex and recalcitrant forms of organic C that are important for enhancing soil C use efficiency, biomass development, and C capture in soils (Brown & Chang, 2014; Datta et al. 2017; Kamimura et al. 2017; Lee et al. 2019; Silva et al. 2021; Vĕtrovsky et al. 2014; Whilhelm et al. 2019). Specifically, fungi have been shown to be more critical than bacteria for decomposition of such complex organic C compounds as lignin, lignocelluloses, polyaromatic compounds, suberins, resins, and others, whose complex biproducts of these oxidations enhance the soil organic C, biomass C, and C use efficiency (Datta et al. 2017; Hattaka & Hammel, 2010; Rahman et al. 2013; Thevenot et al. 2010). As such, changes in the fungal community of complex organic C decomposers should be considered a potential indicator of later stages of advanced decomposition and soil recovery, as well as the efficacy of remediation activities after forest disturbance.
Tropical forest leguminous trees and their soil microbiomes in either natural or assisted reforestation practices provide the principal pathways for recovery of the tropical forest soil N and C cycle dynamics following deforestation (Amazonas et al. 2011; Batterman et al. 2013; Davidson et al. 2007; Groppo et al. 2015; Lojka et al. 2010; Macedo et al. 2008). These abilities have led to the common use of these trees in tropical forest restoration strategies (Lojka et al. 2010; Macedo et al. 2008; Piotto, 2008). Members of the genus Inga are leguminous trees common throughout the tropical Americas, are ecologically important for enhancing accumulation of both soil N and C, are presumed to influence the soil microbial communities associated with these biogeochemical cycles and are commonly used in restoration practices (Leblanc et al. 2005, Lojka et al. 2010). However, little is known of how Inga spp. influence the soil microbiota, particularly the critical soil fungal community of decomposers. Such information would be valuable for the development of more efficacious restoration strategies using Inga to help restore damaged tropical forests in order to enhance recovery of the soil C cycle dynamics, and to assess the success of such strategies.
In a Monteverde, Costa Rica reforestation site, I. punctata trees were planted over time within a previously intact premontane wet forest area that had been cleared, used for agriculture and pasture, then abandoned. An earlier study (Eaton & Hamilton, 2022) showed these tree soils, along a tree age gradient from planted to older forest I. punctata trees, had increasing levels of total organic C (TOC), biomass C, and abundance and complexity of the lignin degrading bacteria increased over time. However, missing from this work was a comparison of the tree soil microbial communities to those in the adjacent pasture (which were part of the same the pasture into which the I.punctata trees were planted,) , how the trees influenced the soil fungal communities and the efficiency of microbial C utilization, and whether these latter two components could serve as indicators of more advanced soil microbial community succession and soil recovery.
Thus, purpose of this study was to assess whether the planting of I. punctata in abandoned pasture soils enhanced the soil fungal community, converting the soils from C sources to C sinks. To do so, the following objectives were used to further analyze the earlier collected DNA sequences and the soil environmental data for additional information not previously published to determine: 1) if the tree soil ecosystems of increasing ages of planted I. punctata were associated with enhanced microbial efficiency of converting organic C into biomass C (as decreased qCO2 levels); 2) if the tree soil ecosystems of increasing ages of planted I. punctata were associated with enhancement of the general fungal community, or the fungal community associated with complex organic C decomposition (CCDec fungal taxa); 3) if there were changes in levels of critical fungal taxa associated with the differences in the soil C metrics; and 4) if different fungal taxa became more or less prominent in the soils along the I. punctata tree age gradient.
2. Materials and Methods
2.1. Inga punctata Reforestation Site Description and Soil Sample Collection
A plan to restore abandoned pasture in the Monteverde Cloud Forest Region of Costa Rica has been implemented by the Fundación Conservacionista Costarricense (FCC) and Monteverde Institute (MVI) since 1998 and 2016, respectively (
https://monteverde-institute.org/reforestation-program.html). This effort included planting seedling
I. punctata in three restoration sites within the former farm site Finca Rodríguez Ecological Reserve (FRER), located in Monteverde (10⁰ 18′ 55.3” N, 84⁰ 50′ 29.8” W) in 2008, 2011, and 2015 in specific plots within the FRER (Hamilton, 2022). All plots were on land previously used for coffee, sugar cane, and dairy cattle pasture, then abandoned for four years before each planting. Prior to planting, manual clearing of all grasses was performed, while after planting all shrub and natural regeneration in the plots was minimized with weeding of the planted seedling plots, which continued until the year of this study. In August 2019, soil was collected from the base of 6 replicate
I. punctata trees from each of four age classes: those planted 4, 8, and 11 years prior to sampling (called the Inga 4, 8 and 11-year-old soils), and from older (> 50 years old) I.
punctata trees (called the Old Inga soils) within the FRER. The stands of any of the four tree age classes were 50-300 m apart. The soil collected from any of the planted trees was at least 5 m from any other planted tree in that stand, and the soil collected from the Old Inga trees were at least 15 m apart from other Old Inga trees. For comparative purposes, soil was also collected from 6 pasture plots (25 m x 20 m each) within the FRER (called PAS soils) that were adjacent to the planted
I. punctata stands, and separated by at least 25 m. The extremely small spatial scale of soil microbial communities is such that soil samples separated by several centimeters to several meters is equivalent to an ecosystem level separation for above-ground communities, and those that are greater than several meters are equivalent to a landscape-level separation of above ground communities (Ettema, 2002; Turbé et al. 2010; Bardgett and van der Putten, 2014). Given this, the soils collected for this study represent 6 true soil replicates from each tree age stand and 6 from the pasture.
Soils were collected from beneath the
I. punctata trees using a combination of methods that are now characterized by Addison et al. (2023) which control for neighbor tree and other environmental effects and ensure all samples are from the same functional location of all trees. This allows for comparison of the potential influence of the aging tree on the specific tree soil fungal community compositions. Specifically, for each of the 6 trees used per tree age class, the 10% distance from the base of the tree to its canopy edge was calculated and considered as the 10% dripline region. Two soil profiles (7.5 cm x 1.25 cm x 15 cm) were collected (approximately 15 cm depth) at each of the four cardinal points at the 10% dripline distance point for each tree, providing 8 soil cores per tree, which were pooled into a single sterile bag per tree. This provided 6 replicate 8-core pooled soil samples from each tree age stand for analysis. Using this method, no other tree canopy radius, shrubs or regenerating trees were within the 10% dripline zone sampled for each
I. punctata tree. There were 6 soil cores collected from each of the PAS plots using the same soil profile corer, and using the stratified, block, systematic plot study design recommended for studies of damaged forest lands (see
www.forestandrange.gov and
www.epa.gov/sites/default/files/2015-06/documents/g5s-final). The 6 soil cores from each PAS plot were collected using a pre-determined sample location strategy and placed in a single sterile bag per plot, providing 6 composite, independent replicate soil samples from the pasture region. Prior to all soil collection, any surface leaf litter was carefully removed to expose the upper organic horizon. In addition, all soil samples were aseptically collected by disinfecting all gloves and collection tools with 70% ethanol between trees and between PAS plots. The samples were passed through sterilized 5-mm sieve at field moist conditions to homogenize the soil sample prior to analysis. After sampling, following the recommendations of Lucas et al. (2020), the field moist soils were kept in semi-open plastic bags under refrigeration at 4 °C for no more than 1 week prior to analysis, as these authors showed that holding tropical soils under these conditions for up to 12 days did not affect microbial activity results.
2.2. Soil Respiration, Biomass-C, and qCO2
Subsamples of 200 g of field and sieved moist soil per replicate soil sample were analyzed by the Centro Agronómico Tropical de Investigación y Enseñanza (CATIE) in Turrialba, Costa Rica for all C cycle metrics. The levels of Respiration (as CO2) were determined by standard closed vessel methods, and the microbial Biomass C determined by standard chloroform-fumigation methods (Alef & Nannipieri, 1995). Many studies have discussed how the Metabolic Quotient, or qCO2, has been used for decades as an indicator of environmental impacts on microbial catabolic activity, the efficiency with which soil microbial communities convert organic C into biomass C, and enhanced soil ecosystem microbial community development, such that at maturity of microbial diversity in a soil system there should be a decreased level of respiration per unit of biomass C generated (for example: Anderson & Domsch, 2010; He et al., 202; Martínez-García et al., 2018; Six et al., 2006; Spohn, 2015; Wallenstein & Hall, 2011). Given this, we used the qCO2 (the ratio of Respiration to Biomass C) with decreasing values to indicate development of a more mature and advanced fungal community of microbes with increased efficiency of converting soil organic C into biomass C. Differences in all mean values were assessed by one-way analysis of variance (ANOVA) followed by Tukey’s HSD or Dunnett’s T3 post-hoc tests, as appropriate, in SPSS (v.27, Armonk, NY, USA). Prior to ANOVA, the Levene’s test and Shapiro-Wilk test were performed in SPSS to determine homogeneity normality of all the data to support the use of ANOVA.
2.3. DNA Extraction, Sequencing, and Bioinformatics
The soil DNA was extracted from three 0.33g soil per each replicate soil sample using the MoBio PowerSoil DNA Isolation Kit (MO BIO Laboratories Inc., Carlsbad, CA, USA), and the concentration and purity determined using a NanoDrop 1000 spectrophotometer (ThermoFisher Scientific, Waltham, MA) prior to downstream. All methods for PCR, DNA sequencing, quality control,Illumina MiSeq sequencing, and bioinformatics used to identify the fungal taxa have been previously explained in detail by McGee et al. (2019). Briefly, the eDNA extracts were amplified by 2-step PCR reactions, targeting the nuclear internal transcribed spacer (ITS) ribosomal RNA gene region for fungi using the ITS1F and ITS4 primers (Gardes & Bruns, 1993), with the resulting amplicons sequenced in Illumina MiSeq runs using a V3 MiSeq sequencing kit (FC-131-1002 and MS-102-3003). The ITS DNA sequences were processed using semi-automated pipelines to generate operational taxonomic units (OTUs) that were taxonomically assigned using the RDP classifier with the UNITE fungal ITS set for fungi (Liu et al. 2012). All sequencing data was submitted to the NCBI-Gene Expression Omnibus (GEO) repository on August 1, 2019 (Submission: SUB6145149, BioProject: PRJNA559202).
There were 6,395 unique fungal OTUs identified in the PAS soils, 2,792 in the Inga 4 soils, 3,189 in the Inga 8 soils, 6,379 in the Inga 11 soils, and 5,928 in the Old Inga soils. These unique OTUs were categorized into 685 specific fungal taxa, generally at the genus level. The library size of each taxon was normalized for each soil sample by determining the mean percentage of the sequences (MPS) for each taxon, as recommended by Weiss et al. (2017), to account for differences in number of sequence hits between samples. The fungal taxa were categorized as being saprobes (SAP), associated with decomposition of less complex organic C decomposers (Dec), root associated (RA), associated with complex carbon decomposition (CCDec), endoparasites (PAR), and wood rot fungi (WRT) using the databases
www.genome.jp/kegg/, https://fungidb.org/, FungalTraits 1.2 ver 16 (Põlme et al. 2020) ,
http://rdp.cme.msu.edu/hierarchy/, www.zoology.ubc.ca/louca/ FAPROTAX/ lib/php/index.php, and
https://www.ars.usda.gov/ (USA National Fungus Collection), and other literature sources (Table S1). The fungal taxa found to be capable of decomposing complex forms of C or were wood rot fungi, are hereafter referred to as the CCDec Fungal taxa. Both the Total Fungal taxa and the CCDec Fungal taxa were used for further analysis. This approach has been recently used by Eaton et al. (2019, 2020 a, b, 2021 a, b, 2024) and Eaton and Hamilton (2022) to link microbial taxa to proposed functional activity within soils from other studies in Costa Rica.
2.4. Differences in Fungal Community Compositions
Differences in fungal community composition were assessed by comparing the MPS levels, the Margalef’s Richness levels, and by ANOSIM and CAP analyses. The MPS values of two subsets of the Total Fungal community were used to compare mean differences in taxa between the soil groups by the Kruskal – Wallis analysis conducted in SPSS (v. 26): the subset of Total Fungi with MPS values > 1.0% (called the Most Common Taxa) and the representatives from this group identified as CCDec Fungal taxa. The MPS data were 4th root transformed as recommended by Anderson et al. (2008) to diminish the influence of extremely abundant or rare taxa and used to determine the Margelef’s Richness indices using PRIMER-E v.6 (Clarke & Gorley, 2006). The richness levels were assessed for differences between soils by ANOVA in SPSS.
Overall taxonomic compositional differences of both the Total Fungal taxa and all CCDec Fungal taxa between the soil habitats were assessed using the ANOSIM routine in the PRIMER-E package (Anderson et al. 2008) applied to 4th root transformed MPS data that was then converted into Bray-Curtis matrices. ANOSIM provides Global and Pairwise R statistics and p values for the main and comparative tests that are used to identify differences in mean values between groups. The strength of any community compositional differences between soil groups identified by ANOSIM was determined using the Canonical Analysis of Principal Coordinates (CAP) ordination method (Anderson & Willis, 2003; Clarke and Gorley, 2006) in the PRIMER-E package with the add-on PERMANOVA+ applied to the same Bray-Curtis similarity matrices mentioned above. The resulting CAP axis squared canonical correlations (R2) provide an approximation of the strength of differences in community compositions between soil samples. Strong differences are indicated by R2 values ≥ 0.7, moderate differences by R2 = 0.5 to 0.69, weak differences by R2 = 0.20 to < 0.5, and no differences by R2 < 0.20.
2.5. Indicators of Fungal Community Successional Development
Soil ecosystems in older established forests, recovering from damage, or undergoing restoration often demonstrate several community compositional patterns linked to microbial community successional development that were assessed in this study. During succession in soil communities, there is often an increase in abundance of critical microbiota associated with the decomposition of more complex organic C compounds (Eaton et al., 2020a, Luis et al., 2004, Peng et al., 2008; Ren et al., 2019). For the current study, this was assessed by comparing the MPS levels of the different taxa for each soil group as described above and determining the changes in levels of the CCDec taxa along the gradient from PAS to Old Inga soils. Another indicator of successional development in the soils is that there may often be decreases in taxonomic richness due to competitive exclusion processes, which result in certain taxa becoming more common or typical of a soil habitat as they become more dominant and better fit for the developing soil niches (Allison and Martiny 2008; Atkinson et al., 2022; Eaton et al., 202a; Galand et al., 2016; Louca & Doebeli, 2016; Louca et al. 2018). Here, the Margalef’s richness levels were determined for each fungal group group and compared for differences by ANOVA. As well, the multivariate SIMPER routine was performed on the transformed data in PRIMER-Ev.6 (Clarke & Gorley, 2006) to determine the percent contribution of each fungal taxa to their respective total community compositions. From this, the more typical or characteristic taxa for a soil group were determined to be those that provided the greatest percent contribution to the community composition for that soil group.
These successional-related processes ultimately result in an increasing level of taxonomic stability as communities develop into a ones demonstrating a more homeostatic relationship with the environmental conditions of the more established and developed soils (Grimm and Wissel 1997; Tilman et al. 2006; Ruijven and Berendse 2007; Isbell et al. 2009; Liu et al., 2020; Shade et al. 2012). We assessed the taxonomic stability of the Total Fungal and CCDec Fungal communities by calculating the Stability S index of the MPS and the richness values as the mean value divided by the standard deviation of that mean, with greater S values suggesting greater stability of that community metric (Griffiths & Philipott 2013; Grimm &Wissel 1997; Shade et al. 2012; Tilman et al. 2006; van Rujiven & Berendse, 2007). The S index has previously been used in microbial community studies (Eaton & Hamilton, 2022; Eaton et al., 2024; Griffiths & Philippot, 2013; Shade et al.2012).
2.6. Potential Influence of Fungal Taxa on the Carbon Metrics
The distance-based linear model (DistLM) analysis (Primer E v.6 and PERMANOVA+) was implemented to determine the degree to which any of the Most Common Fungal taxa might influence the patterns of the soil Biomass-C, Respiration, or qCO2. The MPS data of the Most Common Fungal Taxa were used as predictor variables, and the log (x + 1) transformed C metrics were used as the response variables after converting the transformed C data into Euclidean similarity matrices (Anderson et al. 2008). A stepwise selection process was used, along with an AICc (Akaike’s Information Criterion Corrected) selection criterion and 9,999 permutations (Anderson et al. 2008) for the DistLM analysis.
3. Results
3.1. Differences in C Metrics
There were differences observed in the levels of Biomass-C, Respiration, and
qCO
2 between some of the different soils (
Table 1). The Biomass-C and Respiration were greater in the PAS soils than in the Inga 4 and 8 soils (
p < 0.0001), the Inga 11 soils (
p = 0.0001 and 0.022). However, the Biomass-C levels were greater in the Old Inga than in any other soils (
p range < 0.0001 to 0.031), but the Respiration levels were only somewhat greater in the Old Inga than the Inga 4 and 8 soils ((
p = 0.054 and 0.051), not different between the Old Inga and Inga 11 soils, and less than in the PAS soils (
p = 0.008). There were no differences in the
qCO
2 levels between the PAS, Inga 4, and Inga 8 soils, however the levels were lower in the Inga 11 and Old Inga soils than in the PAS, Inga 4 and Inga 8 soils (
p range = 0.0002 to 0.034). and were also lower in the Old Inga soils than in the Inga 11 soils (
p = 0.015).
3.2. Differences in Fungal Community Compositions
There were 17 of the Total Fungal taxa from the soil groups with MPS levels of ≥ 1% in one or more of the soil groups, which were called the Most Common Fungal taxa, 11 of which were identified as the CCDec Fungal taxa, those with either CCDec or CCDecWRT capability (
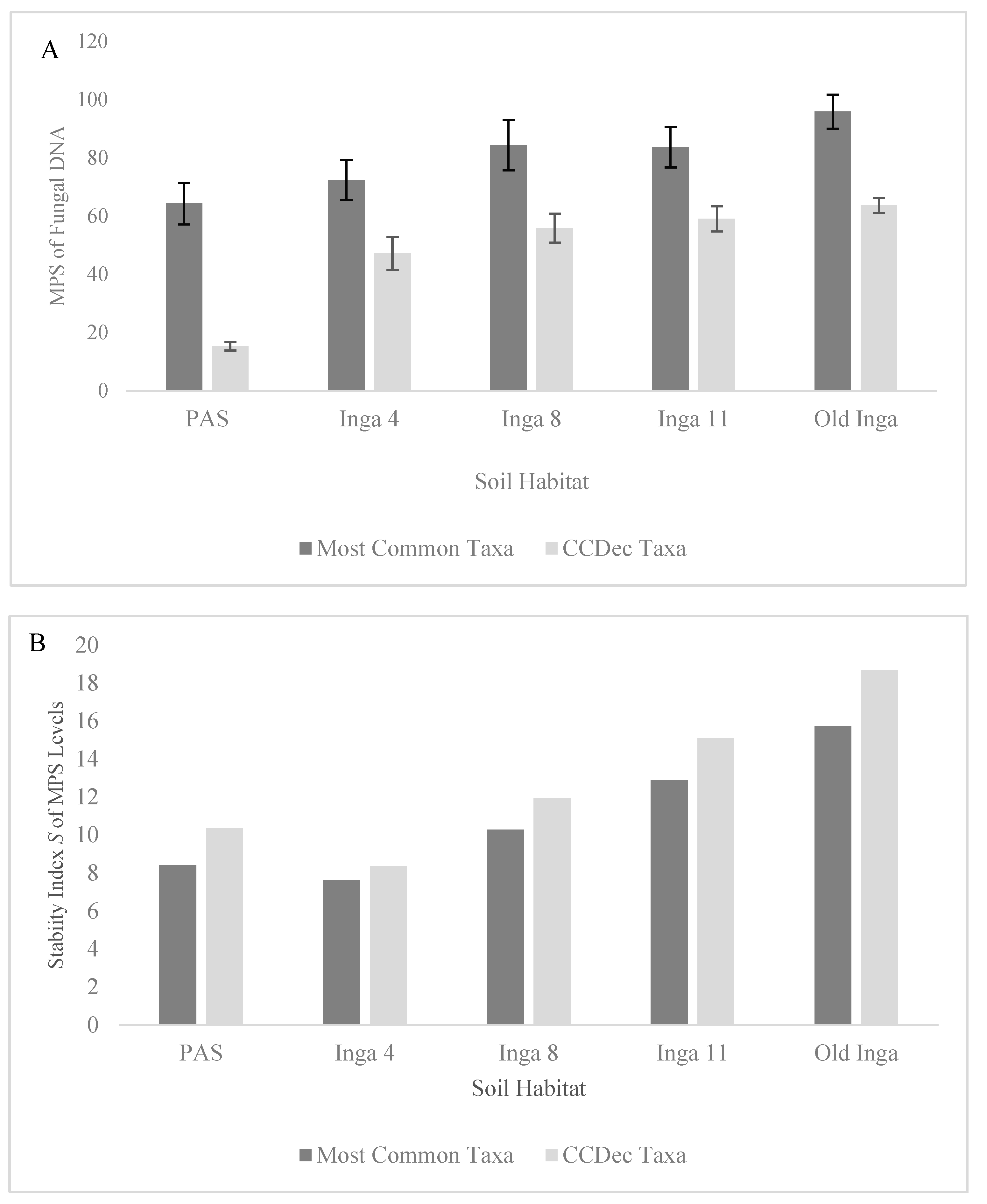
Table 2). The MPS levels of the Most Common Fungal taxa (
Figure 1A) increased along the tree age gradient, as they were lowest in the PAS soils and increased in the Inga 4 soils (64.3% to 72.4%,
p = 0.052), increased from the Inga 4 to Inga 8 and 11 soils (72.4% to 84.4% and 83.73%,
p values < 0.02), and increased in the Old Inga soils (95.9%,
p < 0.025). The MPS levels of the CCDec Fungal taxa (
Figure 1A) followed the same pattern. These MPS levels were far lower in the PAS soils (15.3%) than in any other soils (47.16%-65.04%; all
p values < 0.0001), then increased from the Inga 4 to Inga 8 and 11 soils (45.16% to 55.86% and 59.04%,
p = 0.017), and increased again to the levels in the Old Inga soils (59.04% to 63..65%,
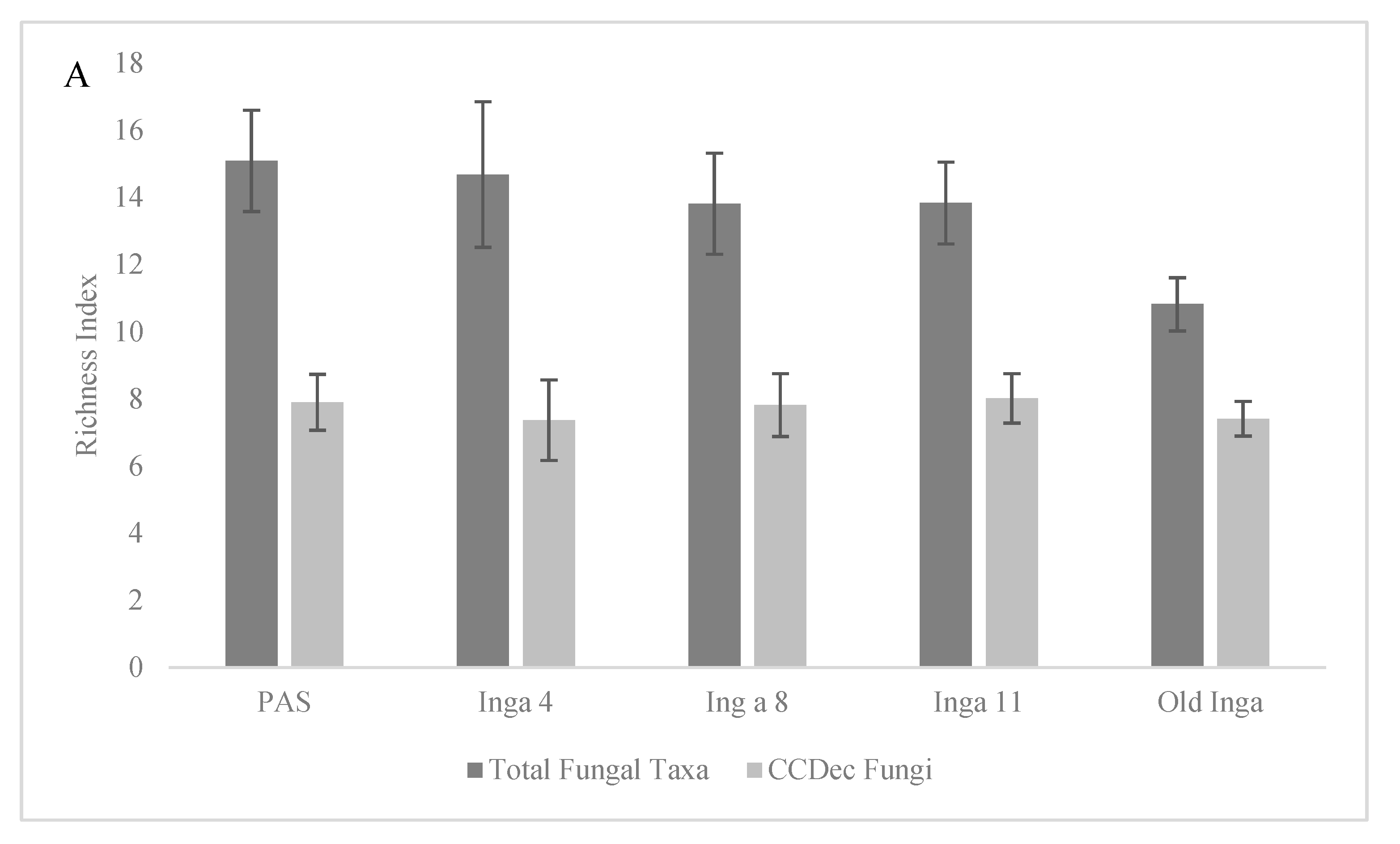
p = 0.048). Although there was a trend observed of decreasing richness of the Total Fungal taxonomic community along the tree age gradient (
Figure 2A), the Margalef’s richness indices were not different (p > 0.237) between the PAS, Inga 4, 8 and 11 soils (d = 13.8 to 15.09) but were the lowest in Old Inga soils (
d = 10.8; p values < 0.002). There were no differences (p > 0.284) in the Margalef’s richness indices of the CCDec Fungal community between any of the 5 soils in this study (
d = 7.37 to 8.02).
The ANOSIM and CAP assessments showed similar results for the differences in composition of both the Total Fungal and all the CCDec Fungal taxonomic communities (
Table 3). These community compositions were separated into 3 levels of difference that approximately coincided with either time between the PAS to planted tree age, or the time between different planted tree ages. There were weak to no differences in either of the two fungal community compositions between the Inga 4 and Inga 8 soils (ANOSIM R = 0.097 and 0.027; CAP R
2 = 0.044 and to 0.019) and the Inga 8 and Inga 11 soils (ANOSIM R = 0.323 to 0.386; CAP R
2 = 0.449 to 0.233). There were moderate differences in the two fungal community compositions between the PAS and Inga 4, PAS and Inga 8, and the Inga 8 and Old Inga soils (ANOSIM R = 0.360 to 0.503; all CAP R
2 values = 0.568). The greatest differences in both fungal community compositions were found between the PAS and Inga 11, PAS and Old Inga, the Inga 4 and Inga 11, and Inga 4 and Old Inga soils which demonstrated strong differences between the soils (ANOSIM R = 0.564 to 0.669; CAP R
2 = 0.745 to 0.791).
3.3. Indicators of Fungal Community Successional Development
The indicators of community successional development used in this study were the MPS and richness levels of the fungal groups, the Stability
S index of both metrics for these two groups, and the presence of taxa typical of the different soils. As mentioned above, there were significant increases in the MPS of both the Most Common Fungal taxa and the CCD Fungal taxa along the tree age gradient from the PAS to the Old Inga tree soils (
Table 2,
Figure 1A). As well, also as mentioned above, there were no significant differences in richness of the CCDec taxa across the soils, and the richness of the Total Fungal Taxa was not different between the PAS to the Inga 11 soils, but was less in the Old Inga soils (
Figure 2A). However, the Stability
S index values for the MPS values for both Most Common Fungal and the CCDec Fungal taxonomic communities followed similar patterns (
Figure 1B), as they decreased from that in the PAS (8.4 and 10.4) to the Inga 4 soils (7.6 and 8.4), then increased in the Inga 8 soils (10.3 and 11.9), increased in the Inga 11 soils (12.9 and 15.1), and increased again in the Old Inga soils (15.7 and 18.7). Interestingly, the Stability
S indices of the MPS and the Margalef’s Richness values for both the Most Common Fungal and the CCDec Fungal communities showed the same pattern (
Figure 2B). These
S values decreased between the PAS (10.0 and 9.5, respectively) and the Inga 4 soils (6.8 and 6.2, respectively), then increased each year in the Inga 8 soils (9.2 and 8.3, respectively), Inga 11 soils (11.4 and 10.9, respectively), and the Old Inga soils (13.6 and 14.4, respectively).
There were specific taxa that showed significant differences in the MPS levels and percent contribution to the Total Fungal community composition between the different soils (
Table 2 and
Table 4). . This suggests these taxa are more typical of the respective soils and may serve as indicators of the different stages of soil succession or recovery over time as the planted trees age. The percent contribution of the CCDec taxon
Saitozyma to the Total Fungal community composition was greater in the Inga 4 and 8 (16.48% and 15.35%) than in the Inga 11 and Old Inga (10.95% and 10.24%) or PAS (0.17%) soils, as was the MPS (29.42% and 28.89% vs. < 9% for other soils;
p < 0.042). The percent contribution of the CCDec taxon
Apiotrichum was far greater in the Inga 11 and Old Inga (22.38% and 24.53%) than in the Inga 4 and 8 (0.06% and 6.99%) and PAS (9.48%) soils, as were the MPS values (43.61% and 41.96% vs. < 6.5% for all other soils,
p <0.016). The percent contribution of the CCDec taxon
Tremella was greater in the Old Inga soils than in any other soils (10.23% vs < 0.2% for all others), as was the MPS values (8.21% vs. < 1.6% for all others,
p < 0.035). The saprobe fungus
Starmerella was also a greater contributor to the community composition in the Inga 11 and Old Inga soils than in all other soils (10.45% and 14.77% vs < 5.2% for all others) and had a greater MPS in the Inga 11 and Old Inga soils than in the other soils (8.13% and 11.44% vs < 3% for all others, < 0.051). Members of the CCDec fungal family Sordariaceae were greater contributors to the community composition of the Inga 4 and Inga 8 than in the other soils (9.39% and 9.85% vs < 5.84% for the others), also with a greater MPS than the other soils (7.98% and 11.6% vs < 1.5% for all others,
p < 0.024). The root-associated
Archaeorhyzomyces had a much greater percent contribution in the PAS than in the other soils (18.22% vs < 11.5% in all others), and with a greater MPS value (32.11% vs <5.5% for all others,
p < 0.015) in the PAS than the other soils. The endoparasite of fungi and oomycetes,
Rozella, also had a greater percent contribution in the PAS soils than in other soils (13.69% vs < 5.5% in all others), a greater MPS value (7.97% vs < 2% for all others,
p < 0.037). The arbuscular mycorrhizal fungal group Glomeromycota were found in all the soils, but the PAS soils had the lowest percent contribution of this group (9.45% vs > 16.2% in all others) and MPS (4.8% vs > 12.7% in all others,
p < 0.046) than in the other soils. In addition, the % Contribution of the total CCDec taxa to the overall composition of the Total Fungal taxonomic community increased in each soil sample along the tree age gradient from 25.11% in the PAS soils, to 42.41% in the Inga 4 soils, to 47.026% in the Inga 8 soils, to 55.13% in the Inga 11 soils, and to 58.76% in the Old Inga soils (
Table 4). As mentioned above, this occurred with the significant increases in total CCDec MPS levels from the PAS soils (15.3%), to the Inga 4 soils (47.16%), to the Inga 8 and Inga 11 soils (55.86% and 59.04%, respectively), and to the Old Inga soils (63.65%).
3.4. Potential Influence of Fungal Taxa on the Carbon Metrics
The DistLM analysis (
Table 5) showed that
Saitozyma and
Apiotrichum were the most probable fungi influencing the patterns of the Biomass-C, Respiration, and
qCO
2 as these two taxa explained 43.97% of the combined patterns of the Biomass-C, Respiration, and
qCO
2 metrics combined. These two taxa also explained 54.82% and 41.99% of the individual patterns of the Respiration and qCO2 values, respectively, while
Saitozyma was the only taxon that explained the individual patterns of Biomass-C values, as it explained 48.67% of the patterns of the Biomass-C values.
4. Discussion
The results from this project indicate that the Total Fungal community and the two analyzed subsets (the Most Common Fungal and CCDec Fungal communities) are becoming more successionally advanced over time in the I. punctata tree soils compared to the pasture soils, which is occurring concurrently with increases in soil Biomass-C, Respiration and enhanced efficiency of converting organic C into Biomass-C (lower qCO2). This indicates that planting I. punctata enhances the tree soil fungal microbiome and associated C-cycle activities resulting in the soil becoming more of a C sink, as opposed to being more of a C source, as is the case with the pasture soils (high respiration rate and high qCO2). There are several specific lines of evidence for this from the study.
The ANOSIM and CAP results showed the Total Fungal and CCDec community compositions were different between the different soils, with both the level and strength of difference coinciding with the difference in time of separation between the PAS and planted trees or by separation of the ages of the planted trees themselves. Specifically, the Inga 4 and 8 and the Inga 11 and Old Inga trees were the least separated by time since planting and had community compositions that were the most similar between them. The PAS and Inga 11, PAS and Old Inga, the Inga4 and 11, Inga 4 and Old Inga, and the Inga 8 and Old Inga soils were the most separated by time since planting or tree age and both fungal communities demonstrated the greatest compositional differences between these soils. These compositional differences were associated with increasing levels of the MPS and the Stability S index of the MPS of the Most Common Fungal and the CCDec Fungal communities, and Margalef’s richness and the Stability S index of the richness of both the Total Fungal and CCDec fungal communities along the tree age gradient. This suggests the taxa within these fungal groups are becoming more dominant and stable over time in the soils, which would be expected of microbial communities in soils undergoing succession or recovery from damage (Allison & Martiny 2008; Eaton et al. 2020a; Griffiths and Philippot 2013; Liu et al. 2020; Louca et al. 2016; Shade et al. 2012 ).
It is known that different microbial taxa will become more typical of a soil habitat undergoing succession or recovery from damage as competitive exclusion occurring within the soils facilitates selection of taxa better fit for the different niches that develop over time (Allison & Martiny, 2008; Louca & Doebeli, 2016; Louca et al. 2018). In the current study, the CCDec taxa Saitozyma and Sordariaceae were more typical of the Inga 4 and Inga 8 soil communities, while the CCDec taxon Apiotrichum and the plant saprobe Starmerella were more typical of the Inga 11 and Old Inga soil communities, and the CCDec taxon Tremella was more typical of the Old Inga tree soil. It may be that greater MPS levels of Saitozyma and Sordariaceae are indicators of early stages of soil ecosystem recovery after damage, while greater MPS levels of Apiotrichum and Starmerella are indicators of later stages of complex organic C compound decomposition and advanced soil ecosystem recovery after damage. The DistLM results support this in that Saitozyma and Apiotrichum were the greatest predictors of the differences found in the Biomass-C, Respiration and qCO2, and suggests that they are important for differentially decomposing complex organic C materials in the early and later stages of soil recovery post-tree-planting.
The PAS soil also had several distinguishing taxonomic characteristics. Archaeorhizomyces and Rozella were greater in the PAS soils that the other soils. This is consistent with evidence in the literature as Archaeorhizomyces and Rozella have both been found to be common in pastures (Pinto-Figueroa et al. 2019; Rosling et al. 2011 and 2013; Tedersoo et al. 2017; Turatsinze et al. 2021). The greater presence of Archaeorhizomyces in the pasture soils may be because it is a non-mycorrhizal root-associated saprotroph that should thrive in the thick root mass of pasture soils (Pinto-Figueroa et al. 2019; Rosling et al. 2011), however, it is not evident why would the endoparasite Rozella would be dominant in the pasture soils. The levels of the arbuscular mycorrhizal (ARM) Glomeromycota were much less abundant in the PAS soils than in any of the other soils, which supports the recent review that found Glomeromycota taxa to be somewhat more common in tropical and subtropical forest soils than in the pastures (Stürmer & Kemmelmeier 2021), which may be due to the greater abundance and diversity of woody plants as ARM hosts in the forested areas. This point needs further work to clarify what could be occurring within the Glomeromycota community with these different soil groups. Nonetheless, in addition to increases in levels of Saitozyma and Sordariaceae and increases in levels of Apiotrichum and Starmerella serving as indicators of early and then later stages of recovery, respectively, it may also be that greater levels of Glomeromycota may also be an indicator of soils undergoing recovery.
The increase observed in the CCDec fungal community MPS, stability, successional development, and percent contribution to the Total Fungal community composition along the tree age gradient suggests the importance of the fungal CCDec activity within all tree soil age groups. Further, it appears that after about 8-11 years post-planting of I. punctata, this taxonomic community reaches a level of homeostasis of taxonomic distribution within the soils that results in an increase in community compositional stability. This is also consistent with a pattern of functional redundancy that may be occurring in the soils associated with CCDec taxa, as these fungal taxa possessing similar metabolic capabilities are experiencing community compositional changes while living within similar niches that are undergoing successional changes over time (Allison & Martiny 2008; Nemergut et al. 2013; Louca et al. 2018). Though more common in bacterial communities, functional redundancy has been shown to occur in fungal communities, although the mechanisms are not clear (Mori et al. 2015; Bannerjee et al. 2016; Wagg et al. 2019). Regardless, it appears that the fungal CCDec community has undergone a rapid successional development in the soils post-tree planting which warrants further study as it suggests this group may be an indicator of soil recovery.
5. Conclusions
The present study showed that following deforestation of a tropical Cloud Forest area and it’s conversion to agricultural use, planting I. punctata in the abandoned pastures appears to facilitate: 1) the successional development of soil fungal community taxa that are experiencing an increase in the MPS levels and the stability of the MPS levels and taxonomic richness of the fungal communities; 2) an increase in the abundance and importance (as % Contribution) of the fungal CCDec Fungal taxa that are critical for complex organic C decomposition in the soil; and 3) the presence of certain fungal taxa that are possibly characteristic of earlier and later stages of soil recovery. All of these changes were associated with increases in the soils capacity to serve as a C sink that was indicated by the increase in soil Biomass-C development, Respiration, and efficiency of conversion of organic C to biomass (qCO2).
This information could be fundamental to understanding the role of tropical leguminous trees in enhancing the soil fungal community and soil C capture and recovery post-disturbance. This is especially timely now as some have questioned whether tropical leguminous trees have positive (Xu et al. 2019) or negative (Taylor et al. 2017; Lai et al. 2018) influences on tree regrowth patterns in reforested tropical areas. It is likely that there are many other co-varying biotic components, including the influence these trees have on the soil fungal microbiomes, that are important drivers of tropical forest regeneration pathways (Arroyo-Rodríguez et al. 2017; Eaton et al. 2020b). Thus, a more comprehensive understanding of the role that dominant leguminous trees such as I. punctata have on recovery of the tropical forest soil fungal communities critical to the C cycle is needed. This information could help in the development of more efficacious tropical reforestation and forest management plans and provide more insight into what components are driving individual tree species regeneration pathways during restoration (Wagg et al. 2011; Mendes et al. 2018; Eaton et al. 2020b; Eaton & Hamilton, 2022). In any case, this current study indicates that planting I. punctata should be part of future restoration and reforestation strategies used in this cloud forest region of Costa Rica to repair the soil ecosystem decomposition activities that are critical for recuperation of healthy C cycle activities and overall forest recovery.
Funding
This work was funded by the Pace University Dyson College Dean, the Dyson Faculty Research Grant Committee, and the Provost Office for research support, and by the Monteverde Institute.
Data Availability Statement
All DNA sequencing data was submitted to the NCBI-Gene Expression Omnibus (GEO) repository on August 1, 2019 (Submission: SUB6145149, BioProject: PRJNA559202). All other data is available upon request.
Acknowledgments
The authors thank Luisa Moreno, Monteverde Institute for assistance in permit acquisition, the FCC and MVI staff for their restoration efforts, Patricia Soteropoulos and Alex Lemenze for the Rutgers University Genomics Center for DNA sequencing, and Pace University students James Day, Brenda Hernandez, Frankie Iglesias, Stephanie Mafra, and Natalie Wayland for assistance collecting soil. We especially wish to thank Patricia Leandro and the staff at the CATIE lab for their soil C analysis. This work was conducted through the Costa Rican permit CONAGEBIO MINAE # R-047-2019-OT-CONAGEBIO.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Addison, S., Daley, K., 2023. Getting to the Root of Tree Soil Microbiome Sampling. Phytobiomes, 2023, ISSN: 2471-2906.
- Aide, T.M., Clark, M.L., Ricardo Grau, H., Lopez-Carr, D., et al., 2013. Deforestation and reforestation of Latin America and the Caribbean (2001–2010). Biotropica 45,262–271.
- Allison, S.D., Martiny, J.B.H.,2008. Colloquium paper: resistance, resilience, and redundancy.
- in microbial communities. PNAS 105,11512–11519.
- Amazonas, N.T., Martinelli, L.A., Piccolo, M.D.C., Rodrigues, R.R., (2011) Nitrogen dynamics during ecosystem development in tropical forest restoration. For. Ecol. Manag. 262,1551–1557.
- Anderson, M.J., Gorley, R.N., Clarke, K.R., 2008. PERMANOVA+ for PRIMER: guide to software and statistical methods PRIMER-E Ltd. Plymouth UK.
- Anderson, M.J., Willis, T.J. 2003. Canonical analysis of principal coordinates: a useful method of constrained ordination for ecology. Ecology 84, 511–525.
- Arai, H., Hadi, A., Darung, U., Limin, S.H., 2014. Land use change affects microbial biomass and fluxes of carbon dioxide and nitrous oxide in tropical peatlands. Soil Sci. Plant Nutr. 60, 423–434.
- Banerjee, S., Kirkby, C.A., Schmutter, D., Bissett, A.,2016. Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol. Biochem, 97,188-198.
- Bardgett, R.D., van der Putten, W.H. 2014. Belowground biodiversity and ecosystem functioning. Nature 515, 505–511.
- Batterman, S.A., Hedin, L.O., van Breugel, M., Ransijn, J.,, et al., 2013. Key role of symbiotic dinitrogen fixation in tropical forest secondary succession. Nature 502,224–227.
- Berg, G., Smalla, K. 2009. Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere: Plant species, soil type and rhizosphere communities. FEMS Microb. Ecol. 68, 1–13.
- Bredeweg, E.L. , Baker, S.E. 2020. Horizontal Gene Transfer in Fungi. In: Nevalainen, H. (eds) Grand Challenges in Fungal Biotechnology. Grand Challenges in Biology and Biotechnology. Springer, Cham. [CrossRef]
- Chazdon, R.L. 2008. Beyond Deforestation: Restoring Forests and Ecosystem Services on Degraded Lands. Science 3201458-1460.
- Clarke, K.R., Gorley, R.N. 2006. PRIMER v6: User manual PRIMER-E Ltd. Plymouth UK.
- Davidson, E.A., de Carvalho, C.J.R., Figueira, A.M., Ishida, F.Y.,et al., 2007. Recuperation of nitrogen cycling in Amazonian forests following agricultural abandonment. Nature 447, 995–998. [CrossRef]
- Eaton, W.D., McGee, K.M., Donnelly, R., Lemenze, A., Karas, O., Hajibabaei, M., 2019. Differences in the soil microbial community and carbon--use efficiency following development of Vochysia guatemalensis tree plantations in unproductive pastures in Costa Rica. Restor. Ecol. [CrossRef]
- Eaton, W.D., McGee, K.M., Alderfer, K., Jimenez, A.R., Hajibabaei, M. 2020a. Increase in abundance and decrease in richness of soil microbes following Hurricane Otto in three primary forest types in the Northern Zone of Costa Rica. Special Topics Issue: “Microbial Ecology of Changing Environments”, PLoS ONE 15: e0231187. [CrossRef]
- Eaton, W.D., McGee, K.M., Hoke, E., Lemenze, A., Hajibabaei, M. 2020b. Influence of Two Important Leguminous Trees on Their Soil Microbiomes and Nitrogen Cycle Activities in a Primary and Recovering Secondary Forest in the Northern Zone of Costa Rica. Soil Syst. 4,65. [CrossRef]
- Eaton, W.D., Hamilton, D.A. 2022. Enhanced carbon, nitrogen and associated bacterial community compositional complexity, stability, evenness, and differences within the tree-soils of Inga punctata along an age gradient of planted trees in reforestation plots. Plant Soil 484, 327–346.
- Eaton, W.D., Hamilton, D.A., Chen, W., Lemenze, A., Soteropoulos, P. 2024. Use of high throughput DNA analysis to characterize the nodule-associated bacterial community from four ages of Inga punctata trees in a Costa Rican cloud forest. AIMS Microbiol.10, 572-595. [CrossRef]
- Ettema. C., 2002. Spatial soil ecology. Trends Ecol.Evol.17, 177–183.
- . [CrossRef]
- Gardes, M. , Bruns, T.D. 1993. ITS primers with enhanced specificity for basidiomycetes -- application to the identification of mycorrhizae and rusts. Mol. Ecol. 2, 113–118.
- Griffiths, B.S., Philippot, L. 2013. Insights into the resistance and resilience of the soil microbial community. FEMS Microb. Rev. 37,112–129.
- Grimm, V., Wissel, C. 1997 Babel, or the ecological stability discussions: an inventory and analysis of terminology and a guide for avoiding confusion. Oecologia 109,323–334.
- Groppo, J.D., Lins, S.R.M., Camargo, P.B., Assad, E.D., Pinto, H.S., et al., 2015. Changes in soil carbon, nitrogen, and phosphorus due to land-use changes in Brazil. Biogeosci. 12,4765–4780.
- Hamilton, D., 2022. Offsetting Destruction: The Important Functional Contribution of Carbon Sequestration in the Restoration of a Tropical Forest in Monteverde, Costa Rica. Reference Module in Earth Systems and Environmental Sciences, Elsevier, 2022. ISBN 9780124095489. [CrossRef]
- He, H., Liu, Y., Hu, Y., Zhang, M., et al., 2020. Soil Microbial Community and Its Interaction with Soil Carbon Dynamics Following a Wetland Drying Process in Mu Us Sandy Land. Int J Environ Res Public Health 17, 4199.
- Kardol, P., Wardle, D.A. 2010. How understanding aboveground-belowground linkages can assist restoration ecology. Trends Ecol.Evol. 25, 670–679.
- Krych, L., Hansen, C.H.F., Hansen, A.K., van den Berg, F.W.J., Nielsen, D.S. 2013. Quantitatively Different, yet Qualitatively Alike: A meta-analysis of the mouse core gut microbiome with a view towards the human gut microbiome. PLoS ONE 8: e62578.
- Lai, H.R., Hall, J.S., Batterman, S.A., Turner, B.L., Breugel, M., 2018. Nitrogen fixer abundance has no effect on biomass recovery during tropical secondary forest succession. J. Ecol. 106,1415–1427.
- Lammel, D.R., Feigl, B.J., Cerri, C.C., Nüsslein, K., 2015. Specific microbial gene abundances and soil parameters contribute to C, N, and greenhouse gas process rates after land use change in Southern Amazonian Soils. Front Microbio 6. [CrossRef]
- Leblanc, H.A., McGraw, R.L., Nygren, P., Roux, C.L. 2005. Neotropical legume tree Inga edulis forms N2-fixing symbiosis with fast-growing Bradyrhizobium strains. Plant Soil 275,123–133.
- Liu, K-L., Porras-Alfaro, A., Kuske, C.R., Eichorst, S.A., et al., 2012. Accurate, rapid taxonomic classification of fungal large-subunit rRNA genes. Appl. Environ. Microbiol. 78,1523–1533.
- Liu, J. , Jia, X., Yan, W., Zhong, Y., Shangguan, Z. 2020. Changes in microbial community structure during secondary succession. Land Degrad. Develop. 1–16. [CrossRef]
- Lojka, B., Dumas, L., Preininger, D., Polesny, Z., Banout, J., 2010. The use and integration of Inga edulis in agroforestry systems in the amazon – review article. Agricult. Trop. Subtrop. 43, 352–359.
- Louca, S., Doebeli, M. (016. Transient dynamics of competitive exclusion in microbial communities. Environ Microbiol 18,1863-1874.
- Louca, S., Jacques, S.M.S., Pires, A.P.F., Leal, J.S., et al., 2016. High taxonomic variability despite stable functional structure across microbial communities. Nature Ecol. Evol. 1,0015.
- Louca, S., Polz, M.F., Mazel, F., Albright, M.B.N., et al., 2018. Function and functional redundancy in microbial systems. Nature Ecol. Evol. 2, 936–943.
- Lucas, Y., Santin, R.C., Da Silva, W.T.L., Merdy, P., et al., 2020. Soil sample conservation from field to lab for heterotrophic respiration assessment. MethodsX. 7:101039.
- Macedo, M.O., Resende, A.S., Garcia, P.C., Boddey, R.M., et al., 2008. Changes in soil C and N stocks and nutrient dynamics 13 years after recovery of degraded land using leguminous nitrogen-fixer trees. For. Ecol. Manag. 255,1516–1524.
- Martínez-García, L. B. , Korthals, G., Brussaard, L., Jørgensen, H. B., et al., 2018. Organic management and cover crop species steer soil microbial community structure and functionality along with soil organic matter properties. Agric. Ecosyst. Environ. 263, 7–17.
- Mori, A.S., Isbell, F., Fujii, S., Makoto, K., Matsuoka, S., Osono, T. 2015. Low multifunctional redundancy of soil fungal diversity at multiple scales. Ecol. Lett. 19, 249–259.
- Nemergut, D.R., Schmidt, S.K., Fukami, T., O’Neill, S.P., et al., 2013. Patterns and Processes of Microbial Community Assembly. Microbioland MolecBiol Rev 77, 342–356.
- Ortiz-Álvarez, R., Fierer, N., de los Ríos, A., Casamayor, E.O., Barberán, A., 2018. Consistent changes in the taxonomic structure and functional attributes of bacterial communities during primary succession. ISME Journal 12:1658–1667.
- Pinto-Figueroa, E.A. , Seddon, E., Yashiro, E., Buri, A., et al., 2019. Archaeorhizomycetes Spatial Distribution in Soils Along Wide Elevational and Environmental Gradients Reveal Co-Abundance Patterns with Other Fungal Saprobes and Potential Weathering Capacities. Front Microbiol 10, 656.
- Piotto, D., Montagnini, F., Ugalde, L., Kanninen, M. 2003. Performance of forest plantations in small and medium-sized farms in the Atlantic lowlands of Costa Rica. For. Ecol. Manag. 175,195–204.
- Põlme, S. , Abarenkov, K., Henrik, N.R., Lindah, B.D., et al., 2020. FungalTraits: a user-friendly traits database of fungi and funguslike stramenopiles. FungDivers 105,1–16.
- Rodrigues, J.L.M., Pellizari, V.H., Mueller, R., Baek, K., et al., 2013. Conversion of the Amazon rainforest to agriculture results in biotic homogenization of soil bacterial communities. PNAS 110, 988-993.
- Rosling, A. , Cox, F., Cruz--Martinez, K., Ihrmark, K., et al., 2011. Archaeorhizomycetes: unearthing an ancient class of ubiquitous soil fungi. Science 333: 876–879.
- Rosling, A. , Timling, I., Taylor, D.L. 2013. “Archaeorhizomycetes: patterns of distribution and abundance in soil,” in Genomics of Soil- and Plant-Associated Fungi. Soil Biology, eds B. Horwitz, P. Mukherjee, M. Mukherjee, and C. Kubicek (Heidelberg: Springer), 333–349. [CrossRef]
- Saiya-Cork, K.R., Sinsabaugh, R.L., Zak, D.R., 2002. The effects of long-term nitrogen deposition on extracellular enzyme activity in an Acer saccharum forest soil. Soil Biol. Biochem. 34,1309-1315.
- Schelhas, J., Sánchez-Azofeifa, G.A., 2006. Post-frontier forest change adjacent to Braulio Carrillo National Park Costa Rica. Human Ecol. 34, 407-431.
- Shade, A., Peter, H., Allison, S.D., Baho, D.L., et al., 2012. Fundamentals of microbial community resistance and resilience. Front. Microbiol. 3,417.
- Shim, J.E., Lee, T., Lee, I., 2017. From sequencing data to gene functions: co-functional network approaches. Animal Cells Syst 21, 77–83.
- Sinsabaugh, R.L., 2010. Phenol oxidase, peroxidase and organic matter dynamics of soil. Soil Biol. Biochem. 42, 391–404.
- Six, J., Frey, S.D., Thiet, R.K., Batten, K.M. 2006. Bacterial and Fungal Contributions to Carbon Sequestration in Agroecosystems. Soil Sci. Soc. Am. J., 70: 555-569.
- Spohn, M. 2015. Microbial respiration per unit microbial biomass depends on litter layer carbon-to-nitrogen ratio, Biogeosci. 12, 817–823.
- Stürmer, S.L. , Kemmelmeier, K., 2021. The Glomeromycota in the Neotropics. Front. Microbiol.11,553679.
- Taylor, B.N., Chazdon, R.L., Bachelot, B., Menge, D.N.L., 2017. Nitrogen-fixer trees inhibit growth of regenerating Costa Rican rainforests. PNAS 114, 8817–8822.
- Tedersoo, L. , Bahram, M., Puusepp, R., Nilsson, R.H., et al., 2017. Novel Soil-Inhabiting Clades Fill Gaps in the Fungal Tree of Life. Microbiome 5,42.
- Tilman, D., Reich, P.B., Knops, J.M., 2006. Biodiversity and ecosystem stability in a decade-long grassland experiment. Nature 441,629-32.
- Turatsinze, A.N. , Kang, B., Zhu, T., Hou, F., et al., 2021. Soil Bacterial and Fungal Composition and Diversity Responses to Seasonal Deer Grazing in a Subalpine Meadow. Diversity 13:84.
- Turbé, A., De Toni, A., Benito, P., Lavelle, P., et al., 2010. Soil biodiversity: functions, threats and tools for policy makers. Bio Intelligence Service, IRD, and NIOO, Technical Report 2010-49 for European Commission (DG Environment).
- van der Heijden, M.G.A., Bardgett, R.D., van Straalen, N.M., 2008. The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol. Lett. 11, 296–310.
- Wagg, C., Jansa, J., Schmid, B., van der Heijden, M.G., 2011. Belowground biodiversity effects of plant symbionts support aboveground productivity. Ecol. Lett. 14.1001-9.
- Wallenstein, M.D., Burns, R.G., 2011. Ecology of extracellular enzyme activities and organic matter degradation in soil: a complex community-driven process. In: Dick R (ed) Methods of Soil Enzymology. 9:35-55.
- Wallenstein, M.D., Hall, E.K. 2011. A trait-based framework for predicting when and where microbial adaptation to climate change will affect ecosystem functioning. Biogeochem. 109: 35–47.
- Weiss, S., Xu, Z.Z., Peddada, S., Amir, A., et al., 2017. Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 5. [CrossRef]
- Xu, C.Y., Du, C., Jian, J.S., Hou, L., et al., 2021. The interplay of labile organic carbon, enzyme activities and microbial communities of two forest soils across seasons. Science Reports 11: 5002.
- Xu, H., Detto, M., Li, Y.,He, F., Fang, S., 2019. Do N-fixing legumes promote neighbor diversity in the tropics? J Ecol. 107, 229–239.
Figure 1.
Differences in the MPS of the DNA sequences and the Stability Index
S index of the Most Common Fungal Taxa (subset of Total Fungi with MPS > 1.0%) and the Complex C Decomposer (CCDec) Taxa within soils of 4 year-old, 8 year-old 11 year-old and old secondary
I. punctata trees (Inga 4, 8, 11 and Old Inga) and an adjacent pasture (PAS) within a reforestation site in Monteverde, Costa Rica.
Figure 1A) shows the MPS of the DNA Sequences and
Figure 1B) shows the Stability
S index for the two communities.
Figure 1.
Differences in the MPS of the DNA sequences and the Stability Index
S index of the Most Common Fungal Taxa (subset of Total Fungi with MPS > 1.0%) and the Complex C Decomposer (CCDec) Taxa within soils of 4 year-old, 8 year-old 11 year-old and old secondary
I. punctata trees (Inga 4, 8, 11 and Old Inga) and an adjacent pasture (PAS) within a reforestation site in Monteverde, Costa Rica.
Figure 1A) shows the MPS of the DNA Sequences and
Figure 1B) shows the Stability
S index for the two communities.
Figure 2.
Analysis of the Margalef’s Richness index
d (± standard deviations) and the Stability index
S of the Richness
d index the community composition of the Total Fungi and the Most Common Fungi within soils of 4 year-old, 8 year-old 11 year-old and old secondary
I. punctata trees (Inga 4, 8, 11 and Old Inga) and an adjacent pasture (PAS) within a reforestation site in Monteverde, Costa Rica.
Figure 2A) shows the Margalef’s richness and
Figure 2B) shows the Stability
S index for the two communities.
Figure 2.
Analysis of the Margalef’s Richness index
d (± standard deviations) and the Stability index
S of the Richness
d index the community composition of the Total Fungi and the Most Common Fungi within soils of 4 year-old, 8 year-old 11 year-old and old secondary
I. punctata trees (Inga 4, 8, 11 and Old Inga) and an adjacent pasture (PAS) within a reforestation site in Monteverde, Costa Rica.
Figure 2A) shows the Margalef’s richness and
Figure 2B) shows the Stability
S index for the two communities.
Table 1.
Mean values (± standard deviations) of the Biomass-C, Respiration (Resp) and qCO2 from soils of Inga punctata trees planted 4, 8, and 11 years before sampling, Old Inga trees (> 50 years old) and an adjacent pasture (PAS) within a restoration area in Monteverde, Costa Rica (1A). Results of ANOVA tests for differences between the means of these metrics between the different soils (1B).
Table 1.
Mean values (± standard deviations) of the Biomass-C, Respiration (Resp) and qCO2 from soils of Inga punctata trees planted 4, 8, and 11 years before sampling, Old Inga trees (> 50 years old) and an adjacent pasture (PAS) within a restoration area in Monteverde, Costa Rica (1A). Results of ANOVA tests for differences between the means of these metrics between the different soils (1B).
| A |
| Habitat |
Biomass-C (µgCO2/g soil) |
Resp (µgCO2/g soil) |
qCO2 (Resp/Biomass-C) |
| PAS |
806.32 ± 64.57 |
261. 04 ± 22.58 |
0.33 ± 0.06 |
| Inga 4 |
547.50 ± 31.05 |
180.67 ± 19.49 |
0.33 ± 0.04 |
| Inga 8 |
540.67 ± 17.13 |
181.33 ± 14.17 |
0.34 ± 0.03 |
| Inga 11 |
725.33 ± 58.29 |
201.65 ± 28.90 |
0.28 ± 0.03 |
| Old Inga |
934.83 ± 70.06 |
209.83 ± 30.47 |
0.22 ± 0.04 |
| B |
| B |
p values for comparisons of means |
| Comparisons |
Biomass C |
Respiration |
qCO2
|
| PAS to Inga 4 |
< 0.0001 |
< 0.0001 |
1 |
| PAS to Inga 8 |
< 0.0001 |
< 0.0001 |
1 |
| PAS to Inga 11 |
0.0001 |
0.022 |
0.034 |
| PAS to Old Inga |
0.031 |
0.008 |
0.006 |
| Old Inga to Inga 4 |
< 0.0001 |
0.054 |
0.0002 |
| Old Inga to Inga 8 |
< 0.0001 |
0.051 |
0.0005 |
| Old Inga to Inga 11 |
0.0002 |
0.644 |
0.015 |
| Inga 11 to Inga 4 |
0.0001 |
0.145 |
0.006 |
| Inga 11 to Inga 8 |
0.0001 |
0.143 |
0.033 |
| Inga 8 to Inga 4 |
0.6472 |
0.942 |
0.635 |
Table 2.
The 17 Total Fungal taxa with MPS > 1%, considered the Most Common Fungal taxa, within soils of 4 year-old, 8 year-old 11 year-old and old secondary I. punctata trees (Inga 4, 8, 11 and Old Inga) and an adjacent pasture (PAS), and the CCDec within a reforestation site in Monteverde, Costa Rica. The CCDec and WRT taxa were collectively called “CCDec”. (Key To Functions: Dec: simple C decomposer; RA: root associated; SAP: saprobe; CCDec: complex C decomposer; PAR: endoparasite; WRT: wood rot).
Table 2.
The 17 Total Fungal taxa with MPS > 1%, considered the Most Common Fungal taxa, within soils of 4 year-old, 8 year-old 11 year-old and old secondary I. punctata trees (Inga 4, 8, 11 and Old Inga) and an adjacent pasture (PAS), and the CCDec within a reforestation site in Monteverde, Costa Rica. The CCDec and WRT taxa were collectively called “CCDec”. (Key To Functions: Dec: simple C decomposer; RA: root associated; SAP: saprobe; CCDec: complex C decomposer; PAR: endoparasite; WRT: wood rot).
| Taxa |
Function |
MPS PAS |
MPS Inga 4 |
MPS Inga 8 |
MPS Inga 11 |
MPS Old Inga |
| Apiotrichum |
CCDec |
5.37 |
0.19 |
6.16 |
43.61 |
41.96 |
| Archaeorhizomyces |
RADec |
32.11 |
3.74 |
5.31 |
2.02 |
3.53 |
| Chaetomium |
CCDecWRT |
0.29 |
1.73 |
0.99 |
0.3 |
0.57 |
| Dipodascus |
CCDec |
4.21 |
0 |
0.56 |
2.02 |
0.54 |
| Geotrichum |
CCDecWRT |
0.18 |
0 |
0.07 |
1.28 |
0.22 |
| Glomeromycota |
ARM |
4.8 |
19.29 |
18.41 |
12.65 |
15.93 |
| Leohumicola |
SAPDec |
2.46 |
0.05 |
0.17 |
0 |
0.38 |
| Lipomyces |
CCDec |
0 |
0.23 |
1.54 |
1.01 |
0.99 |
| Mortierella |
CCDec |
0.37 |
1.15 |
0.38 |
0.02 |
0.45 |
| Phialocephala |
CCDec |
1.87 |
3.56 |
2.23 |
1.84 |
0.89 |
| Pleosporales |
SAPDec |
1.64 |
0.46 |
0.38 |
0.28 |
0.2 |
| Pyrenochaetopsis |
CCDec |
0.79 |
0.23 |
1.94 |
0.77 |
0 |
| Rozella |
PAR |
7.97 |
0.76 |
1.92 |
1.61 |
0.84 |
| Saitozyma |
CCDec |
0.74 |
29.42 |
28.89 |
7.68 |
8.77 |
| Sordariaceae |
CCDec |
1.44 |
7.98 |
11.6 |
0.29 |
1.05 |
| Starmerella |
SAPDec |
0 |
2.95 |
2.35 |
8.13 |
11.44 |
| Tremella |
CCDec |
0.04 |
0.67 |
1.5 |
0.22 |
8.21 |
| MPS of CCDec Taxa |
|
15.30 |
47.16 |
55.86 |
59.04 |
63.65 |
Table 3.
Differences in the fungal community compositions were determined by ANOSIM and Canonical Analysis of Principal Coordinates (CAP) methods, applied to the MPS data of the Total Fungal taxa and all the CCDec Fungal taxa from soils of Inga punctata trees planted 4, 8, and 11 years before sampling, Old Inga trees (> 50 years old) and an adjacent pasture (PAS) within a restoration area in Monteverde, Costa Rica.
Table 3.
Differences in the fungal community compositions were determined by ANOSIM and Canonical Analysis of Principal Coordinates (CAP) methods, applied to the MPS data of the Total Fungal taxa and all the CCDec Fungal taxa from soils of Inga punctata trees planted 4, 8, and 11 years before sampling, Old Inga trees (> 50 years old) and an adjacent pasture (PAS) within a restoration area in Monteverde, Costa Rica.
| ANOSIM of Total Fungal Taxa |
|
CAP of Total Fungal Taxa |
| Global R = 0.404 |
|
|
|
CAP model p value = 0.0024 |
| Global p value 0.0001 |
|
|
|
|
|
| Pairwise Groups |
R Statistic |
p Values |
|
Comparisons |
R2 Value |
Strength of Diff |
| PAS and Inga 4 |
0.369 |
0.015 |
|
PAS and Inga 4 |
0.568 |
moderate |
| PAS and Inga 8 |
0.439 |
0.002 |
|
PAS and Inga 8 |
0.568 |
moderate |
| PAS and Inga 11 |
0.576 |
0.002 |
|
PAS and Inga 11 |
0.791 |
strong |
| PAS and Old Inga |
0.564 |
0.009 |
|
PAS and Old Inga |
0.791 |
strong |
| Inga 4 and Inga 8 |
0.097 |
0.182 |
|
Inga 4 and Inga 8 |
0.044 |
No difference |
| Inga 4 and Inga 11 |
0.669 |
0.002 |
|
Inga 4 and Inga 11 |
0.791 |
strong |
| Inga 4 and Old Inga |
0.564 |
0.016 |
|
Inga 4 and Old Inga |
0.791 |
strong |
| Inga 8 and Inga 11 |
0.323 |
0.019 |
|
Inga 8 and Inga 11 |
0.449 |
weak |
| Inga 8 and Old Inga |
0.503 |
0.053 |
|
Inga 8 and Old Inga |
0.568 |
moderate |
| Inga 11 and Old Inga |
0.201 |
0.037 |
|
Inga 11 and Old Inga |
0.044 |
No difference |
| |
|
|
|
|
|
|
| ANOSIM of CCDec Fungal Taxa |
|
CAP of CCDec Fungal Taxa |
| Global R = 0.473 |
|
|
|
CAP model p value = 0.0006 |
|
| Global p value 0.0001 |
|
|
|
|
|
| Pairwise Groups |
R value |
p Value |
|
Pairwise Groups |
R2 Value |
Strength of Diff |
| PAS and Inga 4 |
0.379 |
0.019 |
|
PAS and Inga 4 |
0.582 |
moderate |
| PAS and Inga 8 |
0.533 |
0.004 |
|
PAS and Inga 8 |
0.687 |
moderate |
| PAS and Inga 11 |
0.654 |
0.002 |
|
PAS and Inga 11 |
0.745 |
strong |
| PAS and Old Inga |
0.616 |
0.006 |
|
PAS and Old Inga |
0.745 |
strong |
| Inga 4 and Inga 8 |
0.027 |
0.359 |
|
Inga 4 and Inga 8 |
0.019 |
No difference |
| Inga 4 and Inga 11 |
0.853 |
0.002 |
|
Inga 4 and Inga 11 |
0.745 |
strong |
| Inga 4 and Old Inga |
0.648 |
0.024 |
|
Inga 4 and Old Inga |
0.745 |
strong |
| Inga 8 and Inga 11 |
0.368 |
0.039 |
|
Inga 8 and Inga 11 |
0.233 |
weak |
| Inga 8 and Old Inga |
0.607 |
0.002 |
|
Inga 8 and Old Inga |
0.582 |
moderate |
| Inga 11 and Old Inga |
0.320 |
0.017 |
|
Inga 11 and Old Inga |
0.019 |
No difference |
Table 4.
The Percent (%) Contribution of the top 10 Fungal taxa, their functions, and the CCDec Fungal Taxa (% Contrib CCDec) to the Total Fungal community composition within soils of 4, 8, and 11 year-old and Old I. punctata trees (Inga 4, 8, 11 and Old Inga) and adjacent pasture (PAS) within a reforestation site in Monteverde, Costa Rica.
Table 4.
The Percent (%) Contribution of the top 10 Fungal taxa, their functions, and the CCDec Fungal Taxa (% Contrib CCDec) to the Total Fungal community composition within soils of 4, 8, and 11 year-old and Old I. punctata trees (Inga 4, 8, 11 and Old Inga) and adjacent pasture (PAS) within a reforestation site in Monteverde, Costa Rica.
| % Contribution of Fungal Taxa to the Total Fungal Community Composition in PAS Soils |
|
% Contribution of Fungal Taxa to the Total Fungal Community Composition in Inga 4 Soils |
| Species |
% Contribution |
Function |
|
Species |
% Contribution |
Function |
| Archaeorhizomyces |
18.22 |
RADec |
|
Glomeromycota |
22.63 |
ARM |
| Rozella |
13.69 |
PAR |
|
Saitozyma |
16.48 |
CCDec |
| Apiotrichum |
9.48 |
CCDec |
|
Phialocephala |
9.84 |
RACCDec |
| Glomeromycota |
9.45 |
ARM |
|
Sordariaceae |
9.39 |
CCDec |
| Dipodascus |
7.32 |
CCDEC |
|
Archaeorhizomyces |
6.25 |
RADec |
| Leohumicola |
3.91 |
SAPDec |
|
Starmerella |
5.96 |
SAPDec |
| Pleosporales |
3.58 |
SAPDec |
|
Chaetomium |
3.69 |
CCDecWRT |
| Sordariaceae |
3.48 |
CCDec |
|
Mortierella |
3.01 |
CCDec |
| Phialocephala |
2.98 |
CCDec |
|
Rozella |
2.61 |
PAR |
| Pyrenochaetopsis |
1.85 |
CCDec |
|
Pleosporales |
1.78 |
SAPDec |
| Total % Contr CCDec |
25.11 |
|
|
Total % Contr CCDec |
42.41 |
|
| % Contribution of Fungal Taxa to the Total Fungal Community Composition In Inga 8 Soils |
|
% Contribution of Fungal Taxa to the Total Fungal Community Composition in Inga 11 Soils |
| Species |
% Contribution |
Function |
|
Species |
% Contribution |
Function |
| Glomeromycota |
18.65 |
ARM |
|
Apiotrichum |
22.38 |
CCDec |
| Saitozyma |
15.35 |
CCDec |
|
Glomeromycota |
16.28 |
ARM |
| Archaeorhizomyces |
11.12 |
RADec |
|
Saitozyma |
10.95 |
CCDec |
| Sordariaceae |
9.85 |
CCDec |
|
Starmerella |
10.45 |
SAPDec |
| Apiotrichum |
6.99 |
CCDec |
|
Dipodasus |
8.16 |
CCDec |
| Phialocephela |
6.06 |
CCDec |
|
Archaeorhizomyces |
6.24 |
RADec |
| Starmerella |
5.18 |
SAPDec |
|
Phialocephala |
5.16 |
CCDec |
| Rozella |
5.11 |
PAR |
|
Rozella |
4.82 |
PAR |
| Pyrenochaetopsis |
4.63 |
CCDec |
|
Geotrichum |
4.35 |
CCDecWRT |
| Lipomyces |
4.14 |
CCDec |
|
Lipomyces |
4.13 |
CCDec |
| Total % Contr CCDec |
47.02 |
|
|
Total % Contr CCDec |
55.13 |
|
| % Contribution of Fungal Taxa to the Total Fungal Community Composition in Old Inga Soils |
|
|
|
|
| Species |
% Contribution |
Function |
|
|
|
|
| Apiotrichum |
25.53 |
CCDec |
|
|
|
|
| Glomeromycota |
17.62 |
ARM |
|
|
|
|
| Starmerella |
14.77 |
SAPDec |
|
|
Key To Functions |
| Saitozyma |
10.24 |
CCDec |
Dec: simple C decomposer |
| Tremella |
10.23 |
CCDec |
|
|
RA: root associated |
| Archaeorhizomyces |
8.19 |
RADec |
|
|
SAP: saprobe |
| Sordariaceae |
5.84 |
CCDec |
|
|
CCDec: complex C decomposer |
| Rozella |
4.24 |
PAR |
|
|
PAR: endoparasite |
| Lipomyces |
3.85 |
CCDec |
|
|
WRT: wood rot |
| Phialocephala |
3.07 |
CCDec |
|
|
|
| Total % Contr CCDec |
58.76 |
|
|
|
|
|
Table 5.
The results of the distance-based linear modeling (DistLM) sequential tests showing the Fungal taxa having the greatest effects on the Biomass-C, Respiration, and qCO2 within the soils beneath 4 year-old, 8 year-old 11 year-old and old secondary I. punctata trees and in an adjacent pasture within a reforestation site in Monteverde, Costa Rica. Shown with the most influential taxa are the DistLM Pseudo-F and p values, and the percent variation (% Var) and cumulative percent variation (Cumul. % Var) of the carbon metrics best explained by the specific taxa.
Table 5.
The results of the distance-based linear modeling (DistLM) sequential tests showing the Fungal taxa having the greatest effects on the Biomass-C, Respiration, and qCO2 within the soils beneath 4 year-old, 8 year-old 11 year-old and old secondary I. punctata trees and in an adjacent pasture within a reforestation site in Monteverde, Costa Rica. Shown with the most influential taxa are the DistLM Pseudo-F and p values, and the percent variation (% Var) and cumulative percent variation (Cumul. % Var) of the carbon metrics best explained by the specific taxa.
| Effects on All C metrics |
AICc |
Pseudo-F |
p value |
% Var |
Cumul. % Var |
| Saitozyma |
24.793 |
11.03 |
0.0001 |
29.79 |
29.79 |
| Apiotrichum |
20.996 |
6.3272 |
0.0093 |
14.18 |
43.97 |
| |
|
|
|
|
|
| Effects on Biomass-C |
AICc |
Pseudo-F |
p value |
% Var |
Cumul. % Var |
| Saitozyma |
16.549 |
24.656 |
0.0003 |
48.67 |
48.67 |
| |
|
|
|
|
|
| Effects on Respiration |
AICc |
Pseudo-F |
p value |
% Var |
Cumul. % Var |
| Saitozyma |
16.362 |
17.637 |
0.0003 |
40.42 |
40.42 |
| Apiotrichum |
11.137 |
7.9661 |
0.0096 |
14.40 |
54.82 |
| |
|
|
|
|
|
| Effects on qCO2
|
AICc |
Pseudo-F |
p value |
% Var |
Cumul. % Var |
| Apiotrichum |
7.7003 |
9.1392 |
0.0055 |
26.00 |
26.00 |
| Saitozyma |
3.4048 |
6.8898 |
0.0153 |
15.99 |
41.99 |
| |
|
|
|
|
|
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).