1. Introduction
Despite significant improvements in overall outcomes for patients with congenital heart diseases (CHDs), neonates and infants undergoing cardiac surgery still experience high rates of postoperative complications and mortality. The Kids' Inpatient Database (KID) reports an in-hospital mortality rate of 6.9% for neonates and infants, compared to 1.28%, 0.67%, and 0.83% for the age groups 1-5, 6-12, and 13-17 years old, respectively [
1]. Longer cardiopulmonary bypass (CPB) time [
2] and larger blood transfusion requirements [
3,
4,
5] are established clinical risk factors. These factors contribute to the development of thrombotic complications and subsequent organ dysfunction/failure, which are major contributors to worse outcomes [
6]. Understanding the mechanisms underlying these complications is therefore critical.
Extracellular vesicles (EVs) are membrane-enclosed nanoparticles that act as messengers between cells. These spheres encapsulate a diverse cargo of functional proteins, nucleic acids, lipids, and other biochemical molecules, enabling them to influence various cellular processes [
7]. Their pleiotropic role has been implicated in a range of diseases [
8,
9,
10].
Among different RNA species encapsulated within EVs, circular RNAs (circRNAs) represent a recently discovered class of non-coding RNAs with exciting potential for disease diagnosis [
11]. Their unique closed-loop structure confers exceptional stability, making them resistant to degradation and facilitating their detection in biological samples [
12]. Additionally, the tissue-specific expression patterns of circRNAs offer a remarkable advantage for pinpointing the origin of pathological processes within the body [
13]. CircRNAs harbor a variety of functions including microRNA sponging, cell cycle regulation, and cellular communications [
14]. Thus, in this study, we examined the expression profiles of plasma EV circRNAs in neonates and infants who developed organ dysfunction (respiratory failure) following congenital cardiac surgery and explored different associations of circRNAs with the pathomechanism of this condition.
2. Material and Methods
2.1 Study Design and Setting
This single-center prospective cohort study was conducted at a quaternary academic pediatric medical center. The study was approved by the Institutional Review Board at Boston Children's Hospital, and written informed consent was obtained from a parent or a legal guardian. We followed Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) statement for cohort studies in the preparation of this manuscript [
19].
2.2. Patient Selection and Perioperative Course
We included neonates and infants scheduled for congenital cardiac surgery on CPB. Exclusion criteria included patients who did not necessitate CPB, presented with an active infection, received chronic steroid therapy, had immunodeficiency such as human immunodeficiency virus (HIV) infection, or a history of malignancy. The patients were enrolled from May 31, 2022 to February 22, 2023.
All patients in the study underwent general anesthesia with endotracheal intubation and arterial and central venous line placement. After surgical dissection, patients were heparinized and underwent aortic and venous cannulation for CPB. The CPB circuit was primed with one unit of packed red blood cells (pRBC) and one unit of fresh frozen plasma (FFP), maintaining a target hematocrit level greater than 30% per our institutional standard protocol. The use of circulatory arrest or regional perfusion, temperature management, and modified ultrafiltration (MUF) was determined on a case-by-case basis. In cases of non-surgical microvascular bleeding, platelets, and cryoprecipitate were administered. Neonates and infants typically did not receive FFP transfusion after CPB in our institution. Following surgery, patients were kept intubated and transferred to the intensive care unit (ICU) for postoperative care.
2.3. Clinical Data Collection
Demographic information, comorbidities, diagnosis, procedures, laboratory values, CPB details, type and volume of blood products administered, postoperative complications, respiratory support, vital signs, as well as hospital and ICU lengths of stay were extracted from the electronic medical record for the analysis of clinical data. The presence of organ dysfunction/failure and/or thrombosis was collected as postoperative complications. Due to the absence of a standardized definition for organ dysfunction following congenital cardiac surgery, we adopted previously published criteria established by others [
20,
21]. Organ dysfunctions included (1) cardiovascular dysfunction (low cardiac output syndrome, reliance on a vasoactive drug to maintain blood pressure, or two of the followings; metabolic acidosis, elevated arterial lactate, oliguria, or prolonged capillary refill), (2) respiratory dysfunction (arterial oxygen tension/fraction of inspired oxygen (PaO
2/FiO
2) < 300, arterial carbon dioxide tension (PaCO
2) > 65 torr or 20 mmHg over baseline PaCO
2, need for > 50% FiO
2 to maintain oxygen saturation ≥ 92%, or need for non-elective mechanical ventilation, prolonged mechanical ventilation ≥ 5 days, (3) renal dysfunction (the presence of creatinine of > 114 μmol/L, urine output of < 1 mL/kg/hr despite diuretic administration, or the requirement for ultrafiltration or hemodialysis, (4) coagulopathy or bleeding complication requiring chest exploration for bleeding or removal of clots, intracranial hemorrhage, prothrombin or partial thromboplastin time was three times normal, or > 30 mL/kg of blood products were infused during a 24-hr period, (5) central nervous system dysfunction (development of a new intracranial infarct or hemorrhage, evidence of hypoxic-ischemic injury by clinical examination or computed tomography of the head, or brain death), 6) hepatic dysfunction (bilirubin concentration of > 2 mg/dL and/or increase of hepatic cellular enzymes two or more times normal). Thrombosis was defined as the presence of any vascular thrombosis detected using ultrasound diagnostic imaging. In addition, we also calculated Pediatric Logistic Organ Dysfunction-2 (PELOD-2) score to assess organ dysfunction as previously used in the CHD cohort [
22], although recognizing its limitation because this system cannot differentiate between therapy and severity of diseases from cardiovascular standpoint [
23].
2.4. Blood Sample Collection
Blood samples were collected from patients through existing central venous catheters. The blood was obtained at four-time points; 1) immediately after anesthesia induction (pre-surgery), 2) on postoperative day 1 (post-surgery). Once blood samples were received, they were subjected to centrifugation at 1,000 x g for 10 min to obtain plasma. Plasma was immediately stored at -80oC.
2.5. Plasma Collection and EVs Isolation
EVs were isolated from 200-300 µL plasma samples using the exoEasy Midi kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. In brief, plasma samples were thawed at 37°C and centrifuged at 10,000 x g for 12 minutes at room temperature to remove cellular debris. Plasma samples were mixed with XBP buffer and added to the exoEasy spin column. Columns were washed with XWP buffer, and then EVs were eluted from the column membrane using XE buffer.
2.6. Nanoparticle Tracking Analysis (NTA)
NTA was performed using the ZetaVIEW equipment (Particle Metrix, Ammersee, Germany). Initially, the instrument was rinsed using 5 mL of filtered water to remove any remaining particles. Auto alignment was done by injecting 2 mL of polystyrene particles into the instrument, which was rinsed again afterwards. As optimal particle per frame values ranges from 140 to 200 particles/frame, plasma EV samples were diluted 1:500 in pre-filtered PBS. Subsequently, 1 mL of each diluted EV sample was injected into the instrument which was carefully rinsed after each measurement.
The manufacturer’s default software settings for EVs, liposomes or nanospheres were selected accordingly. For each measurement, two video cycles were recorded by scanning 11 cell positions and capturing 30 frames per position with the following settings: Focus: autofocus; Camera sensitivity for all samples: 75; Shutter: 70; Scattering intensity: detected automatically; Cell temperature: 25°C; pH: 7.0. After capture, the videos were analyzed by the in-built ZetaView Software v.8.05.05 SP2 with specific analysis parameters: Max size: 1000; Min size: 5; Min brightness: 25. Hardware: embedded laser: 40 mW at 488 nm; camera: CMOS.
2.7. EV Proteomic Antibody Array
EV markers including Golgi matrix protein 130 (GM130), CD63, CD81, epithelial cellular adhesion molecule (EpCAM), annexin A5 (ANXA5), tumor susceptibility gene 101 (TSG101), flotillin 1 (FLOT1), intercellular adhesion molecule 1 (ICAM1), and ALG-2 interacting protein X (ALIX) were measured using a western blot-based method; Exo-Check Exosome Antibody Array Kit, Human (System Biosciences, Palo Alto, CA, USA). In brief, according to the manufacturer’s instruction, 50 µg of eluted EVs were lysed, labelled, and incubated overnight with a panel-antibody-coated membrane. On the next day, the membrane was incubated with a detection antibody and developed with a 1:1 substrate mixture KwikQuant Western Blot Detection kit (Kindle Biosciences, Greenwich, Connecticut, USA). Subsequently, the membrane was visualized with KwikQuant Pro Imager (Kindle Biosciences).
2.8. EV RNA Extraction
Total RNA including small RNAs was extracted from EVs using miRNeasy kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. RNA yield was measured by Qubit™ microRNA Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA). RNA size distribution and quality were assessed by Bioanalyzer Small RNA Analysis kit (Agilent Technologies, Santa Clara, CA, USA).
2.9. RNA Library Preparation and Small RNA Sequencing
Small RNA libraries were constructed using NEBNext Small RNA Library Prep Set for Illumina (New England Biolabs, Ipswich, MA, USA), according to the manufacturer’s protocol, with minor modifications for the low RNA input. Briefly, 3 ng of RNA was used for the library preparation. The 3′ SR Adapter, SR RT Primer, and 5′ SR Adapter were diluted 1:4, and the RNA was ligated with both adapters, and was reverse transcribed, barcoded, and amplified for 15 cycles. The generated libraries were cleaned up using AMPure XP Beads (Beckman Coulter, Brea, CA, USA) and quantified using the Qubit™ dsDNA HS Assay (Thermo Fisher Scientific) and the Bioanalyzer High Sensitivity DNA Analysis kit (Agilent Technologies) prior to sequencing on a HiSeq4000 platform (Illumina, San Diego, CA, USA) with High Output Kit v2.5 and 50 bases single-reads, according to the manufacturer’s instructions.
2.10. Bioinformatic Analysis
CircRNA sequences and annotations were retrieved from the circBase 0.1 database [
15] for the human genome assembly hg19. Small RNA-Seq reads were processed using miRMaster 2.0 [
16]. This included quality control checks to assess read quality and the removal of Illumina adapter sequences. Subsequently, meticulously clean reads were mapped to the reference genome (hg19 assembly) and counted to quantify transcript abundance. To filter out low abundant circRNAs, expression cut-off was applied > 5 counts for at least 4 samples. Differential expression analysis was conducted with Deseq2 R package (version 1.38.3). Density analysis were computed using the average expression of the top 500 variable circRNAs per each sample and visualized with ggdensity R package (version 1.0.0). Functional analysis including biological pathways and processes were curated with the implementation of different databases such as Reactome [
17,
18], Biological process (Gene Ontology) [
18], NCATS BioPlanet [
19] by utilizing clusterProfiler package (Version 4.6.2). Cell type and tissue specificity were retrieved from HuBMAP ASCTplusB augmented 2022 database [
20]. Corrplot R package (version 0.92) was utilized to create correlation analysis and visualization.
2.11. Statistical Analysis
Data were shown as mean ± S.D. Student’s unpaired t-test were used for statistical significance with p value * < 0.05, ** < 0.005, and *** < 0.001.
3. Results
3.1. Altered Plasma EV circRNA Expression Profiles in Individuals with Respiratory Failure Following Pediatric Congenital Heart Surgery
Our current work included four neonates and infants who developed organ dysfunction (OD), primarily respiratory failure, following congenital cardiac surgery, and as controls we enrolled five ones who did not develop organ dysfunction (NOD). The characteristics of all neonates and infants enrolled for this study were shown in
Table 1.
To probe the complex pathomechanisms associated with postoperative OD, we employed EV circRNAs due to their versatile biological functions and their intriguing tissue specificity.
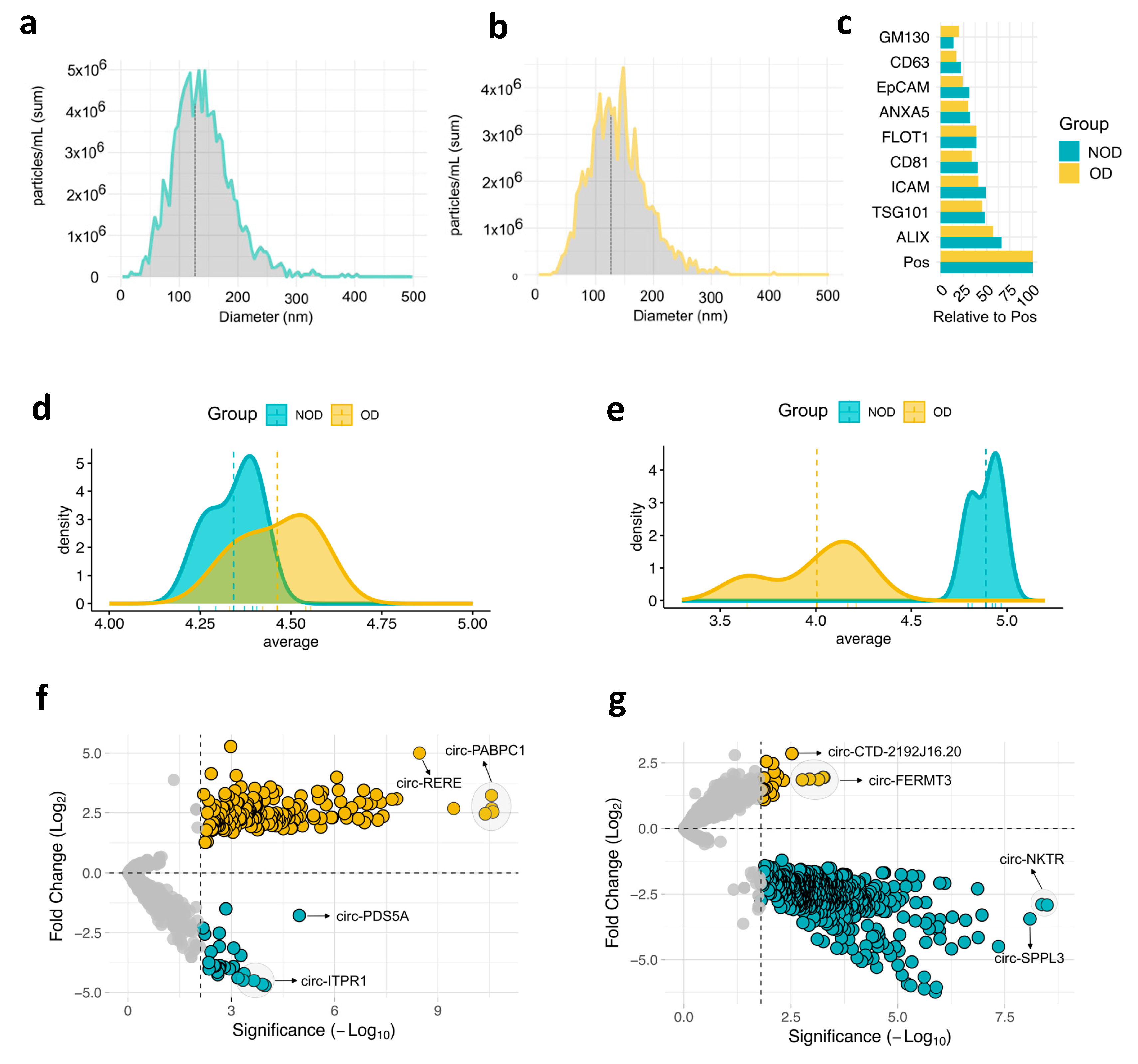
Next, we isolated EVs from plasma of all individuals at two time points; pre-surgery: after anesthesia induction (baseline), and post-surgery: on postoperative day 1. Subsequently, we characterized EVs size and protein markers. Nanoparticle analysis (NTA) revealed a median EV size less than 150 nm, which falls within the typical range for exosomes [
21] (
Figure 1a,b). This was further observed by examining a panel of EV of established EVs protein markers (
Figure 1c and Supplementary Figure S1). All markers were detected in both study groups, with ALIX and TSG101 (exosomes markers) showing the highest expression.
We then conducted small RNA-Sequencing (RNA-Seq) analysis to comprehensively examine the circRNA cargo within these vesicles. Analysis of average circRNA expression revealed a remarkable overlap between the OD and NOD groups in the pre-surgery samples (
Figure 1c). However, a dramatic shift in the post-surgery OD group was observed, leading to a clear distinction from the post-surgery NOD group (
Figure 1d). This indicates a potential association between the surgical procedure and a negative outcome, including the possibility of postoperative OD.
To address the differences between OD and NOD at the two time points of the congenital cardiac surgery, we computed the differentially expressed (DE) circRNAs for the two pairwise comparisons (
Figure 1f-g and Supplementary Table S1). By setting the threshold for false discovery rate (FDR) < 0.05, OD versus NOD revealed 229 upregulated and 36 downregulated circRNAs in pre-surgery (
Figure 1f), and 39 upregulated and 643 downregulated circRNAs in post-surgery (
Figure 1g), mirroring the previously observed post-surgical decline in circRNA expression profiles within the OD group (
Figure 1d).
To account for the larger transfusion volumes in the OD group (
Table 1), we evaluated the housekeeping microRNAs (miRNAs) expression levels from the sequencing data [
22,
23] (Supplementary Table S2). No significant changes in miRNA expression were observed between OD and NOD groups, suggesting that the transfusion volume did not introduce a dilution bias.
3.2. Biological Pathways and Processes Associated with the Altered EV circRNAs
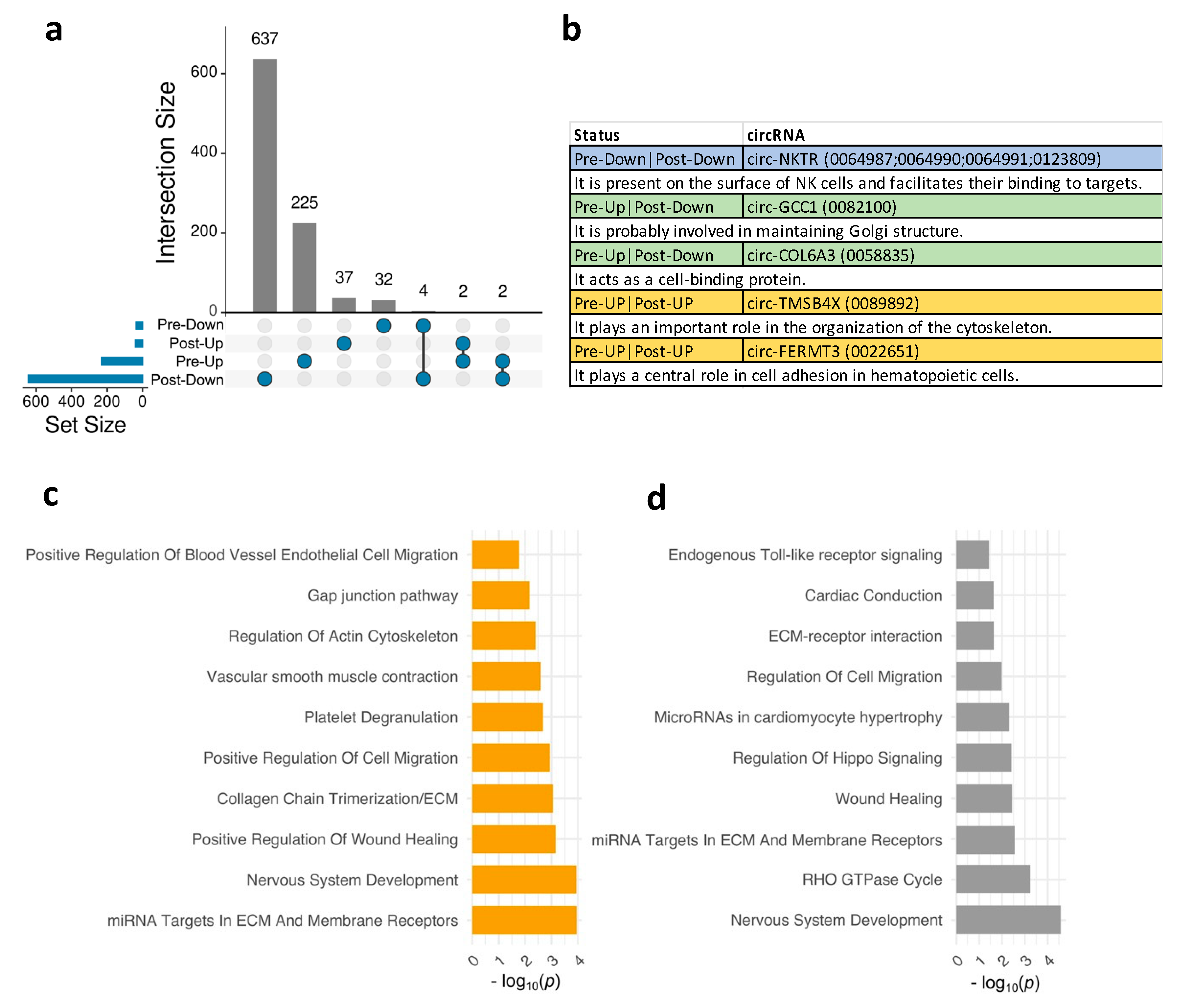
Next, we aimed to explore the shared and unique DE circRNAs in OD compared to NOD at pre- and post-surgical time points. An Upset diagram (
Figure 2a) visually depicted the limited overlap of DE circRNAs in OD versus NOD groups at pre- and post-surgery. Interestingly, among the few shared downregulated DE circRNAs were four originating from the NKTR gene (Natural Killer Cell Triggering Receptor). This gene encodes a surface receptor on natural killer (NK) cells, crucial for their binding to target cells [
24] (
Figure 2b). Other shared circRNAs included circ-TMSB4X (Thymosin Beta 4 X-Linked) and circ-FERMT3 (FERM Domain Containing Kindlin 3) upregulated at both time points (
Figure 2a,b). Upregulated circRNAs in pre-surgery that were downregulated in post-surgery were circ-GCC1 (GRIP And Coiled-Coil Domain Containing 1) and circ-COL6A3 (Collagen Type VI Alpha 3 Chain).
Then, we assessed the biological pathways and processes associated with the DE circRNAs in OD versus NOD at pre- and post-surgery (
Figure 2c-d). Enriched post-surgery pathways primarily revolved around vascular endothelial cell remodelling and migration (
Figure 2c), including gap junction pathway, positive regulation of blood vessel endothelial cell migration, and regulation of actin cytoskeleton. This might indicate vascular leak or injury. While some pathways related to vascular function remained enriched in the post-surgery DE circRNAs (
Figure 2d), there was a notable emergence of distinct pathways, such as RHO GTPase cycle, endogenous Toll-like receptor signalling, regulation of hippo signalling. The latter is recognized to impact cell proliferation and tissue repair after injury [
25].
3.3. Tissue Specificity of the Altered EV circRNAs and Their Potential Clinical Associations
A key advantage of circRNAs is their ability to harbor tissue-specific molecular signatures [
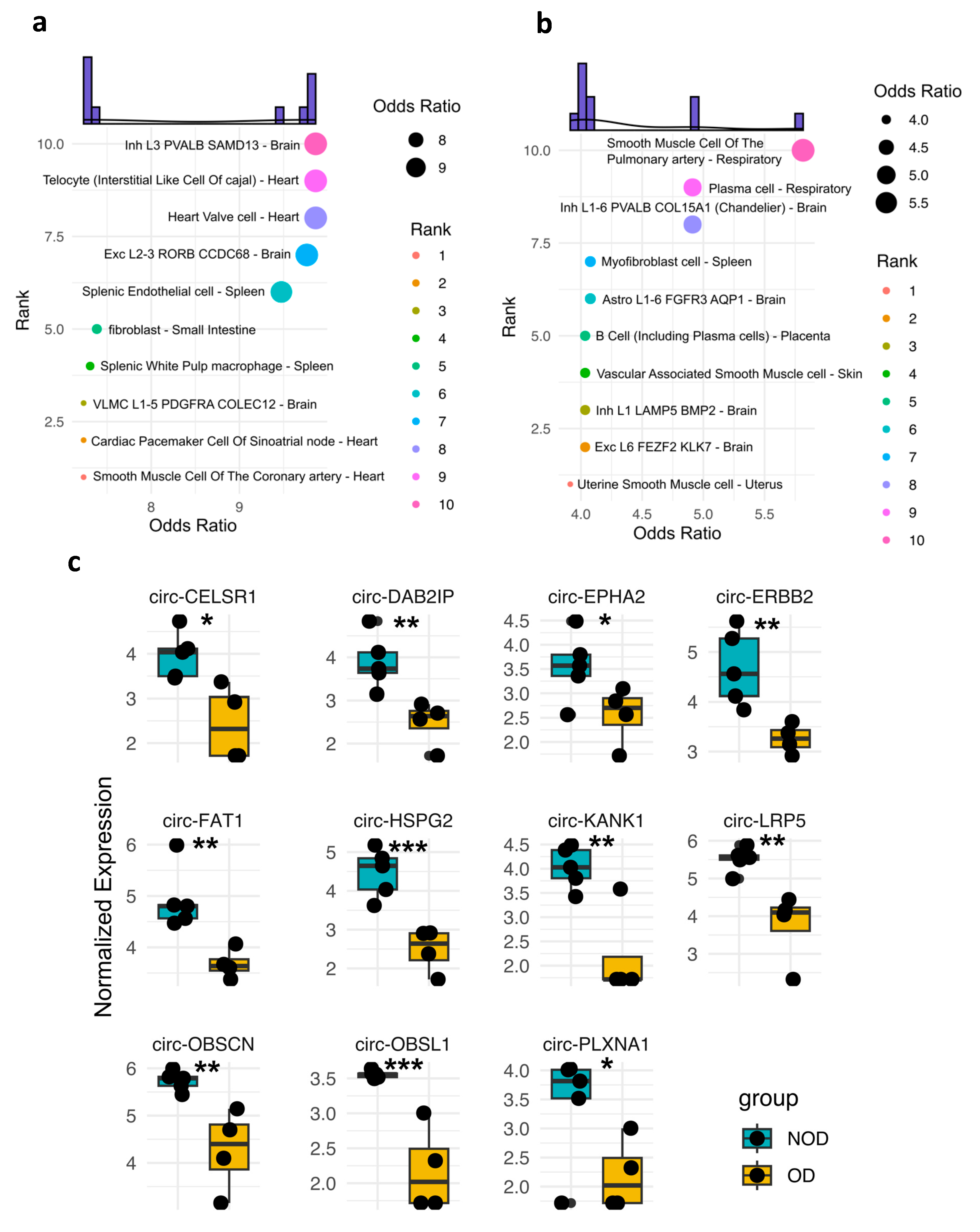
13], making them potentially valuable for diagnostic purposes. Thus, we sought to examine the potential tissue/cellular sources of DE circRNAs in OD versus NOD at pre- and post-surgery samples (
Figure 3a-b). The pre-surgery DE circRNAs were predominantly predicted to originate from specific cell types within the heart and brain (
Figure 3a). A striking observation emerged in the post-surgery DE circRNAs analysis (
Figure 3b). The top two predicted tissue sources shifted to include the respiratory system, which is particularly noteworthy in the context of postoperative respiratory failure observed in the OD group. This finding suggests a potential involvement of the respiratory system-derived circRNAs in the post-surgical response, particularly for individuals who develop respiratory failure.
We then overlaid the expression profiles of the DE circRNAs associated with the two predicted cellular sources of the respiratory system (
Figure 3c). As anticipated, all those circRNAs displayed significantly lower expression levels in OD compared to NOD, matching the decline in the respiratory function in the OD group.
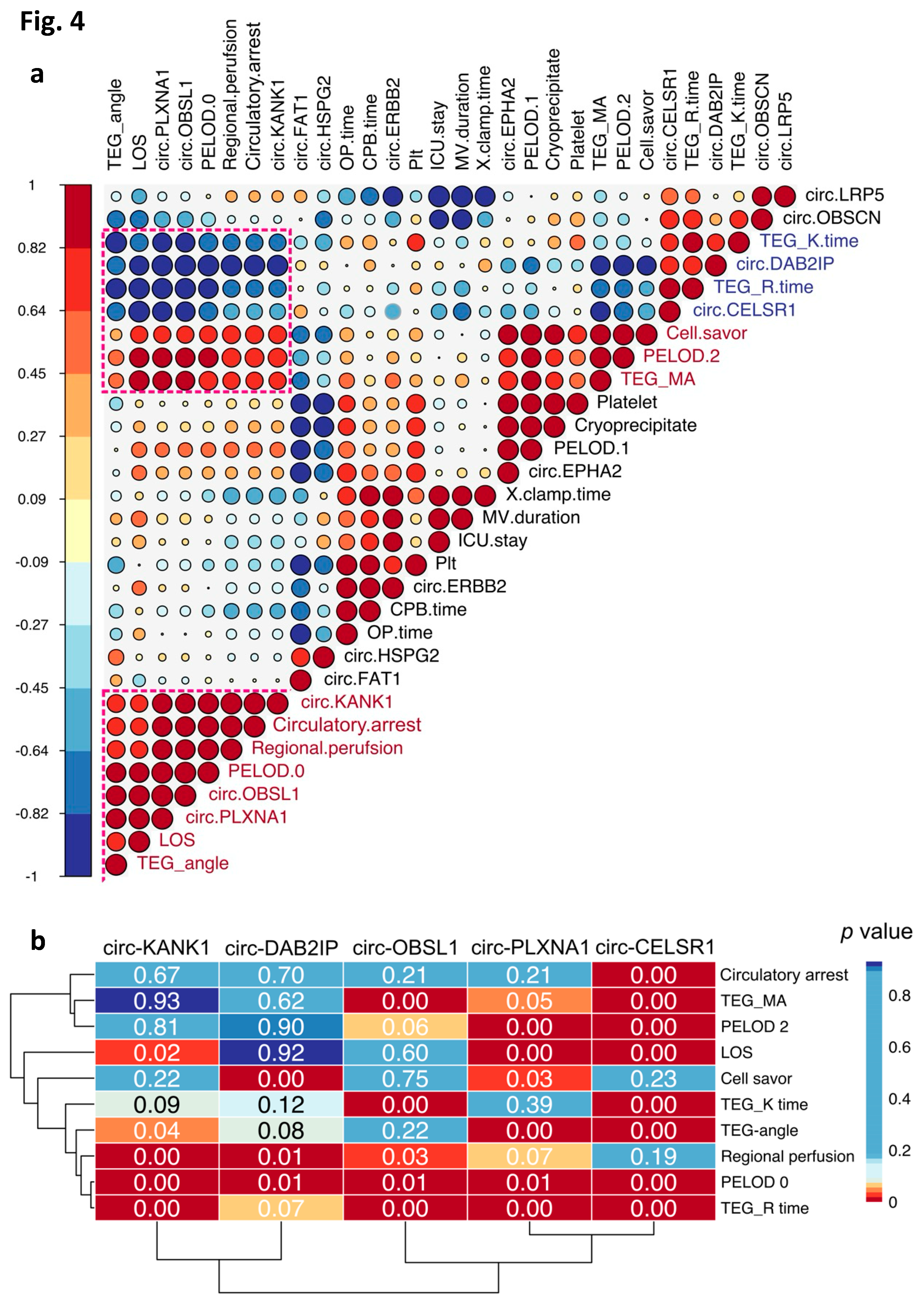
To infer potential clinical associations of these respiratory system-derived circRNAs, we conducted correlation analysis between these circRNAs and a panel of clinical and surgical parameters (
Figure 4a). Notably, the first cluster from the bottom contained the highest number of circRNAs and exhibited robust positive and negative correlations with circRNAs in two other clusters.
We further investigated the correlation significance between the circRNAs and clinical parameters within the highlighted correlation blocks (
Figure 4a,b). Interestingly, circ-CELSR1 (Cadherin EGF LAG Seven-Pass G-Type Receptor 1) displayed significant negative correlations with most of the clinical and surgical parameters. This suggests a potential role for circ-CELSR1 in the pathophysiology of postoperative respiratory failure. Additionally, we observed significant correlation between PELOD 0 (Pediatric Logistic Organ Dysfunction) with the majority of the respiratory system circRNAs (
Figure 4b), indicating potential links between these clinical parameters and the overall circRNA expression profile in the respiratory system following surgery.
4. Discussion
In our current study involving individuals with OD, primarily respiratory failure, we revealed that post-surgery circRNAs were predominantly expressed in specific cell types within the respiratory system, particularly smooth muscle cells of the pulmonary artery and plasma cells. Identifying these cell-type-specific circRNAs holds promise for developing rapid diagnostic tools and targeted therapies to minimize organ injury. Our observations were further supported by a striking shift in the expression profiles of the circRNAs between pre- and post-surgical time points. Given the half-life of circRNAs can be as short as eight hours [
26], this dramatic shift could strongly suggest a link between the surgical procedure and the development of OD. However, we cannot rule out the possibility that pre-existing vascular endothelial cell injury, as indicated by the enriched pre-surgery pathways, was exacerbated by the surgical procedure.
The correlation pattern between respiratory system-derived circRNAs and surgical/clinical parameters may shed light on potential causes of OD/respiratory failure. All five candidate circRNAs significantly correlated with PELOD, indicating a possible association with OD. Additionally, TEG measurements and regional perfusion using CBP seemed to be key risk factors influencing surgical outcomes, including respiratory failure.
The parental gene of circ-CELSR1 (Cadherin EGF LAG Seven-Pass G-Type Receptor 1), significantly correlated with all the candidate surgical/clinical parameters, was required for fetal lung devolvement and its absence caused lung morphogenesis defects [
27]. Additionally, circ-CELSR1 sponges miR-598 [
28]. Given miR-598 attenuates tissue proliferation, the downregulation of circ-CELSR1 could have increase the role of miR-598. Similarly, circ-PLXNA1 can sponge miR-214 [
29]. Because the inhibition of miR-214 attenuates lung injury [
30], the reduction of circ-PLAXNA1 can aggravate lung injury by harboring miR-214.
The observed downregulation of the regulation of hippo signalling after surgery further supports the functional significance of circRNAs. This signalling pathway is well-recognized for its critical role in tissue regeneration following injury [
31]. Yes-associated protein (YAP), the major component of hippo signalling, has been shown to ameliorate the self-renewal of type II alveolar epithelial cells (AECIIs) and their differentiation into type I alveolar epithelial cells (AECIs) to rescue gas exchange following lung injury [
32,
33,
34].
We acknowledge the limited number of individuals enrolled in this study and lack of mechanistic experiments. However, this study underscores the potential of EV circRNAs as prognostic and therapeutic tools for organ dysfunction following the congenital cardiac surgery.
Author Contributions
Conceptualization, F.A. and K.Y.; methodology, F.A. and K.Y.; data curation, F.A. and K.Y.; investigation, F.A. and K.Y.; formal analysis, F.A.; resources, K.Y.; writing – original draft, F.A.; review and editing, F.A. and K.Y.; funding acquisition, K.Y.
Funding
This study was supported by NICHD R21 HD109119 (K.Y.).
Data Availability Statement
Raw sequencing data will deposited on a public repository server upon publication acceptance.
Acknowledgments
We thank the patients and their parents for the participation in this study. We also thank Drs. Wiriya Maisat and Juan Ibla (all Boston Children’s Hospital) for the technical assistance.
Conflicts of Interest
Authors declared no conflict of interest.
References
- Marelli, A.; Gauvreau, K.; Landzberg, M.; Jenkins, K. (2010). Sex differences in mortality in children undergoing congenital heart disease surgery: a United States population-based study. Circulation 122. [CrossRef]
- Brown, K.L.; Ridout, D.; Pagel, C.; Wray, J.; Anderson, D.; Barron, D.J.; Cassidy, J.; Davis, P.J.; Rodrigues, W.; Stoica, S.; et al. (2019). Incidence and risk factors for important early morbidities associated with pediatric cardiac surgery in a UK population. J Thorac Cardiovasc Surg 158, 1185-1196.e7. [CrossRef]
- Salvin, J.W.; Scheurer, M.A.; Laussen, P.C.; Wypij, D.; Polito, A.; Bacha, E.A.; Pigula, F.A.; McGowan, F.X.; Costello, J.M.; Thiagarajan, R.R. (2011). Blood transfusion after pediatric cardiac surgery is associated with prolonged hospital stay. Ann Thorac Surg 91, 204–210. [CrossRef]
- Redlin, M.; Kukucka, M.; Boettcher, W.; Schoenfeld, H.; Huebler, M.; Kuppe, H.; Habazettl, H. (2013). Blood transfusion determines postoperative morbidity in pediatric cardiac surgery applying a comprehensive blood-sparing approach. J Thorac Cardiovasc Surg 146, 537–542. [CrossRef]
- Iyengar, A.; Scipione, C.N.; Sheth, P.; Ohye, R.G.; Riegger, L.; Bove, E.L.; Devaney, E.J.; Hirsch-Romano, J.C. (2013). Association of complications with blood transfusions in pediatric cardiac surgery patients. Ann Thorac Surg 96, 910–916. [CrossRef]
- Faraoni, D.; Nasr, V.G.; DiNardo, J.A. (2016). Overall Hospital Cost Estimates in Children with Congenital Heart Disease: Analysis of the 2012 Kid’s Inpatient Database. Pediatr Cardiol 37, 37–43. [CrossRef]
- Alhamwe, B.A.; Potaczek, D.P.; Miethe, S.; Alhamdan, F.; Hintz, L.; Magomedov, A.; Garn, H. (2021). Extracellular Vesicles and Asthma-More Than Just a Co-Existence. Int J Mol Sci 22. [CrossRef]
- Scrimgeour, L.A.; Potz, B.A.; Aboul Gheit, A.; Shi, G.; Stanley, M.; Zhang, Z.; Sodha, N.R.; Ahsan, N.; Abid, M.R.; Sellke, F.W. (2019). Extracellular Vesicles Promote Arteriogenesis in Chronically Ischemic Myocardium in the Setting of Metabolic Syndrome. J Am Heart Assoc 8. [CrossRef]
- Alhamdan, F.; Greulich, T.; Daviaud, C.; Marsh, L.M.; Pedersen, F.; Thölken, C.; Pfefferle, P.I.; Bahmer, T.; Potaczek, D.P.; Tost, J.; et al. (2023). Identification of extracellular vesicle microRNA signatures specifically linked to inflammatory and metabolic mechanisms in obesity-associated low type-2 asthma. Allergy 78, 2944–2958. [CrossRef]
- Schindler, V.E.M.; Alhamdan, F.; Preußer, C.; Hintz, L.; Alhamwe, B.A.; Nist, A.; Stiewe, T.; von Strandmann, E.P.; Potaczek, D.P.; Thölken, C.; et al. (2022). Side-Directed Release of Differential Extracellular Vesicle-associated microRNA Profiles from Bronchial Epithelial Cells of Healthy and Asthmatic Subjects. Biomedicines 10. [CrossRef]
- Ikeda, Y.; Morikawa, S.; Nakashima, M.; Yoshikawa, S.; Taniguchi, K.; Sawamura, H.; Suga, N.; Tsuji, A.; Matsuda, S. (2023). CircRNAs and RNA-Binding Proteins Involved in the Pathogenesis of Cancers or Central Nervous System Disorders. Noncoding RNA 9. [CrossRef]
- Huang, S.; Yang, B.; Chen, B.J.; Bliim, N.; Ueberham, U.; Arendt, T.; Janitz, M. (2017). The emerging role of circular RNAs in transcriptome regulation. Genomics 109, 401–407. [CrossRef]
- Wu, W.; Zhang, J.; Cao, X.; Cai, Z.; Zhao, F. (2022). Exploring the cellular landscape of circular RNAs using full-length single-cell RNA sequencing. Nature Communications 2022 13:1 13, 1–14. [CrossRef]
- Sharma, A.R.; Bhattacharya, M.; Bhakta, S.; Saha, A.; Lee, S.S.; Chakraborty, C. (2021). Recent research progress on circular RNAs: Biogenesis, properties, functions, and therapeutic potential. Mol Ther Nucleic Acids 25, 355. [CrossRef]
- Glažar, P.; Papavasileiou, P.; Rajewsky, N. (2014). circBase: a database for circular RNAs. RNA 20, 1666–1670. [CrossRef]
- Fehlmann, T.; Kern, F.; Laham, O.; Backes, C.; Solomon, J.; Hirsch, P.; Volz, C.; Müller, R.; Keller, A. (2021). miRMaster 2.0: multi-species non-coding RNA sequencing analyses at scale. Nucleic Acids Res 49, W397–W408. [CrossRef]
- Gillespie, M.; Jassal, B.; Stephan, R.; Milacic, M.; Rothfels, K.; Senff-Ribeiro, A.; Griss, J.; Sevilla, C.; Matthews, L.; Gong, C.; et al. (2022). The reactome pathway knowledgebase 2022. Nucleic Acids Res 50, D687–D692. [CrossRef]
- Aleksander, S.A.; Balhoff, J.; Carbon, S.; Cherry, J.M.; Drabkin, H.J.; Ebert, D.; Feuermann, M.; Gaudet, P.; Harris, N.L.; Hill, D.P.; et al. (2023). The Gene Ontology knowledgebase in 2023. Genetics 224. [CrossRef]
- Huang, R.; Grishagin, I.; Wang, Y.; Zhao, T.; Greene, J.; Obenauer, J.C.; Ngan, D.; Nguyen, D.T.; Guha, R.; Jadhav, A.; et al. (2019). The NCATS BioPlanet – An integrated platform for exploring the universe of cellular signaling pathways for toxicology, systems biology, and chemical genomics. Front Pharmacol 10, 437284. [CrossRef]
- Consortium, H. (2019). The human body at cellular resolution: the NIH Human Biomolecular Atlas Program. Nature 2019 574:7777 574, 187–192. [CrossRef]
- Ortega, F.G.; Roefs, M.T.; de Miguel Perez, D.; Kooijmans, S.A.; de Jong, O.G.; Sluijter, J.P.; Schiffelers, R.M.; Vader, P. (2019). Interfering with endolysosomal trafficking enhances release of bioactive exosomes. Nanomedicine 20, 102014. [CrossRef]
- Duran-Sanchon, S.; Vila-Navarro, E.; Marcuello, M.; Lozano, J.J.; Muñoz, J.; Cubiella, J.; Diez, M.S.; Bujanda, L.; Lanas, A.; Jover, R.; et al. (2019). Validation of miR-1228-3p as Housekeeping for MicroRNA Analysis in Liquid Biopsies from Colorectal Cancer Patients. Biomolecules 2020, Vol. 10, Page 16 10, 16. [CrossRef]
- Ragni, E.; Orfei, C.P.; De Luca, P.; Colombini, A.; Viganò, M.; Lugano, G.; Bollati, V.; de Girolamo, L. (2019). Identification of miRNA Reference Genes in Extracellular Vesicles from Adipose Derived Mesenchymal Stem Cells for Studying Osteoarthritis. Int J Mol Sci 20. [CrossRef]
- Zhao, M.; Qin, T.; Huang, D. (2022). ACT001 inhibits the proliferation of non-small cell lung cancer cells by upregulating NKTR expression. Thorac Cancer 13, 1772–1782. [CrossRef]
- Zhong, Z.; Jiao, Z.; Yu, F.X. (2024). The Hippo signaling pathway in development and regeneration. Cell Rep 43, 113926. [CrossRef]
- Fischer, J.W.; Leung, A.K.L. (2017). CircRNAs: a regulator of cellular stress. Crit Rev Biochem Mol Biol 52, 220–233. [CrossRef]
- Yates, L.L.; Schnatwinkel, C.; Murdoch, J.N.; Bogani, D.; Formstone, C.J.; Townsend, S.; Greenfield, A.; Niswander, L.A.; Dean, C.H. (2010). The PCP genes Celsr1 and Vangl2 are required for normal lung branching morphogenesis. Hum Mol Genet 19, 2251–2267. [CrossRef]
- Zeng, X.Y.; Yuan, J.; Wang, C.; Zeng, D.; Yong, J.H.; Jiang, X.Y.; Lan, H.; Xiao, S.S. (2020). circCELSR1 facilitates ovarian cancer proliferation and metastasis by sponging miR-598 to activate BRD4 signals. Molecular Medicine 26, 1–14. [CrossRef]
- Wang, L.; Liang, W.; Wang, S.; Wang, Z.; Bai, H.; Jiang, Y.; Bi, Y.; Chen, G.; Chang, G. (2020). Circular RNA expression profiling reveals that circ-PLXNA1 functions in duck adipocyte differentiation. PLoS One 15. [CrossRef]
- He, K.; Han, S.; An, L.; Zhang, J. (2021). Inhibition of MicroRNA-214 Alleviates Lung Injury and Inflammation via Increasing FGFR1 Expression in Ventilator-Induced Lung Injury. Lung 199, 63–72. [CrossRef]
- Fu, M.; Hu, Y.; Lan, T.; Guan, K.L.; Luo, T.; Luo, M. (2022). The Hippo signalling pathway and its implications in human health and diseases. Signal Transduction and Targeted Therapy 2022 7:1 7, 1–20. [CrossRef]
- Zhou, B.; Flodby, P.; Luo, J.; Castillo, D.R.; Liu, Y.; Yu, F.X.; McConnell, A.; Varghese, B.; Li, G.; Chimge, N.O.; et al. (2018). Claudin-18–mediated YAP activity regulates lung stem and progenitor cell homeostasis and tumorigenesis. J Clin Invest 128, 970. [CrossRef]
- Jia, X.; Wu, B.; Huang, J.; Fan, L.; Yang, M.; Xu, W. (2021). YAP and Wnt3a independently promote AECIIs proliferation and differentiation by increasing nuclear β-catenin expression in experimental bronchopulmonary dysplasia. Int J Mol Med 47, 195–206. [CrossRef]
- Aspal, M.; Zemans, R.L. (2020). Mechanisms of ATII-to-ATI Cell Differentiation during Lung Regeneration. Int J Mol Sci 21. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).