1. Introduction
Type 2 Diabetes Mellitus is a chronic metabolic disease, defined by hyperglycaemia caused by abnormalities in insulin secretion, insulin action, or both. Diabetes is a major risk to health due to its characteristic long-term complications [
1] with the International Diabetes Association estimating a prevalence of 537 million people living with diabetes worldwide in 2021 and a projected increase of 46% by 2045 [
1].
Frequently, Diabetes Mellitus leads to several health complications. Those can be grouped in “microvascular diseases” (due to damage to small blood vessels) and “macrovascular diseases” (due to damage to the arteries) [
2]. Microvascular complications include retinopathy, nephropathy, and neuropathy [
2]. The major macrovascular complications include accelerated cardiovascular disease resulting in myocardial infarction and cerebrovascular disease manifesting as strokes [
2]. In opposite, diabetic ketoacidosis from exceptionally hyperglycaemia and coma as the result of hypoglycaemia can be fatal [
3]. Type 2 Diabetes Mellitus is also characterized by comorbidities such as obesity and cardiovascular diseases.
Type 2 Diabetes Mellitus has an established screening and diagnostic protocol, either by the haemoglobin A1C criteria or plasma glucose concentration (after 12h fasting or 2-hour plasma glucose) [
4]. After a diagnosis of Diabetes Mellitus, the patients require self-monitoring of plasmatic glucose concentrations, regularly. The estimation of plasmatic glycated haemoglobin, and of lipid levels are also necessary [
4] for the evaluation of disease progression. Although the diagnosis of diabetes and blood glucose monitoring are well established, monitoring the many complications of diabetes remains a challenge [
2].
Saliva is being addressed as a relevant biofluid for diagnostics/prognosis since it is safe and easy to collect, its collection is non-invasive, and it contains several blood and systemics molecules frequently used as biomarkers of diagnosis [
5]. Although salivary diagnostics is increasingly being considered, monitoring pre- and established diabetes and its complications via saliva is yet to be validated [
6]. The identification of markers of prognosis could support the development of “point-of-care” tests that a physician could use for patient follow up and alert for behavioral changes or adjustment of diabetes control medication.
The aim of this study was to propose biomarkers in saliva that allow monitoring of diabetes and its complications. We hypothesize that saliva provides the biological information – proteins – to identify characteristic proteome changes induced by Type 2 Diabetes Mellitus.
2. Results
We hypothesized that Type 2 Diabetes Mellitus induces characteristic proteome changes detectable in saliva. We used a methodological approach for the functional analysis of altered salivary proteins identified by mass spectrometry to point out or clarify molecular changes associated with Type 2 Diabetes Mellitus.
Analysis of the 10 saliva samples (Type 2 Diabetes Mellitus and Healthy) resulted in the identification of 917 proteins (
Table S1) with a 5% normalized FDR, of which 622 were quantified with a 1% normalized FDR (
Table S2). PANTHER gene list analysis has successfully mapped 611 from the 622 proteins participating in 16 biological processes (
Figure S1).
2.1. Protein enrichment analysis
Enrichment analysis of the 622 proteins quantified revealed 185 proteins involved in 11 disrupted biological processes. The fold change of proteins involved in the enriched biological processes is represented as follows: 112 proteins are increased, while 73 proteins are decreased in the saliva of patients with Type 2 Diabetes Mellitus.
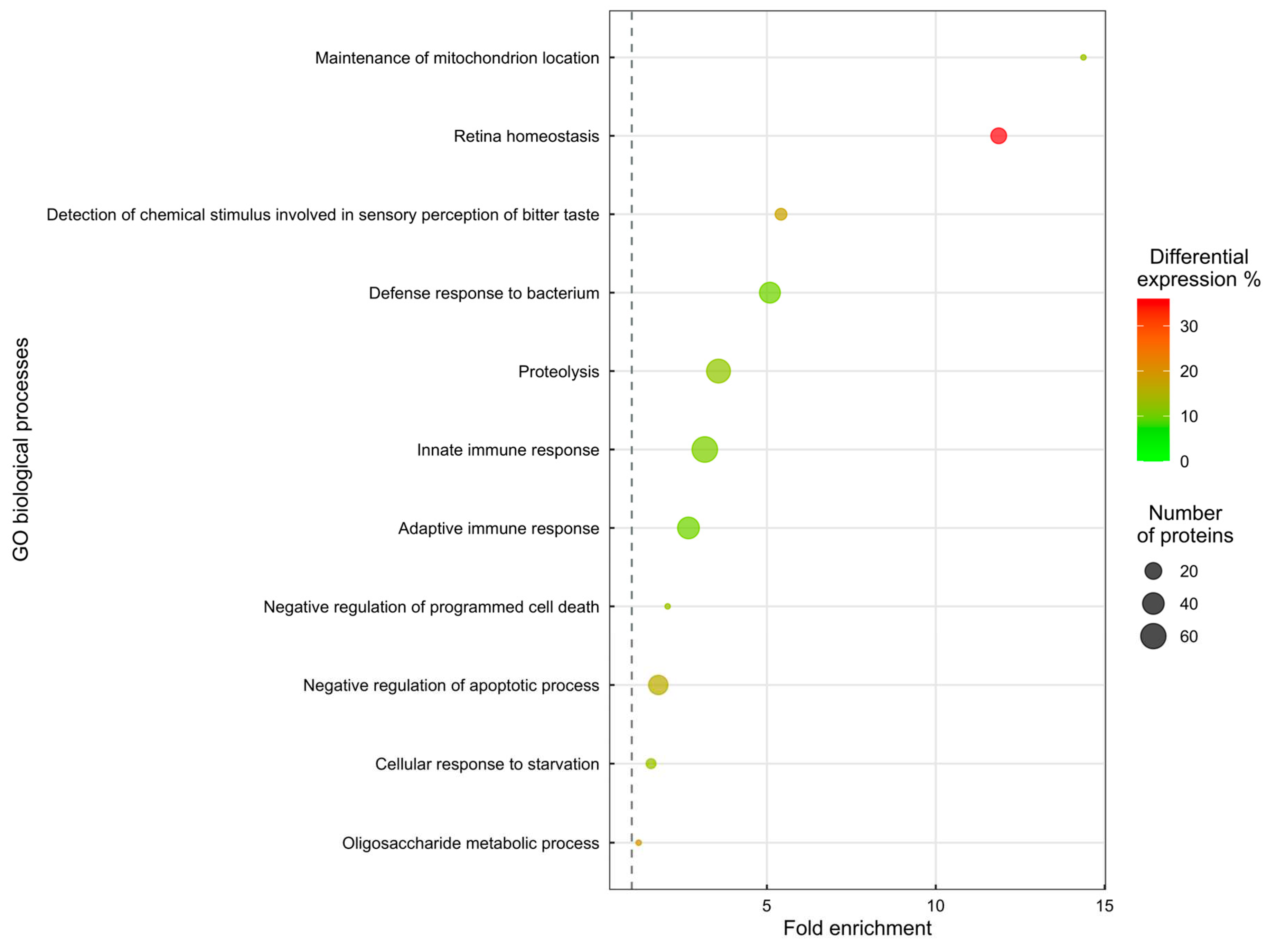
Eleven biological processes are significantly altered (enrichment analysis, Biological Process Enrichment,
Figure 1). In a summary assessment of the enrichment analysis, the mechanism most affected is inflammatory response (124 proteins). Other mechanisms altered in DMT2 patients are Detection of chemical stimulus involved in sensory perception of bitter taste (7 proteins), Cellular response to starvation (4 proteins), Maintenance of mitochondrion location (1 protein) and Oligosaccharide metabolic process (1 protein).
3. Discussion
Despite the high prevalence of diabetes Mellitus type 2, the molecular mechanisms underlying the development and progression of this disease are still not fully understood. Researchers have focused on identifying biomarkers that can facilitate the diagnosis, monitoring, and treatment of Type 2 Diabetes Mellitus. One potential source of biomarkers is saliva, which contains proteins that can reflect changes in the body's metabolic state. We investigated whether Diabetes Mellitus type 2 induces characteristic proteome in saliva.
3.1. Retina homeostasis
Retina homeostasis, related to retinopathy (one of the most well-known complications of diabetes) is dysregulated in Type 2 Diabetes Mellitus. We identified 9 proteins related to retina homeostasis (
Table 1).
Retina requires a healthy choroid to nourish the different outer retinal layers. Lactotransferrin (LTF), present in the human retina, has been successful used in the treatment of lesions in choroidal neovascularization in mice and, in parallel, when coated, demonstrated to be efficient on the treatment of the dry eye [
9], due it is including iron-transferring, anti-inflammatory, antioxidant, antimicrobial, and anticancer activities [
10]. We showed that LTF is more abundant in Type 2 Diabetes Mellitus saliva samples (FC = 5.25) than in control samples, being one of the protein contributors to the separation of study groups (VIP = 2.02). Although LTF has been associated to diabetes before, some researchers report elevated concentrations while others report a decreased concentration of LFT in diabetic patients [
11].
Heat shock protein beta-1 (HspB1) is upregulated upon optic nerve injury [
12]. This suggests that HspB1 might have a protective value for retinal ganglion cells [
12]. In a similar perspective, HspB1 has been shown to play a vital role in maintaining the integrity of retinal capillary endothelial cells. Proinflammatory cytokines are generally upregulated in the diabetic retina, reducing the levels of HspB1 in retinal capillary endothelial cells, which could lead to their apoptosis in diabetic retinopathy [
13]. This reduction was observed in our results with HspB1 being less abundant on Type 2 Diabetes Mellitus saliva samples (FC=-2.49) than in control samples.
We identified an increase in Lipocalin-1 (LCN1) concentration on Type 2 Diabetes Mellitus saliva samples (FC=6.02). To our knowledge, this is the first report of an association between LCN1 and Type 2 Diabetes Mellitus. Lipocalin-1 belongs to a family of transport proteins and scavenges lipids from the corneal surface [
14]. Lipocalin-2 (LCN2, suggested as being an upstream regulator of apoptosis of retinal degeneration) levels are significantly increased in the whole retinae of rats after induction of retinal degeneration, compared to the controls [
15]. LCN2 is instrumental for the progression of diabetic complications like encephalopathy and peripheral neuropathy [
15]. The LCN2 upregulation observed after the retinal degeneration can be correlated with the LCN1 increased concentration on our Type 2 Diabetes Mellitus samples.
Albumin (ALB), previously suggested as diabetic retinopathy biomarker in serum [
16], is also more abundant (FC=2.23) in Type 2 Diabetes Mellitus saliva. Increased levels of albumin have been related to the disruption of the retinal barrier that occurs in diabetes [
17].
3.2. Negative regulation of apoptotic processes
Apoptotic processes are involved in the dynamic regulation of pancreatic β-cell mass which are directly linked to insulin production. Hyperplasia of β-cells may be related to active replication and/or neogenesis. This increase is balanced by the reduction of the number of cells via cell death resultant from necrosis and/or apoptosis. It is apoptosis that plays a critical role in the reduction of β-cell mass occurring in patients with Type 2 Diabetes Mellitus and subsequent progressive worsening of glycaemic control. The negative regulation of apoptotic process is dysregulated in Type 2 Diabetes Mellitus saliva samples (
Table 1).
Lactotransferrin promotes apoptosis and the elevated levels have been related with neutrophil dysfunction in Type 2 Diabetes Mellitus [
18]. In fact, LTF has been suggested as a biomarker to monitor the development or progression of diabetes and its complications [
18]. On the opposite side, Endoplasmic reticulum chaperone BiP (HSPA5) is suggested to be a protector against ER stress, required for β-cells dysregulated apoptosis and consequent Type 2 Diabetes Mellitus development [
19].
It can be hypothesized that a failure of the negative regulation of apoptotic process can lead to necrosis of β-cells. We show that Lactotransferrin is more abundant on Type 2 Diabetes Mellitus saliva samples (FC = 5.25) sustaining the assumptions of previous authors [
18] that LTF promotes the neutrophil dysfunction in Type 2 Diabetes Mellitus. HSPA5, identified as protector of ER, is less abundant on Type 2 Diabetes Mellitus saliva samples (FC = -2.48), which corroborates the previous assumption.
It is important to point out that the negative regulation of programmed cell death process is dysregulated as well (
Figure 1). Expansion (neogenesis, replication and hypertrophy) and involution (apoptosis, necrosis, and atrophy) processes are related since both may contribute to a unfunctional equilibrium in the β-cell mass and consequent glycaemic control. Albumin (ALB), that plays a role as a negative inflammation biomarker, was found elevated on Type 2 Diabetes Mellitus saliva samples (FC = 2.23) suggesting an active inflammatory status [
20].
3.3. Cellular response to starvation
Starvation is one of the earliest forms of insulin resistance and is associated with increased lipid metabolism molecules (lipolysis, plasma fatty acid, ketone bodies and intramyocellular lipids). This increased lipid load is related with reduced insulin sensitivity and consequently with Type 2 Diabetes Mellitus, but the related mechanisms remain unclear [
21]. Our data shows that the negative regulation of cellular response to starvation is altered in Type 2 Diabetes Mellitus (
Figure 1).
Glutamine synthetase (GLUL) is involved in the catalysis of glutamate to glutamine. Glutamate is directly dependent of leptin to be up taken in astrocytes in a time-dependent manner. The rapid glutamate capture by astrocytes indicates that leptin could reduce the stimulatory effects of glutamate at nearby synapses, thereby reducing appetite. The rapid rise of insulin with feeding promotes uptake and storage of ingested energy in fat and muscle. The rapid fall of insulin with starvation promotes catabolism [
22]. Both leptin and insulin play key roles in signaling the physiologic transition from energy sufficiency to deficient energy/starvation [
22]. We show that GLUL is less abundant in saliva (FC = -1.62) of Type 2 Diabetes Mellitus patients which can be a consequence of starvation stimulus dysregulation.
3.4. Maintenance of mitochondrion location
Type 2 Diabetes Mellitus is characterized by mitochondrial dysfunction and consequent homeostasis disruption, with elevated production of reactive oxygen species (ROS) and low levels of ATP. Downregulation of genes involved in mitochondrial biogenesis and oxidative phosphorylation has been observed in Type 2 Diabetes Mellitus [
23]. The maintenance of mitochondrion location is characterized by two processes: the conservation of specific location within a cell and prevention from moving elsewhere.
The link between the levels of Albumin and the maintenance of mitochondrion location are not well established yet. In the context of Type 2 Diabetes Mellitus, albumin levels may be influenced by several factors, including mitochondrial oxidative stress increase that leads to a change in mitochondrial function [
24]. In saliva of patients with Type 2 Diabetes Mellitus, albumin is increased (FC = 2.23). Being a marker of the oxidative stress characteristic of diabetes [
25] a preventive monitorization of Albumin could avoid this microvascular disease appearance.
3.5. Detection of chemical stimulus involved in sensory perception of bitter taste
Taste receptors in the gustatory system are expressed in the gastrointestinal tract and influence nutrients intake. Different flavors stimulate specific taste receptors that release metabolic hormones, modulating insulin expression with impact on glucose homeostasis. In Type 2 Diabetes Mellitus, some taste receptors are altered, and this is linked to glucose and insulin dysregulation. The detection of the chemical stimulus involved in sensory perception of bitter taste seems to be dysregulated, as suggested by the enrichment analysis (
Figure 1).
Cystatin-SN (CST1) is more abundant in saliva of individuals experiencing bitter tastes [
26]. In Type 2 Diabetes Mellitus, Cystatin-SN is up-regulated in patients with burn mouth sensation, attributed to diabetic peripheral neuropathy [
27]. This could reflect a defensive reaction against an on-going inflammation [
27]. In our study, CST1 is less abundant (FC = -2.18) in the saliva of Type 2 Diabetes Mellitus patients, which is contrary to that found previously [
26].
Zinc-alpha-2-glycoprotein (AZGP1) has a direct relation with sensory perception of bitter taste. AZGP1 gene has been suggested as a potential obesity gene that may predispose to the development of obesity [
28]. AZGP1 plays a role in the regulation of adipose tissue mass, stimulating lipolysis, inhibiting lipid accumulation in adipose tissue, regulating serum lipid values, and influencing the secretion of other adipokines [
29]. AZGP1 is less abundant (FC = -1.56) in Type 2 Diabetes Mellitus saliva samples than in control samples.
Carbonic anhydrase 6 (CA6) is a key factor responsible for the reduction of hepatic glucose production and has been suggested as a marker for several complications of diabetes, namely retinopathy, kidney disease, neuropathy, and cardiovascular disease [
30]. In saliva, CA6 was more abundant (FC = 10.52) in Type 2 Diabetes Mellitus patients, in agreement with previous studies [
30].
3.6. Adaptative and Innate immune response
Recent research suggests that both innate and adaptive immunity play a critical role in the pathogenesis of Type 2 Diabetes Mellitus, with evidence pointing to the abnormal differentiation of immune system components as a key factor in the development of insulin resistance. Alterations in the proliferation of T cells and macrophages, and impairment in the function of NK and B cells are frequent phenomena in individuals with Type 2 Diabetes Mellitus [
31].
Enrichment analysis revealed a dysregulation of the adaptative and innate immune response (
Figure 1,
Table 1).
Fibrinogen alpha chain (FGA) and Fibrinogen beta chain (FGB) are involved in the disruption of the adaptative and innate immune responses. Fibrinogen is a crucial modulator of haemostatic balance and inflammatory processes in diabetes [
32]. In our study FGA and FGB were found elevated in saliva of the patients with Type 2 Diabetes Mellitus (FC = 2.11 and FC = 4.06), in consonance with previous fundings that show that Fibrinogen is increased in blood in different moments of Type 2 Diabetes Mellitus: before the onset of Type 2 Diabetes Mellitus, and during the transition from pre-Type 2 Diabetes Mellitus to Type 2 Diabetes Mellitus [
33].
The bactericidal permeability-increasing protein (BPI) is an endogenous protein that in Type 2 Diabetes Mellitus can reduce inflammation [
34]. In saliva, BPI was more abundant in patients with Type 2 Diabetes Mellitus than in healthy subjects (FC = 14.99), probably a reflection of the body's attempt to fight the inflammatory state characteristic of diabetes.
3.7. Defense response to bacterium
In Diabetes, insulin deficiency and hyperglycaemia cause the disruption of immune system mechanisms such as cytokine production, immune cells function and phagocytosis [
35]. The defense response to bacteria is dysregulated in patients with Type 2 Diabetes Mellitus (
Figure 1,
Table 1).
These immune alterations can affect the production and activity of immunoglobulins, reducing their levels in saliva [
36]. A compromised immune response may also impact the expression and secretion of histatin-1, increasing the susceptibility to infections [
37]. We found that, in saliva, Histatin-1 is less abundant (FC = -4.52), as in other studies [
36,
37]. It’s possible to assume a direct link between these factors and an increased risk of oral infections and oral health complications in individuals with T2DM.
We found that, in saliva of diabetic patients, Neutrophil elastase (NE) is more abundant (FC = 8.02) supporting the link with defense response to bacteria dysregulation in Type 2 Diabetes Mellitus.
Diabetic foot ulcer is characterized by the unbalance of the immune system and is a common complication in Type 2 Diabetes Mellitus individuals. Open wounds are an entry for different pathogens that take advantage of a compromised immune system. Although the underlying pathophysiological mechanisms of the diabetic foot are still unclear, the relation between S100-A8 and S100-A9 protein levels in serum and the healing process was reported [
38,
39]. The data is not consensual, as the levels of both proteins in serum were lower in Type 2 Diabetes Mellitus individuals [
39] or higher depending on the technology used to detect them: proteomics or immune-specific quantitative assays [
38]. In saliva of Type 2 Diabetes Mellitus patients, both S100-A8 and A9 were more abundant: S100-A8 (FC = 1.38) and A9 (FC = 1.79). This upregulation can stimulate the inflammatory response, causing the release of proinflammatory cytokines, and preventing the normal wound cicatrisation [
39].
3.8. Proteolysis
Insulin resistance of Type 2 Diabetes Mellitus patients leads to an increase in proteolysis and a decrease in protein synthesis. Enrichment analysis shows that proteolysis is dysregulated in Type 2 Diabetes Mellitus saliva (
Figure 1,
Table 1).
Type 2 Diabetes Mellitus results from a reduction in the ability of insulin to stimulate glucose utilization (insulin resistance) and an inadequate pancreatic β-cell insulin secretion in response to hyperglycaemia [
40]. Matrix mettaloproteinase-9 is upregulated in plasma of Type 2 Diabetes Mellitus individuals exhibiting an increased proteolytic activity [
40]. In saliva of Type 2 Diabetes Mellitus patients MMP-9 is more abundant than in control saliva.
3.9. Oligosaccharide metabolic process
Functional oligosaccharides play a positive role in the prevention of diabetes, including improvement of pancreas function, α-glucosidase inhibition, the relief of insulin and leptin resistance, anti-inflammatory effects, regulation of gut microbiota and gut hormone, and intervention of diabetic risk factors [
41].
We show that the oligosaccharide metabolic process is dysregulated (
Figure 1). This process has the carbohydrate metabolic process as its ancestor. Alpha-amylase 1A, the protein contributing to this dysregulation, is produced primarily in the salivary glands and pancreas and is responsible for breaking down complex carbohydrates, including starch and glycogen, into smaller sugar molecules such as oligosaccharides [
42]. Some studies have shown that individuals with Diabetes Mellitus type 2 may have higher salivary Alpha-amylase activity compared to individuals without diabetes [
43]. Nonetheless, our data show a decrease on the amount of Alpha-amylase in saliva of type 2 Diabetes Mellitus patients (FC = -2.14), which may be explained by the method of saliva collection under fasting or non-fasting conditions [
44].
3.10. Additional protein analysis
In addition to the proteins involved in enriched biological processes, we addressed the role of proteins with VIP>1. Thirty-eight proteins with a VIP > 1 (
Figure 2), but that do not account for the enrichment of the biological processes are listed in the supplementary
Table S3.
Basic salivary proline-rich protein 1 (PRB1) is expressed in large amounts in parotid glands. In saliva, PRB1 is much less abundant in the saliva of diabetes patients than in healthy subjects (FC = -20), which is correlated with a lower oral health typical of diabetic patients [
45].
Alpha-2-macroglobulin-like protein 1 (A2ML1) was suggested as a potential biomarker for diabetic retinopathy and other diabetic complications [
46]. A2ML1 is upregulated in the plasma of diabetic patients with retinopathy [
46]. In our sample group, A2ML1 was less abundant in the saliva samples of Type 2 Diabetes Mellitus patients (FC = -2.78).
Alpha-1-antitrypsin (SERPINA1), a defense response protein involved in the pro-inflammatory immune response, has been associated with Type 2 Diabetes Mellitus, and is also increased in proliferative diabetic retinopathy [
47]. In saliva, this protein was also found more abundant in the saliva of Type 2 Diabetes Mellitus patients (FC = 2.75).
Hemopexin (HPX), a scavenger of heme, is more concentrated in the saliva of patients with proliferative diabetic retinopathy [
48] and in the urine of uncontrolled diabetes patients [
49]. In our study this protein was found more abundant in the saliva of diabetic patients (FC = 3.38).
It is important to point out that both Hemoglobin subunits (alpha and beta) were identified in the saliva of diabetic patients, which surely can be related with the common oral manifestation of diabetes such as periodontitis [characterized by bleeding gums [
50]].
4. Materials and Methods
4.1. Ethical Statement
This study was carried out in accordance with the Helsinki Declaration and the Oviedo Convention. Saliva samples were collected following the guidelines for the protection of personal data and ethical issues in use at the Biobank of the Institute of Molecular Medicine (IMM) of the Faculty of Medicine, University of Lisbon (
http://biobanco-imm.biobanco.pt/documents). Written informed consent including the purpose of the study, data confidentiality, rights of participants, and the right to withdraw from the study at any time was provided by every participant or by their legal representatives or guardians before study enrolment.
4.2. Participant Enrolment
Five saliva samples (3 females and 2 males aged 69 to 82) of a cohort of Type 2 Diabetes Mellitus patients from Centro de Saúde Dão Lafões, Portugal, were collected. Saliva samples from five healthy subjects (3 females and 2 males aged 8 to 39) showing no evidence of oral and systemic pathologies after clinic evaluation, were also collected.
4.3. Saliva Collection
Unstimulated saliva samples were collected with two sterile cotton rolls placed under the tongue, for 2 minutes. The cotton rolls were placed inside a 15 mL sterile plastic tube with a sterile 100 mL pipette tip in the bottom to facilitate saliva collection by centrifugation (10,000 × g; 10 minutes; 4 ºC). Samples were then refrigerated and inactivated using 1% Triton X-100. Whole saliva was centrifuged (10,000 × g; 10 min; 4 °C). The supernatant was aliquoted and stored (80 °C) until analysis.
4.4. Sample preparation and LC-MS methodology
Protein concentration was determined using the Pierce BCA assay kit (ThermoFisher Scientific), according to the manufacturers’ instructions and adjusted to 50 mg/mL. Saliva was analyzed by capillary electrophoresis (Experion™ automated capillary electrophoresis system, Bio-Rad Laboratories), and compared to the in-house profiles to confirm sample integrity (data not shown).
Sample preparation for data-dependent acquisition (DDA) experiments and data-independent acquisition (DIA), as well as LC-MS methodology [protein identification and relative quantification of proteins (SWATH-MS)] was performed as described previously [
7].
4.5. Protein functional analysis
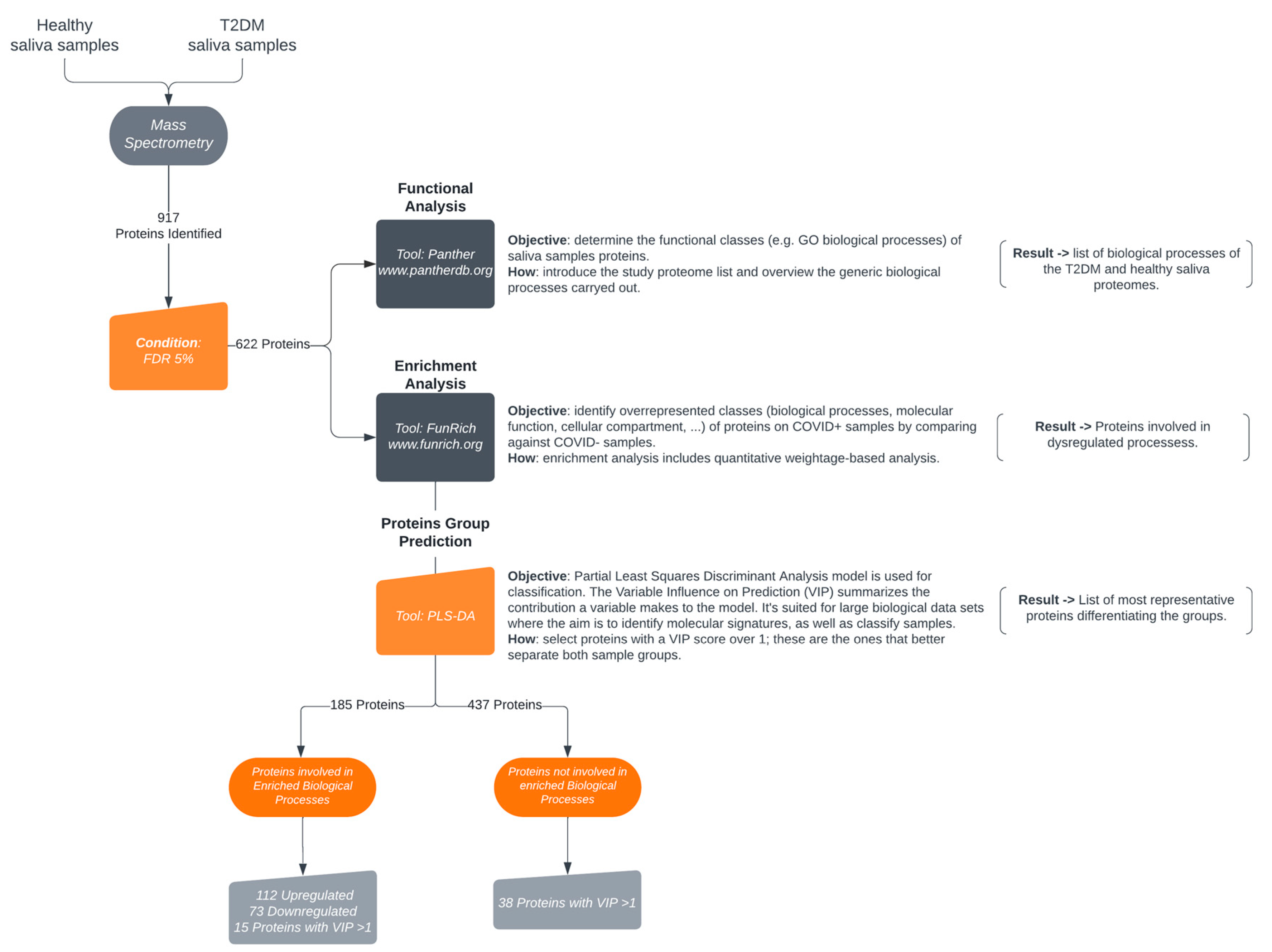
Functional analysis of salivary proteins in Type 2 Diabetes Mellitus identified by mass spectrometry was performed according to
Figure 2.
Narrowed down proteome biological processes classification was obtained via the PANTHER Gene List Analysis (PANTHER V 17.0) (
http://www.pantherdb.org/). Only the PANTHER GO-SLIM biological processes were considered. The dataset followed an enrichment analysis of biological processes’ using FunRich tool against UniProt curated database as background (
http://www.funrich.org/). Selected biological processes enriched with a minimum of 10% of differential expression were plotted using an adapted script as described in Bonnot et al., 2019 [
8] within the ggplot2 package version 3.4.0 in RStudio version 2022.12.0.353.
PLS-DA model, based on variable importance projection (VIP) values, was used to select the proteins capable to classify the 2 groups of samples.
4.6. Statistical analysis
The Mann-Whitney test was used to select the proteins statistically different between the Type 2 Diabetes Mellitus and Healthy samples. The multivariate analysis was performed in MetaboAnalyst as previously described [
7].
5. Conclusions
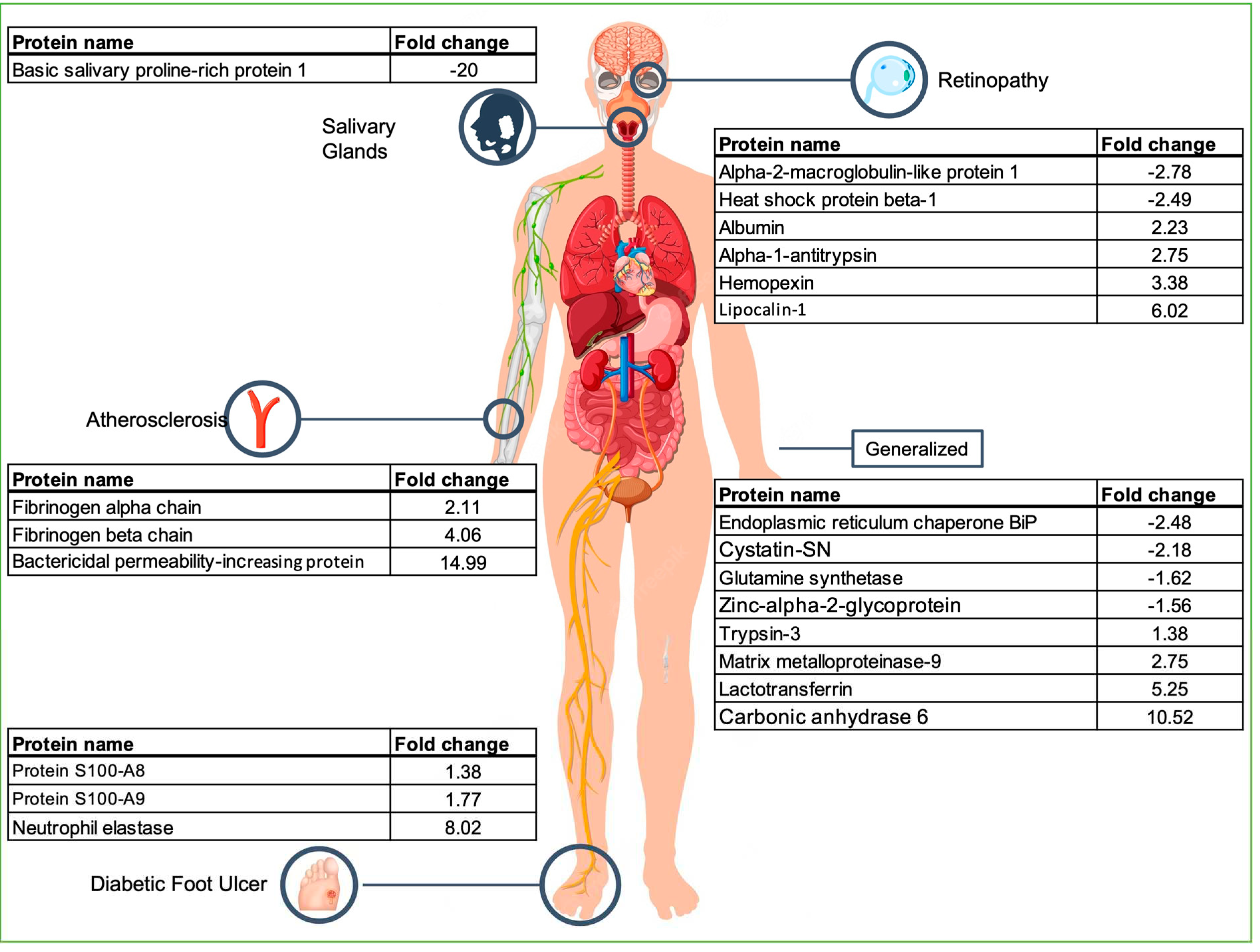
This study shows the enormous potential of saliva as a diagnostic fluid. We have identified 622 proteins altered in patients with Type 2 Diabetes Mellitus, highlighting twenty three that reflect conditions dispersed for different body locations: arteries (atherosclerosis), eye (retinopathy), diabetic foot ulcer (foot), metabolic syndrome (generalized), insulin resistance (generalized), molecular impact of glucose and insulin homeostasis dysregulation (generalized), haemostatic balance and inflammatory processes (generalized), protein catabolism (generalized) and salivary gland function (salivary glands) (
Figure 3). These proteins may be indicative of complications resulting from the disease and therefore we propose them as (potential) biomarkers for diabetes complications and prognosis. The proposed panel of biomarkers of diabetes complications, includes both Albumin and Lactotransferrin that are common to some of these processes. Therefore, these proteins can be considered more generic indicators of diabetes complications but not specific markers of one of these complications.
The identification of a panel of biomarkers for Type 2 Diabetes Mellitus complications in saliva is a pilot study that needs validation in a larger number of samples allowing a stronger correlation of the molecular data with patient complications. These results can serve as a basis for the development of new tests to support a more complete diagnosis of diabetes and its complications, and a point-of-care for close monitoring and providing frequent data to the physician.
Supplementary Materials
Table S1: LC-MS identification table with protein summary (Protein FDR Summary); Table S2: Proteins identified with an FDR 5% and p < 0.05; Table S3: List of proteins with a VIP score > 1 not listed in the enrichment analysis. These are the proteins that better differentiate Type 2 Diabetes Mellitus and Healthy samples. Shown are the UniProtKB accession number (AC), protein names, gene names, fold change (Type 2 Diabetes Mellitus /Healthy) and the variable importance projection (VIP) score defined by the PLS-DA analysis; Figure S1: PANTHER Biological processes of saliva proteins. Box size follows the number of proteins involved in each biological process. 1 – Cellular process (297 proteins); 2 – Metabolic process (226 proteins); 3 - Biological regulation (132 proteins); 4 – Response to stimulus (129 proteins); 5 – Localization (78 proteins); 6 – Immune system process (72 proteins); 7 – Signaling (51 proteins); 8 – Biological process involved in interspecies interaction between organisms (34 proteins); 9 – Multicellular organismal process (32 proteins); 10 – Developmental process (31 proteins); 11 – Locomotion (14 proteins); 12 – Biological adhesion (14 proteins); 13 – Growth (6 proteins); 14 – Reproductive process (2 proteins); 15 – Reproduction (2 proteins) and 16 – Biomineralization (1 protein).
Author Contributions
Conceptualization, E.E., M.J.C., A.C.E. and N.R.; methodology, E.E., M.J.C., A.L., D.A., A.C.E., N.R., B.M. and V.M.M.; validation, E.E., B.M. and V.M.M.; formal analysis, E.E., N.R., B.M. and V.M.M.; investigation, E.E., N.R., B.M. and V.M.M.; resources, N.R. and M.B.; data curation, E.E., A.C.E. and N.R.; writing—original draft preparation, E.E.; writing—review and editing, E.E., V.M.M., B.M., A.L., D.A., A.C.E., M.J.C., M.B., L.B. and N.R.; visualization, E.E., A.C.E. and N.R.; supervision, A.C.E., L.B. and N.R.; project administration, A.C.E. and N.R.; funding acquisition, N.R. and M.B. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by grants from the Fundação para a Ciência e Tecnologia for the support of CESAM (UIDP/50017/2020+UIDB/50017/2020+LA/P/0094/2020), CIIS (UIDB/04279/2020). The authors also wish to acknowledge the financial support of the National Mass Spectrometry Network (RNEM) (POCI-01-0145-FEDER-402-022125 Ref. ROTEIRO/0028/2013).
Institutional Review Board Statement
This study was carried out in accordance with the Helsinki Declaration and the Oviedo Convention. Saliva samples were collected following the guidelines for the protection of personal data and ethical issues in use at the Biobank of the Institute of Molecular Medicine (IMM) of the Faculty of Medicine, University of Lisbon (
http://biobanco-imm.biobanco.pt/documents).
Informed Consent Statement
Written informed consent including the purpose of the study, data confidentiality, rights of participants, and the right to withdraw from the study at any time was provided by every participant or by their legal representatives or guardians before study enrolment.
Data Availability Statement
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD042424.
Acknowledgments
We would like to express our sincere gratitude to the volunteer saliva donors who participated in this study. We would also like to acknowledge the USF Grão Vasco Medical Centre for their support and assistance in the volunteer’s recruitment and clinical management.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Sun H, Saeedi P, Karuranga S, et al. IDF Diabetes Atlas: Global, regional and country-level diabetes prevalence estimates for 2021 and projections for 2045. Diabetes Res Clin Pract 2022;183. [CrossRef]
- Roden M, Shulman GI. The integrative biology of type 2 diabetes. Nature 2019;576:51–60. [CrossRef]
- Forbes JM, Cooper ME. Mechanisms of Diabetic Complications. Physiol Rev 2013;93:137–88. [CrossRef]
- Goyal R, Jialal I. Diabetes Mellitus Type 2. In: StatPearls. Treasure Island (FL): : StatPearls Publishing 2022.
- Abd-Elraheem SE, EL saeed A mohammed, Mansour HH. Salivary changes in type 2 diabetic patients. Diabetes Metab Syndr Clin Res Rev 2017;11:S637–41. [CrossRef]
- Fouani M, Basset CA, Jurjus AR, et al. Salivary gland proteins alterations in the diabetic milieu. J Mol Histol 2021;52:893–904. [CrossRef]
- Esteves E, Mendes VM, Manadas B, et al. COVID-19 Salivary Protein Profile: Unravelling Molecular Aspects of SARS-CoV-2 Infection. J Clin Med 2022;11:5571. [CrossRef]
- Bonnot T, Gillard M, Nagel D. A Simple Protocol for Informative Visualization of Enriched Gene Ontology Terms. BIO-Protoc 2019;9. [CrossRef]
- López-Machado A, Díaz-Garrido N, Cano A, et al. Development of Lactoferrin-Loaded Liposomes for the Management of Dry Eye Disease and Ocular Inflammation. Pharmaceutics 2021;13:1698. [CrossRef]
- Kanwar JR, Roy K, Patel Y, et al. Multifunctional Iron Bound Lactoferrin and Nanomedicinal Approaches to Enhance Its Bioactive Functions. Molecules 2015;20:9703–31. [CrossRef]
- Delcheva G, Stefanova K, Stankova T, et al. IDF21-0107 Association between Serum Lactoferrin Levels and Hyperglycemia in Type 2 Diabetes. Diabetes Res Clin Pract 2022;186. [CrossRef]
- Schmidt T, Fischer D, Andreadaki A, et al. Induction and phosphorylation of the small heat shock proteins HspB1/Hsp25 and HspB5/αB-crystallin in the rat retina upon optic nerve injury. Cell Stress Chaperones 2016;21:167–78. [CrossRef]
- Rajeswaren V, Wong JO, Yabroudi D, et al. Small Heat Shock Proteins in Retinal Diseases. Front Mol Biosci 2022;9.
- Foulks GN, Borchman D. Chapter 17 - Abnormalities of eyelid and tear film lipid. In: Levin LA, Albert DM, eds. Ocular Disease. Edinburgh: : W.B. Saunders 2010. 131–7. [CrossRef]
- Ghosh S, Stepicheva N, Yazdankhah M, et al. The role of lipocalin-2 in age-related macular degeneration (AMD). Cell Mol Life Sci 2020;77:835–51. [CrossRef]
- Wang G-X, Fang Z-B, Li J-T, et al. The correlation between serum albumin and diabetic retinopathy among people with type 2 diabetes mellitus: NHANES 2011–2020. PLoS ONE 2022;17:e0270019. [CrossRef]
- Byrne EM, Llorián-Salvador M, Lyons TJ, et al. Tofacitinib Ameliorates Retinal Vascular Leakage in a Murine Model of Diabetic Retinopathy with Type 2 Diabetes. Int J Mol Sci 2021;22:11876. [CrossRef]
- Alhalwani AY. A Review of Lactoferrin Inflammatory Role in Type 2 Diabetes Mellitus with Neutrophil Dysfunction. J Pharm Res Int 2021;:377–90. [CrossRef]
- Laybutt DR, Preston AM, Åkerfeldt MC, et al. Endoplasmic reticulum stress contributes to beta cell apoptosis in type 2 diabetes. Diabetologia 2007;50:752–63. [CrossRef]
- Wang P, Yuan D, Zhang C, et al. High fibrinogen-to-albumin ratio with type 2 diabetes mellitus is associated with poor prognosis in patients undergoing percutaneous coronary intervention: 5-year findings from a large cohort. Cardiovasc Diabetol 2022;21:46. [CrossRef]
- Frank P, Katz A, Andersson E, et al. Acute exercise reverses starvation-mediated insulin resistance in humans. Am J Physiol-Endocrinol Metab 2013;304:E436–43. [CrossRef]
- Flier JS. Starvation in the Midst of Plenty: Reflections on the History and Biology of Insulin and Leptin. Endocr Rev 2019;40:1–16. [CrossRef]
- Rovira-Llopis S, Bañuls C, Diaz-Morales N, et al. Mitochondrial dynamics in type 2 diabetes: Pathophysiological implications. Redox Biol 2017;11:637–45. [CrossRef]
- Miller DJ, Cascio MA, Rosca MG. Diabetic Retinopathy: The Role of Mitochondria in the Neural Retina and Microvascular Disease. Antioxidants 2020;9:905. [CrossRef]
- Paramasivan S, Adav SS, Ngan SC, et al. Serum albumin cysteine trioxidation is a potential oxidative stress biomarker of type 2 diabetes mellitus. Sci Rep 2020;10:6475. [CrossRef]
- Dsamou M, Palicki O, Septier C, et al. Salivary protein profiles and sensitivity to the bitter taste of caffeine. Chem Senses 2012;37:87–95. [CrossRef]
- Castillo-Felipe C, Franco-Martínez L, Tvarijonaviciute A, et al. Proteomics-Based Identification of Salivary Changes in Patients with Burning Mouth Syndrome. Biology 2021;10:392. [CrossRef]
- Banaszak M, Górna I, Przysławski J. Zinc and the Innovative Zinc-α2-Glycoprotein Adipokine Play an Important Role in Lipid Metabolism: A Critical Review. Nutrients 2021;13:2023. [CrossRef]
- Pearsey HM, Henson J, Sargeant JA, et al. Zinc-alpha2-glycoprotein, dysglycaemia and insulin resistance: a systematic review and meta-analysis. Rev Endocr Metab Disord 2020;21:569–75. [CrossRef]
- Sulaiman AH, Ghassan ZI, Omar TN. Biochemical Evaluation of Carbonic Anhydrase and Some Antioxidant Markers in Patients with Diabetes Complications. Arch Razi Inst 2022;77:169–78. [CrossRef]
- Zhou T, Hu Z, Yang S, et al. Role of Adaptive and Innate Immunity in Type 2 Diabetes Mellitus. J Diabetes Res 2018;2018:e7457269. [CrossRef]
- Liu S-L, Wu N-Q, Shi H-W, et al. Fibrinogen is associated with glucose metabolism and cardiovascular outcomes in patients with coronary artery disease. Cardiovasc Diabetol 2020;19:36. [CrossRef]
- Bryk-Wiązania AH, Undas A. Hypofibrinolysis in type 2 diabetes and its clinical implications: from mechanisms to pharmacological modulation. Cardiovasc Diabetol 2021;20:191. [CrossRef]
- Sun Q, Li T, Li Y, et al. Bactericidal/Permeability-Increasing Protein Improves Cognitive Impairment in Diabetic Mice via Blockade of the LPS-LBP-TLR4 Signaling Pathway. Front Physiol 2021;11:718. [CrossRef]
- Berbudi A, Rahmadika N, Tjahjadi AI, et al. Type 2 Diabetes and its Impact on the Immune System. Curr Diabetes Rev 2020;16:442–9. [CrossRef]
- Chorzewski M, Orywal K, Sierpinska T, et al. Salivary protective factors in patients suffering from decompensated type 2 diabetes. Adv Med Sci 2017;62:211–5. [CrossRef]
- Olayanju OA, Mba IN, Awah NE, et al. Oral innate immunity in patients with type 2 diabetes mellitus in a tertiary hospital in Ibadan Nigeria: a cross-sectional study. Pan Afr Med J 2022;43:134. [CrossRef]
- Trøstrup H, Holstein P, Christophersen L, et al. S100A8/A9 is an important host defence mediator in neuropathic foot ulcers in patients with type 2 diabetes mellitus. Arch Dermatol Res 2016;308:347–55. [CrossRef]
- Krisp C, Jacobsen F, McKay MJ, et al. Proteome analysis reveals antiangiogenic environments in chronic wounds of Diabetes Mellitus Type 2 patients. Proteomics 2013;13:2670–81. [CrossRef]
- Modestino AE, Skowronski EA, Pruitt C, et al. Elevated Resting and Postprandial Digestive Proteolytic Activity in Peripheral Blood of Individuals With Type-2 Diabetes Mellitus, With Uncontrolled Cleavage of Insulin Receptors. J Am Coll Nutr 2019;38:485–92. [CrossRef]
- Zhu D, Yan Q, Liu J, et al. Can functional oligosaccharides reduce the risk of diabetes mellitus? FASEB J 2019;33:11655–67. [CrossRef]
- des Gachons CP, Breslin PAS. Salivary Amylase: Digestion and Metabolic Syndrome. Curr Diab Rep 2016;16:102. [CrossRef]
- Shah VS, Pareikh D, Manjunatha BS. Salivary alpha-amylase–biomarker for monitoring type II diabetes. J Oral Maxillofac Pathol JOMFP 2021;25:441–5. [CrossRef]
- Pérez-Ros P, Navarro-Flores E, Julián-Rochina I, et al. Changes in Salivary Amylase and Glucose in Diabetes: A Scoping Review. Diagnostics 2021;11:453. [CrossRef]
- Lone MA, Shaikh S, Lone MM, et al. Association of salivary gland hypofunction with diabetes mellitus and drugs among the elderly in Karachi, Pakistan. J Investig Clin Dent 2017;8. [CrossRef]
- Lu C-H, Lin S-T, Chou H-C, et al. Proteomic analysis of retinopathy-related plasma biomarkers in diabetic patients. Arch Biochem Biophys 2013;529:146–56. [CrossRef]
- Kalis M, Kumar R, Janciauskiene S, et al. α 1-antitrypsin enhances insulin secretion and prevents cytokine-mediated apoptosis in pancreatic β-cells. Islets 2010;2:185–9. [CrossRef]
- Kowluru RA, Chan P-S. Oxidative Stress and Diabetic Retinopathy. J Diabetes Res 2007;2007:e43603. [CrossRef]
- Bencharit S, Baxter SS, Carlson J, et al. Salivary proteins associated with hyperglycemia in diabetes: a proteomic analysis. Mol Biosyst 2013;9:2785–97. [CrossRef]
- Leite RS, Marlow NM, Fernandes JK. Oral Health and Type 2 Diabetes. Am J Med Sci 2013;345:271–3. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).