1. Introduction
Cyanobacteria prosper in many water resources worldwide, both in fresh and brackish water systems, where they represent a nuisance and a threat to public health, including in Swedish lakes and the Baltic Sea [
1,
2]. Research on cyanobacterial blooms (cyanoblooms) from recent years also shows that they occur earlier and have become more extensive than 40 years ago [
3]. The increasing frequency and intensity of cyanoblooms in lakes, rivers and sea has been linked to input of nutrients, like nitrogen and phosphorus (eutrophication) due to urban, industrial and agricultural activities, in combination with elevated average water temperatures (global warming) [
4]. The proliferation of cyanobacteria during a bloom formation increases the cyanobacterial biomass over a relatively short period of time (days to weeks) and is usually dominated by one or a few cyanobacteria taxa of the phytoplankton community [
5,
6]. This causes major problems by straining the water treatment plants, but also for the recreational and tourism activities by impelling bathing places to close.
In a phytoplankton community, toxic and non-toxic cyanobacteria co-exist, although during a cyanobloom toxic taxa might proliferate producing potent cyanotoxins that are released to the surrounding water after cell death [
7]. As cyanoblooms may occur in different fresh water sources, there is a range of unintentional exposure routes where both humans and animals can be affected if the proliferating species produce cyanotoxins. Since cyanobacteria might produce various potent toxins [
8] severe cyanotoxin outbreaks represent a versatile problem for human, domestic animals as well as ecosystems where wild animals can suffer from serious diseases or even die after ingesting water that contains toxin-producing cyanobacteria [
9,
10,
11,
12,
13]. For humans, the illness associated with the exposure to cyanotoxins is manifested by headaches and nausea with vomiting and diarrhea [
14]. Some of cyanotoxins cause liver damage and are tumor-promoters [
15,
16] while others are neurotoxins that have a paralytic effect [
17,
18,
19]. A toxin-producing cyanobloom becomes thus an extensive societal issue when it hits the surface water reservoirs that are used for the production of drinking water. Due to the underlined toxicity, cyanotoxins that are of main concern to follow are microcystins (MCs), nodularin (NOD), cylindrospermopsin (CYN), anatoxins (ATXs, anatoxin-a and homoanatoxin-a, hATX) and saxitoxins (STXs). As the knowledge about the prevalence of cyanotoxins and the diversity of the producing species in cyanoblooms in Sweden still is limited, the aim of this study was to identify their occurrence and the toxin profiles to improve the insight into the bloom characteristics.
There are several types of methods that can be applied to evaluate cyanoblooms. Traditionally, light microscopy is often used to assess cyanobacterial composition but not to identify toxic taxa [
20,
21]. More recently, 16S rRNA amplicon sequencing is used for the same purpose to determine the taxonomic composition of cyanoblooms [
22,
23,
24]. The second most commonly applied approach is the biochemical assay, Enzyme-Linked Immunosorbent Assay (ELISA) [
20,
25,
26] to screen for cyanotoxin presence in water samples, a method that possesses a higher sensitivity but is known for lower specificity in comparison to chemical methods [
27]. Molecular methods based on polymerase chain reaction (PCR or quantitative PCR, qPCR) represent an alternative technique in investigation of cyanoblooms to confirm the presence of genes encoding specific toxins. The limitation in using PCR-methods for this purpose, however, is that the confirmed presence of a toxin gene does not guarantee that the gene actually has been expressed, i.e. that the cyanobacterium has produced the toxin [
28].
In general, mass spectrometry (MS) based methods have become more commonly applied in the analysis of cyanotoxins in recent years compared to other techniques [
29,
30,
31,
32,
33], where Ultra-Performance Liquid Chromatography (UPLC-MS/MS) has been most often preferred in the detection with quantification of cyanotoxins in various sample matrices [
34,
35,
36,
37].
In the present study, cyanoblooms that occurred in Sweden during the summers of 2016 and 2017 are studied. Ninety eight cyanobloom samples were collected from a wide spatial range representing different types of aquatic environments (lakes, bays, rivers, swamps, harbors and bathing places), including four sources of drinking water. The sampling survey was community-driven effort that involved advertising through various media such as conferences, advertisements and through Facebook´s advanced location targeting.
For the aim of a selective and quantitative chemical analysis of the most interesting cyanotoxin congeners UPLC-MS/MS and Ultra-Performance Hydrophilic Interaction Liquid Chromatography with tandem mass spectrometry (UP-HILIC-MS/MS) were applied for all samples in this study [
2,
38]. In addition, the taxonomic compositions of cyanobacteria was studied with 16S rRNA amplicon sequencing, to determine the taxonomic composition of the bloom, including the cyanobacterial taxa [
39]. Microscopy was also used for species identification, mostly to facilitate the initial assessments carried out by staff in drinking water production [
40]. Using the combined approaches presented in this study a temporal pattern in profiles of cyanotoxins and species is revealed. Toxin groups and cyanobacteria from previously unmonitored surface water areas were successfully studied. The data achieved helps to promote safe drinking water production as well as to elucidate the geographical distribution of the cyanobacterial blooms in Sweden. Finally, the obtained results will also be of benefit to future studies of cyanoblooms and their toxins in Swedish surface waters towards the tailoring of efficient mitigating and preventive measures and handling strategies.
2. Results and Discussion
About half of the drinking water in Sweden is produced from lakes and rivers. Blooms of cyanobacteria in lakes are a recurring feature during the summer news and occur throughout Sweden, from south to north, and can last from a few hours to several weeks [
41]. In some lakes toxic blooms of cyanobacteria are more frequent than in others. Toxic blooms can also occur in infiltration ponds from which surface water is infiltrated (purified through the ground). Before raw water becomes drinking water, it is purified in the water treatment plants with various treatment steps that help to reduce and remove particles and organic material. The present study was planned as the need has been recognized to increase the knowledge about the cyanobacteria diversity and cyanotoxin profiles in different surface water bodies throughout Sweden. The results would serve to create a basis for bringing forth recommendations in order to handle risks related to cyanotoxins in drinking water production.
2.1. Study Sites Selection and the Sampling Strategy
As the sampling strategy was composed during this study it is considered to belong to the results section of this article (
Supplementary Material 2).
Often studies on cyanobacterial blooms collect samples from a single site in a bloom assuming an even distribution of cyanobacterial species and cyanotoxins across the bloom. However, during a cyanobloom the cell densities of cyanobacteria are very dependent on the current drift and the wind, which often causes high spatial variability across the water surface [
42,
43,
44]. Within short time periods cell and toxin concentrations at a site can change dramatically. Furthermore, toxin concentrations within a cyanobloom are influenced by a complex interplay of thermal decomposition, photolysis via UV radiation and microbial degradation which all can vary between different locations within the same water body [
45,
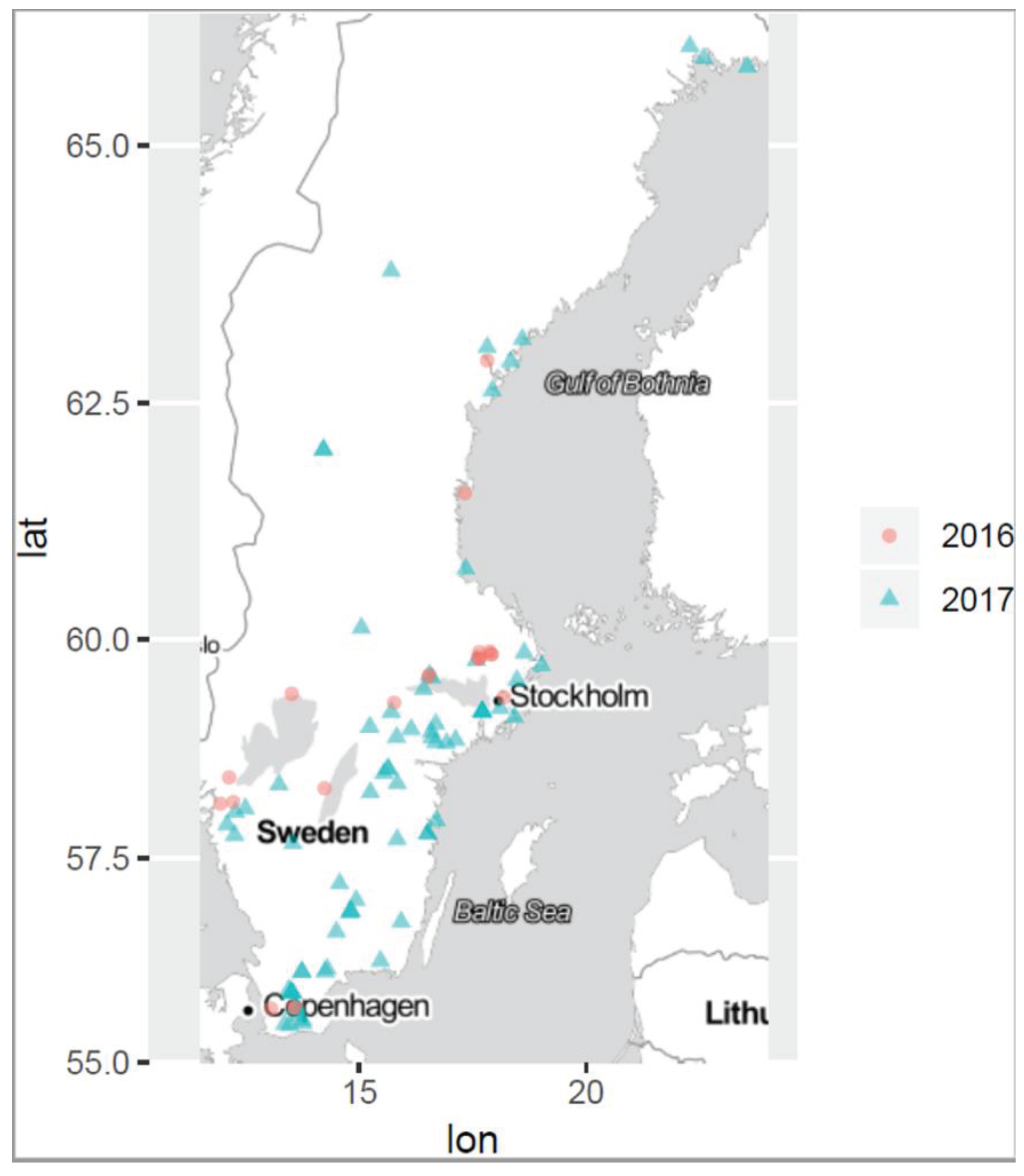
46]. Consequently, a single sample cannot be representative of the entire bloom and therefore cannot provide accurate information on the cyanobloom characteristics. Hence, multiple sampling within a cyanobloom is required for a better understanding and to ensure robust results why three separate samples were taken in each bloom in this study. The sites are depicted in the site-map in
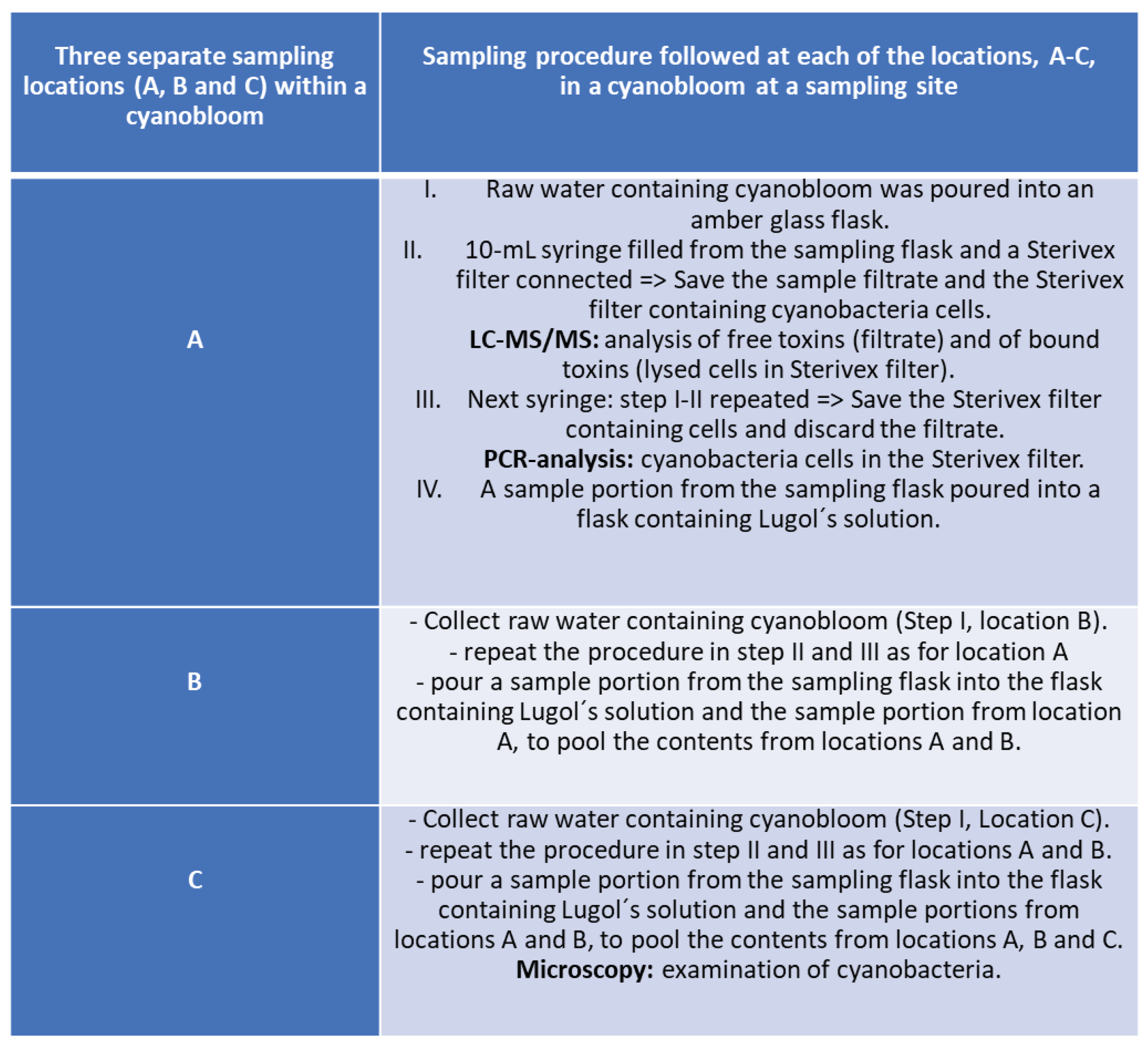
Figure 1. The sampling was dictated by ongoing blooms regardless of geographical location or time point during the summer seasons, June-October 2016 and 2017. For this study a specific sampling protocol was developed which is briefly described in the flowchart of
Figure 2, while the entire sampling procedure instruction is presented in Supplementary Material 2 and the microscopy examination followed the sampling, as a pre-analytical support to LC-MS/MS analysis, is presented in
Table S3 (Supplementary Material 1).
2.2. Diverisity of Cyanotoxin Congeners
Cyanotoxins were detected in 98 out of more than one hundred samples collected in the study. A detailed overview of the results for each sample and sampling site is presented in
Table S2 in Supplementary Material 1, which the sample names will be referring to throughout sections 2.2 and 2.3. Microcystin RR (MC-RR), was, by far, the most often detected congener, in 70% of samples and in 19% of samples it was present only as free toxin, as shown in
Table 1. The standard deviation calculated for the total found levels of MC-RR testifies that this toxin was occurring within a wide concentration range. Next most frequently occurring toxin was microcystin-LR (MC-LR) that was found in 55% of samples and in 10% of samples as a free toxin. MC-LR also stands out as the only congener detected in high concentrations as free toxin (average 148.5 µg/L). Microcystin-RR [Dha7] (MC-RR [Dha7]) is the congener that with the largest share occurs as free toxin (in 20% of samples as free toxin compared to 46% of the total share of samples in which it was detected). Other microcystines such as MC-YR and MC-RR [D-Asp3] + MC-RR [D-Asp3, (E)-Dhb7] seem to be represented in the samples to almost equal extent (in 36% and 39% of samples, respectively), although MC-YR is detected as a single and MC-RR [D-Asp3] + MC-RR [D-Asp3, (E)-Dhb7] as a double congener, since the method lacks specific m/z-transitions for each of the two congeners in the pair. Among the remaining eleven microcystins detected are: MC-LR [DAsp3] (22%), MC-WR (19%), MC (N-methyl-L)R (17%), MC-HilR (16%), MC-HtyR (13%), MC-LY (11%), MC-HphR [D-Asp3, (E)-Dhb7] (10%), MC-HtyR [D-Asp3,(E)-Dhb7] (9%), MC-LW and MC-LF with a share of 8% each, and MC-LA with the lowest frequency of occurrence was detected in 3% of samples. All of the MC-congeners were also detected as free toxins to different extents. The data on microcystins presented in
Table 1 reflects their dominance as the most common and diverse group of cyanotoxins in Swedish freshwaters of this study, represented in 82% of samples in which they were present either as single or multiple congeners together.
The frequency of occurrence was relatively low among the hydrophilic cyanotoxins such as anatoxin-a (ATX), homoanatoxin-a (h-ATX) and Micropepetin 1106 as they were present in 1-6% of samples, in free form or in total levels. The highly polar cylindrospermopsin (CYN) was present in 10% of samples, and in free form in as many as 9%. These data are in coherence with what can be expected for CYN as a very stable toxin persistent in aquatic environments that, unlike other cyanotoxins, is mainly present as a free toxin (extracellular, up to 90%) [
47,
48]. In this study, CYN occurred in samples with toxins from one or more of the other cyanotoxin groups (MCs, STXs, ATXs), but never together with nodularin (NOD), in this study. NOD was present in 13% of samples most of which were collected along the coast of the Baltic Sea or in bays connected to the Baltic Sea, from where it is known to be a typical brackish water toxin produced by, amongst others,
Nodularia spumigena. Among toxins in the saxitoxin group, the main congener, saxitoxin (STX), dominates with its presence in 45% of samples, and occurs as free toxin in a proportion of 36%. Consequently, STX is the second most abundant toxin in Swedish surface water samples and hence often detected in the same samples as MCs, although there are samples where STX is found as the only toxin (sites Hå1, 17-08, 17-13, 17-18, 17-42, 17-73,
Table S2) or together with ATX, or ATX and NOD without any MCs (sites 17-11, 17-75,
Table S2). Furthermore, within this study decarbamoylsaxitoxin (dcSTX) was detected for the first time in Swedish surface water samples at four sampling sites (17-16, 17-22, 17-33 and 17-37), two of which are among the northernmost and the southernmost sampling sites of the study (Granön, Båtskärsnäs in the Baltic Sea and Ellestadssjön, Sjöbo). Besides of MCs, dcSTX was present together with STX as the dominating toxin analogue in all four samples, and with additional presence of CYN in one of these samples (site 17-37). Presence of hATX was modest in five samples (four sampling sites), mostly in single replicates of three samplings where it co-occurred with ATX-a.
2.3. Cyanotoxin Quantities
Toxin release from a cyanobloom increases during the terminal phase of the cyanobacterial growth and during the stationary phase of the cyanobloom. When a cyanobloom collapses meaning that the cyanobacteria die, an extensive toxin release can occur [
47,
49]. Hence, the concentration of the toxins found in a sample, including the variations of the free and the bound toxin levels, depends on the time point at which the sample was taken during a cyanobloom. Consequently, for a study with non-continuous sampling at a certain sampling site over a longer period of time the toxin concentration data found is temporary. At some of the sampling sites comprised by this study, in which the cyanoblooms are frequent and recurring annually, sampling was carried out more than once (e.g. at sites 17-57, 17-68 and 17-60, 17-62), and the toxin quantity data are presented per sampling date. In addition, sampling was carried out at more than one hundred different sites, although it could not always be approved in accordance with the sampling protocol, resulting in samples inadequate for analysis and excluded from the study. The cyanotoxin quantities presented in
Table S2 (Supplementary material 1), are average values of the toxin concentrations found at three sampling locations in the same cyanobloom (sampling site). For some of the sampling sites where the toxin quantification was achieved above the LOQ (0.1 µg/L) in less than three locations within the same cyanobloom, the average toxin concentration was calculated applying “0” for the locations in which the toxin was below the LOQ or not detected. In this way exclusion of single positive sample replicates from a cyanobloom that are still relevant to visualize in the study was avoided to obtain a more representative toxin concentration data. Consequently, some of the toxin average values shown in
Table S2 are lower than the LOQ of the method. In cases where toxins were detected below the LOQ for all three locations in the cyanobloom are presented as <LOQ for the entire cyanobloom. Samples in which the toxins were detected beyond the calibration range of the method were diluted and reanalysed. The highest concentrations measured in this study for the microcystin group peaks at 26 mg/L (Så1), and decreases as written at sites 17-23, VSB1, 17-60 and 17-17. Other, total concentrations found for MCs range between 0.1-220 µg/L, and between 0.06-1400 µg/L as free toxins. Saxitoxins, mostly represented by the main analogue, STX, as described in section 2.2, were generally present in concentrations below 25 µg/L, although higher concentrations were measured at three sites 63 and 420 µg/L (site 17-21 and 17-22), including the highest concentration found för STX, 1890 µg/L (site 17-33). CYN was detected in concentrations up to 9.5 µg/L (site 17-37), while ATXs were present up to 4.2 µg/L (Så1), of which hATX was detected in five samples and always at levels ≤LOQ of the method (up to 0.1 µg/L, calculated as average of three samplings within the same cyanobloom).
The study shows a wide diversity among cyanotoxin analogues represented in Swedish surface waters, which makes the cyanotoxin profile complex even though it is differentiated between the studied sampling sites. The study also shows that the free toxin levels are generally much lower than the bound levels, being an important aspect of the drinking water production as the cyanobacteria cells are removed before the water enters the water treatment plants. However, the few quantitative data described above for MCs and STXs show that the free-toxin levels also can be high posing a challenge in drinking water production. Even though CYN and ATXs, have been detected in lower concentrations, their water solubility and stability together with the high toxicity potential make them not less important to monitor in drinking water production in comparison to other cyanotoxins found in this study.
2.3. Molecular Analysis
2.3.1. Total Community Composition of the Samples
Based on 16S rRNA amplicon analysis a total of 19.665 unique OTU´s were identified in these fresh and brackish water cyanoblooms (Supplementary material 3). However, only 516 OTU´s had a relative abundance that was higher than 1% in at least one of the samples. The most ubiquitous taxon (OTU_12), present in 91% of the samples was an Alphaproteobacteria of the family Sphingomonadaceae, i.e. not cyanobacteria. Yet in only 40% of the cyanoblooms its relative abundance was higher than 1%. In a sample from Lake Anten (17-21, replicate 3) the relative abundance of this taxon was as high as 80%. However, in another sample from the same cyanobloom its relative abundance was only 8%. The most species rich sample was taken from Lake Vomb (VS1), in which 4981 unique OTU´s were detected. Heterotrophic bacteria are expected to benefit from the organic carbon produced by cyanobacteria during blooms.
2.3.2. Cyanobacterial Community Composition
The cyanobacterial taxon OTU_10, annotated as
Aphanizomenon-NIES 81, was present in 88% of all freshwater blooms making this the most ubiquitous cyanobacterial taxon across all sampled cyanoblooms, followed by
Cyanobium PCC-6307 (OTU_13563) and
Microcystis PCC-7914 (OTU_1),
Table 2. The bloom with the most cyanobacterial species was Hornsundssjön (17-49) with an average of 212 unique cyanobacterial OTU´s. The most abundant species in Hornsundssjön was
Aphanizomenon-NIES 81 (OTU_10) with a relative abundance of 17%. Seven cyanobacterial species in this bloom had an abundance greater than 1%. Overall the community composition differed between the three samples taken from different locations within the same cyanobloom. One cyanobloom contained 10 different cyanobacterial OTU´s of which all had a relative abundance higher than 1%. However, more often one or two cyanobacteria species would dominate the bloom. For instance, in Lake Vomb where more than 70% of the bloom consisted of Microcystis PCC-7914 (OTU_1) whereas the 61 other cyanobacterial taxa were present below 1% relative abundance (combined relative abundance 3%). A study by Jankowiak et al. [
50] also showed that
Microcystis PCC-7914 was the dominating species in all the samples in their study.
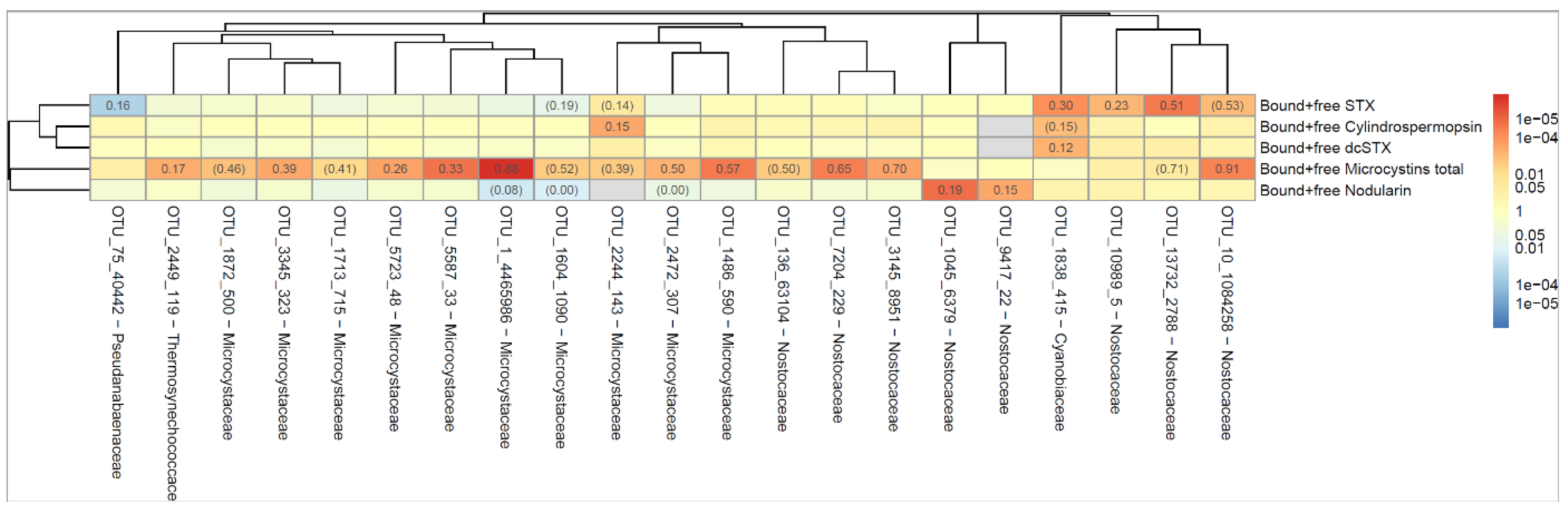
The dendrogram in
Figure 3 shows the correlation between occurrence of cyanobacterial taxa (indicated as operational taxonomic unit, OTU) and cyanotoxin. The numbers indicate the proportion of all samples where both the OTU and the toxin could be detected. For example, 'Bound and free Microcystins' were present at the same time as OTU_10_1084258 in 88% of all samples. Red color indicates that the correlation between OTU and toxin is positive and blue color if the correlation is negative. The darker the color, the lower the p-value (stronger association between OTU and toxin) as calculated according to Spearman's correlation test. The dendrogram shows how different the OTU´s and toxins are, where similarity is measured by Spearman's correlation between the level of the different bacteria/toxins. A strong correlation was seen between OTU_1 (
Microcystis PCC-7914) and MCs, and between OTU_13732 (
Aphanizomenon NIES-81) and STX,
Figure 3. Both taxa are known to produce MCs and STX, respectively [
51].
2.4. Morphological Analysis of Cyanobacteria
Morphological analyses of cyanobacteria were done on 103 samples of which 96 had presence of cyanobacteria. In total 53 cyanobacterial taxa were found from 19 genera. Doli-chospermum was the most common genus present in 67 of the samples represented by 12 species of which D. lemmermannii was the most common and abundant. Microcystis were the second most common genus in the samples present in 63 samples with M. aeruginosa as the most abundant species. The third most common genus was Aphanizomenon present in 52 of the samples and five species of which A. gracile and A. yezoense was most abun-dant. The site with highest cyanobacterial diversity based on morphological identification was Fjällnorabadet, in Lake Trehörningen, Uppland with 17 taxa identified. In 20 samples only one cyanobacterial taxon was present showing blooms with almost a monoculture of cyanobacteria. Cyanobacteria on genus level from each site is presented in
Table S3.
3. Materials and Methods
3.1. Chemicals and Reference Standards
The reference standards of cyanotoxins included in this survey were ordered from several sources (
Table S1, Supplementary materials 1). When possible, standards were purchased as solution. For the substances that were only available as solid standards, stock solutions of 5000 µg/L in methanol were prepared in-house. All stock solutions were stored dark at −20 °C. From the stock solutions three separate standard solution mixtures (A, B and C) were prepared, in methanol (MeOH) at a concentration of 625 µg/L, in which the cyanotoxins were divided between the solutions, as shown in
Table S1 (Supplementary Materials 1). Solvents used for mobile phase preparation and all other chemicals were of LC-MS grade, acetonitrile (ACN, Fisher Scientific, Loughborough, UK), methanol (LiChrosolve) and formic acid 98–100% (Merck, Darmstadt, Germany). LC–MS grade water was purified with Milli-Q purification system (Millipore, Solna, Sweden). An internal standard (IS) solution of deuterium labeled microcystin LF, D
5-MC-LF, (Gold Standards Diagnostics Horsham Inc. Warminster PA, USA) was prepared in 100 μg/L concentration by adding 100 μl of D
5-MC-LF (concentration 10 μg/mL) to 9900 μl of Milli-Q water/methanol (97 % + 3 % v/v). Lugol’s iodine solution (2 g potassium iodide and 1 g iodide in 100 mL distilled water) supplemented with acetic acid was used.
3.2. Materials
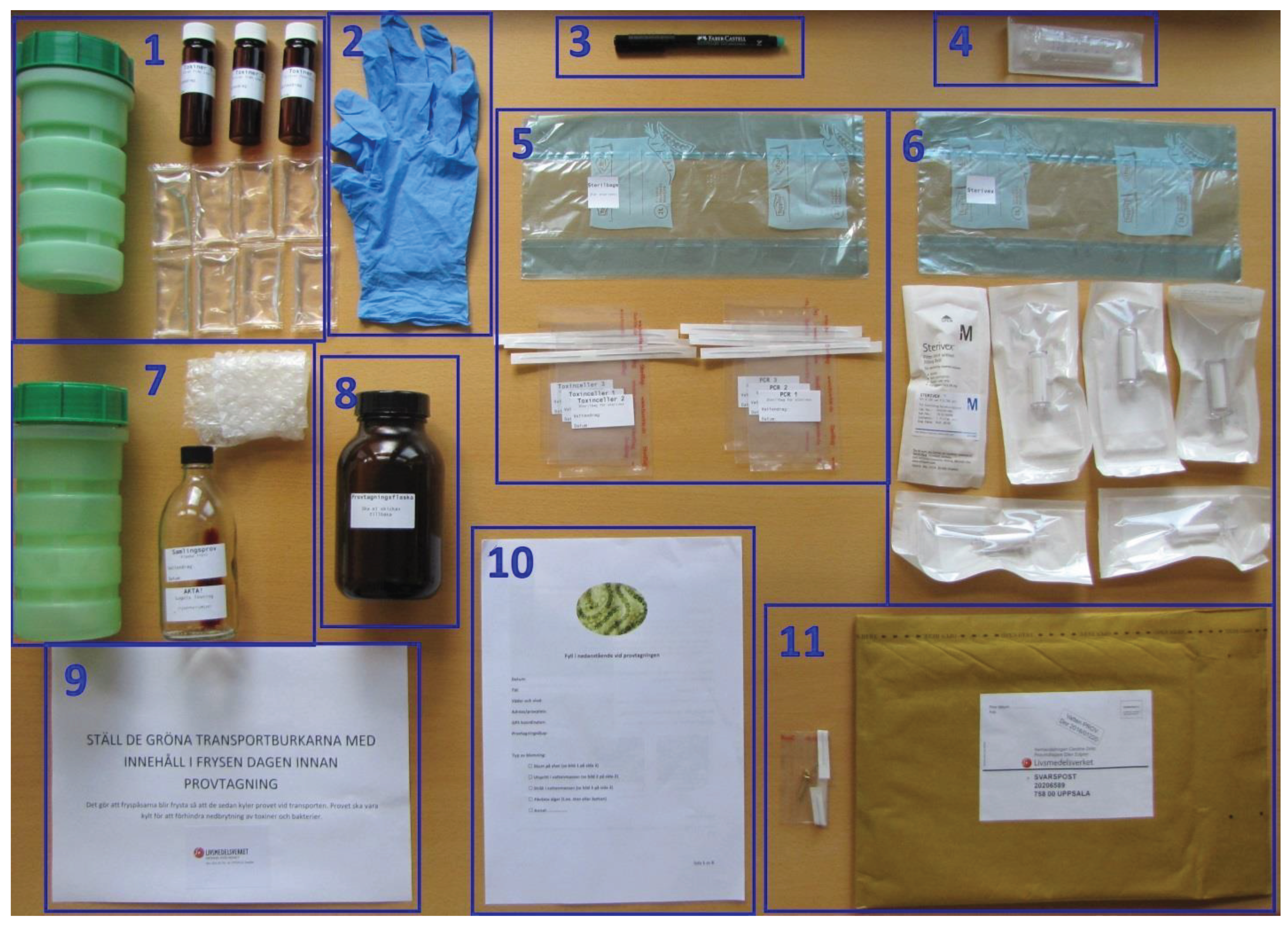
For sampling purposes, a sampling kit was prepared at SFA according to
Figure 4, containing a written protocol for sampling and sample handling at the sampling site (
Supplementary Material 2). Amber glass bottle with a wide opening (1000 mL) was used as sampling vessel to take the samples. Polypropylene syringe (Discardit II 10 mL, Beckton Dickinson and Company, Franklin Lakes, NJ, USA) was used for aspiration of sample from the sampling vessel. Sterivex filter (0.45 μm, MilliPoreSigma, Fisher Scientific GTF AB) was used to filter raw water sample containing cyanobacteria into an amber glass vial (20 mL, Skandinaviska Genetec). For further sample preparation in lab a Vortex Genie 2 Scientific Industries, a Thermo Scientific Haraeus Multi-fuge 3SR+ centrifuge and an Ultrasonication device were used. Mass spectrometry (MS) analyses were performed using a Waters Xevo TQ-S triple quadrupole mass spectrometer coupled to a Waters Acquity UPLC i-Class with a flow through needle sample manager.
3.3. Sampling and Sample Preparation
Pelagic and benthic bloom samples were collected June-October 2016 and 2017,
Table 3, from more than one hundred cyanoblooms in surface water throughout Sweden including the brackish east coast. The most northern sampling location was at 65.9150371N, 22.273922E and the most southern location at 55.4675428N, 13.4682083E (distance 1255 km), as illustrated by
Figure 1. An envelope containing sampling material according to
Figure 4 was sent by mail to each sampler (e.g. fishermen, private initiatives, water treatment plants and other voluntary samplers, besides the sampling carried out by the staff at the SFA). The samplers followed a written sampling protocol enclosed in the envelope together with the sampling material. Briefly, on arrival to the sampler, the envelope was placed in a freezer (-20 °C) over night to prepare the freezer bags for optimal transport of samples on return to the lab. The principle of the sampling procedure was based on separating cyanobacteria cells from free toxins in the raw water sample using a Sterivex filter, immediately after the sample was taken, in order to ensure that the sample would reflect its toxin profile at the time of sampling. Before the elution of the toxins from the Sterivex filter, containing the retarded cyanobloom material, 15 µL of the IS-solution (D
5-MC-LF) was added to the top of the Sterivex filter whereafter 1485 µL of 50% methanol (1:1, MeOH:Milli-Q) was applied using a 10 mL syringe pressure to the Sterivex filter until the entire applied volume was filtered. Lugol’s iodine solution (as described in
Section 3.1) was used to preserve the samples for microscopic phytoplankton examination.
3.3.1. Free Toxins
Separation of free cyanotoxins was done at the sampling site. Samples were taken from 3 different location in the cyanobloom by filling a large (1 L) amber glass bottle (sampling vessel). A portion of the sample was poured into a bottle containing Lugol's solution to prepare a sample pool of cyanobacteria cells from each of the three sampling locations in the bloom. From the remaining sample volume in the sampling bottle, 20 mL of the bloom culture was aspirated with a syringe. The sample volume in the syringe was filtered through a 0.45 µM Sterivex filter and the filtrate was collected in a 20 mL amber glass vial. This filtering procedure was applied to each of the three samples taken in the bloom, and in each bloom included in this study. At the end of a sampling at a bloom site there were 6 Sterivex filters containing biomass material, 3 vials containing filter-eluates with free toxins and a Lugol flask containing a sample pool culture from the 3 locations in the bloom. The Sterivex filters containing the bloom biomass were packed individually in sterile zip-bags and sent to the lab by mail along with vials containing the sample filtrates and the Lugol-sample pool, on the same day as the samples were collected. In addition, the sample package included notes from the sampler with the information regarding sampling date, time, weather and wind, name of the sampling location, GPS coordinates and the type of bloom collected (e.g. foam on the surface, bloom distributed in the body of water, streaks in the body of water, algae growth from rock / bottom or additional information about the sample´s look and origin that was not specifically requested in the sampling protocol).
3.3.2. Cell-Bound Toxins
For sample preparation upon sample arrival to the lab, the sterile zip-bag with the sample in Sterivex filter was placed in freezer at -80 °C. The zip-bag was removed from the freezer after 20 minutes and thawed in a room temperate water bath. The freeze-thaw procedure was repeated twice. The Sterivex filter was spiked with 15 μl of D5-microcystin LF directly on top of the filter using an automatic pipette. 1485 μl of 50% methanol in water (1:1 methanol:Milli-Q) was added to the Sterivex filter and filtered using a 10 ml syringe. The filtrate containing cell-bound toxins was collected in an amber LC vial. When the samples were prepared one day before analysis they were placed in an ultrasonic bath for 5 min prior to analysis. Both free toxins and cell-bound toxins were analyzed using the LC-MS/MS methods described in section 3.4.
3.4. Analysis of Cyanotoxins with LC-MS/MS
Two different LC-MS/MS methods were used: a reverse-phase (RP, C18) UPLC-MS/MS method for the microcystins (18), cylindrospermopsin, nodularin and anatoxins (2), according to Pekar et.al. [
2] and an UP-HILIC MS/MS method for the polar saxitoxin and the analogues (ionized in ESI+ and in ESI- mode) according to Boundy et al. [
38]. All mass spectral analyses were performed using a Waters Xevo TQ-S triple quadrupole mass spectrometer coupled to a Waters Acquity UPLC i-Class with a flow-through needle sample manager. For quantitative analysis a specific calibration curve was built for each of the toxin analogues using Targetlynx v 4.1 software (Waters, Milford, MA, USA, 2011).
3.4.1. LC-MS/MS Analysis with C18 Column
For the RP-UPLC-MS/MS analysis, the method published by Pekar et al. [
2] was applied without further modifications with respect to the chromatographic separation and MS-MS detection. This method was accredited in-house for the analysis of cyanotoxins in raw water and drinking water for 22 cyanotoxin congeners. For this study, the sample preparation procedure of the method was extended by introducing the Sterivex filter to allow for a separation of free and cell-bound toxins. A brief description on how the analysis was carried out according to the method is as follows: for the chromatographic separation an ACQUITY BEH C18 UPLC column, 2.1 × 100 mm fitted with a pre-column from VanGuard ACQUITY BEH C18 UPLC, 2.1 × 5 mm, both with a particle size of 1.7 µm (Waters, Manchester, United Kingdom). The temperature over the columns was 35 °C during analysis, the injection volume was 100 µL. Mobile phase A contained 0.1% formic acid (FA) in MiliQ water and mobile phase B 0.1% FA in acetonitrile (ACN). The gradient elution was performed as follows: 0–0.7 min, 2% B, flow 0.3 mL/min; 0.80 min, 2% B, from here the flow started to increase to 0.45 mL/min; 9.0 min, 70% B; 9.1 min, 90% B; 10.0 min, 90% B; 10.1 min, 2% B; 12.0 min, 2% B. Quantification of cyanotoxins was performed in multiple reaction monitoring (MRM) mode and positive electrospray ionization (ESI + ), with a capillary voltage of 3.0 kV. The source offset was 50 V and the source temperature 150 °C. Nitrogen (N2) was used as desolvation and cone gas at flows of 650 and 150 L/Hr, respectively. The desolvation gas temperature was 350 °C. The nebulizing gas was also N
2 at a pressure of 7.0 bars. Argon was used as collision gas at a flow of 0.15 mL/min. The compound specific MS-parameters such as cone voltage (CV), collision energy (CE) and mass transitions (m/z values) were according to Pekar et al. [
2].
3.4.2. LC-MS/MS with HILIC Column
For the UP-HILIC-MS/MS analysis the method was applied as published by Boundy et al. without further modifications. The extension in sample preparation with the Sterivex filter to separate free from cell-bound toxins was the same as in section 3.4.1, with the exception that the eluates from Sterivex-filter containing the free or the cell-bound toxins were prepared by diluting the eluate into a diluent of 70% acetonitrile. The chromatographic gradient settings were briefly as follows; mobile phases A1: water/formic acid/NH
4OH (500:0.075:0.3v/v/v); B1: acetonitrile/water/formic acid (700:300:0.1 v/v/v); A2: water/formic acid (200:1 v/v); B2: Methanol. The initial conditions consisted of 5:95 A1 and B1 at 0.4 mL/min, held for 4 min, thereafter from 5:95 to 50:50 in a linear gradient over 3.5 min. The mobile phase composition was then held while the flow rate was linearly increased to 0.6 mL/min over 1.5 min. The column was then re-equilibrated using a linear gradient to 5:95 with 0.8 mL/min over 0.5 min and then held for 0.6 min. Finally, the flow rate was decreased back to 0.4 mL/min and held for 0.4 min. The ionization parameters were as follows: capillary voltage 3.0 kV, source desolvation temperature 600 °C, and source ion block temperature of 150 °C. Nitrogen (≥95%) desolation gas flow rate was 1000 L/h, and nebulizer gas at 7.00 Bar. The collision gas flow rate of argon was set at 0.15 mL/min. A minimum of two transitions were used for each toxin analogue in the MRM analysis [
2,
38].
3.5. DNA Extraction, Purification and 16S rRNA Amplicon Sequencing
A total of 70 filter samples were collected in 2016 and 209 filter samples during 2017. The Sterivex filter was opened and part of the filter placed in a PowerBead Tube and disrupted with MP FastPrep®-24 using following settings: speed 6, CY 24×2 for 45 seconds. DNA was then extracted using DNeasy®PowerLyzer®PowerSoil® Kit from Qiagen according to the manufacturer instructions. A two-step PCR method was used to sequence bacterial-specific SSU rRNA amplicon from the DNA samples. The first PCR reaction amplified 570 bp in the variable region V3-V5 [
52] of the 16s rRNA gene using primers 357F (CCTACGGGAGGCAGCAG) and 926R (CCGTCAATTCMTTTRAGT).The PCR conditions were as follows: DNA polymerase heat activation at 95 °C (15 min), then 28 cycles containing four steps: 94 °C (60 s), a step-down to 70 °C (1 s), a ramping rate of 0.4 °C/s to 50 °C (60 s), and a ramping rate of 0.8 °C/s to 72 °C (60 s), this followed by a final extension at 72 °C (10 min). To account for random PCR drift [
53] the reactions were done in triplicate. Purification of the PCR products was done with magnetic AMPure XP beads (Agincourt). PCR products from each sample were pooled and libraries constructed with index adaptor sequence from the TruSeq DNA LT Sample Prep Kit (Illumina).
3.6. Sequence Analysis
Paired-end-sequencing (2×300bp) was performed on an Illumina MiSeq (SciLifeLab, Uppsala, Sweden). To remove forward and reverse primer sequences, raw MiSeq run fastq reads were treated with cutadapt [
54] and filtered to a MINLENGTH:100. The 3’ ends were trimmed to a Phred Quality score of 10 and forward and reverse reads were merged using VSEARCH [
55] v. 1.11.1 with “- -fastq-minovlen” option set to 16. Reads were the de-replicated (- -derep full length) and clustered into centroid OTU’s at a cut-off threshold of 97% using VSEARCH reads. Chimeras were detected and removed using UCHIME [
56] with the SILVA123.1_SSUref_tax:99 database [
57]. The LCA Clasifier [
58] 2.0 with the SILVAMOD_106 database was used to assign taxonomy.
3.7. Microscopy
Within each cyanobloom site raw water samples were collected from three locations and mixed to form a pool-sample from which a subsample was taken for analysis of phytoplankton, according to the flowchart in
Figure 2. Samples were preserved with Lugol’s solution and cyanobacteria were identified to finest possible taxon using an inverted light microscope and a modified Utermöhl technique commonly used in Scandinavia, according to Olrik et al. [
40].
4. Conclusions
This study presents a survey of cyanobacterial toxins and their geographical diversity and distribution in Swedish surface water during cyanobacterial blooms. The geographical scope of the study´s sampling and analysis of cyanotoxins is the largest conducted in Sweden to date. Two different chemical approaches (UPLC-C18 and UP-HILIC MS/MS) were applied to analyze 24 cyanotoxin congeners in 98 samples collected over a spatial distance of 1255 km. Molecular methods were used to determine cyanobacterial composition and cyanobacterial operational taxonomic unit (OTU) with the highest abundance and prevalence in collected samples. The results from the LC-MS/MS analysis showed an overall high variability of cyanotoxins with microcystins as the most commonly occurring cyanotoxins, dominated by MC-RR, followed by saxitoxins as the second most commonly detected group of cyanotoxins. In these groups the highest toxin concentrations were also measured (at single sampling sites). The detection of dcSTX at four sampling sites is the first report on its presence in surface waters in Sweden. The study further confirms that nodularin belongs in brackish water samples from the Baltic Sea, although it has recently also been detected in oysters from the west coast of Sweden [
35].
The results from the molecular methods show that the OTU with highest abundance is annotated to Aphanizomenon NIES-81, followed by the second most abundant taxon annotated to Microcystis PCC-7914.
The sampling approach developed and effectively applied in this study, where the public was interested and engaged in the sample collection, shows the potential of society to contribute to science and knowledge of our waters.
As a result of this study, a handbook has been produced with recommendations for managing risks with cyanotoxins in drinking water [
59]. The handbook is aimed for drinking water producers and other drinking water providers, like municipal companies, as well as control authorities. The purpose of the handbook is to be a support in making the necessary decisions in preventing high levels of cyanotoxins posing a health risk through the drinking water consumption.
The results contribute to increased knowledge of the presence and variation of cyanotoxins in studied sampling sites and will be of benefit to tackle future cyanobacterial blooms in fostering water quality for benefit of public health and production of drinking water. The data presented could be useful in further research studies on cyanobloom characteristics in Swedish surface waters.
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on
Preprints.org, Supplementary material 1_Dirks et al. Table S1: Reference standards of cyanotoxins used in LC-MS/MS analysis. Table S2: Toxin findings in samples from 98 individual sampling sites (cyanoblooms). Table S3: Morphological characterization of cyanobacteria from 98 sampling sites sampled during 2016 and 2017 during bloom conditions.
Author Contributions
“Conceptualization, H.P., C.D. and A.Z.M.; methodology, H.P., S.D., C.D., A.Z.M.; formal analysis, C.D., P.C., S.D., M.B., S.E., M.J., M.P.; investigation, H.P., C.D: P.C., A.Z.M., S.D.; data curation, H.P., P.C., C.D. A.Z.M.; writing—original draft preparation, A.Z.M.; writing—review and editing, A.Z.M., P.C., C.D. H.P. S.D, M.B.; visualization, A.Z.M., H.P., P.C and C.D.; project administration and funding acquisition, H.P. All authors have read and agreed to the published version of the manuscript.
Funding
This study was in part funded by the Swedish Civil Contingency Service through the project ”Förstärkt förmåga hos dricksvattenproducenterna till faroanalys och riskhantering vid toxisk algblomning i vattentäkt”, year 2016-2019. Experimental data has been produced with funding from project Strengthened capacity of drinking water producers (“Förstärkt förmåga hos drickvattenproducenterna), MSB2015-2090.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Manubolu, M.; Eklund, S.; Dutta, P.C.; Malmlöf, K. Variable Exposure and Responses to Cyanotoxins in Cattle Grazing on Pastures in the Coastal Zone of the Baltic Sea: A field Study. Int. J. Environ. Res. 2014, 8, 733–740. [Google Scholar]
- Pekar, H.; Westerberg, E.; Bruno, O.; Lääne, A.; Persson, K.M.; Sundström, L.F.; Thim, A.M. Fast, rugged and sensitive ultra high pressure liquid chromatography tandem mass spectrometry method for analysis of cyanotoxins in raw water and drinking water--First findings of anatoxins, cylindrospermopsins and microcystin variants in Swedish source waters and infiltration ponds. J Chromatogr A 2016, 1429, 265–276. [Google Scholar] [CrossRef]
- Kahru, M.; Elmgren, R. Multidecadal time series of satellite-detected accumulations of cyanobacteria in the Baltic Sea. Biogeosciences 2014, 11, 3619–3633. [Google Scholar] [CrossRef]
- Mantzouki, E.; Lürling, M.; Fastner, J.; de Senerpont Domis, L.; Wilk-Woźniak, E.; Koreivienė, J.; Seelen, L.; Teurlincx, S.; Verstijnen, Y.; Krztoń, W.; et al. Temperature Effects Explain Continental Scale Distribution of Cyanobacterial Toxins. Toxins (Basel) 2018, 10. [Google Scholar] [CrossRef]
- Moreira, C.; Vasconcelos, V.; Antunes, A. Cyanobacterial Blooms: Current Knowledge and New Perspectives. Earth 2022, 3, 127–135. [Google Scholar] [CrossRef]
- Rigosi, A.; Carey, C.C.; Ibelings, B.W.; Brookes, J. The interaction between climate warming and eutrophication to promote cyanobacteria is dependent on trophic state and varies among taxa. Limnology and Oceanography 2014, 59. [Google Scholar] [CrossRef]
- Codd, G.A.; Morrison, L.F.; Metcalf, J.S. Cyanobacterial toxins: risk management for health protection. Toxicol Appl Pharmacol 2005, 203, 264–272. [Google Scholar] [CrossRef]
- Bashir, F.; Bashir, A.; Bouaïcha, N.; Chen, L.; Codd, G.A.; Neilan, B.; Xu, W.L.; Ziko, L.; Rajput, V.D.; Minkina, T.; et al. Cyanotoxins, biosynthetic gene clusters, and factors modulating cyanotoxin biosynthesis. World J Microbiol Biotechnol 2023, 39, 241. [Google Scholar] [CrossRef]
- Moreira, C.; Gomes, C.; Vasconcelos, V.; Antunes, A. Cyanotoxins Occurrence in Portugal: A New Report on Their Recent Multiplication. Toxins 2020, 12, 154. [Google Scholar] [CrossRef]
- Bloch, R.A.; Faulkner, G.; Hilborn, E.D.; Wismer, T.; Martin, N.; Rhea, S. Geographic Variability, Seasonality, and Increase in ASPCA Animal Poison Control Center Harmful Blue-Green Algae Calls-United States and Canada, 2010-2022. Toxins (Basel) 2023, 15. [Google Scholar] [CrossRef]
- Turner, A.D.; Turner, F.R.I.; White, M.; Hartnell, D.; Crompton, C.G.; Bates, N.; Egginton, J.; Branscombe, L.; Lewis, A.M.; Maskrey, B.H. Confirmation Using Triple Quadrupole and High-Resolution Mass Spectrometry of a Fatal Canine Neurotoxicosis following Exposure to Anatoxins at an Inland Reservoir. Toxins (Basel) 2022, 14. [Google Scholar] [CrossRef] [PubMed]
- Menezes, C.; Nova, R.; Vale, M.; Azevedo, J.; Vasconcelos, V.; Pinto, C. First description of an outbreak of cattle intoxication by cyanobacteria (blue-green algae) in the South of Portugal. Bov. Pract. 2019, 53, 66–70. [Google Scholar] [CrossRef]
- Wood, R. Acute animal and human poisonings from cyanotoxin exposure - A review of the literature. Environ Int 2016, 91, 276–282. [Google Scholar] [CrossRef] [PubMed]
- Lad, A.; Breidenbach, J.D.; Su, R.C.; Murray, J.; Kuang, R.; Mascarenhas, A.; Najjar, J.; Patel, S.; Hegde, P.; Youssef, M.; et al. As We Drink and Breathe: Adverse Health Effects of Microcystins and Other Harmful Algal Bloom Toxins in the Liver, Gut, Lungs and Beyond. Life (Basel) 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, B.Y.; Zhu, X.; Nagata, M.; Loo, L.; Chan, O.; Wong, L.L. Cyanotoxin exposure and hepatocellular carcinoma. Toxicology 2023, 487, 153470. [Google Scholar] [CrossRef]
- Niture, S.; Gadi, S.; Qi, Q.; Rios-Colon, L.; Khatiwada, S.; Vandana; Fernando, R.A.; Levine, K.E.; Kumar, D. Cyanotoxins Increase Cytotoxicity and Promote Nonalcoholic Fatty Liver Disease Progression by Enhancing Cell Steatosis. Toxins (Basel) 2023, 15, 411. [Google Scholar] [CrossRef] [PubMed]
- Metcalf, J.S.; Tischbein, M.; Cox, P.A.; Stommel, E.W. Cyanotoxins and the Nervous System. Toxins 2021, 13, 660. [Google Scholar] [CrossRef] [PubMed]
- Aráoz, R.; Molgó, J.; Tandeau de Marsac, N. Neurotoxic cyanobacterial toxins. Toxicon 2010, 56, 813–828. [Google Scholar] [CrossRef] [PubMed]
- Nugumanova, G.; Ponomarev, E.D.; Askarova, S.; Fasler-Kan, E.; Barteneva, N.S. Freshwater Cyanobacterial Toxins, Cyanopeptides and Neurodegenerative Diseases. Toxins 2023, 15, 233. [Google Scholar] [CrossRef] [PubMed]
- Sanseverino, I.; Conduto António, D.; Loos, R.; Lettieri, T. Cyanotoxins: methods and approaches for their analysis and detection. Joint Research Center (European Commission) Technical Reports, EUR 28624 2017. [Google Scholar] [CrossRef]
- Lund, J.W.G.; Kipling, C.; Le Cren, E.D. The inverted microscope method of estimating algal numbers and the statistical basis of estimations by counting. Hydrobiologia 1958, 11, 143–170. [Google Scholar] [CrossRef]
- Parulekar, N.N.; Kolekar, P.; Jenkins, A.; Kleiven, S.; Utkilen, H.; Johansen, A.; Sawant, S.; Kulkarni-Kale, U.; Kale, M.; Sæbø, M. Characterization of bacterial community associated with phytoplankton bloom in a eutrophic lake in South Norway using 16S rRNA gene amplicon sequence analysis. PLoS ONE 2017, 12, e0173408. [Google Scholar] [CrossRef]
- Gobler, C.J.; Jankowiak, J.G. Dynamic Responses of Endosymbiotic Microbial Communities Within Microcystis Colonies in North American Lakes to Altered Nitrogen, Phosphorus, and Temperature Levels. Front Microbiol 2021, 12, 781500. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Lee, C.S.; Cho, J.Y.; Bae, M.J.; Kim, E.J. Comparison of Bacterial Assemblages Associated with Harmful Cyanobacteria under Different Light Conditions. Microorganisms 2022, 10. [Google Scholar] [CrossRef]
- Metcalf, J.S.; Bell, S.G.; Codd, G.A. Production of novel polyclonal antibodies against the cyanobacterial toxin microcystin-LR and their application for the detection and quantification of microcystins and nodularin. Water Research 2000, 34, 2761–2769. [Google Scholar] [CrossRef]
- Zeck, A.; Eikenberg, A.; Weller, M.G.; Niessner, R. Highly sensitive immunoassay based on a monoclonal antibody specific for [4-arginine]microcystins. Analytica Chimica Acta 2001, 441, 1–13. [Google Scholar] [CrossRef]
- Metcalf, J.S.; Beattie, K.A.; Pflugmacher, S.; Codd, G.A. Immuno-crossreactivity and toxicity assessment of conjugation products of the cyanobacterial toxin, microcystin-LR. FEMS Microbiol Lett 2000, 189, 155–158. [Google Scholar] [CrossRef]
- Dittmann, E.; Fewer, D.P.; Neilan, B.A. Cyanobacterial toxins: biosynthetic routes and evolutionary roots. FEMS Microbiology Reviews 2013, 37, 23–43. [Google Scholar] [CrossRef]
- Aparicio-Muriana, M.D.M.; Lara, F.J.; Olmo-Iruela, M.D.; García-Campaña, A.M. Determination of Multiclass Cyanotoxins in Blue-Green Algae (BGA) Dietary Supplements Using Hydrophilic Interaction Liquid Chromatography-Tandem Mass Spectrometry. Toxins (Basel) 2023, 15. [Google Scholar] [CrossRef] [PubMed]
- Klijnstra, M.D.; Faassen, E.J.; Gerssen, A. A Generic LC-HRMS Screening Method for Marine and Freshwater Phycotoxins in Fish, Shellfish, Water, and Supplements. Toxins (Basel) 2021, 13. [Google Scholar] [CrossRef]
- Jacinavicius, F.R.; Valverde Campos, T.G.; Passos, L.S.; Pinto, E.; Geraldes, V. A rapid LC-MS/MS method for multi-class identification and quantification of cyanotoxins. Toxicon 2023, 234, 107282. [Google Scholar] [CrossRef]
- Sundaravadivelu, D.; Sanan, T.T.; Venkatapathy, R.; Mash, H.; Tettenhorst, D.; L, D.A.; Frey, S.; Tatters, A.O.; Lazorchak, J. Determination of Cyanotoxins and Prymnesins in Water, Fish Tissue, and Other Matrices: A Review. Toxins (Basel) 2022, 14. [Google Scholar] [CrossRef] [PubMed]
- Haddad, S.P.; Bobbitt, J.M.; Taylor, R.B.; Lovin, L.M.; Conkle, J.L.; Chambliss, C.K.; Brooks, B.W. Determination of microcystins, nodularin, anatoxin-a, cylindrospermopsin, and saxitoxin in water and fish tissue using isotope dilution liquid chromatography tandem mass spectrometry. J Chromatogr A 2019, 1599, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Turner, A.D.; Waack, J.; Lewis, A.; Edwards, C.; Lawton, L. Development and single-laboratory validation of a UHPLC-MS/MS method for quantitation of microcystins and nodularin in natural water, cyanobacteria, shellfish and algal supplement tablet powders. J Chromatogr B Analyt Technol Biomed Life Sci 2018, 1074-1075, 111–123. [Google Scholar] [CrossRef]
- España Amórtegui, J.C.; Pekar, H.; Retrato, M.D.C.; Persson, M.; Karlson, B.; Bergquist, J.; Zuberovic-Muratovic, A. LC-MS/MS Analysis of Cyanotoxins in Bivalve Mollusks-Method Development, Validation and First Evidence of Occurrence of Nodularin in Mussels (Mytilus edulis) and Oysters (Magallana gigas) from the West Coast of Sweden. Toxins (Basel) 2023, 15. [Google Scholar] [CrossRef] [PubMed]
- Van Hassel, W.H.R.; Masquelier, J.; Andjelkovic, M.; Rajkovic, A. Towards a Better Quantification of Cyanotoxins in Fruits and Vegetables: Validation and Application of an UHPLC-MS/MS-Based Method on Belgian Products. Separations 2022, 9, 319. [Google Scholar] [CrossRef]
- Van Hassel, W.H.R.; Ahn, A.-C.; Huybrechts, B.; Masquelier, J.; Wilmotte, A.; Andjelkovic, M. LC-MS/MS Validation and Quantification of Cyanotoxins in Algal Food Supplements from the Belgium Market and Their Molecular Origins. Toxins 2022, 14, 513. [Google Scholar] [CrossRef] [PubMed]
- Boundy, M.J.; Selwood, A.I.; Harwood, D.T.; McNabb, P.S.; Turner, A.D. Development of a sensitive and selective liquid chromatography-mass spectrometry method for high throughput analysis of paralytic shellfish toxins using graphitised carbon solid phase extraction. J Chromatogr A 2015, 1387, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Martijn, J.; Lind, A.E.; Schön, M.E.; Spiertz, I.; Juzokaite, L.; Bunikis, I.; Pettersson, O.V.; Ettema, T.J.G. Confident phylogenetic identification of uncultured prokaryotes through long read amplicon sequencing of the 16S-ITS-23S rRNA operon. Environ Microbiol 2019, 21, 2485–2498. [Google Scholar] [CrossRef] [PubMed]
- Olrik, K.; Blomqvist, P.; Brettum, P.; Cronberg, G.a.E.P. Methods for quantitative assessment of phytoplankton in freshwaters. part I: Sampling, processing, and application in freshwater environmental monitoring programmes. Rapport No. 4860, Naturvardsverket (Sweden) 1998.
- Edler, L.; Willén, E.; Willén, T.; Ahlgren, G. Skadliga alger i sjöar och hav. Rapport No. 4447, Naturvårdsverket (Sweden) 1995.
- Wang, H.; Zhang, Z.; Liang, D.; du, H.; Pang, Y.; Hu, K.; Wang, J. Separation of wind's influence on harmful cyanobacterial blooms. Water Res 2016, 98, 280–292. [Google Scholar] [CrossRef]
- Vander Woude, A.; Ruberg, S.; Johengen, T.; Miller, R.; Stuart, D. Spatial and temporal scales of variability of cyanobacteria harmful algal blooms from NOAA GLERL airborne hyperspectral imagery. Journal of Great Lakes Research 2019, 45, 536–546. [Google Scholar] [CrossRef]
- Hunter, P.D.; Tyler, A.N.; Willby, N.J.; Gilvear, D.J. The spatial dynamics of vertical migration by Microcystis aeruginosa in a eutrophic shallow lake: A case study using high spatial resolution time-series airborne remote sensing. Limnology and Oceanography 2008, 53, 2391–2406. [Google Scholar] [CrossRef]
- Christoffersen, K.; Lyck, S.; Winding, A. Microbial activity and bacterial community structure during degradation of microcystins. Aquatic Microbial Ecology 2002, 27, 125–136. [Google Scholar] [CrossRef]
- Holst, T.; Jørgensen, N.O.G.; Jørgensen, C.; Johansen, A. Degradation of microcystin in sediments at oxic and anoxic, denitrifying conditions. Water Research 2003, 37, 4748–4760. [Google Scholar] [CrossRef] [PubMed]
- Boopathi, T.; Ki, J.S. Impact of environmental factors on the regulation of cyanotoxin production. Toxins (Basel) 2014, 6, 1951–1978. [Google Scholar] [CrossRef]
- Lee, J.; Lee, S.; Jiang, X. Cyanobacterial Toxins in Freshwater and Food: Important Sources of Exposure to Humans. Annu Rev Food Sci Technol 2017, 8, 281–304. [Google Scholar] [CrossRef]
- Kaplan, A.; Harel, M.; Kaplan-Levy, R.N.; Hadas, O.; Sukenik, A.; Dittmann, E. The languages spoken in the water body (or the biological role of cyanobacterial toxins). Front Microbiol 2012, 3, 138. [Google Scholar] [CrossRef]
- Jankowiak, J.G.; Gobler, C.J. The Composition and Function of Microbiomes Within Microcystis Colonies Are Significantly Different Than Native Bacterial Assemblages in Two North American Lakes. Front Microbiol 2020, 11, 1016. [Google Scholar] [CrossRef]
- Wu, D.; Xu, Z.; Min, S.; Wang, J.; Min, J. Characteristics of microbial community structure and influencing factors of Yangcheng Lake and rivers entering Yangcheng Lake during the wet season. Environ Sci Pollut Res Int 2024, 31, 9565–9581. [Google Scholar] [CrossRef] [PubMed]
- Consortium, C.o.H.M.P. A framework for human microbiome research. Nature 2012, 486, 215–221. [Google Scholar] [CrossRef]
- Acinas, S.G.; Sarma-Rupavtarm, R.; Klepac-Ceraj, V.; Polz, M.F. PCR-induced sequence artifacts and bias: insights from comparison of two 16S rRNA clone libraries constructed from the same sample. Appl Environ Microbiol 2005, 71, 8966–8969. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.Journal 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Research 2012, 41, D590–D596. [Google Scholar] [CrossRef]
- Lanzén, A.; Jørgensen, S.L.; Huson, D.H.; Gorfer, M.; Grindhaug, S.H.; Jonassen, I.; Øvreås, L.; Urich, T. CREST--classification resources for environmental sequence tags. PLoS ONE 2012, 7, e49334. [Google Scholar] [CrossRef]
- Handbok Dricksvattenrisker - Cyanotoxiner i driksvatten. Livsmedelsverket (Sweden), 2018, ISSN 1104-7089.
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).