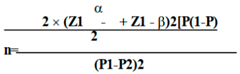Introduction
Today, arbovirus diseases are considered as one of the most important diseases transmitted by arthropods and have caused concerns in the health debate [
1,
2]. Three-day fever is a viral disease transmitted by Sandfly and Symptoms similar to influenza such as headache, high fever, back pain when moving the eye, conjunctival hyperemia, contusion, lethargy, nausea and decreased white blood cell count [
3,
4].The cause of three-day fever is transmitted by
Phlebotomus of phleboviruses (family of psychodia) [
5,
6]. Phleboviruses belong to the phenoviride family. There are 53 regions of viral antigens in the genus Phleboviruses, and 9 species have been defined by the International Report of the Committee for Classification of Viruses [
7,
8]. Sandfly carry leishmaniasis, Three-day fever and carrion disease and are important [
9,
10]. The main Vector of this disease is
Phlebotomus papatasi, which is present in different parts of the world and is the cause of the disease in humans [
10,
11,
12]. Iran, characterized by diverse climatic conditions ranging from arid deserts to humid coastal areas, provides a conducive environment for various species of sandfly. The most significant Sandfly species in Iran, with respect to disease transmission, include Phlebotomus papatasi and Phlebotomus sergenti. Northern Regions: The Caspian Sea littoral, encompassing provinces like Mazandaran and Gilan, exhibits a high prevalence of Phlebotomus species due to its humid climate. Central and Southern Regions: Desert and semidesert areas, such as those in the Yazd, Kerman, and Fars provinces, also support substantial Sandfly populations. Western and Eastern Regions: The mountainous and highland areas, including Kurdistan and Sistan and Baluchestan, respectively, present moderate to high densities of Sandfly populations. Seasonal Distribution: Sandfly activity in Iran is highly seasonal, with peak populations typically observed from late spring to early autumn. Central and Southern Iran: These regions report a higher incidence of Papatasi fever, correlating with the high density of Phlebotomus papatasi. Western and Eastern Iran: There is a lower but notable incidence of Papatasi fever. Vector Control and Prevention Strategies Controlling the spread of sand flies and subsequently Papatasi fever involves a multifaceted approach: Environmental Management: Reducing breeding sites through proper waste management and drainage improvements. Public Health Education: Increasing awareness about Sandfly habitats, peak activity periods, and preventive measures. The prevalence of Papatasi fever is closely linked to the density and activity patterns of Sandfly populations [
13,
14]. and on the other hand, unaware of the status of this disease and its Vector phleboviruses, so this study was performed for molecular monitoring of Papatasi fever virus. Soil mosquitoes were carried out in Robat Karim County, Tehran province in 2023.
Materials and Methods
Sampling Location and Method of Sandfly Hunting
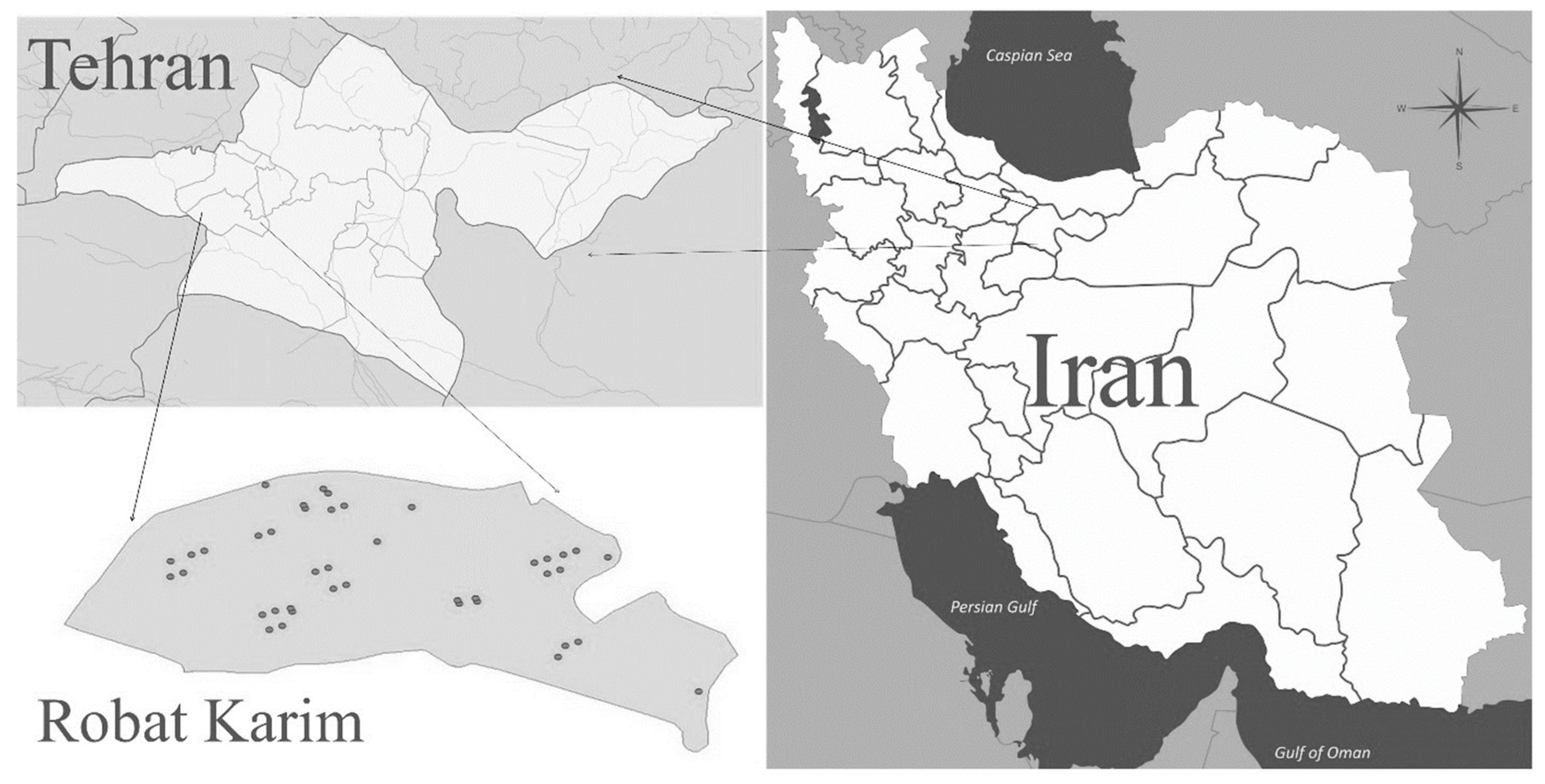
Initially, a detailed study was conducted on the distribution rate and activity season of Sandfly in the study areas in Robat Karim County. In this study, sticky traps, light traps with normal light (UV light) and an aspirator were used to hunt Sandfly. Sandfly hunting operations were carried out from the second half of March to the end of November. The captured samples were transferred to acetone-containing tubes for degreasing for several seconds. After a few minutes, 96% alcohol was added to the samples and then kept in the laboratory for identification and determination of species and refrigerated at 4 °C. The number of Sandfly was recorded by habitat (human and animal) and fishing places (indoor and outdoor), fishing date, Sex of Sandfly and their abdominal status.
Number of Samples and Identify Sandfly
For this study, about 679 Sandfly of different species were hunt, and 500 selected samples were randomly transferred to the laboratory of the Department of Medical Entomology according to the location of the catch. The collected samples were poured into a petri dish and using a drop of purulent medium on the slide and transfer of Sandfly to the slides, the Sandfly were isolated and fixed. Depending on the type of sandfly caught, the placement on the slide was different. For
Phlebotomus, upward and for
sergentomyia, the ventral surface of the head was up and the tentacles were down. Place the fixed slide on a wooden rail at room temperature for 24-48 hours to dry. In order to determine the identity and morphological characteristics of Sandfly, including cyberium teeth and temperaments, the species were studied and then the species were identified with a valid key to identify Iranian Sandfly [
15,
16]. Date, place and time of fishing were recorded along with genus and species of Sandfly. The results of identification as well as the abundance and dispersion of samples in each region were recorded.
Figure 1.
Sampling locations in Rabat Karim city.
Figure 1.
Sampling locations in Rabat Karim city.
Method of Identification and Isolation of Papatasi Fever Virus
Light traps and aspirators were used to identify and isolate Papatasi fever virus from soil mosquitoes. Immediately after catching, the mosquitoes were placed in tubes containing RNA LATER for molecular testing and evaluation of Papatasi fever virus in the transfer medium until extraction at minus 70 degrees.
Real-Time PCR
RNA extraction was performed using Qiagen Company kit instructions and Cat No./ID:85300 and according to the protocol used in the Arbovirology Department of Pasteur Institute of Tehran. In order to identify Papatasi fever virus, Nested RT-PCR technique was used One-Step RT-PCR kits for the first PCR and TaKaRa Ex Tag® kit (Mg2 + free Buffer) for the second PCR. Temperature and time program was considered for Real Time RT-PCR steps. To design the primer and identify the virus genome, sequences similar to the virus genome and different serotypes were extracted from the NCBI site. Then we started alignment using mega 6 Gene Runner 4.0 software and finally the primer was designed based on the protected points. Finally, gene expression analysis was done by Real-time device.
Statistical Analysis
Using the estimated sample size using the following formula, about 322 Sandfly were required and more (about 500 samples) were selected to be sent to the laboratory.
SPSS software version 22 was used to analyse the data of this study, frequency distribution tables, calculation of central indices, mean and calculation of scatter indices (standard deviation).
Result
Finds of Sandfly Fauna
679 Sandfly during 7 months (May, June, July, August, September, October and November) during spring, summer and autumn from 7 regions located in Robat Karim, including: Emamzadeh Abu Taleb, Manjil Abad, Vahn Abad, Hamedanak, Esmail Abad, Saleh Abad and Meymanat Abad was identified. The results showed that the activity of Sandfly in domestic and Outdoor places started in early May and ended with 2 peak activities (one in July and the other in September), in November and the highest percentage of Sandfly catch was reported for Human habitat (62.88%) and indoor places (216) which showed a high abundance of Sandfly in these places (
Table 1). Out of 679 samples of caught Sandfly, Vahn Abad region had the highest percentage of abundance of caught Sandfly (149 = 21.94%). The results of relative abundance of Sandfly by type of human and animal catch (indoor and outdoor) showed the abundance of Sandfly caught in Outdoor places (50.36%). Of the 679 samples taken, 310 (45.65%) were male and 369 (54.35%) were female. The study of sex ratio (310 males versus 369 females) showed that males are not the dominant species in this region and the highest number belongs to females (
Table 1).
Results of Identified Species
After identifying the samples, using a valid key to identify Iranian Sandfly, species were identified in the laboratory under a microscope. Female sex organs (Spermatheca) (Figure 2) and male sex organs (genitalia) (Figure 3) were identified in Sandfly under a microscope after fixation.
Species of Caught Sandfly
7 species including 3 species of Phlebotomus Sandfly and 4 species of Sergentomyia were identified among the samples. The identified species belonged to the genus Phlebotomus including: Phlebotomus papatasi, Phlebotomus alexanderi and Phlebotomus ansarri. Also identified species belonging to the genus Sergentomyia are: Sergenotomyia antennata, Sergentomyia sintoni and Sergentomyia clydei. Phlebotomus papatasi and Phlebotomus Ansari species had the highest and lowest percentages, respectively.
Results of Sandfly Infection with Phleboviruses
The results of Real-time PCR showed that out of a total of 500 samples sent to the laboratory, no positive cases of Papatasi virus infection were observed in the study areas.
Gender and Location Distribution
Gender: Distribution of males and females for each species.
Human/Animal Indoor/Outdoor: Indicates the number of sandflies found in different environments—human indoor places, human outdoor places, animal indoor places, and animal outdoor places.
Discussion
Key Observations
Species Distribution: P. papatasi and P. sergenti are generally the most abundant species across all sites. P. alexandri shows significant presence in some sites like Vahn Abad. S. sintoni and S. dentata are less common but present in certain locations.
Diversity Indices
Shannon and Simpson indices vary across sites, indicating different levels of species diversity.
Golestan Saleh Abad has the highest Shannon index (1.756), indicating the greatest species diversity, while Meymanat Abad has the lowest (1.386).
Evenness
Evenness values are generally high, indicating a relatively balanced distribution of species at most sites.
Gender Distribution
In most sites, there is a mix of male and female sandflies, though certain species may have more pronounced gender distributions.
Environmental Preferences
Sandflies are found in various environments, with some species showing preferences for indoor or outdoor, human or animal environments.
Table 1. provides valuable insights into the ecology and distribution of sandflies in Robat Karim County, which can inform vector control strategies and public health interventions for diseases like Papatasi fever.
The results of the study in Jask city showed two peaks of sandfly activity in May to June and October to November [
16].The results of the study in Kalaleh County showed two peaks of sandfly activity in July and September [
17].In this study, it was also shown that Sandfly had 2 activity peaks (in July and in September).In a study showed that male Sandfly were the predominant species in their target areas[
18].According to previous studies, male Sandfly are caught early in the maturing season and female Sandfly are caught in the late maturing season[
19,
20]..In the present study, it was shown that out of a total of 679 samples, 310 (45.65%) were male and 369 (54.35%) were female. The study of sex ratio (310 males versus 369 females) showed that males are not the dominant species in this region, and the highest number belongs to females. Also, 9 species of phlebotomus and 12 species of sergentomyia have been identified in the study of soil mosquito fauna of Khash city[
21]. In the present study, 7 species including 3 species of
Phlebotomus and 4 species of
Sergenotomy were identified. the highest abundance of Sandfly in indoor and outdoor places belonged to
P. papatasi, which was consistent with other studies conducted in different parts of the country. Other species identified are
P. alexanderi and
P. ansarri. Also identified species belonged to the genus
Sergentomyia including
S. antennata,
S. clydei and
S. syntoni.
P. ansarri was not found in previous studies, which differed from the findings of the present study. Previous studies of Phleboviruses in Sandfly and rodents have shown viral infection in these specimens[
22,
23].The first report of Corfu virus in four species of Sandfly in Iran was detected by Real Time RT-PCR [
24].In the present study, no positive case of Papatasi virus infection was found in molecular studies.
Conclusion
The results of this study showed that Sandfly, especially Phlebotomus species, are widely distributed in the region and Robat Karim is one of the sporadic areas of infectious diseases such as leishmaniasis. Due to climate change in this region, which provides the conditions for the growth and reproduction of Sandfly, especially Phlebotomus papatasi as a vector of sandfly fever. Since mosquito fever is not vaccinated, preventive measures should be taken to control and reduce mosquito-borne diseases.
Contributions of the authors
E.A. designed and collected the ticks, and identified tick species, recorded geographic coordinates and area information, wrote the manuscript, and confirmed and sent the articles.
Funding
This study received no grants from commercial, public, or nonprofit entities.
Availability of data and materials
All data obtained from this research are included in the article main text.
Acknowledgments
The authors thank Tehran province Veterinary Office and the Research Vice-chancellor of Shiraz University of Medical Sciences for their cooperation.
Competing interests
The authors declare that they have no competing interests.
Ethical approval and consent to participate
Not applicable.
Consent to participate
Not applicable.
Consent to publication
The author fully consents to the publication of the article.
References
- Alkan C, Bichaud L, de Lamballerie X, Alten B, Gould EA, Charrel RN. Sandfly-borne phleboviruses of Eurasia and Africa: epidemiology, genetic diversity, geographic range, control measures. Antiviral research. 2013;100(1):54-74. [CrossRef]
- Musso D, Rodriguez-Morales AJ, Levi JE, Cao-Lormeau V-M, Gubler DJ. Unexpected outbreaks of arbovirus infections: lessons learned from the Pacific and tropical America. The Lancet Infectious Diseases. 2018;18(11):e355-e61. [CrossRef]
- Billioud G, Tryfonos C, Richter J. The prevalence of antibodies against sandfly fever viruses and West Nile virus in Cyprus. Journal of arthropod-borne diseases. 2019;13(1):116. [CrossRef]
- Dehghani R, Kassiri H, Khodkar I, Karami S. A comprehensive overview on sandfly fever. Journal of Acute Disease. 2021;10(3):98. [CrossRef]
- Wang J, Fu S, Xu Z, Cheng J, Shi M, Fan N, et al. Emerging Sandfly–Borne Phlebovirus in China. Emerging infectious diseases. 2020;26(10):2435.
- Ready PD. Biology of phlebotomine sand flies as vectors of disease agents. Annual review of entomology. 2013;58:227-50. [CrossRef]
- Xu F, Chen H, da Rosa APT, Tesh RB, Xiao S-Y. Phylogenetic relationships among sandfly fever group viruses (Phlebovirus: Bunyaviridae) based on the small genome segment. Journal of General Virology. 2007;88(8):2312-9. [CrossRef]
- King AM, Lefkowitz E, Adams MJ, Carstens EB. Virus taxonomy: ninth report of the International Committee on Taxonomy of Viruses: Elsevier; 2011.
- Tesh RB. Phlebotomus fevers. The arboviruses: epidemiology and ecology: CRC Press; 2019. p. 15-28.
- Maroli M, Feliciangeli M, Bichaud L, Charrel R, Gradoni L. Phlebotomine Sandfly and the spreading of leishmaniases and other diseases of public health concern. Medical and veterinary entomology. 2013;27(2):123-47. [CrossRef]
- Moriconi M, Rugna G, Calzolari M, Bellini R, Albieri A, Angelini P, et al. Phlebotomine Sandfly–borne pathogens in the Mediterranean Basin: Human leishmaniasis and phlebovirus infections. PLoS neglected tropical diseases. 2017;11(8):e0005660.
- Ayhan N, Charrel RN. Emergent Sandfly–Borne Phleboviruses in the Balkan Region. Emerging Infectious Diseases. 2018;24(12):2324.
- Parhizgari N, Piazak N, Mostafavi E. Vector-borne diseases in Iran: epidemiology and key challenges. Future Microbiology. 2021;16(1):51-69. [CrossRef]
- Yaghoobi-Ershadi M. Phlebotomine sand flies (Diptera: Psychodidae) in Iran and their role on Leishmania transmission. Journal of arthropod-borne diseases. 2012;6(1):1.
- Nadim A, Javadian E. Key for species identification of Sandfly (Phlebotominae; Diptera) of Iran. 1976.
- Rassi Y, Hanafi Bojd A. Sandfly, leishmaniasis vectors. Tehran: Noavaraneelm Publication(Persian). 2006.
- Sofizadeh A, Vatandoost H, Rassi Y, Hanafi-Bojd AA, Rafizadeh S. Spatial analyses of the relation between rodent's active burrows and incidence of zoonotic cutaneous leishmaniasis in Golestan province, northeastern of Iran. Journal of arthropod-borne diseases. 2016;10(4):569.
- Kassiri H, Javadian E. Faunistic study and abundance of the phlebotomine sand flies in Saravan County, South-East Iran. Hormozgan Medical Journal. 2013;16(6):467-75.
- PARVIZI P, Amirkhani A. Mitochondrial DNA characterization of Sergentomyia sintoni populations and finding mammalian Leishmania infections in this sandfly by using ITS-rDNA gene. 2008.
- Javadian E, Nadim A. Studies on cutaneous leishmaniasis in Khuzestan, Iran. Part II. The status of Sandfly. Bulletin de la Societe de Pathologie Exotique et de ses Filiales. 1975;68(5):467-71.
- Kassiri H, Javadian E. Composition of the Sandfly fauna in Khash County, Southeast Iran. Journal of Insect Science. 2012;12(1):132.
- Tchouassi DP, Marklewitz M, Chepkorir E, Zirkel F, Agha SB, Tigoi CC, et al. Sandfly–Associated Phlebovirus with Evidence of Neutralizing Antibodies in Humans, Kenya. Emerging infectious diseases. 2019;25(4):681.
- Badakhshan M, Yaghoobi-Ershadi MR, Moin-Vaziri V, Charrel R, Hanafi-Bojd AA, Rezaei F, et al. Spatial distribution of phlebotomine sand flies (Diptera: Psychodidae) as phlebovirus vectors in different areas of Iran. Journal of medical entomology. 2018;55(4):846-54. [CrossRef]
- Alkan C, Moin Vaziri V, Ayhan N, Badakhshan M, Bichaud L, Rahbarian N, et al. Isolation and sequencing of Dashli virus, a novel Sicilian-like virus in Sandfly from Iran; genetic and phylogenetic evidence for the creation of one novel species within the Phlebovirus genus in the Phenuiviridae family. PLoS neglected tropical diseases. 2017;11(12):e0005978.
Table 1.
Sandfly fauna caught according to habitat and fishing location in Robat Karim County, 2023.
Table 1.
Sandfly fauna caught according to habitat and fishing location in Robat Karim County, 2023.
| No. |
Parts |
Sites |
Species |
Number |
Pi |
Log Pi
(Pi Ln)
|
Pi (Log Pi) |
Gender |
Human |
Animal |
| Male |
Female |
Indoor Places |
Outdoor Places |
Indoor Places |
Outdoor Places |
1
|
Central
|
Emamzadeh Abu Taleb
N=94
Shannon=1.715
Evenness=0.881
Simpson=0.205
1 – D =0.795 |
P. papatasi |
34 |
0.361 |
1.018 |
0.367 |
18 |
16 |
4 |
17 |
4 |
9 |
| P. sergenti |
13 |
0.138 |
1.980 |
0.273 |
10 |
3 |
2 |
8 |
3 |
0 |
| P. alexandri |
18 |
0.191 |
1.655 |
0.316 |
7 |
11 |
7 |
7 |
3 |
1 |
| S. sintoni |
5 |
0.053 |
2.937 |
0.155 |
2 |
3 |
0 |
4 |
1 |
0 |
| S. dentata |
8 |
0.085 |
2.465 |
0.209 |
7 |
1 |
2 |
3 |
2 |
1 |
| S. baghdadis |
12 |
0.127 |
2.063 |
0.262 |
4 |
8 |
1 |
4 |
3 |
4 |
| S. antennata |
4 |
0.042 |
3.170 |
0.133 |
3 |
1 |
0 |
2 |
0 |
2 |
2
|
Manjil Abad
N=104
Shannon=1.627
Evenness=0.908
Simpson=0.207
1 – D =0.793 |
P. papatasi |
28 |
0.269 |
1.313 |
0.353 |
9 |
19 |
5 |
12 |
7 |
4 |
| P. sergenti |
30 |
0.288 |
1.244 |
0.358 |
21 |
9 |
11 |
8 |
6 |
5 |
| P. alexandri |
17 |
0.163 |
1.814 |
0.295 |
3 |
14 |
3 |
8 |
2 |
4 |
| S. sintoni |
4 |
0.038 |
3.270 |
0.124 |
0 |
4 |
0 |
2 |
0 |
2 |
| S. dentata |
16 |
0.153 |
1.877 |
0.287 |
11 |
5 |
3 |
5 |
2 |
6 |
| S. baghdadis |
9 |
0.086 |
2.453 |
0.210 |
2 |
7 |
0 |
6 |
0 |
3 |
| S. antennata |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
3
|
Vahn Abad
N=149
Shannon=1.443
Evenness=0.581 Simpson=0.259
1 – D =0.741 |
P. papatasi |
36 |
0.241 |
1.422 |
0.342 |
9 |
27 |
12 |
9 |
6 |
9 |
| P. sergenti |
41 |
0.275 |
1.290 |
0.354 |
16 |
25 |
20 |
6 |
10 |
5 |
| P. alexandri |
52 |
0.348 |
1.055 |
0.367 |
33 |
19 |
24 |
13 |
3 |
12 |
| S. sintoni |
13 |
0.087 |
2.441 |
0.212 |
2 |
11 |
4 |
3 |
2 |
4 |
| S. dentata |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| S. baghdadis |
2 |
0.013 |
4.342 |
0.056 |
0 |
2 |
0 |
1 |
0 |
1 |
| S. antennata |
5 |
0.033 |
3.411 |
0.112 |
3 |
2 |
0 |
2 |
3 |
0 |
| 4 |
Bostan
|
Hamedanak
N=74
Shannon=1.506
Evenness=0.840
Simpson=0.219
1 – D =0.781 |
P. papatasi |
17 |
0.229 |
1.207 |
0.276 |
12 |
5 |
6 |
2 |
4 |
5 |
| P. sergenti |
23 |
0.310 |
1.171 |
0.363 |
14 |
9 |
8 |
4 |
10 |
1 |
| P. alexandri |
10 |
0.135 |
2.002 |
0.270 |
1 |
9 |
1 |
5 |
2 |
2 |
| S. sintoni |
18 |
0.243 |
1.414 |
0.343 |
6 |
12 |
7 |
4 |
5 |
2 |
| S. dentata |
4 |
0.054 |
2.918 |
0.157 |
1 |
3 |
0 |
1 |
3 |
0 |
| S. baghdadis |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| S. antennata |
2 |
0.027 |
3.611 |
0.097 |
0 |
2 |
0 |
1 |
0 |
1 |
| 5 |
Esmail Abad
N=99
Shannon=1.555
Evenness=0.868
Simpson=0.234
1 – D =0.766 |
P. papatasi |
8 |
0.080 |
2.525 |
0.202 |
3 |
5 |
1 |
2 |
1 |
4 |
| P. sergenti |
35 |
0.353 |
1.041 |
0.367 |
22 |
13 |
7 |
11 |
13 |
4 |
| P. alexandri |
27 |
0.272 |
1.301 |
0.353 |
6 |
21 |
17 |
4 |
5 |
1 |
| S. sintoni |
15 |
0.151 |
1.890 |
0.285 |
5 |
10 |
3 |
5 |
1 |
6 |
| S. dentata |
11 |
0.111 |
2.198 |
0.243 |
2 |
9 |
2 |
4 |
0 |
5 |
| S. baghdadis |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| S. antennata |
3 |
0.030 |
3.506 |
0.105 |
1 |
2 |
0 |
0 |
0 |
3 |
| 6 |
Golestan
|
Saleh Abad
N=73
Shannon=1.756
Evenness=0.902
Simpson=0.174
1 – D =0.826 |
P. papatasi |
12 |
0.164 |
1.807 |
0.296 |
5 |
7 |
3 |
4 |
0 |
5 |
| P. sergenti |
19 |
0.260 |
1.347 |
0.350 |
6 |
13 |
11 |
6 |
1 |
1 |
| P. alexandri |
15 |
0.205 |
1.584 |
0.324 |
14 |
1 |
6 |
2 |
4 |
3 |
| S. sintoni |
5 |
0.068 |
2.688 |
0.182 |
4 |
1 |
0 |
1 |
3 |
1 |
| S. dentata |
1 |
0.013 |
4.342 |
0.056 |
1 |
0 |
0 |
1 |
0 |
0 |
| S. baghdadis |
8 |
0.109 |
2.216 |
0.241 |
2 |
6 |
0 |
2 |
6 |
0 |
| S. antennata |
13 |
0.178 |
1.725 |
0.307 |
4 |
9 |
3 |
4 |
4 |
2 |
| 7 |
Meymanat Abad
N=86
Shannon=1.386
Evenness=0.861
Simpson=0.279
1 – D =0.721 |
P. papatasi |
29 |
0.337 |
1.087 |
0.366 |
13 |
16 |
18 |
9 |
0 |
2 |
| P. sergenti |
33 |
0.383 |
0.959 |
0.367 |
17 |
16 |
20 |
6 |
2 |
5 |
| P. alexandri |
10 |
0.116 |
2.154 |
0.249 |
4 |
6 |
2 |
6 |
0 |
2 |
| S. sintoni |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| S. dentata |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
| S. baghdadis |
8 |
0.093 |
2.375 |
0.220 |
2 |
6 |
1 |
6 |
0 |
1 |
| S. antennata |
6 |
0.069 |
2.673 |
0.184 |
5 |
1 |
2 |
1 |
0 |
3 |
| Total |
679 |
|
310 |
369 |
216 |
211 |
121 |
131 |
| 679 |
427 |
252 |
| 679 |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).