1. Introduction
The p53 tumor suppressor protein impacts on functioning of a cell by acting as an activator of at least hundreds of genes. The early-identified p53-regulated genes coded for proteins promoting apoptosis, cell cycle arrest and DNA repair. However, as many more genes directly activated by p53 have been discovered, it was realized that p53 regulates wide spectrum of biological functions including metabolism, aging, angiogenesis, immunity, and more [
1,
2,
3]. The p53 binds as a tetramer to its response element (RE) RRRCWWGYYYRRRCWWGYYY (R – A or G, Y – C or T, W – A or T) consisting of two decameric half sites divided further into four pentameric quarter sites (RRRCW or WGYYY). Thus, each p53 monomer binds to each quarter site of the p53 RE. This arrangement allows for some flexibility in the selection of p53 binding sites, which sometime can deviate from the consensus sequence [
4].
Recently, we published the transcriptomic data showing changes in gene expression in A549 lung cancer cell line exposed to actinomycin D and nutlin-3a (henceforth abbreviated to A+N) [
5]. Earlier, we noticed that these two substances synergize in activation of p53 and in activation of some p53-regulated genes [
6,
7]. The mechanism of synergy is not known. Actinomycin D inhibits RNA polymerase I, what induces nucleolar stress and release of some ribosomal proteins out of nucleolus. Some of these proteins bind to MDM2, which is the negative regulator of p53. MDM2, bound by ribosomal proteins, does not destabilize p53, which accumulates and activates its target genes [
8]. However, probably it is only a part of the mechanism, by which actinomycin D activates p53 because we noticed, that this drug also promotes p53 phosphorylation by an undefined kinase [
9]. Phosphorylation of p53 by various kinases on different amino acids is crucial for stimulation of p53 as a transcription regulator [
10]. MDM2 inhibits p53 by promoting its degradation but also by concealing its transcription-activating domain [
11,
12]. Nutlin-3a is a molecule, which binds to MDM2 and fills its p53 interaction pocket, what prevents p53-MDM2 interaction, what in turn results in p53 stabilization and activation of p53-target genes [
13]. We hypothesize, that when actinomycin D and nutlin-3a are applied together they synergize because actinomycin D activates p53-phosphorylating kinases and nutlin-3a prevents MDM2 from covering p53, what gives the kinases easy contact with p53 and promotes its strong phosphorylation. By employing this p53 activation mode (A+N) and a cell line (A549) never used before in high-throughput searches for p53-regulated genes, we were able to identify new candidate p53-target genes [
5].
One of the genes strongly activated by A+N is
DUSP13. We selected it for detailed study not only because of its activation by A+N (55-fold by RNA-Seq), but also because its expression, as suggested by the location of mapped sequencing reads, starts from an alternative promoter in intron [
5]. We hypothesized that p53 activated by A+N, induces the expression of
DUSP13 from this alternative promoter.
Initially
DUSP13 was cloned as a gene coding for a protein abundantly expressed in testis [
14]. The regulation of
DUSP13 is very complicated and unusual so we considered it interesting to learn how p53 contributes to this regulation. According to early studies
DUSP13 codes for two isoforms of proteins, which function as dual-specificity phosphatases removing phosphate groups from serines/threonines as well as from tyrosines.
DUSP13 codes these isoforms utilizing the alternative open reading frames (ORF) what is extremely rare in eukaryotes. Exons 1-3 of
DUSP13 code for muscle-restricted dual specificity phosphatase (MDSP), whereas exons 7-9 encode testis and skeletal muscle specific dual specificity phosphatase (TMDP). However, the GeneCards data base considers the DNA fragments coding for
DUSP13 isoforms as a separate genes -
DUSP13A (MDSP) and
DUSP13B (TMDP). On the other hand, the human genome assembly GRCh38p14 and IGV genome viewer display
DUSP13 as a single gene. This ambiguity is due to the complicated expression of this locus. For instance, the transcript variants 3 and 7, which encode
DUSP13B isoform named TMDP-L1 share the exons with
DUSP13A (
Figure 1A). The RT-PCR analysis of RNA from skeletal muscles revealed the existence of mRNAs able to code isoform TMDP-L1 and another one named TMDP-L2, however, the Western blotting analysis of proteins extracted from muscles did not reveal the presence of these proteins [
15]. These results generate several questions, e.g., are these mRNA forms translated, was the employed Western blotting method sensitive enough to detect their presence, and if expressed, what is their physiological function?
Our RNA-Seq data [
5] revealed that activation of p53 by A+N in lung cancer cells may induce the expression of a DUSP13 isoform related to TMDP. This is unexpected observation, because so far the expression of
DUSP13 was detected only in skeletal muscle and testis. Our results suggested that the p53 has the potential to activate expression of
DUSP13 in other tissues. Interestingly, TMDP was found to inactivate stress-activated kinases named MAPKs [
16] what suggests that p53 may modulate the stress response of a cell by activation of this DUSP13 isoform.
2. Materials and Methods
2.1. Cell Culture and Treatment
A549 (lung adenocarcinoma, American Type Culture Collection - ATCC, Manassas, VA, USA), NCI-H292 (mucoepidermoid pulmonary carcinoma, ATCC) and U-2 OS (osteosarcoma, ATCC) cells were grown as previously described [
6]. NCI-H460 (lung cancer, ATCC) were cultured in RPMI-1640 supplemented with 2 mM L-glutamine, 4.5 g/L glucose and 1 mM sodium pyruvate, and 10% heat-inactivated fetal bovine serum (FBS; Invitrogen, Carlsbad, CA, USA). NCI-H1299 large cell carcinoma of the lung cell line (ATCC) was cultured in low-glucose DMEM (with 10% FBS), A375 melanoma cell line (ATCC) was cultured in high-glucose DMEM (with 10% FBS), AGS gastric adenocarcinoma cell line (ATCC) was cultured in McCoy’s 5A medium (with 10% FBS), MCF7 breast adenocarcinoma cell line (ATCC) was cultured in DMEM F12 medium (with 10% FBS) and NCI-H23 lung adenocarcinoma cell line (ATCC) was cultured in RPMI-1640 (with 10% FBS). All culture media were supplemented with 1% penicillin–streptomycin (Sigma-Aldrich, St. Louis, MO, USA). The cells were grown at 37 °C/5% CO
2.
The stock solutions of chemicals were prepared in DMSO: actinomycin D (10 µM; Sigma-Aldrich, St. Louis, MO, USA), camptothecin (10 mM; Calbiochem-Merck, Darmstadt, Germany), nutlin-3a (10 mM; Selleck Chemicals LLC, Houston, TX, USA), idasanutlin (10 mM; MedChemExpress, Monmouth Junction, NJ, USA) and RG7112 (10 mM; MedChemExpress). Stock solutions were diluted in culture medium to the following concentrations: 5 nM actinomycin D, 5 µM nutlin-3a, 5 µM camptothecin, 5 µM idasanutlin and 5 µM RG7112. Control cells were mock-treated with medium containing DMSO.
The generation of p53-deficient A549 and U-2 OS cells using CRISPR/Cas9 technology was described earlier [
5].
2.2. Semi-Quantitative Real-Time PCR
Total RNA samples were isolated from cells using the RNeasy mini kit (Qiagen, Hilden, Germany). The cDNA was synthesized with MuLV reverse transcriptase and random hexamers (Applied Biosystems, Foster City, CA, USA). Measurements of mRNA levels were performed using Real-Time 2x PCR Master Mix SYBR (A&A Biotechnology, Gdynia, Poland). We used the following primers to measure the expression of DUSP13 by RT-PCR: 5’-GAT ACA TCC GAG CTG CCC TC-3’ and 5’-GCC TCT ACC AGC GTC ATG TT-3’. The primers for internal control ACTB were as follows: 5’-GCA AGC AGG AGT ATG ACG AG-3’ and 5’-CAA ATA AAG CCA TGC CAA TC-3’. Amplifications were performed on a CFX96 Real-Time System (Bio-Rad, Hercules, CA, USA). In each PCR run, cDNA samples were amplified in triplicate. Relative quantitation of mRNA was carried out using the ΔΔCT method with ACTB as a reference. Mean and standard deviation were calculated from three or four biological replicates.
2.3. Western Blotting
The preparation of the whole-cell lysates using IP buffer, supplemented with protease and phosphatase inhibitors as well as preparation of the concentrated conditioned medium were described previously [
7]. Aliquots of lysates (35–50 µg) were separated by SDS-PAGE on 8% or 13% gels and electro-transferred onto PVDF membranes. Before incubation with primary antibody, the membranes were incubated for 1 h at room temperature in blocking solution (5% skim milk in PBS with 0.1% Tween-20). The anti-phospho-Ser37 p53 and anti-phospho-Ser392 p53 antibodies were from Cell Signaling Technology (Danvers, MA, USA). Anti-p53 (DO-1), and loading control anti-HSC70 (B-6) antibodies were obtained from Santa Cruz Biotechnology (Dallas, TX, USA). Anti-DUSP13 antibody (rabbit polyclonal) was from Proteintech (Rosemont, IL, USA). The HRP-conjugated anti-GFP (green fluorescent protein) antibody (B-2) was from Santa Cruz Biotechnology. All incubations with primary antibodies were performed overnight at 4 °C in blocking solution. HRP-conjugated secondary antibodies (anti-mouse, anti-rabbit) were detected by chemiluminescence (SuperSignal West Pico or SuperSignal West Femto chemiluminescent substrate, Thermo Fisher Scientific, Waltham, MA, USA).
2.4. Molecular Cloning, Site-Directed Mutagenesis and Luciferase Reporter Assay
The alternative promoter region of DUSP13 was cloned into the pGL3-Basic reporter vector, which encodes firefly luciferase (Promega, Madison, WI, USA). The sequences of primers used to amplify the promoter are: 5’-TTTT ACG CGT CCA CCT CTG CTT CCT CTA CA-3’ and 5’-TTTT CTC GAG CCA GCT CTG GAA GAG AGA TGA-3’. The primers were designed to contain the restriction sites for MluI and XhoI (underlined). Amplified DNA was ligated into the respective sites of pGL3-Basic plasmid. PCR was performed with PfuPlus! DNA polymerase mix (EURx, Gdańsk, Poland) to ensure high fidelity DNA amplification. The inserted DNA was sequenced to ensure that the clones contained no mutations.
The mutations of CWWG (W - A or T) sequence in the putative p53 response element from DUSP13 promoter were created using GeneArt Site-Directed Mutagenesis PLUS kit (Life Technologies, Carlsbad, CA, USA) using following oligonucleotides: DUSP13 forward (5'-GGTGACTGGCCTGGGGCGTCTTGGGAGCTGGAAC-3'), DUSP13 complementary reverse (5'- GTTCCAGCTCCCAAGACGCCCCAGGCCAGTCACC-3'), (the sites of mutation are underlined).
The luciferase reporter assay was performed as described recently [
7]. In short, U-2 OS cells were co-transfected using FuGENE 6 (Promega, Madison, WI, USA) with a combination of reporter vector, encoding firefly luciferase under the control of tested promoter (wild type or mutant), and expression vector pC53-SN3, encoding wild-type p53 or pC53-SCX3 encoding Val143Ala p53 mutant (a gift from Dr. Bert Vogelstein and Dr. Kenneth W. Kinzler from Johns Hopkins University, Baltimore, MD, USA [
17]). As a negative control, the p53 plasmid was replaced by empty vector. The transfection mixture also contained pRL-TK vector, encoding
Renilla sp. luciferase under the control of herpes simplex virus thymidine kinase (HSV-TK) promoter (internal control). The next day, the cells were washed with culture medium and incubated with fresh medium for an additional 24 hours. The cells were lysed with PLB buffer from the Dual Luciferase Reporter Assay system (Promega, Madison, WI, USA) and the activities of the luciferases were measured. Firefly luciferase activity was normalized against
Renilla sp. luciferase activity what produced normalized firefly luciferase activity (NFLA). The NFLA in control cells was set as 1 and the activity in experimental cells was expressed as its fold-change. Each transfection was performed in triplicate in three independent experiments.
To test how the endogenous p53 impacts on the activity of the cloned DUSP13 promoter, the cells were transfected with the DUSP13 reporter vector and the abovementioned pRL-TK control vector. After 24 hours, the medium with the transfection mixture (Fugene 6 + DNA) was removed and the cells were exposed either to control medium or the medium with A+N to activate the endogenous p53. After 24 hours the activities of both luciferases were measured and normalized firefly luciferase activity was calculated as described in the previous paragraph.
To generate DUSP13-EGFP chimeric cDNA, we first isolated RNA from A549 cells exposed to A+N and then we generated cDNA as described above. We employed a triple-ligation method to clone cDNA of the DUSP isoform taking advantage of the ClaI restriction site naturally occurring in the middle of its cDNA sequence. The 5’ part of cDNA was amplified with the following primers: 5’-TTTT AAG CTT ACA GAG CTC ATC TCT CTT CC-3’ and 5’- CAG ACC TCA TCG ATA TGG TTC-3’. This PCR product, containing the ClaI siste (bold), was digested with HinDIII and ClaI restriction enzymes. The HinDIII was created by PCR primer (underlined). The 3’ part of cDNA was amplified with the following primers: 5’-AAC CAT ATC GAT GAG GTC TGG-3’, and 5’-AGG GTC AGG GAT CCT GGC T-3’. This PCR product was digested with ClaI and BamHI (underlined). These products were ligated into HinDIII and BamHI sites of pcDNA3.1(+) plasmid. The whole insert was sequenced to ensure that the clone without mutations was selected. In this way we generated expression vector coding for unmodified isoform of DUSP13. Subsequently we amplified cDNA of the DUSP13 isoform from this plasmid using the following primers: 5’-TTTTGCTAGCCCTGCCATGGGGCTCTGCCAC-3’ and 5’- TTTTAAGCTTTCCGAACCGCCCCGTCTCCCG-3’. In the first primer the NheI restriction site was created (underlined) and the location of start codon is marked in bold. In the second primer the HinDIII site was created (underlined) and the last codon of DUSP13 isoform (Phe, in reverse) is marked by bold font. The PCR product was ligated into NheI and HinDIII sites of pcDNA3.1(+) plasmid containing EGFP cDNA cloned into BamHI and EcoRI sites. In this way we ligated DUSP13 cDNA in frame with EGFP cDNA. The last PCR step was needed to remove the stop codon of DUSP13 isoform. The correctness of the constructed chimeric gene was confirmed by sequencing. To find out if the fusion protein is expressed, we transfected the plasmids coding for chimeric protein or for only EGFP to U-2 OS cells using FuGene 6. The next day, cells were harvested and the lysates were prepared. The expression of proteins was examined by Western blotting as described above. In order to observe the cellular localization of the fusion protein DUSP13-EGFP (or control EGFP), the U-2 OS cells were seeded on chambered coverglasses and on the next day, the cells were transfected with the plasmids using FuGENE 6. The localization of proteins in living cells was observed starting 24 hours post-transfection with Zeiss confocal microscope.
3. Results
The results of high-throughput sequencing of mRNA molecules (RNA-Seq) can easily reveal the alternative splicing and the use of alternative promoters of genes. In our recently-published RNA-Seq results [
5], we noticed that the transcription of
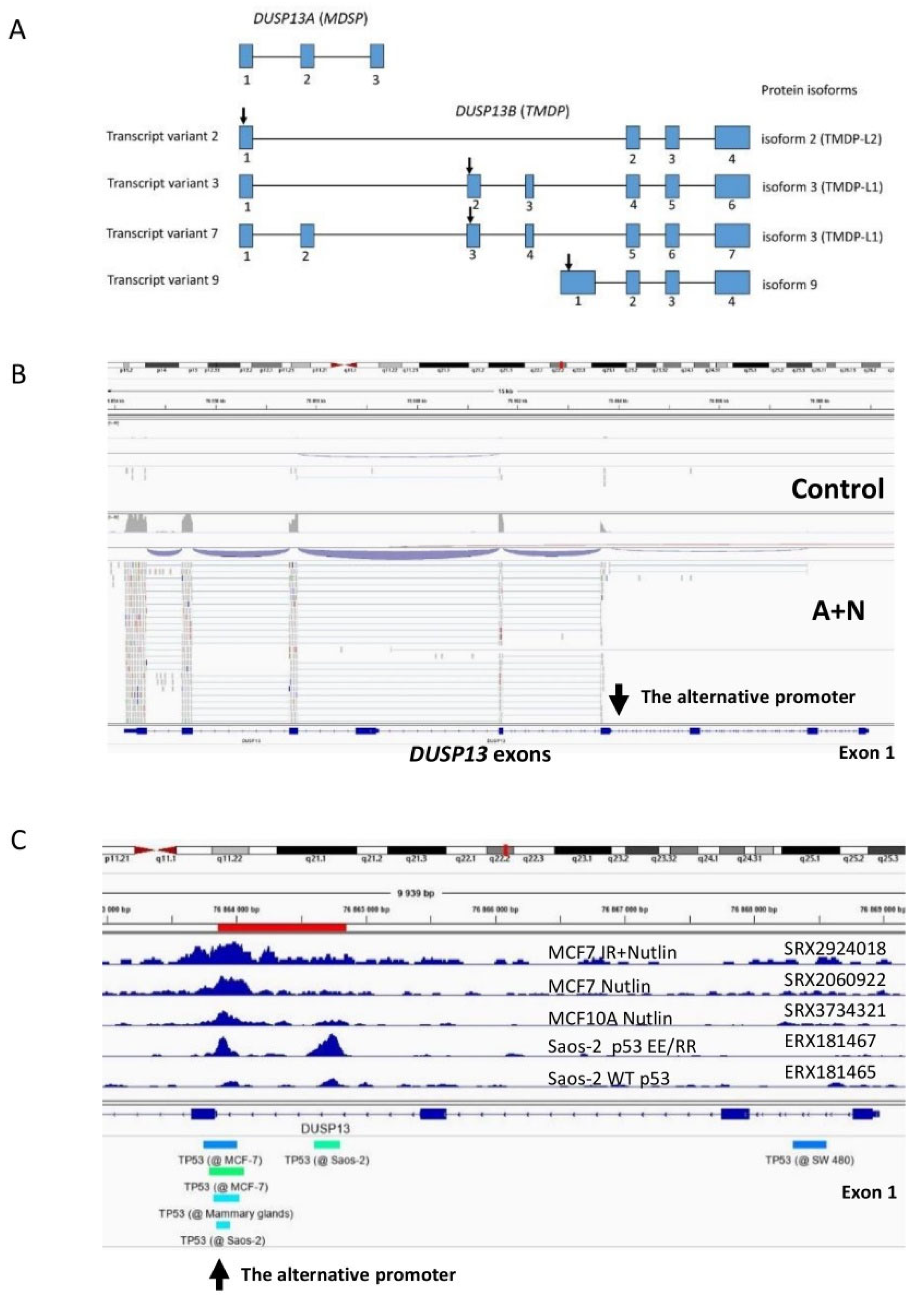
DUSP13 gene in A549 cells exposed to A+N starts from the alternative promoter located in intron (
Figure 1B). Based on the pattern of exon splicing (
Figure 1B), we conclude that the mRNA in the cells exposed to A+N codes for the isoform of DUSP13B called TMDP-L1. The known transcript variants 3 and 7 of
DUSP13B (
Figure 1A), which encode TMDP-L1 contain the long 5’UTR coded by one (variant 3) or two (variant 7) upstream exons shared with
DUSP13A. In case of A549 cells exposed to A+N, the transcription starts just upstream the exon with translation start codon, so the upstream codons for long 5’UTR are not included in this transcript. We observed similar expression pattern in RNA-Seq data from three other biological replicates of the experiment (A549 cells exposed to A+N for 30h) and in three other cell lines (A375, U-2 OS and NCI-H460) treated in the same fashion. The raw sequencing data are deposited in Sequence Read Archive under accession numbers PRJNA831359 and PRJNA837373.
The location of p53 binding sites in DNA can be detected using sequencing of DNA isolated from chromatin immunoprecipitated with anti-p53 antibody (ChIP-Seq). The results of these investigations performed on various cells growing in control conditions or exposed to various stress factors were published in numerous papers. They can be viewed by the ChIP-Atlas platform [
18]. Interestingly, we noticed two p53 ChIP-Seq peaks within intron of
DUSP13, however, these two peaks were detected only in Saos-2 cells ectopically expressing engineered p53 molecules (p53EE/RR) with strong cooperative binding of p53 monomers (
Figure 1C). Wild-type p53 expressed in Saos-2 did not bind to this locus (
Figure 1C) [
19]. Interestingly, the locus close to the exon was found to be bound by p53 in three data sets: in MCF7 cell line exposed to nutlin-3a and ionizing radiation, in MCF7 cells exposed to nutlin-3a and in non-cancerous breast epithelial cells (MCF 10A) exposed to nutlin-3a. Thus, in epithelial cells derived from breast, p53 activated by nutlin-3a is able to bind to the sequence close to the indicated
DUSP13 exon. The ChIP-Atlas tool does not show p53 binding to this locus in any other examined cell line (except Saos-2 expressing the p53EE/RR mutant). Thus, p53 binds to this site in cell-type specific manner, preferentially in cells derived from breast epithelium. To find out if
DUSP13 was found in transcriptomic studies to be activated by nutlin-3a in MCF7 cells, we browsed the database published by Fisher et al. [
20]. In five transcriptomic databases
DUSP13 was found to be activated by nutlin-3a in MCF7 cells. In three of the studies the upregulation was statistically significant. Thus, in MCF7 cells, nutlin-3a induces both upregulation of
DUSP13 mRNA and p53 binding to the alternative promoter detected by us.
In order to find out if this fragment of intron can function as a promoter activated by p53, we cloned it into the reporter vector. The cloned fragment encompasses the region of the two p53 ChIP-Seq peaks as marked on
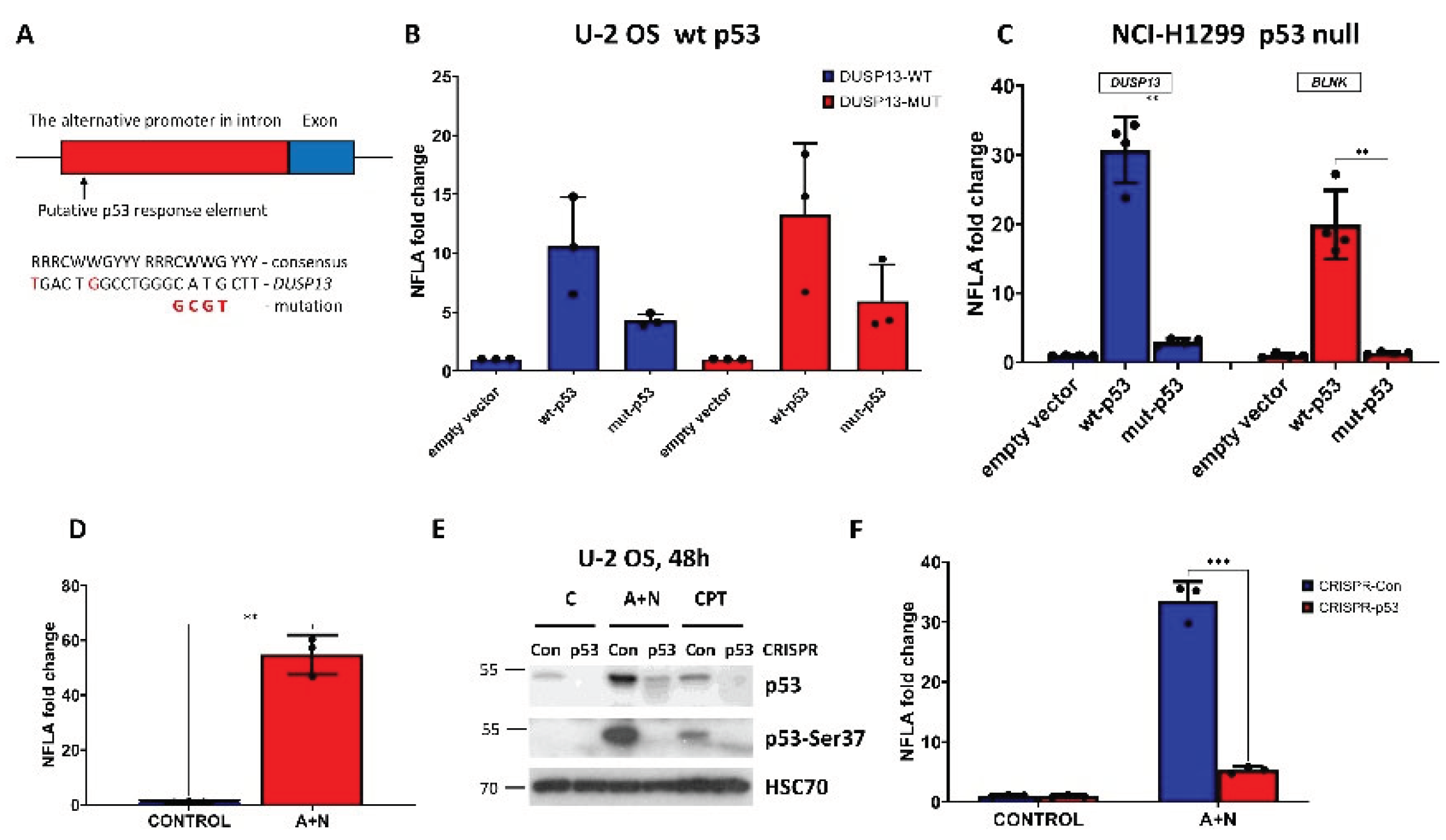
Figure 1C (red line). Moreover, we noticed that there is a sequence closely matching the consensus site of p53 response element and overlapping the distal peak (
Figure 2A). The
DUSP13 sequence shows only two mismatches, what makes this sequence highly probable p53 response element (
Figure 2A). We mutated its four critical sequence positions as shown on
Figure 2A. The reporter assays performed on U-2 OS cell line demonstrated that wild-type p53 stimulated the wild-type promoter approximately 10-fold (
Figure 2B). The activation of the promoter by the mutant p53 was significantly weaker. However, the mutant promoter still can be activated by wild-type p53, hence the p53-response element must be located elsewhere in the cloned region, probably within the proximal ChIP-Seq peak. Because U-2 OS cells have endogenous wild-type p53, which could interfere with the ectopically expressed p53 molecules, we performed the reporter assay on a cell line, NCI-H1299, which does not express endogenous p53. Moreover, for comparison, we employed a reporter plasmid with other p53-activated promoter cloned by us earlier from
BLNK gene [
5]. This gene was also identified by others as p53-regulated gene [
21]. The reporter test (
Figure 2C) performed on p53-null cell line confirmed our conclusion that ectopically expressed wild-type p53 strongly activated cloned
DUSP13 promoter and the mutant p53 lost this ability. Moreover, we found that activation of
DUSP13 promoter was stronger than
BLNK promoter.
Next we tested if cloned
DUSP13 promoter can be activated by the endogenous p53. To this end, U-2 OS cells were transfected with the reporter vector containing wild-type
DUSP13 promoter controlling firefly luciferase gene (
Figure 2D). As an internal control, we transfected a reporter vector with the gene for the luciferase from
Renilla sp. transcriptionally regulated by HSV-TK promoter to control for transfection efficiency and stress induced by A+N. Twenty four hours after transfection, control cells were mock-treated, whereas the experimental cells were exposed to A+N for 24 hours. Subsequently, the cells were harvested and the activities of both luciferases were measured. The firefly luciferase activity was divided by
Renilla luciferase activity producing the normalized firefly luciferase activity (NFLA). In cells exposed to A+N the value of NFLA increased more than 50-fold indicating that exposure to A+N stimulates the activity of the cloned promoter, probably by activating the endogenous p53 (
Figure 2D). To test the involvement of p53, we performed this experiment on p53-deficient U-2 OS cells and their controls prepared using CRISPR/Cas9 technology as described by us earlier [
5]. To show that these cells are p53-deficient (
Figure 2E) we exposed them to A+N or camptothecin (a precursor of anticancer drugs – topotecan and irinotecan) and we examined the expression of total p53 or its form with phosphorylated Ser37 - the amino acid located in transcription activating domain, a fragment, which is a target of CRISPR/Cas9-generated DNA break. The Ser37 is phosphorylated on activated p53. The p53-deficient cells express low amount of total p53 and p53 with the activating phosphorylation on Ser37 (
Figure 2E). In control cells for knockdown, the exposure to A+N activates
DUSP13 promoter more than 30-fold, whereas in p53-deficient cells only five-fold (
Figure 2F). Thus, the activity of p53 is required to strongly activate the alternative promoter of
DUSP13 following exposure to A+N.
To find out if the activation of endogenous
DUSP13 gene is controlled by p53, we performed experiment with p53-deficient A549 cells prepared by CRISPR/Cas9 technology as described previously [
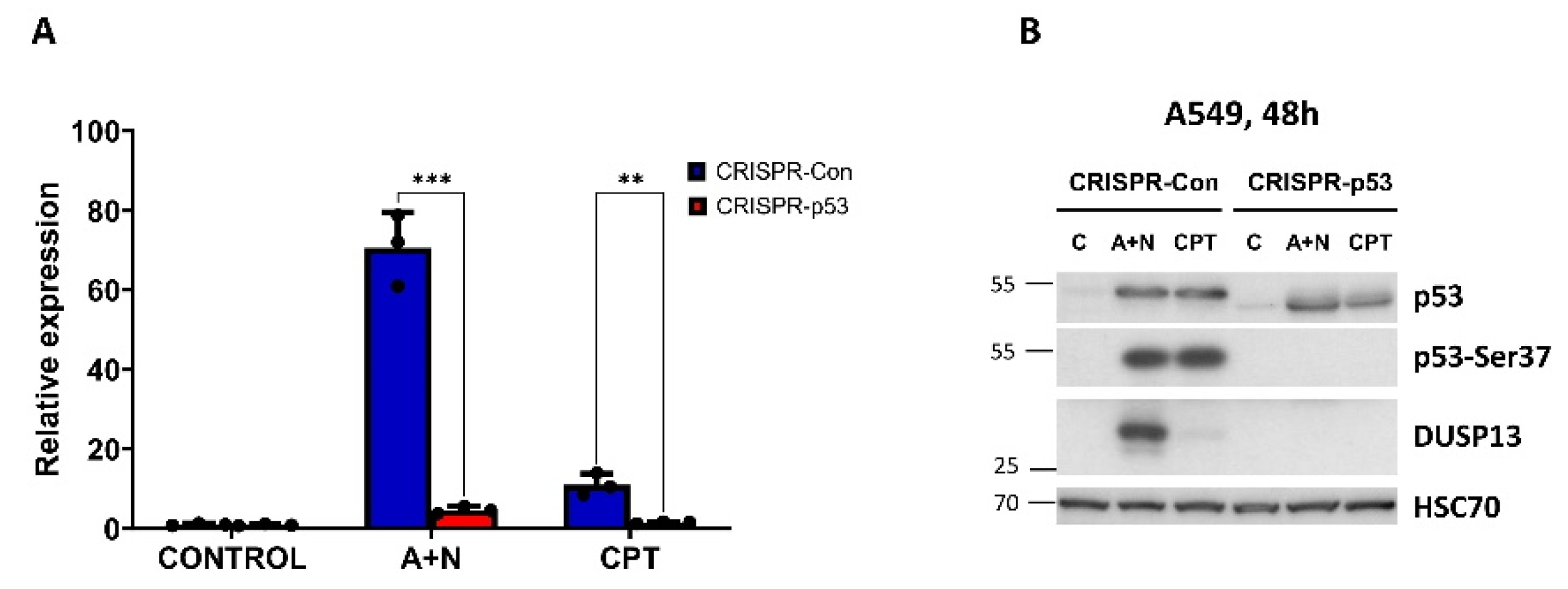
5]. This is a mixture of p53-null clones and clones with deletion in the transcription activating domain of p53. The p53-deficient cells and their controls were exposed to A+N or to another strong activator of p53 – camptothecin. After exposing cells to these compounds either RNA samples were isolated for RT-PCR or protein lysates were prepared for Western blotting. Both analyses gave concordant results, namely that A+N induces stronger activation of
DUSP13 than camptothecin and that wild-type p53 is indispensable for activation of
DUSP13 by either compound (
Figure 3A,B). This Western blot also shows that the protein coded by the alternative mRNA of
DUSP13 is actually produced in cells exposed to A+N (or camptothecin) and can be detected by the antibody employed in our experiments (
Figure 3B).
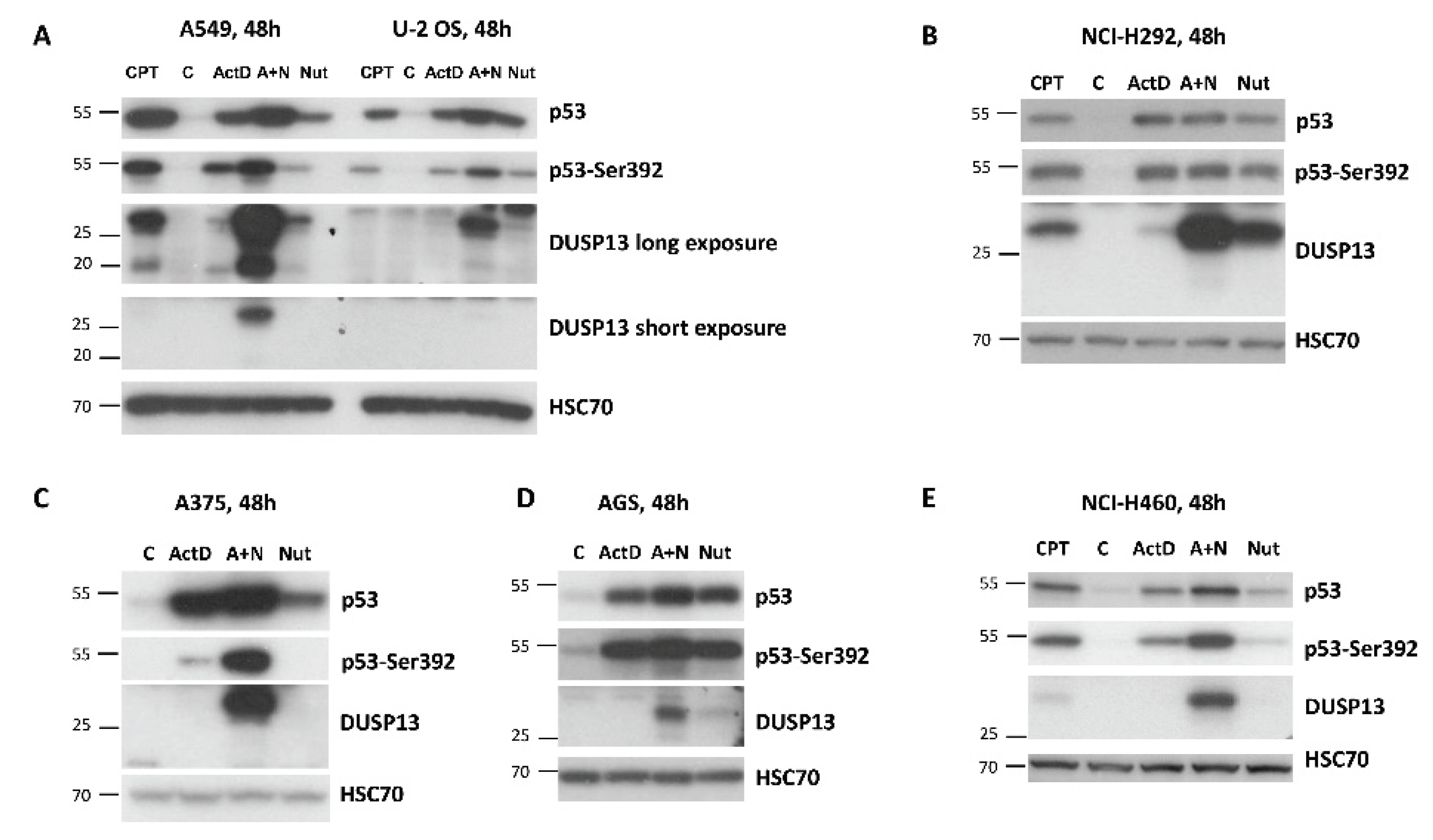
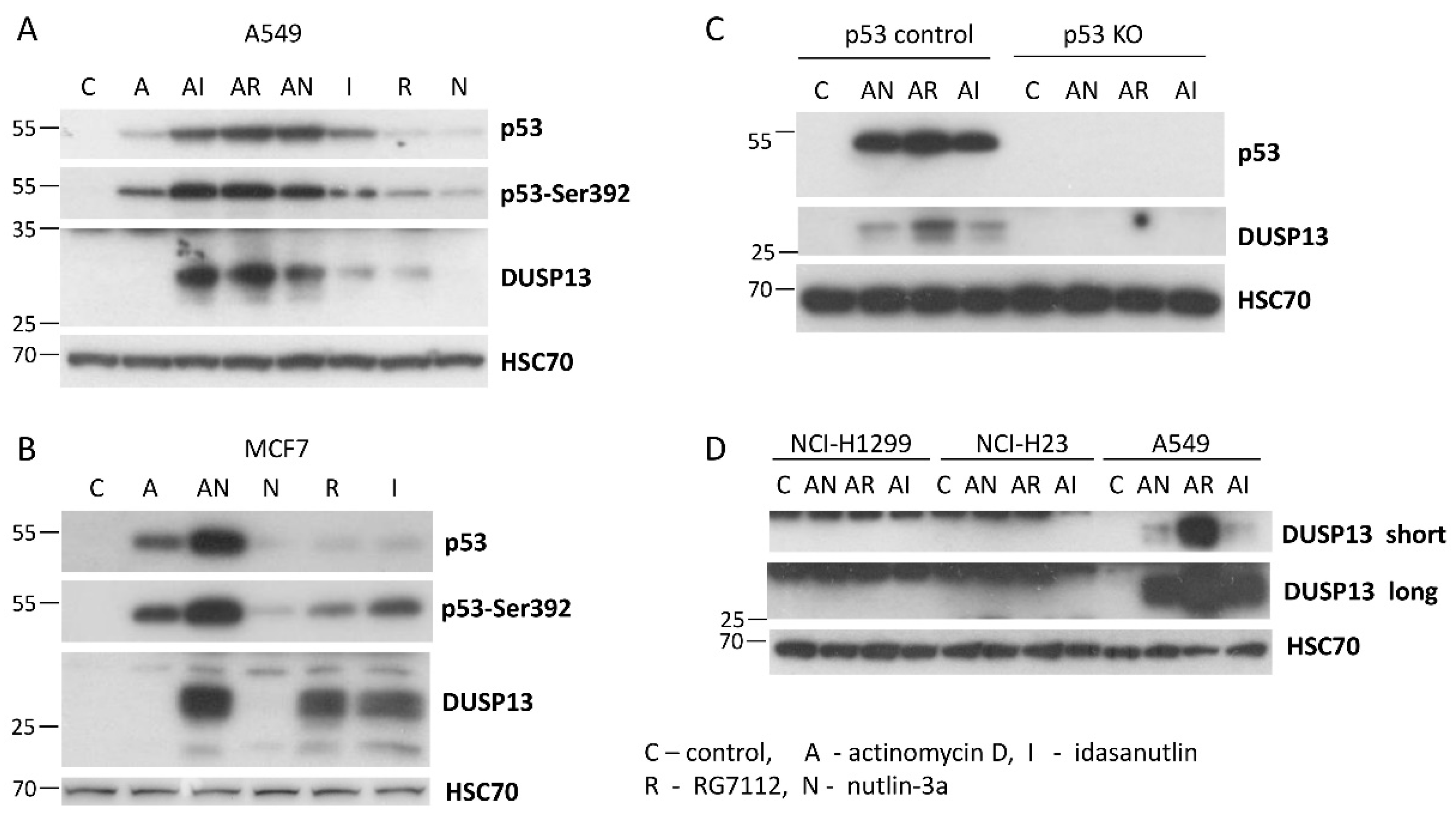
Subsequently, using Western blotting, we examined expression of DUSP13 in various cancer cell lines exposed to actinomycin D, nutlin-3a, A+N and camptothecin. We selected three lung cancer cell lines (A549, NCI-H460, NCI-H292), osteosarcoma cell line (U-2 OS), melanoma cell line (A375) and gastric adenocarcinoma cell line (AGS). All cell lines had wild-type gene coding for p53 protein. The cells were exposed to the compounds for 48 hours (
Figure 4). As a surrogate marker of p53 activation we used the phosphorylation status of Ser392 of p53. In our previous study in A549 cells, the amount of phospho-Ser392-p53 correlated well with the expression level of the examined p53 target genes [
7]. What is clearly visible on the Western blots is very strong synergy between actinomycin D and nutlin-3a in activation of
DUSP13 gene in all cell lines tested and weaker activation of the gene by camptothecin. DUSP13 protein is barely detectable in cells exposed to actinomycin D or nutlin-3a acting alone, whereas the accumulation of DUSP13 in cells exposed to both compounds is very strong. The exception is NCI-H292 cell line showing relatively high expression of DUSP13 protein even in cells exposed to nutlin-3a acting solo. This is consistent with the aforementioned observations made on MCF7 cells that in some cell types, this gene can be upregulated by “standard” activation of p53 with nutlin-3a. In some cells we observed striking correlation between phosphorylation of p53 on Ser392 and the expression of DUSP13 (A375) whereas in other cells the correlation was weak or absent. So the phosphorylation status of Ser392 is not a perfect indicator of the capacity of p53 to activate
DUSP13 gene. In order to explain the divergent ability of p53 to activate
DUSP13 gene one would need to perform detailed analysis of the abundant posttranslational modifications of p53, which modulate its activity [
22]. The general understanding of the mechanisms that determine the cell specificity of gene activation by p53 is very limited.
DUSP13 may be one of the models for such studies.
Regular exposure of our blots detects DUSP13 form with the size of approximately 30 kDa (
Figure 4). After long exposure of the signals on the blot, another protein is detected by the employed antibody in A549 cells treated with A+N. This protein form has a molecular mass of approximately 20 kDa compared with the approximately 30 kDa of the major form (
Figure 4A). We suspect that it may be DUSP13 translated from an AUG codon downstream the major translation start site (see below). There is possibility, that the shorter protein is the TMDP-L2 isoform of DUSP13B, which has calculated size 5 kDa smaller compared with TMDP-L1. However, this possibility appears very unlikely for the following reason. The translation of TMDP-L2 starts from the codon located in the first exon (
Figure 1A). However, our RNA-Seq data show that this exon is not transcribed in cells exposed to A+N (no sequence reads mapped to this exon,
Figure 1B). Similar pattern was observed in the aforementioned RNA-Seq data from additional biological replicates of the treated A549 cells and from treated other cell lines (U-2 OS, A375 and NCI-H460).
To find out if other antagonists of p53-MDM2 interaction have similar impact on DUSP13 expression as nutlin-3a we exposed A549 cells to idasanutlin or RG7112 (reviewed by Kocik et al., [
23]) either alone or in combination with actinomycin D. As is demonstrated on
Figure 5A these two compounds also synergize with actinomycin D in activation of DUSP13. Moreover, at the concentration used in the experiment (5 µM) they induce DUSP13 also when acting alone (
Figure 5A).
Because the published high-throughput experiments demonstrated that nutlin-3a can activate
DUSP13 in MCF7 cells [
20], we exposed them to actinomycin D alone or in combination with nutlin-3a as well as to the MDM2 antagonists acting unassisted (nutlin-3a, RG7112 and idasanutlin) (
Figure 5B). Expectedly, A+N combination upregulated DUSP13, however nutlin-3a had no detectable effect, whereas both RG7112 and idasanutlin were able to stimulate the expression of DUSP13 protein. This observation indicates that the isoform of DUSP13, which we detect by the employed antibody, can be induced not only by the drug combination like A+N, but also by at least two MDM2 antagonists acting solo. Moreover, as in the previous experiment, it is clear that the more advanced versions of MDM2 antagonists, idasanutlin and RG7112, are better activators of p53 target DUSP13 than nutlin-3a.
Next, we used these advanced antagonists of p53-MDM2 interaction to test if DUSP13 protein can be upregulated in cells in which p53 was inactivated by various mechanisms. First, we tested a p53 knockout clone isolated from the mixture of p53-deficient clones used for the experiment presented on
Figure 3. The control clone expresses p53 (
Figure 5C). In p53 knockout clone neither drug combination upregulated the expression of DUSP13 protein. Neither DUSP13 was upregulated in lung cancer cell lines, which are either p53 null (NCI-H1299) or express mutant p53 (NCI-H23) (
Figure 5D) [
24]. Thus, DUSP13 is not upregulated in engineered p53-deficient cells (
Figure 3), in p53-null-cells (
Figure 5C) nor in naturally occurring p53-null or p53-mutated cancer cell lines (
Figure 5D).
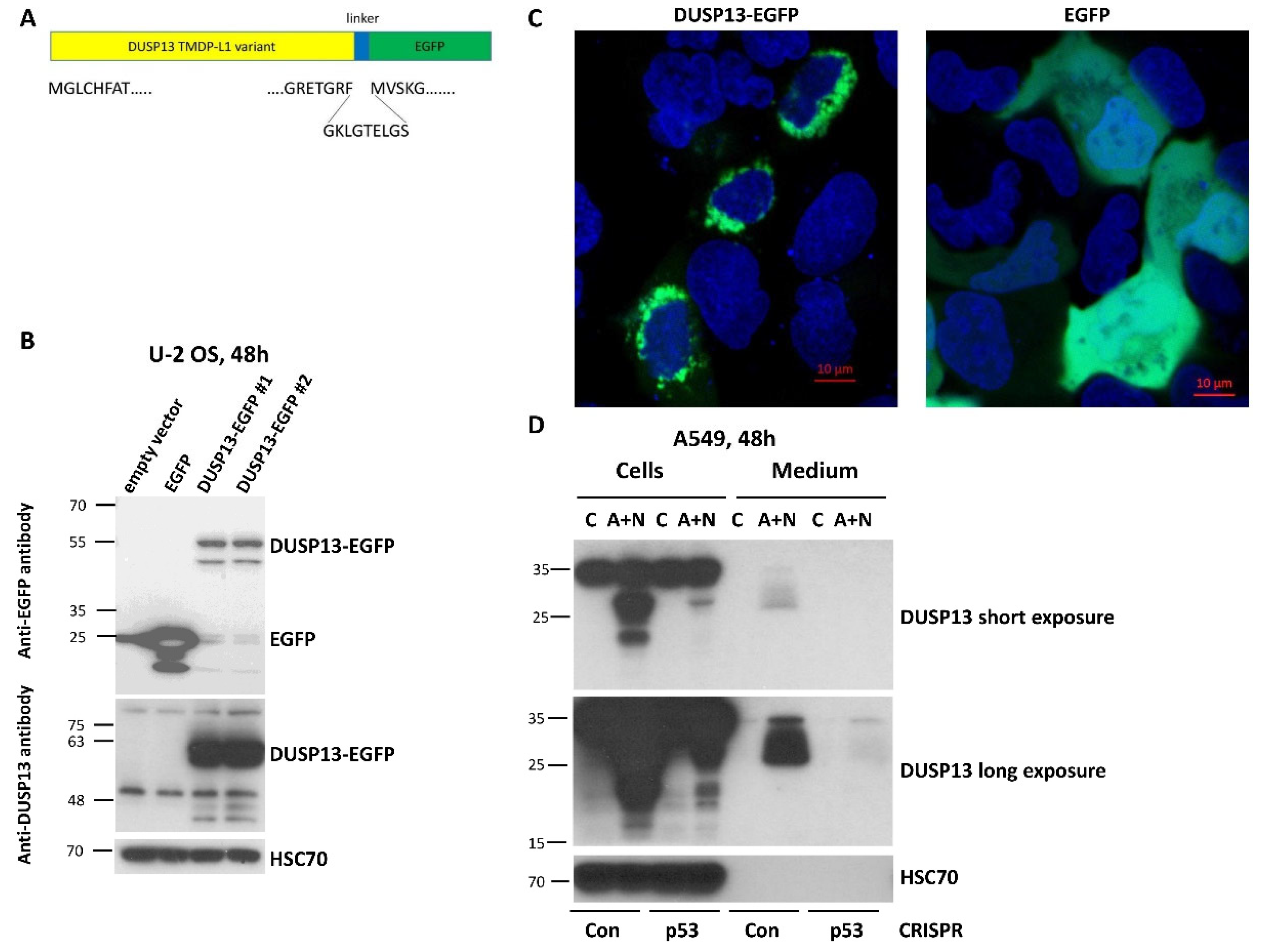
To gain insight into biological function of TMDP-L1 isoform of DUSP13, we amplified its cDNA sequence from A549 cells exposed to A+N and cloned it into an expression vector in front of the coding sequence of the green fluorescent protein EGFP (
Figure 6A). This chimeric cDNA codes for the chimeric protein with EGFP attached to the C-terminal fragment of TMDP-L1. We wanted to keep the native form of the N-terminus of DUSP13, because it may contain a signaling sequence guiding the protein to its destination within a cell or extracellular space. Subsequently, we transfected the vector with the chimeric gene to U-2 OS cells and we observed the molecular weight of the fusion protein by Western blotting (
Figure 6B) and its localization by confocal microscopy of living cells (no fixation) (
Figure 6C). According to the Western blotting (
Figure 6B) the size of the DUSP13-EGFP fusion protein (55 kDa) is about 30 kDa higher when compared to the size of EGFP alone (25 kDa), what is in agreement with the expected size of this version of DUSP13 (32 kDa). The fusion protein was detected by antibody directed against EGFP as well as by antibody directed against DUSP13 used in this work (
Figure 6B). The microscopic observations demonstrated that the DUSP13-EGFP fusion protein is localized in cytoplasmic region around the nucleus, what contrasts with the localization pattern of EGFP, which localizes in cytoplasm and nucleus without any aggregation in perinuclear region (
Figure 6C).
The N-terminus of TMDP-L1 contains a sequence, which was predicted with high probability as a signal peptide MGLCHFATLALILLVLLEALAQAD [
25]. This peptide targets a protein to the secretory pathway encompassing endoplasmic reticulum, Golgi apparatus, trans Golgi network, secretory vesicles and plasma membrane. To find out if this protein is secreted, we analyzed both – the cell lysates and conditioned media from control cells and cells exposed to A+N for 48 hours. We performed the experiment on the aforementioned p53-deficient A549 cells and their controls. The proteins from cell lysates and concentrated media were separated by electrophoresis, transferred to membranes and probed with anti-DUSP13 antibody. In the lysates the antibody detected two protein forms (approximately 20 kDa and 30 kDa) but only in p53-proficient cells exposed to A+N (
Figure 6D). In the medium, the smear of protein with the size of about 30 kDa and higher was detected, but again, only in case of p53-proficient A+N-treated cells. In the medium, the 20 kDa protein was not visible even after overexposure of detection film (
Figure 6D). If this shorter protein is translated from an alternative, downstream start codon, then it lacks the signal peptide for secretion. Moreover, the lack of 20 kDa protein in the medium suggests that DUSP13 does not passively spill to extracellular space – in this case both forms would be detected. In conclusion, a fraction of the endogenous DUSP13 protein from A549 cells exposed to A+N is secreted to the extracellular space.
4. Discussion
The expression of DUSP13 dual specificity phosphatase is best-studied in mice. Exons 1-3 code for the open reading frame of MDSP isoform expressed in diaphragm and skeletal muscles, whereas exons 7-9 encode open reading frame of TMDP isoform expressed in testis. The transcript for MDSP also contains exons 4, 5, 7, 8, 9, which encode long 3’-UTR [
15]. Genes encoding alternative ORFs producing two distinct proteins are rare in higher eukaryotes. The isoform, expressed in A549 cells exposed to A+N is similar to TMDP. This transcript was found for the first time in muscles and its protein product was named TMDP-L1, however it was not detected by Western blotting [
15]. We found that the expression of TMDP-L1 is induced in A549 cells exposed to A+N. In these conditions, A549 cells produce a protein with the size expected for TMDP-L1 (approximately 30 kDa), which can be detected by the antibody employed in our experiments. The DUSP13 mRNA and protein can also be upregulated by another substance strongly activating p53 – camptothecin, although in this case the upregulation is significantly weaker. Moreover, in some cancer cell lines the isoform can be strongly upregulated by nutlin-3a or by more advanced forms of MDM2 antagonists acting alone. Activation of
DUSP13 measured at mRNA or protein level is strongly attenuated in p53-deficient cells (or prevented in p53-null cells) what indicates that p53 plays role in this process, apparently in direct fashion because both ectopically expressed and endogenous p53 is able to activate the cloned alternative promoter of
DUSP13. However, there must be something special in the interaction between p53 and the alternative promoter of
DUSP13 driving the expression of TMDP-L1. The data published by Schlereth et al. [
19] demonstrated that in Saos-2 cells this DNA fragment shows two p53 binding sites, one closer to first exon (proximal site) and one more distant from the exon (distal site). However, these sites are occupied in Saos-2 cells not by wild-type p53 but only by artificially prepared mutant version of this protein (
Figure 1C), which promotes cooperative binding between p53 monomers. Interestingly, ChIP-Seq analyses performed by others demonstrated that the proximal site is occupied by endogenous p53 in MCF7 breast cancer cell line exposed to nutlin-3a or in non-cancerous breast epithelial cell line (MCF 10A) exposed to nutlin-3a. We performed a mutation of a plausible p53 response element overlapping the distal site identified
in silico by Tebaldi et al. [
4], however the mutation did not destroy the ability of the cloned promoter to respond to p53. Thus, this cloned promoter contains other, less obvious p53 binding site, probably overlapping the proximal ChIP-Seq peak occupied by p53 in cells derived from breast epithelium exposed to nutlin-3a. In all examined cell lines actinomycin D and nutlin-3a strongly synergized in stimulation of the expression of DUSP13 protein. Actinomycin D or nutlin-3a acting alone did not stimulate the expression of DUSP13, the exception being NCI-H292 cells, where this isoform is induced by nutlin-3a (
Figure 4B). Thus, “regular” activation of p53 is able to stimulate the expression of DUSP13 only in subset of cells (MCF7, MCF 10A, NCI-H292). Thus, to activate the expression of this isoform of DUSP13, p53 requires a special set of modifications or interactions with other proteins expressed only in some cells. It is also plausible, that this alternative promoter is in closed chromatin in most cells and only the treatment with A+N is able to open this chromatin for p53 binding. Interestingly, a recent review by Fischer et al. [
20] provides a summary of transcriptomic studies searching for p53-regulated genes. In these analyses,
DUSP13 was identified as p53 target only in 12 out of 57 studies. This confirms that this gene is regulated by p53 in specific conditions, what does not depreciate its role in cellular physiology.
DUSP13 is not a well-studied protein. As of this writing “DUSP13” phrase in PubMed returns only 19 papers (as of 19th February 2024). The gene was cloned by Nakamura et al. [
14]. The crystal structure of TMDP isoform was determined [
26]. Interestingly, TMDP variant when overexpressed with DUSP4 in A549 cell line attenuates TGFβ1-induced migration and drug resistance [
27]. Recently,
DUSP13 was found to belong to three-gene signature that can accurately distinguish COVID-19 patients from healthy controls [
28]. This finding indicates that DUSP13 may participate in some immune-related activity. Another recently published data indicate that DUSP13 is a protein, which can be detected in blood plasma and may be associated with the increased risk of atrial fibrillation [
29]. The expression of DUSP13 was reduced after epithelial-mesenchymal transition of ovarian cancer cell line triggered by TGFβ1 [
30]. The oncology research focusing on DUSP13 is very limited [27, 31, 32, 33]. Some researchers stumbled upon
DUSP13 during various high-throughput studies. In the latest of these reports, the authors found that a regulatory axis involving lncRNA PVT1, miR-378c and
DUSP13 is involved in microvascular invasion in hepatocellular carcinoma. Moreover, based on various types of data analyses, the authors suggested that DUSP13 may be involved in lipid metabolism, glycosyl compound metabolic process and xenobiotic metabolic process [
33]. Our analysis suggests that
DUSP13 easily responds to activation of p53 in cells derived from breast epithelium.
The physiological role of TMDP-L1 isoform upregulated by p53 is not known. The TMDP expressed in testis inhibits stress-activated MAPK kinases and suppresses AP-1-dependent gene expression [
16]. Whether a similar role is played by TMDP-L1 variant induced by A+N in p53-dependent manner remains to be determined. Our data clearly indicate that this isoform of DUSP13 is a part of the p53-regulated stress-response system, what may prompt other researchers to better study this form of DUSP13. One of the major findings of this paper is that DUSP13 expression is not limited to testis or muscles but can be found in cancer cells with wild-type p53 exposed to the clinically relevant substances (e.g. camptothecin), experimental drug combination (A+N) or antagonists of MDM2-p53 interaction (nutlin-3a, idasanutlin and RG7112). This line of research may be continued because there is an indispensable tool – the antibody, which can detect TMDP-L1 expression. It is important because, in our experience, there is scarcity of commercially available and usable antibodies recognizing poorly studied proteins.