1. Introduction
Sinonasal undifferentiated carcinoma (SNUC) is a rare and aggressive tumor arising in the nasal cavity and paranasal sinuses and harboring a mutation in codon 172 of the isocitrate dehydrogenase 2 (
IDH2) gene in the majority of reported cases[
1]. Mutations of the IDH1 and
IDH2 genes have been reported in a variety of malignancies, including acute myeloid leukaemias, gliomas, and cholangiocarcinoma. Somatic IDH mutations always occur in residues within the active site of those genes, with hotspots in one of three arginine residues critical for isocitrate binding (
IDH1 R132,
IDH2 R140, and
IDH2 R172) [
2]. These mutations interfere with normal enzyme activity and catalyse isocitrate to 2-hydroxyglutarate (2-HG), instead of alpha-ketoglutarate (
-KG), thus leading to neomorphic enzyme activity[
3,
4]. 2-HG accumulation then leads to the inhibition of histone demethylases and TET family 5-methylcytosine hydroxylases, producing a global DNA hypermethylation. Commonly reported mutations of codon 172 of the
IDH2 gene include R172S, R172T, R172G, or R172M[
3]. In this article, we present a case of SNUC with a never-before-described
IDH2 mutation, R172A. Although the substitution affects the same codon as other previously described mutations, it is not uncommon for different single amino acid replacements affecting the same site to induce widely different consequences on protein function. Examples include R248W vs R248Q mutations of
TP53 [
5,
6], and G12D vs G12V mutations of
KRAS[
7]. It is therefore relevant to characterise the effect of a newly discovered mutation, even though different mutations occurring on the same codon have been frequently described. Given that all previously reported mutations affecting codon 172 of
IDH2 are associated with neomorphic enzyme activity, ultimately leading to hypermethylation, in this report, we investigate this possible scenario for our sample using Illumina Methylation Arrays. The findings are then compared to similar data extracted from the Gene Expression Omnibus (GEO) database.
2. Case Description
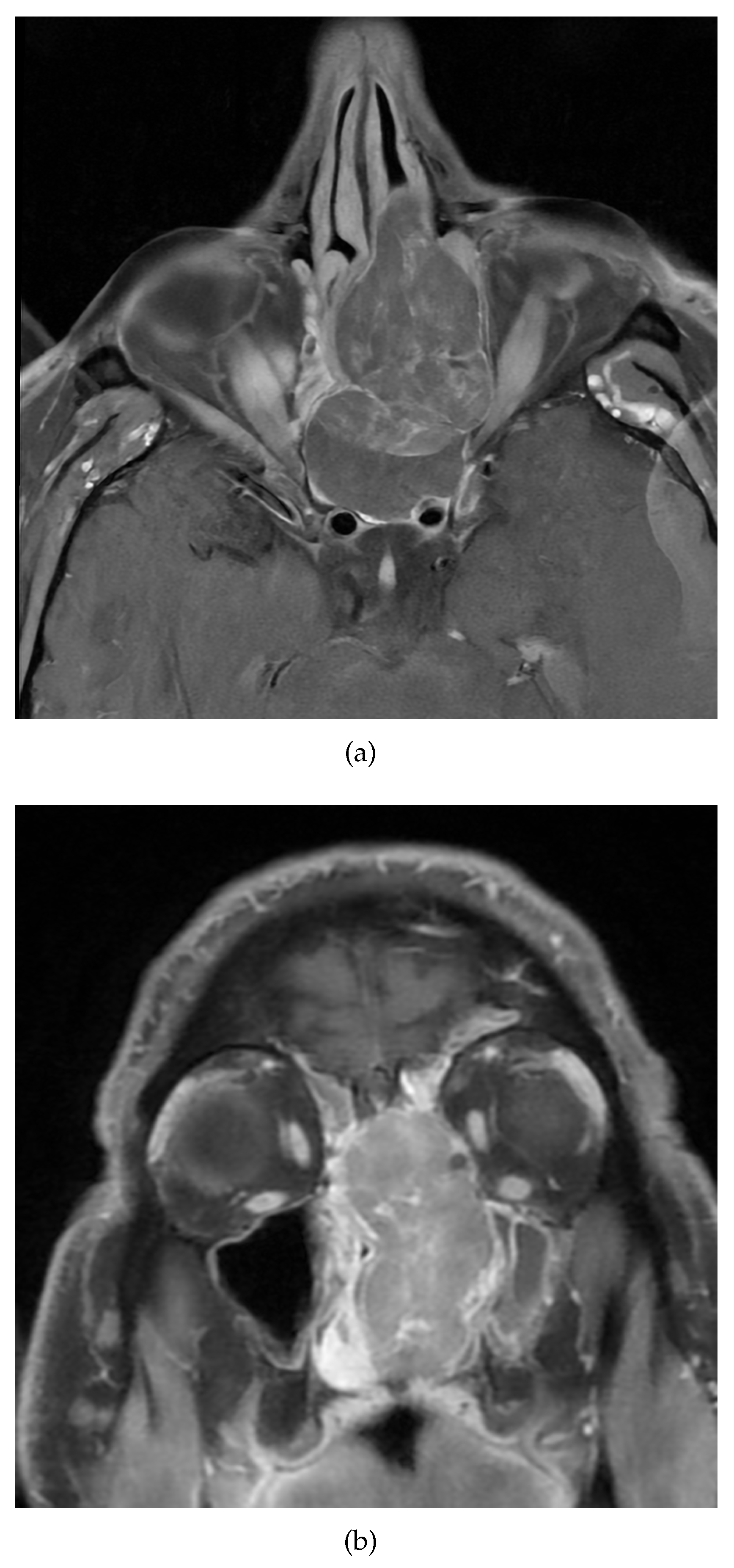
A 48-year-old non-smoker male patient presented with nasal obstruction for over a year. Computed tomography (CT) scan and magnetic resonance imaging (MRI) revealed a large, infiltrating mass involving the left nasal cavity (
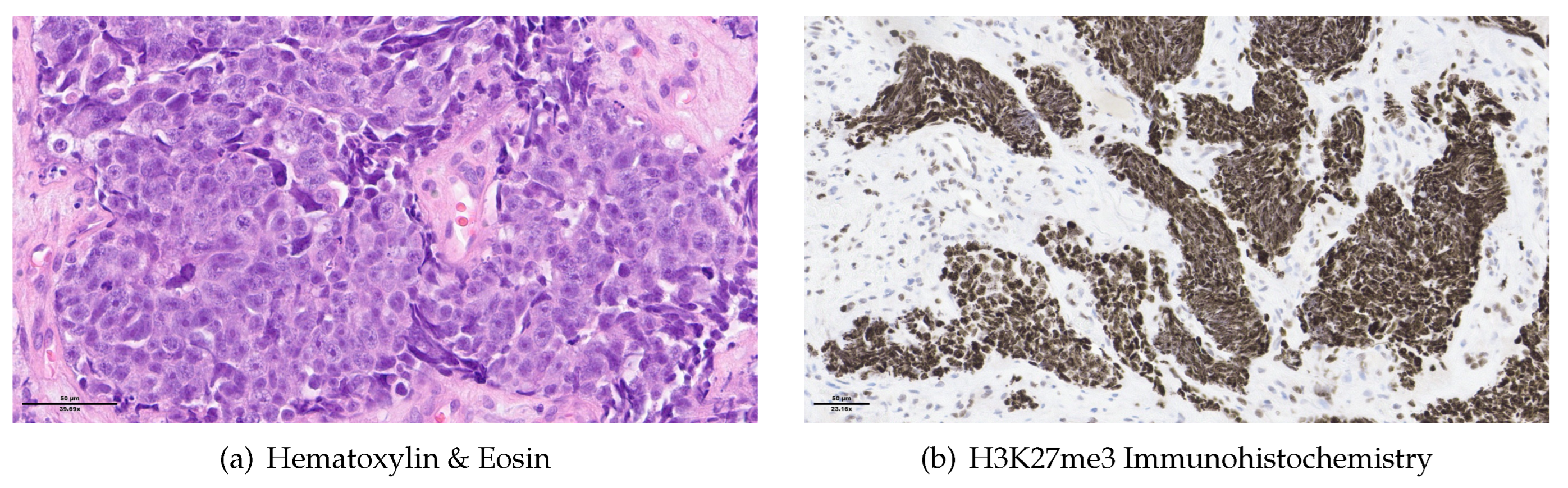
Figure 1) and ethmoid sinus, extending into the periorbital fat and brain parenchyma at the level of the right gyrus. Biopsy and histopathologic examination led to the diagnosis of SNUC (
Figure 3(a)) with increased H3K27 trimethyla tion (
Figure 3(b)), compatible with hypermethylated phenotype. It was classified as stage cT4b based on the radiological findings. Molecular profiling on the biopsy sample showed a previously not reported mutation of
IDH2 (R172A) and a common
TP53 mutation (R273H). A positron emission tomography (PET) using [
18F]fluorodeoxyglucose ([
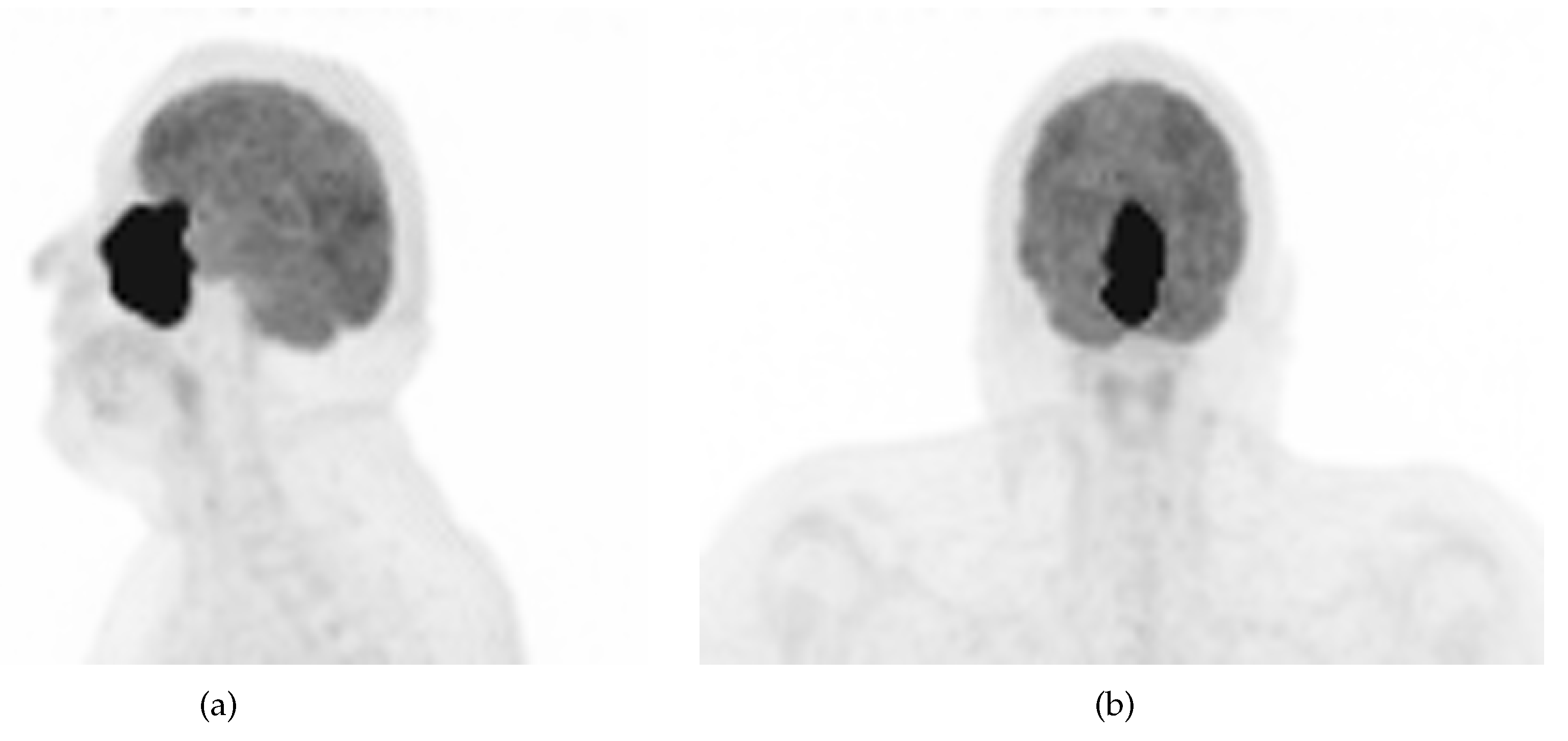
18F]FDG) highlighted an intense hypermetabolism of the lesion (
Figure 2) without evidence of metastasis. Following the diagnosis, the patient underwent 3 cycles of neoadjuvant chemotherapy composed of a combination of docetaxel, carboplatin, and 5-fluorouracil (TCF). The lesion was thereafter surgically removed after 3 months of treatment. Pathological examination of the surgical specimen showed important post-treatment remodelling indicating a good response to chemotherapy and showing only a few residual tumour cells in the nasal septum, spanning 0.6 mm on its longest axis (ypT1). No evidence of loco-regional or distant metastasis was found.
3. Results & Discussion
3.1. Molecular analysis
Targeted NGS revealed a known pathogenic variant c.818G>A (p.R273H) in exon 8 of the
TP53 gene, at a variant allele frequency (VAF) of 76%. Another yet to be classified variant,
, encoding the substitution of Arginine to Alanine at position 172 (p.R172A), was identified in exon 4 of
IDH2, with a VAF of 40%. The latter mutation is not registered in any of the databases we explored (COSMIC[
8], cBioPortal[
9], ClinVar[
10], OncoKB[
11] and Jax-CKB[
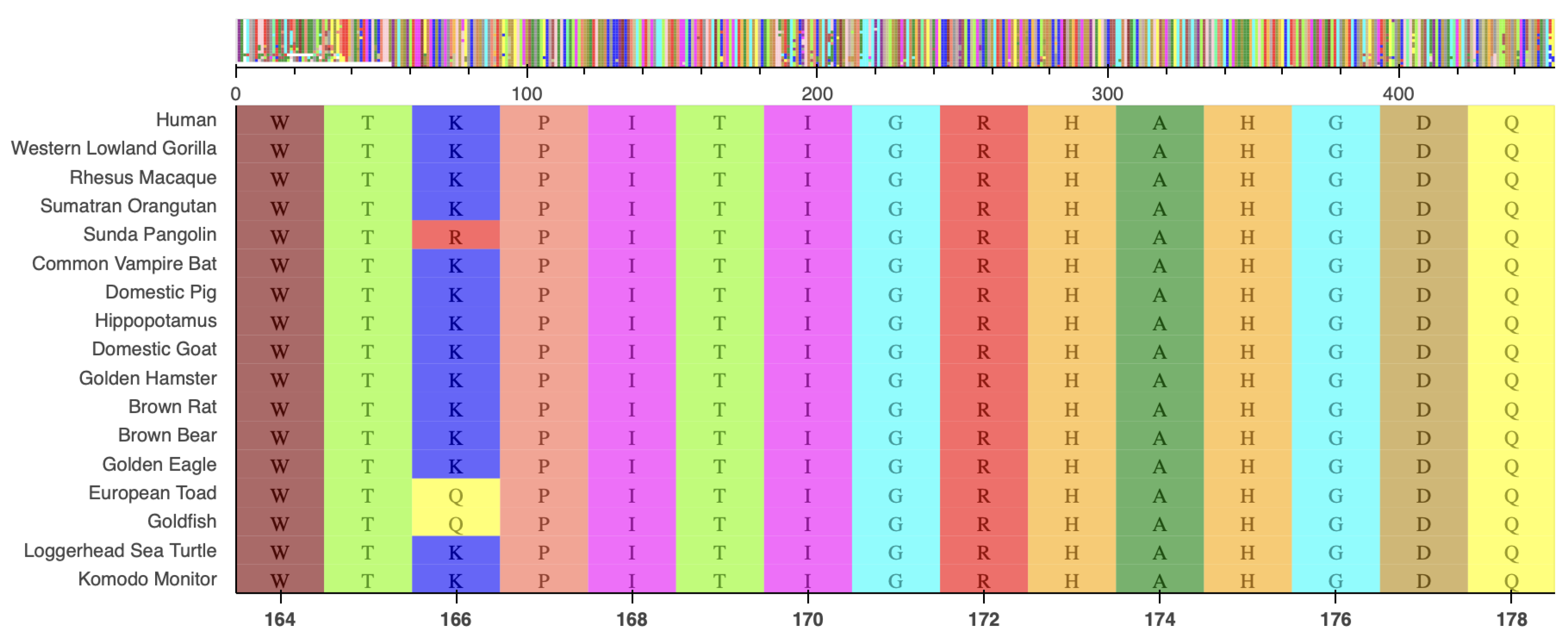
12]), and not reported in the literature. This mutation alters a known hotspot with an arginine residue critical for isocitrate binding[
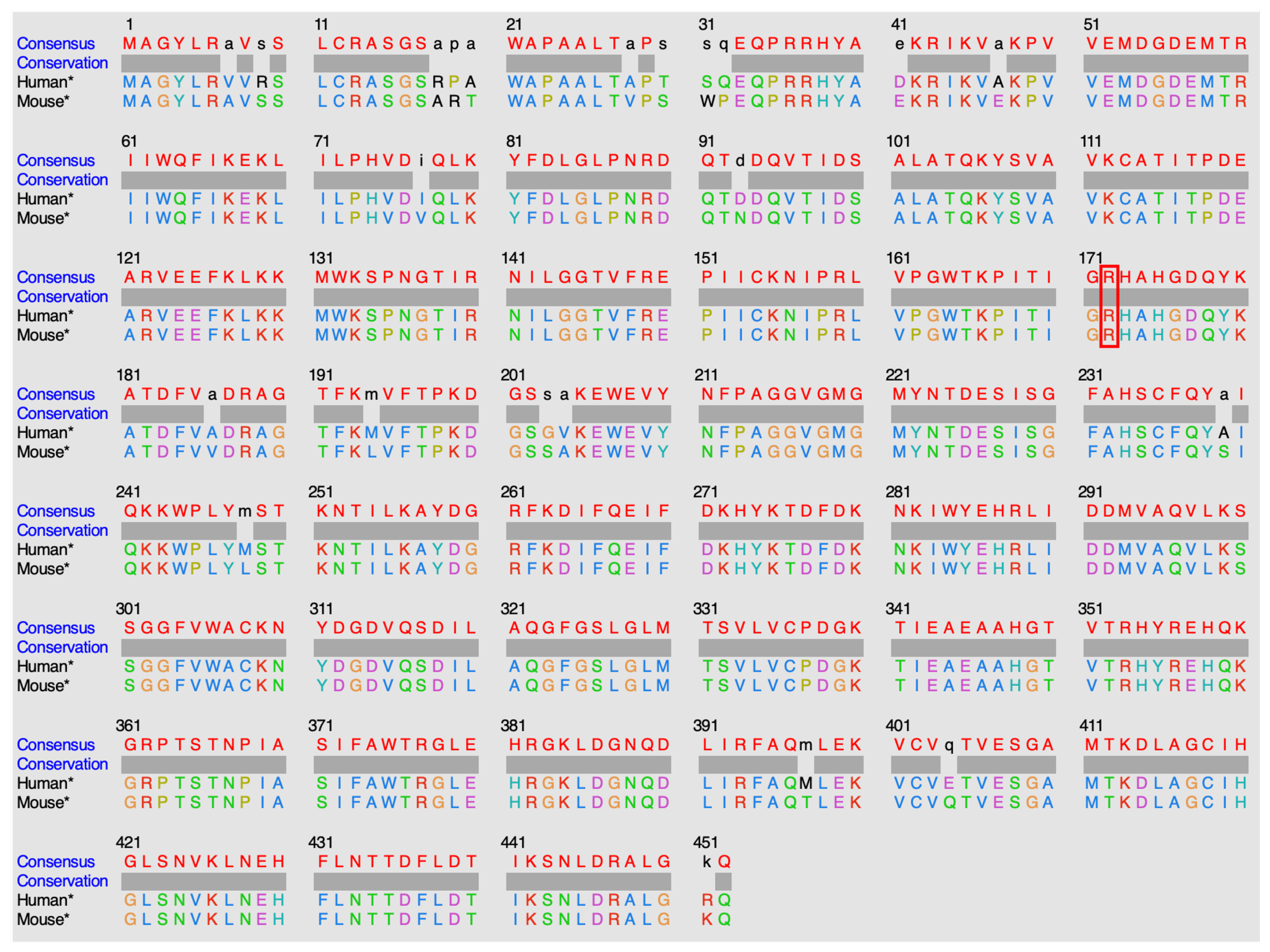
2], located in a highly conserved region (
Figure 4).
IDH2 is a mitochondrial enzyme that plays a crucial role in cellular metabolism and is involved in the Krebs cycle, which is a central metabolic pathway that occurs in the mitochondria. IDH2 allows the conversion of NADP+ into NADPH in the presence of D-threo-isocitrate. Several IDH2 experimental structures are available in the Protein Data Bank[
13] (PDB). However, no human wild-type experimental structure showing the position of residue R172, nor the substrate in the protein active site, exists. An experimental structure of IDH2 from mice, co-crystallized in the presence of isocitrate (ICT) and Mg2+, has been released. IDH2 of Homo sapiens and Mus musculus are highly similar, with a sequence identity of 95%. Both are composed of 452 amino acids, with the arginine of interest in position 172 (
Figure 5). Of note, all residues in the vicinity of R172 and the substrate are conserved. The high similarity between the two proteins in these organisms enables structural analysis of IDH2 using experimental murine structures. Thus, the murine structure with the PDB ID 5h3f[
14] is used in this study, after renumbering the residues based on the human sequence for easier discussions.
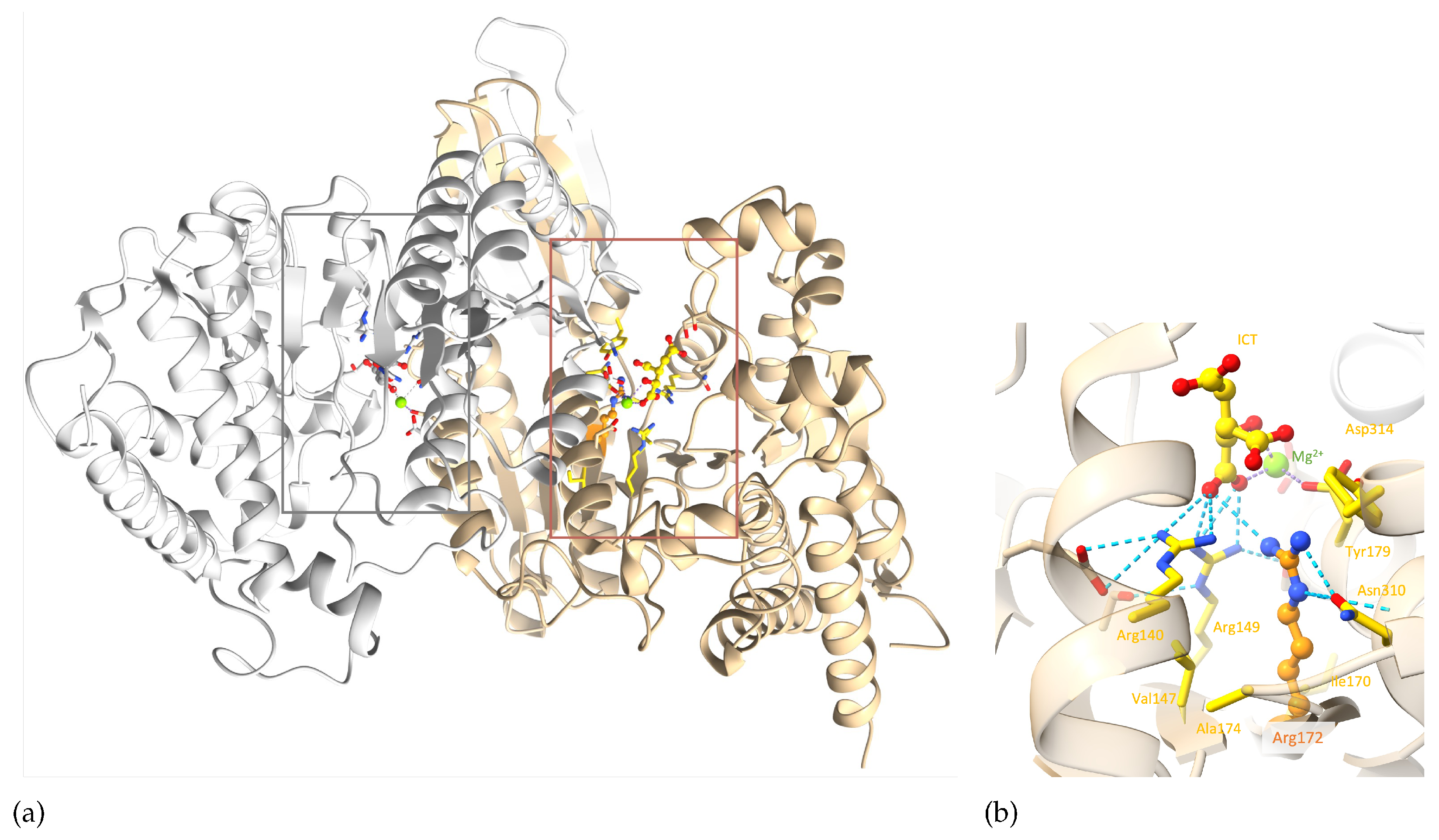
IDH2 is a homodimer protein (
Figure 6a). R172 is situated within the protein active site, more precisely within the ICT anchoring region, and participates in various interactions with the ICT substrate and neighbouring residues. Arginine is a positively charged, large and amphiphile residue, allowing it to interact with both polar and nonpolar functions. This allows R172 to participate in (i) hydrophobic interactions with V147, A174, I170, (ii) hydrogen bonds with N310 and ICT, (iii) salt bridges with ICT and D314, and (iv) a cation-
interaction with Y179 (
Figure 6b). R172 therefore plays an important role in IDH2 structural stability around the active site. Noticeably, ICT chelates to the Mg2+ on one side, while two of its carboxylate functions make ionic interactions with three highly conserved arginines at positions 140, 149 and 172 on the other side. It seems that this specific interaction scheme is crucial to enable optimal enzymatic activity, as mutations of R140 and R172 are known to decrease the ICT conversion into
-KG and increase the conversion of
-KG into R-2-hydroxyglutarate, which is known as an onco-metabolite[
15,
16,
17]. Of note, no mutation involving R149 has been characterised yet.
Structural analysis reveals that replacing any of the three arginine residues is not feasible without affecting the protein structure in the vicinity, as both the size and positive charge of these residues play crucial roles. Even a positively charged amino acid like Lysine falls short in reproducing arginine interactions due to its smaller size. Mutations of R172 to lysine, glycine, methionine and serine[
18,
19] [
20] lead to a gain of function. A structural model was created for each of these mutations, including the mutation of interest (
IDH2 p.R172A), to analyse their impact on the molecular interactions existing in the wild-type system.
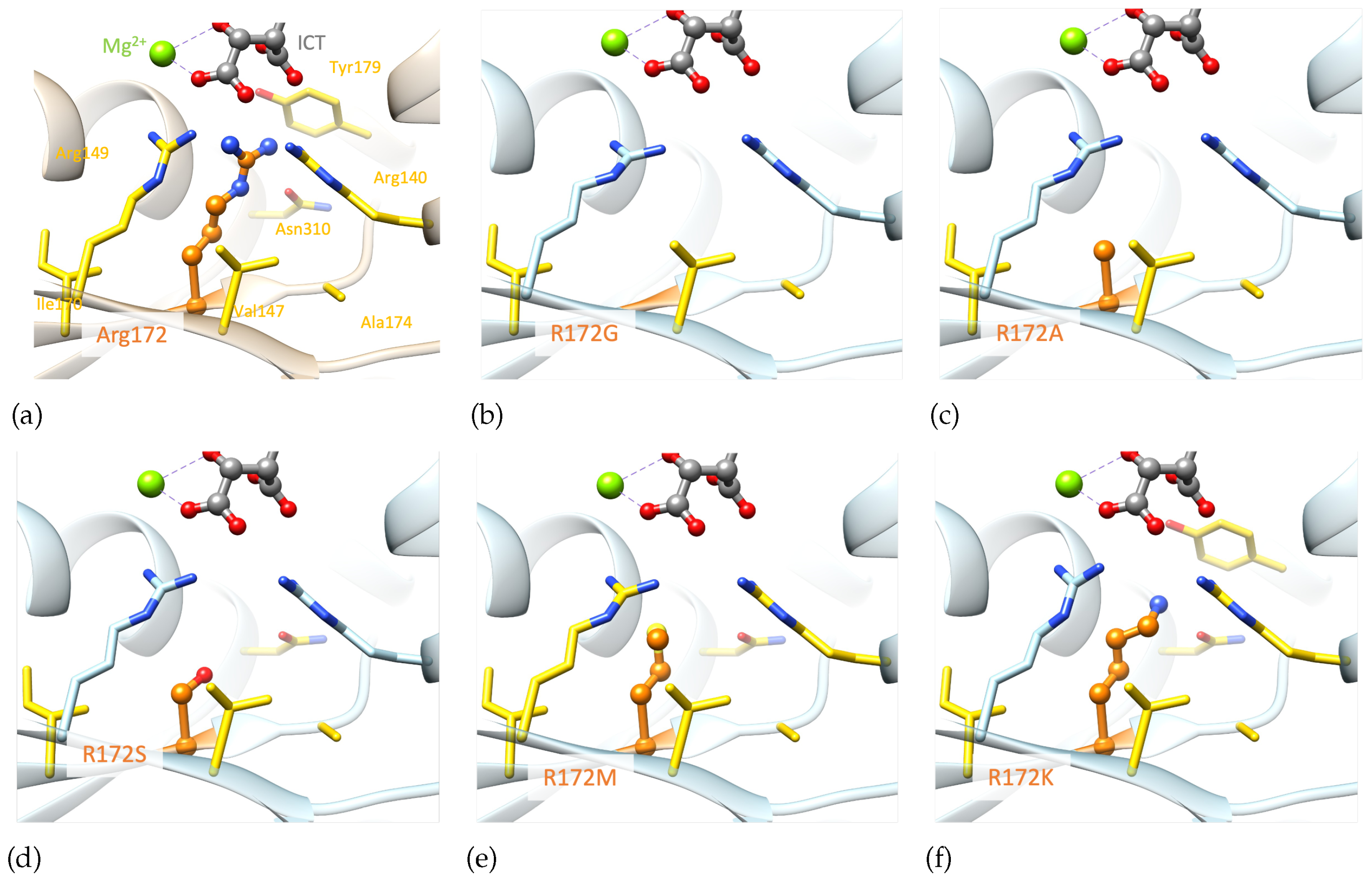
Examining the generated models highlights the importance of both the size and charge of the wild-type residue (
Figure 7), which are not reproduced by alanine. The
IDH2 p.R172A mutation is predicted to have an important structural impact as it can only reproduce a few wild-type hydrophobic interactions with V147, A174 and I170. The smaller the mutated residue, the greater the impact on the structural stability (p.R172G/A/S,
Figure 7b-d.). Alternatively, mutated residues of larger size could stabilize the region via stronger hydrophobic interactions (p.R172M/K,
Figure 7e-f.). Nevertheless, these mutated residues cannot interact with the substrate, which may affect the binding of the latter. In summary, none of the generated mutants, including p.R172K, can maintain the optimal environment for the enzymatic reaction observed in the wild-type protein.
3.2. Methylation analysis
The Brain classifier v11b4[
21] couldn’t classify our sample using the Illumina array. It was however, classified as
IDH2-mutant with a calibrated score of 0.99 using the newer Brain Tumour classifier v12.5 (
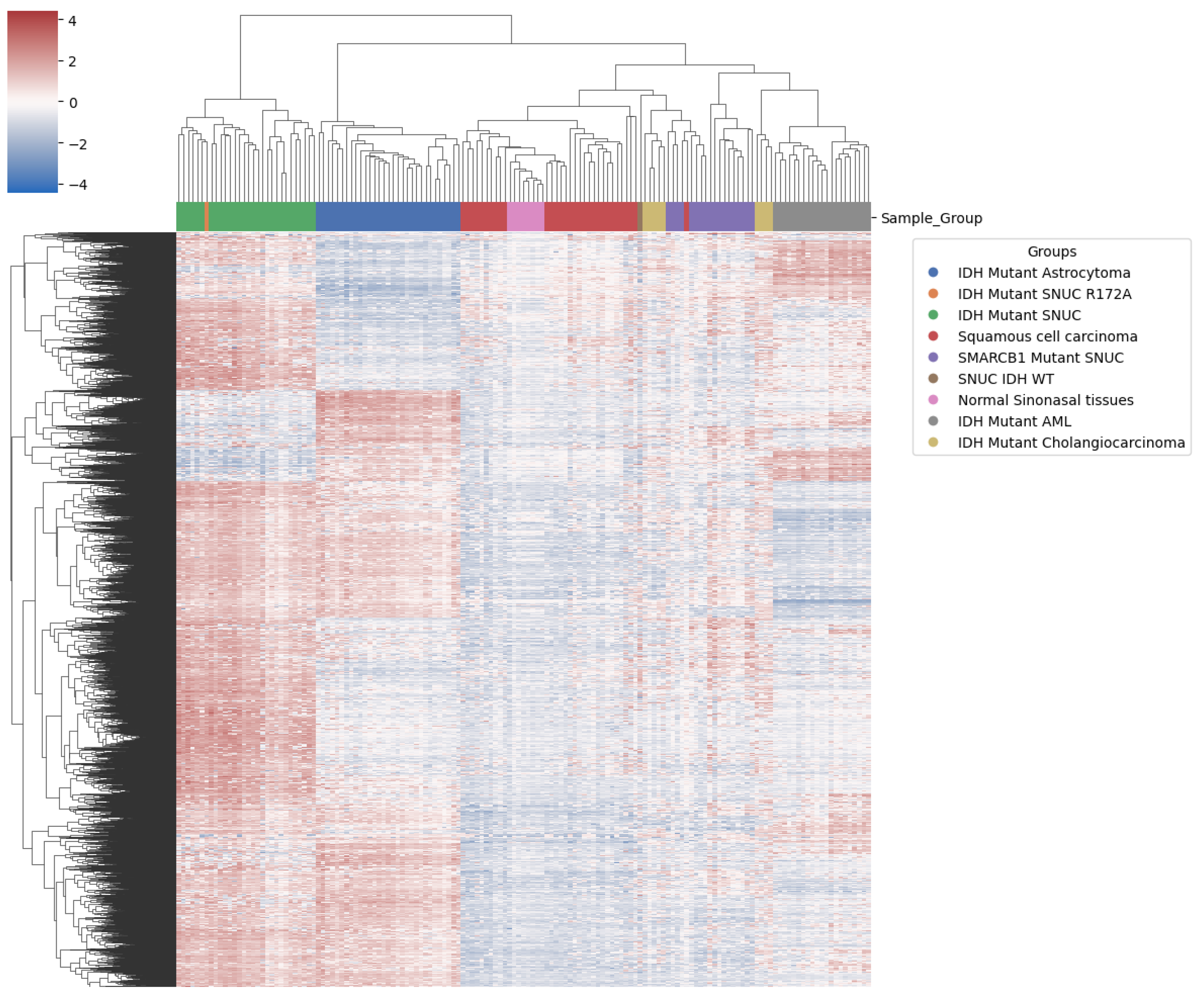
unpublished) which has this tumor class in its reference while the version v11b4 does not. Hierarchical clustering using the 10 000 most differentially expressed probes on the methylation data arrays showed proximity of our sample to other IDH mutant SNUCs (
Figure 8).
These most differentially expressed probes were thus enough to differentiate IDH mutant lesions and cell lineage. They were predominantly located in gene body regions, a pattern that follows the same proportion as in the overall probes kept for analysis (
Table 1).
This clustering also clearly differentiates
SMARCB1 mutant SNUC. Other IDH mutant samples showed clustering according to pathology findings. Interestingly, however, it didn’t differentiate clearly between normal sinonasal tissue and squamous cell carcinoma of the sinonasal region, suggesting a lack of specific methylation patterns in squamous cell carcinoma of the sinonasal region.
IDH1/2 and
SMARCB1 mutated sinonasal samples however, both show a clear differentiation, indicating significant methylation changes compared to normal samples of the same anatomical region. We also observe (
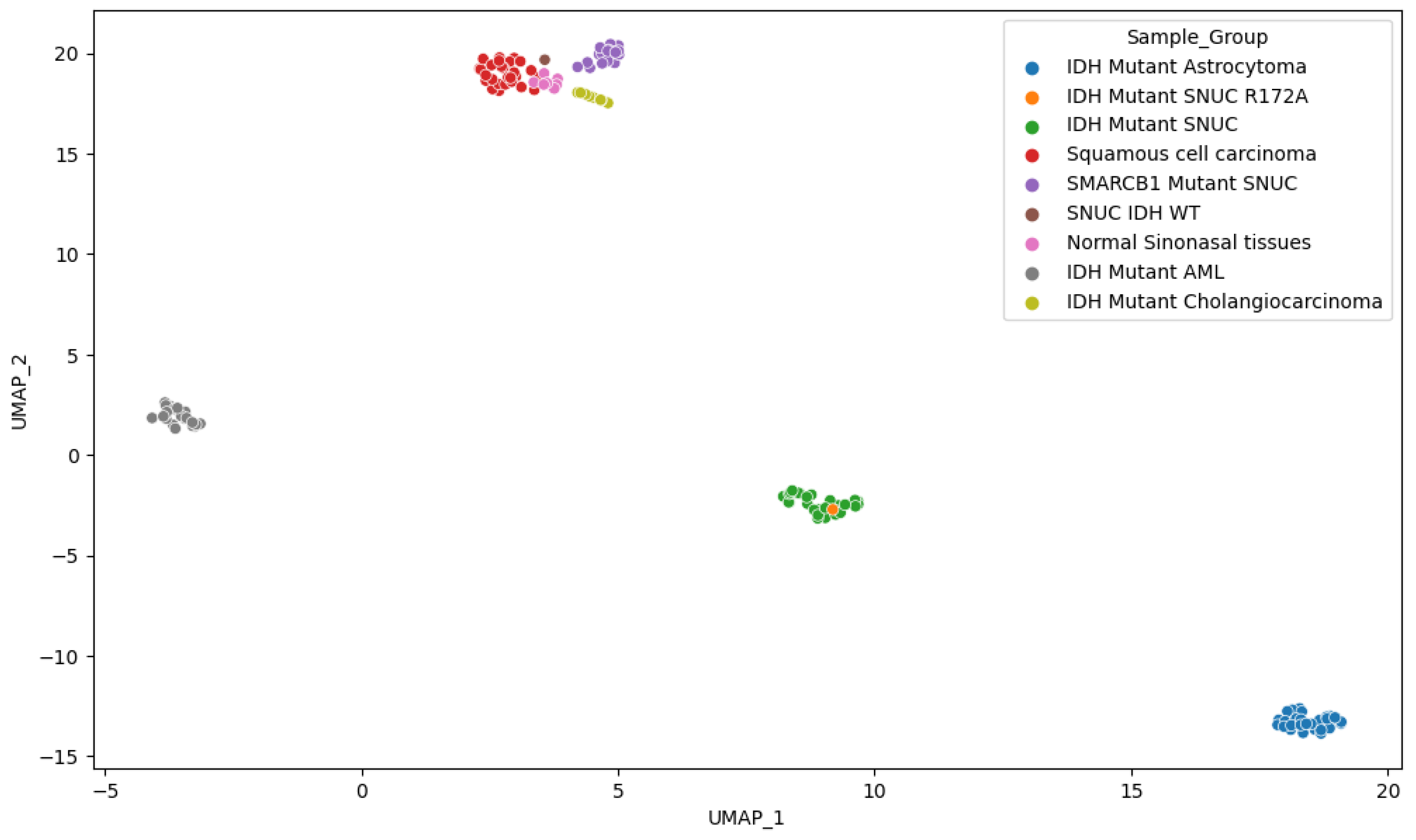
Figure 9) that although IDH mutations are shared among several samples, they still cluster predominately according to the tissue of origin as previously reported13. This suggests that IDH mutations act on a methylation background and that the overall methylation profile remains highly dependent on cellular origin. By utilizing the first 100 components, PCA explained 94.5% of the variance and positioned our sample within 1.02 standard deviation (SD) of the other IDH mutant SNUC cases.
From those analyses, we see that our sample displays a methylation pattern similar to other IDH mutant SNUCs. Given that the methylation phenotype of the IDH mutant samples is determined, among others, by the accumulation of 2-HG produced through neomorphic enzyme activity of the
IDH1/2 alleles, this suggests that the R172A mutation on
IDH2 confers a gain of function similar to other reported
IDH1/2 mutants in SNUC. Interestingly, the position of the samples in the principal components space is also highly dependent on the cell population origin. This would indicate that, although the IDH mutant samples show an increased methylation pattern, the background methylation state associated with cell lineage differentiation is still determinant. The cellular origin of lesions can therefore still be classified using methylation data, even if they have mutations widely increasing overall methylation levels. The findings of our study come with some limitations. The fact that this mutation is found in only one individual lesion is the most obvious. However, given that the mutation had never been reported previously, this is to our knowledge the only available sample, and we still believe it is worth reporting the findings associated with it. Additionally, although the external datasets from the GEO database offer an important insight into methylation patterns according to tissues of origin and lesion types, we couldn’t obtain additional information on which of the isocitrate dehydrogenase was affected in mutated samples (
IDH1 or
IDH2), limiting the precision of our analysis. This point should be assessed in further studies with access to more comprehensive datasets. Furthermore, as the external data come from different institutions, it is possible that differences in sample preparation could influence measurements provided by the methylation arrays. However, the clustering analysis couldn’t find any such trend, and other studies[
22] have also successfully used some of the same data from several labs without any obvious biases. However, this possible limitation should be mentioned and possibly further investigated if methylation-based analysis becomes more widespread in the future. Finally, a plurality of most differentially expressed probes, and therefore most determinant for clustering, were located on gene body regions. The regulatory effect of DNA methylation of these regions is inadequately understood, and interpretation is therefore limited.
4. Materials and Methods
Sequencing and methylation analyses were performed on tumor DNA extracted from the formalin-fixed paraffin-embedded biopsy samples. Targeted next-generation sequencing (NGS) was carried out using a custom amplicon-based panel covering the hotspots of 52 cancer-related genes, including
IDH1 and
IDH2[
23]. Sequencing was performed on the Ion GeneStudio S5 System (Ion Torrent, Thermo Fisher Scientific). Immunohistochemistry for H3K27me3 (clone C36B11) was performed on the Ventana Benchmark Ultra platform (Ventana Medical Systems Inc., Tucson, AZ, USA) using Ventana Optiview detection kit. The human and mouse IDH2 protein sequences were retrieved from UniProt[
24]. Sequence alignments were performed using Clustal Omega[
25]. UCSF Chimera v.1.17.3 was used for the structural and sequence visualizations and analysis[
26]. Models were generated using FoldX v.5[
27]. The analysis of methylation sites was performed using the Infinium Methylation EPIC BeadChip array from Illumina. The methylation data was loaded on the Brain Tumour classifier from Heidelberg University. Our sample was also compared with other IDH mutant SNUC (n=29), IDH wild type (WT) SNUC (n=1), SMARCB1 mutant SNUC (n=18), Normal sinonasal tissues (n=8), Sinonasal squamous cell carcinomas (n=31), IDH mutant Astrocytoma (n=31), IDH mutant Cholangiocarcinoma (n=9) and IDH mutant Acute Myeloid Leukaemia (AML, n=21). Raw methylation data were obtained from the GEO database (GSE124617[
28] & GSE196228[
22]). Methylation data preprocessing was performed using R version 4.2.2. Raw methylation data were preprocessed with the minfi[
29] package, which was used to subset overlapping probes between 450K and 850K arrays using the combineArrays function. The same package was used to verify the quality of the samples by calculating the mean detection p-values. Sex probes were removed. For array normalization, preprocessSWAN was chosen to correct for differences between type I and II probes. M-values were extracted for each sample and used for further analysis, as they have been shown to be more statistically valid than the Beta-value for differential analysis of methylation levels[
30]. Differential analysis was performed in Python v3.8.10 using SciPy[
31] and statsmodel[
32]. Hierarchical clustering was performed using only the 10,000 most differentially methylated probes, which maximized standard deviation. Principal Component Analysis (PCA) was performed on all probes from M-values using scikit-learn[
33]. Uniform Manifold Approximation and Projection (UMAP) was performed on M-values using umap-learn 20. Probes were separated into gene body regions, promoter regions (TSS200 & TSS1500), and enhancer regions using the UCSC gene region information provided for each probe by Illumina. For visualization, we used the Matplotlib[
34] and Seaborn[
35] libraries.
5. Conclusions
This work demonstrates the evaluation of new mutations from a readily available methylation array compared to results freely available on external databases using computational tools. Our findings suggest that the DNA methylation pattern associated with the R172A mutation is consistent with the already described IDH mutant SNUCs. The findings are therefore consistent with R172A mutation on IDH2 conferring a gain of function similar to other reported IDH2 mutants in SNUC. We also observe that IDH mutant SNUCs form a distinct methylation cluster, different from the methylation pattern observed in other IDH1/2 mutated tumours, consistent with IDH mutant samples remaining highly dependent on cell origin. Using a methylation array, samples can therefore still be clustered according to cellular origin even for those harbouring IDH mutations, which significantly affect methylation levels. This highlights the promising perspective of methylation and molecular modelisation analysis for diagnostic and phenotype assessment.
Author Contributions
E.H. managed the project. S.B. and K.E. performed the methylation and statistical analysis. F.S.K. and V.Z. performed the molecular modelling analysis. M.M. and B.B. performed the NGS analysis. A.R., S.S. and S.B. contributed to the case description. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Informed Consent Statement
Informed consent was obtained from the patient involved in the study.
Acknowledgments
The authors would like to thank Dr. Pierre Bady, who helped and advised the study participants on the tools and statistical methods best suited for DNA methylation analysis.
Conflicts of Interest
The authors declare no conflicts of interest.
Sample Availability
Samples of the compounds ... are available from the authors.
References
- Dogan, S.; Vasudevaraja, V.; Xu, B.; et al. DNA methylation-based classification of sinonasal undifferentiated carcinoma. Mod. Pathol. 2019, 32, 1447–1459.
- Losman, J.; Kaelin, W.J. What a difference a hydroxyl makes: mutant IDH, (R)-2-hydroxyglutarate, and cancer. Genes Dev. 2013, 27, 836–852.
- Guo, J.; Zhang, R.; Yang, Z.; et al. Biological Roles and Therapeutic Applications of IDH2 Mutations in Human Cancer. Front. Oncol. 2021, 11, 644857.
- Showalter, M.R.; Hatakeyama, J.; Cajka, T.; et al. Reproducibility Project: Cancer Biology. Replication Study: The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting alpha-ketoglutarate to 2-hydroxyglutarate. eLife 2017, 6, e26030.
- Nakazawa, S.; Sakata, K.I.; Liang, S.; et al. Dominant-negative p53 mutant R248Q increases the motile and invasive activities of oral squamous cell carcinoma cells. Biomed. Res. 2019, 40, 37–49.
- Yoshikawa, K.; Hamada, J.i.; Tada, M.; Kameyama, T.; et al. Mutant p53 R248Q but not R248W enhances in vitro invasiveness of human lung cancer NCI-H1299 cells. Biomed. Res. 2010, 31, 401–411.
- Monticone, M.; Biollo, E.; Maffei, M.; Donadini, A.; et al. Gene expression deregulation by KRAS G12D and G12V in a BRAF V600E context. Mol. Cancer 2008, 7, 92.
- Tate, J.G.; Bamford, S.; Jubb, H.C.; et al. COSMIC: the Catalogue Of Somatic Mutations In Cancer. Nucleic Acids Res. 2019, 47, D941–D947.
- de Bruijn, I.; Kundra, R.; Mastrogiacomo, B.; et al. Analysis and Visualization of Longitudinal Genomic and Clinical Data from the AACR Project GENIE Biopharma Collaborative in cBioPortal. Cancer Res. 2023, 83, 3861–3867.
- Landrum, M.J.; Lee, J.M.; Benson, M.; et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 2018, 46, D1062–D1067.
- Chakravarty, D.; Gao, J.; Phillips, S.M.; et al. OncoKB: A Precision Oncology Knowledge Base. JCO Precis. Oncol. 2017, 2017, 2473–4284.
- Patterson, S.E.; Liu, R.; Statz, C.M.; et al. The clinical trial landscape in oncology and connectivity of somatic mutational profiles to targeted therapies. Hum. Genom. 2016, 4, 1479–7364.
- Berman, H.M.; Westbrook, J.; Feng, Z.; et al. The Protein Data Bank. Nucl. Acids Res 2000, 28, 235–242.
- Xu, Y.; Liu, L.; Nakamura, A.; et al. Studies on the regulatory mechanism of isocitrate dehydrogenase 2 using acetylation mimics. Sci. Rep. 2017, 7, 9785.
- Yang, H.; Ye, D.; Guan, K.; Xiong, Y. IDH1 and IDH2 mutations in tumorigenesis: mechanistic insights and clinical perspectives. Clin. Cancer Res. 2012, 18, 5562–5571.
- Mondesir, J.; Willekens, C.; Touat, M.; et al. IDH1 and IDH2 mutations as novel therapeutic targets: current perspectives. J. Blood Med. 2016, 7, 171–180.
- Koh, J.; Cho, H.; Kim, H.; et al. IDH2 mutation in gliomas including novel mutation. Neuropathology 2015, 35, 236–244.
- Ward, P.; Patel, J.; Wise, D.; et al. The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting alpha-ketoglutarate to 2-hydroxyglutarate. Cancer Cell 2010, 17, 225–234.
- Jin, G.; Reitman, Z.J.; Spasojevic, I.; et al. 2-hydroxyglutarate production, but not dominant negative function, is conferred by glioma-derived NADP-dependent isocitrate dehydrogenase mutations. PLoS One 2011, 6, e16812.
- Chen, J.Y.; Lai, Y.S.; Tsai, H.J.; et al. The oncometabolite R-2-hydroxyglutarate activates NF-κB-dependent tumor-promoting stromal niche for acute myeloid leukemia cells. Sci. Rep. 2016, 6, 32428.
- Capper, D.; Jones, D.; Sill, M.; et al. DNA methylation-based classification of central nervous system tumours. Nature 2018, 555, 469–474.
- Jurmeister, P.; Glöß, S.; Roller, R.; et al. DNA methylation-based classification of sinonasal tumors. Nat. Commun. 2022, 13, 7148.
- Sciarra, A.; Missiaglia, E.; Trimech, M.; et al. Gallbladder Mixed Neuroendocrine-Non-neuroendocrine Neoplasm (MiNEN) Arising in Intracholecystic Papillary Neoplasm: Clinicopathologic and Molecular Analysis of a Case and Review of the Literature. Endocr. Pathol. 2020, 31, 84–93.
- Consortium, U. UniProt Consortium. UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Res. 2023, 6, D523–D531.
- Sievers, F.; Wilm, A.; Dineen, D.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 11, 539.
- Pettersen, E.; Goddard, T.; Huang, C.; et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–12.
- Schymkowitz, J.; Borg, J.; Stricher, F.; et al. The FoldX web server: an online force field. Nucleic Acids Res. 2005, 1, W382–8.
- Bledea, R.; Vasudevaraja, V.; Patel, S.; Stafford, J.; Serrano, J.; Esposito, G.; Tredwin, L.M.; Goodman, N.; Kloetgen, A.; Golfinos, J.G.; et al. Functional and topographic effects on DNA methylation in IDH1/2 mutant cancers. Sci. Rep. 2019, 9, 16830.
- Aryee, M.J.; Jaffe, A.E.; Corrada-Bravo, H.; et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 2014, 30, 1363–1369.
- Du, P.; Zhang, X.; Huang, C.C.; et al. Comparison of Beta-value and M-value methods for quantifying methylation levels by microarray analysis. BMC Bioinformatics 2010, 11, 587.
- Virtanen, P.; Gommers, R.; Oliphant, T.E.; et al. SciPy 1.0: Fundamental Algorithms for Scientific Computing in Python. Nat. Methods 2020, 17, 261–272.
- Seabold, S.; Perktold, J. statsmodels: Econometric and statistical modeling with python. In Proceedings of the 9th Python in Science Conference, 2010.
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830.
- Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 2007, 9, 90–95.
- Waskom, M.L. seaborn: statistical data visualization. J. Open Source Softw. 2021, 6, 3021.
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).