Submitted:
03 August 2024
Posted:
06 August 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Materials and Method
Plant Materials
Experimental Condition
Growth Conditions
Measurements to Determine Drought Intensity
Soil Water Conditions
Relative Water Content of the Leaf Blade
Phenotypic Evaluation
Phenotypic Data Analysis
QTL Analysis
Comparison between Top- and Bottom-DRI Progeny
Review of Rice DRI QTL Papers
Selection of Putative Genes for Drought Resistance
Results
Soil Water Conditions and Relative Water Content of the Leaf Blade
Phenotypic Analysis
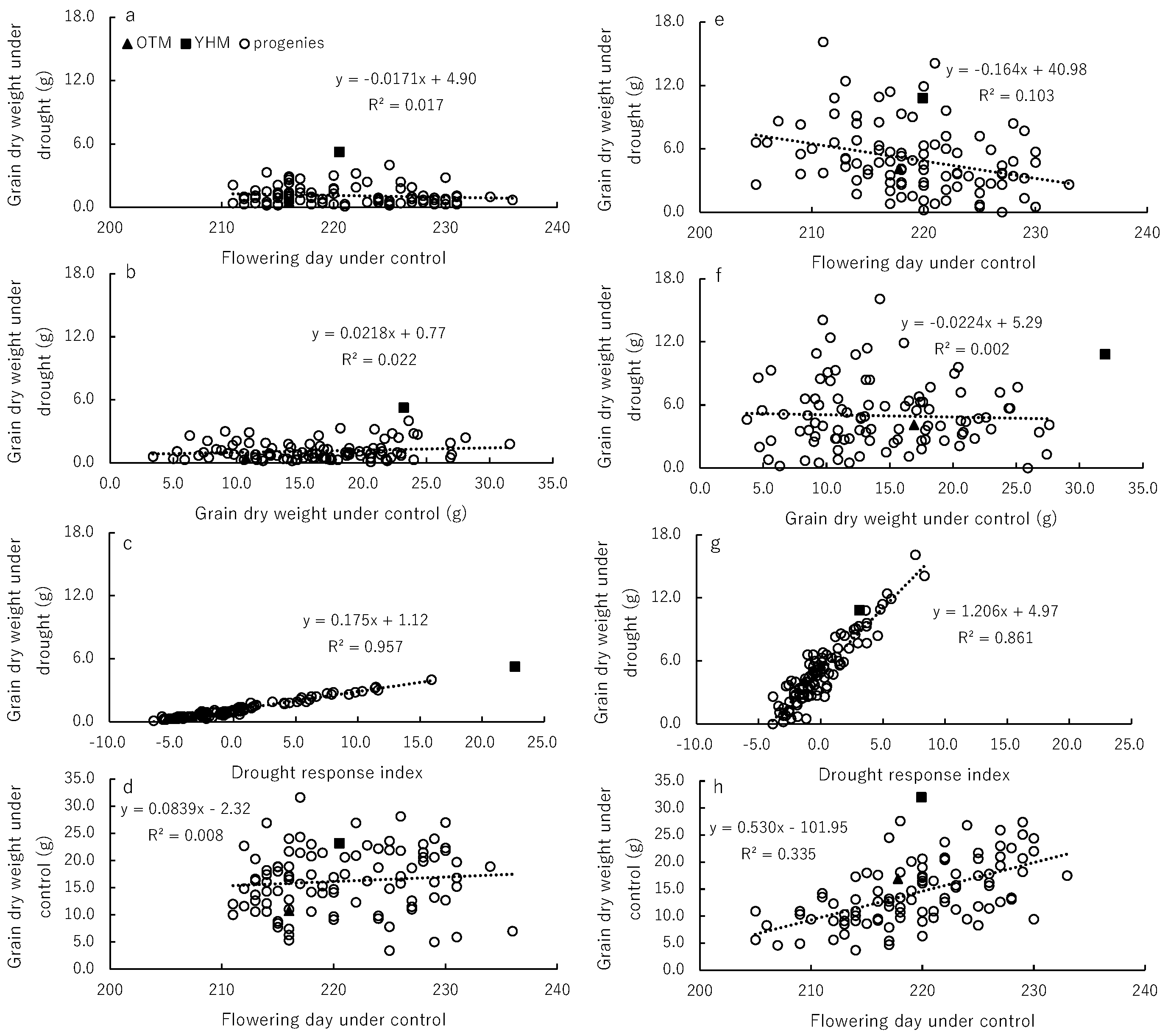
Grain Dry Weight and Drought Response Index
Yield Formation
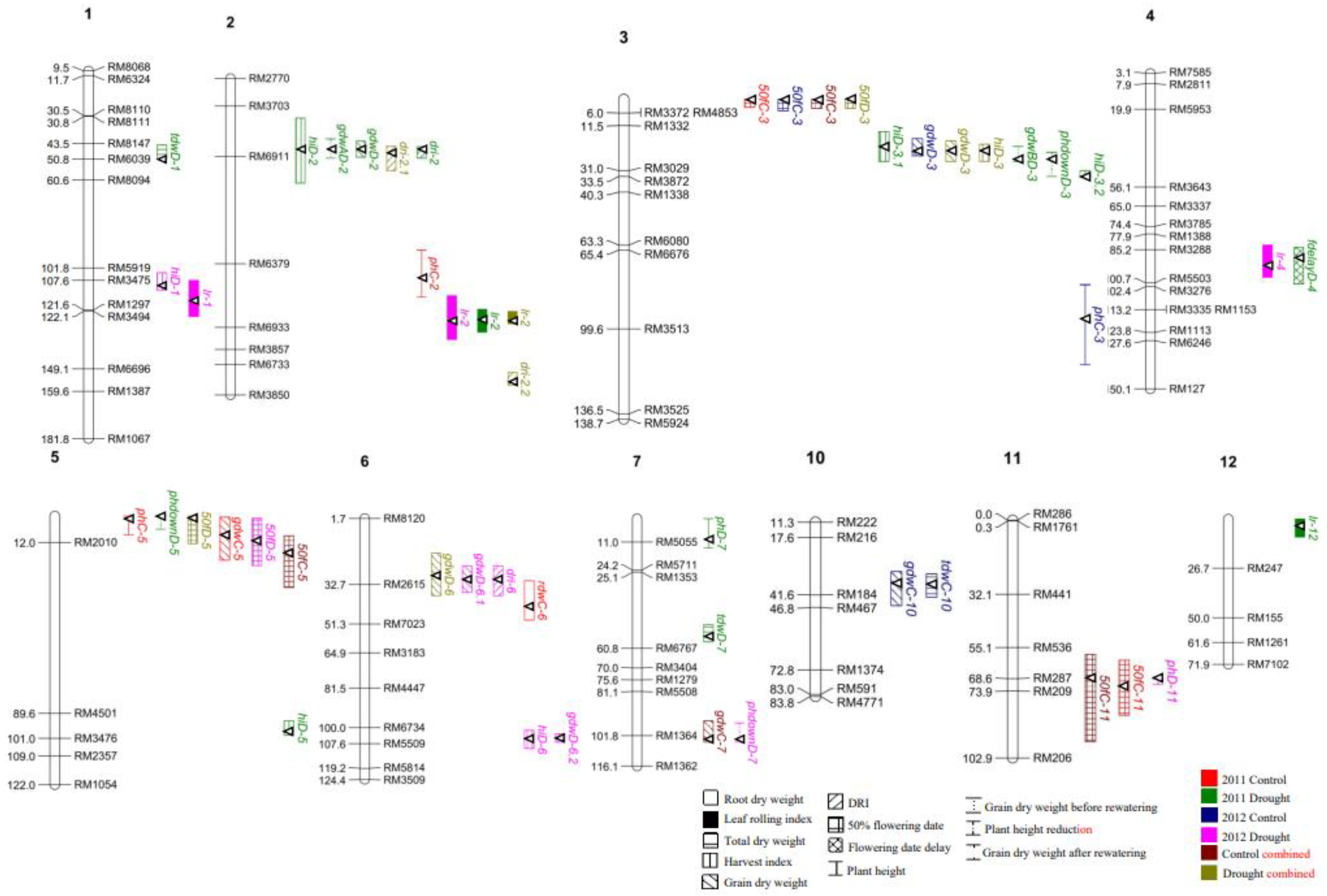
Genomic Analysis
Main-Effect QTLs for DRI
QTL-by-Environment Interaction
Performances of Top and Bottom 10 DRI Lines
Review of Previous Studies on QTLs for DRI
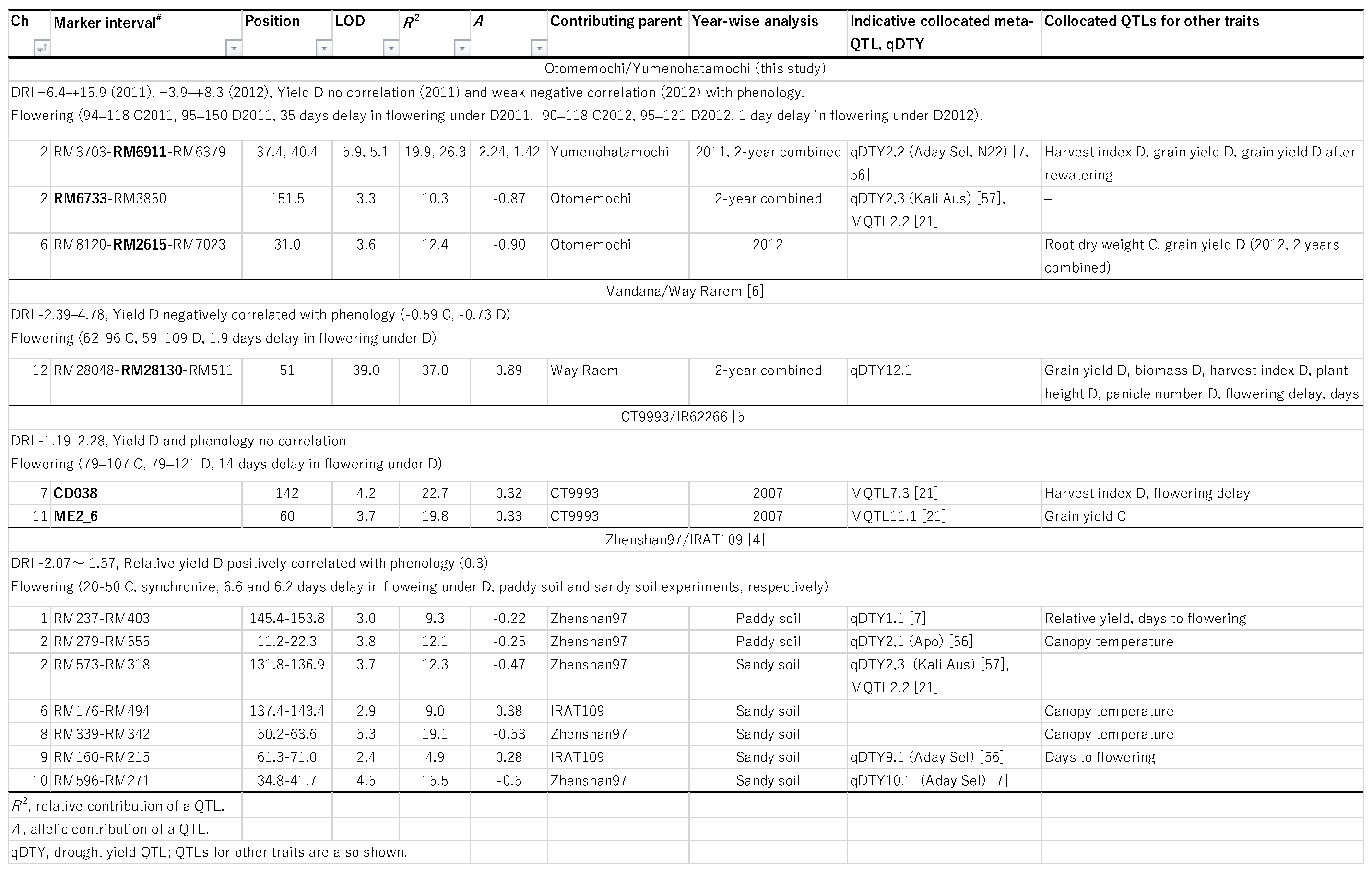 |
Putative Genes for Drought Resistance
Discussion
Phenotypic Assessment of DRI
Genetic Assessment of DRI with QTLs for Other Yield-Related Traits and Drought Resistance Genes
Supplementary Materials
Acknowledgments
Disclosure statement
References
- Ouk, M.; Basnayake, J.; Tsubo, M.; Fukai, S.; Fischer, K.S.; Cooper, M.; Nesbitt, H. Use of drought response index for identification of drought tolerant genotypes in rainfed lowland rice. Field Crops Res. 2006, 99, 48–58. [CrossRef]
- Bidinger, F.; Mahalakshmi, V.; Rao, G. Assessment of drought resistance in pearl millet (Pennisetum americanum (L.) Leeke). II. Estimation of genotype response to stress. Aust. J. Agric. Res. 1987, 38, 49–59. [CrossRef]
- Silim, S.N.; Saxena, M.C. Adaptation of spring-sown chickpea to the Mediterranean Basin. I. Response to moisture supply. Field Crops Res. 1993, 34, 121–136. [CrossRef]
- Yue, B.; Xiong, L.; Xue, W.; Xing, Y.; Luo, L.; Xu, C. Genetic analysis for drought resistance of rice at reproductive stage in field with different types of soil. Theor. Appl. Genet. 2005, 111, 1127–1136. [CrossRef]
- Sellamuthu, R.; Liu, G.F.; Ranganathan, C.B.; Serraj, R. Genetic Analysis and validation of quantitative trait loci associated with reproductive-growth traits and grain yield under drought stress in a doubled haploid line population of rice (Oryza sativa L.). Field Crops Res. 2011, 124, 46–58. [CrossRef]
- Bernier, J.; Kumar, A.; Ramaiah, V.; Spaner, D.; Atlin, G. A Large-effect QTL for grain yield under reproductive-stage drought stress in upland rice. Crop Sci. 2007, 47, 507–518. [CrossRef]
- Kumar, A.; Dixit, S.; Ram, T.; Yadaw, R.B.; Mishra, K.K.; Mandal, N.P. Breeding high-yielding drought-tolerant rice: genetic variations and conventional and molecular approaches. J. Exp. Bot. 2014, 65, 6265–6278. [CrossRef]
- Kamoshita, A.; Babu, R.C.; Boopathi, N.M.; Fukai, S. Phenotypic and genotypic analysis of drought-resistance traits for development of rice cultivars adapted to rainfed environments. Field Crops Res. 2008, 109, 1–23. [CrossRef]
- Lafitte, H.R.; Champoux, M.C.; McLaren, G.; O’Toole, J.C. Rice root morphological traits are related to isozyme group and adaptation. Field Crops Res. 2001, 71, 57–70. [CrossRef]
- Ikeda, H.; Kamoshita, A.; Manabe, T. Genetic analysis of rooting ability of transplanted rice (Oryza sativa L.) under different water conditions. J. Exp. Bot. 2007, 58, 309–318. [CrossRef]
- Hirasawa, H.; Nemoto, H.; Suga, R.; Ishihara, M.; Hirayama, M.; Okamoto, K.; Miyamoto, M. Breeding of a new upland rice variety ‘Yumenohatamochi’ with high drought resistance and good eating quality. Breed. Sci. 1998, 48, 415–419. [CrossRef]
- Kato, Y.; Abe, J.; Kamoshita, A.; Yamagishi, J. Genotypic variation in root growth angle in rice (Oryza sativa L.) and its association with deep root development in upland fields with different water regimes. Plant Soil 2006, 287, 117–129. [CrossRef]
- Kato, Y.; Kamoshita, A.; Yamagishi, J.; Imoto, H.; Abe, J. Growth of rice (Oryza sativa L.) cultivars under upland conditions with different levels of water supply 3. root system development, soil moisture change and plant water status. Plant Prod. Sci. 2007, 10, 3–13. [CrossRef]
- IRRI. Standard Evaluation System (SES) for Rice, International Rice Research Institute, 2002.pp56. http://www.knowledgebank.irri.org/images/docs/rice-standard-evaluation-system.pdf (retrieved on 23 March 2024).
- Bidinger, F.R.; Mahalakshmei, V.; Talukdar, B.S.; Alagarswamy, G. Improvement of drought resistant in pearl millet. In: Drought Resistance in crops with emphasis on rice. 1982, IRRI, Los Banos, Philippines, 357-375.
- Manabe, T.; Hirayama, M.; Miyamoto, M.; Okamoto, K.; Okano, K.; Ishii, T. Mapping of quantitative trait loci associated with deep root dry weight in rice. In: Proceedings of the 10th International Congress of SABRAO 2005, Tsukuba, Japan, 22–23 August 2005; D-18.
- Wang, S.; Basten, C. J.; Zeng, Z. B. Windows QTL Cartographer 2.5, 2012. Department of Statistics, North Carolina State University, Raleigh, NC. (http://statgen.ncsu.edu/qtlcart/WQTLCart.htm).
- Voorrips, R.E. Mapchart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [CrossRef]
- Wang, D.L.; Zhu, J.; Li, Z.K.; Paterson, A.H. Mapping QTLs with epistatic effects and QTL × environment interactions by mixed linear model approaches. Theor. Appl. Genet. 1999, 99, 1255–1264. [CrossRef]
- Nguyen, T. H. A.; Kamoshita, A.; Ramalingam, P.; Y, P. Genetic analysis of root vascular traits in a population from two temperate japonica rice ecotypes. Plant Prod. Sci. 2022, 25, 320–336. [CrossRef]
- Selamat, N.; Nadarajah, K.K. Meta-analysis of quantitative traits loci (QTL) identified in drought response in rice (Oryza sativa L.). Plants 2021, 10, 716. [CrossRef]
- Song, X.J.; Huang, W.; Shi, M.; Zhu, M.Z.; Lin, H.X. A QTL for rice grain width and weight encodes a previously unknown RING-Type E3 ubiquitin ligase. Nat. Genet. 2007, 39, 623–630. [CrossRef]
- Yamaguchi, K.; Yamamoto, T.; Segami, S.; Horikawa, M.; Chaya, G.; Kitano, H.; Iwasaki, Y.; Miura, K. Gw2 mutation increases grain width and culm thickness in rice (Oryza sativa L.). Breed. Sci. 2020, 70, 456–461. [CrossRef]
- Piao, R.; Jiang, W.; Ham, T.H.; Choi, M.S.; Qiao, Y.; Chu, S.H.; Park, J.H.; Woo, M.O.; Jin, Z.; An, G.; et al. Map-based cloning of the ERECT PANICLE 3 gene in rice. Theor. Appl. Genet. 2009, 119, 1497–1506. [CrossRef]
- Li, M.; Tang, D.; Wang, K.; Wu, X.; Lu, L.; Yu, H.; Gu, M.; Yan, C.; Cheng, Z. Mutations in the F-Box gene LARGER PANICLE improve the panicle architecture and enhance the grain yield in rice. Plant Biotechnol. J. 2011, 9, 1002–1013. [CrossRef]
- Coudert, Y.; Bès, M.; Van Anh Le, T.; Pré, M.; Guiderdoni, E.; Gantet, P. Transcript profiling of Crown Rootless 1 mutant stem base reveals new elements associated with crown root development in rice. BMC Genomics 2011, 12, 1–12. [CrossRef]
- Xu, L.; Zhao, H.; Ruan, W.; Deng, M.; Wang, F.; Peng, J.; Luo, J.; Chen, Z.; Yi, K. ABNORMAL INFLORESCENCE MERISTEM 1 functions in salicylic acid biosynthesis to maintain proper reactive oxygen species levels for root meristem activity in rice. Plant Cell 2017, 29, 560–574. [CrossRef]
- Airoldi, C.A.; Della Rovere, F.; Falasca, G.; Marino, G.; Kooiker, M.; Altamura, M.M.; Citterio, S.; Kater, M.M. The Arabidopsis BET bromodomain factor GTE4 is involved in maintenance of the mitotic cell cycle during plant development. Plant Physiol. 2010, 152, 1320–1334. [CrossRef]
- Richmond, T.A.; Bleecker, A.B. A defect in β-oxidation causes abnormal inflorescence development in Arabidopsis. Plant Cell 1999, 11, 1911–1923. [CrossRef]
- Kawakatsu, T.; Takaiwa, F. Differences in transcriptional regulatory mechanisms functioning for free lysine content and seed storage protein accumulation in rice grain. Plant Cell Physiol. 2010, 51, 1964–1974. [CrossRef]
- Goto, F.; Yoshihara, T.; Shigemoto, N.; Toki, S.; Takaiwa, F. Iron fortification of rice seed by the soybean Ferritin gene. Nat. Biotechnol. 1999, 17, 282–286. [CrossRef]
- Tian, X.; Wang, Z.; Li, X.; Lv, T.; Liu, H.; Wang, L.; Niu, H.; Bu, Q. Characterization and functional analysis of Pyrabactin resistance-like abscisic acid receptor family in rice. Rice 2015, 8. [CrossRef]
- Qi, Y.; Yamauchi, Y.; Ling, J.; Kawano, N.; Li, D.; Tanaka, K. Cloning of a Putative Monogalactosyldiacylglycerol Synthase gene from rice (Oryza Sativa L.) plants and its expression in response to submergence and other stresses. Planta 2004, 219, 450–458. [CrossRef]
- Lian, H.L.; Yu, X.; Ye, Q.; Ding, X.S.; Kitagawa, Y.; Kwak, S.S.; Su, W.A.; Tang, Z.C. The role of Aquaporin RWC3 in drought avoidance in rice. Plant Cell Physiol. 2004, 45, 481–489. [CrossRef]
- Liu, S.; Fukumoto, T.; Gena, P.; Feng, P.; Sun, Q.; Li, Q.; Matsumoto, T.; Kaneko, T.; Zhang, H.; Zhang, Y.; et al. Ectopic expression of a rice plasma membrane intrinsic protein (OsPIP1;3) promotes plant growth and water uptake. Plant J. 2020, 102, 779–796. [CrossRef]
- Duan, P.; Rao, Y.; Zeng, D.; Yang, Y.; Xu, R.; Zhang, B.; Dong, G.; Qian, Q.; Li, Y. SMALL GRAIN 1, which encodes a mitogen-activated protein kinase 4, influences grain size in rice. Plant J. 2014, 77, 547–557. [CrossRef]
- Xu, R.; Duan, P.; Yu, H.; Zhou, Z.; Zhang, B.; Wang, R.; Li, J.; Zhang, G.; Zhuang, S.; Lyu, J.; et al. Control of grain size and weight by the OsMKKK10-OsMKK4-OsMAPK6 signaling pathway in rice. Mol. Plant 2018, 11, 860–873. [CrossRef]
- Yang, X.; Ren, Y.; Cai, Y.; Niu, M.; Feng, Z.; Jing, R.; Mou, C.; Liu, X.; Xiao, L.; Zhang, X.; et al. Overexpression of OsbHLH107, a member of the basic helix-loop-helix transcription factor family, enhances grain size in rice (Oryza sativa L.). Rice 2018, 11. [CrossRef]
- Chen, X.; Jiang, L.; Zheng, J.; Chen, F.; Wang, T.; Wang, M.; Tao, Y.; Wang, H.; Hong, Z.; Huang, Y.; et al. A missense mutation in Large Grain Size 1 increases grain size and enhances cold tolerance in rice. J. Exp. Bot. 2019, 70, 3851–3866. [CrossRef]
- Ren, D.; Rao, Y.; Wu, L.; Xu, Q.; Li, Z.; Yu, H.; Zhang, Y.; Leng, Y.; Hu, J.; Zhu, L.; et al. The pleiotropic ABNORMAL FLOWER AND DWARF 1 affects plant height, floral development and grain yield in rice. J. Integr. Plant Biol. 2016, 58, 529–539. [CrossRef]
- Peng, P.; Liu, L.; Fang, J.; Zhao, J.; Yuan, S.; Li, X. The rice TRIANGULAR HULL1 protein acts as a transcriptional repressor in regulating lateral development of spikelet. Sci. Rep. 2017, 7, 1–15. [CrossRef]
- Hossain, M.A.; Cho, J. Il; Han, M.; Ahn, C.H.; Jeon, J.S.; An, G.; Park, P.B. The ABRE-binding BZIP transcription factor OsABF2 is a positive regulator of abiotic stress and ABA signaling in rice. J. Plant Physiol. 2010, 167, 1512–1520. [CrossRef]
- Tang, N.; Zhang, H.; Li, X.; Xiao, J.; Xiong, L. Constitutive activation of transcription factor OsbZIP46 improves drought tolerance in rice. Plant Physiol. 2012, 158, 1755–1768. [CrossRef]
- Xu, M.; Zhu, L.; Shou, H.; Wu, P. A PIN1 Family Gene, OsPIN1, involved in auxin-dependent adventitious root emergence and tillering in rice. Plant Cell Physiol. 2005, 46, 1674–1681. [CrossRef]
- Sun, H.; Tao, J.; Bi, Y.; Hou, M.; Lou, J.; Chen, X.; Zhang, X.; Luo, L.; Xie, X.; Yoneyama, K.; et al. OsPIN1b is involved in rice seminal root elongation by regulating root apical meristem activity in response to low nitrogen and phosphate. Sci. Rep. 2018, 8, 1–11. [CrossRef]
- Barrôco, R.M.; Peres, A.; Droual, A.M.; De Veylder, L.; Nguyen, L.S.L.; De Wolf, J.; Mironov, V.; Peerbolte, R.; Beemster, G.T.S.; Inzé, D.; et al. The cyclin-dependent kinase inhibitor Orysa;KRP1 plays an important role in seed development of rice. Plant Physiol. 2006, 142, 1053–1064. [CrossRef]
- Kasai, K.; Kanno, T.; Akita, M.; Ikejiri-Kanno, Y.; Wakasa, K.; Tozawa, Y. Identification of three shikimate kinase genes in rice: Characterization of their differential expression during panicle development and of the enzymatic activities of the encoded proteins. Planta 2005, 222, 438–447. [CrossRef]
- Nojiri, H.; Sugimori, M.; Yamane, H.; Nishimura, Y.; Yamada, A.; Shibuya, N.; Kodama, O.; Murofushi, N.; Omori, T. Involvement of jasmonic acid in elicitor-induced phytoalexin production in suspension-cultured rice cells. Plant Physiol. 1996, 110, 387–392. [CrossRef]
- Lee, S.; Choi, S.C.; An, G. Rice SVP-group MADS-box proteins, OsMADS22 and OsMADS55, are negative regulators of brassinosteroid responses. Plant J. 2008, 54, 93–105. [CrossRef]
- Tanaka, A.; Nakagawa, H.; Tomita, C.; Shimatani, Z.; Ohtake, M.; Nomura, T.; Jiang, C.J.; Dubouzet, J.G.; Kikuchi, S.; Sekimoto, H.; et al. Brassinosteroid Upregulated 1, encoding a helix-loop-helix protein, is a novel gene involved in brassinosteroid signaling and controls bending of the lamina joint in rice. Plant Physiol. 2009, 151, 669–680. [CrossRef]
- Dong, H.; Zhao, H.; Li, S.; Han, Z.; Hu, G.; Liu, C.; Yang, G.; Wang, G.; Xie, W.; Xing, Y. Genome-wide association studies reveal that members of BHLH subfamily 16 share a conserved function in regulating flag leaf angle in rice (Oryza Sativa). PLoS Genet. 2018, 14, 1–20. [CrossRef]
- Pantuwan, G.; Fukai, S.; Cooper, M.; Rajatasereekul, S.; O’Toole, J.C. Yield Response of Rice (Oryza Sativa L.) Genotypes to different types of drought under rainfed lowlands part 1. Grain yield and yield components. Field Crops Res. 2002, 73, 153–168. [CrossRef]
- Pantuwan, G.; Fukai, S.; Cooper, M.; Rajatasereekul, S.; O’Toole, J.C. Yield Response of Rice (Oryza Sativa L.) Genotypes to drought under rainfed lowlands 2. Selection of drought resistant genotypes. Field Crops Res. 2002, 73, 169–180. [CrossRef]
- Pantuwan, G.; Fukai, S.; Cooper, M.; Rajatasereekul, S.; O’Toole, J.C.; Basnayake, J. Yield response of rice (Oryza Sativa L.) genotypes to drought under rainfed lowlands: 3. Plant factors contributing to drought resistance. Field Crops Res. 2004, 89, 281–297. [CrossRef]
- Henry, A.; Gowda, V.R.P.; Torres, R.O.; McNally, K.L.; Serraj, R. Variation in root system architecture and drought response in rice (Oryza sativa): Phenotyping of the OryzaSNP panel in rainfed lowland fields. Field Crops Res. 2011, 120, 205–214. [CrossRef]
- Dixit, S.; Swamy, B.P.M.; Vikram, P.; Ahmed, H.U.; Sta Cruz, M.T.; Amante, M.; Atri, D.; Leung, H.; Kumar, A. Fine mapping of QTLs for rice grain yield under drought reveals sub-QTLs conferring a response to variable drought severities. Theor. Appl. Genet. 2012, 125, 155–169. [CrossRef]
- Palanog, A.D.; Swamy, B.P.M.; Shamsudin, N.A.A.; Dixit, S.; Hernandez, J.E.; Boromeo, T.H.; Cruz, P.C.S.; Kumar, A. Grain yield QTLs with consistent-effect under reproductive-stage drought stress in rice. Field Crops Res. 2014, 161, 46–54. [CrossRef]
- Venuprasad, R.; Dalid, C.O.; Del Valle, M.; Zhao, D.; Espiritu, M.; Sta Cruz, M.T.; Amante, M.; Kumar, A.; Atlin, G.N. Identification and characterization of large-effect quantitative trait loci for grain yield under lowland drought stress in rice using bulk-segregant analysis. Theor. Appl. Genet. 2009, 120, 177–190. [CrossRef]
- Swamy B. P., M.; Ahmed, H.U.; Henry, A.; Mauleon, R.; Dixit, S.; Vikram, P.; Tilatto, R.; Verulkar, S.B.; Perraju, P.; Mandal, N.P.; et al. Genetic, physiological, and gene expression analyses reveal that multiple QTL enhance yield of rice mega-variety IR64 under drought. PLoS One 2013, 8, 719–729. [CrossRef]
- Y, P.; Kamoshita, A.; Norisada, M.; Deshmukh, V. Eco-physiological evaluation of stele transversal area 1 for rice root anatomy and shoot growth. Plant Prod. Sci. 2020, 23, 202–210. [CrossRef]
- Sun, X.; Xiong, H.; Jiang, C.; Zhang, D.; Yang, Z.; Huang, Y.; Zhu, W.; Ma, S.; Duan, J.; Wang, X.; et al. Natural Variation of DROT1 confers drought adaptation in upland rice. Nat. Commun. 2022, 13, 1–17. [CrossRef]
- Panda, D.; Mishra, S.S.; Behera, P.K. Drought tolerance in rice: Focus on recent mechanisms and approaches. Rice Sci. 2021, 28, 119–132. [CrossRef]
- Li, Z.; Mu, P.; Li, C.; Zhang, H.; Li, Z.; Gao, Y.; Wang, X. QTL Mapping of root traits in a doubled haploid population from a cross between upland and lowland japonica rice in three environments. Theor. Appl. Genet. 2005, 110, 1244–1252. [CrossRef]
- Yi, K.; Wu, Z.; Zhou, J.; Du, L.; Guo, L.; Wu, Y.; Wu, P. OsPTF1, a novel transcription factor involved in tolerance to phosphate starvation in rice. Plant Physiol. 2005, 138, 2087–2096. [CrossRef]
- Monna, L.; Lin, H.X.; Kojima, S.; Sasaki, T.; Yano, M. Genetic dissection of a genomic region for a quantitative trait locus, Hd3, into two loci, Hd3a and Hd3b, controlling heading date in rice. Theor. Appl. Genet. 2002, 104, 772–778. [CrossRef]
- Begum, H.; Spindel, J.E.; Lalusin, A.; Borromeo, T.; Gregorio, G.; Hernandez, J.; Virk, P.; Collard, B.; McCouch, S.R. Genome-wide association mapping for yield and other agronomic traits in an elite breeding population of tropical rice (Oryza sativa). PLoS One 2015, 10, 1–19. [CrossRef]
- Courtois, B.; Ahmadi, N.; Khowaja, F.; Price, A.H.; Rami, J.F.; Frouin, J.; Hamelin, C.; Ruiz, M. Rice root genetic architecture: Meta-analysis from a drought QTL database. Rice 2009, 2, 115–128. [CrossRef]
- Singh, S.; Kumar, A.; Panda, D.; Modi, M.K.; Sen, P. Identification and characterization of drought responsive miRNAs from a drought tolerant rice genotype of Assam. Plant Gene 2020, 21, 100213. [CrossRef]
- Li, P.; Zhang, Y.; Yin, S.; Zhu, P.; Pan, T.; Xu, Y.; Wang, J.; Hao, D.; Fang, H.; Xu, C.; et al. QTL-by-environment interaction in the response of maize root and shoot traits to different water regimes. Front. Plant Sci. 2018, 9, 229. [CrossRef]
| Experiment 1 | ||||||||
| Trait | Otomemochi | Yumenohatamochi | Progeny | |||||
| Average | Minimum | Maximum | N | SD | P value | |||
| Control | ||||||||
| Date of 50% flowering | 8/4/2011 | 8/9/2011 | 8/9/2011 | 7/31/2011 | 8/24/2011 | 97 | 6.3 | <0.001 |
| Grain dry weight per hill (g) | 10.8 | 23.2 | 16.2 | 3.4 | 31.6 | 97 | 5.8 | <0.001 |
| Estimated yield (g/m2) | 269 | 580 | 407 | 92 | 790 | 97 | 145 | <0.001 |
| Total dry weight per hill (g) | 28.8 | 46.6 | 48.1 | 15.9 | 81.9 | 97 | 14.2 | <0.001 |
| Harvest index | 0.39 | 0.50 | 0.34 | 0.12 | 0.58 | 97 | 0.07 | <0.001 |
| Plant height (cm) | 79 | 94 | 80 | 57 | 96 | 97 | 8.0 | <0.001 |
| Dry root weight (g) | 2.7 | 2.7 | 2.8 | 0.9 | 6.5 | 97 | 1.1 | <0.001 |
| Drought | ||||||||
| Date of 50% flowering | 9/14/2011 | 9/20/2011 | 9/13/2011 | 8/1/2011 | 9/25/2011 | 97 | 12 | <0.001 |
| Grain dry weight per hill (g) | 0.9 | 6.2 | 1.5 | 0.1 | 6.5 | 97 | 1.2 | <0.001 |
| Estimated yield (g/m2) | 23 | 154 | 37 | 1 | 162 | 97 | 31 | <0.001 |
| Grain dry weight after rewatering (g) | 0.7 | 4.9 | 1.1 | 0.0 | 6.4 | 97 | 1.01 | <0.001 |
| Total dry weight per hill (g) | 9.9 | 24.8 | 17.3 | 5.0 | 35.2 | 97 | 6.4 | <0.001 |
| Harvest index | 0.05 | 0.21 | 0.07 | 0.01 | 0.21 | 97 | 0.04 | <0.001 |
| Plant Height (cm) | 56 | 66 | 61 | 47 | 76 | 97 | 6.1 | <0.001 |
| Root dry weight (g) | 2.5 | 2.4 | 2.7 | 1.2 | 5.5 | 97 | 0.92 | <0.001 |
| Leaf rolling score | 6.2 | 1.4 | 3.1 | 0.0 | 9.2 | 97 | 2.2 | <0.001 |
| Flowering delay (days) | −41 | −42 | −35 | 0 | −52 | 97 | 11 | <0.001 |
| Plant height reduction (cm) | −23 | −28 | −18 | −34 | 4 | 97 | 8 | <0.001 |
| Drought response index | −3.6 | 22.6 | 0.0 | −6.4 | 15.9 | 97 | 4.7 | – |
| Experiment 2 | ||||||||
| Trait | Otomemochi | Yumenohatamochi | Progeny | |||||
| Average | Minimum | Maximum | N | SD | P value | |||
| Control | ||||||||
| Date of 50% flowering | 8/5/2012 | 8/7/2012 | 8/6/2012 | 7/23/2012 | 8/20/2012 | 97 | 6.3 | <0.001 |
| Grain dry weight per hill (g) | 16.9 | 32.0 | 14.3 | 3.7 | 27.6 | 97 | 5.8 | <0.001 |
| Estimated yield (g/m2) | 422 | 800 | 357 | 92 | 690 | 97 | 145 | <0.001 |
| Total dry weight per hill (g) | 46.5 | 73.3 | 36.1 | 9.6 | 73.6 | 97 | 12.3 | <0.001 |
| Harvest index | 0.44 | 0.37 | 0.39 | 0.22 | 0.56 | 97 | 0.08 | <0.001 |
| Plant height (cm) | 77 | 96 | 88 | 70 | 112 | 97 | 9.5 | <0.001 |
| Drought | ||||||||
| Date of 50% flowering | 8/5/2012 | 8/7/2012 | 8/7/2012 | 7/28/2012 | 8/23/2012 | 97 | 6.6 | <0.001 |
| Grain dry weight per hill (g) | 4.1 | 10.8 | 5.0 | 0 | 16.1 | 97 | 3.2 | <0.001 |
| Estimated yield (g/m2) | 102 | 270 | 125 | 0 | 402 | 97 | 80 | <0.001 |
| Total dry weight per hill (g) | 36.4 | 59.3 | 37.5 | 12.7 | 63.1 | 97 | 10.8 | <0.001 |
| Harvest index | 0.11 | 0.18 | 0.14 | 0 | 0.47 | 96 | 0.09 | <0.001 |
| Plant height (cm) | 66 | 82 | 67 | 48 | 89 | 97 | 8.4 | <0.001 |
| Leaf rolling | 6.2 | 4.4 | 5.5 | 2.4 | 8.3 | 97 | 1.3 | 0.02 |
| Flowering delay (days) | −0 | 0 | 1 | −11 | 10 | 97 | 4 | <0.001 |
| Plant height reduction (cm) | −31 | 4 | −20 | −46 | 10 | 97 | 13 | <0.001 |
| Drought response index | −1.3 | 3.6 | 0.0 | −3.9 | 8.3 | 97 | 2.5 | <0.001 |
| Experiment | 50% flowering | Plant height | Total dry weight | Harvest index | Grain dry weight | Grain weight before rewatering | Grain weight after rewatering | Root dry weight | Leaf rolling | Flowering delay | Plant height reduction |
| Experiment 1 | 0.04 | 0.10 | 0.48** | 0.77** | 0.98** | 0.42** | 0.76** | 0.28* | −0.34** | 0.04 | 0.22 |
| Experiment 2 | −0.13 | 0.13 | 0.03 | 0.72** | 0.93** | – | – | - | −0.05 | −0.23* | 0.03 |
| Experiment(s) | Chr. | Trait | Marker interval a | Position b | LOD c | R2 d | Ae |
| 1 | 2 | DRI | RM3703–RM6911 | 37.4 | 5.9 | 19.9 | 2.24 |
| 2 | 6 | DRI | RM8120–RM7023 | 31.0 | 3.6 | 12.4 | −0.90 |
| 1 + 2 combined | 2 | DRI | RM3703–RM6379 | 40.4 | 5.1 | 26.3 | 1.42 |
| 1 + 2 combined | 2 | DRI | RM6733–RM3850 | 151.5 | 3.3 | 10.3 | −0.87 |
| Trait | Chr | Interval | LOD | A | P | H2 | AE |
| Drought response index | 2 | RM6911–RM6379 | 7.96 | 1.08 | 0.33 | 0.08 | 0.16 |
| 50% flowering C | 3 | RM3372–RM4853 | 9.08 | −2.58 | 0.00 | 0.23 | 0.02 |
| 11 | RM536–RM287 | 3.40 | −1.56 | 0.00 | |||
| Plant height C | 5 | RM2010–RM4501 | 4.10 | −1.90 | 0.29 | 0.04 | 0.06 |
| Total dry weight C | 10 | RM216–RM184 | 3.69 | 3.69 | 0.11 | 0.07 | 0.04 |
| Harvest index C | 6 | RM2615–RM7023 | 3.96 | 0.02 | 0.03 | 0.05 | 0.02 |
| Grain yield C | 10 | RM216–RM184 | 4.14 | 1.63 | 0.24 | 0.08 | 0.09 |
| Plant height D | 11 | RM536–RM287 | 5.80 | −1.56 | 0.49 | 0.04 | 0.15 |
| Harvest index D | 11 | RM441–RM536 | 3.39 | 0.02 | 0.20 | 0.06 | 0.06 |
| Grain yield D | 3 | RM1332–RM3029 | 2.60 | 0.51 | 0.15 | 0.06 | 0.08 |
| 6 | RM4447–RM6734 | 4.34 | −0.56 | 0.41 | |||
| Leaf rolling D | 1 | RM1387–RM1067 | 2.44† | −0.27 | 0.30 | 0.14 | 0.09 |
| 2 | RM6933–RM3857 | 11.56 | −0.70 | 0.09 | |||
| 4 | RM3288–RM5503 | 2.64 | −0.35 | 0.00 |
| Group | DRI | Control (combined) | Drought (combined) | Experiment 1 | [20] | ||||||
| 50% flowering | Plant height (cm) | Grain dry weight (g) | 50% flowering | Plant height (cm) | Grain dry weight (g) | Leaf rolling | Root weight change under drought (g) | Stele transversal area (STA) (μm2) | %STA ratio to root transversal area | ||
| Top-DRI lines | 4.7 | 220 | 83 | 16.7 | 240 | 66 | 5.5 | 3.7 | 0.69 | 52.7 | 6.4 |
| Bottom-DRI lines | −3.4 | 220 | 85 | 16.4 | 239 | 64 | 1.4 | 5.0 | −0.49 | 39.4 | 5.4 |
| P | ** | ns | ns | ns | ns | Ns | ** | + | * | * | * |
| Gene | Chr. | Encoded proteins and their functions | Start | End | Location | Conditions and references |
| GW2 | 2 | RING-type E3 ubiquitin ligase, Negative regulation of grain width and size | 8115223 | 8121651 | LOC_Os02g14720.1-2 | [22]Paddy [23] |
| OsGTE4 | 2 | Bromo-domain-containing protein, Homologous to Arabidopsis GTE4, Maintenance of root meristem, Cell cycle regulation | 8490392 | 8498003 | LOC_Os02g15220.1 | Petri dish, 1/2 strength MS [28]7-day-old plantlets [26] |
| RPBF | 2 | Dof zinc finger transcription activator (factor), Grain filling, GA response in aleurone cell | 8590318 | 8593735 | LOC_Os02g15350.1 | Greenhouse [30,31] |
| PYL/RCAR3 | 2 | ABA receptor, Pyrabactin resistance-like (PYL) ABA receptor family protein, Survival rate under cold and drought stress, ABA-mediated inhibition of seed germination | 8801325 | 8805273 | LOC_Os02g15640.1 | 3 weeks after sowing, water withhold for 10 days, rehydrated for 7 days (drought) and cold treatment [32] |
| EP3 | 2 | Erect panicle, Panicle branching, Tillering, Culm mechanical strength | 9071132 | 9075127 | LOC_Os02g15950.1 | Greenhouse and field [24]Paddy standard condition [25] |
| AIM1 | 2 | 3-Hydroxyacyl-CoA dehydrogenase, Salicylic acid biosynthesis, Maintenance of root meristem activity, Beta-oxidation of fatty acids | 10001543 | 10010228 | LOC_Os02g17390.1 | Continuous illumination [29]Hydroponic in JA, DIECA, CA BA [27] |
| LKR/SDH | 2 | Lysine ketoglutarate reductase/saccharopine dehydrogenase, Lysine-degrading enzyme in developing grain | 33253386 | 33264913 | LOC_Os02g54254.1-4 | Greenhouse [30,31] |
| SMG1 | 2 | Mitogen-activated protein kinase 4, Defense response, Cell proliferation, Grain growth | 33442070 | 33443948 | LOC_Os02g54600.1 | Field, natural condition [36,37] |
| MGD2 | 2 | Monogalactosyldiacylglycerol synthase, upregulated under stress | 34223738 | 34227369 | LOC_Os02g55910.1-2 | Pot with salt, submerged, drought, cold treatments [33] |
| LGS1 | 2 | Basic helix-loop-helix transcription factor, Regulation of grain size | 34353730 | 34356787 | LOC_Os02g56140.1 | Field, natural conditions [38,39] |
| BLS1 | 2 | DUF640 domain-containing protein, ALOG domain-containing nuclear protein, Regulation of plant height, floral development and grain yield, Spikelet morphogenesis | 34687624 | 34689723 | LOC_Os02g56610.1 | Field, natural conditions [40]Paddy, natural conditions [41] |
| PIP1;3 | 2 | Plasma membrane intrinsic protein, Higher water potential and root hydraulic conductivity under water stress, Taller plants with more leaves | 35349673 | 35351221 | LOC_Os02g57720.1 | Water deficit at –0.82 MPa [34]Water deficit at SRWC 85% [35] |
| BZIP46 | 6 | bZIP transcription factor, Positive regulator of ABA signaling, Survival rate | 5677157 | 5681985 | LOC_Os06g10880.1-3 | Growth chamber, 3-week-old with drought, salinity, oxidative stress [42]Pot, drought, cold, and heat stress [43] |
| KRP2 | 6 | Cyclin-dependent kinase inhibitor 2, KIP-related protein, Grain filling, Seed germination, Seed morphogenesis | 5791466 | 5795810 | LOC_Os06g11050.1 | Greenhouse, 60%–70% relative humidity [46] |
| MADS55 | 6 | Short vegetative phase group MADS-box protein, Negative regulation of brassinosteroid responses, Dwarfism, Leaf angle, Brassinosteroid sensitivity | 5952599 | 5963309 | LOC_Os06g11330.1 | Paddy field, glasshouseBL, GA, BR treatment [49] |
| SK2 | 6 | Shikimate kinase 2, Defense response, Development (panicle) | 6495212 | 6497797 | LOC_Os06g12150.1-2 | Medium with elicitor (N-acetylchitoheptaose) treatment [47,48] |
| BU1 | 6 | Leaf angle, Grain size, Brassinosteroid sensitivity | 6556697 | 6557748 | LOC_Os06g12210.1 | Greenhouse [50]Field, natural conditions [51] |
| PIN1B | 6 | PIN protein, Auxin efflux carrier, Auxin transport and signaling, Development (root, shoot, and inflorescence) | 6866393 | 6869521 | LOC_Os02g50960.1-2 | Hydroponic [44]Greenhouse and N, P deficit treatment [45] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




