Submitted:
02 January 2024
Posted:
03 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. Results
3.1. Sequence Identification and Evolution Analysis of MoMyb13
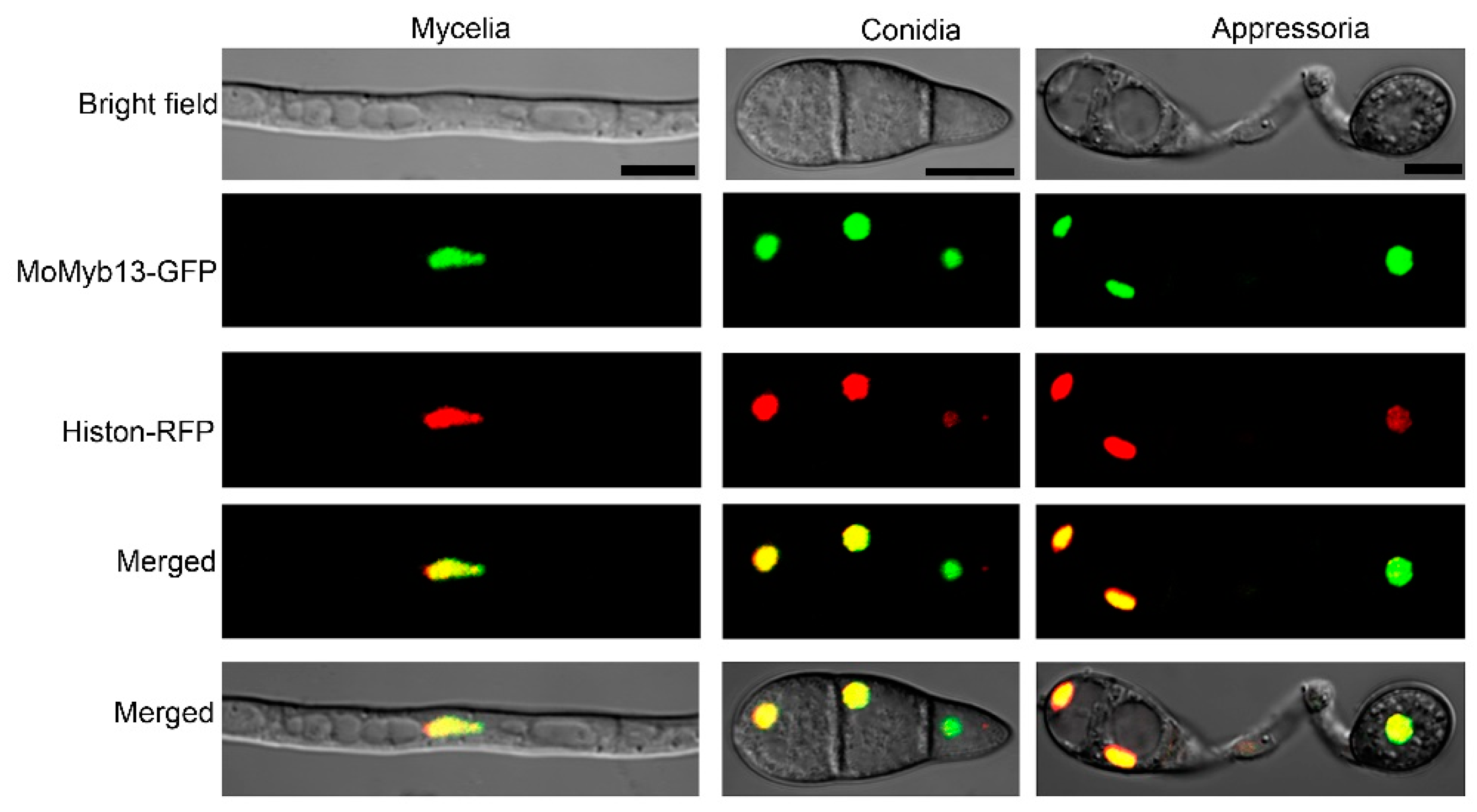
3.2. MoMyb13 Is Located in the Nuclei of M. oryzae
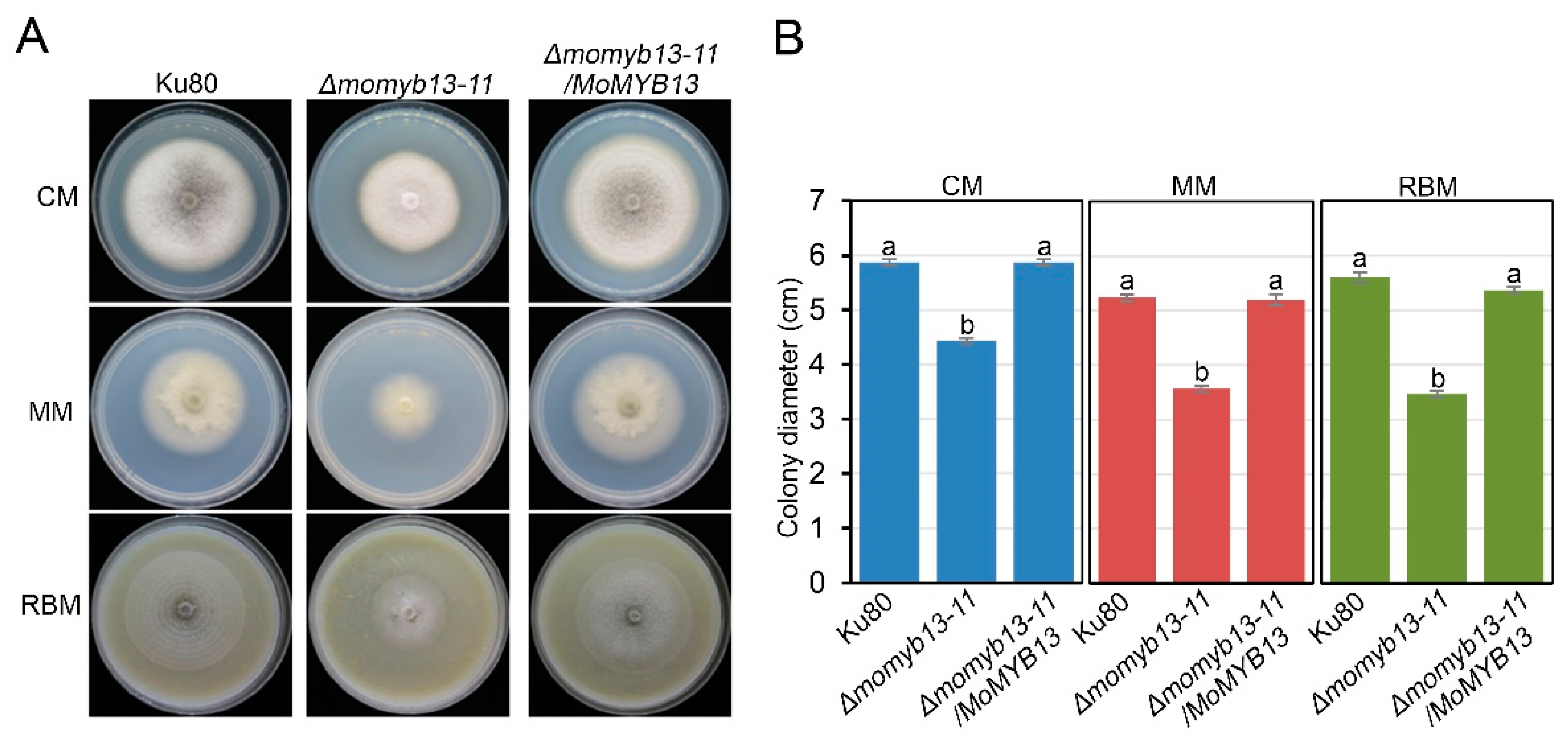
3.3. MoMyb13 Is Required for the Growth of M. oryzae
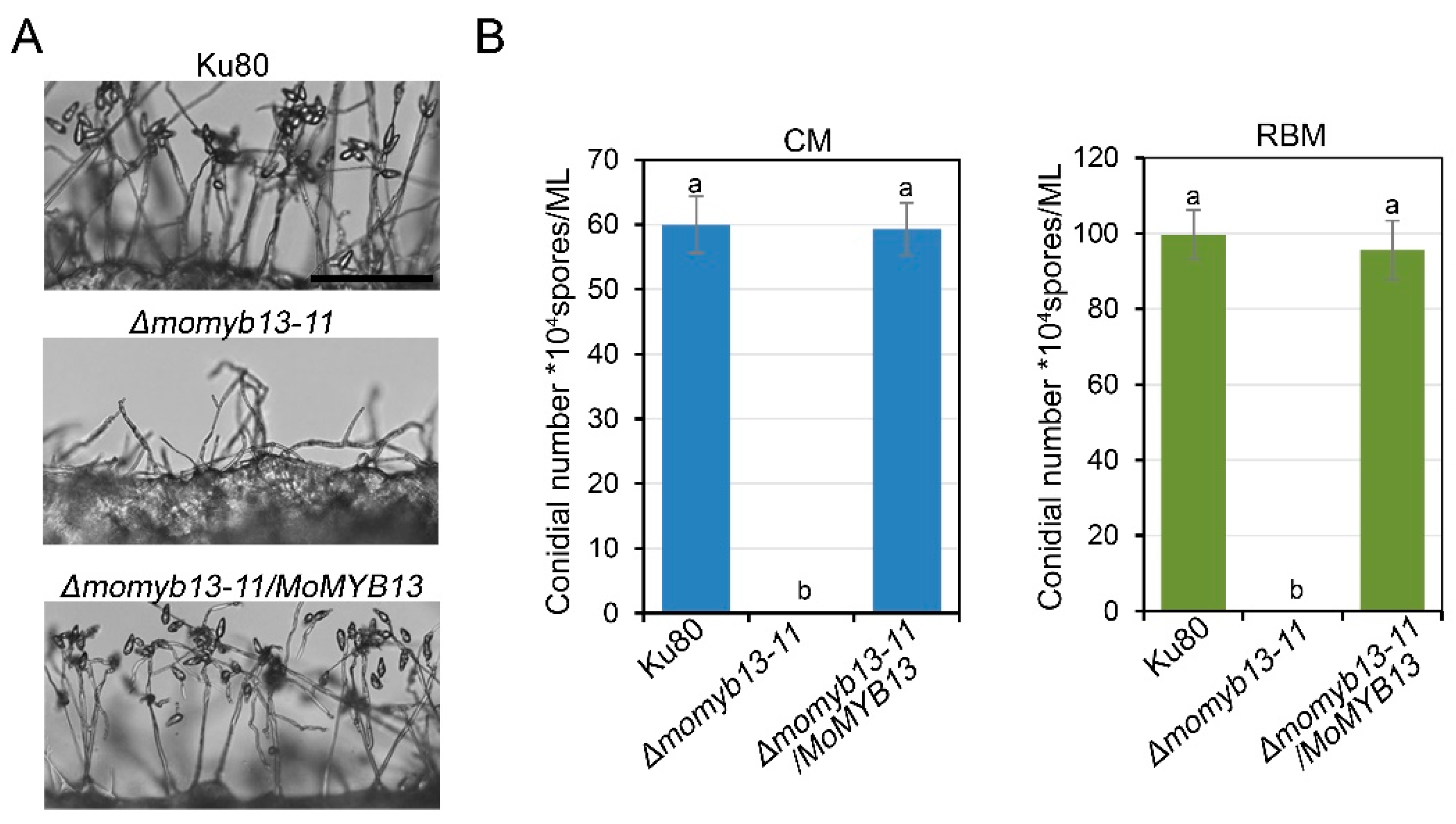
3.4. MoMyb13 Is Essential to the Conidiation of M. oryzae
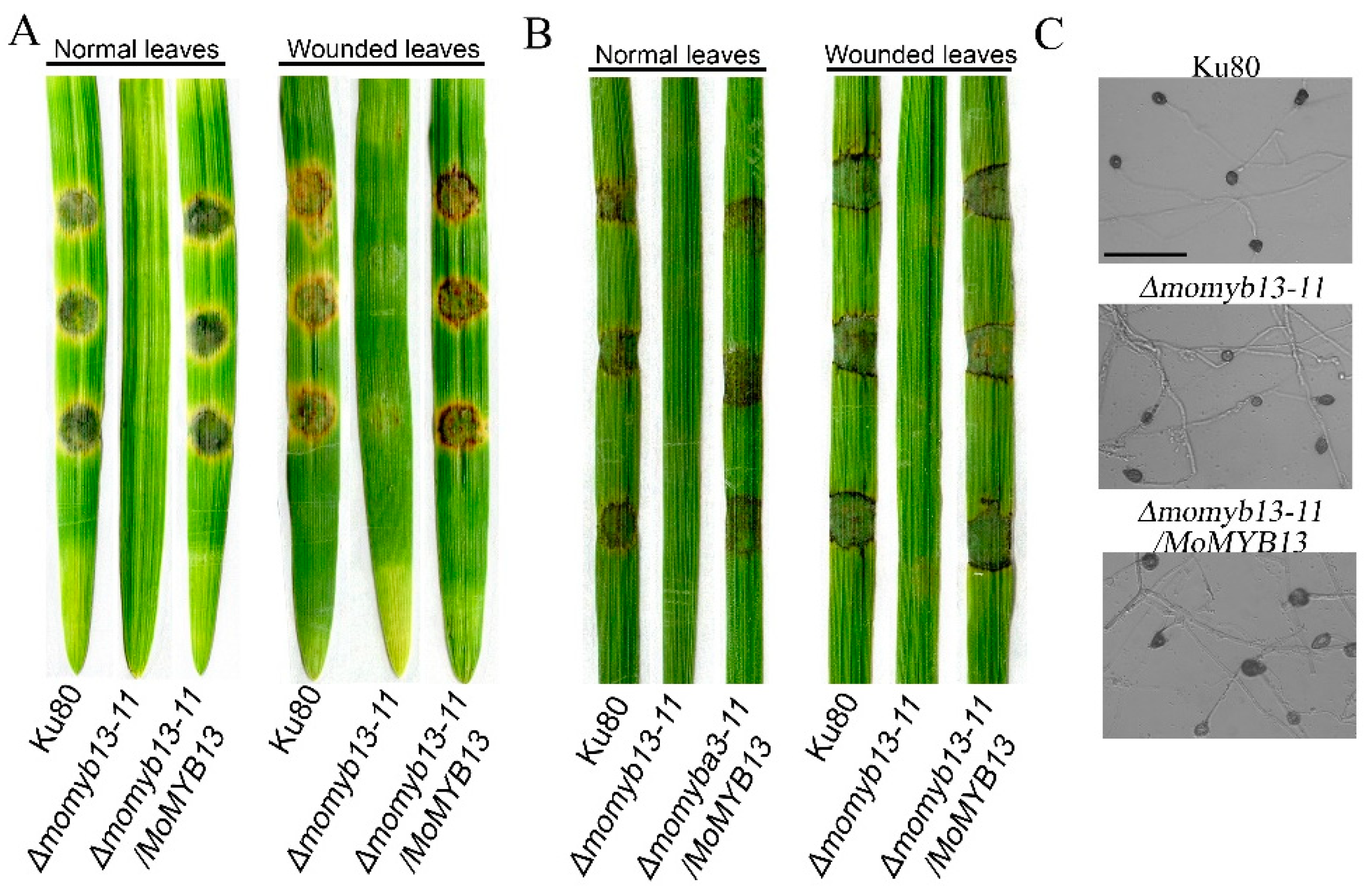
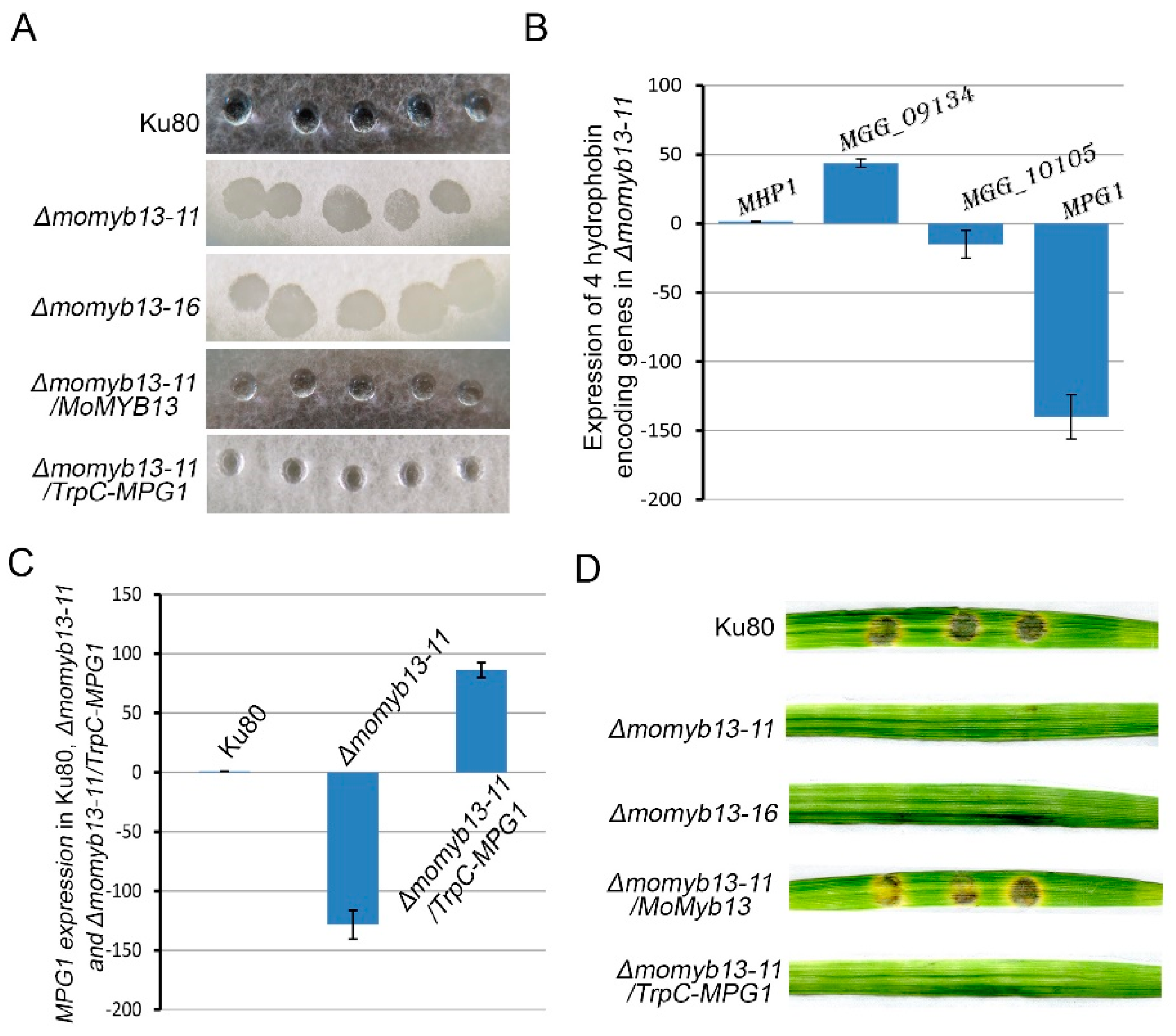
3.5. MoMyb13 Is Essential to the Pathogenicity of M. oryzae
3.6. MoMyb13 Is Involved in Regulating Hydrophobins
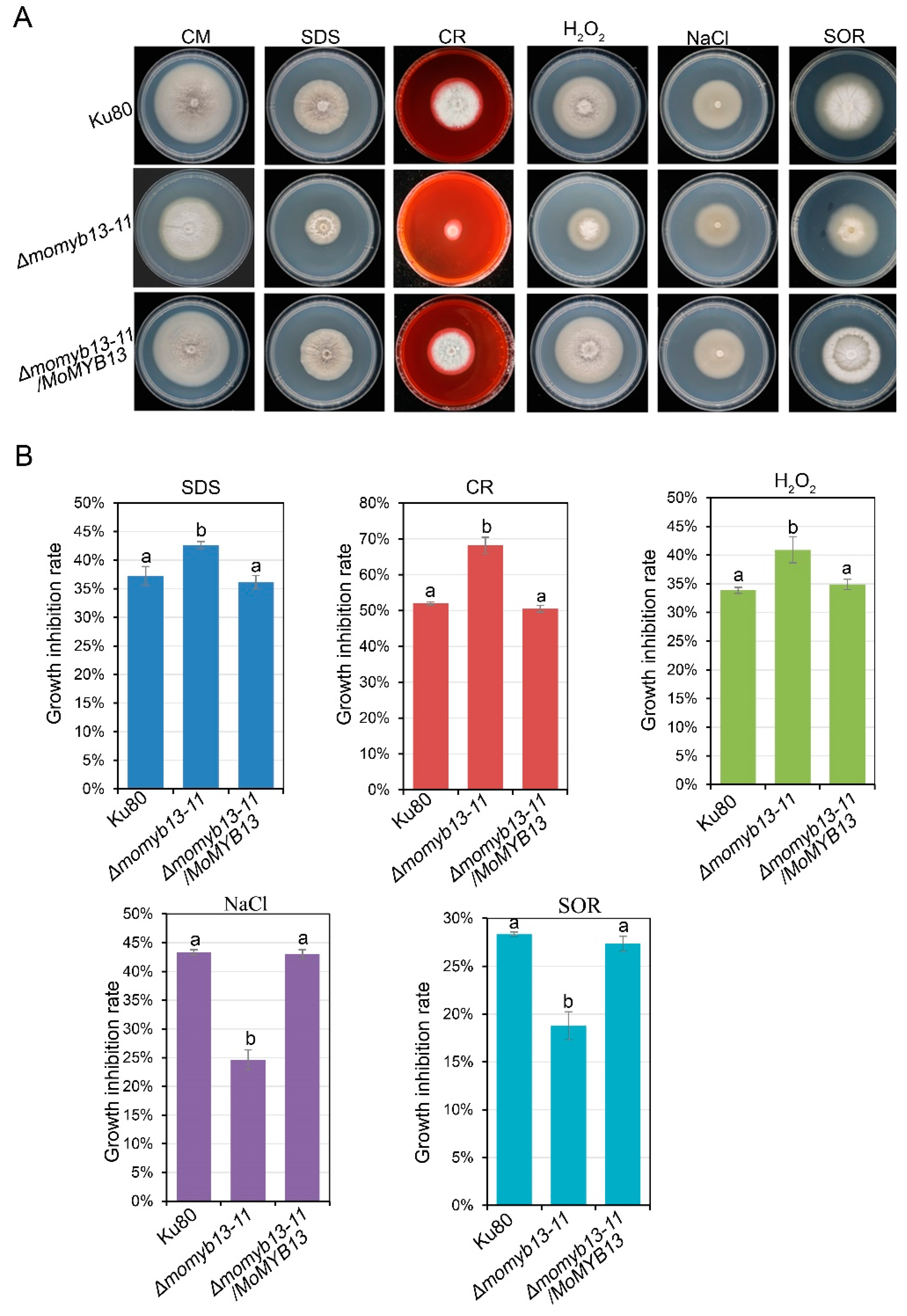
3.7. MoMyb13 Is Involved in Stress Response
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pennisi, E. Armed and Dangerous. Science 2010, 327, 804–805. [Google Scholar] [PubMed]
- Howard, R.J.; Ferrari, M.A.; Roach, D.H.; Money, N.P. Penetration of hard Substrates by a Fungus employing Enormous Turgor Pressures. Proceedings of the National Academy of Sciences 1991, 88, 11281–11284. [Google Scholar] [CrossRef] [PubMed]
- Kankanala, P.; Czymmek, K.; Valent, B. Roles for Rice Membrane Dynamics and Plasmodesmata during Biotrophic Invasion by the Blast Fungus. The Plant Cell 2007, 19, 706–724. [Google Scholar] [CrossRef] [PubMed]
- Sakulkoo, W.; Osés-Ruiz, M.; Oliveira Garcia, E.; Soanes, D.M.; Littlejohn, G.R.; Hacker, C.; et al. A Single Fungal MAP Kinase controls Plant Cell-to-cell invasion by The Rice Blast Fungus. Science 2018, 359, 1399–1403. [Google Scholar] [CrossRef] [PubMed]
- Gladieux, P.; Condon, B.; Ravel, S.; Soanes, D.; Maciel, J.L.N.; Nhani, A.; et al. Gene Flow between Divergent Cereal- and Grass-Specific Lineages of the Rice Blast Fungus Magnaporthe oryzae. mBio 2018, 9, e01219-17. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.T.; Croll, D.; Gladieux, P.; Soanes, D.M.; Persoons, A.; Bhattacharjee, P.; et al. Emergence of Wheat Blast in Bangladesh was caused by a South American Lineage of Magnaporthe oryzae. BMC Biol 2016, 14, 84. [Google Scholar] [CrossRef] [PubMed]
- Malaker, P.K.; Barma, N.C.D.; Tiwari, T.P.; Collis, W.J.; Duveiller, E.; Singh, P.K.; et al. First Report of Wheat Blast Caused by Magnaporthe oryzae Pathotype triticum in Bangladesh. Plant Disease 2016, 100, 2330–2330. [Google Scholar] [CrossRef]
- Dean, R.A.; Talbot, N.J.; Ebbole, D.J.; Farman, M.L.; Mitchell, T.K.; Orbach, M.J.; et al. The Genome Sequence of the Rice Blast Fungus Magnaporthe grisea. Nature 2005, 434, 980–986. [Google Scholar] [CrossRef]
- Ebbole, D.J. Magnaporthe as a Model for Understanding Host-Pathogen Interactions. Annu Rev Phytopathol 2007, 45, 437–456. [Google Scholar] [CrossRef]
- Huang, P.Y.; Wang, J.; Li, Y.; Wang, Q.; Huang, Z.C.; Qian, H.; Liu, X.H.; Lin, F.C.; Lu, J.P. Transcription factors Vrf1 and Hox7 coordinately regulate appressorium maturation in the rice blast fungus Magnaporthe oryzae. Microbiol Res 2022, 263, 127141. [Google Scholar] [CrossRef]
- Verma, S.; Gazara, R.K.; Verma, P.K. Transcription Factor Repertoire of Necrotrophic Fungal Phytopathogen Ascochyta rabiei: Predominance of MYB Transcription Factors As Potential Regulators of Secretome. Front Plant Sci 2017, 8, 1037. [Google Scholar] [CrossRef] [PubMed]
- Du, H.; Zhang, L.; Liu, L.; Tang, X.F.; Yang, W.J.; Wu, Y.M.; et al. Biochemical and Molecular Characterization of Plant MYB Transcription Factor Family. Biochemistry Moscow 2009, 74, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Prouse, M.B. and Campbell, M.M. The Interaction Between MYB Proteins and their Target DNA binding Sites. Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms 2012, 1819, 67–77. [Google Scholar] [CrossRef] [PubMed]
- Roy, S. Function of MYB Domain Transcription Factors in Abiotic Stress and Epigenetic Control of Stress Response in Plant Genome. Plant Signaling & Behavior 2016, 11, e1117723. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Gao, W.; Wu, X.; Zhao, M.; Qu, J.; Huang, C.; Zhang, J. Genome-Wide Characterization and Expression Analyses of Pleurotus ostreatus MYB Transcription Factors during Developmental Stages and under Heat Stress Based on de novo Sequenced Genome. International journal of molecular sciences 2018, 19, 2052. [Google Scholar] [CrossRef] [PubMed]
- Ambawat, S.; Sharma, P.; Yadav, N.R.; Yadav, R.C. MYB transcription factor genes as regulators for plant responses: an overview. Physiology and molecular biology of plants 2013, 19, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Eme, L.; Spang, A.; Lombard, J.; Stairs, C.W.; Ettema, T.J.G. Archaea and the Origin of Eukaryotes. Nature reviews. Microbiology 2017, 15, 711–723. [Google Scholar] [CrossRef] [PubMed]
- Allan, A.C. and Espley, R.V. MYBs Drive Novel Consumer Traits in Fruits and Vegetables. Trends in Plant Science 2018, 23, 693–705. [Google Scholar] [CrossRef] [PubMed]
- Baldoni, E.; Genga, A.; Cominelli, E. Plant MYB Transcription Factors: Their Role in Drought Response Mechanisms. International journal of molecular sciences 2015, 16, 15811–15851. [Google Scholar] [CrossRef]
- Cao, Y.; Li, K.; Li, Y.; Zhao, X.; Wang, L. MYB Transcription Factors as Regulators of Secondary Metabolism in Plants. Biology 2020, 9, 61. [Google Scholar] [CrossRef]
- Dubos, C.; Stracke, R.; Grotewold, E.; Weisshaar, B.; Martin, C.; Lepiniec, L. MYB Transcription Factors in Arabidopsis. Trends in Plant Science 2010, 15, 573–581. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Han, G.; Sun, C.; Sui, N. Research Advances of MYB Transcription Factors in Plant Stress Resistance and Breeding. Plant Signaling & Behavior 2019, 14, 1613131. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Osbourn, A.; Ma, P. MYB Transcription Factors as Regulators of Phenylpropanoid Metabolism in Plants. Molecular Plant 2015, 8, 689–708. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Constabel, C.P. Repressors as Regulators of Phenylpropanoid Metabolism in Plants. Trends in Plant Science 2019, 24, 275–289. [Google Scholar] [CrossRef] [PubMed]
- Yu, E.Y.; Yen, W.F.; Steinberg-Neifach, O.; Lue, N.F. Rap1 in Candida albicans: an Unusual Structural Organization and a Critical Function in Suppressing Telomere Recombination. Mol Cell Biol 2010, 30, 1254–1268. [Google Scholar] [CrossRef]
- Hogues, H.; Lavoie, H.; Sellam, A.; Mangos, M.; Roemer, T.; Purisima, E.; et al. Transcription Factor Substitution during the Evolution of Fungal Ribosome Regulation. Molecular Cell 2008, 29, 552–562. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Sánchez, L.; Rodríguez-López, M.; García, Z.; Tenorio-Gómez, M.; Schvartzman, J.B.; Krimer, D.B.; Hernández, P. The fission yeast rDNA-binding protein Reb1 regulates G1 phase under nutritional stress. Journal of Cell Science 2011, 124, 25–34. [Google Scholar] [CrossRef]
- Mukherjee, K.; Pandey, D.M.; Vidyarthi, A.S. Molecular Dynamics Simulation of Rap1 Myb-type domain in Saccharomyces cerevisiae. Bioinformation 2012, 8, 881–885. [Google Scholar] [CrossRef]
- Ramos-Sáenz, A.; González-Álvarez, D.; Rodríguez-Galán, O.; Rodríguez-Gil, A.; Gaspar, S.G.; Villalobo, E.; et al. Pol5 is an essential ribosome biogenesis factor required for 60S ribosomal subunit maturation in Saccharomyces cerevisiae. RNA 2019, 25, 1561–1575. [Google Scholar] [CrossRef]
- Matheis, S.; Yemelin, A.; Scheps, D.; Andresen, K.; Jacob, S.; Thines, E.; Foster, A.J. Functions of the Magnaporthe oryzae Flb3p and Flb4p transcription factors in the regulation of conidiation. Microbiological Research 2017, 196, 106–117. [Google Scholar] [CrossRef]
- Valsecchi, I.; Sarikaya-Bayram, Ö.; Wong Sak Hoi, J.; Muszkieta, L.; Gibbons, J.; Prevost, M.C.; et al. MybA, a transcription factor involved in conidiation and conidial viability of the human pathogen Aspergillus fumigatus: A transcription factor regulating conidiation in Aspergillus. Molecular Microbiology 2017, 105, 880–900. [Google Scholar] [CrossRef]
- Wieser, J.; Adams, T.H. flbD encodes a Myb-like DNA-binding protein that coordinates initiation of Aspergillus mdulans conidiophore development. Genes & Development 1995, 9, 491–502. [Google Scholar] [CrossRef]
- Mateos, L.; Jiménez, A.; Revuelta, J.L.; Santos, M.A. Purine Biosynthesis, Riboflavin Production, and Trophic-Phase Span Are Controlled by a Myb-Related Transcription Factor in the Fungus Ashbya gossypii. Appl Environ Microbiol 2006, 72, 5052–5060. [Google Scholar] [CrossRef]
- Mitra, P. Transcription Regulation of MYB: a Potential and Novel Therapeutic Target in Cancer. Ann Transl Med 2018, 6, 443–443. [Google Scholar] [CrossRef]
- Pattabiraman, D.R.; Gonda, T.J. Role and Potential for Therapeutic targeting of MYB in Leukemia. Leukemia 2013, 27, 269–277. [Google Scholar] [CrossRef]
- Kim, Y.; Kim, H.; Son, H.; Choi, G.J.; Kim, J.C.; Lee, Y.W. MYT3, A Myb-Like Transcription Factor, Affects Fungal Development and Pathogenicity of Fusarium graminearum. PloS one 2014, 9, e94359. [Google Scholar] [CrossRef]
- Lin, Y.; Son, H.; Lee, J.; Min, K.; Choi, G.J.; Kim, J.C.; Lee, Y.W. A Putative Transcription Factor MYT1 Is Required for Female Fertility in the Ascomycete Gibberella zeae. PloS one 2011, 6, e25586. [Google Scholar] [CrossRef]
- Lin, Y.; Son, H.; Min, K.; Lee, J.; Choi, G.J.; Kim, J.C.; Lee, Y.W. A Putative Transcription Factor MYT2 Regulates Perithecium Size in the Ascomycete Gibberella zeae. PloS one 2012, 7, e37859. [Google Scholar] [CrossRef]
- Dong, Y.; Zhao, Q.; Liu, X.; Zhang, X.; Qi, Z.; Zhang, H.; et al. MoMyb1 is Required for Asexual Development and Tissue-specific Infection in the Rice Blast Fungus Magnaporthe oryzae. BMC Microbiol 2015, 15, 37. [Google Scholar] [CrossRef]
- Lee, S.; Völz, R.; Song, H.; Harris, W.; Lee, Y.H. Characterization of the MYB Genes Reveals Insights Into Their Evolutionary Conservation, Structural Diversity, and Functional Roles in Magnaporthe oryzae. Front Microbiol 2021, 12, 721530. [Google Scholar] [CrossRef] [PubMed]
- Villalba, F.; Collemare, J.; Landraud, P.; Lambou, K.; Brozek, V.; Cirer, B.; et al. Improved gene targeting in Magnaporthe grisea by inactivation of MgKU80 required for non-homologous end joining. Fungal Genetics and Biology 2008, 45, 68–75. [Google Scholar] [CrossRef]
- Li, Y.; Que, Y.; Liu, Y.; Yue, X.; Meng, X.; Zhang, Z.; Wang, Z. The Putative Gγ Subunit Gene MGG1 is required for Conidiation, Appressorium Formation, mating and Pathogenicity in Magnaporthe oryzae. Curr Genet 2015, 61, 641–651. [Google Scholar] [CrossRef]
- Li, Y.; Yue, X.; Que, Y.; Yan, X.; Ma, Z.; Talbot, N.J.; Wang, Z. Characterisation of Four LIM Protein-Encoding Genes Involved in Infection-Related Development and Pathogenicity by the Rice Blast Fungus Magnaporthe oryzae. PLoS ONE 2014, 9, e88246. [Google Scholar] [CrossRef]
- Li, Y.; Yan, X.; Wang, H.; Liang, S.; Ma, W.B.; Fang, M.Y.; et al. MoRic8 Is a Novel Component of G-Protein Signaling During Plant Infection by the Rice Blast Fungus Magnaporthe oryzae. Molecular plant-microbe interactions 2010, 23, 317–331. [Google Scholar] [CrossRef]
- Li, Y.; Zheng, X.; Zhu, M.; Chen, M.; Zhang, S.; He, F.; Chen, X.; Lv, J.; Pei, M.; Zhang, Y.; Zhang, Y.; Wang, W.; Zhang, J.; Wang, M.; Wang, Z.; Li, G.; Lu, G. MoIVD-Mediated Leucine Catabolism Is Required for Vegetative Growth, Conidiation and Full Virulence of the Rice Blast Fungus Magnaporthe oryzae. Frontiers in microbiology 2019, 10, 444. [Google Scholar] [CrossRef]
- Cao, Y.; Chen, J.; Xie, X.; Liu, S.; Jiang, Y.; Pei, M.; Wu, Q.; Qi, P.; Du, L.; Peng, B.; Lan, J.; Wu, F.; Feng, K.; Zhang, Y.; Fang, Y.; Liu, M.; Jaber, M.Y.; Wang, Z.; Olsson, S.; Lu, G.; Li, Y. Characterization of two infection-induced transcription factors of Magnaporthe oryzae reveals their roles in regulating early infection and effector expression. Molecular plant pathology 2022, 23, 1200–1213. [Google Scholar] [CrossRef]
- Li, Y.; Liang, S.; Yan, X.; Wang, H.; Li, D.; Soanes, D.M.; et al. Characterization of MoLDB1 Required for Vegetative Growth, Infection-Related Morphogenesis, and Pathogenicity in the Rice Blast Fungus Magnaporthe oryzae. Molecular plant-microbe interactions 2010, 23, 1260–1274. [Google Scholar] [CrossRef]
- Livak, K.J. and Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Berger, B.W. and Sallada, N.D. Hydrophobins: Multifunctional Biosurfactants for Interface Engineering. Journal of biological engineering 2019, 13, 10. [Google Scholar] [CrossRef]
- Bayry, J.; Aimanianda, V.; Guijarro, J.I.; Sunde, M.; Latgé, J.P. Hydrophobins—Unique Fungal Proteins. PLoS Pathog 2012, 8, e1002700. [Google Scholar] [CrossRef]
- Beckerman, J.L. MPG1, a Gene Encoding a Fungal Hydrophobin of Magnaporthe grisea, Is Involved in Surface Recognition. Molecular plant-microbe interactions 1996, 9, 450. [Google Scholar] [CrossRef] [PubMed]
- Kou, Y.; Qiu, J.; Tao, Z. Every Coin Has Two Sides: Reactive Oxygen Species during Rice—Magnaporthe oryzae Interaction. International journal of molecular sciences 2019, 20, 1191. [Google Scholar] [CrossRef] [PubMed]
- Collado, J.; Gonzalez, A.; Platas, G.; Stchigel, A.M.; Guarro, J.; Pelaez, F. Monosporascus ibericus sp. nov., an Endophytic Ascomycete from Plants on Saline Soils, with Observations on the Position of the Genus based on Sequence Analysis of the 18S rDNA. Mycological Research 2002, 106, 118–127. [Google Scholar] [CrossRef]
- Liu, K.; Ding, X.; Deng, B.; Chen, W. Isolation and Characterization of Endophytic Taxol-producing Fungi from Taxus chinensis. Journal of industrial microbiology & biotechnology 2009, 36, 1171–1177. [Google Scholar] [CrossRef]
- Al-Khawaldeh, M.M.; Araj, S.-E.; Alananbeh, K.M.; Antary, T.M.A. Wheat Cultivable Fungal Endophytes in Jordan. Fresenius Environmental Bulletin 2020, 29, 13. [Google Scholar]
- Geisen, S.; Hooven, F.C.; Kostenko, O.; Snoek, L.B.; Putten, W.H. Fungal Root Endophytes Influence Plants in a Species-specific Manner that depends on Plant’s Growth Stage. J Ecol 2021, 109, 1618–1632. [Google Scholar] [CrossRef]
- Sarsaiya, S.; Jain, A.; Fan, X.; Jia, Q.; Xu, Q.; Shu, F.; et al. New Insights into Detection of a Dendrobine Compound From a Novel Endophytic Trichoderma longibrachiatum Strain and Its Toxicity Against Phytopathogenic Bacteria. Frontiers in microbiology 2020, 11, 337. [Google Scholar] [CrossRef]
- Gao, H.; Pan, M.; Tian, C.; Fan, X. Cytospora and Diaporthe Species Associated with Hazelnut Canker and Dieback in Beijing, China. Frontiers in cellular and infection microbiology 2021, 11, 664366. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Mao, L.; Zhao, N.; Xia, C.; Liu, J.; Kubicek, C.P.; et al. Verification of TRI3 Acetylation of Trichodermol to Trichodermin in the Plant Endophyte Trichoderma taxi. Frontiers in microbiology 2021, 12, 731425. [Google Scholar] [CrossRef]
- Zhu, S.; Yan, Y.; Qu, Y.; Wang, J.; Feng, X.; Liu, X.; Lin, F.; Lu, J. Role refinement of melanin synthesis genes by gene knockout reveals their functional diversity in Pyricularia oryzae strains. Microbiological Research 2021, 242, 126620. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



